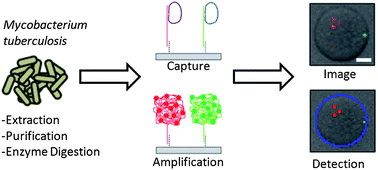
Mycobacterium tuberculosis detection via rolling circle amplification†
Abstract
Hybridization-based assays for DNA detection often use
* Corresponding authors
a
Department of Mechanical and Aerospace Engineering, California NanoSystems Institute, University of California, Los Angeles, California, USA
E-mail:
yongchen@seas.ucla.edu
b Department of Microbiology, Immunology and Molecular Genetics, University of California, Los Angeles, California, USA
Hybridization-based assays for DNA detection often use

 Please wait while we load your content...
Something went wrong. Try again?
Please wait while we load your content...
Something went wrong. Try again?
E. Schopf, Y. Liu, J. C. Deng, S. Yang, G. Cheng and Y. Chen, Anal. Methods, 2011, 3, 267 DOI: 10.1039/C0AY00529K
To request permission to reproduce material from this article, please go to the Copyright Clearance Center request page.
If you are an author contributing to an RSC publication, you do not need to request permission provided correct acknowledgement is given.
If you are the author of this article, you do not need to request permission to reproduce figures and diagrams provided correct acknowledgement is given. If you want to reproduce the whole article in a third-party publication (excluding your thesis/dissertation for which permission is not required) please go to the Copyright Clearance Center request page.
Read more about how to correctly acknowledge RSC content.
 Fetching data from CrossRef.
Fetching data from CrossRef.
This may take some time to load.
Loading related content
