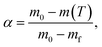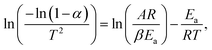Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Ribonucleic acid as an intrinsically synergistic bio-derived flame retardant
Sang Yun Jeong†
a,
Jong Seok Lim† a,
Abdullah Malika,
Jae Chul Roa,
Hui Hun Cho*ab,
Jun Hyuk Heo
a,
Abdullah Malika,
Jae Chul Roa,
Hui Hun Cho*ab,
Jun Hyuk Heo *ab and
Jung Heon Lee
*ab and
Jung Heon Lee *abcd
*abcd
aSchool of Advanced Materials Science and Engineering, Sungkyunkwan University (SKKU), Suwon 16419, South Korea. E-mail: saegusa@skku.edu; jhlee7@skku.edu
bResearch Center for Advanced Materials Technology, SKKU, Suwon 16419, South Korea
cBiomedical Institute for Convergence at SKKU, SKKU, Suwon 16419, South Korea
dSKKU Advanced Institute of Nanotechnology, SKKU, Suwon 16419, South Korea. E-mail: studisthun@skku.edu
First published on 11th May 2026
Abstract
Ribonucleic acid (RNA) was investigated as an intrinsically synergistic biomolecular flame retardant, exhibiting distinct thermal behavior and an intumescent char-forming ability. Despite the growing interest in bio-derived flame retardants, the flame-retardant potential of RNA has not been previously explored. Owing to its intrinsically high carbon, nitrogen, and phosphorus content, together with the presence of uracil and a 2′-hydroxyl group, RNA undergoes efficient dehydration and promotes effective intumescent char formation during combustion. Compared with the structurally similar deoxyribonucleic acid (DNA), RNA exhibited distinct thermal transitions and superior char yield. When applied as a coating on polyurethane foam (PUF), RNA increases the residual mass from 0.8% to 37.5%, elevates the limiting oxygen index (LOI) to 29.3%, and enables a UL-94 HF-1 classification. Cone calorimetry further shows a 49.4% reduction in peak heat release rate (p-HRR). These findings demonstrate that RNA functions as a sustainable and inherently synergistic bio-derived flame retardant by suggesting combined condensed-phase protection with likely gas-dilution/gas-phase contributions.
1 Introduction
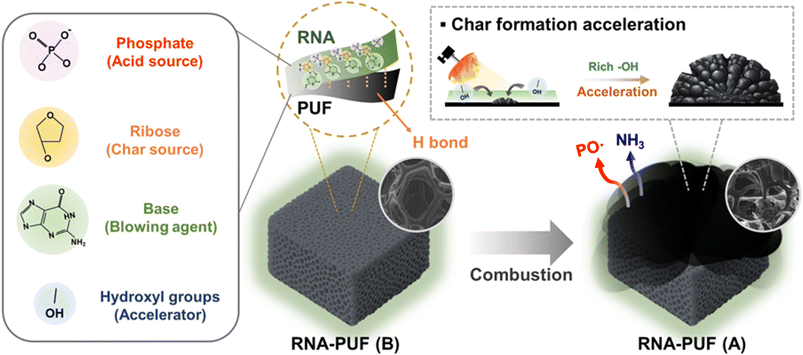
The tightening of global fire-safety standards and environmental regulations has drawn increasing attention to the limitations of conventional halogenated and high-loading phosphorus-based flame retardants (FRs).1–4 Although these materials have been widely used to suppress polymer flammability, their long-term viability is increasingly questioned due to the generation of corrosive and toxic combustion byproducts, as well as the degradation of polymer properties at high additive loadings. These limitations are particularly critical for lightweight foams and mechanically sensitive engineering polymers, underscoring the need for flame-retardant strategies that combine environmental safety, material compatibility, and high fire-suppression efficiency.5–7In response to these challenges, biomolecule-derived flame retardants (Bio-FRs) have attracted increasing attention as sustainable materials.8–10 Their inherent biocompatibility, renewable origin, and intrinsically optimized molecular structures enable functionalities that conventional FR systems cannot readily achieve.11,12 Many biomolecules contain carbon-, nitrogen-, and phosphorus-rich domains that allow them to operate as intrinsically self-contained intumescent systems. Phosphate groups initiate dehydration and promote condensed-phase char formation; polysaccharide or polymeric backbones generate thermally stable carbonaceous layers; and nitrogen-containing moieties release non-combustible gases that dilute flammable volatiles and inhibit flame propagation.13,14 These synergistic, molecularly embedded mechanisms allow Bio-FRs to deliver flame-retardant performance at relatively low additive loadings, providing a mechanistically distinct platform for environmentally benign fire-safety technologies.15–18
Molecules such as deoxyribonucleic acid (DNA) and adenosine triphosphate (ATP) uniquely integrate the essential components of an intumescent system, including an acid source, carbon source, and blowing agent, within a single molecular architecture.18–21 In the case of DNA, the phosphate backbone generates phosphoric acids that catalyze dehydration, while the deoxyribose moiety carbonizes to form a protective char layer. Meanwhile, the nucleobases release inert and non-flammable gases that promote char expansion and dilute combustible volatiles.19 These multifunctional characteristics have established DNA as a benchmark biomolecule-derived flame retardant (Bio-FR).
However, despite the progress enabled by DNA-based systems, inherent structural constraints suggest opportunities to identify alternative biomolecules with more favorable thermal stability and carbonization behavior. In particular, the presence of methylated thymine (T) in DNA facilitates early N-glycosidic bond cleavage and volatilization into low-molecular-weight fragments during thermal decomposition, resulting in premature degradation and reduced char yield.22
Ribonucleic acid (RNA), although traditionally regarded as a genetic information carrier rather than a functional material, has recently been re-evaluated for its diverse functional roles in bioengineering. RNA-based structure-switching aptamers and light-up sensors have demonstrated exceptional performance in detecting viral proteins and enzymatic activities.23–26 Beyond sensing, RNA-based platforms integrated with nanomaterials have advanced microRNA diagnostics and therapeutic applications through precisely engineered probe designs.27,28 Furthermore, advanced microfluidic systems for mRNA-encapsulated lipid nanoparticles underscore the growing clinical importance of RNA in gene delivery and vaccine development.29–31
Expanding this functional paradigm beyond biotechnological applications, we demonstrate that RNA possesses a molecular configuration inherently favorable for intumescent flame retardancy. Similar to other biomolecule-derived flame retardants, RNA intrinsically integrates the essential components of an intumescent system, including an acid source, carbon source, and blowing agent. In particular, RNA contains a higher density of hydroxyl groups in its ribose units than DNA. The presence of a 2′-OH group enhances dehydration-driven carbonization and facilitates the early formation of cohesive char structures. Furthermore, the substitution of uracil (U) for methylated thymine suppresses volatilization pathways that limit the char yield of DNA.32 The polyanionic phosphate backbone of RNA functions as a strong intrinsic acid source, while its nitrogen-containing nucleobases contribute to gas-phase flame inhibition. In addition, the abundant hydroxyl groups of RNA improve interfacial adhesion and coating uniformity on polymer substrates such as polyurethane foam.33 Despite these distinct structural advantages, RNA has not yet been explored as a flame-retardant material, leaving a notable gap in the rapidly expanding field of Bio-FRs.
Here, we systematically investigate the combustion behavior, condensed-phase carbonization, and flame-suppression performance using polyurethane foam as a representative flammable model substrate.34 By demonstrating the advantages of RNA relative to existing biomolecule-derived systems, this work expands the molecular design space for sustainable flame-retardant technologies and highlights the potential of RNA for future fire-safe material applications.
2 Experimental section
2.1 Materials
Thymine (T), thymidine (Thd), uracil (U), and uridine (Urd) were purchased from Tokyo Chemical Industry (Tokyo, Japan). Deoxyribonucleic acid (DNA, low molecular weight, salmon sperm) was obtained from Sigma-Aldrich (St. Louis, MO, USA), and ribonucleic acid (RNA, Cat. No. 10109223001) was purchased from Roche Diagnostics GmbH (Mannheim, Germany). Deionized (DI) water with a resistivity of 18.2 MΩ·cm was prepared using an Arium® Pro Ultrapure Water System (Sartorius, Germany). Commercial polyether-based polyurethane foam (PUF) (open-cell, 100 pores per inch (ppi), 16.2 kg m−3) was purchased from Wonchang Corporation (Gyeonggi, Korea).2.2 Preparation of nucleic acid solutions
Aqueous solutions of RNA and DNA were prepared at concentrations of 2.5, 5.0, 7.5, and 10 wt% by dissolving the corresponding powders in deionized water. The appropriate amount of RNA or DNA powder was gradually added to preheated deionized water maintained at 55 °C under continuous stirring at 200 rpm. Stirring and temperature were maintained for 2 hours until complete dissolution was achieved.2.3 Preparation of nucleic acid-coated PUF
PUF was cut into the required dimensions for subsequent analyses. The volume of the nucleic acid solution was first adjusted to a 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 volume-to-weight (v/w) ratio based on the mass of the pre-weighed PUF, and subsequently, the foam samples were immersed in the solution for 2 h. After impregnation, the samples were incubated at 50 °C for 6 h to facilitate adsorption of the nucleic acids, followed by oven drying at 50 °C for an additional 3 h. The final mass of each sample was measured to determine the add-on percentage of the nucleic acids on PUF. These measurements were performed on five independent samples to ensure accuracy. Furthermore, to evaluate the FR performance and environmental durability of the coated PUF under various conditions, ambient aging, heat aging, and water immersion tests were conducted. For ambient aging, the samples were conditioned at 25 ± 1 °C with a relative humidity of 20 ± 5% for 168 h. For heat aging, the samples were maintained at 125 °C for 16 h, in accordance with the ISO 2440
1 volume-to-weight (v/w) ratio based on the mass of the pre-weighed PUF, and subsequently, the foam samples were immersed in the solution for 2 h. After impregnation, the samples were incubated at 50 °C for 6 h to facilitate adsorption of the nucleic acids, followed by oven drying at 50 °C for an additional 3 h. The final mass of each sample was measured to determine the add-on percentage of the nucleic acids on PUF. These measurements were performed on five independent samples to ensure accuracy. Furthermore, to evaluate the FR performance and environmental durability of the coated PUF under various conditions, ambient aging, heat aging, and water immersion tests were conducted. For ambient aging, the samples were conditioned at 25 ± 1 °C with a relative humidity of 20 ± 5% for 168 h. For heat aging, the samples were maintained at 125 °C for 16 h, in accordance with the ISO 2440![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2019 (E) standard. For the immersion test, the samples were fully submerged in tap water for 24 h and tested only after complete drying. The resulting nucleic acid-coated PUF samples were then used for subsequent characterization and flammability tests.
2019 (E) standard. For the immersion test, the samples were fully submerged in tap water for 24 h and tested only after complete drying. The resulting nucleic acid-coated PUF samples were then used for subsequent characterization and flammability tests.
2.4 Characterization and measurements
The conversion degree (α) was calculated from the TGA mass loss data according to:
For a first-order reaction, the Coats–Redfern equation can be expressed as:
where A is the pre-exponential factor, R is the gas constant, β is the heating rate, and Ea is the activation energy. The activation energy was determined from the slope of the linear fit of ln[−(1−α)/T2] versus 1/T, according to:
| Ea = −(slope) × R. |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2012 standard to assess the flame retardancy of the PUF specimens with dimensions of 150 × 50 × 10 mm3. In accordance with the protocol for cellular plastics, the samples were placed horizontally and subjected to a flame with a height of 38 mm for 60 s at one end. Burning behavior was recorded using an infrared (IR) camera (TG297, FLIR systems, Inc).
2012 standard to assess the flame retardancy of the PUF specimens with dimensions of 150 × 50 × 10 mm3. In accordance with the protocol for cellular plastics, the samples were placed horizontally and subjected to a flame with a height of 38 mm for 60 s at one end. Burning behavior was recorded using an infrared (IR) camera (TG297, FLIR systems, Inc).![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2017 standard to determine the minimum oxygen concentration required for sustained combustion. Each sample was tested 5 times, and average values were reported.
2017 standard to determine the minimum oxygen concentration required for sustained combustion. Each sample was tested 5 times, and average values were reported.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2015 standard. Specimens (100 × 100 × 10 mm3) were mounted horizontally in a specimen holder using a retainer frame without a grid and then exposed to an external heat flux of 35 kW m−2. Key parameters, including time to ignition (TTI), heat release rate (HRR), total heat released (THR), total smoke release (TSR), effective heat of combustion (EHC), and char yield were recorded.
2015 standard. Specimens (100 × 100 × 10 mm3) were mounted horizontally in a specimen holder using a retainer frame without a grid and then exposed to an external heat flux of 35 kW m−2. Key parameters, including time to ignition (TTI), heat release rate (HRR), total heat released (THR), total smoke release (TSR), effective heat of combustion (EHC), and char yield were recorded.3 Results and discussion
3.1 Structural origin of the distinct thermal properties between RNA and DNA
Both RNA and DNA share a phosphate–sugar-base backbone and contain the same nucleobases, adenine (A), guanine (G), and cytosine (C). Their structural differences arise primarily from the substitution of uracil (U) for thymine (T) in RNA, as well as from the presence of an additional 2′-OH group on the ribose units of RNA (Fig. S1). To examine the molecular similarities and distinctions between the two nucleic acids, Fourier transform infrared (FTIR) spectroscopy was performed (Fig. S2).The in-plane ring vibrations of A, G, C, and U were observed at approximately 1690 cm−1, while the characteristic cytosine ring vibration appeared at 1466 cm−1. Stretching modes associated with C–N, C–H, and N–H were detected at 1413 cm−1. The asymmetric and symmetric stretching vibrations of PO2− and P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O groups were identified at 1222 and 1059 cm−1, respectively. The C–O stretching vibration of the ribose moiety appeared at 963 cm−1, and the broad band between 878 and 821 cm−1 was assigned to ribose–phosphate backbone vibrations.
O groups were identified at 1222 and 1059 cm−1, respectively. The C–O stretching vibration of the ribose moiety appeared at 963 cm−1, and the broad band between 878 and 821 cm−1 was assigned to ribose–phosphate backbone vibrations.
Notably, the only distinguishable difference between the FTIR spectra was observed in the 2875–2960 cm−1 region, corresponding to methyl C–H stretching vibrations that appear exclusively in DNA due to the presence of thymine.35,36 These FTIR results confirm a high degree of molecular structural similarity between RNA and DNA.
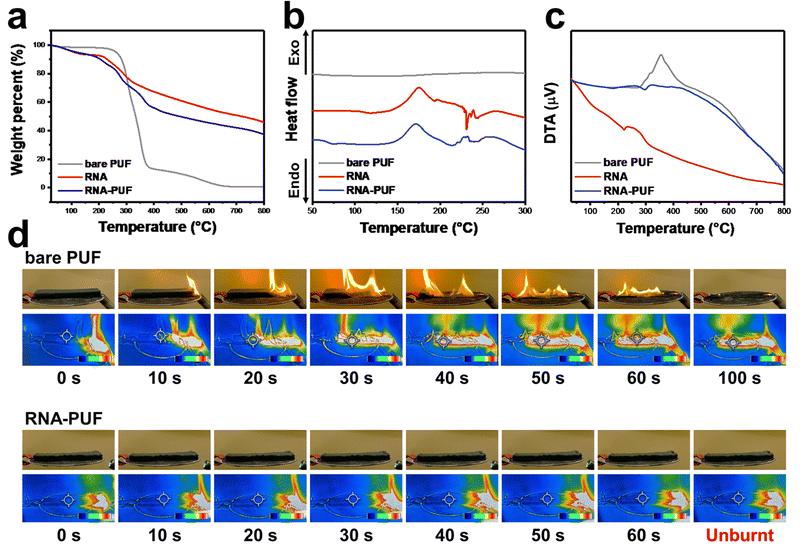
Despite this high structural similarity, RNA exhibits distinct thermal degradation behavior compared to DNA, as revealed by TGA (Fig. 1a). The thermal degradation profiles of RNA and DNA begin to diverge near 200 °C, and the final char residues reach 46.0% and 43.6% for RNA and DNA, respectively.
To elucidate the molecular origin of this behavior, TGA was further performed on uridine (Urd) and thymidine (Thd), the corresponding nucleosides of RNA and DNA, respectively (Fig. 1d). Unlike the isolated nucleobases U and T, which exhibit only subtle differences in thermal behavior (Fig. S3a), Urd and Thd show a clear divergence in thermal degradation profiles, reflecting synergistic interactions between the sugar moiety and the nucleobase. Urd exhibits markedly enhanced thermal stability, with major decomposition initiating near 300 °C and yielding a substantial char residue of 13.1%. In contrast, Thd undergoes rapid volatilization and leaves negligible residue. This contrasting behavior arises from the presence of the 2′-OH group in Urd, which promotes dehydration, intermolecular cross-linking, and early-stage carbonization. Conversely, the methyl group in thymine facilitates premature N-glycosidic bond cleavage and volatilization during thermal decomposition, thereby suppressing char formation.
Further differences between RNA and DNA are revealed from DSC analysis (Fig. 1b). RNA exhibits a distinct exothermic transition at approximately 170 °C, which is attributed to the molecular rearrangement and cross-linking reactions between the 2′-OH groups of the ribose units and adjacent phosphate species. The close spatial proximity of these groups in RNA lowers the activation barrier for condensation, thereby facilitating early-stage network formation. Subsequently, RNA exhibits a sharp endothermic peak at approximately 230 °C, corresponding to the cleavage of C–N bonds and partial structural decomposition. The resulting fragments then undergo phosphate-assisted recombination and further condensation, giving rise to a secondary exothermic peak near 240 °C. In contrast, DNA displays significantly weaker and broader thermal transitions between 130 and 230 °C, followed by a sharp endothermic peak at approximately 240 °C. These observations suggest that DNA undergoes less efficient dehydration and delayed carbonization compared to the rapid, pre-activated charring mechanism of RNA.
To further clarify the origin of these differences, DSC analyses of Urd and Thd were performed (Fig. 1e). Urd and Thd exhibit endothermic peaks at approximately 170 and 190 °C, respectively, corresponding to their melting transitions, which for Urd are accompanied by onset of 2′-OH-mediated dehydration-condensation as evidenced by the subsequent char-forming behavior in TGA. Following melting, the presence of the 2′-OH groups in Urd facilitates intermolecular cross-linking reactions, which act as precursor steps toward char formation. Consistent with this trend, the isolated nucleobases U and T also exhibit distinct DSC behaviors, with U showing a stronger endothermic response than T (Fig. S3b).
Differential thermal analysis (DTA) further reinforces these trends (Fig. 1c). RNA exhibits a pronounced endothermic peak at approximately 200 °C, followed by a broad exothermic transition, consistent with early dehydration and enhanced condensed-phase char formation. In contrast, DNA displays weaker endothermic behavior accompanied by delayed exothermic activity, suggesting less efficient dehydration and slower carbonization. To further elucidate the origin of these differences, DTA analyses of Urd and Thd were conducted (Fig. 1f). Urd exhibits an earlier endothermic peak at around 170 °C, which can be attributed to dehydration-associated structural reorganization. In comparison, Thd shows an endothermic peak near 200 °C, likely related to methyl-group-assisted bond cleavage and volatilization. DTA analyses of U and T also reveal distinct thermal transition behaviors (Fig. S3c). U exhibits a pronounced endothermic peak at approximately 340 °C, whereas T shows a corresponding peak at a lower temperature near 325 °C. This earlier peak of T is associated with methyl group-induced bond cleavage and volatilization. Collectively, these results indicate that the absence of the methyl group in U inherently contributes to enhanced thermal stability and promotes more favorable char formation pathways during combustion. Consequently, this structural difference demonstrates the superior flame-retardant performance of RNA compared to DNA.
To further quantitatively assess the activation energy required for dehydration and carbonization of RNA and DNA, non-isothermal kinetic analysis was performed by dividing the thermal degradation process into two characteristic stages (Table 1). The first stage corresponds to dehydration in the temperature range from 150 to 200 °C, with a conversion range of 0.050–0.948. The second stage represents the onset of carbonization occurring between 200 and 250 °C, accompanied by a conversion range of 0.052–0.950. During the dehydration stage, RNA exhibited a markedly lower activation energy of 159.6 kJ mol−1 compared to 364.6 kJ mol−1 for DNA, corresponding to an approximate 56% reduction in the energy barrier for thermal degradation. In the carbonization onset stage, RNA exhibited a lower activation energy (267.5 kJ mol−1) compared to DNA (339.7 kJ mol−1), representing an approximate 21% reduction. The pronounced reductions in activation energy observed during the two stages highlight the critical role of the hydroxyl groups in RNA in facilitating early dehydration and phosphate-assisted condensation reactions. These kinetic results further demonstrate that the structural features unique to RNA promote more energetically favorable pathways toward dehydration-driven carbonization and char formation.
| Stage | Sample | Conversion range (α) | R2 | Slope | Activation energy (kJ mol−1) |
|---|---|---|---|---|---|
| I | RNA | 0.050–0.948 | 0.991 | −19194.31 | 159.6 |
| DNA | 0.993 | −43857.53 | 364.6 | ||
| II | RNA | 0.052–0.950 | 0.996 | −32179.85 | 267.5 |
| DNA | 0.994 | −40853.71 | 339.7 |
To further compare the inherent char-forming ability between RNA and DNA, a pellet combustion test was conducted (Fig. 1g and Video S1). Upon exposure to flame, the RNA pellet rapidly expanded and developed into a highly swollen, elongated, and long worm-like intumescent char structure. This efficient intumescent char formation originates from the intrinsic coexistence of carbon, nitrogen, and phosphorus within the RNA, which function synergistically as the char source, blowing agent, and acid source, respectively, in an intumescent system.1,37 When evaluated under identical conditions, DNA produced a relatively lower char residue with limited intumescent char development. This distinctive char-forming behavior is due to the structural differences. The abundant 2′-OH groups in RNA actively promote early dehydration and robust intermolecular cross-linking, facilitating the rapid formation of a highly flexible carbonaceous framework. Furthermore, the absence of a methyl group in uracil suppresses premature volatilization during combustion.22,38
The structural superiority of the RNA-derived char was further corroborated by Raman spectroscopy (Fig. 1h). The spectra exhibit two characteristic peaks, the D-band at approximately 1350 cm−1, associated with disordered or defective carbon structures, and the G-band at approximately 1580 cm−1, corresponding to the sp2-hybridized graphitic carbon network. The intensity ratio of the D-band to the G-band (ID/IG) serves as a reliable indicator of the degree of graphitization. RNA and DNA chars display ID/IG ratios of 2.66 and 3.56. The significantly lower ratio of RNA than DNA demonstrates that RNA forms a relatively more graphitized and structurally more robust carbonaceous layer compared to DNA char. The superior condensed-phase stabilization of RNA can be attributed to its abundant 2′-OH groups, which facilitate efficient dehydration and intensive cross-linking, yielding a dense and thermally stable protective char barrier.
Overall, RNA exhibits intrinsically enhanced thermal endurance and char-forming capacity compared to DNA, despite their similar molecular frameworks. These advantages originate from key structural features unique to RNA, namely the presence of the 2′-OH group, which accelerates dehydration and promotes the formation of cross-linked char structures, and the absence of a methyl group in uracil, which suppresses premature volatilization during thermal decomposition. As a result, RNA achieves enhanced condensed-phase stabilization, positioning it as a structurally optimized, next-generation bio-derived flame-retardant molecule.
3.2 Chemical-state evolution of the RNA-driven char
To clarify the char-forming mechanism of RNA, wide-scan XPS was performed before and after combustion (Fig. 2a). After burning, the carbon content increased markedly from 49.61% to 57.32%, while the nitrogen content increased from 14.01% to 16.74%, indicating substantial carbonization and enrichment of nitrogen-containing species within thermally stable condensed structures. In contrast, the oxygen content decreased from 32.05% to 21.77%, consistent with extensive dehydration during combustion. Notably, the phosphorus content remained nearly unchanged, suggesting that phosphate moieties were retained in the condensed phase rather than volatilized. In addition, slight shifts of the core-level peaks toward higher binding energies were observed after combustion, reflecting oxidation-related structural rearrangements that favor the formation of P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O– and C
O– and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O–rich condensed structures. These chemically stabilized char components are expected to strengthen the protective char layer and contribute to the enhanced flame-retardant performance.39,40
O–rich condensed structures. These chemically stabilized char components are expected to strengthen the protective char layer and contribute to the enhanced flame-retardant performance.39,40
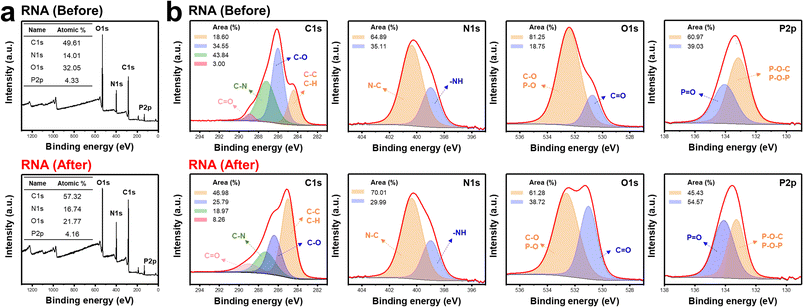 | ||
| Fig. 2 Chemical-state evolution of RNA before and after combustion. (a) Wide-scan X-ray photoelectron spectroscopy (XPS) spectra and (b) narrow-scan XPS spectra of RNA before and after combustion. | ||
To further elucidate the chemical changes occurring during combustion, narrow-scan XPS spectra were analyzed before and after burning (Fig. 2b). In the C 1s spectra of pristine RNA, peaks corresponding to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, C–N, C–O, and C–C/C–H bonds were observed at binding energies of 288.9, 287.2, 286.0, and 284.4 eV, respectively41–44 After combustion, these peaks shifted to higher binding energies of 289.0, 287.3, 286.4, and 284.8 eV, indicating chemical restructuring within the condensed phase. Quantitative analysis revealed that the relative area of the C
O, C–N, C–O, and C–C/C–H bonds were observed at binding energies of 288.9, 287.2, 286.0, and 284.4 eV, respectively41–44 After combustion, these peaks shifted to higher binding energies of 289.0, 287.3, 286.4, and 284.8 eV, indicating chemical restructuring within the condensed phase. Quantitative analysis revealed that the relative area of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O component, mainly associated with nucleobase-derived structures, increased from 3.00% to 8.26%. In contrast, the C–N component decreased from 43.84% to 18.97%, reflecting thermal degradation and transformation of nitrogen-containing moieties. Similarly, the C–O component, primarily originating from the ribose units, decreased from 34.55% to 25.79% due to dehydration and structural rearrangement of the sugar backbone. Conversely, the C–C/C–H component, indicative of char formation, increased markedly from 18.60% to 46.98%, consistent with extensive dehydration and carbonization during combustion.
O component, mainly associated with nucleobase-derived structures, increased from 3.00% to 8.26%. In contrast, the C–N component decreased from 43.84% to 18.97%, reflecting thermal degradation and transformation of nitrogen-containing moieties. Similarly, the C–O component, primarily originating from the ribose units, decreased from 34.55% to 25.79% due to dehydration and structural rearrangement of the sugar backbone. Conversely, the C–C/C–H component, indicative of char formation, increased markedly from 18.60% to 46.98%, consistent with extensive dehydration and carbonization during combustion.
In the N 1s spectra, peaks corresponding to N–C and –NH bonds were observed at binding energies of 400.0 and 399.0 eV, respectively.45,46 After combustion, these peaks shifted slightly to 400.4 and 399.1 eV. Quantitative analysis revealed an increase in the N–C component from 64.89% to 70.01%, suggesting the stabilization of nitrogen-containing species within the condensed phase, while the –NH component decreased from 35.11% to 29.99% due to the loss or transformation of amine- and amide-related functionalities during combustion.47
In the O 1s spectra, peaks corresponding to C–O/P–O and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bonds appeared at 532.4 and 530.7 eV, respectively.48–51 After combustion, these peaks shifted to 532.6 and 531.0 eV. The relative contribution of the C–O/P–O component decreased from 81.25% to 61.28%, reflecting the structural reorganization of ribose and phosphate groups, whereas the C
O bonds appeared at 532.4 and 530.7 eV, respectively.48–51 After combustion, these peaks shifted to 532.6 and 531.0 eV. The relative contribution of the C–O/P–O component decreased from 81.25% to 61.28%, reflecting the structural reorganization of ribose and phosphate groups, whereas the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O component increased from 18.75% to 38.72%, indicative of enhanced oxidation within the condensed char structure.
O component increased from 18.75% to 38.72%, indicative of enhanced oxidation within the condensed char structure.
In the P 2p spectra, peaks corresponding to P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and P–O–C/P–O–P bonds were observed at 134.0 and 133.1 eV, respectively,52,53 and shifted slightly to 134.1 and 133.3 eV after combustion. The relative area of the P
O and P–O–C/P–O–P bonds were observed at 134.0 and 133.1 eV, respectively,52,53 and shifted slightly to 134.1 and 133.3 eV after combustion. The relative area of the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O component increased from 39.03% to 45.43%, while that of P–O–C/P–O–P decreased from 60.97% to 54.57%, indicating the conversion of organic phosphate linkages into thermally stable condensed P
O component increased from 39.03% to 45.43%, while that of P–O–C/P–O–P decreased from 60.97% to 54.57%, indicating the conversion of organic phosphate linkages into thermally stable condensed P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O structures during combustion.54
O structures during combustion.54
Consequently, the morphological evolution and chemical transformations revealed by XPS demonstrate that RNA undergoes pronounced intumescent expansion, substantial carbon retention, stabilization of nitrogen-containing species, and the formation of condensed phosphate-rich networks. These synergistic processes contribute to the development of a highly stable protective char layer. Accordingly, RNA is not merely a biomolecule with incidental flame–retardant properties, but rather a structurally optimized, intrinsically synergistic bio-derived FR capable of providing robust condensed-phase fire protection.
3.3 Thermal behavior and flammability of RNA-PUF
The optimal RNA coating concentration was determined based on TGA results. The thermal degradation behavior of RNA-coated polyurethane foam (RNA-PUF) was systematically evaluated using TGA, DTG, DSC, and DTA analyses. To identify the optimal RNA loading for PUF coating, TGA was selected as the principal analytical tool, as the final char residue is a direct indicator of char-forming capability, a key parameter governing flame-retardant performance (Fig. S4). Bare PUF exhibited rapid mass loss starting at approximately 250 °C and left only 0.8% residue at 800 °C, consistent with its intrinsically high flammability. In contrast, RNA-PUF samples containing 2.5, 5.0, 7.5, and 10 wt% RNA showed substantially improved char formation, yielding final residues of 23.5, 37.0, 37.1, and 37.5%, respectively. Notably, while the char residue increased markedly when the RNA content increased from 2.5 to 5.0 wt%, the differences among 5.0, 7.5, and 10 wt% RNA were negligible. This trend indicates that the char yield reached a plateau at an RNA coating concentration of 5.0 wt%. At this optimal concentration, RNA-PUF and DNA-PUF exhibited add-on percentages of 32.40 ± 2.29% and 23.59 ± 3.34%, respectively, indicating the higher coating availability of RNA on the PUF matrix compared to DNA, which is attributed to the abundant 2′-OH groups on RNA ribose units. These hydroxyl groups facilitate the formation of a robust hydrogen-bonding network with the PUF substrate, significantly enhancing interfacial compatibility and adhesion. The higher coating efficiency of RNA-PUF would lead to superior thermal properties compared to DNA-PUF.At this optimal coating concentration, the thermal properties of RNA-PUF were further examined through advanced thermal analyses. To evaluate the thermal degradation and char-forming behavior of RNA-PUF, TGA was performed (Fig. 3a). Pure RNA yielded a high char residue of 46.2%, reflecting its intrinsically strong carbonization capability. Notably, RNA-PUF exhibited a final residue of 37.5%, which is substantially higher than that of bare PUF, indicating the formation of a protective RNA-derived char layer that remains in the condensed phase during combustion. In comparison, DNA-PUF showed a significantly lower final residue of 33.0%, highlighting the superior condensed-phase stability of RNA relative to DNA (Fig. S5a).
DSC analysis provided further insight into the thermal transitions governing the protective behavior of RNA-PUF (Fig. 3b). A pronounced exothermic peak appeared at approximately 160 °C, attributable to 2′-OH-driven dehydration and condensation reactions of RNA that initiate early carbonaceous network formation. The near-identical position and intensity of this exotherm compared with that observed in RNA (Fig. 1b) indicate that the intrinsic 2′-OH-driven network formation of RNA is preserved within the PUF matrix, with possible additional contribution from interfacial condensation between RNA's 2′-OH groups and urethane functionalities of the PUF. In the temperature range of 200–250 °C, the thermal transition pattern of RNA-PUF was distinctly modified compared with RNA. Notably, the endothermic peak characteristic of pure RNA at around 230 °C was replaced by a complex exothermic transition in RNA-PUF. This change suggests that RNA follows a modified thermal degradation pathway in the presence of the PUF substrate, likely involving strong interfacial interactions or cross-linking-like reactions. Such interactions alter the decomposition mechanism and facilitate char formation, thereby enhancing the flame-retardant effect. In contrast, DNA-PUF (Fig. S5b) exhibited an early, broad, and shallow exothermic feature starting near 50 °C, consistent with the gradual disruption of weaker and more heterogeneous hydrogen-bonding interactions associated with the absence of the 2′-OH group in DNA.
Additional mechanistic insight was obtained from DTA (Fig. 3c). Bare PUF exhibited a sharp and intense exothermic peak centered at approximately 350 °C, corresponding to rapid thermal decomposition and polymer chain scission. In contrast, pure RNA did not show a pronounced exothermic behavior; instead, it showed a continuous downward deviation in the DTA signal, indicative of dominant endothermic dehydration and char-forming processes. Notably, RNA-PUF effectively suppressed the characteristic exothermic peak observed for bare PUF, demonstrating that the presence of RNA strongly inhibits the thermal oxidation of the polymer through the formation of a protective condensed-phase char layer. In comparison, DNA-PUF retained a prominent exothermic peak similar to that of bare PUF, indicating substantially weaker condensed-phase stabilization (Fig. S5c). These observations are consistent with the DTG results (Fig. S6), which show that RNA-coated PUF undergoes delayed and moderated mass-loss stages relative to both bare PUF and DNA-PUF.
The small-scale flammability behavior of RNA-PUF was evaluated using the UL-94 horizontal burning foamed material (HBF) test, an internationally recognized method for assessing early-stage ignition behavior and flame-spread resistance (Fig. 3d and Video S2). Bare PUF ignited immediately and was completely consumed within 100 s, exhibiting continuous flame propagation accompanied by rapid collapse of its framework. In contrast, RNA-PUF retained its macroscopic structure throughout the test and successfully achieved a UL-94 HF-1 rating, indicating effective thermal insulation and flame-spread suppression. In addition, IR thermal imaging provided direct visualization of the thermal-shielding role of the RNA-derived char layer. During combustion of bare PUF, localized hot spots rapidly exceeded the upper detectable temperature limit of the IR sensor, appearing as bright yellow-white regions and reflecting severe localized heating and rapid flame propagation. By comparison, RNA-PUF maintained significantly lower surface temperatures throughout the burning process. The thermal images were dominated by blue-green regions, indicating reduced heat intensities with only minor and transient hot spots observed near the ignition zone. The surface temperature of RNA-PUF remained well below that of bare PUF, confirming that RNA forms a thermally insulating protective char barrier that effectively suppresses heat transfer and inhibits flame spread across the foam surface. Similarly, DNA-PUF maintained its original shape for 60 s, achieving an HF-1 rating, demonstrating its excellent FR performance on PUF (Fig. S7 and Video S3).
To evaluate the FR performance of RNA-PUF under conditions simulating environmental durability, UL-94 tests were conducted on aged samples (Fig. S8 and Video S4). Following ambient aging and heat aging, RNA-PUF maintained its structural integrity against a harsh flame for 60 s, successfully achieving an HF-1 rating. The robust performance demonstrates the anti-aging capabilities and practical applicability of the RNA coating. However, after the water immersion test, the flame on the RNA-PUF sample was completely extinguished at 85 s, but only after prolonged burning, resulting in a non-rating (NR). This reduction in performance may be attributed to water molecules competitively disrupting the hydrogen bonds between the RNA and the PUF matrix, leading to partial leaching of the RNA coating. Similarly, DNA-PUF maintained an HF-1 rating after both ambient and heat aging. However, following the water immersion test, the flame extinguished at 95 s, also achieving non-rating (Fig. S9 and Video S5). When comparing the two immersed samples, the shorter burning time of RNA-PUF relative to DNA-PUF demonstrates a higher retention of the FR properties.
Overall, these results demonstrate that RNA acts as a highly effective bio-derived flame retardant through its 2′-OH-driven dehydration, robust carbonization, and intrinsic cross-linking-like interactions. The RNA coating enables early-stage formation of a protective char layer, limits heat and mass transfer, suppresses combustion kinetics, and ultimately allows PUF to satisfy international UL-94 fire-safety requirements. However, further research to enhance the water-washing resistance of the RNA coating on the PUF matrix is necessary.
3.4 Quantitative analyses of flame retardant properties in RNA-PUF
To quantitatively evaluate the flame-retardant performance of RNA-PUF, limiting oxygen index (LOI) measurements and cone calorimetry tests were conducted. LOI is a widely used metric for assessing intrinsic flammability, as it reflects the minimum oxygen concentration required to sustain combustion, whereas cone calorimetry provides a comprehensive evaluation of heat release behavior, smoke production, ignition characteristics, and degradation kinetics under an external heat flux. Together, these complementary techniques enable a rigorous assessment of both ignitability and overall fire behavior, which is essential for validating the practical applicability of RNA as a bio-derived flame-retardant system.LOI testing revealed a substantial enhancement in oxygen resistance following RNA coating (Table 2). Bare PUF exhibited a low LOI value of 17.8 ± 0.1%, consistent with its intrinsically high flammability. DNA-PUF showed an increased LOI of 28.0 ± 0.4%, while RNA-PUF exhibited an even higher LOI value of 29.3 ± 0.5%. This superior performance is attributed to the enhanced compatibility of RNA with the PUF matrix, facilitated by a robust hydrogen-bonding network via its abundant 2′-OH groups. Consistent with this enhancement, pronounced char formation was observed for RNA-PUF during LOI testing (Fig. S10), further confirming the strong condensed-phase stabilization and protective function of RNA. To further investigate the practical flame-retardant durability of the coated foams, LOI tests were conducted following ambient aging, heat aging, and water immersion (Table S1). After ambient aging, the LOI of RNA-PUF remained remarkably stable at 29.2 ± 0.4%. After heat aging, the LOI value was reduced to 26.6 ± 0.8%, which is attributed to the partial thermal decomposition of RNA under the elevated aging temperature. Similarly, DNA-PUF exhibited a negligible decrease to 27.8 ± 0.5% after ambient aging and a moderate reduction to 24.3 ± 0.7% after heat aging, following the same trend of thermal degradation. However, following the water immersion test, the LOI values decreased significantly to 18.8 ± 0.1% and 17.9 ± 0.1% for RNA-PUF and DNA-PUF, respectively. This pronounced reduction is consistent with the corresponding UL-94 test results and is fundamentally attributed to water molecules competitively disrupting the hydrogen bonds between the nucleic acids and the PUF matrix, leading to the loss of the coatings. Consequently, while the RNA coating demonstrates exceptional flame retardancy under ambient and heat-aged conditions, its susceptibility to water-induced degradation remains a limitation.
| Bare PUF | RNA-PUF | DNA-PUF | |
|---|---|---|---|
| UL94 HBF test | NR | HF-1 | HF-1 |
| LOI (%) | 17.8 ± 0.1 | 29.3 ± 0.5 | 28.0 ± 0.4 |
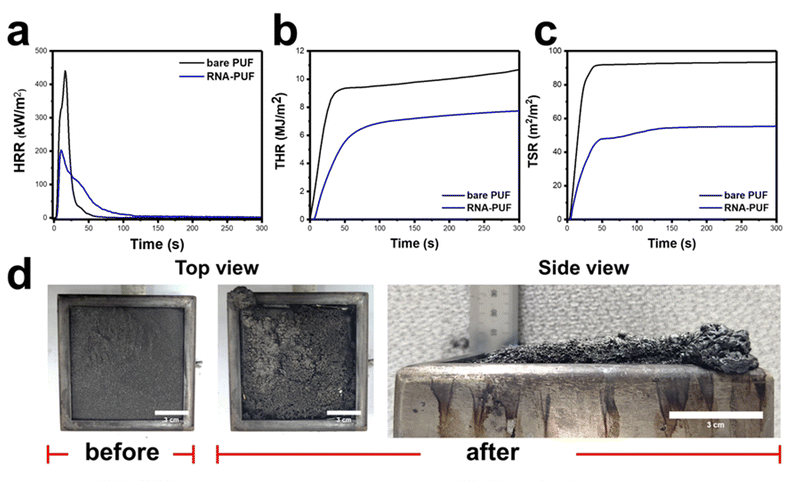
Cone calorimetry results further demonstrated the superior fire safety performance of RNA-PUF (Table 3). As shown in the heat release rate (HRR) profiles (Fig. 4a), RNA-PUF exhibited consistently lower heat release compared to bare PUF. The peak HRR (p-HRR) was significantly reduced by 49.4%, from 440.9 ± 18.5 to 223.3 ± 21.3 kW m−2, demonstrating that the RNA coating effectively mitigates combustion intensity. Additionally, the total heat release (THR) decreased by 28.0%, from 10.7 ± 0.3 to 7.7 ± 0.5 MJ m−2, confirming that RNA suppresses both the rate and overall magnitude of energy generation. Furthermore, RNA-PUF exhibited a significantly more gradual slope in its THR curve compared to bare PUF, indicating the robust RNA-derived char layer effectively delays the combustion process and limits rapid fire spread (Fig. 4b). RNA-PUF also displayed enhanced ignition resistance, with the time to ignition (TTI) increasing from 2.2 ± 0.7 s to 4.0 ± 0.6 s. Furthermore, the total smoke release (TSR) profiles exhibited a substantial reduction by 41.6%, from 94.9 ± 3.7 to 55.4 ± 9.8 m2 m−2 (Fig. 4c), and the effective heat of combustion (EHC) dropped sharply from 29.0 ± 0.2 to 21.7 ± 0.1 MJ kg−1. These results, coupled with an increase in char yield from 0.3 ± 0.2% to 22.6 ± 0.4%, indicate that the dense RNA-derived char layer successfully traps flammable volatiles and disrupts combustion through radical capturing. Consistent with these data, post-burn morphological analysis revealed that RNA-PUF retained a cohesive and expanded char barrier, preserving the structural integrity of PUF (Fig. 4d). In the case of DNA-PUF, FR improvements were also observed, including a delayed TTI from 3.4 ± 0.5 s and a 33.5% reduction in p-HRR to 293.1 ± 20.4 kW m−2. Furthermore, the THR profile displayed a gentler slope compared to bare PUF, indicating effective heat release suppression. However, when compared to RNA-PUF, DNA-PUF exhibited a distinct double-peak in its HRR curve (Fig. S11a) and yielded relatively higher THR of 9.7 ± 0.6 MJ m−2 (Fig. S11b) and TSR of 89.6 ± 4.9 m2 m−2 (Fig. S11c). Char residue after combustion also showed the relatively lower char-forming ability of DNA-PUF compared to RNA-PUF (Fig. S11d). This difference is attributed to the inferior structural stability of the DNA-derived char layer. While the DNA coating initially forms a protective char that corresponds to the first HRR peak, the structurally defective layer eventually cracks under continuous heat flux. The breakdown exposes the underlying PUF and the remaining DNA coating to complete combustion, generating the second HRR peak and ultimately resulting in a greater total heat release. Moreover, these structural cracks act as direct escape pathways for pyrolyzed volatiles, resulting in the higher TSR. Nevertheless, the reduced EHC of 24.6 ± 1.9 MJ kg−1 and increased char yield of 18.3 ± 0.5% demonstrate that DNA still provides a meaningful FR improvement over bare PUF.
| Sample | TTI (s) | p-HRR (kW m−2) | THR (MJ m−2) | TSR (m2 m−2) | EHC (MJ kg−1) | Char yield (%) |
|---|---|---|---|---|---|---|
| Bare PUF | 2.2 ± 0.7 | 440.9 ± 18.5 | 10.7 ± 0.3 | 94.9 ± 3.7 | 29.0 ± 0.2 | 0.3 ± 0.2 |
| RNA-PUF | 4.0 ± 0.6 | 223.3 ± 21.3 | 7.7 ± 0.5 | 55.4 ± 9.8 | 21.7 ± 0.1 | 22.6 ± 0.4 |
| DNA-PUF | 3.4 ± 0.5 | 293.1 ± 20.4 | 9.7 ± 0.6 | 89.6 ± 4.9 | 24.6 ± 1.9 | 18.3 ± 0.5 |
Taken together, the combined LOI and cone calorimetry results provide strong evidence that RNA, through its 2′-OH-driven dehydration, robust carbonization, and efficient intumescent char formation, significantly enhances both the ignition resistance and overall fire performance of PUF. Notably, RNA-PUF outperforms many conventional phosphorus-based flame retardants reported for PUF (Table S2), underscoring the unique potential of RNA as an intrinsically synergistic, bio-derived FR material.
3.5 Mechanism of the RNA-derived flame-retardant system
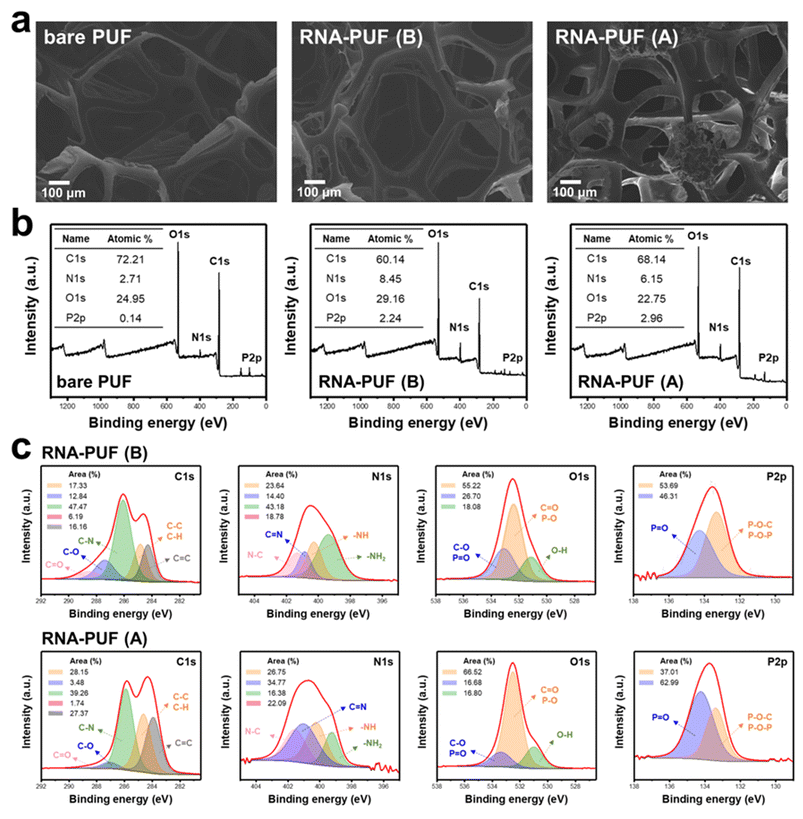
To elucidate the coating characteristics of RNA on PUF and the combustion behavior of RNA-PUF, SEM was performed (Fig. 5a). SEM images revealed that RNA was uniformly deposited on the PUF structures while preserving the original porous microstructure of the foam. EDS analysis further confirmed the presence and enrichment of nitrogen and phosphorus, characteristic elements of RNA, verifying the successful coating of RNA onto PUF (Fig. S12). Consistent with these observations, RNA-coated PUF exhibited enhanced mechanical performance in tensile tests conducted in accordance with ISO 1798![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2008. The tensile strain of RNA-PUF reached 154.8%, which can be attributed to the formation of a flexible hydrogen-bonding network facilitated by the abundant 2′-OH groups of RNA and its good interfacial compatibility with the PUF matrix (Fig. S13). In contrast, DNA-PUF displayed significantly lower tensile strain, likely due to the stronger base–base intermolecular interactions and poorer interfacial compatibility with the PUF matrix arising from the absence of 2′-OH groups. The recovery behavior under compressive loading (3000 kPa) was further evaluated (Fig. S14). Both bare PUF and RNA-PUF largely retained their original shapes after compression, indicating that the RNA coating does not compromise the intrinsic elastic recovery of the foam. In contrast, DNA-PUF exhibited noticeable deformation, reflecting reduced structural resilience associated with its rigid intermolecular interactions.
2008. The tensile strain of RNA-PUF reached 154.8%, which can be attributed to the formation of a flexible hydrogen-bonding network facilitated by the abundant 2′-OH groups of RNA and its good interfacial compatibility with the PUF matrix (Fig. S13). In contrast, DNA-PUF displayed significantly lower tensile strain, likely due to the stronger base–base intermolecular interactions and poorer interfacial compatibility with the PUF matrix arising from the absence of 2′-OH groups. The recovery behavior under compressive loading (3000 kPa) was further evaluated (Fig. S14). Both bare PUF and RNA-PUF largely retained their original shapes after compression, indicating that the RNA coating does not compromise the intrinsic elastic recovery of the foam. In contrast, DNA-PUF exhibited noticeable deformation, reflecting reduced structural resilience associated with its rigid intermolecular interactions.
These results collectively indicate that the RNA coating preserves the intrinsic elastic recovery of PUF while maintaining mechanical integrity under compression. After combustion, a continuous micro-scale char layer was observed on RNA-PUF, providing direct morphological evidence of its strong char-forming capability and condensed-phase flame-retardant mechanism.
XPS was employed to examine the surface composition of RNA-PUF before and after combustion. In the wide-scan spectra, the phosphorus content, a characteristic element of RNA, increased to 2.24% compared with bare PUF, in which phosphorus was not detected, verifying the successful deposition of RNA on the PUF surface (Fig. 5b). The presence of RNA on PUF was further supported by FTIR analysis (Fig. S15). After combustion, the carbon and phosphorus contents increased from 60.14% to 68.14% and from 2.24% to 2.96%, respectively, indicating the formation of carbon- and phosphorus-rich condensed char layer that contributes to the flame-retardant protection of RNA-PUF.
To further investigate the coating characteristics and combustion-induced chemical evolution, narrow-scan XPS spectra were analyzed before and after burning (Fig. 5c). Comparison of the narrow-scan spectra of bare PUF and RNA-PUF revealed the emergence of characteristic signals corresponding to N–C, –NH2, O–H, P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, and P–O–C/P–O–P bonds, confirming the presence of RNA on the PUF surface (Fig. S16).
O, and P–O–C/P–O–P bonds, confirming the presence of RNA on the PUF surface (Fig. S16).
In the C 1s spectra of RNA-PUF prior to combustion, peaks attributed to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, C–O, C–N, C–C/C–H, and C
O, C–O, C–N, C–C/C–H, and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds were observed at 288.6, 287.4, 286.1, 284.8, and 284.3 eV, respectively.55–57 After combustion, the relative areas of the C
C bonds were observed at 288.6, 287.4, 286.1, 284.8, and 284.3 eV, respectively.55–57 After combustion, the relative areas of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and C–O components decreased, reflecting thermal degradation of RNA, while the C–N contribution slightly declined due to the decomposition of nucleobases. In contrast, the C–C/C–H and C
O and C–O components decreased, reflecting thermal degradation of RNA, while the C–N contribution slightly declined due to the decomposition of nucleobases. In contrast, the C–C/C–H and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C components, the key indicators of carbonaceous char, significantly increased from 17.33% to 28.15% and from 16.16% to 27.37%, respectively. These changes confirm that RNA effectively promotes carbonization even in the PUF-coated configuration.
C components, the key indicators of carbonaceous char, significantly increased from 17.33% to 28.15% and from 16.16% to 27.37%, respectively. These changes confirm that RNA effectively promotes carbonization even in the PUF-coated configuration.
In the N 1s spectra, components attributed to N–C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N, –NH, and –NH2 species were observed at 401.3, 400.9, 400.3, and 399.4 eV, respectively, and shifted slightly to 401.6, 401.0, 400.4, and 399.5 eV after combustion58–61 Notably, the relative contribution of the –NH2 component decreased markedly from 43.18% to 16.38%, which can be attributed to dehydration and condensation reactions leading to nitrogen reorganization within the char matrix. Correspondingly, the C
N, –NH, and –NH2 species were observed at 401.3, 400.9, 400.3, and 399.4 eV, respectively, and shifted slightly to 401.6, 401.0, 400.4, and 399.5 eV after combustion58–61 Notably, the relative contribution of the –NH2 component decreased markedly from 43.18% to 16.38%, which can be attributed to dehydration and condensation reactions leading to nitrogen reorganization within the char matrix. Correspondingly, the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N component increased from 14.40% to 34.77%, indicating the formation of more thermally stable nitrogen-containing structures.
N component increased from 14.40% to 34.77%, indicating the formation of more thermally stable nitrogen-containing structures.
In the O 1s spectra, peaks corresponding to C–O/P–O, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O/P
O/P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, and O–H species appeared at 533.1, 532.4, and 531.1 eV, respectively, and exhibited minor shifts to 533.2, 532.5, and 531.0 eV after combustion.62,63 The relative contributions of the C–O/P–O and O–H components decreased from 26.70% to 16.68% and from 18.08% to 16.80%, respectively, reflecting dehydration and partial oxygen loss. In contrast, the C
O, and O–H species appeared at 533.1, 532.4, and 531.1 eV, respectively, and exhibited minor shifts to 533.2, 532.5, and 531.0 eV after combustion.62,63 The relative contributions of the C–O/P–O and O–H components decreased from 26.70% to 16.68% and from 18.08% to 16.80%, respectively, reflecting dehydration and partial oxygen loss. In contrast, the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O/P
O/P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O component increased from 55.22% to 66.52%, consistent with dehydration- and oxidation-induced transformation of functional groups originating from ribose units and nucleobases. The enrichment of C
O component increased from 55.22% to 66.52%, consistent with dehydration- and oxidation-induced transformation of functional groups originating from ribose units and nucleobases. The enrichment of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O/P
O/P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O species is expected to contribute to the formation of a compact and thermally robust char layer.
O species is expected to contribute to the formation of a compact and thermally robust char layer.
In the P 2p spectra, peaks assigned to P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and P–O–C/P–O–P bonds were detected at 134.3 and 133.3 eV, respectively, and showed only negligible shifts after combustion.64 The relative area of the P
O and P–O–C/P–O–P bonds were detected at 134.3 and 133.3 eV, respectively, and showed only negligible shifts after combustion.64 The relative area of the P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O component increased from 46.31% to 62.99%, while that of the P–O–C/P–O–P component decreased from 53.69% to 37.01%, indicating the conversion of organic phosphate structures into thermally stable condensed P
O component increased from 46.31% to 62.99%, while that of the P–O–C/P–O–P component decreased from 53.69% to 37.01%, indicating the conversion of organic phosphate structures into thermally stable condensed P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O structures during combustion.
O structures during combustion.
Finally, to investigate the gas-phase emissions during the thermal degradation of RNA, STA-MS analysis was conducted (Fig. S17). The emissions of highly flammable methane (CH4, m/z = 16) and toxic hydrogen cyanide (HCN, m/z = 27) showed lower intensities compared to other gaseous species throughout the entire temperature range. This significant suppression is attributed to the fact that RNA predominantly releases nitrogen in non-toxic forms and rapidly forms a robust early-stage char barrier that traps combustible volatile fragments. Simultaneously, the intensive emission of non-combustible gases was observed. Water vapor (H2O, m/z = 18) displayed prominent peaks at approximately 240 °C and 270 °C, providing direct evidence of the early and intensive dehydration reactions driven by the abundant 2′-OH groups on the RNA ribose units. Moreover, H2O can act as a cooling agent against combustible volatiles in the gas phase. Ammonia (NH3, m/z = 17) exhibited pronounced peaks in the same temperature range, originating from the deamination and thermal decomposition of the nitrogen-rich nucleobases. Furthermore, carbon dioxide (CO2, m/z = 44) was abundantly released at 270 °C and 310 °C, resulting from the thermal cleavage and decomposition of the ribose sugar rings and nucleobases. Together, the continuous release of these non-flammable gases acts as a powerful blowing agent to expand the char and provides a significant dilution effect. The facilitated formation of the RNA-derived char further inhibited the release of carbon monoxide (CO, m/z = 28). While the CO emission was kept remarkably low around 270 °C compared to the non-combustible gases, the release sharply increased only after reaching extreme temperatures above 650 °C. This delayed CO emission is likely associated with the progressive aromatization and high-temperature pyrolytic decomposition of residual oxygen-containing functional groups within the highly crosslinked carbonaceous network under an inert atmosphere. These results clearly indicate that the effective char layer formed by RNA can endure extreme thermal stress, serving as an exceptional bio-derived flame retardant.
Overall, RNA coating on PUF promotes rapid char formation primarily through dehydration and condensation reactions of its functional groups, particularly the abundant hydroxyl groups present in RNA (Scheme 1). This process results in the formation of a carbon- and phosphorus-rich protective layer that effectively suppresses flame propagation by limiting heat and mass transfer. The resulting char barrier acts as a thermal shield, reducing heat flux from the flame to the PUF substrate and thereby slowing internal thermal degradation during combustion. In addition to condensed-phase protection, the high phosphorus and nitrogen contents in RNA can contribute to gas-phase flame inhibition. Phosphorus-containing species are reported to generate PO˙ radicals, which effectively scavenge O˙ and OH˙ radicals and interrupt chain-propagation reactions in the flame.65–67 Meanwhile, the release of nitrogen-containing gases, such as NH3, enables dilution of combustible volatiles, further enhancing the overall flame-retardant performance.68
4 Conclusion
In summary, the thermal behavior and flame-retardant performance of ribonucleic acid (RNA), a bio-derived macromolecule, were systematically investigated to assess its potential as an intrinsically synergistic biomolecular flame retardant. RNA exhibited high thermal stability and pronounced intumescent char-forming capability, arising from its intrinsically high carbon, nitrogen, and phosphorus content, acting respectively as the carbon source, blowing agent, and acid source during combustion. Compared with the structurally analogous biomolecule deoxyribonucleic acid (DNA), RNA displayed distinctly different degradation pathways, attributable to the presence of uracil and an additional 2′-OH group in its ribose units, which promote early dehydration and significantly enhance condensed-phase char formation.When applied as a coating on PUF, RNA markedly increased the residual char yield from 0.8% to 37.5% in TGA, demonstrating its strong char-promoting capacity. RNA-PUF achieved an LOI of 29.3% and attained a UL-94 HF-1 classification, confirming its effective flammability suppression. Cone calorimetry further revealed a 49.4% reduction in the peak heat-release rate (p-HRR) compared with bare PUF. Moreover, RNA-PUF successfully retained its excellent flame-retardant performance after both ambient and heat aging. However, it exhibited reduced flame retardancy following water immersion. This decline is attributed to water molecules competitively disrupting the intermolecular hydrogen bonds between the RNA and the PUF substrate, leading to coating leaching.
Consequently, these results identify RNA as a sustainable and intrinsically synergistic bio-based flame retardant that integrates char formation, gas dilution, and radical quenching within a single molecular framework. RNA thus represents a highly promising platform material for the development of next-generation fire-safety technologies, however, the water-washing resistance remains a challenge for future research.69–71
Author contributions
Sang Yun Jeong: conceptualization; investigation; visualization; writing – original draft; Jong Seok Lim: conceptualization; investigation; visualization; writing – original draft; Abdullah Malik: investigation, methodology; Jae Chul Ro: methodology, analysis, funding acquisition, writing – review & editing; Hui Hun Cho: supervision, writing – review & editing; Jun Hyuk Heo: conceptualization, funding acquisition, supervision, writing – review & editing; Jung Heon Lee: conceptualization, funding acquisition, supervision, writing – review & editing.Conflicts of interest
The authors declare that they have no competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Additional data are available from the corresponding authors upon reasonable request. Supplementary information: (i) molecular structure comparison of RNA and DNA highlighting the key chemical differences relevant to flame-retardant behavior; (ii) comparison of the chemical state between RNA and DNA via FTIR; (iii) thermal behaviors of uracil and thymine. (a) TGA, (b) DSC, and (c) DTA; (iv) optimization of RNA concentration through TGA analysis; (v) comparative thermal analyses of RNA-PUF and DNA-PUF. (a) TGA, (b) DSC, and (c) DTA; (vi) comparison of thermal degradation behavior between RNA-PUF and DNA-PUF through DTG curves; (vii) UL-94 HBF test for DNA-PUF; (viii) UL-94 HBF test for RNA-PUF after aging and water immersion; (ix) UL-94 HBF test for DNA-PUF after aging and water immersion; (x) photographs of RNA-PUF before and after conduction of LOI test; (xi) cone calorimetry analysis of DNA-PUF. (a) Heat-release rate (HRR), (b) total heat release (THR), and total smoke release (TSR) curves of bare PUF and DNA-PUF measured under an external heat flux of 35 kW m−2. (d) Top-view and side-view photographs of DNA-PUF before and after cone calorimetry testing; (xii) EDS analysis results of bare PUF and RNA-PUF (B); (B) means before combustion; (xiii) tensile stress–strain curves of bare PUF, RNA-PUF, and DNA-PUF, along with the photographs of RNA-PUF during the tensile test; (xiv) investigation of recovery behavior of bare PUF, RNA-PUF, and DNA-PUF under a compression load of 3000 kPa; (xv) comparative FTIR analysis among bare PUF, RNA, and RNA-PUF; (xvi) XPS narrow-range spectra for bare PUF and RNA-PUF; (xvii) STA-MS spectra of RNA thermal decomposition; (xviii) UL94 HBF and LOI test results for RNA-PUF and DNA-PUF after aging and immersion (Table S1); (xix) comparison of flame retardancy evaluation of FR applied PUFs (Table S2); (xx) burning tests on DNA and RNA pellets (Video S1); (xxi) UL-94 horizontal burning (HBF) tests of bare PUF and RNA-PUF (Video S2); (xxii) UL94 HBF test of DNA-PUF (Video S3); (xxiii) UL-94 HBF test of RNA-PUF after aging and immersion (Video S4); (xxiv) UL-94 HBF test of DNA-PUF after aging and immersion (Video S5). See DOI: https://doi.org/10.1039/d6ta00548a.Acknowledgements
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF), funded by the Ministry of Education (No. RS-2019-NR040077), and by the Technology Innovation Program (TIPS) funded by the Ministry of SMEs and Startups (MSS, Republic of Korea) (No. RS-2025-25464395). J.H.H. acknowledges additional support from the Basic Science Research Program through the National Research Foundation of Korea (NRF), funded by the Ministry of Education (No. RS-2022-NR072200). This work was also supported by the GRRC program of Gyeonggi Province [GRRC Sungkyunkwan(S) 2023-B02] and the Korea Basic Science Institute (National Research Facilities and Equipment Center) funded by the Ministry of Education (No. RS-2025-02308784).References
- S. H. Jeong, C. H. Park, H. Song, J. H. Heo and J. H. Lee, J. Clean. Prod., 2022, 368, 133241 CrossRef CAS.
- C. Zhang, Y. Jiang, S. Li, Z. Huang, X.-Q. Zhan, N. Ma and F.-C. Tsai, Heliyon, 2022, 8, e11225 CrossRef CAS PubMed.
- S. D. Shaw, A. Blum, R. Weber, K. Kannan, D. Rich, D. Lucas, C. P. Koshland, D. Dobraca, S. Hanson and L. S. Birnbaum, Rev. Environ. Health, 2010, 25, 261–306 CAS.
- M.-J. Chen, Z.-B. Shao, X.-L. Wang, L. Chen and Y.-Z. Wang, Ind. Eng. Chem. Res., 2012, 51, 9769–9776 CrossRef CAS.
- H. Song, C. H. Park, S. H. Jeong, J. H. Heo and J. H. Lee, ACS Appl. Polym. Mater., 2023, 5, 4388–4399 CrossRef CAS.
- K. A. Salmeia, S. Gaan and G. Malucelli, Polymers, 2016, 8, 319 CrossRef PubMed.
- S. Yoon, H. Song, S. H. Jeong, Y. Kim, J. H. Heo and J. H. Lee, ACS Appl. Polym. Mater., 2023, 5, 8284–8295 CrossRef CAS.
- M. M. Velencoso, A. Battig, J. C. Markwart, B. Schartel and F. R. Wurm, Angew. Chem., Int. Ed., 2018, 57, 10450–10467 CrossRef CAS PubMed.
- F. Xu, L. Zhong, C. Zhang, P. Wang, F. Zhang and G. Zhang, ACS Sustain. Chem. Eng., 2019, 7, 13999–14008 CrossRef CAS.
- B. Hou, B. Liu, H. Shen and M. Zhang, Chem. Eng. J., 2025, 524, 169171 CrossRef CAS.
- P. Jiang, X. Gu, S. Zhang, S. Wu, Q. Zhao and Z. Hu, Ind. Eng. Chem. Res., 2015, 54, 2974–2982 CrossRef CAS.
- B. Jiang, J. Wu, Y. Huang, W. Yu, S. Zhang and Y. Zhang, Chem. Eng. J., 2025, 521, 166541 CrossRef CAS.
- H. Duan, Y. Chen, S. Ji, R. Hu and H. Ma, Chem. Eng. J., 2019, 375, 121916 CrossRef CAS.
- Z. Chen, B. Yang, N. Song, T. Chen, Q. Zhang, C. Li, J. Jiang, T. Chen, Y. Yu and L. X. Liu, Chem. Eng. J., 2023, 455, 140547 CrossRef CAS.
- S. Y. Jeong, H. Song, K. Koo, J. S. Lim, H. H. Cho, J. H. Heo and J. H. Lee, Adv. Ind. Eng. Polym. Res., 2025, 8, 539–551 CAS.
- H. Nabipour and S. Rohani, J. Environ. Chem. Eng., 2025, 13, 116932 CrossRef CAS.
- K. Wazarkar, M. Kathalewar and A. Sabnis, Prog. Org. Coat., 2015, 87, 75–82 CrossRef CAS.
- S. H. Jeong, J. H. Heo, J. W. Lee, M. J. Kim, C. H. Park and J. H. Lee, ACS Appl. Mater. Interfaces, 2021, 13, 22935–22945 CrossRef CAS.
- J. Alongi, R. A. Carletto, A. D. Blasio, F. Carosio, F. Bosco and G. Malucelli, J. Mater. Chem. A, 2013, 1, 4779–4785 RSC.
- F. Carosio, A. D. Blasio, J. Alongi and G. Malucelli, Polymer, 2013, 54, 5148–5153 CrossRef CAS.
- J. Alongi, R. A. Carletto, A. D. Blasio, F. Cuttica, F. Carosio, F. Bosco and G. Malucelli, Carbohydr. Polym., 2013, 96, 296–304 CrossRef CAS PubMed.
- Z. J. Devereaux, Y. Zhu and M. Rodgers, Eur. J. Mass Spectrom., 2018, 25, 16–29 CrossRef PubMed.
- J. S. Paige, K. Y. Wu and S. R. Jaffrey, Science, 2011, 333, 642–646 CrossRef CAS PubMed.
- C. A. Kellenberger, C. Chen, A. T. Whiteley, D. A. Portnoy and M. C. Hammond, J. Am. Chem. Soc., 2015, 137, 6432–6435 CrossRef CAS PubMed.
- H. Han, J. H. Jeung, Y. Cho and J. K. Ahn, Biochip J., 2025, 19, 109–116 CrossRef CAS.
- H.-M. Kim and J. Park, Biochip J., 2024, 18, 622–632 CrossRef CAS.
- S. W. Shin, C. Baek, D. Luo, S. H. Um and J. Min, Biochip J., 2024, 18, 566–575 CrossRef CAS.
- A. R. Chandrasekaran, J. A. Punnoose, L. Zhou, P. Dey, B. K. Dey and K. Halvorsen, Nucleic Acids Res., 2019, 47, 10489–10505 CrossRef CAS PubMed.
- J. Han, J. Lim, C.-P. J. Wang, J.-H. Han, H. E. Shin, S.-N. Kim, D. Jeong, S. H. Lee, B.-H. Chun, C. G. Park and W. Park, Nano Converg., 2023, 10, 36 CrossRef CAS PubMed.
- X. Hou, T. Zaks, R. Langer and Y. Dong, Nat. Rev. Mater., 2021, 6, 1078–1094 CrossRef CAS PubMed.
- D. Jung, S. Jang, D. Park, N. H. Bae, C. S. Han, S. Ryu, E.-K. Lim and K. G. Lee, Biochip J., 2025, 19, 79–90 CrossRef CAS.
- L. Costes, F. Laoutid, S. Brohez and P. Dubois, Mater. Sci. Eng. R Rep., 2017, 117, 1–25 CrossRef.
- P. Zhang, Y. Zhou, H. Su, H. Lin, S. Tian, Y. Chen, J. Yan, Y. He and H. Fan, Polym. Int., 2017, 66, 1598–1609 CrossRef CAS.
- H.-B. Chen, P. Shen, M.-J. Chen, H.-B. Zhao and D. A. Schiraldi, ACS Appl. Mater. Interfaces, 2016, 8, 32557–32564 CrossRef CAS PubMed.
- P. Routh, A. Garai and A. K. Nandi, Phys. Chem. Chem. Phys., 2011, 13, 13670–13682 RSC.
- D. Kapusuz, Colloid Polym. Sci., 2021, 299, 63–72 CrossRef CAS.
- J. Wang, L. Qian, B. Xu, W. Xi and X. Liu, Polym. Degrad. Stab., 2015, 122, 8–17 CrossRef CAS.
- T. P. Haider, O. Suraeva, I. Lieberwirth, P. Paneth and F. R. Wurm, Chem. Sci., 2021, 12, 16054–16064 RSC.
- J. Mäkelä, A. Lahti, M. Tuominen, M. Yasir, M. Kuzmin, P. Laukkanen, K. Kokko, M. P. J. Punkkinen, H. Dong, B. Brennan and R. M. Wallace, Sci. Rep., 2019, 9, 1462 CrossRef PubMed.
- B. Schartel, Materials, 2010, 3, 4710–4745 CrossRef CAS PubMed.
- P. Wang, L. Xia, R. Jian, Y. Ai, X. Zheng, G. Chen and J. Wang, Polym. Degrad. Stab., 2018, 149, 69–77 CrossRef CAS.
- A. I. A. Soliman, J. A. D. Baca and P. Fatehi, Carbohydr. Polym., 2023, 308, 120619 CrossRef CAS PubMed.
- H. Wang, Dinesh and J. Kim, Carbohydr. Polym., 2024, 326, 121616 CrossRef CAS PubMed.
- D. Hu, A. Wang, X. Qi, J. Yang, D. Sun and Y. Wang, J. Mater. Chem. A, 2025, 13, 22425–22444 RSC.
- S. R. Dugasani, B. Gnapareddy, M. R. Kesama, S. Jeon, J.-H. Jeong and S. H. Park, AIP Adv., 2019, 9, 015011 CrossRef.
- D. Lachowicz, A. Kmita, R. Wirecka, K. Berent, M. Szuwarzyński, S. Zapotoczny, A. Pajdak, G. Cios, N. Mazur-Panasiuk, K. Pyrc and A. Bernasik, Carbohydr. Polym., 2023, 312, 120756 CrossRef CAS PubMed.
- Y. V. Lim, Y. Wang, D. Kong, L. Guo, J. I. Wong, L. K. Ang and H. Y. Yang, J. Mater. Chem. A, 2017, 5, 10406–10415 RSC.
- S. Kokkiligadda, S. R. Dugasani, E. P. Komarala, S. Jeon, J.-H. Jeong and S. H. Park, J. Phys. D: Appl. Phys., 2021, 54, 375401 CrossRef CAS.
- Y. Dong, B. Liu, S. H. Lee, W. C. Lum, Y. Ren, X. Zhou, H. Wang, B. Zhou and J. Zhang, Chem. Eng. J., 2024, 495, 153614 CrossRef CAS.
- F. Yu, Z. Ba, Z. Gao, Y. Wang, Y. Xie, H. Wang, Z. Qiu and Z. Xiao, Chem. Eng. J., 2024, 493, 152827 CrossRef CAS.
- X. Qian, L. Song, B. Yu, B. Wang, B. Yuan, Y. Shi, Y. Hu and R. K. K. Yuen, J. Mater. Chem. A, 2013, 1, 6822–6830 RSC.
- P. M. A. Sherwood, Surf. Sci. Spectra, 2002, 9, 62–66 CrossRef CAS PubMed.
- W. Ali, O. Zilke, D. Danielsiek, A. Salma, B. Assfour, V. Shabani, S. Caglar, H. M. Phan, L. Kamps, R. Wallmeier, Y. Feng, T. Textor, J. S. Gutmann and T. Mayer-Gall, Cellulose, 2023, 30, 2627–2652 CrossRef CAS.
- W. Liu, D.-Q. Chen, Y.-Z. Wang, D.-Y. Wang and M.-H. Qu, Polym. Degrad. Stab., 2007, 92, 1046–1052 CrossRef CAS.
- S. Kumaravel, P. Thiruvengetam, K. Karthick, S. S. Sankar and S. Kundu, ACS Sustain. Chem. Eng., 2019, 7, 18463–18475 CrossRef CAS.
- C. Y. Lee, S. Y. Jeong, D.-H. Kim, M. J. Kim, J. H. Heo and J. H. Lee, Sens. Actuators, B Chem., 2024, 419, 136358 CrossRef CAS.
- S. Sui, H. Quan, J. Wang, Y. Lu, Y. Yang, Y. Sheng, Z. Sun and Y. Zhang, Adv. Sci., 2025, 12, 2410938 CrossRef CAS PubMed.
- E. Korin, N. Froumin and S. Cohen, ACS Biomater. Sci. Eng., 2017, 3, 882–889 CrossRef CAS.
- F. Bian, R. Huang, X. Li, J. Hu and S. Lin, Adv. Sci., 2024, 11, 2307482 CrossRef CAS PubMed.
- Y. Zeng, Z. Li, Y. Feng, M. Chi, H. Lv and G.-Y. Yang, J. Mater. Chem. A, 2025, 13, 9830–9840 RSC.
- X. Jin, J. Sun, J. S. Zhang, X. Gu, S. Bourbigot, H. Li, W. Tang and S. Zhang, ACS Appl. Mater. Interfaces, 2017, 9, 24964–24975 CrossRef CAS PubMed.
- J. W. Lee, B. Lee, C. H. Park, J. H. Heo, T. Y. Lee, D. Lee, J. Bae, P. R. Sundharbaabu, W. K. Yeom, S. Chae, J. Lim, S. Lee, J. Choi, H. Bae, J. Choi, E. Lee, D. S. Yoon, G. Y. Yeom, H. Shin and J. H. Lee, Adv. Mater., 2024, 36, e2406179 CrossRef PubMed.
- W. Yi, W. Tang, W. Su, K. Shi, Q. Xiao, Z. Wang, X. Li, J. Mu, H. Yao and Z. Peng, Adv. Sci., 2025, e16139 Search PubMed.
- M. Ramstedt and A. Shchukarev, Surf. Sci. Spectra, 2023, 30, 014011 CrossRef CAS.
- M. Yu, Y. Chu, W. Xie, L. Fang, O. Zhang, M. Ren and J. Sun, Chem. Eng. J., 2024, 480, 148167 CrossRef CAS.
- G. Zhu, J. Gao, Y. Huang, Z. Wang, Y. Wang and Z. Zhu, Constr. Build. Mater., 2026, 509, 145224 CrossRef CAS.
- Y. Huang, G. Zhu, Z. Wang, Y. Wang and Z. Zhu, Chem. Eng. J., 2026, 527, 171625 CrossRef CAS.
- Z. Zhu, J. Gao, G. Zhu, Y. Huang, Z. Wang, J. Feng, Y. Wang and P. Song, Composites, Part B, 2026, 312, 113344 CrossRef CAS.
- H. Song, S. H. Jeong, C. H. Park, M. J. Kim, H. Kim, J. H. Heo and J. H. Lee, Chem. Eng. J., 2024, 497, 154406 CrossRef CAS.
- S. F. A. Zaidi, A. Saeed, V.-C. Ho, J. H. Heo, H. H. Cho, N. Sarwar, N.-E. Lee, J. Mun and J. H. Lee, Int. J. Biol. Macromol., 2023, 234, 123725 CrossRef CAS PubMed.
- C. H. Park, H. Song, D.-H. Kim, J. Kim, S. H. Jeong, J. B. B. Rayappan, J. H. Heo and J. H. Lee, Chem. Eng. J., 2025, 506, 159912 CrossRef CAS.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |