Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Grotthuss-type proton transport governed by oxygen coordination environment in VO2 polymorphs
Sunghyun Park a,
Shin-ichi Nishimura
a,
Shin-ichi Nishimura a,
Jinshi Lia,
Muyuan Lia and
Atsuo Yamada
a,
Jinshi Lia,
Muyuan Lia and
Atsuo Yamada *ab
*ab
aDepartment of Chemical System Engineering, School of Engineering, The University of Tokyo, Hongo 7-3-1, Bunkyo-ku, Tokyo 113-8656, Japan. E-mail: yamada@chemsys.t.u-tokyo.ac.jp
bInstitute of Energy Science & Technology (SIEST), Sungkyunkwan University, Suwon 16419, Korea
First published on 24th April 2026
Abstract
Proton-insertion coupled electron transfer (PICET) offers fast-charge energy storage; however, predictive links between oxide crystal structures and proton mobility remain limited. Here, we establish a dynamics-based structure–transport framework by comparing three stoichiometrically identical VO2 polymorphs—VO2(A), VO2(B), and rutile VO2(R)—using molecular dynamics driven by a fine-tuned Universal Model for Atoms (UMA) machine-learned interatomic potential. We show that proton mobility is governed by a hierarchy of structural descriptors: (i) the availability of low-coordination oxygen sites that stabilize proton binding, (ii) the connectivity of reorientation–hopping motifs that enable pathway percolation, and (iii) oxygen–oxygen separation that controls hydrogen-bond-assisted transfer barriers. VO2(A) supports a percolating one-dimensional rotation–direct-hop pathway with single-file-like signatures and the lowest activation energy (≈20.7 kJ mol−1), whereas VO2(B) exhibits anisotropic transport that requires intermittent edge hopping and a higher activation energy (≈39.6 kJ mol−1). The lack of favorable sites and connected motifs in VO2(R) leads to localized proton distributions and strongly suppressed diffusion. These results translate polymorph-dependent PICET behavior into transferable design rules for engineering oxides capable of fast and reversible proton intercalation.
Introduction
Faradaic proton intercalation has emerged as a compelling energy-storage mechanism because it combines the intrinsically fast kinetics of aqueous proton transfer in protic electrolytes with the possibility of framework-stabilized insertion in selected inorganic host lattices operated within appropriate potential and chemical windows.1,2 Unlike protic electrolytes, where transport proceeds through ionic conduction, a non-faradaic process, proton intercalation electrodes operate through the coupled transfer of protons and electrons, known as proton-insertion coupled electron transfer (PICET), within redox-active host lattices.2 Although this coupling is essential for electrochemical operation, the underlying transport properties are ultimately governed by the migration and redistribution of protons within the solid lattice.3,4 Therefore, understanding how these structural characteristics influence proton motion is central to the rational design of proton-intercalation materials.Experimental techniques provide reliable and often quantitative insights into proton incorporation and mobility in solid oxides.5–7 For example, solid-state nuclear magnetic resonance spectroscopy and muon spin spectroscopy can probe proton environments and dynamics over relevant timescales, whereas neutron diffraction and neutron spectroscopy can track structural responses to proton insertion and dynamics, respectively.8,9 Another reasonable starting point is molecular dynamics simulations using empirical force fields, which enable large system sizes and long timescales. However, such models typically lack the fidelity required to describe proton transfer and hydrogen-bond rearrangements, particularly in systems involving transition-metal redox chemistry.10,11 More fundamentally, proton migration involves the making and breaking of O–H bonds and a local electronic response of the host framework, which cannot be captured reliably by fixed-charge descriptions. Ab initio molecular dynamics (AIMD) addresses this limitation by deriving interatomic forces directly from electronic structure calculations and has provided valuable insights into elementary proton transfer events. Nevertheless, its computational cost severely restricts the accessible time scale and system size, limiting the ability to capture long-range diffusion and transport governed by pathway connectivity and collective lattice fluctuations.10,12,13 To overcome these constraints, we employ a machine-learned interatomic potential based on the Universal Model for Atoms (UMA) that is fine-tuned to first-principles reference data for the present VO2–H system. This fine-tuned UMA enables molecular dynamics simulations with near–ab initio accuracy while accessing extended spatiotemporal scales.14–17 In this framework, electronic structure effects are considered implicitly through the training targets (energies and forces), allowing efficient sampling of proton dynamics coupled to lattice fluctuations within the validated chemical and configurational spaces.
As a model oxide system, we focus on three VO2 polymorphs—VO2(A), VO2(B), and VO2(R)—which share the same stoichiometry but differ fundamentally in framework topology and oxygen coordination environment. The selection of these specific phases allows for a systematic comparison between open frameworks (A and B) and a more densely packed structure (R), providing a clear contrast in proton-insertion behavior. Indeed, our previous experimental and computational study showed that electrochemical proton intercalation in VO2 is strongly structure-dependent: VO2(A) and VO2(B) accommodate substantial proton contents approaching HxVO2 (x ≈ 0.8–1.0), whereas VO2(M), a monoclinic form of rutile-type VO2(R), exhibits only subtle proton uptake.18 This contrast was traced to local structural factors, including the availability of less-transition-metal-coordinated oxygen sites, short O–O separations that favor extended O–H⋯O connectivity, and framework flexibility associated with corner-sharing motifs, which together are expected to promote Grotthuss-like transport.19 By comparing these three polymorphs, we aim to resolve how their distinct lattice geometries dictate the transition between efficient long-range diffusion and localized trapping.
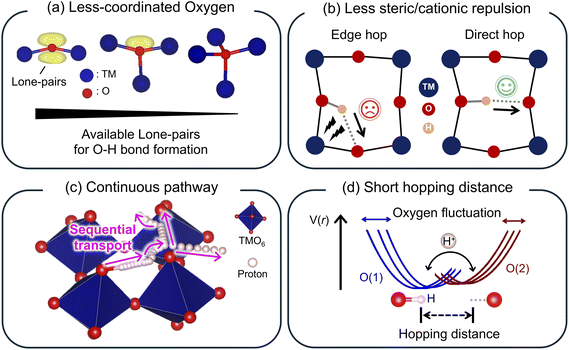
Building on these insights, we move beyond static insertion energetics to explicitly resolve proton dynamics and identify the structural descriptors that control long-range transport. As illustrated schematically in Scheme 1a–d, we examine four key characteristics: (i) local coordination environments of oxygen that define proton-binding sites, (ii) local steric and electrostatic constraints imposed by the atomic arrangements, (iii) pathway connectivity that enables sequential couples of reorientation-hopping, and (iv) the geometric arrangement and spacing of neighboring oxygen atoms that determine feasible hopping geometries and barriers. By directly tracking proton trajectories and concurrent lattice fluctuations across structurally distinct yet compositionally identical VO2 polymorphs, we demonstrate how subtle differences in framework geometry produce qualitatively different transport regimes, ranging from efficient long-range diffusion to anisotropic transport and dynamic trapping. These results provide a dynamics-resolved, structure-based perspective on solid-state proton transport and offer practical design principles for oxide frameworks that support fast and reversible proton intercalation.
Results and discussions
Binding site preference of proton in VO2 polymorphs
Fig. 1a–c compares the crystal structures of VO2(A), VO2(B), and VO2(R), which share VO6 octahedra as common building units but differ markedly in their crystal structural architecture. For clarity, crystallographically distinct oxygen sites are denoted as Oi, with their coordination environments indicated as Oi(nTM), where n indicates the number of transition metal atoms coordinated to each oxygen site. VO2(A) crystallizes in a tetragonal structure composed of dense 2 × 2 × ∞ blocks of corner- and edge-sharing VO6 octahedra. These blocks are interconnected by bridging oxygens, O2(2TM), whereas the block interiors are connected by highly coordinated O1(4TM) sites. VO2(B) adopts a shear-type layered structure consisting of double-layered V4O10 slabs stacked along the c-axis, in which four distinct oxygen sites coexist, including bridging oxygens (O1(2TM)) between slabs and higher-coordinated oxygen sites (O2(4TM), O3(3TM), and O4(3TM)) within the layers. In contrast, VO2(R) forms a rutile-type framework composed of edge-sharing VO6 octahedral chains in which all oxygen atoms are crystallographically equivalent and three-coordinated (O1(3TM)). Notably, VO2(R) lacks less-coordinated oxygen sites such as O(2TM), in sharp contrast to the A- and B-type polymorphs. In contrast, VO2(R) contains only three-coordinated oxygen sites (3TM).In our previous study, density functional theory-based calculations showed that less-coordinated oxygen sites, particularly two-coordinated (2TM) pillar oxygen atoms, are energetically preferred for proton binding, leading to pronounced site selectivity.18,19 These 2TM pillar oxygen atoms possess two lone pairs in an sp3-like electronic configuration, enabling protonation to form hydroxyl species, while simultaneously acting as hydrogen-bond acceptors. This dual role promotes the formation of extended hydrogen-bonding networks within the framework of the material.
This energetic preference is directly reflected in the dynamic proton distributions obtained from molecular dynamics simulations of the protonated VO2 polymorphs. Molecular dynamics simulations at a proton concentration of H1/4VO2 revealed a pronounced structure dependence of both proton binding and diffusion. After thermal annealing at proton-dynamical temperatures of 700 K, protons preferentially reside near the energetically favorable O(2TM) atoms in H1/4VO2(A) and H1/4VO2(B). Notably, in H1/4VO2(A), proton binding is essentially confined to the O(2TM) sites, whereas H1/4VO2(B) exhibits a measurable contribution from the O(3TM) site even at the low proton concentration of 1/4, consistent with its more diverse oxygen local environments (Fig. 1d and e). In contrast, VO2(R) lacks proton-binding preferences owing to the symmetrical equivalence of its oxygen atoms (Fig. 1f). Collectively, these results demonstrate that the local environment of oxygen atoms, specifically the transition metal coordination number, governs the dynamic distribution of protons among the binding sites, providing a unified framework for understanding the site-dependent proton behavior in VO2 polymorphs.
Anisotropic proton diffusion in VO2 polymorphs
Direction-resolved mean squared displacements (MSDs) of protons, obtained from molecular dynamics simulations in the temperature range of 550–780 K, exhibit significant anisotropy in proton dynamics, reflecting the intrinsic anisotropy of their crystal frameworks (Fig. 2 and S1–S3). For all polymorphs, the MSD magnitude systematically increases with increasing temperature, consistent with thermally activated proton motion. In H1/4VO2(A), proton transport is essentially one-dimensional along the z-axis (Fig. 2a). The MSD increases nearly linearly at short lag times before transitioning to a sub-linear regime, a hallmark of single-file diffusion commonly observed in narrow one-dimensional channels where mobile species cannot bypass one another.20–22 H1/4VO2(B) also displays anisotropic proton diffusion, with a dominant contribution along the y-direction and a secondary component along thex-direction, consistent with proton migration through multiple interconnected diffusion pathways (Fig. 2b). In contrast, H1/4VO2(R) exhibits a markedly smaller MSD in the same time window (Fig. 2c), indicating strongly suppressed proton mobility. These results demonstrate that the presence and dimensionality of proton-conducting channels—one-dimensional in VO2(A) and quasi-two-dimensional in VO2(B)—critically govern proton transport, whereas their absence in VO2(R) leads to suppressed proton transport. The systematic trend in proton diffusivity is in good agreement with experimental trend in electrochemical capacity and rate-capability in our previous study.18Grotthuss-type hopping mechanism in solid oxides
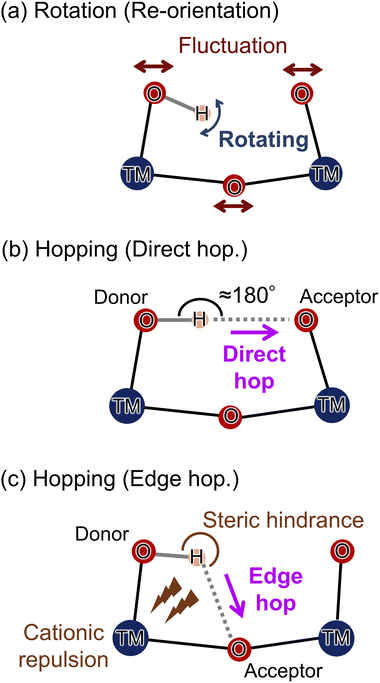
In solid oxides, proton migration proceeds through a coupled sequence of O–H bond reorientation (local hydroxyl rotation) and inter-oxygen proton transfer (hopping), and long-range transport emerges only when this reorientation–hopping motif percolates through the entire structure to form a continuous pathway (Scheme 2).23–25 When neighboring oxygen atoms can adopt a favorable O–H⋯O geometry through reorientation of the O–H bond, proton transfer occurs via a direct hop along a near-linear hydrogen bond. This process exhibits a low activation barrier because a near-linear O–H⋯O alignment maximizes the directional overlap between the acceptor's lone pair and the O–H σ* orbital.26,27 In contrast, when such alignment is geometrically inaccessible, proton motion proceeds via an edge-hopping route that circumvents a metal–oxygen polyhedron; this pathway is typically penalized by steric constraints and short-range cation-proton electrostatic repulsion, leading to a higher barrier.28In this context, MD simulations show that proton migration in VO2 polymorphs is governed by the ability of each crystal structure to sustain connected rotation–hopping pathways, as visualized by proton density distributions and quantified by the corresponding free-energy landscapes for protons (Fig. 3). These results demonstrate that the proton mobility in VO2 polymorphs is fundamentally governed by the arrangement of the oxygen network in each crystal structure, which supports continuous rotation–hopping pathways.
Among the three polymorphs, VO2(A) supports continuous long-range proton transport along a one-dimensional wavy channel (Fig. 3a) defined by an alternating zigzag array of O2(2TM) atoms. Protons preferentially bind to these O2(2TM) sites, and the resulting hydroxyl O–H units reorient toward the neighboring O2(2TM) oxygen atoms that serve as hydrogen-bond acceptors, thereby forming favorable O–H⋯O configurations. Proton migration proceeds via a repetitive sequence of local O–H reorientation, which renews the hydrogen-bond geometry, followed by direct proton transfer to the adjacent oxygen site (Fig. 3b). Consistent with this mechanism, the free-energy profile exhibits a low activation barrier for the direct hop (≈6.8 kJ mol−1; 0.07 eV), enabling uninterrupted reorientation-hopping events and thus, efficient one-dimensional proton conduction (Fig. 3c).
In contrast, the energy landscape in VO2(B) is fragmented by the coexistence of multiple oxygen environments (Fig. 3d). Although VO2(B) also utilizes the reorientation-transfer motif, the locally feasible direct hops cannot be concatenated into a long-range percolating network because of the lack of successive favorable O–H⋯O configurations. The most stable O1–H⋯O1 configurations form a relatively deep double-well, trapping the protons (Fig. 3d). Long-range transport requires intermittent edge-hopping events that bridge preferential proton-trapping motifs, with O4(3TM) serving as the key connector (Fig. 3e). The OH reorientation toward this key connector is the bottleneck step with a moderate free-energy barrier of 22.7 kJ mol−1 (0.25 eV) (Fig. 3f), which is higher than that in VO2(A), but still allows for appreciable conduction.
The transport limitations are even more pronounced in VO2(R), where the proton density remains highly localized between adjacent O1(3TM) sites, indicating a lack of long-range connectivity (Fig. 3g). In this rutile framework, macroscopic transport must rely almost exclusively on edge-hopping events rather than a facile direct-hopping network (Fig. 3h). Consistent with this picture, the free-energy profile exhibits substantially higher energy barriers of 45.0 kJ mol−1 (0.47 eV), resulting in suppressed proton diffusion, even at 700 K (Fig. 3i).
These distinct energetic landscapes for proton conduction are directly translated into macroscopic transport kinetics, as evidenced by the Arrhenius analyses of the self-diffusion coefficients (Fig. 4 and S1–S3). The calculated activation energies (Ea) follow the hierarchy of the bottleneck barriers in the free-energy landscapes. To further examine the geometric constraints, we analyzed the statistical distribution of O–O distances (dO–O) from the MD trajectories (Fig. S4). According to the classical Lippincott–Schroeder (LS) model, shortening the donor–acceptor separation monotonously lowers the energy barrier for proton transfer and approaches zero at the critical distance for hydrogen-bond symmetrization.29 In this context, the edge-hops are expected to show lower barrier because they tend to have shorter dO–O than the direct-hops, as evidenced by the statistical distribution in Fig. S4. However, edge-hops are inherently unfavorable with high barrier because of their geometrically unstable hydrogen environment, where the hydrogen bond is far from collinear with the OH-bond and is imposed with steric penalties, as discussed above (Scheme 2c).
Taken together, these results show that short O–O separation is a necessary but insufficient descriptor for proton transfer in redox-active oxides: short donor–acceptor distances facilitate low-barrier transfer only when the local geometry supports a near-linear O–H⋯O configuration. Accordingly, the highest mobility in VO2(A) arises from a percolating network of short O–O pairs that enable successive direct hops, whereas VO2(B) and VO2(R) are limited by non-percolating connectivity and edge-hopping steps that are geometrically frustrated despite comparable O–O distances. This framework reconciles the Arrhenius activation energies with the free-energy bottlenecks and highlights pathway connectivity and hydrogen-bond geometry, in addition to dO–O, as decisive design criteria for fast solid-state proton transport.
Conclusion
An atomistic understanding of proton conduction in VO2 polymorphs—VO2(A), VO2(B), and VO2(R)— was established by leveraging machine-learning interatomic potential molecular dynamics simulations. Among the three polymorphs, VO2(A) unequivocally exhibits the highest proton conductivity, sustained by a continuous hydrogen-bond network consisting of less-coordinated oxygen sites. VO2(B) supports appreciable but anisotropic proton transport, where conduction is intermittently enabled through edge-sharing pathways, whereas VO2(R) is intrinsically unfavorable for proton migration owing to the absence of suitable oxygen coordination environments and extended hydrogen-bond connectivity. Across all polymorphs, proton migration follows a Grotthuss-type mechanism involving coupled O–H bond reorientation and proton transfer, which is accommodated by the lattice flexibility. Proton mobility in oxides is largely influenced by the local coordination environment, steric and electrostatic constraints, and—most critically—the continuity of hydrogen-bond-assisted hopping pathways, even with an identical chemical composition.Conflicts of interest
There are no conflicts to declare.Data availability
All data generated or analyzed during this study are included in this published article and its associated supplementary information (SI). Any additional raw data that support the findings of this study are available from the corresponding author upon reasonable request. Supplementary information is available. See DOI: https://doi.org/10.1039/d6ta01847e.Acknowledgements
This work was supported by the Japan Science and Technology Agency (JST) CREST Grant Number JPMJCR21O6. Part of the computational work was performed using the facilities of the Supercomputer Center, Institute for Solid State Physics, University of Tokyo.References
- D. G. Nocera, J. Am. Chem. Soc., 2022, 144, 1069–1081 CrossRef CAS PubMed.
- N. P. Holzapfel, V. Augustyn and V. Balland, ACS Energy Lett., 2025, 10, 1143–1164 CrossRef CAS.
- H. Guo and C. Zhao, Small Methods, 2024, 8, 2300699 CrossRef CAS PubMed.
- C. Huang, W. Zhang and W. Zheng, Energy Storage Mater., 2023, 61, 102913 CrossRef.
- H. Ahn, D. Kim, M. Lee and K. W. Nam, Commun. Mater., 2023, 4, 37 CrossRef.
- Y. Liu, Y. Zhu and Y. Cui, Nat. Energy, 2019, 4, 540–550 CrossRef.
- S. Wu, H. Guo and C. Zhao, Adv. Funct. Mater., 2024, 34, 2405401 CrossRef CAS.
- J. K. Park, H.-J. Kwon and C. E. Lee, Sci. Rep., 2016, 6, 23378 CrossRef CAS.
- F. L. Pratt, P. J. Baker and S. J. Blundell, J. Phys.:Conf. Ser., 2023, 2462, 012044 CrossRef CAS.
- E. Paquet and H. L. Viktor, Adv. Chem., 2018, 2018, 9839641 Search PubMed.
- D. W. M. Hofmann, L. N. Kuleshova and B. D'Aguanno, J. Power Sources, 2010, 195, 7743–7750 CrossRef CAS.
- R. Vuilleumier and D. Borgis, J. Chem. Phys., 1999, 111, 4251–4266 CrossRef CAS.
- R. Iftimie, P. Minary and M. E. Tuckerman, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 6654–6659 CrossRef CAS PubMed.
- O. T. Unke, S. Chmiela, H. E. Sauceda, M. Gastegger, I. Poltavsky, K. T. Schütt, A. Tkatchenko and K.-R. Müller, Chem. Rev., 2021, 121, 10142–10186 CrossRef CAS PubMed.
- D. M. Anstine and O. Isayev, J. Phys. Chem. A, 2023, 127, 2417–2431 CrossRef CAS PubMed.
- B. Mortazavi, X. Zhuang, T. Rabczuk and A. V. Shapeev, Mater. Horiz., 2023, 10, 1956–1968 RSC.
- M. Grunert, M. Großmann, J. Hänseroth, A. Flötotto, J. Oumard, J. L. Wolf, E. Runge and C. Dreßler, J. Phys. Chem. C, 2025, 129, 9662–9669 CrossRef CAS.
- S. Park, S. Nishimura, J. Li, A. Kitada, S. Lee, W.-S. Yoon and A. Yamada, J. Am. Chem. Soc., 2026, 148, 7931–7940 CrossRef CAS PubMed.
- S. Park, S. Nishimura, A. Kitada and A. Yamada, ACS Appl. Energy Mater., 2024, 7, 4347–4352 CrossRef CAS.
- A. Taloni and F. Marchesoni, Phys. Rev. Lett., 2006, 96, 020601 CrossRef.
- K. Hahn and J. Kärger, J. Phys. Chem., 1996, 100, 316–326 CrossRef CAS.
- X. Yang, M. Wu, Z. Qin, J. Wang and T. Wen, J. Appl. Phys., 2009, 106, 084905 CrossRef.
- K.-D. Kreuer, Chem. Mater., 1996, 8, 610–641 CrossRef CAS.
- K. D. Kreuer, Annu. Rev. Mater. Res., 2003, 33, 333–359 CrossRef CAS.
- K. D. Kreuer, Solid State Ionics, 1997, 97, 1–15 CrossRef CAS.
- N. Agmon, Chem. Phys. Lett., 1995, 244, 456–462 CrossRef CAS.
- S. Scheiner, in Intermolecular Interactions in Crystals: Fundamentals of Crystal Engineering, ed. J. J. Novoa, The Royal Society of Chemistry, Cambridge, 2017, pp. 410–452. Search PubMed.
- M. S. Islam, S. Wang, A. T. Hall and Y. Mo, Chem. Mater., 2022, 34, 5938–5948 CrossRef CAS.
- E. R. Lippincott and R. Schroeder, J. Chem. Phys., 1955, 23, 1099–1106 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |