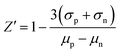Graphene oxide-based fluorescent biosensor for high-throughput screening to discover SARS-CoV-2 RdRp inhibitors
Hojeong
Shin†
 a,
Jiwon
Woo†
a,
Jiwon
Woo†
 a,
Soeun
Yoo
a,
Soo Min
Lee
a,
So Young
Kim
a,
Hyung Jin
Cha
a,
Soeun
Yoo
a,
Soo Min
Lee
a,
So Young
Kim
a,
Hyung Jin
Cha
 b,
Seounghun
Kang
c,
Ji-Seon
Lee
b,
Seounghun
Kang
c,
Ji-Seon
Lee
 a,
Seho
Lee
a,
Seho
Lee
 a,
Yoo-Bin
Kwon
a,
Yoo-Bin
Kwon
 a,
Jae-Sung
Woo
b and
Dal-Hee
Min
a,
Jae-Sung
Woo
b and
Dal-Hee
Min
 *a
*a
aDepartment of Chemistry, Seoul National University, Seoul 08826, Republic of Korea. E-mail: dalheemin@snu.ac.kr
bDepartment of Life Sciences, Korea University, Seoul 02841, Republic of Korea
cDepartment of Chemistry, Soongsil University, Seoul, 06978, Republic of Korea
First published on 30th December 2025
Abstract
The COVID-19 pandemic caused by SARS-CoV-2 underscored the global need for rapid, efficient drug discovery platforms to combat emerging viral threats. Conventional antiviral screening methods are often time-consuming and low-throughput, making them insufficient for timely therapeutic development during acute outbreaks. RNA-dependent RNA polymerase (RdRp), a key enzyme in viral replication, represents a validated antiviral target for RNA viruses, including SARS-CoV-2. However, few assays directly monitor RdRp activity in a high-throughput format. To address this gap, we developed a fluorescence-based assay for real-time monitoring of RdRp activity using graphene oxide nanomaterials. Here, we designed a graphene oxide-based RdRp assay that transduces polymerase activity into measurable fluorescence intensity changes. The assay is rapid, homogeneous, and compatible with multi-well plate formats for high-throughput screening. Using this platform, we screened a library of FDA-approved small molecules and identified fingolimod, an immunomodulatory drug for multiple sclerosis, as a potential RdRp inhibitor. In vitro cell-based assays confirmed that fingolimod significantly reduced SARS-CoV-2 replication without cytotoxicity at therapeutic concentrations. This result supports fingolimod's potential as a repurposed direct-acting antiviral agent. The assay's robustness highlights its applicability in antiviral drug discovery, enabling rapid responses to future viral outbreaks. This graphene oxide-based RdRp assay provides a versatile tool for antiviral screening and demonstrates the feasibility of repurposing approved drugs as direct-acting antivirals. The platform's adaptability and rapid readout capability make it well-suited for pandemic preparedness and therapeutic discovery against emerging viral threats.
1. Introduction
The COVID-19 pandemic, caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has emerged as the most significant global health crisis in recent decades, resulting in over a million fatalities and infecting millions more worldwide.1,2 The continuing spread of the virus underscores the urgent need for effective therapeutic strategies to reduce both morbidity and mortality, particularly among vulnerable populations. Current pharmacological interventions are limited, with few treatments like Remdesivir receiving U.S. Food and Drug Administration (FDA) approval for treating COVID-19.2 As the pandemic persists, the discovery and development of novel antiviral agents remain a critical priority to address emerging challenges such as viral resistance and variant proliferation.SARS-CoV-2 is a positive-sense, single-stranded RNA virus belonging to the betacoronavirus genus, which also includes other significant human pathogens like SARS-CoV and MERS-CoV.3 The virus encodes several non-structural proteins (nsps), crucial for its replication cycle. Among these, the RNA-dependent RNA polymerase (RdRp) is a key enzyme responsible for viral RNA synthesis. Due to its essential role in viral replication and absence of closely related homologs in humans, RdRp is an attractive target for antiviral drug development.4 Moreover, SARS-CoV-2 RdRp is highly conserved among coronaviruses, with 96% homology to SARS-CoV and 70% to MERS-CoV.5 Notably, spontaneous mutations are less frequent in the RdRp region compared to the spike protein region among SARS-CoV-2 variants.6,7 This conservation suggests that targeting RdRp could yield prophylactic and therapeutic drugs effective against multiple SARS-CoV-2 variants, providing a broad-spectrum antiviral strategy.
In general, the rapid mutation rate of RNA viruses poses ongoing challenges for drug development, further emphasizing the need for robust methods to discover effective antivirals. However, drug discovery faces significant challenges, as conventional RdRp assays, such as gel electrophoresis or radioisotope labeling, are labor-intensive, time-consuming, and costly, making them impractical for high-throughput applications. These limitations hinder the rapid identification of promising drug candidates, which is critical during fast-spreading viral outbreaks.
To overcome these challenges, we developed a nano-graphene oxide (NGO)-based biosensor designed to detect the enzymatic activity of SARS-CoV-2 RdRp. NGO offers several advantages for biosensing applications, including a large surface area, excellent electrical conductivity, and abundant oxygen-containing functional groups.8 The ability of NGO to quench fluorescence and interact with nucleic acids through π–π stacking interactions and hydrogen bonding can be exploited to develop sensitive and selective biosensor platforms.9 By converting enzymatic activity into quantifiable fluorescence intensity, our NGO-based biosensor simplifies the detection process and enhances throughput, making it appropriate for high-throughput drug screening. Using this platform, we conducted extensive screening of 1443 FDA-approved drugs to identify potential RdRp inhibitors. Among the candidates tested, fingolimod—a drug originally approved for treating multiple sclerosis—demonstrated the strongest inhibitory effect on RNA replication. Subsequent cell-based evaluations indicated that fingolimod may act as a direct-acting antiviral agent by inhibiting RdRp activity, offering a promising avenue for repurposing existing drugs against SARS-CoV-2. Beyond SARS-CoV-2, the versatility of our NGO-based biosensor provides a scalable platform for discovering RdRp inhibitors effective against other RNA viruses, addressing the need for rapid responses to future large-scale outbreaks.
2. Experimental
2.1. Synthesis and characterization of NGO
NGO was synthesized using a modified Hummers’ method.10,11 Briefly, graphite powder, sodium nitrate, and sulfuric acid were mixed under vigorous stirring in an ice bath. Then, KMnO4 was slowly added to the mixture. The flask was then heated at 35 °C for 1 hour. Subsequently, DW was added while maintaining it in an ice bath. The flask was returned to the oil bath and heated at 95 °C for 30 minutes. Residual reactive substances were quenched by adding H2O2. The resulting graphene oxide solution was purified by filtration and dialyzed.Fourier transform infrared (FT-IR) spectra were recorded using an FTIR-7600 spectrometer (Lambda Scientific Systems, Inc.). Raman spectroscopy was performed using a HEDA system (WEVE) equipped with a 532 nm diode laser, focused through an integrated microscope (BX53M, Olympus) with a 20× objective lens (MPLFLN20×, Olympus). The size and morphology were examined with an atomic force microscope (AFM; NX10, Park Systems). Hydrodynamic size and zeta potential were measured using a Zetasizer Nano ZS (Malvern). Ultraviolet-visible (UV-Vis) absorption spectra were obtained using a GENESYS 150 spectrophotometer (Thermo Scientific).
2.2. Protein purification of SARS-CoV-2 nsp12
The gene encoding the SARS-CoV-2 nsp12 was fused to the coding sequence of the C-terminal His-tagged mCherry tag and 3C cleavage site, and inserted into the pX vector.12 The plasmid was transfected into HEK293E cells as described previously.10,12 The transfected cells were grown at 33 °C for 3 days and harvested using a centrifuge at 500g for 15 min. The harvested cells were resuspended with a buffer (20 mM Tris, pH 7.5, 300 mM NaCl, 10% glycerol, 2 mM β-mercaptoethanol, 1 mM MgCl2, ribonuclease A, 0.1 mM PMSF, and protease inhibitor cocktail (Thermo Fisher Scientific, catalog no. 88666)). After lysis by sonication, the lysate was centrifuged at 35![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 1 hour. The supernatant was loaded onto a Ni-NTA resin pre-equilibrated with buffer A (20 mM Tris, pH 7.5, 300 mM NaCl, 2 mM β-mercaptoethanol, and 1 mM MgCl2). The bound proteins were washed using buffer A supplemented with 40 mM imidazole and eluted using buffer A supplemented with 300 mM imidazole and 5% glycerol. After treatment with 3C protease, the cleaved proteins were further purified using a Superdex 200 increase 10/300 column (GE Healthcare) equilibrated with a buffer (20 mM HEPES, pH 7.5, 300 mM NaCl, 1 mM TCEP, and 1 mM MgCl2, and 5% glycerol). The peak fractions were pooled, concentrated, flash frozen and stored at −80 °C prior to the biochemical assay.
000g for 1 hour. The supernatant was loaded onto a Ni-NTA resin pre-equilibrated with buffer A (20 mM Tris, pH 7.5, 300 mM NaCl, 2 mM β-mercaptoethanol, and 1 mM MgCl2). The bound proteins were washed using buffer A supplemented with 40 mM imidazole and eluted using buffer A supplemented with 300 mM imidazole and 5% glycerol. After treatment with 3C protease, the cleaved proteins were further purified using a Superdex 200 increase 10/300 column (GE Healthcare) equilibrated with a buffer (20 mM HEPES, pH 7.5, 300 mM NaCl, 1 mM TCEP, and 1 mM MgCl2, and 5% glycerol). The peak fractions were pooled, concentrated, flash frozen and stored at −80 °C prior to the biochemical assay.
2.3. Protein purification of SARS-CoV-2 nsp7 and nsp8
The sequence encoding SARS-CoV-2 nsp7 or nsp8 was inserted into a pET28a expression vector as a fusion construct with an N-terminal His-tagged superfolder GFP tag and TEV protease cleavage site (ENLYFQ) to produce intact nsp7 or nsp8. nsp7 and nsp8 proteins were expressed and purified separately with the same procedure, as mentioned below. Transformed E. coli BL21(DE3) cells were grown in the Terrific Broth medium supplemented with 30 µg mL−1 kanamycin at 37 °C. When OD600 reached 1.2–1.4, we added 0.5 mM IPTG into the media to overexpress the desired proteins. After the cells were further grown at 18 °C for 12–18 hours, the cells were harvested using a centrifuge at 5500g for 15 min. The harvested cells were resuspended in buffer B (20 mM Tris, pH 7.5, 150 mM NaCl, and 2 mM β-mercaptoethanol) supplemented with 4 mM MgCl2, 10% glycerol, 0.1 mM PMSF, ribonuclease A, and protease inhibitor cocktail. The resuspended cells were lysed by sonication and the lysate was centrifuged at 35![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 g for 1 hour. For purification, the supernatant was loaded on a Ni-NTA resin and washed with buffer B supplemented with 40 mM imidazole. The resin-bound proteins were eluted using buffer B supplemented with 300 mM imidazole and 5% glycerol. After treatment with TEV protease, the cleaved proteins were loaded onto a HiTrapQ column (GE Healthcare). The proteins were eluted with a 50–500 mM NaCl gradient in a buffer containing 20 mM Tris, pH 8.0, 1 mM DTT, 1 mM MgCl2, and 5% glycerol. The eluted proteins were further purified using a Superdex 75 increase 10/300 column (GE Healthcare) in a final buffer (20 mM HEPES, pH 7.5, 150 mM NaCl, 1 mM TCEP, 1 mM MgCl2, and 5% glycerol). The peak fractions were pooled and stored at −80 °C prior to the biochemical assay.
000 g for 1 hour. For purification, the supernatant was loaded on a Ni-NTA resin and washed with buffer B supplemented with 40 mM imidazole. The resin-bound proteins were eluted using buffer B supplemented with 300 mM imidazole and 5% glycerol. After treatment with TEV protease, the cleaved proteins were loaded onto a HiTrapQ column (GE Healthcare). The proteins were eluted with a 50–500 mM NaCl gradient in a buffer containing 20 mM Tris, pH 8.0, 1 mM DTT, 1 mM MgCl2, and 5% glycerol. The eluted proteins were further purified using a Superdex 75 increase 10/300 column (GE Healthcare) in a final buffer (20 mM HEPES, pH 7.5, 150 mM NaCl, 1 mM TCEP, 1 mM MgCl2, and 5% glycerol). The peak fractions were pooled and stored at −80 °C prior to the biochemical assay.
2.4. NGO-based RdRp activity assay
The reaction mixture for measuring RdRp activity consisted of 0.2 µM RNA substrate, 0.5 µM SARS-CoV-2 RdRp complex (comprising 0.5 µM nsp12, 1.5 µM nsp7, and 1.5 µM nsp8), and NTPs (67 µM ATP, and 33 µM each of CTP, UTP, and GTP) in RdRp buffer (20 mM HEPES, pH 7.5; 10 mM KCl; 1 mM MgCl2; 1 mM DTT). The mixture was incubated at 37 °C for 1 hour. After incubation, NGO was added to each sample at a final concentration of 5 µg mL−1. RdRp activity was measured by monitoring the change in fluorescence intensity using a Synergy MX (BioTek).2.5. Z′-factor determination
To evaluate the assay's performance, the Z′-factor was calculated using positive and negative controls. For the positive control, the reaction mixture included 0.2 µM RNA substrate, 0.5 µM SARS-CoV-2 RdRp and NTPs as described above. For the negative control, nuclease-free water (NFW) replaced the NTP solution. After 1 hour of incubation at 37 °C, NGO was added to each well at a final concentration of 5 µg mL−1. Fluorescence intensity was measured immediately after NGO addition. The Z′ factor is calculated using the following equation:where μ and σ represent the mean and standard deviation, respectively, of the positive (p) and negative (n) controls.
2.6. High-throughput RdRp inhibitor screening
High-throughput screening was conducted using 1443 compounds from an FDA-approved chemical library (Selleckchem). Each compound was added to the RdRp activity reaction mixture at a final concentration of 100 µM under the optimized conditions described above. For the negative control, NTPs were replaced with NFW. After 1 hour of incubation at 37 °C, NGO was added at a final concentration of 5 µg mL−1. The final DMSO concentration was maintained at 2% (v/v). Fluorescence intensity was measured immediately after NGO addition.Hits were identified as compounds exhibiting inhibition greater than 3 standard deviations from the mean of the positive controls. To eliminate false positives, primary hits were subjected to a counter-screen in the absence of the RdRp enzyme. Assay robustness was monitored by calculating the Z′-factor and coefficient of variation (CV) for each plate.
2.7. Determination of IC50
To determine the half-maximal inhibitory concentration (IC50) of fingolimod, reaction mixtures were prepared containing 0.2 µM RNA substrate, 0.5 µM SARS-CoV-2 RdRp, NTPs (67 µM ATP and 33 µM each of CTP, UTP, and GTP), and varying concentrations of fingolimod (0 to 40 µM) in RdRp buffer (20 mM HEPES, pH 7.5; 10 mM KCl; 1 mM MgCl2; 1 mM DTT). After 1 hour of incubation at 37 °C, NGO was added at a final concentration of 5 µg mL−1, and fluorescence intensities were measured.To evaluate target selectivity, the inhibitory activity of fingolimod was tested against the Dengue virus RdRp using the same assay platform described previously.10
2.8. Enzyme kinetic analysis
To determine the mechanism of inhibition, RdRp activity assays were performed with varying concentrations of NTPs in the absence or presence of 50, 100 µM fingolimod. The initial velocity V0 was determined based on the fluorescence quenching efficiency. Kinetic parameters, including the Michaelis constant (Km) and maximum velocity (Vmax), were calculated by fitting the data to the Michaelis–Menten equation using non-linear regression analysis in GraphPad Prism.2.9. Cells and viruses
HCoV-OC43 (KBPV-VR-8) was purchased from the Korea Bank for Pathogenic Viruses (KBPV). Vero E6 was kindly provided by Professor K. Ahn from the Department of Biological Sciences, Seoul National University. SARS-CoV-2 virus (NCCP no. 43326, βCoV/Korea/KCDC03/2020) was obtained from the Korea Centers for Disease Control and Prevention (KCDC). Vero cells were sourced from the American Type Culture Collection (ATCC-CCL81).2.10. In vitro virus infection test using HCoV-OC43
Vero E6 cells were seeded at a density of 3 × 104 cells per well in 96-well plates. After overnight incubation at 37 °C with 5% CO2, the cells were infected with HCoV-OC43 virus in DMEM supplemented with 2% fetal bovine serum (FBS) and penicillin–streptomycin (P/S). The virus was added to the cell monolayers and incubated for 1 hour, with gentle rocking every 30 minutes to ensure even distribution. Fingolimod solutions were prepared in complete culture medium containing methylcellulose at serially diluted concentrations, with a final DMSO concentration of 0.1%. After the infection period, wells were overlaid with 100 µL of DMEM containing 10% FBS, P/S, and methylcellulose, and incubated for 2 days.Following incubation, cell monolayers were fixed with 4% paraformaldehyde and blocked with a blocking solution for 1 hour. Samples were then incubated with a primary antibody specific for HCoV-OC43 (Millipore Sigma, MAB9013), followed by a fluorescein isothiocyanate (FITC)-labeled goat anti-mouse IgG secondary antibody (Sigma-Aldrich, F0257). Fluorescence was measured to assess viral infection levels.
2.11. Determination of the stage of antiviral action
To identify the specific stage of the viral life cycle inhibited by fingolimod, time-of-addition assays were performed using HCoV-OC43. The cells were treated with 25 µM fingolimod under three conditions: (i) full-time: drug present during pre-treatment (1 hour), viral infection (1 hour), and the subsequent incubation period; (ii) entry: drug present only during pre-treatment (1 hour) and viral infection (1 hour), followed by incubation in a drug-free medium; and (iii) post-entry: drug added only after the viral infection period. Viral infection was quantified by immunofluorescence staining as described above.2.12. In vitro virus infection test using SARS-CoV-2
Experiments involving live SARS-CoV-2 were conducted under contract at the Institut Pasteur Korea to comply with BSL-3 regulations. Vero cells were seeded at 1.2 × 104 cells per well in 384-well tissue culture plates. The following day, compounds dissolved in DMSO were serially diluted twofold, with a maximum concentration of 50 µM, and added to the cells. Approximately 1 hour after compound treatment, cells were infected with SARS-CoV-2 at a multiplicity of infection (MOI) of 0.025 in a BSL-3 facility and incubated at 37 °C for 48 hours.The cells were then fixed with 4% paraformaldehyde and permeabilized. After blocking, cells were incubated with the anti-SARS-CoV-2 N protein primary antibody (Sino Biological), followed by the Alexa Fluor 488-conjugated goat anti-rabbit IgG secondary antibody (Molecular Probes) and Hoechst 33342 for nuclear staining. Fluorescent images were acquired using the Operetta high-content imaging system (PerkinElmer). Image analysis was performed with Columbus software.
2.13. Statistical analysis
Concentration-dependent assay data were analyzed using GraphPad Prism. Results are presented as mean ± standard deviation (SD), with error bars indicating the variability across replicates.3. Results and discussion
3.1. Development of the NGO-based RdRp activity assay
The NGO-based RdRp activity assay exploits the differential interaction of NGO with single-stranded RNA (ssRNA) and double-stranded RNA (dsRNA) (Fig. 1). In the assay, a fluorescent dye-labeled ssRNA substrate is incubated with the RdRp complex. In the absence of RdRp activity, ssRNA substrate remains single-stranded and interacts strongly with NGO through π–π interactions and hydrogen bonding. This close association brings the fluorophore into proximity with NGO, resulting in efficient quenching of fluorescence. In contrast, when the RdRp is active and synthesizes the complementary RNA strand, the ssRNA is converted into dsRNA. In dsRNA, the nucleobases are sequestered within the helical structure, reducing their interaction with NGO. This structural change reduces the quenching effect. Therefore, the fluorescence intensity correlates with RdRp activity, allowing for quantitative analysis of enzyme activity. | ||
| Fig. 1 Schematic illustration of NGO-based SARS-CoV-2 RdRp activity assay for high-throughput drug screening. | ||
To develop the NGO-based RdRp activity assay, we first synthesized the NGO. The NGO was prepared using a modified Hummers' method, ensuring the production of nanoscale sheets. Fourier transform infrared spectroscopy (FT-IR) confirmed the presence of oxygen-containing functional groups, with characteristic peaks observed at 3428 cm−1 (O–H stretching), 1718 cm−1 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching), and 1078 cm−1 (C–O stretching) (Fig. S1A). Raman spectroscopy revealed prominent D and G bands at 1342 cm−1 and 1591 cm−1, respectively, indicating the presence of disordered and graphitic carbon structures (Fig. S1B). X-ray photoelectron spectroscopy (XPS) analysis further characterized the chemical composition, revealing a C/O atomic ratio of 2.16 (Fig. S1C). The high-resolution C 1s spectrum of NGO was deconvoluted into four distinct peaks centered at 284.2, 286.2, 287.7, and 288.7 eV, corresponding to C–C/C
O stretching), and 1078 cm−1 (C–O stretching) (Fig. S1A). Raman spectroscopy revealed prominent D and G bands at 1342 cm−1 and 1591 cm−1, respectively, indicating the presence of disordered and graphitic carbon structures (Fig. S1B). X-ray photoelectron spectroscopy (XPS) analysis further characterized the chemical composition, revealing a C/O atomic ratio of 2.16 (Fig. S1C). The high-resolution C 1s spectrum of NGO was deconvoluted into four distinct peaks centered at 284.2, 286.2, 287.7, and 288.7 eV, corresponding to C–C/C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C–O, C
C, C–O, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, and O–C
O, and O–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bonds, respectively (Fig. S1D). Quantitative analysis indicated relative compositions of 44.4%, 43.0%, 6.9%, and 5.7% for these functional groups, respectively (Fig. S1E). The morphological and dimensional characteristics of the NGO were further analyzed. Dynamic light scattering (DLS) revealed a mean hydrodynamic diameter of 20.2 nm (Fig. S2A). The zeta potential was measured to be −45.2 ± 1.9 mV, indicating high colloidal stability due to electrostatic repulsion. Atomic force microscopy (AFM) analysis confirmed the synthesis of uniform NGO sheets, with the particle height histogram (Fig. S2B) and the corresponding topographic profile (Fig. S2C) indicating an average thickness of approximately 2 nm. Taken together, it was clearly confirmed that NGO was successfully prepared.
O bonds, respectively (Fig. S1D). Quantitative analysis indicated relative compositions of 44.4%, 43.0%, 6.9%, and 5.7% for these functional groups, respectively (Fig. S1E). The morphological and dimensional characteristics of the NGO were further analyzed. Dynamic light scattering (DLS) revealed a mean hydrodynamic diameter of 20.2 nm (Fig. S2A). The zeta potential was measured to be −45.2 ± 1.9 mV, indicating high colloidal stability due to electrostatic repulsion. Atomic force microscopy (AFM) analysis confirmed the synthesis of uniform NGO sheets, with the particle height histogram (Fig. S2B) and the corresponding topographic profile (Fig. S2C) indicating an average thickness of approximately 2 nm. Taken together, it was clearly confirmed that NGO was successfully prepared.
To investigate the in vitro activity of the SARS-CoV-2 RdRp, we first focused on designing an appropriate RNA template for the polymerase assay. SARS-CoV-2 RdRp has been shown to utilize RNA templates that can fold back on themselves at the 3′ end, forming intramolecular hairpin structures.13 This folding enables the production of near-dimer-sized hairpin products through a mechanism known as copy-back RNA synthesis. Therefore, we engineered a 25-nt RNA template designed to form a stable tetraloop near the 3′ terminus (Fig. 2A).
The SARS-CoV-2 RdRp complex consists of the catalytic subunit nsp12 and the accessory proteins nsp7 and nsp8, which enhance polymerase activity and processivity.13–15 We expressed and purified recombinant nsp12, nsp7, and nsp8 proteins using a mammalian expression system to ensure appropriate folding and post-translational modifications. The in vitro activity of the assembled RdRp complex was assessed using the designed RNA hairpin substrate in a gel-based extension assay, demonstrating RdRp activity (Fig. 2B).
To optimize the NGO-based RdRp assay, we determined the optimal NGO concentration that maximizes the fluorescence quenching of ssRNA while minimizing quenching of dsRNA. We tested NGO concentrations ranging from 1 to 10 µg mL−1 in the presence of fluorescent dye-labeled ssRNA and dsRNA (Fig. 2C and Fig. S3). The fluorescence intensity was measured for each concentration, and an NGO concentration of 5 µg mL−1 was selected as the optimal concentration, as it offered a substantial fluorescence difference between ssRNA and dsRNA without overly quenching the baseline dsRNA signal.
We next validated the assay by comparing fluorescence signals in RdRp reactions with and without NTPs, which are essential for RNA synthesis. RdRp reactions containing NTPs (NTP+) exhibited increased fluorescence compared to those without NTPs (NTP−), indicating successful RNA synthesis and the formation of dsRNA that avoided NGO quenching (Fig. 2D). The significant difference in fluorescence between the two groups confirmed the capability of the assay to detect RdRp activity.
Furthermore, we confirmed the functional reproducibility of the NGO by comparing the assay performance of independently synthesized batches. All tested batches exhibited consistent quenching of ssRNA and preservation of dsRNA fluorescence, demonstrating a stable assay window (Fig. S4).
3.2. High-throughput screening for RdRp inhibitors
To assess the suitability of the assay for high-throughput screening, we calculated the Z′ factor, a statistical parameter that evaluates assay quality.16 A Z′ factor between 0.5 and 1.0 indicates an excellent assay and our assay achieved a Z′ factor of 0.79, demonstrating its robustness and reliability for high-throughput applications. To verify the assay robustness with a pharmacological inhibitor, we validated the assay using remdesivir triphosphate (RTP) at the screening concentration as a positive control. Under these conditions, the assay maintained a robust performance with a recalculated Z′-factor of 0.81.Employing our NGO-based assay, we screened an FDA-approved drug library comprising 1443 compounds to identify potential RdRp inhibitors (Fig. S5 and Table S1). Among the compounds tested, fingolimod emerged as the most potent inhibitor, exhibiting approximately 90% inhibition of RdRp activity at a concentration of 200 µM.
To quantify the inhibitory effect of fingolimod, we performed dose–response experiments using our NGO-based assay. The inhibition curve showed a sigmoidal relationship between fingolimod concentration and RdRp activity, with a calculated half-maximal inhibitory concentration (IC50) of 17.2 µM (Fig. 3A). The inhibitory effect was further validated using a conventional gel-based extension assay, confirming that fingolimod effectively inhibits RdRp-mediated RNA synthesis in a concentration-dependent manner (Fig. 3B). The inhibition curves from both assays overlapped closely, demonstrating that the NGO assay yields results consistent with the standard gel method (Fig. S6).
To further elucidate the mechanism of action, we performed enzyme kinetic studies with varying NTP concentrations. The Michaelis–Menten kinetic analysis revealed that while the maximum velocity (Vmax) remained constant in the presence of fingolimod, the Michaelis constant (Km) exhibited a significant increase (Fig. 3C and D). These kinetic data are consistent with a competitive inhibition mechanism, suggesting that fingolimod may compete with natural nucleotide substrates for the RdRp active site.
3.3. Antiviral activity of fingolimod against HCoV-OC43
Given the structural and functional similarities between SARS-CoV-2 RdRp and those of other betacoronaviruses, we investigated whether fingolimod could inhibit the replication of HCoV-OC43, a human betacoronavirus that causes the common cold. HCoV-OC43 is widely used as a BSL-2 model for studying antiviral agents against SARS-CoV-2.17,18We performed a focus-forming assay (FFA) to evaluate the antiviral efficacy of fingolimod against HCoV-OC43 in Vero E6 cells. Cells were infected with HCoV-OC43 and treated with serial dilutions of fingolimod. Following immunostaining for viral proteins, the number of infectious foci was quantified to determine the extent of viral replication (Fig. 4A). Fingolimod reduced viral replication in a dose-dependent manner, with a half-maximal effective concentration (EC50) of 8.20 µM (Fig. 4B). No cytotoxic effects were observed in Vero E6 cells at concentrations up to 25 µM (Fig. 4C). These results indicate that fingolimod effectively inhibits HCoV-OC43 replication in vitro, supporting its potential as a broad-spectrum antiviral agent targeting betacoronavirus RdRp.
To elucidate the specific stage of inhibition, we performed a time-of-addition assay using HCoV-OC43. While partial inhibition was observed during viral entry (∼40%), fingolimod exhibited significantly stronger antiviral efficacy at the post-entry stage, confirming that its dominant mechanism targets viral replication, consistent with direct RdRp inhibition (Fig. S7).
3.4. Inhibition of SARS-CoV-2 replication by fingolimod
To confirm the antiviral activity of fingolimod against SARS-CoV-2, we conducted experiments at a BSL-3 facility. Vero cells were pre-treated with fingolimod for 1 hour, followed by infection with SARS-CoV-2 (strain βCoV/Korea/KCDC03/2020) at a multiplicity of infection (MOI) of 0.025. After a 48-hour incubation, the cells were fixed and immunostained for the viral nucleocapsid (N) protein (Fig. 5A). Quantitative analysis of infected cells revealed that fingolimod inhibited SARS-CoV-2 infection in a dose-dependent manner, with an EC50 value of 5.73 µM (Fig. 5B). The reduction in viral infection correlated with increased concentrations of fingolimod, demonstrating its effectiveness as an antiviral agent against SARS-CoV-2.To demonstrate the target selectivity of fingolimod, we evaluated its inhibitory effect against an unrelated viral RNA polymerase, the Dengue virus (DENV) RdRp. Unlike SARS-CoV-2 RdRp, fingolimod exhibited no significant inhibition of DENV RdRp activity (Fig. S8). This lack of cross-reactivity confirms that fingolimod does not act through non-specific nucleic acid binding or general interference with polymerase enzymology, but rather possesses selectivity for the SARS-CoV-2 RdRp complex.
The rapid discovery of effective antiviral agents is essential for timely responses to emerging viral outbreaks. Traditional enzyme-based screening methods are often labor-intensive, time-consuming, and not suitable for high-throughput analysis, limiting their utility in urgent situations. In this study, we addressed these challenges by developing an NGO-based RdRp activity assay. This assay exploits the unique properties of NGO, particularly its differential interaction with ssRNA and dsRNA, to enable efficient and direct measurement of RdRp activity. The strong affinity of the NGO for ssRNA results in fluorescence quenching, while its reduced interaction with dsRNA allows fluorescence signals to persist. Validation of the assay demonstrated high reproducibility and reliability, as evidenced by a robust Z′-factor calculation. Utilizing this platform, we screened an FDA-approved small-molecule library to identify potential RdRp inhibitors. Among the molecules tested, we identified fingolimod as a potential inhibitor of SARS-CoV-2 RdRp.
Fingolimod is an immunomodulatory drug approved for the treatment of multiple sclerosis.19 Its known mechanisms include modulation of sphingosine-1-phosphate receptors, which affects lymphocyte trafficking and reduces inflammatory responses. Interestingly, clinical observations have reported that multiple sclerosis patients receiving fingolimod experienced milder COVID-19 symptoms,20,21 suggesting a possible protective effect against SARS-CoV-2 infection. However, prior to our study, potential direct antiviral activity of fingolimod, particularly as an RdRp inhibitor, had not been explored. Our findings reveal that fingolimod directly inhibits SARS-CoV-2 RdRp activity in vitro, suggesting a novel mechanism of action as a direct-acting antiviral agent distinct from its immunomodulatory effects.
In addition, coronaviruses possess a unique 3′-to-5′ exonuclease (ExoN) activity, mediated by nsp14 and activated by nsp10, allowing them to correct replication errors.22 This proofreading mechanism presents a significant challenge for the development of nucleoside analogs (NAs) as antiviral agents, as the erroneous nucleotides introduced by NAs can be removed during replication. In this context, the identification of fingolimod as an RdRp inhibitor offers a promising alternative, as it is structurally distinct from typical NAs and may function independently of ExoN activity.
4. Conclusions
In summary, we developed an NGO-based RdRp activity assay, providing an efficient platform for the discovery of viral RdRp inhibitors. By converting enzyme activity into a fluorescence signal, this assay aims to reduce experimental complexity and increase throughput, making it suitable for high-throughput drug screening. Its application to an FDA-approved drug library led to the identification of fingolimod as a potent direct-acting antiviral candidate by inhibiting SARS-CoV-2 RdRp activity. As the rapid evolution of RNA viruses continues to pose challenges to antiviral therapy, this NGO-based assay may offer a promising approach for accelerating drug discovery efforts, not only against SARS-CoV-2 but also other emerging RNA viruses. By facilitating the rapid identification of viral RdRp-specific inhibitors, our system could contribute to the development of effective treatments at the early stages of the drug discovery process, addressing the ongoing need for versatile and reliable antiviral screening platforms.Author contributions
Hojeong Shin: conceptualization, investigation, formal analysis, visualization, and writing – original draft. Jiwon Woo: investigation, formal analysis, visualization, and writing – original draft. Soeun Yoo: investigation. Soo Min Lee: investigation. So Young Kim: conceptualization, investigation, and formal analysis. Hyung Jin Cha: investigation. Seounghun Kang: investigation. Ji-Seon Lee: conceptualization and investigation. Seho Lee: investigation. Yoo-Bin Kwon: investigation. Jae-Sung Woo: supervision. Dal-Hee Min: conceptualization, supervision, funding acquisition, and writing – review and editing.Conflicts of interest
H. S. and D.-H.M. are named on patents related to this work including a patent for a new use of fingolimod. The remaining authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5tb01764e.Acknowledgements
This work was supported by the Basic Science Research Program (2021R1A2B5B03086506) through the National Research Foundation of Korea (NRF).Notes and references
- B. Hu, H. Guo, P. Zhou and Z. L. Shi, Characteristics of SARS-CoV-2 and COVID-19, Nat. Rev. Microbiol., 2021, 19(3), 141–154 CrossRef CAS.
- J. F. W. Chan, S. F. Yuan, H. Chu, S. Sridhar and K. Y. Yuen, COVID-19 drug discovery and treatment options, Nat. Rev. Microbiol., 2024, 22(7), 391–407 CrossRef CAS.
- K. Dhama, S. K. Patel, K. Sharun, M. Pathak, R. Tiwari and M. I. Yatoo, et al., SARS-CoV-2 jumping the species barrier: Zoonotic lessons from SARS, MERS and recent advances to combat this pandemic virus, Travel Med. Infect. Dis., 2020, 37, 101830 CrossRef PubMed.
- L. Zeng, D. Y. Li, W. D. Tong, T. L. Shi and B. T. Ning, Biochemical features and mutations of key proteins in SARS-CoV-2 and their impacts on RNA therapeutics, Biochem. Pharmacol., 2021, 189, 114424 CrossRef CAS PubMed.
- F. Picarazzi, I. Vicenti, F. Saladini, M. Zazzi and M. Mori, Targeting the RdRp of Emerging RNA Viruses: The Structure-Based Drug Design Challenge, Molecules, 2020, 25(23), 5695 CrossRef CAS PubMed.
- P. Majumdar and S. Niyogi, SARS-CoV-2 mutations: the biological trackway towards viral fitness, Epidemiol. Infect., 2021, 149, e110 CrossRef CAS PubMed.
- M. H. Abbasian, M. Mahmanzar, K. Rahimian, B. Mahdavi, S. Tokhanbigli and B. Moradi, et al., Global landscape of SARS-CoV-2 mutations and conserved regions, J. Transl. Med., 2023, 21(1), 152 CrossRef CAS PubMed.
- Y. Yim, H. Shin, S. M. Ahn and D. H. Min, Graphene oxide-based fluorescent biosensors and their biomedical applications in diagnosis and drug discovery, Chem. Commun., 2021, 57(77), 9820–9833 RSC.
- H. Shin, S. J. Park, J. Kim, J. S. Lee and D. H. Min, A graphene oxide-based fluorescent nanosensor to identify antiviral agents via a drug repurposing screen, Biosens. Bioelectron., 2021, 183, 113208 CrossRef CAS PubMed.
- S. J. Park, J. Kim, S. Kang, H. J. Cha, H. Shin and J. Park, et al., Discovery of direct-acting antiviral agents with a graphene-based fluorescent nanosensor, Sci. Adv., 2020, 6(22), eaaz8201 CrossRef CAS.
- J. Kim, S. J. Park, J. Park, H. Shin, Y. S. Jang and J. S. Woo, et al., Identification of a Direct-Acting Antiviral Agent Targeting RNA Helicase via a Graphene Oxide Nanobiosensor, ACS Appl. Mater. Interfaces, 2021, 13(22), 25715–25726 CrossRef CAS.
- T. A. Nguyen, M. H. Jo, Y. G. Choi, J. Park, S. C. Kwon and S. Hohng, et al., Functional Anatomy of the Human Microprocessor, Cell, 2015, 161(6), 1374–1387 CrossRef CAS PubMed.
- H. S. Hillen, G. Kokic, L. Farnung, C. Dienemann, D. Tegunov and P. Cramer, Structure of replicating SARS-CoV-2 polymerase, Nature, 2020, 584(7819), 154–156 CrossRef CAS PubMed.
- Q. Wang, J. Q. Wu, H. F. Wang, Y. Gao, Q. J. Liu and A. Mu, et al., Structural Basis for RNA Replication by the SARS-CoV-2 Polymerase, Cell, 2020, 182(2), 417–428 CrossRef CAS PubMed.
- J. Chen, B. Malone, E. Llewellyn, M. Grasso, P. M. M. Shelton and P. D. B. Olinares, et al., Structural Basis for Helicase-Polymerase Coupling in the SARS-CoV-2 Replication-Transcription Complex, Cell, 2020, 182(6), 1560–1573 CrossRef CAS PubMed.
- J. H. Zhang, T. D. Y. Chung and K. R. Oldenburg, A simple statistical parameter for use in evaluation and validation of high throughput screening assays, J. Biomol. Screen., 1999, 4(2), 67–73 CrossRef PubMed.
- N. Drayman, J. K. DeMarco, K. A. Jones, S. A. Azizi, H. M. Froggatt and K. Tan, et al., Masitinib is a broad coronavirus 3CL inhibitor that blocks replication of SARS-CoV-2, Science, 2021, 373(6557), 931–936 CrossRef CAS PubMed.
- E. E. Schirtzinger, Y. Kim and A. S. Davis, Improving human coronavirus OC43 (HCoV-OC43) research comparability in studies using HCoV-OC43 as a surrogate for SARS-CoV-2, J. Virol. Methods, 2022, 299, 114317 CrossRef CAS.
- V. Brinkmann, A. Billich, T. Baumruker, P. Heining, R. Schmouder and G. Francis, et al., Fingolimod (FTY720): discovery and development of an oral drug to treat multiple sclerosis, Nat. Rev. Drug Discovery, 2010, 9(11), 883–897 CrossRef CAS.
- G. Mallucci, A. Zito, B. Dal Fabbro and R. Bergamaschi, Asymptomatic SARS-CoV-2 infection in two patients with multiple sclerosis treated with fingolimod, Mult. Scler. Relat. Disord., 2020, 45, 102414 CrossRef PubMed.
- M. Medeiros da Silva, R. Odebrecht de Souza and M. V. Magno Gonçalves, Mild COVID-19 symptoms in a patient with multiple sclerosis in uninterrupted treatment with fingolimod, Neuroimmunol. Rep., 2022, 2, 100071 CrossRef CAS.
- A. Shannon, A. Chazot, M. Feracci, C. Falcou, V. Fattorini and B. Selisko, et al., An exonuclease-resistant chain-terminating nucleotide analogue targeting the SARS-CoV-2 replicase complex, Nucleic Acids Res., 2024, 52(3), 1325–1340 CrossRef CAS PubMed.
Footnote |
| † H. S. and J. W. contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |