 Open Access Article
Open Access ArticleMetagenomic insights into density-driven plastic stratification in open dumpsites: implications for waste management and bioremediation
Anuja
Joseph
 a and
Sudha
Goel
a and
Sudha
Goel
 *b
*b
aSchool of Environmental Science and Engineering, Indian Institute of Technology Kharagpur, Kharagpur, West Bengal 721302, India. E-mail: anujajoseph26@iitkgp.ac.in
bEnvironmental Engineering and Management Laboratory, Department of Civil Engineering, Indian Institute of Technology Kharagpur, Kharagpur, West Bengal 721302, India. E-mail: sudhagoel@civil.iitkgp.ac.in; Tel: +91-03222 283436
First published on 2nd February 2026
Abstract
Open dumpsites in developing countries constitute a critical nexus of plastic contamination and microbial ecology. However, the mechanistic interplay between polymer density-driven stratification and niche-specific microbial adaptation remains unexplored. This study presents the first integrated metagenomic investigation linking density-mediated plastic segregation to depth-stratified microbial community assembly and metabolic functionality in a tropical open dumpsite. Depth profiling revealed that low-density polyethylene dominated the layers closer to the surface, whereas polypropylene and polyethylene terephthalate were concentrated in the bottom layers. A stratification pattern was subsequently detected in adjacent soil microplastic profiles, indicating density-dependent dispersal mechanisms. Metagenomic sequencing revealed distinct microbial niches aligned with plastic layers: surface zones favoured aerobic Proteobacteria and Actinobacteria, whereas anoxic depths enriched anaerobic taxa and pathways for methane metabolism, alternative carbon fixation, and dissimilatory processes. These findings highlight plastic density as an important driver of vertical microplastic migration and indigenous microbial community structuring, informing predictive modelling of contamination risk trajectories and the design of depth-targeted bioremediation strategies using adapted native consortia. This work advances fundamental understanding of density-driven ecological structuring in anthropogenic waste systems while providing an evidence-based framework for optimizing site-specific waste stabilization and developing indigenous microbial solutions for plastic remediation in regions dependent on uncontrolled dumping.
Environmental significanceOpen dumpsites in developing countries are major, yet understudied, sources of plastic pollution that disrupt soil ecosystems and threaten environmental health. Understanding how plastic types and microbial communities distribute and interact vertically within these waste mounds is critical to addressing pollution impacts and devising effective bioremediation strategies. This study reveals significant depth-dependent stratification of plastics and associated bacterial communities accompanied by distinct metabolic functions linked to oxygen gradients. Key findings show that native microbial populations adapt to environmental conditions to potentially degrade plastics, offering promising avenues for targeted bioremediation. These insights highlight the urgent need to incorporate microbial ecology into waste management frameworks, helping mitigate plastic contamination and restore soil ecosystem functions globally, especially in regions burdened by uncontrolled dumping. |
1. Introduction
Plastic pollution is widely recognized as a critical global challenge demanding immediate action. Since 1950, over 9 billion tonnes of plastic including additives and synthetic fibres have been produced,1 yet an estimated 23 million metric tonnes (MMT) entered the aquatic systems since 2016 alone, with projections reaching 53 MMT by 2030.2 Despite worldwide efforts to reduce plastic pollution through initiatives like the “Beat Plastic Pollution” campaign by the United Nations Environment Programme (UNEP), drawing attention towards the root causes of MPW still remains a challenge.The burden of MPW, dominated by packaging waste, is particularly severe in developing regions with inadequate waste management infrastructure.3 Global assessments highlight that land-based sources contribute over 80 percent of marine plastic inputs via riverine and atmospheric pathways.4,5 In India, only 60 percent of the 3.5 MMT of annual plastic waste is collected; the remainder accumulates in over 2400 active dumpsites. According to the annual CPCB report 2021–22, West Bengal generated 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 709 tonnes per day (TPD) of municipal solid waste, of which 9475 TPD remained unprocessed, among the highest unprocessed loads nationally, distributed across 107 active and numerous unreported open dumpsites. By comparison, Maharashtra produces 23
709 tonnes per day (TPD) of municipal solid waste, of which 9475 TPD remained unprocessed, among the highest unprocessed loads nationally, distributed across 107 active and numerous unreported open dumpsites. By comparison, Maharashtra produces 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 531 TPD, Uttar Pradesh produces 14
531 TPD, Uttar Pradesh produces 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 710 TPD, and Bihar produces 8075 TPD, with similarly large unprocessed fractions (Fig. S1). These sites leach complex contaminant mixtures and emit airborne particulates that infiltrate soils and groundwater, altering soil chemistry and disrupting microbial nutrient cycling.6
710 TPD, and Bihar produces 8075 TPD, with similarly large unprocessed fractions (Fig. S1). These sites leach complex contaminant mixtures and emit airborne particulates that infiltrate soils and groundwater, altering soil chemistry and disrupting microbial nutrient cycling.6
Metagenomics, in conjunction with next-generation sequencing, has revolutionised our understanding of the plastisphere, enabling high-resolution characterisation of microbial consortia on plastics and in soils.7 Metagenomic profiling of waste contaminated soil has emerged as a pivotal tool for understanding microbial interactions with plastic waste and identifying novel bioremediation pathways.8 A recent study demonstrated the engineering of a peptidase enzyme (LCCICCG) derived from the information obtained from metagenomic analysis, which achieved a minimum of 90% depolymerization of PET within 10 hours, highlighting the potential for enzyme optimization in industrial applications.9 Landfill soils were enriched with Bacillus and Aspergillus species equipped with genes encoding esterases and hydrolases, which are instrumental in hydrolysing synthetic polymers like polyethylene (PE) and polyurethane (PU).10 Studies of marine ecosystems have shown that bacteria like Alcanivorax, Mainbocher, and Arenibacter have the ability to utilise low-density polyethylene (LDPE) and polyethylene terephthalate (PET) as a sole source of carbon.11 Such insights obtained from metagenomic profiling of soil and plastic waste are crucial for tackling MPW in areas with many open dumpsites. Such approaches help in the identification of “degradation hotspots” and guide targeted bioremediation strategies.
Open dumpsites not only fragment plastics into persistent micro- and nanoplastics but also leach a complex mixture of additives, heavy metals, and organic pollutants into the surrounding environment, contaminating both soils and groundwater. Studies in tropical regions report that leachate percolating through waste mounds elevates concentrations of toxic compounds, elements, and trace metals in adjacent soils, while runoff transports microplastics and associated contaminants into nearby streams and aquifers.12–14 These processes create coupled soil–water pollution gradients that degrade soil fertility, alter microbial nutrient cycling, and threaten drinking-water quality.
Bridging global and local scales, this study explores how heavily dumped sites drive pollution gradients and links plastic layering, leachate chemistry, and microbial shifts in a single regional model. Understanding the interactions between plastics and soil microbial communities at such dumpsites is essential for designing effective, site-specific remediation. This study leverages vertical profiling of plastic waste and metagenomic analysis to (i) map density-driven plastic stratification and adjacent soil microplastic dispersion, (ii) characterize depth-dependent shifts in bacterial diversity and metabolic function, and (iii) identify indigenous consortia and pathways for targeted bioremediation in tropical open dumpsites.
2. Materials and methodology
2.1 Study location
The town of Midnapore was chosen as the sampling location for the state of West Bengal. Constituted in the year 1865, the Midnapore municipality has a population of over 170![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 and comprises 25 wards. Waste dumping activity is carried out near the outskirts of the town at the Dharma dumpsite; it has been accumulating municipal solid waste for over 50 years. The Dharma dumpsite (22°25′18″N 87°20′24″E) has received mixed municipal solid waste since 1970s, including food scraps, paper, textiles, and plastics, and now spans 2.62 ha adjacent to residential and agricultural areas. Soil quality around the Midnapore region varies between sandy loamy soil and red soil.15 The area experiences high wind speeds of 9.6 mph on average during the month of June. Historical waste audits by the municipal authority indicate that 60% of incoming MSW is organic, 25% inert, and 15% plastics, primarily low-density polyethylene (LDPE) and polypropylene (PP).16
000 and comprises 25 wards. Waste dumping activity is carried out near the outskirts of the town at the Dharma dumpsite; it has been accumulating municipal solid waste for over 50 years. The Dharma dumpsite (22°25′18″N 87°20′24″E) has received mixed municipal solid waste since 1970s, including food scraps, paper, textiles, and plastics, and now spans 2.62 ha adjacent to residential and agricultural areas. Soil quality around the Midnapore region varies between sandy loamy soil and red soil.15 The area experiences high wind speeds of 9.6 mph on average during the month of June. Historical waste audits by the municipal authority indicate that 60% of incoming MSW is organic, 25% inert, and 15% plastics, primarily low-density polyethylene (LDPE) and polypropylene (PP).16
2.2 Sample collection and storage
Plastic waste samples were collected from three old open waste dumps about 1 m high in June 2024. Each waste mound was sectioned into 3 parts based on its height. Three sections (i.e. Layer 1, Layer 2, and Layer 3) of waste were segregated. Each section of the mound was approximately 20 cm in height (Fig. 1b). Plastic waste was collected and stored under proper conditions for segregation and metagenomic analysis. Two controls were used for this set-up, control 1 and control 2, plastic packaging samples collected from two household wastes, before disposing of them into the waste collection units.Soil samples were collected from an open dumpsite (DS) and a control site (CS) within a 1 km radius. The control site was an abandoned agricultural field, litter-free and had limited interference from human activity (Fig. 1a). The dumpsite was heavily loaded with municipal waste; mostly plastic waste was visually predominant. Soil samples were collected at 15 cm depth from the surface, and approximately 10 g soil per replicate was collected. At each sampling site, 4 replicate samples were collected within an area of 1 m × 1 m followed by mixing them into a composite sample.
2.3 Sample processing
2.4 Physical characterisation
2.5 Metagenomic analysis
Two grams of 2 soil samples each (n = 2) and 4 grams of plastic waste (n = 5) were used to extract bacterial cells. The microbial community was characterized using 16S rRNA sequencing. Using the primers 16SF (5′AGAGTTTGATGMTGGCTCAG3′) and 16SR (5′TTACCGCGGCMGCSGGCAC3′), the bacterial V3–V4 region of the 16S rRNA gene was amplified and sequenced using the Illumina MiSeq platform, with a read length of 2 × 300 bp paired end. An average read count of ∼0.1 million bp was maintained for all samples. Bioinformatics analysis and quality check of the demultiplexed data were conducted using FastQC and MultiQC, while the adapters and the low-quality reads were trimmed using TRIMGALORE, which includes merging paired-end reads, chimera removal, operational taxonomic unit (OTU) abundance calculation, and estimation correction. This was achieved using Uclust (QIIME 2 pipeline), which enables highly accurate investigations at the genus level. Taxonomic classification was done using the SILVA database. KEGG pathway analysis was carried out using PICRUSt.17Data filtering was done to remove low quality features and improve downstream statistical analysis. Data were further scaled using the Total Sum Scaling (TSS) factor to normalise information. In this study, various biodiversity assays were used to shed light on the microbial communities that colonise and adapt to the plastic surface under different environmental conditions. Alpha diversity in metagenomic studies reflects the bacterial dynamics in the different layers of the open dumpsite and health of a particular ecosystem. Metrics like Abundance-based Coverage Estimator (ACE) and Chao1 were implemented to assess species richness within the groups, while the Shannon diversity index and phylogenetic diversity index were used to assess the diversity within the 2 groups.18,19 Beta diversity metrics compare the differences between bacterial communities in different groups. The assessment explores if the samples share similar taxa or differ significantly in their bacterial communities.18 Here we used Principal Coordinates Analysis (PCoA) to understand how each sample within the groups differs along with PERMANOVA to establish set-significance. The Bray–Curtis method was adopted and the analysis was conducted at the genus level. All assays were conducted with the help of EZ-BioCloud.20
3 Results and discussion
3.1 Soil characterization
The average pH of dumpsite soil was 8.01 while that of the control soil was around 6.67. The electrical conductivity of the dumpsite soil was 824 µS m−1 and that of the control soil was 542 µS m−1.3.2 Spatial profiling of plastic content in the soil samples
The soil samples collected from the dumpsite were abundant in microplastic particles (less than 5 mm) when compared to the control site. Soil was sieved with a 365 µ sieve and plastic particles that were visible in the soil were separated. All plastic particles were characterized with the help of FTIR. A total of 50 microplastic particles were identified. Around 60% of the particles were polyethylene (PE), around 20% samples were polypropylene (PP), and 10% samples were polyethylene terephthalate (PET) whereas the soil sample from the control site had a few PE film particles only (Fig. 2a). Around 5 + 2 particles per g of soil were visualized under the fluorescence microscope (Fig. 2b); however, due to their small size range the polymer type of the particles could not be identified using an FTIR.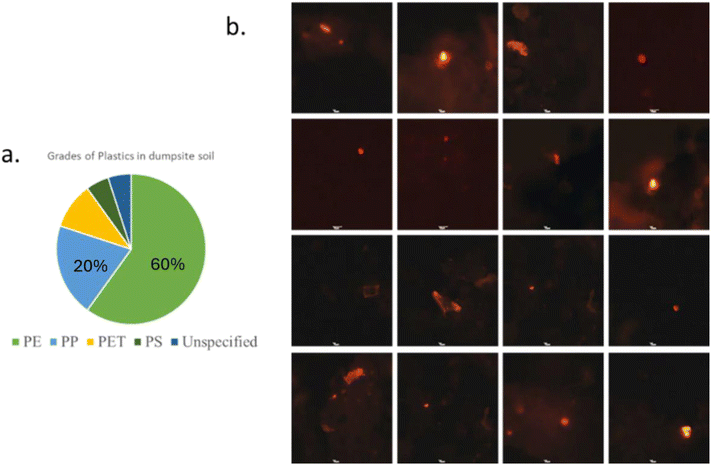 | ||
| Fig. 2 (a) Distribution of plastic grades in dumpsite soil. (b) Microplastic particles in the soil from the dumpsite viewed under a fluorescence microscope at 10× magnification. | ||
3.3 Vertical profiling of plastic waste in the dumpsite
The plastic waste from each layer (20 cm height) was segregated and characterised using an FTIR. The types of plastics identified were polyethylene (PE), polypropylene (PP), polyethylene terephthalate (PET), and polystyrene (PS) (Table 1). One-way ANOVA yielded a p-value of 0.0001 which is <0.05, (d.f. = 2); hence there is a significant difference between the types of plastic in the different layers.| Layer | PE | PP | PET | PS | PVC |
|---|---|---|---|---|---|
| 1 | 8.8 + 0.6 | 1.7 + 0.5 | 0.9 + 0.5 | 0.6 + 0.8 | 0 |
| 2 | 3.8 + 1.4 | 5 + 0.8 | 2.6 + 1.4 | 0.7 + 0.5 | 0.2 + 0.4 |
| 3 | 1.5 + 0.7 | 5.6 + 0.7 | 3.7 + 0.9 | 0.9 + 0.9 | 0.3 + 0.5 |
| Density, g cm−3 | 0.91 to 0.965 | 0.895 to 0.92 | 1.38 | 0.96 to 1.05 | 1.1 to 1.45 |
The correlation matrix reveals pronounced, statistically robust relationships among plastic types and their vertical distribution across the three layers of the dumpsite (Fig. 3b). Most notably, there is a strong negative correlation between PE and layer (−0.94), indicating that the polyethylene (PE) concentration is highest at the surface and sharply declines with depth. In contrast, PP shows a strong positive correlation with layer (+0.89), and PET is moderately positively correlated (+0.69), signifying that polypropylene and PET accumulate preferentially in deeper layers.
 | ||
| Fig. 3 (a) PCA biplot and (b) correlation matrix for the distribution of plastic types across 3 layers. (c) Scree plot for the PCA. | ||
The PCA biplot visually confirms and amplifies these findings (Fig. 3a) The primary axis (PC1), explaining 58.75% of total variance, is structured by positive loadings for the layers, PP, and PET, directly linking these variables to depth. PE is oriented strongly in the negative PC1 direction, confirming its association with upper layers and suggesting exclusive surface enrichment. PC2 (19.05% variance), driven primarily by PS, captures secondary variability but is less ecologically informative in this context.
Spatial profiling of the soil microplastic content showed a higher abundance of PE in the dumpsite soil. It may be inferred that the lightweight PE from the topmost layers of the waste mounds might get displaced and get lodged onto the soil surface, thus integrating itself into the soil.
These results highlight how the composition of plastic waste evolves over time within a dumpsite, as heaps of new waste are dumped on top of the older heaps. These insights can be utilised for waste management practices and material persistence under different environmental conditions.
3.4 Taxonomic composition: bacterial communities thriving on plastic waste
Microbial community composition reveals ecological succession driven by environmental maturation within the waste dump (Table 2). Layer 1 (surface, freshly exposed plastics) is dominated by Proteobacteria (99%), characteristic of pioneer communities with low alpha diversity and high turnover rates typical of newly colonized substrates. In contrast, layers 2 and 3 exhibit significantly reduced Proteobacteria abundance (63% and 75.6% respectively) and substantially increased microbial diversity, indicating community stabilization during ecological succession. This depth-dependent shift reflects reduced oxygen availability and substrate depletion at depth, creating selective pressures for metabolically specialized communities.| Phylum | Layer 1 | Layer 2 | Layer 3 |
|---|---|---|---|
| Proteobacteria | ∼99 | 63 | 75.6 |
| Firmicutes | <0.3 | 17 | 12 |
| Actinobacteria | <0.07 | 10 | 6 |
| Bacteroidetes | <0.06 | 5 | 3 |
| Planctomycetes | — | 2.2 | 1.3 |
| Chloroflexi | — | 0.4 | 0.3 |
| Acidobacteria | — | 0.2 | 0.1 |
The emergence of Firmicutes (17–12% in layers 2–3) and Actinobacteria (10–6% in layers 2–3) in deeper layers signals a functional transition toward anaerobic metabolic pathways. Notably, classes such as Clostridia and Bacilli—obligate and facultative anaerobes capable of fermentative metabolism and polymer hydrolysis, become dominant only in anoxic deeper layers. The exclusive presence of Planctomycetes, Chloroflexi, andAcidobacteria in layers 2–3 indicates niche differentiation and specialization absent in pioneer surface communities. Control samples, dominated by Gammaproteobacteria (∼99%), lack these depth-specialized taxa, confirming that prolonged plastic-soil contact selects for functionally diverse, depth-adapted microbial consortia. This vertical stratification demonstrates ecological succession, i.e., initial colonization by generalist proteobacteria to niche differentiation with depth-dependent functional specialization.
By class level analysis 42 different classes were recognised; the control site was abundant with Acidobacteria (29%), Alphaproteobacteria (29%), Actinobacteria (13%), and Planctomycetia (10%) (Fig. S5). The control site had a greater relative abundance of Acidobacteria (29%) when compared to the dumpsite soil (0.5%) (Fig. S6). Both soils had almost equal relative abundance of Alphaproteobacteria, while the dumpsite soil was rich in Actinobacteria (27%) when compared to the control site (13%). Besides, the dumpsite had a moderately equal distribution of Planctomycetia (4%), Betaproteobacteria (5%) and Gammaproteobacteria (5%).
The top enriched genus of both the control site and dumpsite is presented in Fig. S7. These results indicate that the bacterial communities can help facilitate further degradation of plastic particles to tinier microplastic particles. Highest relative abundance was observed for the phylum Proteobacteria, class Alphaproteobacteria, and order Rhizobiales (Fig. S6).
3.5. Comparative metagenomic insights into bacterial communities on plastic waste
The ACE graph showed significantly higher species richness for samples of plastics collected from the waste dump (median observation = 3371), while the richness was low (median observation = 174) in the samples considered as control (p = 0.019). Similarly, the Chao1 graph also showed elevated species richness within the plastics collected from the dump site (median observation = 3462, p = 0.019). Higher values for the bacterial species collected from the plastic waste suggest the presence of a richer microbial community when compared to the control plastics.
The Shannon index and the phylogenetic index were used to explore the diversity within the samples. The Shannon index for plastics from waste had a higher range of value (median = 3.33) while the values for the control samples had a lower range (median = 0.34) with a p-value of 0.011. Similarly, the phylogenetic diversity index also yielded similar results, indicating higher species richness in the plastic waste (median observation = 2448) with a p-value of 0.033. A p-value <0.05 indicates statistically significant data in all cases.
Beta diversity estimation creates a dissimilarity matrix that contains, for each sample (arranged in the rows), dissimilarity to any other sample (arranged in the columns). PCoA is a non-linear dimension reduction method, which is identical to Principal Component Analysis (PCA) (except for scaling). Principal coordinates 1 and 2 are, hence, the percentage of total variations observed. The first principal coordinate suggests 86.22% variance, while the 2nd principal coordinate accounts for 11.39%, capturing secondary differences in the microbial composition (Fig. 5).
 | ||
| Fig. 5 PCoA (Bray–Curtis difference) for the bacterial communities thriving on the plastic waste vs. control plastics. | ||
The bacterial communities from the plastic waste collected from layers 1, 2, and 3 cluster separately from the control group (control 1 and 2) indicating distinct bacterial genera between the groups. Within the plastic bacteria group, the bacterial genus from layer 1 has a stark difference from that of layers 2 and 3. Layers 2 and 3 are closely related while layer 1 was not. This dissimilarity indicates a distinct change in the bacterial activity with the change in oxygen and light. The tight clustering of control samples indicates limited variability in the microbiota due to less exposure to the outside environment.
The PERMANOVA analysis statistically confirms differences in the composition of the microbiome with 999 permutations. The results with a pseudo-F value of 6.083 and p-value of 0.011 indicated that the differences between the two sets (plastic waste and control) were statistically significant. The pseudo-F value suggested that the inter-group variability is much greater than the intra-group variability.
Therefore, the results demonstrate that dumpsite plastics harbour distinct bacterial communities compared to control plastics. It can thus be inferred that these differences are likely driven by the environmental exposure at dumpsites, creating unique ecological niches for bacterial colonization.
Microbial metabolic capacity demonstrates clear depth-dependent specialization (Fig. 7). Surface layers (layer 1) are characterized by oxidative phosphorylation and carbon fixation, supporting aerobic heterotrophs. Deeper layers (layer 3) show dominant methane metabolism, reflecting anaerobic methanogenesis. Carbohydrate metabolism pathways remain consistently abundant across all depths, indicating continuous organic matter degradation. Amino acid metabolism progressively increases with depth, suggesting enhanced protein utilization under anaerobic conditions. Notably, xenobiotic degradation pathways are present, indicating microbial capacity for plastic-derived compound metabolism. These findings demonstrate oxygen-driven metabolic niches, with implications for bioremediation strategies (Fig. 7).
Methane metabolism activity is significantly higher in layer 3 and this activity is directly related to anaerobic conditions that enable the growth of methanogenic bacteria. In the absence of oxygen, the available CO2 and acetate can be utilized as terminal electron acceptors. Many methanotrophs belong to the phylum Proteobacteria, specifically with the classes Alphaproteobacteria and Gammaproteobacteria.22 Most methanotrophs in an area with low oxygen and high methane availability belong to Gammaproteobacteria, while in areas with low methane and high oxygen availability, methanotrophs belonging to the Alphaproteobacter class are common.23 In the taxonomic context, an increase in Methylobacterium sp. and Methylocystis sp. was significantly evident in layers 2 and 3. Such bacteria are capable of growing on single-carbon compounds that can be derived from different types of polymers.24,25
Oxidative phosphorylation was higher in layers 2 and 3 when compared to layer 1; this suggests crucial adaptation of microbial communities to depth-stratified conditions. The increased oxidative phosphorylation activity in the lower layers (layers 2 and 3) implicates the presence of microbial communities capable of carrying out anaerobic respiration using alternative electron acceptors like nitrate, sulphate, or carbon dioxide.26 The presence of sulphate-reducing bacteria like Desulfovibrio sp. and Desulfovibrio vulgaris in layers 2 and 3 only indicates the utilisation of sulphate as a terminal electron acceptor through modified oxidative phosphorylation pathways under semi-anaerobic and anaerobic conditions.27,28 Layer 1 has been exposed to atmospheric conditions, supporting predominantly aerobic microorganisms; however, the higher oxidative phosphorylation in the bottom layers reflects a vertical stratification of microbial activity. The stratification is evident where anaerobic organisms like Clostridium sp. display significantly higher abundance in layers 2 and 3 when compared to layer 1.29–31 Similarly, Faecalibacterium prausnitzii shows higher abundance in layer 3, followed by layer 2, while it is absent in layer 1, suggesting adaptation to an anaerobic environment.32 Carbon fixation pathways convert inorganic carbon into organic compounds. The highest expression of this pathway was observed in layer 3, indicating the probable dominance of anaerobic autotrophic bacteria. Layer 2 supports moderate activity with limited oxygen availability thus supporting facultative autotrophs. In contrast, layer 1 with higher oxygen abundance facilitates heterotrophs over autotrophs; consequently, carbon fixation pathways are less active compared to the deeper layers.
Pathways like amino sugar and nucleotide sugar metabolism, glycine–serine–threonine metabolism, alanine–aspartate–glutamate metabolism, and nitrogen metabolism had higher expression in layer 2 than in layer 3. This represents a complex biochemical adaptation to a transitional oxygen environment. Nucleotide sugars act as critical donors for sugar residues in glycosylation reactions and act as substrates for glycosyltransferases.33,34 The activated sugars then link glucose metabolism with cellular functions through protein glycosylation.35 Layer 2 represents a transitional zone between aerobic and anaerobic conditions, supporting optimal conditions for facultative anaerobes. Adaptation to such a metabolic strategy is advantageous for rapidly proliferating microbes that require higher amounts of anabolic precursors. The middle layer likely has an abundance of partially degraded carbohydrates from the upper layer, providing sufficient substrates for nucleotide sugar synthesis. However, in layer 3, such carbohydrates may have been depleted. Glycine–serine–threonine metabolism highlights a crucial point in amino acid metabolism. Glycine is derived from serine by the enzyme serine hydroxymethyltransferase (SHMT), while serine is derived from an intermediate in the glycolysis pathway (3-phospho-D-glycerate).36 Some enzymes in this pathway require oxygen to function optimally. The glycine cleavage system releases ammonia and requires NAD+ (nicotinamide adenine dinucleotide) regeneration – such processes function efficiently under semi-aerobic conditions that are available in layer 2.36 Alanine (ALT) and aspartate (AST) aminotransferase play important roles in nitrogen metabolism through transamination reactions.37 ALT catalyses reversible transamination between alanine and 2-oxoglutarate for pyruvate and glutamate, while AST performs similar reactions with aspartate.37 Hence, semi-aerobic conditions in layer 2 provide an optimal environment for transamination reactions that shuttle nitrogen between amino acids and keto acids which is absent in layer 3. The availability of some oxygen in layer 2 facilitates the participation of ALT in gluconeogenesis. Nitrogen metabolism involves the processing of ammonia which is toxic to cells and requires regular detoxification.38 Layer 2 most likely experiences significant ammonia release from protein degradation and amino acid metabolism as discussed earlier. Under semi-aerobic conditions in landfill sites, microbial activity enhances decomposition, releasing nitrogen compounds.39,40 Also, the limited oxygen in this layer may create decentralized pockets in the environment where both nitrification and denitrification can occur simultaneously, leading to more active nitrogen cycling.
The metabolic pathways across the three layers of waste dump reflect distinct microbial adaptations to environmental gradients like oxygen availability, nutrient accessibility, and depth. Layer 3 or the bottom layer consistently shows the highest expression of key pathways, including methane metabolism, carbon fixation, oxidative phosphorylation, and glycolysis/gluconeogenesis. This is representative of specialised microbial communities capable of utilising alternative electron acceptors under strict anaerobic conditions. Layer 2 or the middle layer shows moderate expression of most pathways, reflecting the transitional state between aerobic and anaerobic zones. Facultative anaerobes dominate this layer, contributing to enhanced nucleotide metabolism (purine and pyrimidine), amino acid metabolism (alanine–aspartate–glutamate and serine–glycine–threonine metabolic pathways) and nitrogen metabolism. Layer 1, the topmost layer displays lowest pathway expression overall due to its predominantly aerobic environment. Oxygen availability in this layer favours heterotrophic activity over autotrophic and anaerobic bacterial communities. These findings affirm that stratification of metabolic pathways across layers emphasises the dynamic interplay between microbial communities and environmental conditions within the dumpsite. The dominance of plastics also has an effect on the availability of carbon as a source of energy. The results therefore, underscore the significance of depth-specific microbial adaptation in driving key biogeochemical processes such as methane cycling, carbon fixation and energy metabolism (Fig. 8).
4 Conclusion
This study reveals the vertical and spatial patterns of plastic debris and microbial communities at a major open dumpsite in West Bengal, India. The key findings demonstrate (1) density-driven stratification of plastics, with lighter polymers accumulating at the surface and denser fractions at depth, (2) depth-specialized microbial communities showing functional shifts that may inform bioremediation and gas capture strategies, and (3) off-site PE microplastic dispersal significantly alters soil microbial ecology within a 200 m radius. Limitations of this work lie in the inability to directly measure bulk density, local wind, or hydrodynamic data; hence, results from the literature were incorporated. These novel insights underscore the inadequacy of current waste management practices and highlight the urgent need for revised segregation, site engineering, and monitoring policies. Integrating depth-resolved microbial assessment into waste impact assessments can directly inform remediation strategies, supporting India's solid waste management rules. Region-specific solutions based on these findings offer a practical roadmap toward achieving the United Nations SDGs for sustainable cities and climate action.Author contributions
Anuja Joseph: conceptualisation, methodology, formal analysis, writing – original draft, writing – review and editing, visualisation Sudha Goel: resources, formal analysis, writing – review & editing, supervision.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationship that could have appeared to influence the work reported in this paper.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5va00288e.Acknowledgements
The authors would like to acknowledge the sequencing service provided by BioKart, India. During the preparation of this work the authors used ChatGPT, OpenAI, in order to improve the grammar and language. After using this tool/service, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication.References
- R. Geyer, Production, use, and fate of synthetic polymers, in Plastic Waste and Recycling, Elsevier, 2020, pp. 13–32, https://linkinghub.elsevier.com/retrieve/pii/B9780128178805000025 Search PubMed.
- S. B. Borrelle, J. Ringma, K. L. Law, C. C. Monnahan, L. Lebreton and A. McGivern, et al., Predicted growth in plastic waste exceeds efforts to mitigate plastic pollution, Science, 2020, 369(6510), 1515–1518 Search PubMed . https://www.science.org/doi/10.1126/science.aba3656.
- C. A. Velis and E. Cook, Mismanagement of Plastic Waste through Open Burning with Emphasis on the Global South: A Systematic Review of Risks to Occupational and Public Health, Environ. Sci. Technol., 2021, 55(11), 7186–7207, DOI:10.1021/acs.est.0c08536.
- L. Lebreton, B. Slat, F. Ferrari, B. Sainte-Rose, J. Aitken and R. Marthouse, et al., Evidence that the Great Pacific Garbage Patch is rapidly accumulating plastic, Sci. Rep., 2018, 8(1), 4666 Search PubMed . https://www.nature.com/articles/s41598-018-22939-w.
- W. C. Li, H. F. Tse and L. Fok, Plastic waste in the marine environment: a review of sources, occurrence and effects, Sci. Total Environ., 2016, 566–567, 333–349 CrossRef CAS PubMed . https://www.sciencedirect.com/science/article/pii/S0048969716310154.
- A. Lodh, M. Shafi and S. Goel, Microplastics in municipal solid waste landfill leachate and their removal in treatment units: a perspective of controlled and uncontrolled landfills, Environ. Pollut., 2025, 369, 125853 Search PubMed . https://linkinghub.elsevier.com/retrieve/pii/S026974912500226X.
- N. Tiwari, M. Bansal, D. Santhiya and J. G. Sharma, Insights into microbial diversity on plastisphere by multi-omics, Arch. Microbiol., 2022, 204(4), 216, DOI:10.1007/s00203-022-02806-z.
- A. Joseph, B. Nithin and S. Goel, Biodegradation of Plastics, in Advances in Solid and Hazardous Waste Management, ed. S. Goel, Springer International Publishing, Cham, 2024, pp. 221–240, DOI:10.1007/978-3-031-49144-3_13.
- V. Tournier, C. M. Topham, A. Gilles, B. David, C. Folgoas and E. Moya-Leclair, et al., An engineered PET depolymerase to break down and recycle plastic bottles, Nature, 2020, 580(7802), 216–219 Search PubMed . Available from: https://www.nature.com/articles/s41586-020-2149-4.
- M. Merchant, T. Kahali, D. K. Kumawat, S. S. Mande and P. Sar, Understanding the Structure and Function of Landfill Microbiome Through Genomics, in Microbial Diversity in the Genomic Era, Elsevier, 2024, pp. 669–695, https://linkinghub.elsevier.com/retrieve/pii/B9780443133206000238 Search PubMed.
- A. Delacuvellerie, V. Cyriaque, S. Gobert, S. Benali and R. Wattiez, The plastisphere in marine ecosystem hosts potential specific microbial degraders including Alcanivorax borkumensis as a key player for the low-density polyethylene degradation, J. Hazard. Mater., 2019, 380, 120899 Search PubMed.
- T. O. Ale, T. H. Ogunribido, O. Ademila and A. S. Akingboye, Soil–water contamination assessment due to dumpsite-impacted leachates in a metamorphic environment, J. Afr. Earth Sci., 2024, 216, 105312 CrossRef CAS . https://www.sciencedirect.com/science/article/pii/S1464343X24001456.
- T. O. Ale, T. H. Ogunribido, O. Ademila and A. S. Akingboye, Soil pollution status due to potentially toxic elements in active open dumpsites: insights from different Nigerian geological environments, Environ. Earth Sci., 2024, 83(18), 535, DOI:10.1007/s12665-024-11810-z.
- O. O. Tovide, J. A. O. Oyekunle, O. T. Ore, B. A. Oyebode, E. O. Moseri and A. S. Adekunle, et al., Speciation studies of potentially toxic elements within the vicinity of major dumpsites in Ile-Ife, Osun State, Nigeria, J. Trace Elem. Min., 2025, 11, 100210 CrossRef . https://www.sciencedirect.com/science/article/pii/S2773050625000011.
- West Bengal Government, Livelihood Survey Document, Paschim Medinipur District, 2025, https://dstbt.bangla.gov.in/news_notification/LivelihoodSurveyDocument_PaschimMedinipurDistrict.pdf#page=2.14.
- K. N. Kumar and S. Goel, Characterization of Municipal Solid Waste (MSW) and a proposed management plan for Kharagpur, West Bengal, India, Resour., Conserv. Recycl., 2009, 53(3), 166–174 Search PubMed . https://www.sciencedirect.com/science/article/pii/S0921344908002012.
- M. G. I. Langille, J. Zaneveld, J. G. Caporaso, D. McDonald, D. Knights and J. A. Reyes, et al., Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences, Nat. Biotechnol., 2013, 31(9), 814–821 Search PubMed . https://www.nature.com/articles/nbt.2676.
- J. G. Kers and E. Saccenti, The Power of Microbiome Studies: Some Considerations on Which Alpha and Beta Metrics to Use and How to Report Results, Front. Microbiol., 2022, 12, 796025 Search PubMed . https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2021.796025/full.
- D. R. Finn, A metagenomic alpha-diversity index for microbial functional biodiversity, FEMS Microbiol. Ecol., 2024, 100(3), fiae019, DOI:10.1093/femsec/fiae019.
- M. Chalita, Y. O. Kim, S. Park, H. S. Oh, J. H. Cho and J. Moon, et al., EzBioCloud: a genome-driven database and platform for microbiome identification and discovery, Int. J. Syst. Evol. Microbiol., 2024, 74(6), 006421 Search PubMed.
- S. Dey, A. Kumar Rout, K. Ghosh, S. Dixit, V. Kumar and B. Kumar Das, et al., Unveiling the plastic degrading potential of the beneficial microbiome through plastisphere community diversity and predictive functionality analysis in waste disposal sites in the adjoining areas of Kolkata, West Bengal, India, Curr. Res. Biotechnol., 2024, 8, 100237 Search PubMed . https://www.sciencedirect.com/science/article/pii/S2590262824000637.
- M. G. Kalyuzhnaya, O. A. Gomez and J. C. Murrell, The Methane-Oxidizing Bacteria (Methanotrophs), in Taxonomy, Genomics and Ecophysiology of Hydrocarbon-Degrading Microbes, ed. McGenity T. J., Springer International Publishing, Cham, 2019, pp. 1–34, http://link.springer.com/10.1007/978-3-319-60053-6_10-1 Search PubMed.
- A. Hakobyan and W. Liesack, Unexpected metabolic versatility among type II methanotrophs in the Alphaproteobacteria, Biol. Chem., 2020, 401(12), 1469–1477, DOI:10.1515/hsz-2020-0200/html.
- M. N. Allemann, M. Tessman, J. Reindel, G. B. Scofield, P. Evans and R. S. Pomeroy, et al., Rapid biodegradation of microplastics generated from bio-based thermoplastic polyurethane, Sci. Rep., 2024, 14(1), 6036 CrossRef CAS PubMed , https://www.nature.com/articles/s41598-024-56492-6.
- F. M. de Souza and R. K. Gupta, Bacteria for Bioplastics: Progress, Applications, and Challenges, ACS Omega, 2024, 9(8), 8666–8686, DOI:10.1021/acsomega.3c07372.
- H. Lemieux and P. U. Blier, Exploring Thermal Sensitivities and Adaptations of Oxidative Phosphorylation Pathways, Metabolites, 2022, 12(4), 360 Search PubMed . https://www.mdpi.com/2218-1989/12/4/360.
- K. A. Demin, E. V. Prazdnova, T. M. Minkina and A. V. Gorovtsov, Sulfate-reducing bacteria unearthed: ecological functions of the diverse prokaryotic group in terrestrial environments, Appl. Environ. Microbiol., 2024, 90(4), e01390–23, DOI:10.1128/aem.01390-23.
- P. M. Matias, I. A. C. Pereira, C. M. Soares and M. A. Carrondo, Sulphate respiration from hydrogen in Desulfovibrio bacteria: a structural biology overview, Prog. Biophys. Mol. Biol., 2005, 89(3), 292–329 Search PubMed . https://www.sciencedirect.com/science/article/pii/S0079610704001683.
- J. Subirats, H. Sharpe and E. Topp, Fate of Clostridia and other spore-forming Firmicute bacteria during feedstock anaerobic digestion and aerobic composting, J. Environ. Manage., 2022, 309, 114643 CrossRef CAS PubMed . https://www.sciencedirect.com/science/article/pii/S030147972200216X.
- M. Cyprowski, A. Ławniczek-Wałczyk, A. Stobnicka-Kupiec and R. L. Górny, Occupational exposure to anaerobic bacteria in a waste sorting plant, J. Air Waste Manage. Assoc., 2021, 71(10), 1292–1302 CrossRef CAS PubMed . https://www.tandfonline.com/doi/full/10.1080/10962247.2021.1934185.
- Y. S. Nimonkar, D. R. Ranade, A. Poojary, M. S. Dake and O. Prakash, Comparison of Taxonomic Resolutions of Various Typing Methods for Clostridium and Paraclostridium Species Isolated from Landfill Leachate, Indian J. Microbiol., 2024, 64(4), 1577–1586 Search PubMed . https://link.springer.com/10.1007/s12088-023-01179-1.
- K. Laczi, A. Bodor, T. Kovács, B. Magyar, K. Perei and G. Rákhely, Methanogenesis coupled hydrocarbon biodegradation enhanced by ferric and sulphate ions, Appl. Microbiol. Biotechnol., 2024, 108(1), 449, DOI:10.1007/s00253-024-13278-0.
- J. Rini, J. Esko and A. Varki, Glycosyltransferases and Glycan-processing Enzymes, in Essentials of Glycobiology, ed. Varki A., Cummings R. D., Esko J. D., Freeze H. H., Stanley P. and Bertozzi C. R., Cold Spring Harbor Laboratory Press, Cold Spring Harbor (NY), 2nd edn, 2009, http://www.ncbi.nlm.nih.gov/books/NBK1921/ Search PubMed.
- S. Mikkola, Nucleotide Sugars in Chemistry and Biology, Molecules, 2020, 25(23), 5755 Search PubMed . https://www.mdpi.com/1420-3049/25/23/5755.
- F. Conte, M. J. Noga, M. van Scherpenzeel, R. Veizaj, R. Scharn and J. E. Sam, et al., Isotopic Tracing of Nucleotide Sugar Metabolism in Human Pluripotent Stem Cells, Cells, 2023, 12(13), 1765 CrossRef CAS PubMed . https://www.ncbi.nlm.nih.gov/pmc/articles/PMC10340731/.
- S. L. Geeraerts, E. Heylen, K. De Keersmaecker and K. R. Kampen, The ins and outs of serine and glycine metabolism in cancer, Nat. Metab., 2021, 3(2), 131–141 CrossRef CAS PubMed . https://www.nature.com/articles/s42255-020-00329-9.
- R. Bornstein, M. T. Mulholland, M. Sedensky, P. Morgan and S. C. Johnson, Glutamine metabolism in diseases associated with mitochondrial dysfunction, Mol. Cell. Neurosci., 2023, 126, 103887 CrossRef CAS PubMed . https://www.sciencedirect.com/science/article/pii/S1044743123000817.
- H. G. Hoang, B. T. P. Thuy, C. Lin, D. V. N. Vo, H. T. Tran and M. B. Bahari, et al., The nitrogen cycle and mitigation strategies for nitrogen loss during organic waste composting: A review, Chemosphere, 2022, 300, 134514 CrossRef CAS PubMed . https://www.sciencedirect.com/science/article/pii/S0045653522010074.
- D. Xiao, Y. Chen, W. Xu and L. Zhan, An Aerobic Degradation Model for Landfilled Municipal Solid Waste, Appl. Sci., 2021, 11(16), 7557 CrossRef CAS . https://www.mdpi.com/2076-3417/11/16/7557.
- S. Zhao, Q. Zheng, H. Wang and X. Fan, Nitrogen in landfills: Sources, environmental impacts and novel treatment approaches, Sci. Total Environ., 2024, 924, 171725 CrossRef CAS PubMed . https://www.sciencedirect.com/science/article/pii/S0048969724018680.
| This journal is © The Royal Society of Chemistry 2026 |





