Zinc ions and zinc-embedded carbon quantum dots as competitive inhibitors of fumarase: preferential inhibition of the reverse reaction
Amene
Navaser
a and
Hamid R.
Kalhor
 *ab
*ab
aBiochemistry and Chemical Biology Research Laboratory, Chemistry Department, Sharif University of Technology, Tehran, Iran. E-mail: kalhor@sharif.edu; Fax: +982166165315; Tel: +982166165315
bInstitute for Convergence Science & Technology, Sharif University of Technology, Tehran, Iran
First published on 4th February 2026
Abstract
Targeted modulation of enzyme activity offers a promising strategy for both elucidating catalytic mechanisms and developing novel therapeutics. In this study Zn2+ ions were introduced as an effective competitive inhibitor of fumarase, a pivotal enzyme in the citric acid cycle. Zn2+ binding significantly alters the Michaelis constant (Km) for both L-malate and fumarate, with a pronounced preference for inhibiting the reverse reaction (L-malate to fumarate), a direction relevant to redox homeostasis and anaplerotic flux. A major limitation of the clinical application of many metal-based inhibitors is their poor water solubility. To overcome this challenge and introduce a new class of enzyme inhibitors, zinc-modified carbon quantum dots (Zn–CQDs) were synthesized. Owing to their polar surface, Zn–CQDs interact more effectively with the enzyme, which increases the local concentration of Zn2+ ions at the active site. As a result, these nanomaterials exhibit enhanced water solubility and significantly greater inhibitory potency compared to free Zn2+ ions. Biophysical and kinetic analyses confirmed the competitive inhibition mechanism and demonstrated that Zn–CQDs interact with the enzyme without perturbing its secondary structure. Notably, both Zn2+ ions and Zn–CQDs preferentially inhibited the reverse reaction of fumarase, offering precise control over fumarase activity. Molecular docking and MD simulations elucidated the plausible binding site of Zn2+ within the active site. It was found that Zn2+ interacts with Glu340, a residue previously shown to be involved in binding fumarase inhibitors. These findings establish Zn–CQDs as a novel class of water-soluble fumarase inhibitors, distinguished by their facile synthesis, tunable solubility, and selective inhibition profile. This work highlights the potential of zinc-based nanomaterials in enzyme regulation, offering a powerful alternative to existing inhibitors and developing targeted redox-sensitive therapeutic strategies.
Introduction
Despite infinitesimal RNA catalysis in living organisms, almost all biochemical reactions are catalyzed by enzymes. They facilitate the biochemical processes with a remarkable degree of selectivity as well as specificity.1,2 Over the years, it has been evident that the enzymes have involved in many metabolic disorders and treatments. Hence, the regulation of enzyme activities gives rise to precise control over biochemical pathways.3–5 Enzyme regulation is therefore crucial in modulating enzyme function to maintain metabolic balance and to tame metabolic pathways so that the desired outcomes can be achieved. Inhibitors, for example, can harness enzyme activity by binding to an active site or another regulatory region within the enzyme, thereby altering the enzyme activity through competitive,6 non-competitive,7 and uncompetitive8 inhibition. Among the various strategies reported so far, the use of an inhibitor is a promising means of regulating enzyme activity, which results in several clinical treatments.5 For example, zileuton, a 5-lipoxygenase inhibitor, is an FDA approved organic-based drug for the treatment of chronic asthma.9 As a nucleoside reverse transcriptase inhibitor, zidovudine is another FDA approved organic medication in the prophylaxis of HIV-1.10Fumarate hydratase (fumarase) is known as a homotetrameric hydratase enzyme, which is responsible for the reversible hydration of fumarate to L-malate. Fumarase mainly exists in two cytosolic and mitochondrial isoforms and is expressed in most tissues.11 The mitochondrial fumarase is involved in the citric acid cycle (TCA) and energy production.12 However, apart from its traditional function, the cytosolic fumarase possesses a direct role in DNA damage response and the urea cycle. The enzyme is also indirectly involved in the catabolism of some amino acids.13,14 The dual localization, cytosolic and mitochondrial isoforms, enables the enzyme to function in metabolic pathways in both environments.15 Studies have shown that fumarase is linked to several biological conditions, including fumarate hydratase deficiency16 and certain types of cancer17,18e.g. renal cancer,19 making it an interesting target for the therapeutic approach. Fumarase inactivation presents several potential benefits, particularly in the context of cancer therapy and metabolic regulation. As a case in point, the fumarase inhibition disturbs cellular redox balance, thereby promoting the cell dependency on glycolysis for energy production. Hence, it may have therapeutic potential by enhancing the sensitivity of cancer cells to therapies targeting oxidative stress and metabolic pathways.20 Furthermore, the inactivation of fumarase has been shown to create a synthetic lethality with ferroptosis induction, a form of cell death that can selectively target fumarase-deficient cancer cells, such as those found in hereditary leiomyomatosis and renal cell cancer.21 Therefore, introducing inhibitors for fumarase is driven by the need for a better understanding of its biological function, thereby developing potential treatments for associated disorders. Beyond its implications in tumor biology, fumarase inhibition is also gaining attention as a novel strategy for antimicrobial intervention. Given its essential role in bacterial central metabolism, particularly within the TCA cycle, targeting fumarase can critically impair microbial energy production and redox balance. This metabolic disruption compromises bacterial viability, positioning fumarase as a promising target for antibacterial development. Unlike conventional antibiotics that typically target cell wall synthesis or DNA replication, fumarase inhibition offers a metabolism-based approach to suppress bacterial growth. This strategy may prove especially valuable in the era of rising antibiotic resistance, providing a novel avenue for therapeutic innovation.22 Therefore, the development of fumarase inhibitors not only enables mechanistic insights into its diverse cellular functions but also holds translational potential across oncology and infectious disease therapeutics. Beyond therapeutic applications, fumarase inhibition may also offer strategic advantages in whole cell or cell free extract biocatalytic systems, particularly in organic syntheses where fumarate and L-malate serve as substrates. By selectively downregulating endogenous fumarase activity, these pathways can be redirected or preserved, minimizing undesired metabolic turnover and improving precursor availability or product yields.23 Up to now, several inhibitors have been reported for the effective inhibition of fumarase. Despite those advances, the reported inhibitors are mostly insoluble in water and this, in turn, prevents their further in vivo studies. Additionally, those inhibitors are generally complex organic molecules and therefore challenging and costly synthetic compounds (Scheme 1).20,24–27 The outlined challenges can hinder reproducibility across studies and limit the inhibitors’ scalability for broader research and clinical use. As such, the introduction of a biocompatible inhibitor that works effectively and overcomes the ever-present limitations is highly in demand.
 | ||
| Scheme 1 Structures of fumarase inhibitors. (a) Previously introduced inhibitors. (b) Inhibitor from the present study. | ||
Metal ions have specific chemical and physical properties that make them highly effective for enzyme regulation.28–30 Unlike organic regulators, which often rely on complex molecular structures and specific functional groups to bind to enzyme active sites, metal ions can form stable complexes with enzymes through their ability to act as acceptors.31 This, in turn, creates a three-dimensional architecture that authorizes excellent fitting into the active site.32 Moreover, interaction with various sites of enzymes allows the metal ions to exert different types of inhibition by altering the enzyme's conformations.33 Additionally, earth-abundant metals are relatively inexpensive and highly accessible, making them a practical choice for research and therapeutic applications.34 Many studies showed that the metal ions can act as effective inhibitors. Studies revealed that various metal ions, including Co2+, Zn2+, Ca2+, Fe2+, Mn2+, Cr3+, Sn2+, and Mg2+, significantly affect the enzymatic activity of glutathione reductase in the liver of rainbow trout. The inhibitory effects varied among the metals, with cobalt being identified as the most potent inhibitor.35 Zinc ions inhibit the activity of horseradish peroxidase through a non-competitive mechanism; therefore, zinc ions do not compete with the substrate for binding to the active site and prefer to bind to the enzyme–substrate complex.36 In another study, it was suggested that zinc cooperates with tumor protein p53, and as a result, the mitochondrial aconitase activity could be inhibited by the increased reactive oxygen species concentration in the reaction media.37 It was also reported that Bi3+ could serve as a non-competitive inhibitor for Helicobacter pylori fumarase by lowering the Vmax and Km from 3.85 min−1 and 15.3 mM to 0.56 min−1 and 10.29 mM, respectively. Despite the efficiency of the Bi(NO3)3 as a non-competitive inhibitor for fumarase,38 its poor water-solubility hinders the further in vivo application of the inhibitor.39 Additionally, due largely to the accumulation of bismuth in tissues, which leads to potential side effects such as neurotoxicity and kidney damage, Bi-based compounds often face with human health concerns.40 Considering all the above-mentioned limitations, the exploration of alternative metal ions that may offer more stable, safer, and effective inhibition of fumarase activity is highly in demand. Recently, it was reported that anionic water-soluble zinc porphyrins affected the fumarase activity and inhibited the fumarate production. While both zinc-complexed cationic porphyrin and metal-free cationic porphyrins had no effect on the fumarase activity, the metal-free anionic porphyrins simply decreased the fumarate production.41 The aforementioned study has investigated the inhibitory effects of porphyrins on fumarase, focusing on the activity of both zinc-bound porphyrins and metal-free porphyrins. Nevertheless, despite the advances, porphyrin ligands are expensive; furthermore, the authors missed the effect of zinc ions on the inhibition and its mechanistic details and kinetics upon the inhibition process. Addressing the significant gaps by systematically studying the impact of zinc ions on the inhibition of fumarase by porphyrins, along with a detailed kinetic analysis of the inhibition, is still in demand.
Poor water solubility, which is often accompanied by increased toxicity, has largely been connected with low success of certain introduced therapeutic compounds in their progress through clinical trials.33,42 One effective strategy for making an insoluble metal compound water-soluble is modifying carbon quantum dots with the metal, since carbon quantum dots (CQDs) enjoy high and tunable water-solubility.43,44 They are considered as permeable to cells, biocompatible, and non-toxic carbon-based nanomaterials with high physicochemical stability.45,46 There are various straightforward methods for (mass) production and functionalization of CQDs.47 Thus, CQDs have found application in many areas,48 bioimaging,49 biosensors,50 drug delivery,51 and catalysis,52,53 to name some. CQDs possess a large surface area enriched with polar groups and a hydrophobic core, making well-suited for enzymatic applications. Consequently, they have been widely employed in enzymatic systems like fluorometric assays to monitor enzyme activity.54 In an enzyme inhibition screening system, for instance, a dopamine-functionalized CQD has enabled sensitive monitoring of the tyrosinase activity in the inhibitor screening system. The assay utilizes fluorescence quenching upon tyrosinase catalyzed oxidation of dopamine.55 CQD and gold nanoparticles have also been exploited to monitor acetylcholinesterase inhibition and reactivation by fluorescence quenching/recovery of the CQD in the presence of dispersed/aggregated gold nanoparticles.56 Though interesting sensing behavior of enzyme activities was achieved by CQDs, they have rarely been investigated as inhibitors of the enzymes.
Motivated by the critical role of fumarase in regulating biochemical pathways and its therapeutic relevance in cancer, we aimed to modulate the activity of Saccharomyces cerevisiae fumarase using simple, cost-effective metal cations. Among various salts tested, ZnCl2 emerged as a potent competitive inhibitor, significantly increasing the enzyme's Km. Although ZnCl2 is intrinsically highly soluble, under the near-neutral aqueous conditions required for enzymatic assays, it undergoes slightly hydrolysis and precipitation, limiting its practical applicability and reproducibility.57 To address this, we developed zinc-modified CQDs (Zn–CQDs), which provide more stable aqueous dispersions of Zn2+ ions. Zn–CQDs outperformed ZnCl2, achieving a remarkable increase in Km for fumarate and L-malate by 7.38-fold and 147.07-fold, respectively. Both zinc ions and Zn–CQDs inhibit the reverse reaction more strongly than the forward reaction. These findings position Zn–CQDs as a promising and selective tool for fumarase modulation, with potential implications for manipulating TCA cycle flux and developing therapeutic strategies against cancer-associated metabolic reprogramming.
Results and discussion
Zinc ion inhibits fumarase activity
At the outset of our study, the effect of a series of cations on regulation of the fumarase activity was investigated. The L-malate to fumarate conversion was selected as the model conversion. As shown in Fig. S10, Mn2+, Co2+, Mg2+, and Ca2+ did not affect the fumarase activity. However, it was observed that zinc chloride could inhibit the fumarase activity effectively. As shown in Fig. 1, Zn2+ inhibited the L-malate to fumarate conversion to a great extent. To further confirm the inhibitory role of Zn2+, the experiments were repeated in the presence of ethylenediaminetetraacetic acid (EDTA) as a chelating agent of the zinc cation. As expected, in the presence of EDTA no inhibition was observed for both conversions (Fig. 2). Due to the chelating nature of EDTA, it could form a complex with zinc ions and prevent the cation from binding to the enzyme; therefore, fumarase proceeds with its function without any distributing effect of the cation (Fig. 2). The distinct effects of metal ions on enzyme activity can be attributed to their physicochemical properties, including ionic radius, charge density, and coordination geometry. These factors determine how efficiently a metal ion binds to the active site and whether it supports or disrupts activity.58 Zn2+ possesses a ligand-field stabilization energy of zero making it more flexible to readily adopt diverse coordination geometries, unlike other metals.59Zinc-modified CQD enhanced fumarase inhibition
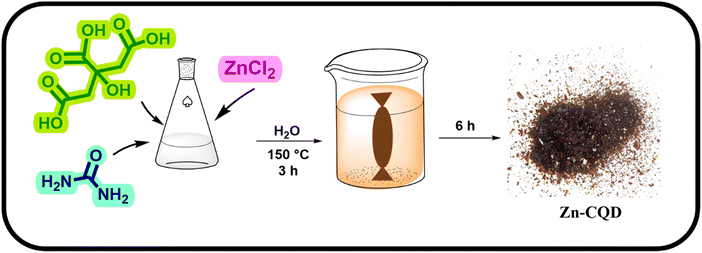
Subsequently, the impact of the ZnCl2 concentration on the regulation of the fumarase activity was investigated. To that end, the inhibitory effect of ZnCl2 was examined at various concentrations ranging from nanomolar to millimolar. At nanomolar concentrations no significant inhibition was observed; yet, within the micromolar concentrations, ZnCl2 demonstrated a noticeable level of inhibition on fumarase activity (Fig. 1). While a clear solution of ZnCl2 was obtained at both nanomolar and micromolar concentrations, at millimolar concentrations, due to the formation of basic zinc chloride, a turbid solution of zinc oxychloride was formed.57,60 Therefore, investigation of the higher concentration of Zn2+ for further improving the fumarase inhibition was found to be a challenge. To overcome this limitation, we hypothesized that modifying CQDs with ZnCl2 (Zn–CQD) could not only yield water-soluble Zn–CQDs but also enhance the inhibitory effect of zinc, even at low ZnCl2 content. The large surface area and high density of polar groups on the CQD surface facilitate significant enzyme–CQD interactions, resulting in stronger binding and an increased local Zn2+ concentration at the protein–CQD interface, thereby making Zn2+ more readily available near the binding site.52,61,62 A mild and green hydrothermal method was used to generate the intended Zn–CQDs (Scheme 2). Zinc chloride along with citric acid, the carbon source of the CQD for shaping the core, and urea were the initial precursors for constructing Zn–CQD. The incorporation of urea in the synthesis of Zn–CQDs was mostly to augment the water-solubility of the synthesized Zn–CQD.In light of the initial content of ZnCl2 used in the composition of various Zn–CQDs, three nanomaterials named Zn(0.5)–CQDs, Zn(1)–CQDs, and Zn(2)–CQDs were synthesized. In doing so, a 0.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio, 1
1 molar ratio, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio and a 2
1 molar ratio and a 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio of ZnCl2 to citric acid were used to construct the Zn(0.5)–CQDs, Zn(1)–CQDs, and Zn(2)–CQDs, respectively (for details, see the SI). Initially, both Zn(0.5)–CQDs and Zn(1)–CQDs showed excellent water-solubility. This enhanced water-solubility of zinc in the context of the synthesized nanomaterials could provide effective application of such zinc complexes in aqueous enzyme inhibition studies, where their inhibitory effects on fumarase could be explored. In contrast, Zn(2)–CQDs formed aggregated particles with poor dispersibility and significantly reduced water solubility; therefore, Zn(2)–CQDs were excluded from further investigations. Accordingly, the primary challenge of exploring Zn2+ as an inhibitor was overcome. As for inhibitory effect of the zinc in Zn–CQD compositions, the L-malate to fumarate conversion was studied in the presence of Zn–CQDs (Fig. 3). As shown in Fig. 3, in comparison to cationic zinc, both Zn(0.5)–CQDs and Zn(1)–CQDs exhibited enhanced inhibition for fumarase activity. Additionally, both Zn(0.5)–CQDs and Zn(1)–CQDs inhibited the enzyme function up to 53.3% and 43.3% with respect to the free enzyme, respectively. It can be concluded that the incorporation of Zn2+ within the CQD matrix enhances its inhibitory potency, most likely due to the hydrogen bond rich surface chemistry of the Zn–CQDs, which results in a stronger enzyme–inhibitor interaction. Furthermore, Zn(1)–CQDs displayed a superior inhibitory effect than Zn(0.5)–CQDs, as a result, Zn(1)–CQDs werr selected as the optimized inhibitor. To investigate the effect of CQD composition on the fumarase inhibition, a N-doped CQD with a 1
1 molar ratio of ZnCl2 to citric acid were used to construct the Zn(0.5)–CQDs, Zn(1)–CQDs, and Zn(2)–CQDs, respectively (for details, see the SI). Initially, both Zn(0.5)–CQDs and Zn(1)–CQDs showed excellent water-solubility. This enhanced water-solubility of zinc in the context of the synthesized nanomaterials could provide effective application of such zinc complexes in aqueous enzyme inhibition studies, where their inhibitory effects on fumarase could be explored. In contrast, Zn(2)–CQDs formed aggregated particles with poor dispersibility and significantly reduced water solubility; therefore, Zn(2)–CQDs were excluded from further investigations. Accordingly, the primary challenge of exploring Zn2+ as an inhibitor was overcome. As for inhibitory effect of the zinc in Zn–CQD compositions, the L-malate to fumarate conversion was studied in the presence of Zn–CQDs (Fig. 3). As shown in Fig. 3, in comparison to cationic zinc, both Zn(0.5)–CQDs and Zn(1)–CQDs exhibited enhanced inhibition for fumarase activity. Additionally, both Zn(0.5)–CQDs and Zn(1)–CQDs inhibited the enzyme function up to 53.3% and 43.3% with respect to the free enzyme, respectively. It can be concluded that the incorporation of Zn2+ within the CQD matrix enhances its inhibitory potency, most likely due to the hydrogen bond rich surface chemistry of the Zn–CQDs, which results in a stronger enzyme–inhibitor interaction. Furthermore, Zn(1)–CQDs displayed a superior inhibitory effect than Zn(0.5)–CQDs, as a result, Zn(1)–CQDs werr selected as the optimized inhibitor. To investigate the effect of CQD composition on the fumarase inhibition, a N-doped CQD with a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio of citric acid to urea and without the incorporation of ZnCl2 was synthesized (for details, see the SI). As a result, the N-doped CQD showed no effect on the fumarase activity, evidencing critical role of the Zn embedding in the inhibition process (Fig. 3). Additionally, the water-solubility of Zn(1)–CQD was quantified. As such, its solubility was found to be 180 mg mL−1 (for details, see the SI).
1 molar ratio of citric acid to urea and without the incorporation of ZnCl2 was synthesized (for details, see the SI). As a result, the N-doped CQD showed no effect on the fumarase activity, evidencing critical role of the Zn embedding in the inhibition process (Fig. 3). Additionally, the water-solubility of Zn(1)–CQD was quantified. As such, its solubility was found to be 180 mg mL−1 (for details, see the SI).
Characterization of Zn(1)–CQD
The optical properties of synthesized Zn(1)–CQDs were obtained from the ultraviolet and visible (UV-vis) absorption and photoluminescence (PL) experiments. The UV-vis absorption spectrum (Fig. 4a) exhibited two distinct peaks around 240 and 340 nm. The peak around 240 was assigned to the π–π* transition of the sp2 carbons in the core of the Zn(1)–CQDs, and that around 340 nm resulted from the transition of n–π* of the amino, hydroxyl, and carboxyl groups associated with the surface of the Zn(1)–CQDs; these peaks suggest that the synthesized Zn(1)–CQDs are functionalized and the surface of the Zn(1)–CQDs perchance play a role in the interacting with the enzyme. A detailed PL spectroscopy study was conducted on the Zn(1)–CQDs with regular excitation intervals ranging from 240 nm to 420 nm and the results are shown in Fig. 4b. The emission intensity displayed a clear excitation-dependent behavior; as the excitation wavelengths increased, the emission increased, reaching a maximum at 340 nm, beyond this wavelength the emission decreased gradually with further increases in excitation wavelength. This trend is typical of carbon quantum dots and is attributed to multiple surface states, arising from diverse functional groups and size-distribution effects, which lead to varied electronic transitions and the radiative recombination pathway.63 The emission maxima were consistently located in the blue region, indicating that the surface functional groups play a central role in stabilizing emissive states and enhancing luminescence. The strong blue emission observed at 340 nm excitation corresponds well with the optical image inset in Fig. 4b, where the Zn(1)–CQD solution exhibits intense blue fluorescence under UV illumination, in contrast to its transparent to brownish appearance under visible light. To gain insights into the surface chemistry of the nanoparticles, the Fourier-transform infrared (FT-IR) spectra were studied in detail. Following Zn embedding into the N-doped CQDs, the obtained Zn(1)–CQDs demonstrated almost the same IR spectral pattern as N-doped CQDs. However a distinct peak at 631 cm−1 attributed to the stretching vibration of the Zn–O bond confirmed the proper incorporation of Zn in the CQD composition.64,65 The peak at 1731 cm−1 corresponds to the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching vibration. A broad peak at 3337 cm−1 is indicative of the stretching vibration of the N–H and O–H bonds, highlighting the high water solubility of Zn(1)–CQDs. The urea incorporation into the composition of Zn(1)–CQDs was demonstrated by the peak at 1430 cm−1 assigned to C–N stretching vibration. Morphological properties of the synthesized Zn(1)–CQD was studied through X-ray diffraction (XRD) analysis. As demonstrated in Fig. 4d, a broad peak centered on 2θ = 20° indicates the spherical morphology of the nanoparticles, which is considered as a distinguishing characteristic for CQD-based nanoparticles. Additionally, peak broadening around 35° could be attributed to the uniform distribution of ZnCl2 in Zn(1)–CQD composition, which leads to disappearance of the typical XRD peaks of zinc salts. Transmission electron microscopy (TEM) was employed to investigate the morphology and size distribution of the Zn(1)–CQDs. The TEM image (Fig. 5) reveals that the Zn(1)–CQDs exhibit a nearly spherical morphology. The inset particle size distribution histogram confirms that the Zn(1)–CQDs possess an average diameter of approximately 5 nm, consistent with the expected CQDs’ nanoscale range. The Zn content in Zn(1)–CQDs was determined by inductively coupled plasma optical emission spectroscopy (ICP-OES). The analysis confirmed that Zn constitutes 22.6 wt% of the Zn(1)–CQDs, and this value was used to calculate the actual Zn2+ concentration applied in the inhibition assays.
O stretching vibration. A broad peak at 3337 cm−1 is indicative of the stretching vibration of the N–H and O–H bonds, highlighting the high water solubility of Zn(1)–CQDs. The urea incorporation into the composition of Zn(1)–CQDs was demonstrated by the peak at 1430 cm−1 assigned to C–N stretching vibration. Morphological properties of the synthesized Zn(1)–CQD was studied through X-ray diffraction (XRD) analysis. As demonstrated in Fig. 4d, a broad peak centered on 2θ = 20° indicates the spherical morphology of the nanoparticles, which is considered as a distinguishing characteristic for CQD-based nanoparticles. Additionally, peak broadening around 35° could be attributed to the uniform distribution of ZnCl2 in Zn(1)–CQD composition, which leads to disappearance of the typical XRD peaks of zinc salts. Transmission electron microscopy (TEM) was employed to investigate the morphology and size distribution of the Zn(1)–CQDs. The TEM image (Fig. 5) reveals that the Zn(1)–CQDs exhibit a nearly spherical morphology. The inset particle size distribution histogram confirms that the Zn(1)–CQDs possess an average diameter of approximately 5 nm, consistent with the expected CQDs’ nanoscale range. The Zn content in Zn(1)–CQDs was determined by inductively coupled plasma optical emission spectroscopy (ICP-OES). The analysis confirmed that Zn constitutes 22.6 wt% of the Zn(1)–CQDs, and this value was used to calculate the actual Zn2+ concentration applied in the inhibition assays.
 | ||
| Fig. 5 TEM image of the Zn(1)–CQD, showing a nearly spherical morphology. The inset displays the particle size distribution histogram, with an average particle size of 5 nm. | ||
Further insights into the composition of Zn(1)–CQD was gained by field-emission scanning electron microscopy (FE-SEM) studies (Fig. 6). As such, energy dispersive spectroscopy (EDS) was performed, and the results confirmed the presence of the Zn in the Zn(1)-CQD composition (Fig. 6b). According to EDS analysis, Zn was incorporated onto the CQD with the final concentration of Zn reaching 27.88 wt%. To obtain additional insights into the surface of the Zn(1)–CQDs, EDS elemental mapping analysis was carried out (Fig. 6c), further exhibiting a uniform distribution of carbon, oxygen, nitrogen, and zinc atoms across the surface of the CQDs. As a result, it could be concluded that the Zn incorporating process was effective and did not disrupt the overall structural integrity of the CQDs, with the Zn being well-distributed throughout the nanoparticles. To quantify the surface functionalities of Zn(1)–CQDs, a back titration was carried out and analyzed using a Gran plot (for details, see the SI). The total number of acid/base sites on the CQD surface was determined to be 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 333 µmol g−1 (Fig. 7). This value is significantly higher than those reported for modified CQDs in the literature, indicating a high degree of surface functionalization.66,67 Such enrichment in polar functional groups suggests that Zn(1)–CQDs possess an enhanced capacity for protein interactions, which may contribute to their inhibitory activity.
333 µmol g−1 (Fig. 7). This value is significantly higher than those reported for modified CQDs in the literature, indicating a high degree of surface functionalization.66,67 Such enrichment in polar functional groups suggests that Zn(1)–CQDs possess an enhanced capacity for protein interactions, which may contribute to their inhibitory activity.
 | ||
| Fig. 7 (a) Back titration of Zn(1)–CQD. (b) Gran plot of Zn(1)–CQD based on back titration of Zn(1)–CQD. | ||
Competitive inhibition of fumarase by Zn2+ ions and Zn–CQDs
The kinetics parameters of the fumarase were calculated by measuring the initial reaction velocities (V0) at varying substrate concentrations, both in the presence and the absence of the inhibitors. The Lineweaver–Burk plot was employed to evaluate changes in the apparent Michaelis constant (Km), maximum velocity (Vmax), inhibition constant (Ki) and the mechanism of the inhibition (for details, see the SI). In doing so, different concentrations of Zn(1)–CQDs were initially utilized to optimize the Zn(1)–CQDs concentration (for details, see the SI and Fig. S3). With the optimized concentration in hand, the Lineweaver–Burk plots for fumarate to L-malate and the L-malate to fumarate conversions were obtained (Fig. 8). Accordingly, for the forward reaction (Fig. 8a), the conversion of fumarate to L-malate, the Km value of the enzyme was calculated to be 1.10 ± 0.12 mM, and it was reached 6.72 ± 0.12 (p < 0.001) and 8.12 ± 0.24 (p < 0.01) mM when the Zn2+ ions and Zn(1)–CQDs were respectively served as the inhibitor of the transformation (Table 1). In essence, the Zn(1)–CQDs increased the Km value by 7.38-fold. In addition, the Ki value for fumarate was calculated as 39.14 ± 5.18 µM and 23.50 ± 4.24 µg mL−1 using Zn2+ and Zn(1)–CQDs, respectively. It was found that both Zn ions and Zn(1)–CQDs acted through a competitive inhibition mechanism for the conversion of fumarate to L-malate since an increase in the apparent Km value proceeded without a notable change in the Vmax amount (Fig. 8a). For the reverse reaction, a competitive inhibition mechanism was also detected (Fig. 8b), in the absence of any inhibitor, the Km value of fumarase was found to be 1.58 ± 0.32 mM; however, the presence of the Zn2+ inhibitor increased the Km value up to 131.95 ± 2.35 (p < 0.001) mM. Interestingly, when Zn(1)–CQDs was used as an inhibitor the Km value was further increased up to 221.32 ± 5.75 (p < 0.001) mM, 1.68 times higher than the Zn2+ (p < 0.001) (Table 2). In addition, the Km value of fumarase was increased by 147.07-fold by using Zn(1)–CQDs for L-malate. The Ki value of Zn2+ ions and Zn(1)–CQDs for L-malate was calculated to be 2.42 ± 0.49 µM and 1.08 ± 0.22 µg mL−1 respectively, suggesting that both inhibitors bond effectively to the enzyme. To further validate these results, detailed statistical analyses were conducted, according to the statistical analyses summarized in Tables S1–S3 (for details, see the SI); the inhibition of fumarase by Zn2+ ions and Zn(1)–CQDs follows a competitive inhibition pattern. Consequently, the Vmax and Kcat values remain nearly constant among the free enzyme, ZnCl2, and Zn(1)–CQD-treated groups, resulting in relatively large p-values (greater than 0.1). In contrast, the Km values differ significantly between the groups, indicating that the inhibitors primarily affect the substrate binding affinity rather than the catalytic turnover rate. Intriguingly, both inhibitors exhibited distinct selectivity toward the reverse reaction of fumarase. Such selectivity offers exciting potential for developing selective inhibitors with therapeutic applications. Interestingly, the size of Saccharomyces cerevisiae fumarase is approximately 11.83 × 4 nm (Fig. S8). The dimensions of the enzyme and Zn–CQDs, together with the presence of polar functional groups on the nanoparticle surface, suggest the potential for direct interactions between the two. These interactions are likely mediated by hydrogen bonding, which may destabilize the Zn–CQD coordination and promote the release of Zn2+ ions at the enzyme–nanoparticle interface. The localized enrichment of Zn2+ in close proximity to fumarase would facilitate delivery of the ions to the active site, thereby enhancing inhibitory efficiency. This mechanism provides a rational explanation for the observed enhanced enzyme inhibition by Zn–CQDs. Previous studies have also found that the high levels of Zn2+ lead to a reduced cellular energy production. The underlying mechanism of such observation have been partially elucidated, where the increased production level of reactive oxygen species leads to the loss of mitochondrial membrane potential or energy production.68,69 However, the effect of Zn2+ ions on the fumarase activity as one of the key enzymes involved in the cellular energy production has been not investigated yet. As such, the results of this work highlight the important role of fumarase in the energy production of cells as the results are in agreement with the reducing energy production.| V max (µmol min−1 mg−1) | K m (mM) | K cat (s−1) | K cat/Km | K i | |
|---|---|---|---|---|---|
| Free enzyme | 347.81 ± 1.96 | 1.10 ± 0.12 | 617.02 ± 19.87 | 560.92 | — |
| ZnCl2 | 351.60 ± 3.0 | 6.72 ± 0.12 | 637.51 ± 18.85 | 94.87 | 39.14 ± 5.18 µM |
| Zn(1)–CQDs | 347.54 ± 7.43 | 8.12 ± 0.24 | 610.74 ± 15.65 | 75.21 | 23.50 ± 4.24 µg mL−1 |
| V max (µmol min−1 mg−1) | K m (mM) | K cat (s−1) | K cat/Km | K i | |
|---|---|---|---|---|---|
| Free enzyme | 338.09 ± 7.46 | 1.58 ± 0.32 | 595.74 ± 14.20 | 377.05 | — |
| ZnCl2 | 334.55 ± 7.39 | 131.95 ± 2.35 | 591.82 ± 13.10 | 4.48 | 2.42 ± 0.49 µM |
| Zn(1)–CQDs | 336.18 ± 6.92 | 221.32 ± 5.75 | 596.07 ± 12.28 | 2.69 | 1.08 ± 0.22 µg mL−1 |
Cellular impact of zinc-based inhibitors on fumarase activity
To assess the biological relevance of fumarase inhibition, bacterial growth assay was performed using untreated E. coli BL21 cells, ZnCl2 treated cells and, Zn(1)–CQD treated cells. As shown in Fig. S5, the control cells exhibited a typical exponential growth pattern, reaching an OD600 of 4.4 after 5 hours. In contrast, bacterial cultures treated with the Zn2+ ions and Zn(1)–CQD displayed markedly reduced growth. Notably, the Zn(1)–CQD group exhibited more significant inhibition, suggesting that Zn(1)–CQDs are cell-permeable and exert a more pronounced antibacterial effect in vivo. To further validate the competitive inhibition mechanism, we evaluated cell growth in the presence of increasing fumarate concentrations for control, ZnCl2-treated, and Zn(1)–CQD–treated cells. Increasing fumarate progressively restored growth in the zinc-treated samples, with near-complete recovery at 15 mM (Fig. S5). Although high fumarate levels slightly reduced growth in control cells due to its known antimicrobial activity, the cell growth restoration of Zn2+ and Zn(1)–CQD–induced inhibition strongly supports a competitive interaction between fumarate and zinc at the level of fumarase. To evaluate the inhibitory effect under more native-like conditions, the impact of zinc-based treatments was investigated. Fumarase-overexpressing cell free extracts were prepared from cultures grown under same conditions. Following normalization based on total protein content, fumarase activity was monitored. As shown in Fig. 9, treatment with ZnCl2 and Zn(1)–CQDs significantly reduced fumarase activity compared to the control. Notably, cells treated with Zn(1)–CQDs exhibited the lowest residual activity, suggesting that Zn–CQDs exert a more potent inhibitory effect than free zinc ions in a cellular environment.Minimal structural alterations in fumarase
To investigate structural effects of inhibitor bonding on fumarase, circular dichroism (CD) spectra of free fumarase, fumarase in the presence of Zn2+ ions, and fumarase in the presence of Zn(1)–CQDs were performed (Fig. 10). In the case of free enzyme, two negative peaks around 210 nm and 220 nm, and one positive peak around 195 nm, indicate the predominant existence of α-helix along with the limited existence of β-sheets, turn and random forms. The CD spectra of fumarase in the presence of inhibitors displayed almost the same pattern as free fumarase, suggesting that no dramatic structural alterations were created by the inhibitors. However, the CD spectra revealed some changes in the enzyme's secondary structure upon the presence of an inhibitor. Thus, it could be concluded that both inhibitors induce the distinct conformational rearrangements in the enzyme's secondary structure. A notable increase in α-helical content, accompanied by a slight decrease in β-sheets, turns, and random coils was observed when Zn2+ ions were presented. These changes suggest that Zn2+ ions stabilize α-helical regions of the enzyme, possibly by promoting a more ordered and compact structure. This shift suggests that Zn2+ stabilizes α-helical domains, possibly restricting the conformational dynamics required for efficient substrate binding and ultimately leading to inhibition. Striking structural changes were observed in the presence of Zn(1)–CQDs, including a further increase in α-helical content, a further decrease in turns and random coil, and nearly complete elimination of β-sheet content. These changes suggest that Zn(1)–CQDs favor a highly helical structure and stabilize or induce specific α-helical conformation. Such stabilization of α-helical regions and suppression of structural flexibility are consistent with a more rigid enzyme conformation. This rigidity, driven by tighter enzyme–nanoparticle interactions and may impair the conformational adaptability essential for fumarase activity. Considering all the data, the CD spectra suggest that both inhibitors potentially reduce the enzyme's flexibility by promoting structural ordering in fumarase and stabilizing α-helix structures. Based on the CD results, the superior inhibitory effect of Zn(1)–CQDs is likely due to its stronger interaction with fumarase, interacting with multiple sites or subunits on the fumarase tetramer, effectively locking the enzyme into a highly rigid, inactive conformation. To further examine the structural effect of inhibitors on the fumarase, native PAGE analysis was performed. As shown in Fig. S9 (for details, see the SI), apo fumarase, Zn2+-treated fumarase, and Zn(1)–CQD-treated fumarase all exhibited a single band with identical mobility, indicating that the tetrameric state of the enzyme is preserved in the presence of the inhibitors. These results, together with the CD data, confirm that inhibitors do not induce significant structural or oligomeric changes in fumarase.Molecular docking and molecular dynamics reveal putative Zn2+ binding sites
It is well-established that enzymes containing histidine, aspartate, and glutamate residues in their active sites readily bind Zn2+ ions.70 Since these residues are already present in the active site of fumarase, which is located within a deep cavity at the interface of the homotetrameric structure,71,72 it is conceivable that the Zn2+ ions are stabilized through coordination with these residues. To investigate this interaction further, molecular docking simulations were performed using the Saccharomyces cerevisiae fumarase structure (for details, see the SI). The docking results were agreement with kinetic studies, strongly supporting the selective binding of Zn2+ to the active site cavity (Fig. 11a). Notably, despite the unbiased docking search covering the entire enzyme structure, Zn2+ predominantly localized within the active site rather than other regions of the protein through various binding poses (Fig. S7). Zn2+ is stabilized through interaction with Asn56 and Asn339, along with binary hydrogen bonds with Glu340 (Fig. 11b). Interestingly, studies on the crystal structure of fumarase with meso-tartrate, a known competitive inhibitor, also revealed an interaction with Glu340.72 These findings provide molecular insights into the putative Zn2+ binding site, supporting its role as a competitive inhibitor of fumarase. However, 10 ns molecular dynamics (MD) simulation was conducted for the first conformation obtained from docking to evaluate the stability of the Zn2+ ion in the active site (for details, see the SI). Significantly, the Zn2+ ion remained within the active site cavity throughout the simulation and exhibited consistent interactions with Asn56 and Asn339. However, while at the beginning of the simulation the Zn2+ ions initially formed two bonds with Glu340, only one bond persisted at the end, suggesting a stable coordination environment (Fig. 11c–f). To dive a little deeper into the stability of Zn2+ coordination, the average distances between the Zn2+ ion and key contributing residues were calculated throughout the simulations. The results indicate stable interactions between the inhibitor and the contributing amino acids of the active site. As such, Zn2+ maintains an average distance of 3.3 Å and 2.2 Å with Glu340 (sites 1 and 2) respectively, and also 2.08 Å with Asn339 as well as and 4.3 Å with Asn56 (Fig. 12a–d). Therefore, a well-defined coordination environment, reinforcing the stability of Zn2+ within the active site cavity could be assumed. Additionally, root mean square deviation (RMSD) analysis revealed that fumarase in the presence of Zn2+ exhibits lower structural deviations compared to the free enzyme (Fig. 12e), suggesting that Zn2+ binding enhances enzyme stability. Furthermore, the radius of gyration (Rg) measurements indicated a more compact structure for the enzyme upon the Zn2+ coordination, as also evidenced by reduced Rg variation (Fig. 12f). Considering all facts, it can be suggested that in the absence of Zn2+ ions, fumarase adopts a more flexible and extended conformation, whereas Zn2+ binding reduces overall dynamicity. More importantly, these results are consistent with CD spectroscopy, further supporting the stabilizing effect of Zn2+ on the enzyme structure.Conclusions
This study establishes Zn2+ ions as potent competitive inhibitors of Saccharomyces cerevisiae fumarase, a key TCA cycle enzyme catalyzing the hydration of fumarate to L-malate, increasing Km to 131.95 mM for L-malate and 6.72 mM for fumarate. However, the limited aqueous solubility of ZnCl2 restricts its practical utility. To address this, we synthesized zinc-modified carbon quantum dots (Zn–CQDs), which combine excellent water solubility with enhanced inhibitory efficacy. Optimized Zn(1)–CQDs (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ZnCl2 to citric acid ratio) further elevated Km to 221.32 mM for L-malate and 8.12 mM for fumarate. Strikingly, both Zn2+ and Zn–CQDs preferentially inhibit the reverse reaction (L-malate to fumarate), enabling selective modulation of fumarase activity. CD analysis confirmed minimal conformational changes in fumarase upon binding, while molecular docking and MD simulations identified the Zn2+ binding site within the enzyme's active site. These findings position Zn–CQDs as a novel, soluble platform for regulating fumarase, with significant potential for therapeutic applications in cancer and metabolic disorders linked to TCA cycle dysregulation. Future studies elucidating Zn–CQD–fumarase interactions could unlock further opportunities for precision therapeutics and metabolic engineering.
1 ZnCl2 to citric acid ratio) further elevated Km to 221.32 mM for L-malate and 8.12 mM for fumarate. Strikingly, both Zn2+ and Zn–CQDs preferentially inhibit the reverse reaction (L-malate to fumarate), enabling selective modulation of fumarase activity. CD analysis confirmed minimal conformational changes in fumarase upon binding, while molecular docking and MD simulations identified the Zn2+ binding site within the enzyme's active site. These findings position Zn–CQDs as a novel, soluble platform for regulating fumarase, with significant potential for therapeutic applications in cancer and metabolic disorders linked to TCA cycle dysregulation. Future studies elucidating Zn–CQD–fumarase interactions could unlock further opportunities for precision therapeutics and metabolic engineering.
Conflicts of interest
There are no conflicts to declare.Data availability
All experimental data, characterization details, and supporting information relevant to this study are provided in supplementary information (SI) associated with this article. Supplementary information is available. See DOI: https://doi.org/10.1039/d5tb02141c.Acknowledgements
The authors would like to thank Davoud Gharailou for his assistance with the TEM analysis. They also gratefully acknowledge the Sharif University of Technology Research Council for partial financial support.References
- A. Navaser, H. R. Kalhor and F. Hayati, Heliyon, 2023, 9, e19315 CrossRef CAS PubMed.
- Y. Li, J. Ding and W. Qin, J. Am. Chem. Soc., 2024, 146, 24389–24397 CrossRef CAS PubMed.
- S. Chaudhury, J. Phys. Chem. B, 2014, 118, 10405–10412 CrossRef CAS PubMed.
- C. Savojardo, D. Baldazzi, G. Babbi, P. L. Martelli and R. Casadio, Sci. Rep., 2022, 12, 17963 Search PubMed.
- A. N. Matthew, F. Leidner, G. J. Lockbaum, M. Henes, J. Zephyr, S. Hou, D. N. Rao, J. Timm, L. N. Rusere and D. A. Ragland, Chem. Rev., 2021, 121, 3238–3270 CrossRef CAS PubMed.
- A. A. Aziz and L. J. Twyman, ACS Appl. Mater. Interfaces, 2019, 11, 44941–44948 CrossRef CAS PubMed.
- Y. Pan, H. Li, F. Shahidi, T. Luo and Z. Deng, Trends Food Sci. Technol., 2022, 124, 38–50 CrossRef CAS.
- R. Arya, Z. Maben, D. Rane, A. Ali and L. J. Stern, ACS Chem. Biol., 2022, 17, 1756–1768 CrossRef CAS PubMed.
- S. E. Wenzel and A. K. Kamada, Ann. Pharmacother., 1996, 30, 858–864 Search PubMed.
- S. Chaudhuri, J. A. Symons and J. Deval, Antiviral Res., 2018, 155, 76–88 Search PubMed.
- M. A. Ajalla Aleixo, V. L. Rangel, J. K. Rustiguel, R. A. de Pádua and M. C. Nonato, FEBS J., 2019, 286, 1925–1940 Search PubMed.
- P. R. Feliciano, C. L. Drennan and M. C. Nonato, ACS Chem. Biol., 2019, 14, 266–275 CrossRef CAS PubMed.
- O. Yogev, O. Yogev, E. Singer, E. Shaulian, M. Goldberg, T. D. Fox and O. Pines, PLoS Biol., 2010, 8, e1000328 CrossRef PubMed.
- L. Valcarcel-Jimenez and C. Frezza, Br. J. Cancer, 2023, 129, 1546–1557 Search PubMed.
- O. Yogev, A. Naamati and O. Pines, FEBS J., 2011, 278, 4230–4242 Search PubMed.
- S. Picaud, K. L. Kavanagh, W. W. Yue, W. H. Lee, S. Muller-Knapp, O. Gileadi, J. Sacchettini and U. Oppermann, J. Inherited Metab. Dis., 2011, 34, 671–676 Search PubMed.
- A. King, M. Selak and E. Gottlieb, Oncogene, 2006, 25, 4675–4682 Search PubMed.
- T. Chen, T. Wang, W. Liang, Q. Zhao, Q. Yu, C.-M. Ma, L. Zhuo, D. Guo, K. Zheng and C. Zhou, Cancer Res., 2019, 79, 1383–1397 Search PubMed.
- S. Sudarshan, W. Linehan and L. Neckers, Br. J. Cancer, 2007, 96, 403–407 CrossRef CAS PubMed.
- T. Takeuchi, P. T. Schumacker and S. A. Kozmin, J. Am. Chem. Soc., 2015, 137, 564–567 CrossRef CAS PubMed.
- M. J. Kerins, J. Milligan, J. A. Wohlschlegel and A. Ooi, Cancer Sci., 2018, 109, 2757–2766 CrossRef CAS.
- J. Harrison and J. A. Cox, Journal, 2019, 62, 10583–10585 CAS.
- J. Zhang, E. Grandi, H. Fu, T. Saravanan, L. Bothof, P. G. Tepper, A. M. W. Thunnissen and G. J. Poelarends, Angew. Chem., 2020, 132, 437–443 Search PubMed.
- M. Kasbekar, G. Fischer, B. T. Mott, A. Yasgar, M. Hyvönen, H. I. Boshoff, C. Abell, C. E. Barry III and C. J. Thomas, Proc. Natl. Acad. Sci. U. S. A., 2016, 113, 7503–7508 Search PubMed.
- J. Greenhut, H. Umezawa and F. B. Rudolph, J. Biol. Chem., 1985, 260, 6684–6686 CrossRef CAS.
- P. Penner and L. Cohen, J. Biol. Chem., 1969, 244, 1070–1075 CrossRef CAS.
- A. J. Whitehouse, M. D. J. Libardo, M. Kasbekar, P. D. Brear, G. Fischer, C. J. Thomas, C. E. Barry III, H. I. Boshoff, A. G. Coyne and C. Abell, J. Med. Chem., 2019, 62, 10586–10604 CrossRef CAS PubMed.
- A. Casini, A. Guerri, C. Gabbiani and L. Messori, J. Inorg. Biochem., 2008, 102, 995–1006 CrossRef CAS PubMed.
- M. J. Knape, L. G. Ahuja, D. Bertinetti, N. C. Burghardt, B. Zimmermann, S. S. Taylor and F. W. Herberg, ACS Chem. Biol., 2015, 10, 2303–2315 Search PubMed.
- Y. Sun, Z. Li, J. Wu, Z. Wang, Y. Dong, H. Wang, J. L. Brash, L. Yuan and H. Chen, J. Mater. Chem. B, 2019, 7, 3260–3267 Search PubMed.
- A. Casini and J. Reedijk, Chem. Sci., 2012, 3, 3135–3144 Search PubMed.
- C.-M. Che and F.-M. Siu, Curr. Opin. Chem. Biol., 2010, 14, 255–261 Search PubMed.
- K. J. Kilpin and P. J. Dyson, Chem. Sci., 2013, 4, 1410–1419 Search PubMed.
- Y. Li, Y. Wang, L. Zhao, M. H. Stenzel and Y. Jiang, Mater. Horiz., 2024, 11, 4275–4310 RSC.
- D. Ekinci and M. Şentürk, J. Enzyme Inhib. Med. Chem., 2013, 28, 11–15 Search PubMed.
- N. Hadizadeh Shirazi, J. Food Biochem., 2019, 43, e12724 Search PubMed.
- Y. N. Xue, Y. N. Liu, J. Su, J. L. Li, Y. Wu, R. Guo, B. B. Yu, X. Y. Yan, L. C. Zhang and L. K. Sun, Cancer Med., 2019, 8, 2462–2473 CrossRef CAS PubMed.
- Z. Chen, Q. Zhou and R. Ge, Biometals, 2012, 25, 95–102 CrossRef CAS PubMed.
- T. Ollevier, A. Jalba and H. Keipour, Encycl. Reagents Org. Synth., 2001, 1–9 Search PubMed.
- L. E. Pelepenko, A. C. P. Janini, B. P. Gomes, A. de-Jesus-Soares and M. A. Marciano, Antibiotics, 2022, 11, 1741 CrossRef CAS PubMed.
- M. Takeuchi and Y. Amao, New J. Chem., 2023, 47, 17679–17684 RSC.
- L. Jiang, Y. Gu, Y. Du and J. Liu, Mol. Pharmaceutics, 2019, 16, 3333–3349 CrossRef CAS PubMed.
- A.-M. Alam, B.-Y. Park, Z. K. Ghouri, M. Park and H.-Y. Kim, Green Chem., 2015, 17, 3791–3797 RSC.
- B. C. Martindale, G. A. Hutton, C. A. Caputo and E. Reisner, J. Am. Chem. Soc., 2015, 137, 6018–6025 Search PubMed.
- W. Kong, J. Liu, R. Liu, H. Li, Y. Liu, H. Huang, K. Li, J. Liu, S.-T. Lee and Z. Kang, Nanoscale, 2014, 6, 5116–5120 RSC.
- Y. Pan, J. Yang, Y. Fang, J. Zheng, R. Song and C. Yi, J. Mater. Chem. B, 2017, 5, 92–101 Search PubMed.
- L. Zhu, D. Shen, C. Wu and S. Gu, Ind. Eng. Chem. Res., 2020, 59, 22017–22039 CrossRef CAS.
- M. Bai, X. Shao, C. Wang, J. Wang, X. Wang, P. Guan and X. Hu, Mater. Horiz., 2025, 12, 673–693 Search PubMed.
- G. Han, J. Zhao, R. Zhang, X. Tian, Z. Liu, A. Wang, R. Liu, B. Liu, M. Y. Han and X. Gao, Angew. Chem., 2019, 131, 7161–7165 Search PubMed.
- Y. Li, Y. Yuan, X. Liang and L. Zhao, ACS Appl. Nano Mater., 2022, 5, 14507–14519 CrossRef CAS.
- X.-W. Hua, Y.-W. Bao and F.-G. Wu, ACS Appl. Mater. Interfaces, 2018, 10, 10664–10677 Search PubMed.
- M. Hasani and H. R. Kalhor, ACS Catal., 2021, 11, 10778–10788 Search PubMed.
- M. Hasani and H. R. Kalhor, J. Org. Chem., 2024, 89, 13836–13846 CrossRef CAS PubMed.
- C. Tang, J. Zhou, Z. Qian, Y. Ma, Y. Huang and H. Feng, J. Mater. Chem. B, 2017, 5, 1971–1979 Search PubMed.
- L. Chai, J. Zhou, H. Feng, C. Tang, Y. Huang and Z. Qian, ACS Appl. Mater. Interfaces, 2015, 7, 23564–23574 Search PubMed.
- J. Korram, L. Dewangan, R. Nagwanshi, I. Karbhal, K. K. Ghosh and M. L. Satnami, New J. Chem., 2019, 43, 6874–6882 RSC.
- S. Peulon and D. Lincot, J. Electrochem. Soc., 1998, 145, 864 Search PubMed.
- J. F. Da Silva and R. J. P. Williams, The biological chemistry of the elements: the inorganic chemistry of life, Oxford University Press, 2001 Search PubMed.
- K. A. McCall, C.-c Huang and C. A. Fierke, J. Nutr., 2000, 130, 1437S–1446S Search PubMed.
- J. C. Peacock and B. L. D. Peacock, J. Am. Pharm. Assoc., 1918, 7, 689–697 Search PubMed.
- Y. Lv, M. Ma, Y. Huang and Y. Xia, Chem. - Eur. J., 2019, 25, 954–960 Search PubMed.
- M. Hasani, H. R. Kalhor, M. Salehi and F. Rahgozar, Org. Biomol. Chem., 2025, 23, 7320–7330 RSC.
- S. E. Elugoke, G. E. Uwaya, T. W. Quadri and E. E. Ebenso, in Carbon dots: recent developments and future perspectives, ACS Publications, 2024, pp. 3–42 Search PubMed.
- S. K. Tammina, Y. Wan, Y. Li and Y. Yang, J. Photochem. Photobiol., B, 2020, 202, 111734 CrossRef CAS PubMed.
- M. Alikhani, E. Khoshkalam, J. Sadeghi, L. Bulgariu and H. Eshghi, RSC Adv., 2024, 14, 24534–24547 RSC.
- G. Filippini, F. Amato, C. Rosso, G. Ragazzon, A. Vega-Peñaloza, X. Companyó, L. Dell’Amico, M. Bonchio and M. Prato, Chem, 2020, 6, 3022–3037 Search PubMed.
- J. A. Boiani, J. Chem. Educ., 1986, 63, 724 Search PubMed.
- A. M. Brown, B. S. Kristal, M. S. Effron, A. I. Shestopalov, P. A. Ullucci, K.-F. R. Sheu, J. P. Blass and A. J. Cooper, J. Biol. Chem., 2000, 275, 13441–13447 Search PubMed.
- K. E. Dineley, T. V. Votyakova and I. J. Reynolds, J. Neurochem., 2003, 85, 563–570 Search PubMed.
- W. Maret, Biometals, 2013, 26, 197–204 Search PubMed.
- A. E. Mechaly, A. Haouz, I. Miras, N. Barilone, P. Weber, W. Shepard, P. M. Alzari and M. Bellinzoni, FEBS Lett., 2012, 586, 1606–1611 Search PubMed.
- T. Weaver, M. Lees, V. Zaitsev, I. Zaitseva, E. Duke, P. Lindley, S. McSweeny, A. Svensson, J. Keruchenko and I. Keruchenko, J. Mol. Biol., 1998, 280, 431–442 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |