Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Rational design of microfluidic templated HA-LPEI nanogels for the targeted delivery of doxorubicin
Emanuele
Limiti†
ab,
Eleonora
D’Alessandro†
 a,
Alessio
Bucciarelli
a,
Alessio
Bucciarelli
 c,
Sofia
Raniolo
a,
Pamela
Mozetic
bde,
Elisa
De Luca
be,
Enrico Domenico
Lemma
c,
Sofia
Raniolo
a,
Pamela
Mozetic
bde,
Elisa
De Luca
be,
Enrico Domenico
Lemma
 ab,
Francesco
Basoli
a,
Giuseppe
Gigli
bef,
Fabio
Pizzetti
g,
Emanuele
Mauri
ab,
Francesco
Basoli
a,
Giuseppe
Gigli
bef,
Fabio
Pizzetti
g,
Emanuele
Mauri
 g,
Filippo
Rossi
g,
Filippo
Rossi
 g,
Marcella
Trombetta
a,
Sara Maria
Giannitelli
*a and
Alberto
Rainer
g,
Marcella
Trombetta
a,
Sara Maria
Giannitelli
*a and
Alberto
Rainer
 *ah
*ah
aDepartment of Engineering, Università Campus Bio-Medico di Roma, via Álvaro del Portillo 21, 00128 Rome, Italy. E-mail: s.giannitelli@unicampus.it; a.rainer@unicampus.it
bInstitute of Nanotechnology (NANOTEC), National Research Council, via Monteroni, 73100 Lecce, Italy
cBIOtech Research Center and European Institute of Excellence on Tissue Engineering and Regenerative Medicine, Department of Industrial Engineering, University of Trento, Via Delle Regole 101, Trento 38123, Italy
dDepartment of Life Science, Health, and Health Professions, Link Campus University, Rome, Italy
eTecnomed Puglia - Tecnopolo per la medicina di precisione (Biotech Lecce Hub) c/o Campus Ecotekne, via Monteroni, Lecce, 73100, Italy
fDepartment of Experimental Medicine, Università del Salento, via Monteroni, 73100 Lecce, Italy
gDepartment of Chemistry, Materials and Chemical Engineering “G. Natta”, Politecnico di Milano, via L. Mancinelli 7, 20131 Milan, Italy
hFondazione Policlinico Universitario Campus Bio-Medico di Roma, via Álvaro del Portillo 200, 00128 Rome, Italy
First published on 8th April 2026
Abstract
Hyaluronan-based nanomaterials represent an intriguing class of materials for drug delivery applications. In this scenario, continuous flow syntheses hold promise for superior control over process parameters, leading to more homogeneous and reproducible nanomaterials. However, the high sensitivity to parameter variation necessitates extensive and time-consuming experiments to ascertain the optimal process conditions. This study explores the implementation of a Response Surface Methodology (RSM) supporting the rational design of hyaluronic acid (HA) - linear polyethyleneimine (LPEI) nanogels (NGs) via droplet-based microfluidics for the intracellular delivery of doxorubicin. The RSM approach facilitated the development of an empirical model predicting NG physico-chemical properties and biological performance as a function of process parameters (namely, the flow rate ratio (FRR) between the internal and external phases and the LPEI/HA monomer molar ratio (MR) of the polymeric precursors) with high accuracy levels. Results indicate that FRR predominantly influences NG size, which ranges between 100 and 400 nm, and polydispersity (0.01–0.1), while MR affects composition, cytocompatibility, cellular uptake and NP-mediated drug effect. In particular, increased HA content resulted in improved cytocompatibility and enhanced cellular uptake. Overall, this work underscores the key role of RSM in advancing the synthesis of hyaluronan-based nanomaterials for drug delivery applications.
1. Introduction
In the last decades, the demand for new methods to increase the efficacy of therapeutics has led to the development of several drug delivery systems (DDSs).1–3 Starting from the visionary work of Vinogradov et al.,4 Nanogels (NGs) have arisen as one of the most promising nanotechnological tools for drug delivery purposes, mainly due to their distinctive features.5 Nanogels are defined as nano-sized three-dimensional networks of chemically or physically cross-linked polymers with a marked swelling behavior. Their capacity to retain substantial amounts of water enhances their biocompatibility; at the same time, their polymeric mesh can be engineered with hydrophilic and hydrophobic domains, enabling the delivery of both hydrophilic and hydrophobic drugs with high loading capacity and controlled release.6–11 Moreover, NGs can be formulated to show stimuli-responsiveness to physico-chemical triggers (such as pH, temperature, or light), and they can be further modified for the recognition of target cells or tissues.7,8,12–15 Polysaccharides are emerging as attractive candidates for NGs synthesis, primarily due to their biocompatibility, biodegradability, availability in nature, and simple chemical derivatization.16,17 Most importantly, many polysaccharides are inherently recognized by specific cell surface molecules, enabling receptor-mediated endocytosis.18–20 As an example, hyaluronic acid (HA) is commonly employed in several cancer therapies to actively target the CD44 receptor, which is highly overexpressed by many cancer cells.20–22Recently, the application of microfluidics has impacted the field of nanomedicine for the in-flow synthesis of DDSs.23,24 The fine manipulation of nano/picoliter fluid flows within microscale channels enables fast reaction kinetics, highly controllable mixing, and thermal homogeneity, giving unrivaled control over reaction conditions when manufacturing nanomaterials.25,26 Indeed, these unique properties have made it possible to address several issues of traditional bulk syntheses, such as poor batch-to-batch reproducibility, high polydispersity, and inhomogeneous material properties.27 Microfluidic approaches have also been optimized for high-throughput production, enabling scalable synthesis of nanomaterials.28 Indeed, literature reports the parallelization of thousands of independent fluidic units working simultaneously, eventually reaching throughputs suitable for commercial and clinical applications.29–32
To date, single-phase flow systems are the most widely investigated microfluidic devices for the synthesis of nanocarriers via in-flow nanoprecipitation or self-assembly processes.24,33–38 Specifically, during nanoprecipitation, precursors are first dissolved in an appropriate solvent and then mixed microfluidically with an antisolvent phase, leading to nanoparticle formation through nucleation, growth and precipitation.39
Despite their significant potential, single-phase flow systems are prone to channel clogging, which can lead to process instability or even failure in nanoparticle synthesis.39 Consequently, droplet-based microfluidics has emerged as a promising strategy to overcome these limitations. Here, the confinement of the reaction within discrete droplets reduces the interaction between reagents and channel walls, limiting the formation of clogging aggregates within the device and leading to a more robust and stable process. However, while this technique has been widely investigated for the synthesis of inorganic nanoparticles,40–43 its application for carbohydrate-based polymeric nanomaterials is still an emerging area of research.
Given the tight control offered by microfluidics over a variety of compositional and fluid-dynamic parameters influencing synthesis outcomes, conducting multi-parametric optimization is essential to identify the most suitable process conditions. In this context, the design of experiment (DoE) approach coupled with response surface methodology (RSM) represents a powerful framework to rationally explore the influence of independent input variables (predictors) on dependent output variables (responses).44 Furthermore, RSM can capture higher-order effects, which are usually overlooked in simple linear approximations.45 Literature reports some early optimization approaches applied to the continuous flow synthesis of nanomaterials based on carbohydrate polymers. For instance, Whiteley et al.46, investigated the synthesis and drug loading of chitosan-based NGs produced in a single-phase flow system through DoE and RSM. However, their study focused on process optimization, with bioperformance aspects remaining unexplored.
In this work, we applied DoE/RSM to the droplet-based microfluidic synthesis of NGs in the hyaluronic acid – linear polyethyleneimine (HA-LPEI) system for the targeted delivery of a chemotherapeutic agent (doxorubicin, DOX).
Hyaluronic acid is a natural polysaccharide composed of repeating disaccharide units, consisting of D-glucuronic acid and N-acetylglucosamine linked with alternating β-(1–4) and β-(1–3) glycosidic bonds.47,48 It is a major component of the extracellular matrix of several biological tissues, known for its excellent viscoelasticity, high moisture retention capacity, and hygroscopic properties. Moreover, HA is extensively used in the biomedical and cosmetic fields due to its excellent biocompatibility and biodegradability.49–51 Its use in bioconjugation strategies for nanogel and microgel synthesis introduces beneficial properties such as cell-targeting capability, reduced immunogenicity, and enhanced support for prolonged 3D cell culture.52–54
Polyethyleneimine (PEI) is a synthetic polycation commonly used for drug and gene delivery.55 PEI offers several key advantages, including: (i) the ability to easily condensate nucleic acid into polyplex nanoparticles;55,56 (ii) the proton-sponge effect promoting endosomal escape after cellular internalization;57,58 and (iii) its amenability for chemical modification, owing to the abundance of amine groups in the polymeric chain.55,56 On the other hand, PEI is limited by its poor biocompatibility, which poses a barrier to its clinical application.59–61
DOX is an anthracycline antibiotic widely used in various cancer types.62 Its mechanism of action involves intercalation between DNA base pairs and stabilization of DNA–topoisomerase II cleavage complexes, thereby inhibiting DNA replication. Additionally, DOX undergoes redox cycling, generating reactive oxygen species (ROS) that induce further DNA damage and trigger apoptosis.63 These combined effects on nucleic acids and oxidative stress underlie DOX potent cytotoxic activity against tumor cells.
To support the development of the HA/LPEI-based nanoDDS, we hypothesized that a DoE/RSM approach could serve as a systematic framework for optimization. Accordingly, we set a two-factor three-level design to investigate the influence of two process parameters, namely FRR and LPEI/HA ratio (expressed as the molar ratio between PEI monomeric units and HA disaccharide units) (MR) on several physicochemical properties (size, PDI, composition, and drug release profile) and biological characteristics (cytocompatibility, cellular uptake, and performance as a DDS). Based on the collected data, we developed a RSM empirical model that enables the adjustment of the two process parameters to achieve targeted physicochemical and biological responses within the space of the investigated input variables. The presented DoE/RSM is functional to the ad hoc synthesis of nanoDDS tailored to specific requirements for future nanomedicine strategies, as presented in Fig. 1.
 | ||
| Fig. 1 Schematic illustration representing the workflow for the potential application of DoE/RSM approach to the rational design of formulated nanocarriers. | ||
2. Experimental
2.1. Materials
Hyaluronic acid sodium salt (HA, ultra-low molecular weight, avg. MW = 13 kDa, Biosynth Carbosynth, Compton, UK) and linear polyethyleneimine (LPEI, MW = 2.5 kDa, Polysciences, Warrington, PA) were used as the starting polymers for NG formulation. SU-8 photoresist (MicroChem, Newton, MA) and polydimethylsiloxane (PDMS, Sylgard 184, Dow Corning, Midland, MI) were used for chip microfabrication. All other chemicals were purchased from Merck KGaA (Darmstadt, Germany) and used as received, without any further purification. Solvents were of analytical grade purity.2.2. Chip design and microfabrication
The microfluidic device was fabricated using standard photo- and soft-lithography techniques. SU-8 2075 photoresist was spin-coated with a thickness of 60 µm on a 3” silicon wafer and patterned by UV exposure through a chrome-on-glass photomask. The obtained master was silanized overnight in a chamber saturated with trimethylchlorosilane (TMCS) vapor to ease demolding.A PDMS solution (10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 w/w of prepolymer to curing agent ratio) was poured onto the silicon master and cured in an oven at 70 °C for 2 h. The cured PDMS replica was demolded, and microfluidic inlets and outlets were created using a biopsy puncher. Subsequently, the PDMS replica was bound to a glass microscope slide via oxygen plasma treatment (Femto Plasma Etcher, Diener electronic, Ebhausen, Germany) to complete chip assembly. Finally, hydrophobic modification of the channel surface was achieved by flowing a stream of TMCS-saturated nitrogen for 15 min, followed by thermal treatment at 140 °C for 10 min.
1 w/w of prepolymer to curing agent ratio) was poured onto the silicon master and cured in an oven at 70 °C for 2 h. The cured PDMS replica was demolded, and microfluidic inlets and outlets were created using a biopsy puncher. Subsequently, the PDMS replica was bound to a glass microscope slide via oxygen plasma treatment (Femto Plasma Etcher, Diener electronic, Ebhausen, Germany) to complete chip assembly. Finally, hydrophobic modification of the channel surface was achieved by flowing a stream of TMCS-saturated nitrogen for 15 min, followed by thermal treatment at 140 °C for 10 min.
The chip design featured a standard Hydrodynamic Flow Focusing (HFF) junction used as a droplet generator, as reported in Fig. 2. The inlet and outlet channels were 200 and 500 µm wide, respectively, while the junction orifice had a width of 80 µm and a length of 100 µm.
2.3. Polymer functionalization
In the second step, the alkyne-modified LPEI was reacted with RhB azide (RhB-N3) through copper(I)-catalyzed azide–alkyne cycloaddition (CuAAC). Derivatization of RhB-N3 is detailed in SI.
Propargyl LPEI (120 mg, 0.048 mmol) and RhB-N3 (15.6 mg, 0.029 mmol) were dissolved in 10 mL of DIW. The catalyst cupric sulfate (0.029 mmol) and the reducing agent sodium ascorbate (0.029 mmol) were added to the mixture, which was left under magnetic stirring for 24 h at 50 °C. After 2 days of dialysis against DIW (membrane MWCO: 100–500 Da), the product (LPEI-RhB) was lyophilized and stored as a purple powder at −20 °C.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) EDC
EDC![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NHS = 1
NHS = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5
5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2. The mixture was stirred for 3 h at RT and used for the microfluidic synthesis.
2. The mixture was stirred for 3 h at RT and used for the microfluidic synthesis.
2.4. NGs microfluidic synthesis
To generate droplets through the HFF microfluidic setup, two immiscible phases are required. A solution of mineral oil with 3 wt% Span 80 was injected as the continuous phase, while the aqueous polymer solution was injected as the dispersed phase (Fig. 2B).Three different MR values (5.75, 11.5, and 17.25) were evaluated in this study. In detail, 1.5 mL of the activated HA were mixed by vortexing for 1 min with 1.5 mL of a pre-prepared solution containing a selected amount (4.4, 8.8, or 13.2 mg) of a mixture of LPEI-RhB and pristine LPEI (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 by weight, adjusted to pH 4.5 with HCl to enhance the solubility of pristine LPEI).
1 by weight, adjusted to pH 4.5 with HCl to enhance the solubility of pristine LPEI).
The final aq. polymer solution and the oil phase were loaded into 2.5 mL and 10 mL gas-tight glass syringes (Hamilton Company, Reno, NV), respectively. The syringes were connected to the HFF device through Teflon tubing and actuated by programmable syringe pumps (Nemesys, CETONI GmbH, Korbussen, Germany).
Three different FRR levels (0.1, 0.2, and 0.4) were established by varying the dispersed phase flow rate (QD) between 1 and 4 µL min−1, while maintaining the continuous phase flow rate (QC) constant at 10 µL min−1.
Hence, MR and FRR defined a two-factor three-level study (FRR = 0.1, 0.2, 0.4; MR = 5.75, 11.5, 17.25). The resulting experimental design, consisting of nine different synthesis conditions obtained by varying both the MR and FRR parameters, is reported in Table 1.
Droplet formation was monitored over time under a Nikon Eclipse Ti–E fluorescence microscope (4× objective) equipped with a high-sensitivity camera (Neo 5.5, Andor Technology, Belfast, UK). To ensure reproducibility and proper device performance throughout the experiment, a time-lapse acquisition was carried out, capturing 1 frame every 10 minutes.
The emulsion collected at the outlet of the microfluidic system was broken after 8 h of mild agitation on a tilting plate and the aqueous phase was extracted in a separating funnel by washing with diethyl ether (3 × 10 mL). The NG suspension was dialyzed against DIW (membrane MWCO: 15 kDa) for two days with daily water replacement. Finally, the samples were lyophilized and stored at −20 °C.
2.5. Nuclear magnetic resonance (NMR)
Chemical composition of pristine polymers, intermediates, and final NGs was analyzed by 1H NMR. Analyses were carried out on a Bruker AC spectrometer (400 MHz, Bruker Corp., Billerica, MA) with deuterium oxide (D2O) as the solvent. Chemical shifts were reported as δ values (ppm) relative to the tetramethylsilane internal peak reference.2.6. Dynamic light scattering (DLS)
NG suspensions in ultrapure water (1 mg mL−1) were analyzed by DLS (Zetasizer Nano ZS, Malvern Panalytical, Malvern, UK) to determine particle size and polydispersity index (PDI). Before each measurement, specimens were sonicated for 10 min to minimize aggregation of the colloidal system. Readings were performed in triplicate.2.7. Drug release
In vitro release experiments were performed using DOX as a model chemotherapeutic drug.DOX-loaded nanogels (DOX_NGs) were prepared by dissolving lyophilized NG samples (1 mg) in 10 µL of a 5 mM DOX solution in DMSO. The system was centrifuged and left at RT for 30 min to complete drug loading and then diluted with PBS to the final NG concentration of 100 µg mL−1. 300 µL of the resulting suspension were loaded inside a microdialysis device (membrane MWCO: 3.5 kDa) and placed into a 96 deep-well plate (Thermo Fisher Scientific) allowing DOX_NGs to exchange against 1.4 mL of PBS at pH 7.4.
The drug release profile was investigated at 37 °C under gentle agitation (rocking shaker at 100 rpm). At each time point, the dialysis buffer was replaced with fresh PBS and the sequential release was quantified (λex = 488 nm, λem = 590 nm) on an Infinite M200 Pro multiplate reader (TECAN, Männedorf, Switzerland), based on an experimental standard curve for DOX (SI, Fig. S1).
2.8. Cell model
The human epithelial ovarian cancer cell line OVCA433 (American Type Culture Collection, ATCC, Manassas, VA) was selected as the biological model of interest, based on its relevance in previous studies.64,65 Cells were cultured in Dulbecco's modified Eagle's medium (DMEM low glucose, GIBCO, Thermo Fisher Scientific), supplemented with 10% fetal bovine serum, 100 U mL−1 penicillin/streptomycin and 1 mM L-glutamine. Cultures were maintained at 37 °C in a humidified atmosphere containing 5% CO2.2.9. Nanogel cytocompatibility
NG cytocompatibility was first assessed on OVCA433 cells using the Vybrant Cytotoxicity Assay (Thermo Fisher Scientific).66 Cells were seeded at a density of 2 × 103 cells per well into a 96-well plate for 24 h and then incubated with NGs at a concentration of 20 µg mL−1. The release of the cytosolic enzyme glucose 6-phosphate dehydrogenase (G6PD) from damaged cells into the surrounding medium was quantified after 24 h of incubation with NGs. 50 µL of supernatant were transferred into a 96-well plate and, after 30 min of incubation with 50 µL of resazurin/reaction mixture at 37 °C in 5% CO2, the fluorescent metabolite of resazurin (resorufin) was detected (λex = 530 nm, λem = 590 nm; TECAN Infinite M200 Pro). Cytotoxicity was calculated as the fluorescence intensity ratio between NG-treated cells and fully lysed control cells (positive control), previously background-corrected by subtracting from each reading the value of untreated cells (negative control). Experiments were performed in triplicate.To further assess NG cytocompatibility, a cell metabolic activity assay using 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) (Merck KGaA, Darmstadt, Germany) was performed. Cells were seeded at a density of 2 × 103 cells per well into a 96-well plate for 24 h and then incubated with NGs at a concentration of 20 µg mL−1 for 24 h. After incubation, the culture medium was removed, and 100 µL of MTT solution (0.5 mg mL−1) in complete cell culture medium were added to each well. Cells were incubated for 3 hours at 37 °C and 5% CO2. Subsequently, the MTT solution was removed, and the resulting formazan crystals were solubilized in 100 µL of dimethyl sulfoxide (DMSO). Absorbance was measured at 570 nm on a TECAN Infinite M200-Pro plate reader. Cell viability was calculated by dividing the absorbance of NG-treated cell cultures by that of the corresponding untreated controls. Experiments were performed in triplicate.
2.10. Flow cytometric analysis of NG internalization
Cells were seeded at a density of 2.4 × 104 cells per well into a 12-well plate and incubated for 24 h before supplementation with a NG suspension at 20 µg mL−1 for further 24 h. Subsequently, cells were analyzed by flow cytometry (CytoFLEX cytometer, Beckman Coulter, Brea, CA). Live cells were gated based on their forward and side scatter parameters; NG signal was recorded in the phycoerythrin (PE-A) channel and quantified as the median fluorescence intensity (MFI) value. Experiments were performed in triplicate.2.11. Confocal and super-resolution microscopy
NG internalization was also characterized using confocal and super-resolution microscopy. Cells were seeded at a density of 1.5 × 104 cells per well into 8-well glass bottom µ-Slide chambers (Ibidi GmbH, Gräfelfing, Germany) and grown for 24 h before the experiment. Cells were incubated with NGs at a concentration of 20 µg mL−1 for 24 h, then fixed with 4% buffered paraformaldehyde (PFA) for 15 min at RT and washed with PBS (3 times). For confocal microscopy, cell membranes were stained with CellMask™ Green Plasma Membrane Stain (Thermo Fisher Scientific; 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 dilution in PBS for 5 min) and nuclei were counterstained with DAPI (Thermo Fisher Scientific; 1
1000 dilution in PBS for 5 min) and nuclei were counterstained with DAPI (Thermo Fisher Scientific; 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 dilution in PBS for 10 min). Specimens were observed under a Nikon A1R+ laser scanning confocal microscope (Nikon Instruments, Tokyo, Japan) with a 20× NA0.7 dry objective. To gain deeper insight into NG internalization at the subcellular level, super-resolution optical microscopy was employed. This technique overcomes the limitations of conventional diffraction-limited optical imaging, allowing for more precise and detailed mapping of NG distribution. Cells were stained with labelled phalloidin (ActinGreen 488 stain, GeneCopoeia, Rockville, MD; 1
1000 dilution in PBS for 10 min). Specimens were observed under a Nikon A1R+ laser scanning confocal microscope (Nikon Instruments, Tokyo, Japan) with a 20× NA0.7 dry objective. To gain deeper insight into NG internalization at the subcellular level, super-resolution optical microscopy was employed. This technique overcomes the limitations of conventional diffraction-limited optical imaging, allowing for more precise and detailed mapping of NG distribution. Cells were stained with labelled phalloidin (ActinGreen 488 stain, GeneCopoeia, Rockville, MD; 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 in PBS for 45 min), and nuclei were counterstained with DAPI. Imaging was performed using a ZEISS ELYRA 7 microscope (ZEISS Microscopy, Jena, Germany) equipped with a 63× NA1.4 oil immersion objective, and raw datasets were processed using the Lattice SIM2 algorithm to enhance spatial resolution.
50 in PBS for 45 min), and nuclei were counterstained with DAPI. Imaging was performed using a ZEISS ELYRA 7 microscope (ZEISS Microscopy, Jena, Germany) equipped with a 63× NA1.4 oil immersion objective, and raw datasets were processed using the Lattice SIM2 algorithm to enhance spatial resolution.
2.12. Drug delivery to cancer cells
DOX_NG systems were prepared as described in Section 2.7, achieving a final DOX concentration of 1 µM while maintaining the NG concentration at 20 µg mL−1. The selected DOX concentration corresponds to a sublethal dose, determined from a dose–response curve evaluated at 24 hours on the OVCA433 cell line (SI, Fig. S2). The DOX_NG suspension was administered to OVCA433 cells seeded at a density of 2 × 103 cells per well into a 96-well plate. NGs without DOX and 1 µM free DOX administration were used as internal controls, alongside untreated cells used as a negative control. The effect of DOX-loaded NGs was quantified after 24 h using the MTT assay, following the protocol detailed in Section 2.9.To distinguish the specific contribution of DOX from the intrinsic cytotoxicity of nanogels (NGs) in the overall residual cell viability, a nanoencapsulation enhancement (NE) parameter was defined as follows:
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DOX are the absorbance levels of untreated cells and cells treated with free DOX, respectively.
DOX are the absorbance levels of untreated cells and cells treated with free DOX, respectively.
2.13. Empirical model
To build an empirical model, an RSM method was adopted, previously applied to various experimental contexts.67–71 The entire statistical analysis was conducted using the programming language R,72 following the statistical strategy described in previous works.67,73–75 An initial exploratory analysis was performed by calculating a correlation matrix, reporting the values of the Pearson's coefficients (r) to identify the potential linear relationships among the variables. Subsequently, a predictive model was constructed by selecting statistically significant factors, as determined by analysis of variance (ANOVA). The significance level was assigned as follows: p ≤ 0.1 (.), p ≤ 0.05 (*), p ≤ 0.01 (**), p ≤ 0.001 (***). The complete model is reported in eqn (1), where A and B represent the MR and FRR parameters, respectively. It should be noticed that not all the terms are necessarily included in the models; only those with p ≤ 0.1 were retained, while the model was considered significant if p ≤ 0.05. The model function (denoted as F in eqn (1)) was selected to fulfill two objectives: (i) normalization of the model residuals and (ii) minimization of residual patterning. To assess the goodness of fit of the model, the coefficient of determination (R2) was calculated, with R2 = 1 indicating a perfect fit. To evaluate the predictive performance of the model, the predicted residual error sum of squares (PRESS) statistic was applied. The resulting predicted R2 (Rpred2) represents a good estimator of the out-of-sample performance of the model.To further refine the model, we chose to restrict the space of acceptable solutions to NG formulations exhibiting high cytocompatibility. To this end, a binary desirability index value was defined, assigning values of 0 or 1 based on NG cytocompatibility. A threshold (t) of 90% cell viability was set as the cutoff criterion (see eqn (2)), ensuring that only formulations meeting this standard were considered desirable.
| F(Y) = c0 + c1A + c2B + c3AB + c4A2 + c5B2 + c6A2B + c7AB2 + c8A2B2 | (1) |
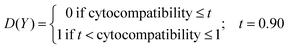 | (2) |
2.14. Statistical analysis
Experimental data were analyzed using Prism ver. 10.2.1 (GraphPad Software, San Diego, CA) and reported as mean ± SD, unless otherwise specified. Statistical significance was assessed by one-way or two-way analysis of variance (ANOVA), as appropriate, with a significance threshold set at the 0.05 level.3. Results
3.1. NG synthesis and characterization
In the present work, a microfluidic synthesis of HA-LPEI NGs based on an HFF droplet-generator device was investigated. Fig. 3A and B illustrate the operation of the HFF microfluidic device as a droplet generator under varying FRR levels: increasing FRR led to the formation of larger emulsion droplets, with the average droplet diameter increasing from approximately 30 to 60 µm over the investigated FRR range.The formation of NGs by chemical crosslinking between HA-NHS ester derivatives (degree of functionalization ≅40.5%, SI Fig. S3) and amine groups in the LPEI backbone (reaction scheme in Fig. 3C) was confirmed by 1H-NMR conducted on each NG formulation as well as on the starting polymers. Representative NMR spectra for each of the three MR levels are shown in Fig. 3D, while the complete set of spectra of all NG formulations is reported in SI, Fig. S4. To quantify the composition of each NG formulation, the weight fraction of LPEI (χLPEI) was estimated by integrating the NMR signals corresponding to the starting materials (HA and LPEI). The resulting χLPEI values are shown in Table 2.
| Sample | χ LPEI [−] | Diameter [nm] | PDI [−] |
|---|---|---|---|
| NG−1,−1 | 0.76 | 109.6 ± 10.5 | 0.105 |
| NG−1,−0.33 | 0.74 | 357.8 ± 12.0 | 0.030 |
| NG−1,+1 | 0.71 | 339.3 ± 33.3 | 0.010 |
| NG0,−1 | 0.82 | 184.5 ± 17.5 | 0.060 |
| NG0,−0.33 | 0.84 | 246.4 ± 21.9 | 0.038 |
| NG0,+1 | 0.81 | 339.2 ± 32.7 | 0.019 |
| NG+1,−1 | 0.84 | 111.7 ± 8.8 | 0.054 |
| NG+1,−0.33 | 0.86 | 190.3 ± 12.3 | 0.012 |
| NG+1,+1 | 0.85 | 292.5 ± 3.5 | 0.010 |
Characterization of NG size distribution was performed by DLS analysis (Table 2). A two-way ANOVA was performed on DLS data to assess the influence of the two main variables (MR and FRR) on the resulting morphological parameters. Fig. 4A and B show the interaction plots for NG size and PDI, respectively.
The effect of FRR was much larger (F(2,18) = 97.5, p < 0.0001, η2 = 76.6%) than that of MR (F(2,18) = 9.82, p = 0.0013, η2 = 7.71%). A significant interaction (F(4,18) = 5.52, p = 0.0044, η2 = 8.67%) was also reported. PDI showed a similar trend, with a significant interaction (F(4,16) = 3.74, p = 0.0248, η2 = 12.2%), and a much larger effect size for FRR (F(2,16) = 40.8, p < 0.0001, η2 = 66.5%) than for MR (F(2,16) = 5.16, p = 0.0186, η2 = 8.4%). These results are in line with the observed large influence of FRR (which dictates the size of the emulsion droplets) on nanogel size and PDI, with higher FRR levels resulting in smaller and more monodisperse nanogels.
NG composition (in terms of χLPEI) was also investigated by two-way ANOVA without replications (interaction plot in Fig. 4C). Results showed a large and significant positive main effect of MR on NG composition (F(2,4)= 49.0, p = 0.0015, η2 = 93.2%), while the effect of FRR was not significant (p = 0.2427).
Overall, these results demonstrate the versatility of droplet-based microfluidics in tuning the composition and size of NGs, enabling reproducible synthesis across diverse molar ratios and diameters.
3.2. Drug release
The selection of the appropriate method to evaluate the drug release profile from engineered nanomaterials is a source of debate in the scientific community.76–79 Despite the reliability of the data obtained by the dialysis method has been questioned in the past, it is by far the most used methodology in the field.76 In conventional dialysis experiments, two stages may influence the apparent release kinetics from the nanomaterial: (1) the release of the cargo from the nanomaterials into the surrounding solution inside the dialysis bag; and (2) the permeation of the cargo through the dialysis membrane. In most cases, this second step is considered negligible, allowing the actual release kinetics to be revealed by measuring the concentration of the cargo in the receiver compartment.However, in the present case, the permeation of free DOX through the membrane cannot be neglected, as shown in SI, Fig. S7A. This behavior can be attributed to the well-documented interaction between DOX and the dialysis membrane,79–83 which can lead to an underestimation of the release constant from NGs. Accordingly, the amount of drug released at infinite time (M∞) was selected as the defining parameter for characterizing the release of drug from NG-DOX systems (Table 3).
| Sample | M∞ (%) |
|---|---|
| NG−1,−1 | 78.9 ± 2.3 |
| NG−1,−0.33 | 49.9 ± 3.0 |
| NG−1,+1 | 63.1 ± 1.0 |
| NG0,−1 | 49.7 ± 2.3 |
| NG0,−0.33 | 50.8 ± 1.2 |
| NG0,+1 | 67.7 ± 4.0 |
| NG+1,−1 | 87.8 ± 6.7 |
| NG+1,−0.33 | 67.7 ± 3.8 |
| NG+1,+1 | 63.3 ± 3.1 |
M ∞ levels were also analyzed by two-way ANOVA (interaction plot in Fig. 4D). Results showed a large interaction (F(4,17) = 12.0, p < 0.0001, η2 = 37.4%), indicating that the impact of each factor was contingent on the level of the other. This interdependence limited the interpretability of main effects.
3.3. Biocompatibility and cellular internalization
To preliminarily validate the NG formulations as an effective DDS for the treatment of ovarian carcinoma, their cytocompatibility and cellular internalization were investigated in combination with OVCA433 cells.First, the cytocompatibility of the NGs was investigated through G6PD and MTT assays following 24 h of cell exposure to NGs at a concentration of 20 µg mL−1. Notably, none of the NG formulations induced a statistically significant reduction in cell viability as measured by the G6PD assay (SI, Fig. S8), which detects the release of glucose 6-phosphate dehydrogenase from damaged or dying cells. However, the evaluation of cellular metabolic activity via the MTT assay revealed a different cytocompatibility profile, as reported in Table 4. These results suggest that, under the tested conditions, NGs did not induce cell death or membrane damage. Rather, LPEI-rich compositions (corresponding to the highest MR level) led to a reduction in cell metabolic activity, with cells remaining viable but metabolically compromised.
| Sample | NG cytocompatibility (%) (MTT assay) |
|---|---|
| NG−1,−1 | 109.3 ± 18.9 |
| NG−1,−0.33 | 99.6 ± 11.1 |
| NG−1,+1 | 102.2 ± 10.2 |
| NG0,−1 | 89.3 ± 16.0 |
| NG0,−0.33 | 105.2 ± 10.6 |
| NG0,+1 | 94.2 ± 4.9 |
| NG+1,−1 | 66.8 ± 11.4 |
| NG+1,−0.33 | 60.5 ± 4.5 |
| NG+1,+1 | 61.8 ± 4.8 |
Results of two-way ANOVA (interaction plot in Fig. 5A) showed that MR was the only factor with a strong and extremely significant negative main effect (F(2,18) = 169, p < 0.0001, η2 = 72.3%) on NG cytocompatibility, despite significance of the interaction term (F(6,18) = 4.37, p = 0.0068, η2 = 5.60%).
Cellular uptake of NGs was characterized using a combination of flow cytometry and fluorescence microscopy (confocal and super-resolution), leveraging the intrinsic fluorescence of labelled LPEI (LPEI-RhB). The raw flow cytometry data were adjusted with a correction coefficient accounting for the different χLPEI values, as measured by NMR. This adjustment compensates for differences in the amount of fluorescent dye present in each formulation due to variations in the total LPEI content (complete discussion has been reported in SI).
Table 5 reports the raw fluorescence data, the corrective factors, and the corrected fluorescence data, which more accurately reflect the amount of cell internalization. Two-way ANOVA showed that the main effect of MR (F(2,18) = 384, p < 0.0001, η2 = 54.9%) overcame that of FRR (F(2,18) = 10.4, p = 0.0010, η2 = 1.48%) on NG uptake, with the two factors having an extremely significant interaction (F(4,18) = 148, p < 0.0001, η2 = 42.3%). Despite the large interaction hampering the interpretation of main effects, it can be observed that the specimens with the highest HA content (NG−1,x) were characterized by the highest internalization values.
| Sample | Raw fluorescence (MFI) | Correction factor | Corrected fluorescence (MFI) |
|---|---|---|---|
| NG−1,−1 | 2.29 ± 0.12 | 1.6 | 3.61 ± 0.19 |
| NG−1,−0.33 | 3.17 ± 0.07 | 1.7 | 5.46 ± 0.13 |
| NG−1,+1 | 1.75 ± 0.02 | 1.9 | 3.35 ± 0.04 |
| NG0,−1 | 2.53 ± 0.06 | 1.2 | 3.05 ± 0.07 |
| NG0,−0.33 | 2.04 ± 0.09 | 1.1 | 2.30 ± 0.10 |
| NG0,+1 | 2.38 ± 0.05 | 1.3 | 3.02 ± 0.06 |
| NG+1,−1 | 2.40 ± 0.04 | 1.1 | 2.66 ± 0.05 |
| NG+1,−0.33 | 2.28 ± 0.05 | 1 | 2.28 ± 0.05 |
| NG+1,+1 | 2.78 ± 0.24 | 1.1 | 2.99 ± 0.26 |
Further confirmation of successful NG uptake was obtained through confocal and super-resolution microscopy (Fig. 5C and D). Red fluorescent NGs exhibited intracellular localization (cell membrane stained in green) (Fig. 5C). Super-resolution micrographs (Fig. 5D) gave a clearer insight into the NG distribution within the cells. Noticeably, the reported figures highlighted the distribution of the NGs around the cell nucleus without evidence of aggregation. These findings suggest that the formulated NGs maintain their structural integrity within the cellular environment up to 24 hours post-administration.
3.4. Drug response
The performance of NGs as drug delivery systems was tested in combination with doxorubicin (DOX, 1 µM) as a model chemotherapeutic agent, using the MTT assay.Fig. 6A shows the residual cellular metabolic activity after 24 h following administration of DOX_NGs. Data were normalized to untreated controls and statistically tested against cells treated with free DOX.
It is worth noting that most of the NG samples showed increased toxicity to cells compared to free DOX. However, these results could be misleading, as they do not account for the intrinsic cytotoxic effect of the NG carriers themselves—particularly in LPEI-rich compositions—on cell viability. Accordingly, a nanoencapsulation-related enhancement (NE%) parameter was defined as described in Section 2.12 to isolate the contribution of drug delivery from carrier-induced effects. The NE% values, which more accurately reflect the enhancement due to nanoencapsulation, are reported in Fig. 6B.
Two-way ANOVA on NE data (in Fig. 6C) revealed a large and significant negative main effect of MR (F(2,18) = 209, p < 0.0001, η2 = 84.9%), a modest but very significant (F(2,18) = 10.3, p = 0.001, η2 = 4.81%) main effect of FRR, and a modest but extremely significant interaction (F(4,18) = 8.97, p = 0.0004, η2 = 7.28%).
3.5. Empirical model
The experimental data collected were used to construct an empirical model based on RSM. As a first step, an exploratory analysis was conducted by calculating a correlation matrix (Pearson's) to identify first-order relationships among variables, including both process parameters and NG properties (Fig. 7). Only variables with |r| ≥ 0.4 and p < 0.05 were considered significantly correlated.FRR showed a strong direct correlation with NG size and a strong inverse correlation with PDI. Expectedly, also NG size and PDI were inversely correlated.
Not surprisingly, MR exhibited a strong correlation with the resulting χLPEI. Both MR and χLPEI showed a strong inverse correlation with NG cytocompatibility, which can be explained by the potential toxicity of the LPEI component.
Moreover, MR and χLPEI were inversely correlated with the cellular uptake. This can be ascribed to the lower hyaluronan content of LPEI-rich formulations (χHA = 1 − χLPEI) which, in turn, can modulate NG internalization in cells expressing the hyaluronan receptor (CD44).
The implementation of the RSM led to the development of an empirical model, represented by the following set of equations, with the two variables (A = MR; B = FRR) expressed in coded units:
| Size (nm) = 2.73 × 102 − 9.25 × 101A + 7.74 × 101B − 1.22 × 101AB + 5.15 × 101A2 − 1.16 × 101B2 + 2.52 × 101A2B + 8.13 × 101AB2 − 1.00 × 102A2B2 | (3) |
| PDI (−) = 1.50 × 10−2 − 1.13 × 10−2A − 2.99 × 10−2B + 1.15 × 10−2AB + 2.82 × 10−2B2 | (4) |
| χLPEI (−) = 8.27 × 10−1 + 5.66 × 10−2A − 3.06 × 10−2A2 | (5) |
| [M∞ (%)]−1 = 1.81 × 10−2 − 2.94 × 10−3A − 2.78 × 10−3B + 4.53 × 10−3A2B + 2.74 × 10−3AB2 − 4.06 × 10−3A2B2 | (6) |
| Cellular uptake (−) = 2.21 − 1.69 A + 1.51 × 10−1AB + 1.76 A2 + 8.30 × 10−1B2 + 1.36AB2 − 1.64 A2B2 | (7) |
| NG cytocompatibility (%) = 1.07 × 102 − 2.03 × 101A − 2.86 × 101A2 − 1.51 × 101B2 − 3.03A2B + 2.19 × 101A2B2 | (8) |
| NE (%) = 1.07 × 102 − 1.15 × 101A − 5.12 × 101B + 1.64 × 101AB + 7.57A2 − 1.93 × 101A2B | (9) |
The complete set of variance and coefficients tables for eqn (3)–(9) are reported in SI (Tables S2–S15), while the surface contour plots are reported in Fig. 8.
With RSM, the dependence of the NG properties on the two process variables was described by polynomials of higher degree, which effectively captured non-linear relationships that were not apparent in the initial linear correlation analysis. To avoid overfitting, the degree of the polynomials was kept as low as possible, while still ensuring a high-quality fit of the experimental data. Moreover, we chose to limit the space of solutions to those formulations maintaining the cellular metabolic activity above 90%. This constraint was implemented via a desirability function (Fig. 8A and B) and the corresponding “non-desirable” region has been hatched in each contour plot (Fig. 8C–H).
To qualify the developed empirical models, the predicted vs. actual plots (Fig. 9) were analyzed. The coefficient of determination (R2) gave an indication of the goodness of fit, resulting in good fit quality for all the equations (R2 > 0.8). Noteworthy, an R2 > 0.9 was reached for NG cytocompatibility (R2 = 0.953), size (R2 = 0.969), χLPEI (R2 = 0.924), and cellular uptake (R2 = 0.987).
Although R2 is a robust indicator of the goodness of fit, it does not quantify the ability of the model to predict responses for new, unseen observations—those not used during model estimation. To address this limitation, we calculated the predicted R2 (Rpred2). For all the equations of the empirical model, the Rpred2 values exceeded 0.7, indicating good predictive quality.
4. Discussion
Recently, the use of microfluidics in the formulation of novel DDSs has gained increasing attention mainly due to the high control over a variety of process parameters, which outperforms traditional batch syntheses. However, the flexibility in modulating the fluid-dynamic and compositional parameters offered by microfluidic setups necessitates a time-intensive optimization to identify the appropriate process conditions. To address this challenge, a DoE approach incorporating RSM was employed to systematically investigate the influence of process parameters over a wide range of physicochemical and biological features of HA-LPEI NGs synthesized via droplet-based microfluidics.4.1. Effects of process parameters on NG physicochemical properties
To validate the application of droplet-based microfluidics for the synthesis of polymeric NGs and to deepen the understanding of the effect that process parameters exert over NGs features, the chemical and physical properties of these nanomaterials were investigated in a range of process parameter values.Comparison of the 1H-NMR spectra of the starting polymers with that of the NG samples (Fig. 3D) revealed signals from both polymers, along with a chemical shift of LPEI, indicative of successful conjugation. Specifically, characteristic peaks of LPEI could be detected at 2.78 ppm (†) (–CH2–CH2–NH–). Upon conjugation, this signal shifted slightly downfield, reflecting changes in the chemical environment and supporting the successful formation of amide bonds (¶¶, §§, ##).84
For hyaluronic acid, the 1H-NMR spectrum in D2O shows a distinct singlet at ∼1.9 ppm (***), attributed to the methyl protons of the N-acetyl group (–NHCOCH3). Additionally, multiple signals appeared in the 3.2–4.0 ppm range (**) corresponding to the ring protons of the glucuronic acid and glucosamine residues. The anomeric proton appeared around 4.4–4.6 ppm (*).
In the NGs, the involvement of the activated carboxyl groups of HA resulted in the shift of the signal ascribable to the protons in the sugar ring from 3.8 ppm to 3.38 ppm (¶, §, #), thus corroborating the bonding between polymers for all NG specimens.
Successful chemical crosslinking between HA and LPEI was further confirmed by FTIR spectroscopy (SI, Fig. S5).
Integration of NMR signals ascribable to HA and LPEI enabled estimation of the χLPEI for each NG formulation, as reported in Table 2. A strong dependence on MR was confirmed (92.3% of the total variance, p = 0.0015), while no significant main effect of FRR was detected. These results are matched by the strong linear correlation between MR and χLPEI (r = 0.92, p < 0.001).
Interestingly, as reported in Section 3.1, a major direct influence of FRR on NGs size can be highlighted by the two-way ANOVA (FRR accounts for 76.6% of the total variance, p < 0.0001). This might be attributed to the reduction of microdroplet size achieved for low FRR levels, which limits the volume of the reaction chamber where the crosslinking reaction takes place. A reduction in the droplet diameter from ca. 60 to 30 µm was observed when the FRR was decreased from 0.4 to 0.1, resulting in a volume change of approximately 87.5% (Fig. 3A and B). MR, on the other side, demonstrated a minor influence on NG size (8.67% of the total variance, p = 0.0044). However, the data revealed a slight inverse proportionality, highlighting the pivotal role of LPEI as a crosslinker. Indeed, NGs with a lower LPEI content exhibited an overall larger diameter, likely due to a less densely crosslinked matrix. NG size ranged from 100 to 360 nm (Table 2), consistently falling within the optimal range for drug delivery applications across all process parameters.
All the experimental conditions resulted in monodisperse nanoparticles with PDI values ranging from 0.01 to 0.11 (Table 2). A strong negative effect of FRR on PDI was evidenced by two-way ANOVA (66.5% of the total variance, p < 0.0001) and by the strong inverse correlation between the two variables (r = −0.76, p = 0.017). This behavior could be explained in light of previous literature findings85 evidencing that the mixing processes occurring within the emulsified dispersed phase are enhanced by an increased droplet velocity. Hence, the narrow size distribution obtained at higher FRR could be ascribed to improved mixing conditions, as reported for other microfluidics-based processes for nanoparticles synthesis.86–90 Based on the above-reported considerations, also NG size and PDI were found to be inversely correlated (r = −0.78, p = 0.013).
Transmission electron microscopy (TEM) served as an ancillary characterization technique, used to analyze dry-state diameters of the nanogels, which appeared reduced compared to their solvated counterparts. Representative micrographs are presented in SI (Fig. S6). To further explore NGs stability, ζ-potential measurements were performed (SI, Table S1). Interestingly, the data suggested a trend in ζ-potential values relative to the MR used during NG synthesis, which itself is closely linked to LPEI content. Specifically, NGs synthesized with a low MR tended to exhibit negative or near-neutral ζ-potentials, whereas those prepared with higher MRs (11.5 or 17.25) showed more positive values.
4.2. Physicochemical properties affect NG bioperformance
In this study, the bioperformance of NGs was assessed by evaluating drug release profiles, cytotoxicity, cellular uptake, and their functionality as DDSs in ovarian cancer cells. Building on the previously established relationship between synthesis parameters and the physicochemical properties of NGs, this section explores how these properties influence the in vitro bioperformance of the nanomaterials.A strong and significant inverse correlation (r = −0.71, p = 0.033) between χLPEI and NG cytocompatibility was evidenced. Indeed, LPEI-rich formulations (NG+1,x) resulted in reduced cell viability (71.2%, 60.5% and 61.8%, respectively) compared to the control group. Linear and branched PEI are known for toxicity and non-degradability, which restrict their clinical applicability.91,92 However, the intrinsic toxicity of LPEI can be mitigated through conjugation with other biopolymers such as HA,93 as inferred from Table 4, where the NGs with higher HA content demonstrated no significant variation in metabolic activity (MTT assay) compared to untreated cells.
χ LPEI was also reported to dictate the cellular uptake of NGs. Data suggested that NGs with a higher LPEI content were less efficiently internalized after 24 hours. This observation is also supported by the strong negative correlation between χLPEI and cellular uptake (r = −0.71, p = 0.032), meaning that NGs with a higher HA fraction facilitated cell internalization. This behavior can be attributed to the interaction between HA and its receptor, CD44, which is highly expressed in ovarian cancer cells and plays a role in mediating NG endocytosis, as already demonstrated for NGs with similar composition.65 Indeed, cell uptake quantification by flow cytometry revealed the highest fluorescence levels for HA-rich formulations (NG−1,x), with NG−1,−0.33 exhibiting approximately a six fold increase compared to untreated cells.
The NE parameter has been defined as a metric of nanoencapsulation-driven enhancement of drug effect, subtracting the account for the effect of pristine NG material on cell viability. It is worth noting that, in determining the NE, a superposition principle has been applied—under the strong assumption that the net response elicited by the two stimuli (pristine NG and released DOX) is equivalent to the sum of the individual responses that each stimulus would produce independently. Nonetheless, this parameter allowed us to compare different NG formulations, which are characterized by distinct cytocompatibility levels. χLPEI was inversely correlated with NE (r = −0.73, p = 0.027), suggesting that HA-rich formulations (low χLPEI), which are both highly biocompatible and readily internalized by cells, exhibit a more pronounced effect from drug nanoencapsulation strategy, surpassing that of free DOX. Expectedly, given the strong dependence of χLPEI on MR, similar trends were observed for NG cytocompatibility, cellular uptake, and NE, all of which showed a direct correlation with MR. It is also worth mentioning that, although the use of linear correlations was helpful in identifying the key dependences among different variables, the nature of the studied phenomena goes far beyond linearity. Thus, an interpretation limited to such a type of relationship may be misleading and may conceal high-order phenomena, which are instead evidenced by the polynomial equations of the response surface model.
4.3. RSM to dictate process parameters selection in microfluidic synthesis of nanomaterials
The RSM-based empirical model developed from the experimental data demonstrated good fit quality, with R2 > 0.81 for all the equations. However, although R2 is an adequate metric for the goodness of fit, it does not extend to evaluating the model's predictive strength for data points outside the training set. Indeed, a regression model could fit the existing data well, but it could not be as good at making predictions. Rpred2 is a statistical tool which helps elucidate the ability of a regression model to make predictions by removing one data point at a time and each time recalculating the regression to test how well the model predicts the missing observation. Rpred2 values were generally satisfactory for all the equations and did not differ markedly from the corresponding R2 (less than 13.1% decrease), suggesting that the empirical model was not overfitting the experimental data. This finding supports the use of RSM as a predictive tool for the microfluidic droplet-based synthesis of HA-LPEI NGs and, more broadly, for the continuous flow synthesis of nano-DDSs, allowing operators to achieve target responses by tuning process parameters.4.4. Future outlook
Despite the promises of NGs in advancing nanomedicine therapeutics, several challenges concerning stability, biodistribution, and pharmacokinetics under in vivo conditions are still open.94–96 After systemic administration, NG interaction with biological fluids may reduce the colloidal stability of the system, and enzymatic degradation may occur. Moreover, the eventual formation of the protein corona may induce opsonization followed by rapid clearance by the mononuclear phagocyte system.95It is well-documented that physicochemical parameters, such as size, shape, surface charge, and surface functionalization, play a crucial role in determining the pharmacokinetic profile and biodistribution of nanosystems. At the same time, pathological conditions produce damage to tissues, inducing the enhanced permeability and retention (EPR) effect, which enables the peripheral accumulation of designed nanocarriers.95,96
Microfluidic platforms and the DoE/RSM approach represent the ideal framework to systematically investigate how the variation of well-defined process parameters may affect in vivo behavior of NGs, ultimately contributing to a deeper understanding and improved design of nanodrugs.
5. Conclusion
A statistical approach was applied to the continuous flow synthesis of HA-LPEI NGs, produced by droplet microfluidics under a broad range of experimental conditions. The resulting panel of NG formulations were characterized in terms of size, polydispersity, composition, cytocompatibility, drug release, cellular uptake, and performance as a DDS on ovarian cancer cells. The fluid dynamic regime, represented by the flow rate ratio (FRR) parameter, was found to govern the resulting size and polydispersity index of the NGs, while compositional parameters (molar ratio between LPEI and HA repeating units, MR) dictated the resulting NG composition and influenced key biological properties, such as NG cytocompatibility, cellular uptake and enhancement of drug efficacy via nanoencapsulation. Using the Response Surface Methodology, we developed an empirical model describing the effects of process parameters on NG physicochemical and biological properties. The model demonstrated strong fit quality and robustness to out-of-sample predictions. In conclusion, this work highlights the potential of RSM to guide and support the continuous flow synthesis of tailored hyaluronan-based nanomaterials responding to specific design requirements for future nanomedicine applications.Author contributions
E. Limiti, E. D’Alessandro: methodology, investigation, visualization, writing – original draft. A. Bucciarelli: methodology, formal analysis, data curation, visualization. S. Raniolo: methodology, investigation. P. Mozetic, E. De Luca, E.D. Lemma, F. Basoli: investigation, writing – review & editing, supervision. G. Gigli: resources, writing – review & editing, supervision. F. Pizzetti: investigation. E. Mauri: conceptualization, methodology, writing – review & editing, supervision. F. Rossi, M. Trombetta, S. M. Giannitelli: resources, writing – review & editing, funding acquisition, supervision. A. Rainer: conceptualization, validation, writing – review & editing, funding acquisition, project administration.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5tb02518d.Acknowledgements
This work was funded by the European Union - Next Generation EU, Italian NRRP M4C2 Investment 1.5 “Rome Technopole”, CUP: C83C22000510001. P. Mozetic and E. De Luca are supported by the “National Center for Gene Therapy and Drug based on RNA Technology” (CN00000041 CN3 RNA), funded by the European Union - Next Generation EU, Italian NRRP M4.C2 - Investment 1.4. The authors are grateful to the “Tecnopolo per la medicina di precisione” (TecnoMed Puglia) - Regione Puglia: DGR n.2117 del 21/11/2018, CUP: B84I18000540002. E. D. Lemma is supported by the Italian Ministry for University and Research (Young Researchers - Seal of Excellence, CUP C83C22001250006). A. Rainer is supported by the Italian Ministry of Research, under the complementary actions to the NRRP “Fit4MedRob - Fit for Medical Robotics” Grant (# PNC0000007)”.References
- R. Langer, Drug delivery and targeting, Nature, 1998, 392(6679 Suppl), 5–10 CAS.
- M. R. Prausnitz and R. Langer, Transdermal drug delivery, Nat. Biotechnol., 2008, 26(11), 1261–1268, DOI:10.1038/nbt.1504.
- M. W. Tibbitt, J. E. Dahlman and R. Langer, Emerging Frontiers in Drug Delivery, J. Am. Chem. Soc., 2016, 138(3), 704–717, DOI:10.1021/jacs.5b09974.
- S. Vinogradov, E. Batrakova and A. Kabanov, Poly(ethylene glycol)–polyethyleneimine NanoGelTM particles: novel drug delivery systems for antisense oligonucleotides, Colloids Surf., B, 1999, 16(1–4), 291–304, DOI:10.1016/S0927-7765(99)00080-6.
- J. Gao, J. M. Karp, R. Langer and N. Joshi, The Future of Drug Delivery, Chem. Mater., 2023, 35(2), 359–363, DOI:10.1021/acs.chemmater.2c03003.
- S. Ahmed, K. Alhareth and N. Mignet, Advancement in nanogel formulations provides controlled drug release, Int. J. Pharm., 2020, 584, 119435, DOI:10.1016/j.ijpharm.2020.119435.
- Brianna, A. Anwar, S. Y. Teow and Y. S. Wu, Nanogel-based drug delivery system as a treatment modality for diverse diseases: Are we there yet?, J. Drug Delivery Sci. Technol., 2024, 91, 105224, DOI:10.1016/j.jddst.2023.105224.
- J. Gupta and G. Sharma, Nanogel: A versatile drug delivery system for the treatment of various diseases and their future perspective, Drug Delivery Transl. Res., 2025, 15(2), 455–482, DOI:10.1007/s13346-024-01684-w.
- I. Neamtu, A. G. Rusu, A. Diaconu, L. E. Nita and A. P. Chiriac, Basic concepts and recent advances in nanogels as carriers for medical applications, Drug Delivery, 2017, 24(1), 539–557, DOI:10.1080/10717544.2016.1276232.
- S. Shah, N. Rangaraj, K. Laxmikeshav and S. Sampathi, Nanogels as drug carriers – Introduction, chemical aspects, release mechanisms and potential applications, Int. J. Pharm., 2020, 581, 119268, DOI:10.1016/j.ijpharm.2020.119268.
- D. Suzuki, Nanogel/Microgel Science and Beyond, Langmuir, 2023, 39(22), 7525–7529, DOI:10.1021/acs.langmuir.3c00560.
- S. Hajebi, N. Rabiee and M. Bagherzadeh, et al., Stimulus-responsive polymeric nanogels as smart drug delivery systems, Acta Biomater., 2019, 92, 1–18, DOI:10.1016/j.actbio.2019.05.018.
- S. Mura, J. Nicolas and P. Couvreur, Stimuli-responsive nanocarriers for drug delivery, Nat. Mater., 2013, 12(11), 991–1003, DOI:10.1038/nmat3776.
- Z. Sun, Z. Yi and H. Zhang, et al., Bio-responsive alginate-keratin composite nanogels with enhanced drug loading efficiency for cancer therapy, Carbohydr. Polym., 2017, 175, 159–169, DOI:10.1016/j.carbpol.2017.07.078.
- Y. Wang, J. Zheng, Y. Tian and W. Yang, Acid degradable poly(vinylcaprolactam)-based nanogels with ketal linkages for drug delivery, J. Mater. Chem. B, 2015, 3(28), 5824–5832, 10.1039/C5TB00703H.
- T. A. Debele, S. L. Mekuria and H. C. Tsai, Polysaccharide based nanogels in the drug delivery system: Application as the carrier of pharmaceutical agents, Mater. Sci. Eng., C, 2016, 68, 964–981, DOI:10.1016/j.msec.2016.05.121.
- H. Zhao, Q. Li, A. Shi, J. Cui, W. Chen and J. Wu, Application of Plant Polysaccharide Nanoparticles as Polymeric Carrier Materials for the Construction of Medicine Carriers, J. Clust. Sci., 2023, 34(5), 2215–2228, DOI:10.1007/s10876-022-02393-5.
- C. Y. Yu, Y. M. Wang and N. M. Li, et al., In Vitro and in Vivo Evaluation of Pectin-Based Nanoparticles for Hepatocellular Carcinoma Drug Chemotherapy, Mol. Pharm., 2014, 11(2), 638–644, DOI:10.1021/mp400412c.
- Y. Liu, Y. Zong and Z. Yang, et al., Dual-targeted controlled delivery based on folic acid modified pectin-based nanoparticles for combination therapy of liver cancer, ACS Sustainable Chem. Eng., 2019, 7(3), 3614–3623, DOI:10.1021/acssuschemeng.8b06586.
- Y. Liang, Y. Wang and L. Wang, et al., Self-crosslinkable chitosan-hyaluronic acid dialdehyde nanoparticles for CD44-targeted siRNA delivery to treat bladder cancer, Bioact. Mater., 2021, 6(2), 433–446, DOI:10.1016/j.bioactmat.2020.08.019.
- J. Li, M. Li and L. Tian, et al., Facile strategy by hyaluronic acid functional carbon dot-doxorubicin nanoparticles for CD44 targeted drug delivery and enhanced breast cancer therapy, Int. J. Pharm., 2020, 578, 119122, DOI:10.1016/j.ijpharm.2020.119122.
- M. C. De Paula, S. G. Carvalho, A. L. P. Silvestre, A. M. Dos Santos, A. B. Meneguin and M. Chorilli, The role of hyaluronic acid in the design and functionalization of nanoparticles for the treatment of colorectal cancer, Carbohydr. Polym., 2023, 320, 121257, DOI:10.1016/j.carbpol.2023.121257.
- Z. Ma, B. Li, J. Peng and D. Gao, Recent Development of Drug Delivery Systems through Microfluidics: From Synthesis to Evaluation, Pharmaceutics, 2022, 14(2), 434, DOI:10.3390/pharmaceutics14020434.
- Y. Liu, G. Yang, Y. Hui, S. Ranaweera and C. Zhao, Microfluidic Nanoparticles for Drug Delivery, Small, 2022, 18(36), 2106580, DOI:10.1002/smll.202106580.
- S. Marre, Y. Roig and C. Aymonier, Supercritical microfluidics: Opportunities in flow-through chemistry and materials science, J. Supercrit. Fluids, 2012, 66, 251–264, DOI:10.1016/j.supflu.2011.11.029.
- D. T. Chiu, A. J. deMello and D. Di Carlo, et al., Small but Perfectly Formed? Successes, Challenges, and Opportunities for Microfluidics in the Chemical and Biological Sciences | Elsevier Enhanced Reader, Chem, 2017, 2(2), 201–223, DOI:10.1016/j.chempr.2017.01.009.
- S. J. Shepherd, D. Issadore and M. J. Mitchell, Microfluidic formulation of nanoparticles for biomedical applications, Biomaterials, 2021, 274, 120826, DOI:10.1016/j.biomaterials.2021.120826.
- S. Yadavali, D. Lee and D. Issadore, Robust Microfabrication of Highly Parallelized Three-Dimensional Microfluidics on Silicon, Sci. Rep., 2019, 9(1), 12213, DOI:10.1038/s41598-019-48515-4.
- J. Wu, S. Yadavali, D. A. Issadore and D. Lee, Ultrahigh Throughput On-Chip Synthesis of Microgels with Tunable Mechanical Properties, Adv. Mater. Technol., 2022, 7(5), 2101160, DOI:10.1002/admt.202101160.
- S. J. Shepherd, C. C. Warzecha and S. Yadavali, et al., Scalable mRNA and siRNA Lipid Nanoparticle Production Using a Parallelized Microfluidic Device, Nano Lett., 2021, 21(13), 5671–5680, DOI:10.1021/acs.nanolett.1c01353.
- S. Nawar, J. K. Stolaroff and C. Ye, et al., Parallelizable microfluidic dropmakers with multilayer geometry for the generation of double emulsions, Lab Chip, 2020, 20(1), 147–154, 10.1039/C9LC00966C.
- J. Wu, S. Yadavali, D. Lee and D. A. Issadore, Scaling up the throughput of microfluidic droplet-based materials synthesis: A review of recent progress and outlook, Appl. Phys. Rev., 2021, 8(3), 031304, DOI:10.1063/5.0049897.
- S. Gimondi, H. Ferreira, R. L. Reis and N. M. Neves, Microfluidic Devices: A Tool for Nanoparticle Synthesis and Performance Evaluation, ACS Nano, 2023, 17(15), 14205–14228, DOI:10.1021/acsnano.3c01117.
- M. Su, Y. Zhu and J. Chen, et al., Microfluidic synthesis of manganese-alginate nanogels with self-supplying H2O2 capability for synergistic chemo/chemodynamic therapy and boosting anticancer immunity, Chem. Eng. J., 2022, 435, 134926, DOI:10.1016/j.cej.2022.134926.
- T. Baby, Y. Liu, G. Yang, D. Chen and C. X. Zhao, Microfluidic synthesis of curcumin loaded polymer nanoparticles with tunable drug loading and pH-triggered release, J. Colloid Interface Sci., 2021, 594, 474–484, DOI:10.1016/j.jcis.2021.03.035.
- H. Chen, A. E. Celik and A. Mutschler, et al., Assembly of Fluorescent Polymer Nanoparticles Using Different Microfluidic Mixers, Langmuir, 2022, 38(26), 7945–7955, DOI:10.1021/acs.langmuir.2c00534.
- A. G. Mares, G. Pacassoni, J. S. Marti, S. Pujals and L. Albertazzi, Formulation of tunable size PLGA-PEG nanoparticles for drug delivery using microfluidic technology, ed J. Das Neves, PLOS ONE, 2021;vol. 16(6), p.e0251821 DOI:10.1371/journal.pone.0251821.
- T. Baby, Y. Liu, A. P. J. Middelberg and C. X. Zhao, Fundamental studies on throughput capacities of hydrodynamic flow-focusing microfluidics for producing monodisperse polymer nanoparticles, Chem. Eng. Sci., 2017, 169, 128–139, DOI:10.1016/j.ces.2017.04.046.
- S. I. Hamdallah, R. Zoqlam and P. Erfle, et al., Microfluidics for pharmaceutical nanoparticle fabrication: The truth and the myth, Int. J. Pharm., 2020, 584, 119408, DOI:10.1016/j.ijpharm.2020.119408.
- O. Kašpar, A. H. Koyuncu, A. Pittermannová, P. Ulbrich and V. Tokárová, Governing factors for preparation of silver nanoparticles using droplet-based microfluidic device, Biomed. Microdevices, 2019, 21(4), 88, DOI:10.1007/s10544-019-0435-4.
- C. D. Ahrberg, J. W. Choi and B. G. Chung, Droplet-based synthesis of homogeneous magnetic iron oxide nanoparticles, Beilstein J. Nanotechnol., 2018, 9, 2413–2420, DOI:10.3762/bjnano.9.226.
- J. W. Choi, Y. J. Kim, J. M. Lee, J. H. Choi, J. W. Choi and B. G. Chung, Droplet-based Synthesis of Homogeneous Gold Nanoparticles for Enhancing HRP-based ELISA Signals, BioChip J., 2020, 14(3), 298–307, DOI:10.1007/s13206-020-4307-z.
- L. Zou, B. Huang and X. Zheng, et al., Microfluidic synthesis of magnetic nanoparticles in droplet-based microreactors, Mater. Chem. Phys., 2022, 276, 125384, DOI:10.1016/j.matchemphys.2021.125384.
- N. Politis, S. Colombo, P. Colombo G and M. Rekkas, Design of experiments (DoE) in pharmaceutical development, Drug Dev. Ind. Pharm., 2017, 43(6), 889–901, DOI:10.1080/03639045.2017.1291672.
- A. I. Khuri and S. Mukhopadhyay, Response surface methodology, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2010, 2(2), 128–149, DOI:10.1002/wics.73.
- Z. Whiteley, H. M. K. Ho and Y. X. Gan, et al., Microfluidic synthesis of protein-loaded nanogels in a coaxial flow reactor using a design of experiments approach, Nanoscale Adv., 2021, 3(7), 2039–2055, 10.1039/D0NA01051K.
- R. C. Gupta, R. Lall, A. Srivastava and A. Sinha, Hyaluronic Acid: Molecular Mechanisms and Therapeutic Trajectory, Front. Vet. Sci., 2019, 6, DOI:10.3389/fvets.2019.00192.
- N. M. Salwowska, K. A. Bebenek, D. A. Żądło and D. L. Wcisło-Dziadecka, Physiochemical properties and application of hyaluronic acid: a systematic review, J. Cosmet. Dermatol., 2016, 15(4), 520–526, DOI:10.1111/jocd.12237.
- J. Cai, J. Fu, R. Li, F. Zhang, G. Ling and P. Zhang, A potential carrier for anti-tumor targeted delivery-hyaluronic acid nanoparticles, Carbohydr. Polym., 2019, 208, 356–364, DOI:10.1016/j.carbpol.2018.12.074.
- M. C. de Paula, S. G. Carvalho, A. L. P. Silvestre, A. M. dos Santos, A. B. Meneguin and M. Chorilli, The role of hyaluronic acid in the design and functionalization of nanoparticles for the treatment of colorectal cancer, Carbohydr. Polym., 2023, 320, 121257, DOI:10.1016/j.carbpol.2023.121257.
- N. G. Kotla, I. L. Mohd Isa and A. Larrañaga, et al., Hyaluronic Acid-Based Bioconjugate Systems, Scaffolds, and Their Therapeutic Potential, Adv. Healthcare Mater., 2023, 12(20), 2203104, DOI:10.1002/adhm.202203104.
- C. Bednarek, U. Schepers, F. Thomas and S. Bräse, Bioconjugation in Materials Science, Adv. Funct. Mater., 2024, 34(20), 2303613, DOI:10.1002/adfm.202303613.
- F. M. Veronese and M. Morpurgo, Bioconjugation in pharmaceutical chemistry, Il Farm., 1999, 54(8), 497–516, DOI:10.1016/S0014-827X(99)00066-X.
- K. Werengowska-Ciećwierz, M. Wiśniewski, A. P. Terzyk and S. Furmaniak, The Chemistry of Bioconjugation in Nanoparticles-Based Drug Delivery System, Adv. Condens Matter Phys., 2015, 2015(1), 198175, DOI:10.1155/2015/198175.
- J. Casper, S. H. Schenk, E. Parhizkar, P. Detampel, A. Dehshahri and J. Huwyler, Polyethylenimine (PEI) in gene therapy: Current status and clinical applications, J. Controlled Release, 2023, 362, 667–691, DOI:10.1016/j.jconrel.2023.09.001.
- A. P. Pandey and K. K. Sawant, Polyethylenimine: A versatile, multifunctional non-viral vector for nucleic acid delivery, Mater. Sci. Eng., C, 2016, 68, 904–918, DOI:10.1016/j.msec.2016.07.066.
- R. V. Benjaminsen, M. A. Mattebjerg, J. R. Henriksen, S. M. Moghimi and T. L. Andresen, The Possible “Proton Sponge” Effect of Polyethylenimine (PEI) Does Not Include Change in Lysosomal pH, Mol. Ther., 2013, 21(1), 149–157, DOI:10.1038/mt.2012.185.
- C. E. Gallops, C. Yu, J. D. Ziebarth and Y. Wang, Effect of the Protonation Level and Ionic Strength on the Structure of Linear Polyethyleneimine, ACS Omega, 2019, 4(4), 7255–7264, DOI:10.1021/acsomega.9b00066.
- A. Beyerle, M. Irmler, J. Beckers, T. Kissel and T. Stoeger, Toxicity Pathway Focused Gene Expression Profiling of PEI-Based Polymers for Pulmonary Applications, Mol. Pharm., 2010, 7(3), 727–737, DOI:10.1021/mp900278x.
- C. Brunot, L. Ponsonnet, C. Lagneau, P. Farge, C. Picart and B. Grosgogeat, Cytotoxicity of polyethyleneimine (PEI), precursor base layer of polyelectrolyte multilayer films, Biomaterials, 2007, 28(4), 632–640, DOI:10.1016/j.biomaterials.2006.09.026.
- V. Kafil and Y. Omidi, Cytotoxic Impacts of Linear and Branched Polyethylenimine Nanostructures in A431 Cells, BioImpacts, 2011, 1(1), 23–30, DOI:10.5681/bi.2011.004.
- Y. Qin, T. Guo, Z. Wang and Y. Zhao, The role of iron in doxorubicin-induced cardiotoxicity: recent advances and implication for drug delivery, J. Mater. Chem. B, 2021, 9(24), 4793–4803, 10.1039/D1TB00551K.
- A. Bisht, D. Avinash, K. K. Sahu, P. Patel, G. Das Gupta and B. D. Kurmi, A comprehensive review on doxorubicin: mechanisms, toxicity, clinical trials, combination therapies and nanoformulations in breast cancer, Drug Delivery Transl. Res., 2025, 15(1), 102–133, DOI:10.1007/s13346-024-01648-0.
- S. M. Giannitelli, E. Limiti and P. Mozetic, et al., Droplet-based microfluidic synthesis of nanogels for controlled drug delivery: tailoring nanomaterial properties via pneumatically actuated flow-focusing junction, Nanoscale, 2022, 14(31), 11415–11428, 10.1039/D2NR00827K.
- E. Limiti, P. Mozetic and S. M. Giannitelli, et al., Hyaluronic Acid–Polyethyleneimine Nanogels for Controlled Drug Delivery in Cancer Treatment, ACS Appl. Nano Mater., 2022, 5(4), 5544–5557, DOI:10.1021/acsanm.2c00524.
- M. Gori, S. M. Giannitelli and M. Torre, et al., Biofabrication of Hepatic Constructs by 3D Bioprinting of aCell-Laden Thermogel: An Effective Tool to AssessDrug-Induced Hepatotoxic Response, Adv. Healthcare Mater., 2020, 9(21), 2001163, DOI:10.1002/adhm.202001163.
- A. Bucciarelli, S. Chiera, A. Quaranta, V. K. Yadavalli, A. Motta and D. Maniglio, A Thermal-Reflow-Based Low-Temperature, High-Pressure Sintering of Lyophilized Silk Fibroin for the Fast Fabrication of Biosubstrates, Adv. Funct. Mater., 2019, 29(42), 1901134, DOI:10.1002/adfm.201901134.
- A. Gaiardo, D. Novel and E. Scattolo, et al., Optimization of a Low-Power Chemoresistive Gas Sensor: Predictive Thermal Modelling and Mechanical Failure Analysis, Sensors, 2021, 21(3), 783, DOI:10.3390/s21030783.
- J. K. W. Lam, Y. Xu, A. Worsley and I. C. K. Wong, Oral transmucosal drug delivery for pediatric use, Adv. Drug Delivery Rev., 2014, 73, 50–62, DOI:10.1016/j.addr.2013.08.011.
- A. Bucciarelli, A. Adami, C. R. Chandaiahgari and L. Lorenzelli Multivariable optimization of inkjet printing process of Ag nanoparticle ink on Kapton. In: 2020 IEEE International Conference on Flexible and Printable Sensors and Systems (FLEPS). IEEE; 2020:1-4 DOI:10.1109/FLEPS49123.2020.9239474.
- A. M. Bossi, A. Bucciarelli and D. Maniglio, Molecularly Imprinted Silk Fibroin Nanoparticles, ACS Appl. Mater. Interfaces, 2021, 13(27), 31431–31439, DOI:10.1021/acsami.1c05405.
- Team RC. R: A Language and Environment for Statistical Computing. Vienna Austria. Published online 2019.
- A. Bucciarelli, C. Reddy Chandraiahgari, A. Adami, V. Mulloni and L. Lorenzelli, Precise dot inkjet printing thought multifactorial statistical optimization of the piezoelectric actuator waveform, Flex Print Electron., 2020, 5(4), 045002, DOI:10.1088/2058-8585/abbb7e.
- A. Bucciarelli, T. Muthukumar and J. S. Kim, et al., Preparation and Statistical Characterization of Tunable Porous Sponge Scaffolds using UV Cross-linking of Methacrylate-Modified Silk Fibroin, ACS Biomater. Sci. Eng., 2019, 5(12), 6374–6388, DOI:10.1021/acsbiomaterials.9b00814.
- A. Bucciarelli, A. Adami, C. R. Chandaiahgari and L. Lorenzelli Multivariable optimization of inkjet printing process of Ag nanoparticle ink on Kapton. In: 2020 IEEE International Conference on Flexible and Printable Sensors and Systems (FLEPS). IEEE; 2020:1-4 DOI:10.1109/FLEPS49123.2020.9239474.
- Y. Zambito, E. Pedreschi and G. Di Colo, Is dialysis a reliable method for studying drug release from nanoparticulate systems? —A case study, Int. J. Pharm., 2012, 434(1–2), 28–34, DOI:10.1016/j.ijpharm.2012.05.020.
- S. K. Paswan and T. R. Saini, Comparative Evaluation of In Vitro Drug Release Methods Employed for Nanoparticle Drug Release Studies, Dissolution Technol., 2021, 28(4), 30–38, DOI:10.14227/DT280421P30.
- S. J. Wallace, J. Li, R. L. Nation and B. J. Boyd, Drug release from nanomedicines: selection of appropriate encapsulation and release methodology, Drug Delivery Transl. Res., 2012, 2(4), 284–292, DOI:10.1007/s13346-012-0064-4.
- M. Yu, W. Yuan, D. Li, A. Schwendeman and S. P. Schwendeman, Predicting drug release kinetics from nanocarriers inside dialysis bags, J. Controlled Release, 2019, 315, 23–30, DOI:10.1016/j.jconrel.2019.09.016.
- D. Curry, H. Scheller and M. Lu, et al., Prevention of doxorubicin sorptive losses in drug delivery studies using polyethylene glycol, RSC Adv., 2015, 5(33), 25693–25698, 10.1039/C5RA01799H.
- D. Missirlis, R. Kawamura, N. Tirelli and J. A. Hubbell, Doxorubicin encapsulation and diffusional release from stable, polymeric, hydrogel nanoparticles, Eur. J. Pharm. Sci., 2006, 29(2), 120–129, DOI:10.1016/j.ejps.2006.06.003.
- B. Manocha and A. Margaritis, Controlled Release of Doxorubicin from Doxorubicin/γ -Polyglutamic Acid Ionic Complex, J. Nanomater., 2010, 2010, 1–9, DOI:10.1155/2010/780171.
- D. C. Wu and C. M. Ofner, Adsorption and Degradation of Doxorubicin from Aqueous Solution in Polypropylene Containers, AAPS PharmSciTech, 2013, 14(1), 74–77, DOI:10.1208/s12249-012-9885-1.
- G. Jiang, K. Park and J. Kim, et al., Hyaluronic acid–polyethyleneimine conjugate for target specific intracellular delivery of siRNA, Biopolymers, 2008, 89(7), 635–642, DOI:10.1002/bip.20978.
- S. Zhao, A. Riaud, G. Luo, Y. Jin and Y. Cheng, Simulation of liquid mixing inside micro-droplets by a lattice Boltzmann method, Chem. Eng. Sci., 2015, 131, 118–128, DOI:10.1016/j.ces.2015.03.066.
- M. Arruebo, L. Uson, M. Miana, I. Ortiz De Solorzano, V. Sebastian and A. Larrea, Continuous synthesis of drug-loaded nanoparticles using microchannel emulsification and numerical modeling: effect of passive mixing, Int. J. Nanomed., 2016, 11, 3397–3416, DOI:10.2147/ijn.s108812.
- F. Sarrazin, K. Loubière, L. Prat, C. Gourdon, T. Bonometti and J. Magnaudet, Experimental and numerical study of droplets hydrodynamics in microchannels, AIChE J., 2006, 52(12), 4061–4070, DOI:10.1002/aic.11033.
- A. Tiribocchi, A. Montessori and M. Lauricella, et al., The vortex-driven dynamics of droplets within droplets, Nat. Commun., 2021, 12(1), 82, DOI:10.1038/s41467-020-20364-0.
- H. Song, M. R. Bringer, J. D. Tice, C. J. Gerdts and R. F. Ismagilov, Experimental test of scaling of mixing by chaotic advection in droplets moving through microfluidic channels, Appl. Phys. Lett., 2003, 83(22), 4664–4666, DOI:10.1063/1.1630378.
- O. Kašpar, A. H. Koyuncu, A. Hubatová-Vacková, M. Balouch and V. Tokárová, Influence of channel height on mixing efficiency and synthesis of iron oxide nanoparticles using droplet-based microfluidics, RSC Adv., 2020, 10(26), 15179–15189, 10.1039/D0RA02470H.
- L. Parhamifar, A. K. Larsen, A. C. Hunter, T. L. Andresen and S. M. Moghimi, Polycation cytotoxicity: a delicate matter for nucleic acid therapy—focus on polyethylenimine, Soft Matter, 2010, 6(17), 4001, 10.1039/c000190b.
- A. P. Pandey and K. K. Sawant, Polyethylenimine: A versatile, multifunctional non-viral vector for nucleic acid delivery, Mater. Sci. Eng., C, 2016, 68, 904–918, DOI:10.1016/j.msec.2016.07.066.
- B. Fortuni, T. Inose and M. Ricci, et al., Polymeric Engineering of Nanoparticles for Highly Efficient Multifunctional Drug Delivery Systems, Sci. Rep., 2019, 9(1), 2666, DOI:10.1038/s41598-019-39107-3.
- X. Du, Y. Gao, Q. Kang and J. Xing, Design and Applications of Tumor Microenvironment-Responsive Nanogels as Drug Carriers, Front. Bioeng. Biotechnol., 2021, 9, DOI:10.3389/fbioe.2021.771851.
- Y. Wei, L. Quan, C. Zhou and Q. Zhan, Factors Relating to the Biodistribution & Clearance of Nanoparticles & Their Effects on In Vivo Application, Nanomedicine, 2018, 13(12), 1495–1512, DOI:10.2217/nnm-2018-0040.
- F. Zhao, J. Wang and Y. Zhang, et al., In vivo Fate of Targeted Drug Delivery Carriers, Int. J. Nanomed., 2024, 19, 6895–6929, DOI:10.2147/ijn.s465959.
Footnote |
| † Authors equally contributed to this work. |
| This journal is © The Royal Society of Chemistry 2026 |