 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Microkinetic characterization of ROS-driven methane production through quantum and classical simulations
Ronghan Zhang *ab,
Zexuan Wang
*ab,
Zexuan Wang *ab,
Kin Tung Michael Ho
*ab,
Kin Tung Michael Ho a,
Nigel J. D. Graham
a,
Nigel J. D. Graham a,
Giin-Yu Amy Tan
a,
Giin-Yu Amy Tan *b and
Po-Heng Henry Lee
*b and
Po-Heng Henry Lee *a
*a
aDepartment of Civil and Environmental Engineering, Imperial College London, London, SW7 2AZ, UK. E-mail: po-heng.lee@imperial.ac.uk
bDepartment of Architecture and Civil Engineering, City University of Hong Kong, Hong Kong SAR, China. E-mail: rzhang343-c@my.cityu.edu.hk; zexuawang4-c@my.cityu.edu.hk; amytan@cityu.edu.hk
First published on 8th April 2026
Abstract
Methane is ubiquitous in natural and engineered settings, but the mechanisms of reactive oxygen species (ROS)-driven methane production in living organisms remain unclear. Unraveling the interactions and microkinetics among ROS, iron species, and organic methyl groups (MET) is crucial for reducing emissions and optimizing energy recovery. In this study, the quantum Harrow–Hassidim–Lloyd (HHL) algorithm and classical Levenberg–Marquardt (LM) algorithm were employed to: (1) investigate the primary pathway of Fe3+ and ascorbic acid (ASC) reactions, (2) determine the kinetic parameters for the reaction between [FeIV![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ and MET, and (3) simulate the ROS-driven methane production pathway under specific abiotic conditions. The results revealed that Fe3+ and ASC are primarily responsible for the redox reaction, with a kinetic rate constant of 1.83 × 10−4 M−1 s−1 for the reaction between [FeIV
O]2+ and MET, and (3) simulate the ROS-driven methane production pathway under specific abiotic conditions. The results revealed that Fe3+ and ASC are primarily responsible for the redox reaction, with a kinetic rate constant of 1.83 × 10−4 M−1 s−1 for the reaction between [FeIV![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ and MET, and a ∼10% conversion rate of MET to methane. Our study establishes a theoretical framework for ROS-driven methane production. While identifying a feasible pathway for engineered and natural systems, further investigation and validation are needed to quantify its specific contribution in complex environments. Furthermore, this work demonstrates the potential of hybrid quantum-classical simulations for microkinetic analysis.
O]2+ and MET, and a ∼10% conversion rate of MET to methane. Our study establishes a theoretical framework for ROS-driven methane production. While identifying a feasible pathway for engineered and natural systems, further investigation and validation are needed to quantify its specific contribution in complex environments. Furthermore, this work demonstrates the potential of hybrid quantum-classical simulations for microkinetic analysis.
Environmental significanceMethane is a double-edged molecule—an important renewable energy source yet a potent greenhouse gas. This study elucidates a previously unclear abiotic and biotic mechanism of reactive oxygen species (ROS)-driven methane formation, revealing how cellular redox reactions involving Fe3+, ascorbic acid, and methylated compounds can generate methane even in oxygenated environments. By integrating quantum (Harrow–Hassidim–Lloyd) and classical (Levenberg–Marquardt) microkinetic simulations, we quantify key rate constants and demonstrate that ferryl ion reactions, rather than hydroxyl radicals, dominate ROS-mediated methane production. These insights advance understanding of methane cycling in both engineered and natural systems, offering new directions for optimising micro-aerated anaerobic digestion and predicting methane emissions from oxygen-exposed ecosystems under changing redox and oxygen conditions. |
1. Introduction
As global warming intensifies, it is of great significance to develop clean energy that can replace fossil fuel energy.1 Methane is a renewable, clean energy source, albeit also a potent greenhouse gas. Harnessing methane production and collection for energy purposes presents a promising approach to mitigate greenhouse gas emissions. Methane production through anaerobic digestion (AD) can be enhanced by 10–30% using micro-aeration (2.63–37.5 mL O2/LR d, where LR is the reactor working volume), which improves hydrolysis and generates reactive oxygen species (ROS) (0.27–0.29 mM H2O2 per cycle.2–8 Contrary to popular belief that methane can only be produced under anaerobic conditions, enhanced methane formation has been observed in natural systems in the presence of oxygen, including oxygen-rich wetlands and oceans, where the addition of oxygen increased methane emissions by 4–10 fold.9,10 The ability to produce methane appears to be deeply rooted in evolution and present in all living organisms. Through coupling oxygen-generated ROS with methane production as part of an organism’s innate oxidative stress mechanism, this mitigates ROS-induced oxidative damage. For example, Bacillus subtilis and Escherichia coli showed a 1.2–3.2 times methane yield increases in the presence of ROS as compared to that of non-ROS condition.3 Leveraging on these emerging insights can provide new avenues for mitigating emissions and improving renewable energy harvesting. However, the precise mechanisms and micro-kinetics underlying the role of ROS in facilitating methane production remain poorly understood.ROS-driven methane production has been linked to the presence of methyl group-containing organosulfur compounds, ROS, Fe2+and reducing agent ascorbic acid (ASC) (Fig. 1a).3,11 Fe2+ and H2O2 (a form of ROS) can undergo Fenton reaction to generate Fe3+, hydroxyl radicals (˙OH) and ferryl ions ([Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+). [Fe(IV)
O]2+). [Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ and ˙OH can react with methyl-containing organosulfur compounds to form methane.11 The Fe3+ produced can be reduced by ASC to regenerate Fe2+, sustaining the Fenton reaction. It is expected that these chemical species are present in natural and AD-engineered systems. Iron, an active metal predominantly coordinated in heme groups and iron-sulfur clusters that drive electron transport chains and participates in redox reactions, is widely present in living organisms including microorganisms.12 Exposure of microbes to oxygen often produces ROS, which damage cellular structures and impair metabolisms.13 ASC may play a crucial role in cellular protective mechanisms from ROS damage by facilitating iron and ROS redox reactions, thereby producing methane as a byproduct. However, there is a discrepancy in the proposed mechanisms for the interaction between ASC and iron species (Fig. 1b). Here, two competing mechanisms are possible: a catalytic mechanism proposed by Buettner (eqn (1)) and a redox mechanism proposed by Buettner and Jurkiewicz (eqn (2)),14,15 as shown below:
O]2+ and ˙OH can react with methyl-containing organosulfur compounds to form methane.11 The Fe3+ produced can be reduced by ASC to regenerate Fe2+, sustaining the Fenton reaction. It is expected that these chemical species are present in natural and AD-engineered systems. Iron, an active metal predominantly coordinated in heme groups and iron-sulfur clusters that drive electron transport chains and participates in redox reactions, is widely present in living organisms including microorganisms.12 Exposure of microbes to oxygen often produces ROS, which damage cellular structures and impair metabolisms.13 ASC may play a crucial role in cellular protective mechanisms from ROS damage by facilitating iron and ROS redox reactions, thereby producing methane as a byproduct. However, there is a discrepancy in the proposed mechanisms for the interaction between ASC and iron species (Fig. 1b). Here, two competing mechanisms are possible: a catalytic mechanism proposed by Buettner (eqn (1)) and a redox mechanism proposed by Buettner and Jurkiewicz (eqn (2)),14,15 as shown below:
| ASC + Fe3+ + O2 = Fe3+ + DHA + H2O2 | (1) |
| ASC + Fe3+ = Fe2+ + A˙− + 2H+ | (2) |
Models proposed by Shen et al. showed that the catalysis degree of the reaction was 4 to 6-fold greater than that of the redox reaction.16 However, experimental data from Elmagirbi et al. indicated that 76% of Fe3+ was converted to Fe2+ in the presence of ASC, suggesting that only 24% of Fe3+ may participate in the catalyst reaction.17 Currently, it is unclear which pathway, whether redox or catalysis, plays a more significant role in promoting ROS-activated microkinetics. If the redox reaction between ASC and Fe3+ dominates, it can produce Fe2+ which promotes the ROS-driven methane production pathway to mitigate oxidative cell damage. In contrast, if the catalytic reaction is dominant, it will produce additional H2O2, which can cause cell damage and inhibit the methane production pathway.
Overall, the interaction between ASC and iron species drives two competing loops (Fig. 1c). The catalytic pathway (blue loop) regenerates H2O2, potentially leading to ROS accumulation. In contrast, the redox pathway (orange loop) reduces Fe3+ to Fe2+, driving the Fenton reaction to generate [Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ and hydroxyl radicals. These intermediates subsequently oxidize methyl-containing organosulfur compounds to produce methane while regenerating Fe3+ for the next cycle. Consequently, determining the dominant microkinetic pathway is essential to distinguish between the ROS regeneration and the methane production loop. Based on this, we hypothesize that methane production proceeds via ferryl ions and methyl group-containing organosulfur compounds such as methionine (MET), through the pathway described in eqn (9) (Table 2).
O]2+ and hydroxyl radicals. These intermediates subsequently oxidize methyl-containing organosulfur compounds to produce methane while regenerating Fe3+ for the next cycle. Consequently, determining the dominant microkinetic pathway is essential to distinguish between the ROS regeneration and the methane production loop. Based on this, we hypothesize that methane production proceeds via ferryl ions and methyl group-containing organosulfur compounds such as methionine (MET), through the pathway described in eqn (9) (Table 2).
The ROS regeneration loop may produce H2O2 continuously and cause cell damage, however, the generation of ˙OH and [Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ as intermediates in the Fenton-driven methane production loop may contribute to methane production. Addressing the counteracting effects of the ROS regeneration loop and the Fenton-driven methane production loop would require the preferred reaction for ROS-driven methane production microkinetics to be established. To this end, microkinetic simulation was used to determine which reaction cycle (the orange, or blue), or both reaction cycles shown in Fig. 1c is more favorable.
O]2+ as intermediates in the Fenton-driven methane production loop may contribute to methane production. Addressing the counteracting effects of the ROS regeneration loop and the Fenton-driven methane production loop would require the preferred reaction for ROS-driven methane production microkinetics to be established. To this end, microkinetic simulation was used to determine which reaction cycle (the orange, or blue), or both reaction cycles shown in Fig. 1c is more favorable.
Microkinetic simulation can be conducted using both quantum and classical computing algorithms, each offering its own set of advantages. Most recently, the Harrow Hassidim and Lloyd (HHL) quantum algorithm has been successfully applied in solving linear microkinetic matrices.18,19 The Levenberg–Marquardt (LM) classical algorithm has already been used to determine the microkinetic parameters with nonlinear matrices.20–22 These studies have demonstrated the versatility of microkinetic simulation and its potential to couple both quantum and classical computing approaches.
In the context of ROS-driven methane production microkinetics, the mass balance of the reaction between ASC and Fe3+ can be formulated as a system of linear equations, whereas that of the associated reactions between ferryl ion and MET can be modelled non-linearly. This study, thus, aims to achieve three objectives: (1) determine the microkinetic model between ASC and Fe3+ using the quantum HHL algorithm, (2) employ the classical LM algorithm to calculate the microkinetics between ferryl ion and MET, and (3) simulate the ROS-driven methane-forming microkinetics using the HHL algorithm. The quantum HHL simulation results were benchmarked against classical computations, with further validation conducted against existing experimental data from relevant studies. Additionally, the microkinetics of ROS scavenging, ASC, oxygen, and iron species for ROS-driven methane production will be discussed. The findings of this study will advance our understanding of ROS-driven methane production in natural and engineered systems. Moreover, the insights for future research studies associated with applying quantum and classical computing simulations in microkinetic models are addressed.
2. Methodology
2.1 Simulation parameters and assumptions
To develop our microkinetic simulation, we utilized experimental data from Althoff et al., who investigated abiotic methane formation from organosulfur compounds (MET) in the presence of iron, ascorbate, and H2O2.11,23 We adopted their reported reactant concentrations and environmental conditions to define the initial settings for our computational model (Table 1).| Reactant | Concentration (mmol L−1) |
|---|---|
| ASC | 50 |
| Ferric ion | 40 |
| MET | 0.5 |
| Hydrogen peroxide | 100 |
| Dissolved oxygen | 160 |
| Condition | Value |
|---|---|
| pH value | 3 |
| Temperature | 25 °C |
| Reaction time | 30 h |
| Reaction volume | 1 mL |
In this simulation, a pH of 3 was selected for the following reasons: (1) previous research indicated a maximum ROS-driven methane production at pH = 3 based on experiments conducted in the pH range of 1–9.11 (2) In our case, ROS-driven methane production in E. coli likely occurs in the E. coli periplasmic cell envelope, where its pH is in the range of pH = 1–3.24 Since such a pathway is applicable to all organisms, it is plausible to have it in the E. coli periplasm.3 Moreover, ASC formation by E. coli has been reported previously, indicating its capability to survive in acidic conditions.25 Furthermore, E. coli, possesses regulatory hydrogenase that can effectively ensure its survival under lower pH conditions (pH ≥ 2).26 (3) Some archaeal species can still be active even at pH = 1–2.27 (4) The cytoplasmic proteins of acetogens remain stable under extremely acidic conditions.28 Therefore, ROS-driven methane production pathways may occur in organisms at low pH.
Table 2 provides a summary of the stoichiometry, reaction kinetics, and thermodynamic constants used for the microkinetic simulations. Here, eqn (1)–(7) were used to describe the interactions between different reactants. The chemical reactions eqn (1)–(4) represent the catalytic and redox reactions between ASC and Fe3+, as well as the interactions between the corresponding products. Eqn (5) and (6) reflect the products of the two Fenton reactions, namely ˙OH and [Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+. Finally, eqn (7) reflects the role of ASC in scavenging ˙OH.
O]2+. Finally, eqn (7) reflects the role of ASC in scavenging ˙OH.
| Reaction | Kinetic parameter (k) (M−1 s−1) | Reference | Gibbs free energy (ΔG0) (kcal mol−1) | Reference | |
|---|---|---|---|---|---|
| a The ΔG0–1 was calculated using the relationship ΔG0 = −nF Ecell, where n = 2 and F is the Faraday constant. Ecell is the change of total reaction reduction potential, which was obtained between E(O2/H2O2) = +0.28V and E(dehydroascorbate/ascorbate) = +0.06V.32,33b The ΔG value was calculated based on the standard Gibbs free energies of formation ΔGf for the individual chemical species.35,42,43c Calculated using the equilibrium constant (K) estimated from the MATLAB kinetic simulation.d Refer to Section 2.1 and SI Table 1.e Enzymatic scavengers (SOD, Catalase) are excluded in this study. Their presence in vivo regulates ROS levels but does not alter the fundamental role of Ferryl iron in driving methane formation. | |||||
| (1) | ASC + Fe3+ + O2 = DHA + Fe3+ + H2O2 | k1 = 4.0 × 105 | 31 | ΔG0–1 = −10.2 | 32 and 33 a |
| (2) | ASC + Fe3+ = Fe2+ + 2H+ + A˙− | k2 = 4.5 × 103 | 34 | ΔG0–2 = 1.1 | 35 |
| (3) | Fe2+ + O2 = Fe3+ + O2˙− | k3 = 1.0 × 10−5 | 36 | ΔG0–3 = −91.6 | 37 |
| (4) | A˙− + O2˙− + H2O = DHA + HO2˙− + OH− | k4 = 2.6 × 108 | 38 | ΔG0–4 = 38.4 | 39 |
| (5) | Fe2+ + H2O2 = Fe3+ + ˙OH + OH− | k5 = 5.7 × 102 | 40 | ΔG5 = 28.7 | b |
| (6) | Fe2+ + H2O2 = [FeIV![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ + H2O O]2+ + H2O |
k6 = 1.0 × 106 | 41 | ΔG6 = −14.9 | c |
| (7) | ASC + ˙OH = A˙− + H2O + H+ | k7 = 7.9 × 109 | 16 | ΔG7 = −30.4 | b |
| (8) | ASC + O2 = 2H+ + O2˙− + A˙− | — | d | — | d |
| (9) | 2[FeIV![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ + MET + H2O2 + 2H+ = 2Fe3+ + HCyA + CH4 + H2O O]2+ + MET + H2O2 + 2H+ = 2Fe3+ + HCyA + CH4 + H2O |
— | d | — | d |
| (10) | ASC + Fe2+ + H2O2 = Fe3+ + A˙− + 2H2O | — | d | — | d |
To establish the microkinetic model, several assumptions were made: (1) the molar concentration of DHA in eqn (1) and (4) shown in Table 2 was taken to be the same as the concentration of HO2˙− and H2O2, according to the stoichiometric ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1; (2) the concentration of DHA has a linear relationship with Fe3+;29 (3) the concentration of O2˙− is 3.9 mM, using a conversion rate of 1.5% for [O2] to [O2˙−] according to previous literature;30 (4) eqn (2) and (3) can be combined as eqn (8) in Table 2, and since eqn (3) is the rate limiting reaction, the kinetic reaction rate constant can be taken as k3, (5) the interaction pathway between [FeIV
1; (2) the concentration of DHA has a linear relationship with Fe3+;29 (3) the concentration of O2˙− is 3.9 mM, using a conversion rate of 1.5% for [O2] to [O2˙−] according to previous literature;30 (4) eqn (2) and (3) can be combined as eqn (8) in Table 2, and since eqn (3) is the rate limiting reaction, the kinetic reaction rate constant can be taken as k3, (5) the interaction pathway between [FeIV![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ and MET, as proposed by Althoff et al., can be summarized as eqn (9) in Table 2.11 (6) Reactions eqn (5) and (7) can be combined as eqn (10) in Table 2, and the kinetic reaction rate constant can be taken as k5 (rate limiting).
O]2+ and MET, as proposed by Althoff et al., can be summarized as eqn (9) in Table 2.11 (6) Reactions eqn (5) and (7) can be combined as eqn (10) in Table 2, and the kinetic reaction rate constant can be taken as k5 (rate limiting).
2.2 Kinetic estimation using HHL algorithm
In this study, the HHL algorithm was used to simulate microkinetic models to determine the mechanism behind ASC, Fe3+, and the ROS-driven methane production (redox and/or catalysis pathway). We constructed the model according to the chemical equilibrium principle that the rates of forward reaction and reverse reaction are equivalent. The forward reaction rate constant (k) and Gibbs free energy (ΔG0) used are shown in Table 2 and in the SI Table 1. Detailed calculations are shown in the SI Note 1.To investigate the mechanism for ROS-driven methane production microkinetics in non-methanogenic microbes at the conditions listed in Table 1, the following system of linear equations was built.
| A·x = b | (11) |
The HHL algorithm requires matrix A to be Hermitian, necessitating the transformation of A into matrix AH.
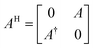 | (12) |
After the conversion of matrix (A) of eqn (11) into the Hermitian matrix (AH) in eqn (12), the unknown term x was calculated using eqn (13) in the HHL algorithm:
| x = b × AH−1 | (13) |
Hence, to investigate whether the redox and/or catalysis pathway drives the reaction between ASC and Fe3+, a microkinetic linear matrix model (eqn (14)) was constructed based on eqn (1), (4) and (8) at steady state. The procedure for establishing eqn (14) is shown in the SI Note 2.
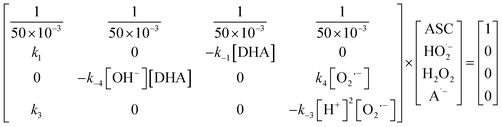 | (14) |
To investigate the ROS-driven methane production pathway, a microkinetic linear matrix (eqn (15)) was built initially to simulate the probability of the Fenton reaction forming [Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ to participate in the ROS-driven methane production pathway. Eqn (15) was constructed based on reactions eqn (2), (6) and (10) at steady state. The process for establishing eqn (15) is given in the SI Note 3.
O]2+ to participate in the ROS-driven methane production pathway. Eqn (15) was constructed based on reactions eqn (2), (6) and (10) at steady state. The process for establishing eqn (15) is given in the SI Note 3.
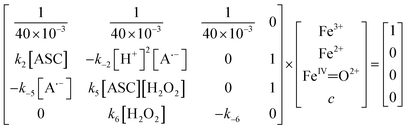 | (15) |
To investigate the methane production of MET oxidated [Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ in the ROS-driven methane production pathway, a microkinetic linear matrix model (eqn (16)) was constructed based on the eqn (9) reaction at steady state. According to the stoichiometric ratio in the eqn (9) reaction, the production of HCyA represents methane production. The process for establishing eqn (16) is shown in the SI Note 3.
O]2+ in the ROS-driven methane production pathway, a microkinetic linear matrix model (eqn (16)) was constructed based on the eqn (9) reaction at steady state. According to the stoichiometric ratio in the eqn (9) reaction, the production of HCyA represents methane production. The process for establishing eqn (16) is shown in the SI Note 3.
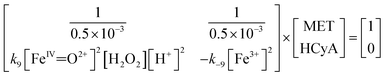 | (16) |
2.3 Kinetic estimation using LM algorithm
The reaction rate constant between MET and [Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ was determined using mass balances derived from reaction eqn (2), (5), (6), (7) and (9). The resulting non-linear matrix (eqn (17)) was solved using MATLAB with a LM algorithm. Further information about the matrix development is given in SI Note 4.
O]2+ was determined using mass balances derived from reaction eqn (2), (5), (6), (7) and (9). The resulting non-linear matrix (eqn (17)) was solved using MATLAB with a LM algorithm. Further information about the matrix development is given in SI Note 4.
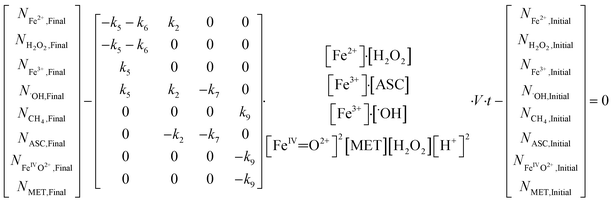 | (17) |
3. Results
3.1 Mechanistic determination analysis
Table 3 summarizes the simulated concentrations of the reaction products, [HO2˙−], [A˙−] and [H2O2] associated with ASC and Fe3+ participation in redox and catalytic reactions, respectively. The results obtained using the quantum HHL algorithm showed good agreement with the classical MATLAB simulation. Notably, regarding the catalytic product [HO2˙−], the classical simulation yielded a value of ∼10−20 mol L−1. It is important to clarify that this non-zero value represents a numerical artifact (asymptotic residue) inherent to classical ODE solvers using floating-point arithmetic, rather than a physical concentration. In contrast, the quantum simulation reported a concentration of 0.00 mol L−1, correctly identifying the absence of this species within the algorithmic resolution.| Unknown parameter | Estimated values (quantum) | Estimated values (classical) | Remarks | Error |
|---|---|---|---|---|
| a Classical results include a numerical artifact from ODE solvers. The residual values (∼10−7) for trace species are negligible in representing physical concentrations.b Products of catalytic/redox reaction indicated in eqn (14).c k9 is the kinetic parameter for eqn (11).d The proportion of iron ions involved in Fenton reaction.e Methane production.f Simulated conversion rate (η) of MET to methane in Google Quantum AI.g Experimentally measured conversion rate (η) of MET to methane conducted by Althoff et al.11 | ||||
| Mechanistic determination by Google Quantum AI & Classical MATLAB | ||||
| [ASC] | 0.00 (mol L−1) | 0.00 (mol L−1) | — | 1.56 × 10−20 |
| [HO2˙−] | 0.00 (mol L−1) | 1.56 × 10−20 (mol L−1) | Redox reactionb | |
| [H2O2] | 0.00 (mol L−1) | 0.00 (mol L−1) | Catalytic reactionb | |
| [A˙−] | 5.00 × 10−2 (mol L−1) | 5.00 × 10−2 (mol L−1) | Redox reactionb | |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Kinetic reaction rate constant determination by Classical MATLAB | ||||
| k9c | — | 1.83 × 10−4 (M−1 s−1) | — | — |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Microkinetic simulation by Google Quantum AI & Classical MATLAB | ||||
| [Fe3+] | 0.00 (mol L−1) | 2.93 × 10−7 (mol L−1) | 0d | 4.14 × 10−7 |
| [Fe2+] | 0.00 (mol L−1) | 1.20 × 10−11 (mol L−1) | 0d | |
[FeIV![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O2+] O2+] |
4.00 × 10−2 (mol L−1) | 4.00 × 10−2 (mol L−1) | 100%d | |
| [MET] | 4.71 × 10−4 (mol L−1) | 4.71 × 10−4 (mol L−1) | — | 1.23 × 10−12 |
| NMethanee | 4.71 × 10−8 mol | 4.71 × 10−8 mol | — | |
| Conversion ratef | 9.42% | This work | — | |
| Conversion rateg | 10.00% | Althoff et al.11 | — | |
Consequently, both simulations confirmed that nearly 100% of the reaction proceeded via the redox pathway. This result was consistent with the fact that the Gibbs free energy for eqn (8) (ΔG = −90.5 kcal mol−1) is significantly more negative than that for eqn (1) (ΔG = −10.2 kcal mol−1), indicating a strong thermodynamic preference for the redox pathway. In the presence of oxygen, ASC, and Fe3+, A˙− generation by eqn (8) is more favorable. Eqn (8) is dependent on the redox reaction between ASC and Fe3+ occurring, rather than the catalytic reaction. These findings aligned well with the results of the studies conducted by Timoshnikov et al. and Buettner & Jurkiewicz, which showed the high degree of Fe3+ reduction by ASC under aerobic conditions and at a pH of 3.15,44 The simulation results showed that ASC is beneficial for promoting methane production in the Fenton-driven methane production loop. Thus, the redox reaction between ASC and Fe3+ plays a decisive role in tilting the microkinetics towards the ROS-driven methane production pathway.
3.2 Kinetic reaction rate constant determination
The kinetic rate constant k9 in the matrix (eqn (17)) was derived as 1.83 × 10−4 M−1 s−1 (Table 3) by implementing the LM algorithm in MATLAB. Comparing the reactions represented by eqn (6) and (9), the reaction rate constant (k9) between [Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ and MET is approximately ten orders of magnitude lower than the reaction rate constant (k6) of the Fenton reaction to produce [Fe(IV)
O]2+ and MET is approximately ten orders of magnitude lower than the reaction rate constant (k6) of the Fenton reaction to produce [Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+, implying that the reaction of [Fe(IV)
O]2+, implying that the reaction of [Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ and MET is the rate-determining step in the ROS-driven methane production pathway. SI Table 2 provides a summary of the MATLAB output results.
O]2+ and MET is the rate-determining step in the ROS-driven methane production pathway. SI Table 2 provides a summary of the MATLAB output results.
3.3 Microkinetic simulation
The results derived from the execution of eqn (15) matrix on Google Quantum AI platform were consistent with classical MATLAB computations (Table 3). It is evident that through the Fenton reaction, iron is prone towards ferryl ion formation (4.00 × 10−2 (mol L−1), with near 100% conversion rate from iron to ferryl ion. This result matched the findings of Deguillaume et al., which showed that at a low pH of 3.5, the concentration of ferryl ion produced by the reaction of Fe2+ and H2O2 was 104 times greater than the concentration of ˙OH produced.45 From the microkinetics perspective, the kinetic reaction rate constant between Fe3+ and H2O2 to produce ferryl ion (k6) is 104 times greater than that of ˙OH formation (k5) in the ROS-driven methane production process. Fe3+ (k2) and ˙OH (k7) exhibit a greater susceptibility to reduction by ASC as compared to k5. As a result, the ferryl ion serves as a primary reactant with MET to produce methane.According to the stoichiometric ratio in eqn (9), the product concentration of HCyA is directly proportional to the methane produced. Therefore, methane production can be estimated using eqn (16) through the HHL algorithm. In addition, the conversion rate (η) of MET to methane can be calculated. The quantum-simulated conversion rate (η) was 9.42% (Table 3), showing strong agreement with the classical calculation (10.00%) and the experimental value reported by Althoff et al.11 The detailed calculations are shown in SI Note 5.
4. Discussion
4.1 Effect of radical kinetics and oxygen levels for methane production
An investigation of the ˙OH concentration in cells is of paramount importance. In the ROS-driven methane production pathway studied in this work (conditions presented in Table 1), ˙OH produced by the Fenton reaction (Fig. 1c), can either react with methyl compounds to form methane or be scavenged by ASC. ASC reportedly has a strong affinity for ˙OH with a high reaction rate of 7 × 109 M−1 s−1.46 Meanwhile, the reaction rate constant of ˙OH attacking intracellular DNA is 3 × 108 M−1 s−1.47 This difference in the order of magnitude indicates that the scavenging effect of ASC occurs at a higher rate than ˙OH-induced cellular damage. As a result, it is proposed that ROS in the methane production pathway does not cause significant damage to cells. Furthermore, the reaction rate constant of methyl-containing compounds and ˙OH is 1.5 × 107 M−1 s−1, which is notably lower than the scavenging rate of ˙OH by ASC (7 × 109 M−1 s−1).48 Therefore, the contribution of ˙OH may not be as significant as that of ferryl ions in the ROS-driven methane production pathway (Fig. 2). | ||
| Fig. 2 Schematic indicating that ˙OH (the product of the Fenton reaction) does not participate in the ROS-driven CH4 production loop and damages intracellular DNA. ka is the reaction rate constant between ASC and ˙OH;46 kb is the reaction rate constant between DNA and ˙OH;47 kc is the reaction rate constant between methyl-containing compounds and ˙OH.48 | ||
Our findings mechanistically demonstrate the feasibility of a ROS-driven methane production pathway under the specific abiotic conditions modeled. This pathway offers a theoretical framework to interpret methane enhancement phenomena not only in micro-aeration anaerobic digestion (MAAD), but also in natural systems, such as wetlands.6,49 While these systems can synthesize H2O2 under oxygen (∼21 µM in wetlands vs. 270–290µM in MAAD), which reacts with Fe2+ to promote the generation of ferryl ions and facilitates the conversion of MET to methane.8,50,51 Extrapolating these microkinetic results to complex environmental matrices introduces inherent uncertainties. Factors such as pH variations, organic ligand complexation, and microbial interactions in natural ecosystems may significantly alter the reaction kinetics compared to our simplified model. Recent studies suggest that ROS can promote Fenton reaction in bacteria and archaea, generating ferryl ions that oxidize methyl-containing donors.3 These biological observations align with our mechanistic findings; however, distinguishing the specific contribution of this ROS-driven pathway from canonical enzymatic processes in diverse oxygen environments requires further in situ validation to bridge the gap between idealized modeling and ecosystem complexity.
However, excessively high oxygen levels may negatively impact the overall methane production in these systems. For instance, in cyanobacteria, the conversion rate of organic MET is affected when the oxygen concentration is close to that of air-saturated water.23,50 Higher intracellular oxygen levels have been shown to lower the conversion rates for both MET (about 10%) and [FeIV![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+.11 In addition, Althoff et al. reported a lack of CH4 formation in an oxygen-saturated environment, most likely due to oxygen’s adverse effects on ROS-driven methane production.23 This might be attributed to the reaction between the methyl radical (˙CH3), the methane precursor, and O2, forming methylperoxyl (CH3OO˙) with a significantly high reaction rate constant of 3.7 × 109 M−1 s−1.51 The formation of methylperoxyl reduces the production of methane. This rate constant for the reaction between ˙CH3 and O2 is notably larger than the calculated rate constant for the methane production reaction of [FeIV
O]2+.11 In addition, Althoff et al. reported a lack of CH4 formation in an oxygen-saturated environment, most likely due to oxygen’s adverse effects on ROS-driven methane production.23 This might be attributed to the reaction between the methyl radical (˙CH3), the methane precursor, and O2, forming methylperoxyl (CH3OO˙) with a significantly high reaction rate constant of 3.7 × 109 M−1 s−1.51 The formation of methylperoxyl reduces the production of methane. This rate constant for the reaction between ˙CH3 and O2 is notably larger than the calculated rate constant for the methane production reaction of [FeIV![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ with MET (1.83 × 10−4 M−1 s−1) investigated in this study. Aside from ROS-driven methane production, oxygen overload also affects methanogenesis by creating an unfavorable environment for methanogens’ survival, lowering the overall methane production rate.2 Hence, ascertaining the oxygen concentration threshold required for striking a balance between ROS-induced and methanogen-based methane production would be advantageous for engineered systems such as MAAD seeking enhanced methane production as well as the control and mitigation of methane emission from natural ecosystems. Future work must bridge the gap between this idealized microkinetic model and in situ environmental conditions to validate these broader implications.
O]2+ with MET (1.83 × 10−4 M−1 s−1) investigated in this study. Aside from ROS-driven methane production, oxygen overload also affects methanogenesis by creating an unfavorable environment for methanogens’ survival, lowering the overall methane production rate.2 Hence, ascertaining the oxygen concentration threshold required for striking a balance between ROS-induced and methanogen-based methane production would be advantageous for engineered systems such as MAAD seeking enhanced methane production as well as the control and mitigation of methane emission from natural ecosystems. Future work must bridge the gap between this idealized microkinetic model and in situ environmental conditions to validate these broader implications.
4.2 Perspectives and limitations of microkinetic using quantum computing
Quantum computing incorporates the superposition and entanglement properties of quantum mechanics and can surpasses classical computation in analyzing and quantifying information under certain conditions. Notably, it can efficiently simulate the energy consumption of the Haber–Bosch process (1–2% of global energy consumption), an energy-intensive process, by analyzing the underlying chemical reaction mechanisms.52 Specifically, the HHL algorithm enables the implementation of chemical microkinetic models on quantum computers, for processes such as CO oxidation, aiding in the resolution of chemical kinetics-related issues.19,53 Furthermore, Becerra et al. used the same quantum algorithm to quantify the uncertainties in Rh(III)-catalyzed CO oxidation pathways, which classical computation struggles to calculate. This emerging trend suggests that quantum computing may be a promising technique for simulating chemical reaction mechanisms and microkinetic models.18,54 It foreshadows the potential mainstream adoption of quantum simulation for predicting molecular properties, activation energies, and reaction rate constants.55,56While classical solvers remain efficient for these small-scale systems, this study establishes a prospective algorithmic advantage. By replacing classical polynomial scaling (e.g., O(N3) for matrix inversion), the HHL algorithm offers an exponential speedup with logarithmic scaling (O(log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) N)).57 To illustrate this, for a hypothetical large-scale network with N = 106 species, classical computational cost would surge to 1018 operations, whereas the quantum complexity grows only to the order of log(106) (approx. 20). Nevertheless, this scaling analysis indicates that quantum solvers will be indispensable for future large-scale biochemical reaction network simulations, where high dimensionality renders classical approaches computationally prohibitive. This capability will enable the study of more complex environments for further investigation and related systems in the future.
N)).57 To illustrate this, for a hypothetical large-scale network with N = 106 species, classical computational cost would surge to 1018 operations, whereas the quantum complexity grows only to the order of log(106) (approx. 20). Nevertheless, this scaling analysis indicates that quantum solvers will be indispensable for future large-scale biochemical reaction network simulations, where high dimensionality renders classical approaches computationally prohibitive. This capability will enable the study of more complex environments for further investigation and related systems in the future.
Variations in the oxygen content of organisms may have an impact on ROS concentration, thereby influencing variations in the methane content.9 Consequently, this study investigated the impact of oxygen concentration in the range of 0–1000 mM in the models. The concentration of O2˙− in the initial model is influenced by the oxygen content, consequently affecting the relative importance of the redox and catalysis reactions. However, the results have shown that the redox reaction dominated within this oxygen concentration range, as detailed in SI Fig. 1. For the microkinetic simulation of ROS-induced methane production modelling, alterations in oxygen concentration ought to influence H2O2. However, the model’s foundation is based on the chemical experimental data presented by Althoff et al.11,23 Hence, the H2O2 concentration remained constant throughout this study. Furthermore, the proportional relationship between the conversion of oxygen to H2O2 within living organisms remains unclear and it is not known how much oxygen can be converted into H2O2 within a cell. A comprehensive investigation into the influence of oxygen concentration on methane production in methanogenic pathways driven by ROS is thus unattainable. Moreover, the simulation of this work was largely based on the experimental data of Althoff et al., obtained through the reactions of pure chemicals at pH 3.11 There is a lack of research on ROS-driven methane production in organisms at higher biological pH. In nature and environmental systems such as anaerobic digesters, which operate at circumneutral pH (6.5–8.0), iron speciation differs significantly, often involving precipitation or complexation with organic ligands.58 These structural changes alter reaction energy barriers and kinetic rate constants (k). To extend our algorithmic framework to these conditions, we propose a hybrid computational strategy: theoretical methods like Density Functional Theory (DFT) or the Variational Quantum Eigensolver (VQE) can be employed to calculate the specific activation energies and rate constants for iron-ligand complexes at neutral pH.59,60 Additionally, experimental techniques such as Electron Paramagnetic Resonance (EPR) spectroscopy could be integrated in future studies to directly detect the transient ferryl iron intermediates predicted by our model.61 These derived parameters can then serve as updated inputs for the HHL solver, ensuring the model’s applicability across diverse biochemical settings.
From a biological perspective, fermentative bacteria, which are known for their ability to survive and function well in a wide pH range (4–8), could serve as better candidates to study this phenomenon.62 The intracellular pH is generally lower than the extracellular pH, reaching 3.5 to 4.5.63 Methods to study methane production in vivo such as microbial electro-methanogenesis, may be used to explore ROS-driven methane production at pH 3.5 to 7.5 in future investigations.64
5. Conclusion
This study employed a hybrid quantum-classical computational framework to investigate the microkinetics of ROS-driven methane production. The results indicate that the redox reaction between ASC and Fe3+ is favored over the catalytic pathway, thereby supporting continued Fenton-driven methane production through Fe2+ regeneration. The reaction between [Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O]2+ and MET was identified as the rate-determining step, and the quantum simulated MET to methane conversion rate showed good agreement with current experimental observations, supporting the proposed mechanistic pathway.
O]2+ and MET was identified as the rate-determining step, and the quantum simulated MET to methane conversion rate showed good agreement with current experimental observations, supporting the proposed mechanistic pathway.
These findings identify a feasible abiotic pathway for ROS-driven methane production and provide a theoretical framework for interpreting methane enhancement phenomena in micro-aeration anaerobic digestion and potentially in natural systems such as wetlands. More broadly, this work demonstrates the potential of the HHL quantum algorithm for microkinetic modelling, particularly in larger biochemical reaction networks where classical computation may become challenging. Further experimental validation under more complex biological conditions will be important for extending the applicability of this framework.
Author contributions
Ronghan Zhang and Zexuan Wang: conceptualization, methodology, software, formal analysis, investigation, writing – original draft, writing – review & editing. Kin Tung Michael Ho: conceptualization, writing – review & editing. Nigel J. D. Graham: writing – review & editing. Giin-Yu Amy Tan: writing – review & editing. Po-Heng Lee: supervision, conceptualization, writing – review & editing.Conflicts of interest
The authors declare no competing financial interest.Data availability
The source code and the detailed model-building procedure are available in the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5va00420a.Acknowledgements
The authors gratefully acknowledge Prof. Aleksandra Szczuka from the Department of Civil and Environmental Engineering, University of Michigan, Ann Habour, for her valuable input in modelling and reviewing the paper.References
- S. Biswas, A. P. Kulkarni, S. Giddey and S. Bhattacharya, A review on synthesis of methane as a pathway for renewable energy storage with a focus on solid oxide electrolytic cell-based processes, Front. Energy Res., 2020, 8, 570112 CrossRef.
- N. Yu, B. Guo, Y. Zhang, L. Zhang, Y. Zhou and Y. Liu, Different micro-aeration rates facilitate production of different end-products from source-diverted blackwater, Water Res., 2020, 177, 115783 CrossRef CAS PubMed.
- L. Ernst, B. Steinfeld, U. Barayeu, T. Klintzsch, M. Kurth, D. Grimm, T. P. Dick, J. G. Rebelein, I. B. Bischofs and F. Keppler, Methane formation driven by reactive oxygen species across all living organisms, Nature, 2022, 603, 482–487 CrossRef CAS PubMed.
- S. Montalvo, F. Ojeda, C. Huiliñir, L. Guerrero, R. Borja and A. Castillo, Performance evaluation of micro-aerobic hydrolysis of mixed sludge: Optimum aeration and effect on its biochemical methane potential, J. Environ. Sci. Health Part A, 2016, 51, 1269–1277 CrossRef CAS PubMed.
- S.-F. Fu, F. Wang, X.-S. Shi and R.-B. Guo, Impacts of microaeration on the anaerobic digestion of corn straw and the microbial community structure, Chem. Eng. J., 2016, 287, 523–528 CrossRef CAS.
- D. Nguyen and S. K. Khanal, A little breath of fresh air into an anaerobic system: How microaeration facilitates anaerobic digestion process, Biotechnol. Adv., 2018, 36, 1971–1983 CrossRef CAS PubMed.
- Q. Chen, W. Wu, D. Qi, Y. Ding and Z. Zhao, Review on microaeration-based anaerobic digestion: State of the art, challenges, and prospectives, Sci. Total Environ., 2020, 710, 136388 CrossRef CAS PubMed.
- X. Liu, D. Huang, C. Zhu, F. Zhu, X. Zhu and D. Zhou, Production of reactive oxygen species during redox manipulation and its potential impacts on activated sludge wastewater treatment processes, Environ. Sci. Technol., 2024, 58, 23042–23052 CrossRef CAS PubMed.
- J. C. Angle, T. H. Morin, L. M. Solden, A. B. Narrowe, G. J. Smith, M. A. Borton, C. Rey-Sanchez, R. A. Daly, G. Mirfenderesgi and D. W. Hoyt, Methanogenesis in oxygenated soils is a substantial fraction of wetland methane emissions, Nat. Commun., 2017, 8, 1567 CrossRef PubMed.
- A. V. Borges, W. Champenois, N. Gypens, B. Delille and J. Harlay, Massive marine methane emissions from near-shore shallow coastal areas, Sci. Rep., 2016, 6, 27908 CrossRef CAS PubMed.
- F. Althoff, K. Benzing, P. Comba, C. McRoberts, D. R. Boyd, S. Greiner and F. Keppler, Abiotic methanogenesis from organosulphur compounds under ambient conditions, Nat. Commun., 2014, 5, 4205 CrossRef CAS PubMed.
- E. R. Frawley and F. C. Fang, The ins and outs of bacterial iron metabolism, Mol. Microbiol., 2014, 93, 609–616 CrossRef CAS PubMed.
- F. Keppler, L. Ernst, D. Polag, J. Zhang and M. Boros, ROS-driven cellular methane formation: Potential implications for health sciences, Clin. Transl. Med., 2022, 12(7), e905 CrossRef CAS PubMed.
- G. R. Buettner, In the absence of catalytic metals ascorbate does not autoxidize at pH 7: ascorbate as a test for catalytic metals, J. Biochem. Biophys. Methods, 1988, 16, 27–40 CrossRef CAS PubMed.
- G. R. Buettner and B. A. Jurkiewicz, Catalytic metals, ascorbate and free radicals: combinations to avoid, Radiat. Res., 1996, 145, 532–541 CrossRef CAS PubMed.
- J. Shen, P. T. Griffiths, S. J. Campbell, B. Utinger, M. Kalberer and S. E. Paulson, Ascorbate oxidation by iron, copper and reactive oxygen species: Review, model development, and derivation of key rate constants, Sci. Rep., 2021, 11, 7417 CrossRef CAS PubMed.
- A. Elmagirbi, H. Sulistyarti and A. Atikah, Study of ascorbic acid as iron (III) reducing agent for spectrophotometric iron speciation, J. Pure Appl. Chem. Res., 2012, 1, 11–17 CrossRef.
- A. Becerra, A. Prabhu, M. S. Rongali, S. C. S. Velpur, B. Debusschere and E. A. Walker, How a quantum computer could quantify uncertainty in microkinetic models, J. Phys. Chem. Lett., 2021, 12, 6955–6960 CrossRef CAS PubMed.
- E. A. Walker and S. A. Pallathadka, How a quantum computer could solve a microkinetic model, J. Phys. Chem. Lett., 2020, 12, 592–597 CrossRef PubMed.
- H. P. Gavin, The Levenberg-Marquardt algorithm for nonlinear least squares curve-fitting problems, Department of Civil and Environmental Engineering Duke University, 2019, vol. 3, pp. 1–23 Search PubMed.
- K. Toch, J. Thybaut, B. Vandegehuchte, C. Narasimhan, L. Domokos and G. Marin, A Single-Event MicroKinetic model for “ethylbenzene dealkylation/xylene isomerization” on Pt/H-ZSM-5 zeolite catalyst, Appl. Catal., A, 2012, 425, 130–144 CrossRef.
- F. Jin, Y. Fan, M. Yuan, F. Min, G. Wu, Y. Ding and G. F. Froment, Single-event kinetic modeling of ethene oligomerization on ZSM-5, Catal. Today, 2018, 316, 129–141 CrossRef CAS.
- F. Althoff, A. Jugold and F. Keppler, Methane formation by oxidation of ascorbic acid using iron minerals and hydrogen peroxide, Chemosphere, 2010, 80, 286–292 CrossRef CAS PubMed.
- D. He, M. Zhang, S. Liu, X. Xie and P. R. Chen, Protease-mediated protein quality control for bacterial acid resistance, Cell Chem. Biol., 2019, 26, 144–150 CrossRef CAS PubMed.
- Z. Zhang, M. Aboulwafa, M. H. Smith and M. H. Saier Jr, The ascorbate transporter of Escherichia coli, J. Bacteriol., 2003, 185, 2243–2250 CrossRef CAS PubMed.
- K. Noguchi, D. P. Riggins, K. C. Eldahan, R. D. Kitko and J. L. Slonczewski, Hydrogenase-3 contributes to anaerobic acid resistance of Escherichia coli, PLoS One, 2010, 5, e10132 CrossRef PubMed.
- C. Baker-Austin and M. Dopson, Life in acid: pH homeostasis in acidophiles, Trends Microbiol., 2007, 15, 165–171 CrossRef CAS PubMed.
- J. A. Francois and T. J. Kappock, Alanine racemase from the acidophile Acetobacter aceti, Protein Expr. Purif., 2007, 51, 39–48 CrossRef CAS PubMed.
- S. J. Campbell, B. Utinger, D. M. Lienhard, S. E. Paulson, J. Shen, P. T. Griffiths, A. C. Stell and M. Kalberer, Development of a physiologically relevant online chemical assay to quantify aerosol oxidative potential, Anal. Chem., 2019, 91, 13088–13095 CrossRef CAS PubMed.
- S. W. Schaffer and M. S. Suleiman, Mitochondria: the Dynamic Organelle, Springer Science & Business Media, 2010 Search PubMed.
- M. T. Khan and A. E. Martell, Metal ion and metal chelate catalyzed oxidation of ascorbic acid by molecular oxygen. I. Cupric and ferric ion catalyzed oxidation, J. Am. Chem. Soc., 1967, 89, 4176–4185 CrossRef CAS PubMed.
- P. M. Wood, The potential diagram for oxygen at pH 7, Biochem. J., 1988, 253, 287 CrossRef CAS PubMed.
- I. H. Segel, Biochemical Calculations: How to Solve Mathematical Problems in General Biochemistry, John Wiley & Sons, 1991 Search PubMed.
- H. Sun, G. Xie, D. He and L. Zhang, Ascorbic acid promoted magnetite Fenton degradation of alachlor: Mechanistic insights and kinetic modeling, Appl. Catal. B Environ., 2020, 267, 118383 CrossRef CAS.
- D. H. Truong, T. C. Ngo, N. T. A. Nhung, D. T. Quang, T. L. A. Nguyen, D. Khiri, S. Taamalli, F. Louis, A. El Bakali and D. Q. Dao, New insights into the competition between antioxidant activities and pro-oxidant risks of rosmarinic acid, RSC Adv., 2022, 12, 1499–1514 RSC.
- W. Stumm and J. J. Morgan, Aquatic Chemistry: Chemical Equilibria and Rates in Natural Waters, John Wiley & Sons, 2013 Search PubMed.
- C. A. Juan, J. M. Pérez de la Lastra, F. J. Plou and E. Pérez-Lebeña, The chemistry of reactive oxygen species (ROS) revisited: outlining their role in biological macromolecules (DNA, lipids and proteins) and induced pathologies, Int. J. Mol. Sci., 2021, 22, 4642 CrossRef CAS PubMed.
- W. Bors and G. Buettner, The Vitamin C Radical and its Reactions, 1997 Search PubMed.
- Y.-J. Tu, D. Njus and H. B. Schlegel, A theoretical study of ascorbic acid oxidation and HOO/O 2- radical scavenging, Org. Biomol. Chem., 2017, 15, 4417–4431 RSC.
- Z. Wang, J. Jiang, S. Pang, Y. Zhou, C. Guan, Y. Gao, J. Li, Y. Yang, W. Qiu and C. Jiang, Is sulfate radical really generated from peroxydisulfate activated by iron (II) for environmental decontamination?, Environ. Sci. Technol., 2018, 52, 11276–11284 CrossRef CAS PubMed.
- G. Tachiev, J. Roth and A. Bowers, Kinetics of hydrogen peroxide decomposition with complexed and “free” iron catalysts, Int. J. Chem. Kinet., 2000, 32, 24–35 CrossRef CAS.
- P. Refait, C. Bon, L. Simon, G. Bourrie, F. Trolard, J. Bessière and J.-M. Génin, Chemical composition and Gibbs standard free energy of formation of Fe (II)-Fe (III) hydroxysulphate green rust and Fe (II) hydroxide, Clay Miner., 1999, 34, 499–510 CrossRef CAS.
- N. Bazhin, Standard and transformed values of gibbs energy formation for some radicals and ions involved in biochemical reactions, Arch. Biochem. Biophys., 2020, 686, 108282 CrossRef CAS PubMed.
- V. A. Timoshnikov, T. V. Kobzeva, N. E. Polyakov and G. J. Kontoghiorghes, Redox interactions of vitamin C and iron: Inhibition of the pro-oxidant activity by deferiprone, Int. J. Mol. Sci., 2020, 21, 3967 CrossRef CAS PubMed.
- L. Deguillaume, M. Leriche and N. Chaumerliac, Impact of radical versus non-radical pathway in the Fenton chemistry on the iron redox cycle in clouds, Chemosphere, 2005, 60, 718–724 CrossRef CAS PubMed.
- B. HALLIWELL, Ascorbic acid and the illuminated chloroplast, in Ascorbic Acid: Chemistry, Metabolism, and Uses, ACS Publications, 1982 Search PubMed.
- M. A. SIDDIQI and E. BOTHE, Single- and double-strand break formation in DNA irradiated in aqueous solution: dependence on dose and OH radical scavenger concentration, Radiat. Res., 1987, 112, 449–463 CrossRef CAS PubMed.
- R. Herscu-Kluska, A. Masarwa, M. Saphier, H. Cohen and D. Meyerstein, Mechanism of the reaction of radicals with peroxides and dimethyl sulfoxide in aqueous solution, Chem.–Eur. J., 2008, 14, 5880–5889 CrossRef CAS PubMed.
- J. L. Wilmoth, J. K. Schaefer, D. R. Schlesinger, S. W. Roth, P. G. Hatcher, J. K. Shoemaker and X. Zhang, The role of oxygen in stimulating methane production in wetlands, Glob. Change Biol., 2021, 27, 5831–5847 CrossRef CAS PubMed.
- S. Kihara, D. A. Hartzler and S. Savikhin, Oxygen concentration inside a functioning photosynthetic cell, Biophys. J., 2014, 106, 1882–1889 CrossRef CAS PubMed.
- A. Sauer, H. Cohen and D. Meyerstein, The methyl (cyclam) nickel (III) dication in aqueous solutions: determination of the equilibrium constant of homolysis, kinetics of oxygen insertion, and methyl transfer to aquated chromium (2+), Inorg. Chem., 1988, 27, 4578–4581 CrossRef CAS.
- C. Tang and S.-Z. Qiao, How to explore ambient electrocatalytic nitrogen reduction reliably and insightfully, Chem. Soc. Rev., 2019, 48, 3166–3180 RSC.
- S. Matera, W. F. Schneider, A. Heyden and A. Savara, Progress in accurate chemical kinetic modeling, simulations, and parameter estimation for heterogeneous catalysis, ACS Catal., 2019, 9, 6624–6647 CrossRef CAS.
- S. Sharma, Quantum algorithms for simulation of quantum chemistry problems by quantum computers: an appraisal, Found. Chem., 2022, 24, 263–276 CrossRef.
- M. Reiher, N. Wiebe, K. M. Svore, D. Wecker and M. Troyer, Elucidating Reaction Mechanisms on Quantum Computers, Proceedings of the national academy of sciences, 2017, vol. 114, pp. 7555–7560 Search PubMed.
- J. D. Whitfield, J. Biamonte and A. Aspuru-Guzik, Simulation of electronic structure Hamiltonians using quantum computers, Mol. Phys., 2011, 109, 735–750 CrossRef CAS.
- A. W. Harrow, A. Hassidim and S. Lloyd, Quantum algorithm for linear systems of equations, Phys. Rev. Lett., 2009, 103, 150502 CrossRef PubMed.
- V. V. Nenonen, R. Kaegi, S. J. Hug, J. Luster, J. Göttlicher, S. Mangold, L. H. Winkel and A. Voegelin, Effects of organic ligands, phosphate and Ca on the structure and composition of Fe (III)-precipitates formed by Fe (II) oxidation at near-neutral pH, Environ. Sci.: Processes Impacts, 2025, 27, 2479–2506 RSC.
- U. Azad and H. Singh, Quantum chemistry calculations using energy derivatives on quantum computers, Chem. Phys., 2022, 558, 111506 CrossRef CAS.
- S.-C. Liu, X.-R. Zhu, D.-Y. Liu and D.-C. Fang, DFT calculations in solution systems: solvation energy, dispersion energy and entropy, Phys. Chem. Chem. Phys., 2023, 25, 913–931 RSC.
- J. Krzystek, J. England, K. Ray, A. Ozarowski, D. Smirnov, L. Que Jr and J. Telser, Determination by high-frequency and-field EPR of zero-field splitting in iron (IV) oxo complexes: Implications for intermediates in nonheme iron enzymes, Inorg. Chem., 2008, 47, 3483–3485 CrossRef CAS PubMed.
- B. F. Staley, F. L. de los Reyes III and M. A. Barlaz, Effect of spatial differences in microbial activity, pH, and substrate levels on methanogenesis initiation in refuse, Appl. Environ. Microbiol., 2011, 77, 2381–2391 CrossRef CAS PubMed.
- M. Flinck, S. Kramer and S. Pedersen, Roles of pH in control of cell proliferation, Acta Physiol., 2018, 223, e13068 CrossRef CAS PubMed.
- G. Pelaz, D. Carrillo-Pena, A. Morán and A. Escapa, Microbial electromethanogenesis for energy storage: Influence of acidic pH on process performance, J. Energy Storage, 2024, 75, 109685 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |

