 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Uncertainty-aware dimensionality prediction of low-dimensional hybrid metal halides by integrating Bayesian modeling and experiments
Migon
Choi
 a,
Dongkyu Derek
Cho
b,
Richard
Sheridan
a,
Dongkyu Derek
Cho
b,
Richard
Sheridan
 a,
David B.
Mitzi
a,
David B.
Mitzi
 ac and
L. Catherine
Brinson
ac and
L. Catherine
Brinson
 *a
*a
aDepartment of Mechanical Engineering and Materials Science, Duke University, Durham, North Carolina 27708, USA. E-mail: cate.brinson@duke.edu
bDepartment of Statistical Science, Duke University, Durham, North Carolina 27708, USA
cDepartment of Chemistry, Duke University, Durham, North Carolina 27708, USA
First published on 9th April 2026
Abstract
Data-driven materials informatics is revolutionizing materials design by uncovering complex structure–property relationships beyond traditional trial-and-error methods. In perovskites and more generally hybrid metal halides (HMHs), comprising metal halide anions and organic spacer cations, variations in the spacer cations can alter the inorganic framework dimensionality, which has direct consequences for optical and electronic properties. In this paper, we explore the factors impacting HMH dimensionality, with a focused experimental study and a broad materials informatics approach. Experimentally, we employed three structurally similar branched alkyl cations. Despite their close similarity, they produced distinct lead iodide hybrid frameworks: one forming a 2D layered structure and two forming 1D chain phases. These results confirm that molecular differences can dictate dimensionality. Expanding from these observations, we curated a dataset of 113 HMHs and applied Bayesian additive regression trees to predict dimensionality from molecular descriptors. Here, we show that Bayesian additive regression trees achieved a strong predictive power (posterior mean area under the curve of around 0.8) while quantifying uncertainty. The results highlight organic cation aspect ratio, polar surface area, and the number of branched points as dominant features for dimensionality prediction. An active learning strategy further enhanced model net improvement by ∼20%, increasing the efficiency of identifying promising candidates compared to random sampling. Together, this study provides both experimental evidence and machine learning rules that clarify how spacer cation structure governs HMH dimensionality, offering a data-driven path to rational design of low-dimensional hybrid semiconductors. By integrating experimental observations with data-driven modeling, this study highlights the potential of materials informatics to guide predictive design across structurally diverse material systems and motivates broader curation of well annotated materials datasets.
1. Introduction
Advances in materials informatics are transforming the way materials are designed and optimized, enabling the discovery of structure–property relationships beyond conventional trial-and-error approaches.1,2 These data-driven methods have shown great promise for tackling complex material systems where chemical variations can lead to drastic changes in structure and performance, such as hybrid perovskites, where molecular variations can lead to dramatic shifts in structure. For instance, a slight change in the organic spacer cation can alter crystal dimensionality (i.e., referring to connectivity of the inorganic framework), switching the material's optoelectronic behavior and long-term stability.3–7Hybrid perovskites represent a class of materials, generally comprising an extended framework of corner-sharing metal halide octahedra alternating with regions of organic cations.8 These semiconductors have shown outstanding potential in optoelectronic devices, due to low processing cost, tunable band gap, and high efficiency.9–12 The structural diversity is remarkable, and their performance depends on the relationship between structure, processing, properties, and performance.13,14 The dimensionality of these frameworks – whether the crystal forms 0D, 1D, 2D, or 3D – impacts optoelectronic properties, such as light absorption and charge transport.15–17 Therefore, controlling dimensionality is critical for efficient and stable devices based on these semiconductors. Given that our study includes octahedra sharing paradigms beyond conventional corner-sharing perovskites (e.g., corner-, edge-, or face-sharing), we collectively refer to these materials as hybrid metal halides (HMHs), a broader class of which hybrid organic–inorganic perovskites represent a special subset.
In low-dimensional HMHs, the organic spacer cation plays a key role in determining the resulting structure. Recent studies have shown that both structural and electronic features of the spacer – such as aromaticity, steric hindrance, and dielectric constant – can significantly influence the HMH's structure, properties, and performance.18–20 Despite these insights, the selection of spacer cations has largely relied on researchers' intuition, and a systematic, data-driven understanding of how molecular features govern HMH dimensionality remains limited. Recent studies demonstrate that machine learning (ML) can extract meaningful design rules for low-dimensional HMHs. Lyu et al. showed that steric and hydrogen-bonding descriptors can distinguish whether a spacer forms 2D frameworks.21 Mai et al. applied descriptor-based regression modeling at the device level, using feature-importance analysis to identify molecular properties most correlated with power conversion efficiency.22 Other works, such as database-driven band-gap modeling23 and ligand screening for stability,24 further highlight the potential of ML in this space. While these studies provide valuable advances, a systemic understanding of how spacer descriptors relate to dimensionality within experimentally accessible, data-limited regimes remains limited. Moreover, most prior ML approaches rely on point-estimate models without addressing predictive uncertainty, which is an important consideration for reliable predictions in the small, heterogeneous datasets common in experimental materials research.
Within this framework, we apply machine learning to investigate how variations in spacer cation structure influence HMH dimensionality. We scope our work to a specific experimental regime defined by Pb–I based frameworks and comparable synthesis protocols, and how molecular-level descriptors of spacer cations can support probabilistic predictions of dimensionality within this constrained space. We utilized Bayesian Additive Regression Trees (BART) as our primary predictive model, with a Random Forest classifier (RF classifier) serving as the baseline. Unlike conventional classifiers, BART provides not only predictions but also posterior predictive uncertainty.25 In data-limited and heterogeneous materials systems, quantifying uncertainty is crucial for identifying overconfident extrapolation and assessing the reliability of model predictions.26–28
To ground our machine learning analysis, we start with experimental observations showing that variations in branched alkyl spacer cations lead to distinct HMH dimensionalities. To capture these trends systemically, we begin with three experimentally realized systems derived from organic spacers based on branched alkyl amines: 3,3-dimethylbutylamine (3,3-DMBA), 2,3-dimethylbutan-2-amine (2,3-DMB2A), and N-methylbutan-2-amine (NMB2A), each combined with PbI2 under identical conditions. Variations in the spacer cations drive different outcomes, yielding a 2D layered phase ((3,3-DMBA·H)2PbI4) versus 1D chain motifs ((2,3-DMB2A·H)PbI3 and (NMB2A·H)PbI3). Motivated by these experimental insights, we subsequently broaden our machine learning analysis to explore a wider chemical space by combining data curated from the literature with an existing data resource. Using the classification models, we predict the dimensionality of HMH structures and systematically investigate the most influential molecular features that govern this outcome. Finally, through a simulated active learning cycle with BART, we highlight how its uncertainty-aware framework can both enhance predictive performance and provide interpretable cluster-level insights, pointing to its potential for future experimental application.
This work presents a framework for the data-driven design of low-dimensional HMH structures, including hybrid perovskites, offering new insight into the relationship between molecular structure and dimensionality. Our goal is not a universally generalizable model, but the development of an uncertainty-aware, data-driven framework that can inform experimental decision-making under realistic data limitations. While demonstrated here for low-dimensional HMHs, the conceptual framework – particularly the emphasis on uncertainty-aware learning in data-scarce regimes – may be informative for other hybrid materials systems, such as Covalent Organic Frameworks (CoFs),29 and van der Waals layered materials,30 where structural diversity complicates rational design. In particular, the uncertainty-aware nature of BART makes it powerful in data-scarce regimes, where experimental throughput is limited, and predictive reliability is crucial.31,32
2. Methods
2.1. Crystal growth and characterization
Single crystals were obtained by slow evaporation of PbI2 with the corresponding organic amines in HI/methanol solutions. Stoichiometries of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 2
1, 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, and 4
1, and 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 spacer-to-lead ratios were explored. Exact reagent quantities and conditions are summarized in SI (1. Single crystal growth). Structural analysis was conducted using PXRD and SCXRD, while optical properties were probed by UV-vis and PL spectroscopy. Thermal stability was assessed by TGA. Full instrument specifications and measurement parameters are provided in SI (2. Characterization).
1 spacer-to-lead ratios were explored. Exact reagent quantities and conditions are summarized in SI (1. Single crystal growth). Structural analysis was conducted using PXRD and SCXRD, while optical properties were probed by UV-vis and PL spectroscopy. Thermal stability was assessed by TGA. Full instrument specifications and measurement parameters are provided in SI (2. Characterization).
2.2. Data handling
A dataset of 113 HMHs was compiled from the Hybrid3 database (available at https://hybrid3.duke.edu/), the literature, and our experimental results. Chemical names and dimensionality labels were obtained directly from these sources, while molecular descriptors were computed separately as described in Section 2.3. Descriptors in methods section. Hybrid3 database provides 11 1D structures, which was supplemented with 26 additional 1D structures, including 2 generated in our experiments and 24 collected from the literature, resulting in a total of 37 1D entries. We also included 76 2D structures, of which 75 came from the Hybrid3 database and 1 from our experiments. To ensure consistency, we only retained lead iodide-based single-crystal samples with 1D or 2D structures, and organic cations that could be converted to SMILES strings (Table S3). The final dataset comprised 113 entries (76 2D and 37 1D). More detailed filtering criteria are described in SI (3. Data handling). The data were split into training and test sets (Train: 84, Test: 29) for Sections 3.2.1 (Model performance and uncertainty quantification) and 3.2.2 (Feature importance); into training, test, and experimental sets (Train: 82, Test: 28, Experimental: 3) for Section 3.2.3 (Demonstration on synthesized structures); and into training, test, and pool sets (Train: 69, Test: 29, Pool: 15) for Section 3.2.4 (Active learning). For active learning, 6 samples with the highest information gain were selected from the pool set and added to the training set. Before machine learning, data were label encoded for categorical descriptors and min–max scaled. Pairwise Controlled Manifold Approximation Project (PaCMAP33) analysis is employed to visualize the high-dimensional data (see Section 4. PaCMAP in SI). The PaCMAP embedding does not preserve interpretable coordinates; rather, it provides a visualization of the sample distribution in reduced space. The relative proximity between points reflects similarity based on the input descriptors. The categorical descriptor (nitrogen atom position) is omitted for PaCMAP analysis.Because the present dataset is restricted to Pb–I frameworks and limited in size, the analysis focuses on spacer-driven dimensional trends; incorporation of inorganic geometric descriptors and connectivity subclasses represents an important direction for future expansion as larger standardized datasets become available.
2.3. Descriptors
To capture the key molecular factors governing HMH dimensionality, we gathered ten chemically interpretable descriptors that reflect steric, electronic, and geometric characteristics of the spacer cations. These include the number of carbon and nitrogen atoms, rotatable bonds, aromatic rings, hydrophobicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), polar surface area (PSA), aspect ratio, van der Waals volume, nitrogen atom position, and number of branched point (see Section 5. Descriptors in SI, Table S3). The selected descriptors were chosen to represent parameters known to influence intermolecular interactions, packing motifs, and hydrogen-bonding environments in HMHs.34,35 Specifically, steric and geometric descriptors (e.g., aspect ratio, number of branched points, van der Waals volume) capture the spatial constraints affecting layer spacing and packing density, while electronic and polarity-related descriptors (e.g., log
P), polar surface area (PSA), aspect ratio, van der Waals volume, nitrogen atom position, and number of branched point (see Section 5. Descriptors in SI, Table S3). The selected descriptors were chosen to represent parameters known to influence intermolecular interactions, packing motifs, and hydrogen-bonding environments in HMHs.34,35 Specifically, steric and geometric descriptors (e.g., aspect ratio, number of branched points, van der Waals volume) capture the spatial constraints affecting layer spacing and packing density, while electronic and polarity-related descriptors (e.g., log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, PSA) account for variations in hydrogen bonding and ionic interactions that modulate the stability and dimensionality of the resulting structures.36–38 While synthesis conditions can impact crystal formation, because of inconsistent reporting of processing steps in the literature and lack of clear annotated metadata on synthesis in the available database, no descriptors related to synthesis conditions are utilized. As discussed later, this represents a significant opportunity for future research.
P, PSA) account for variations in hydrogen bonding and ionic interactions that modulate the stability and dimensionality of the resulting structures.36–38 While synthesis conditions can impact crystal formation, because of inconsistent reporting of processing steps in the literature and lack of clear annotated metadata on synthesis in the available database, no descriptors related to synthesis conditions are utilized. As discussed later, this represents a significant opportunity for future research.
2.4. Machine learning
Machine learning methods, including tree-based regression and deep learning models, have demonstrated strong empirical success when trained on large-scale datasets.39–41 With sufficient data, these models benefit from asymptotic convergence of parameters, yielding stable estimators without explicitly quantifying uncertainty. However, in materials science applications, the conditions for such asymptotic guarantees are rarely met, as experimental datasets are often small and heterogeneous. Under these circumstances, point estimates of predictive performance can be misleading, and it becomes essential to explicitly capture uncertainty. Quantifying uncertainty is therefore essential for identifying when the model is extrapolating and for guiding decisions based on the reliability of each prediction.26–28 Bayesian Additive Regression Trees (BART)25 offers an potential solution for this setting. BART provides a posterior predictive distribution, enabling uncertainty quantification without relying on restrictive parametric assumptions. While traditional ensembles such as Random Forest (RF)42 can achieve comparable accuracy, they deliver only point estimates by default and lack principled measures of uncertainty.BART constructs its final prediction as an ensemble of shallow decision trees, using a regularization prior which prevents overfitting. During training, it uses Markov chain Monte Carlo to sample from the posterior distribution over tree structures and leaf parameters. Each posterior draw corresponds to one model instance, which in turn produces a prediction; aggregating these draws yields the predictive distribution. From this distribution, we can quantify uncertainty, for example, by computing credible intervals. We demonstrate how BART captures per-sample predictive uncertainty through posterior probability intervals and summarizes model-level performance using a posterior distribution of the area under the receiver operating characteristic curve (AUC).43 AUC is a standard metric used to evaluate how well a classification model distinguishes between two classes. An AUC of 0.5 corresponds to random guessing, whereas an AUC of 1.0 indicates perfect separation. In a Bayesian model, each prediction is expressed as a posterior probability, the estimated likelihood that a given sample belongs to a specific class. Instead of providing a single fixed prediction like RF, the model produces a distribution of possible probabilities, from which credible intervals can be derived. We further show how BART can be applied to active learning. Additional details are provided in SI (see Sections 6. Random forest classifier (RF), 7. Bayesian additive regression trees (BART), and 8. Model interpretation and active learning evaluation).
3. Results and discussion
In the following section, we present the key findings of our study. We first grew single crystals of HMHs based on three chemically distinct yet structurally related branched alkyl ammonium spacers. Structural and optical characterizations of these crystals confirmed the expectation that variations in spacer molecular structure can lead to distinct dimensionalities. Motivated by these observations, we expanded our scope using machine learning. After collecting and curating a dataset (details of curation and data splitting are provided in the Methods section, 2.2. Data handling), we applied BART25 to investigate whether variations in spacer cations could predict 1D versus 2D dimensionalities with the training from this larger dataset. We further analyzed the relative importance of molecular descriptors to identify features that most strongly influence dimensionality. Building on these findings, we implemented an active learning framework to demonstrate its potential utility for guiding future spacer cation selection in experimental studies. Finally, we conducted PaCMAP analysis to explore regions of molecular space where experimental exploration would most improve predictive performance.3.1. Dimensional control via spacer cation structure
As an initial demonstration, we investigated three spacer amines: 3,3-dimethylbutylamine, 2,3-dimethylbutan-2-amine, and N-methylbutan-2-amine (Fig. 1). We combined these spacer amines with PbI2 under identical conditions, resulting in their protonation and insertion as spacer cations in the final HMH structures. While all three amines share the similarities of being aliphatic and branched, the HMH structure outcomes diverged: 3,3-dimethylbutylamine yielded a 2D layered structure based on corner-shared octahedra ((3,3-DMBA·H)2PbI4), while the other two produced 1D chain structures with face-sharing connectivity ((2,3-DMB2A·H)PbI3 and (NMB2A·H)PbI3), as confirmed by X-ray diffraction (Fig. 2, S1–S3 and Tables S1, S2). | ||
| Fig. 1 Choice of amines, where (a) 3,3-dimethylbutylamine (3,3-DMBA), (b) 2,3-dimethylbutan-2-amine (2,3-DMB2A), and (c) N-methylbutan-2-amine (NMB2A). | ||
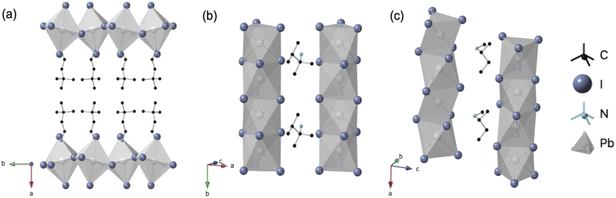 | ||
| Fig. 2 Schematic single-crystal structures of (a) (3,3-DMBA·H)2PbI4, (b) (2,3-DMB2A·H)PbI3, and (c) (NMB2A·H)PbI3. | ||
Optical spectroscopy further reflected this dimensional contrast (see 2.2. Optical properties in SI). The 2D phase exhibited excitonic absorption/emission near 483/491 nm, consistent with typical 2D HMHs.44,45 In contrast, the 1D phases showed blue-shifted absorption/emission (≈385/458 nm and ≈381/414 nm), in line with trends in reduced-dimensional systems46–48 (Fig. S4–S6). Our experiments confirm that systematic variations in spacer molecular structure are sufficient to shift crystal dimensionality and tune optical response. Beyond these primary outcomes, stoichiometry variation may also lead to additional motifs, including corner-sharing and trimer-type 1D phases (Fig. S7–S10; 2.3. Single crystal X-ray diffraction (SCXRD) for additional crystallographic refinements and characterization in SI).
To contextualize the synthesized samples within the overall chemical space, we visualized their locations relative to our entire curated dataset. All data are presented in the PaCMAP visualization (Fig. 3) showing a clear clustering of the complete 113 sample dataset into 3 clusters, where the positions of the synthesized samples are noted by red circles. Fig. S11 shows the molecular feature profiles of the three clusters. Group 1, which includes the three synthesized samples, corresponds to more alkyl-like spacer structures with lower van der Waals volume, fewer aromatic rings, and reduced polar surface area. Group 0 represents molecules with moderate polar surface area, whereas group 2 consists of molecules with higher polar surface areas, multiple nitrogen atoms, and greater flexibility. Although these clusters differ in molecular descriptors and structural descriptors, the distribution of 1D and 2D HMHs is relatively balanced across groups, indicating that the grouping discovered by PaCMAP primarily reflects molecular structural similarity.
Yet, despite the apparent closeness for the 3 synthesized samples, their experimental outcomes diverged. The sensitivity revealed by this discrepancy motivated us to develop a machine learning framework aimed at predicting the dimensionality of HMH phases. It is acknowledged that identical spacer cations can result in different structural configurations (e.g., dimensionality and/or connectivity) depending on the synthesis conditions (e.g., precursor ratio) (Fig. S7 and S8; 2.3. Single crystal X-ray diffraction (SCXRD) in SI for additional crystallographic refinements and characterization). Indeed, in the chemical space examined here, spacer-to-lead stoichiometry represents a experimentally demonstrated control parameter for dimensionality and connectivity in our 3 synthesized samples, as evidenced by the ratio-dependent motif changes observed in Fig. S7–S10. However, because synthesis parameters such as precursor stoichiometry, solvent environment, and temperature are not systematically available across published reports or as tagged metadata in the Hybrid3 database, in the subsequent machine learning analysis synthesis parameters cannot be encoded as features, and we thus assume that each spacer cation corresponds to a single dimensionality. Accordingly, the model should be interpreted as capturing spacer-intrinsic structural tendencies within typical synthetic contexts, rather than predicting a complete synthesis-dependent phase diagram. This assumption of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 cation-dimensionality correspondence may restrict our predictive performance and motivates future efforts toward active data sharing within the community to collect and clearly annotate more comprehensive synthesis and processing information.
1 cation-dimensionality correspondence may restrict our predictive performance and motivates future efforts toward active data sharing within the community to collect and clearly annotate more comprehensive synthesis and processing information.
3.2. Machine learning
In this section, we extend our experimental findings by applying Bayesian additive regression trees. We evaluate model performance and uncertainty quantification (3.2.1. Model performance and uncertainty quantification), analyze feature importance (3.2.2. Feature importance) and apply BART to the synthesized structures (3.2.3. Demonstration on synthesized structures). We further demonstrate how BART can be used to perform active learning (3.2.4. Active learning).To summarize classification performance across all thresholds, we further examined the posterior distribution of the area under the receiver operating characteristic curve (AUC),43 shown in Fig. 4(b). Each value in this distribution corresponds to the AUC computed from one posterior draw of the trained model, reflecting uncertainty in the model's discriminative ability. The posterior mean AUC on the test set was 0.837, with a 95% credible interval of 0.750–0.909.
To qualitatively assess model behavior on the test set, we projected all test samples into a 2D PaCMAP embedding similar to Fig. 3, shown in Fig. S15, coloring points by their ground-truth dimensionality and marking prediction correctness. Note that in this analysis, the newly synthesized 3 experimental samples are simply part of the entire data pool and are thus selected randomly into test or train sets. Although 1D and 2D samples are not completely separated, correct predictions dominate within the major clusters (5/7 for 1D and 20/22 for 2D). Additional diagnostic results suggest satisfactory convergence and mixing across all chains, with effective sample sizes (ESS) generally above 4000 and R-hat values close to 1.00 (Fig. S12 and S13).
The data-driven approach integrates diverse molecular descriptors into a unified quantitative framework, moving beyond intuition-based assessments of individual features, and providing a quantitative context that connects with prior studies emphasizing specific molecular parameters. As PSA increases, the probability of forming a 2D phase rises; this is consistent with reports that N–H⋯I hydrogen bonding and electrostatic interactions between ammonium spacers and halides strengthen interfacial cohesion and interlayer networks in 2D Ruddlesden–Popper phase HMHs.37,45,50 A larger aspect ratio correlates with a higher likelihood of 2D formation: slender, more anisotropic spacer cations promote parallel alignment and dense in-plane packing of the organic bilayers, stabilizing extended 2D slabs; in contrast, bulkier/shorter shapes introduce corrugation and packing frustration that bias chain-like (1D) motifs. This interpretation aligns with studies showing that spacer packing arrangements and orientational order control film structure and energetics, and that linear, longer-chain spacers enhance molecular organization.51,52 In layered HMHs, branching near the ammonium headgroup increases steric demand that can hinder planar tiling and sheet continuity, increasing interlayer spacing and steering Dion–Jacobson and Ruddlesden–Popper structural evolution; in certain chemistries, branched headgroups can also stabilize 1D chains (e.g., isopropylammonium lead iodide).53,54 Thus, while the statistical analysis from these models cannot provide direct mechanistic proof, their ability to capture nonlinear feature interactions and patterns in the data provide researchers with strong evidence to connect to physical mechanisms.
With this data split analysis, the model achieved a posterior mean AUC on the test set of 0.72. Out of the three predictions, two were correct: (3,3-DMBA·H)2PbI4, (2D) and (2,3-DMB2A·H) PbI3 (1D) (Fig. 6). The misclassified case, (NMB2A·H) PbI3 (1D), is noteworthy as the trained model provides an incorrect classification with relatively high confidence. The misclassification of (NMB2A·H)PbI3 highlights an important limitation of relying primarily on steric and geometric descriptors. Although NMB2A exhibits a relatively high aspect ratio and low branching – features that statistically correlate with 2D formation in our dataset – it experimentally stabilizes a 1D face-sharing structure. Notably, NMB2A is the only secondary amine among the three spacers, which may influence hydrogen-bonding geometry and packing interactions. This suggests that steric descriptors alone may not fully capture the balance between organic packing constraints and inorganic framework energetics that governs connectivity selection. Face-sharing motifs may be influenced not only by spacer geometry but also by hydrogen-bonding geometry, lattice strain accommodation, and possible solvent-mediated stabilization. The NMB2A case therefore underscores that dimensionality emerges from a coupled organic–inorganic system, and that spacer-only descriptors capture dominant statistical trends but cannot universally resolve all structural outcomes. Future extensions incorporating additional structural and processing descriptors may help clarify such boundary cases.
These results illustrate the potential practical advantage of uncertainty-driven active learning within this dataset. Reductions in mutual information and log loss correspond to improved confidence and calibration in probabilistic predictions, which in turn enhance the reliability of selecting candidate spacers for experimental validation. In this study, information-gain-based selection identified successful candidates at a higher rate than random sampling across multiple independent trials. While demonstrated within a constrained candidate pool, this result suggests that uncertainty-aware acquisition can improve experimental efficiency in small-data regimes.
To better understand the structural space, we identified representative clusters in Fig. 7 and quantified the net improvement within each cluster (Fig. 8). The cluster feature interpretation is derived from post-hoc descriptor analysis. This analysis was based on the three most influential descriptors based on feature importance (Fig. 5): aspect ratio, polar surface area (PSA), and the number of branch points (branching). Radar plots illustrate the relative magnitude of key descriptors within each cluster. The polygon area and shape highlight which features (e.g., aspect ratio, PSA, and branching) are more dominant in each cluster. The results revealed that clusters A, C, and D exhibited positive net improvements, corresponding to high-aspect ratio molecules (cluster A), structurally balanced molecules (cluster C), and compact, simple molecules (cluster D). In contrast, cluster B, characterized by a high degree of branching and large PSA, showed a negative net improvement. This indicates that targeted experimental sampling within this region of the feature space could enhance the model's representation and predictive performance in subsequent iterations. Within the current dataset, compact, simple, and high-aspect ratio structures appear more favorable for optimization. However, this trend should be interpreted with caution, as highly polar or branched spacers may interact nonlinearly with other descriptors. With more data or expanded features, these spacers could exhibit different behaviors, emphasizing the need to preserve chemical diversity in future active learning cycles. At present, it remains unclear whether such regions reflect intrinsic phase competition or sparse sampling in descriptor space, further underscoring the importance of expanded and condition-aware datasets.
4. Conclusions
In this study, we investigated the dimensionality prediction of low-dimensional Hybrid Metal Halides (HMHs) by combining experimental synthesis and data-driven modeling. Experimentally, we synthesized new low-dimensional HMH crystals based on branched alkyl-based ammonium cations and confirmed that variations in spacer cations influence both the structural dimensionality and the optical properties within this family. Motivated by these observations, we extended the study to a data-centric approach by integrating data from the Hybrid3 repository, literature sources, and newly synthesized structures into a unified dataset. Molecular descriptors were computed using RDKit software and visualized with PaCMAP embedding, which suggested grouping patterns related to spacer-cation properties.Based on this dataset, we developed a machine-learning framework to classify HMH dimensionality. The Bayesian Additive Regression Tree (BART) model achieved a posterior mean AUC of 0.83, which is comparable to that of the random forest baseline. While both models provide similar predictive performance, BART additionally offers posterior probability distributions that quantify prediction confidence. This uncertainty estimation is particularly valuable for small and heterogeneous datasets that are common in experimental materials research. Feature-importance analysis identified three key molecular factors that govern dimensionality: aspect ratio, polar surface area, and the number of branched points. Incorporating an active learning strategy further improved net improvement by approximately 20 percent.
Overall, this uncertainty-aware framework establishes a predictive approach that links molecular structure to emergent dimensionality. The methodology can be extended to larger and more diverse datasets and ultimately provides a foundation for data-driven design of next-generation hybrid materials. The work also motivates a need for more curated experimental data sets with more complete metadata annotation, especially capturing the synthesis and processing features. Given the existence of text based information in the literature, coupled with complexities of defining a standard for data reporting in publications, future work integrating natural language processing-based extraction of synthesis conditions and broader families of spacer cation-inorganic stoichiometries is a promising pathway. Such work would allow inclusion of a richer set of descriptors for future data driven exploration and expand the accessible design space and enhance structural predictability.
Author contributions
M. C.: conceptualization; methodology; software; data curation; formal analysis; investigation; visualization; writing – original draft; writing – review & editing. D. D. C.: methodology; validation; writing – review & editing. R. S.: validation; writing – review & editing. D. B. M.: conceptualization (experimental); supervision (experimental); writing – review & editing. L. C. B.: conceptualization; methodology; supervision; writing – review & editing.Conflicts of interest
There are no conflicts of interest to declare.Data availability
The datasets and machine learning codes supporting this study are available on GitHub at (https://github.com/migonchoi/Perovskite-Dimensionality-Prediction) and Duke Research Data Repository (RDR) (https://doi.org/10.7924/r4n87ht86).CCDC 2490415 ((3,3-DMBA·H)2PbI4), 2490430 ((2,3-DMB2A·H)PbI3) and 2490431 ((NMB2A·H)PbI3) contain the supplementary crystallographic data for this paper.57a–c
Supplementary information (SI): detailed experimental procedures, additional characterization data (XRD, UV-Vis, PL, SCXRD, TGA, and FTIR), crystallographic tables, data processing methods, molecular descriptor definitions, and full descriptions of the machine learning models along with model interpretation and active learning analyses. See DOI: https://doi.org/10.1039/d5ta09980c.
Acknowledgements
M. C. and D. B. M. acknowledge support from National Science Foundation under award number DMR-2323547. R. S. and L. C. B. acknowledge support from National Science Foundation under award number DGE-2022040. We thank Dr Yi Xie for guidance in selecting the three spacer cations and for early support with crystal growth, SCXRD, and structural refinement. We also thank Dr Rayan Chakraborty for assistance with crystal structure refinement.References
- D. Sivan, K. Satheesh Kumar, A. Abdullah, V. Raj, I. I. Misnon, S. Ramakrishna and R. Jose, J. Mater. Sci., 2024, 59, 2602–2643 CrossRef CAS.
- R. Ramprasad, R. Batra, G. Pilania, A. Mannodi-Kanakkithodi and C. Kim, npj Comput. Mater., 2017, 3, 54 CrossRef.
- X. Liu, H. Yan, Z. Shu, X. Cui and Y. Cai, Nanoscale, 2025, 17, 2658–2667 RSC.
- Q. Liu, X. Wu, A. H. Coffey, H. Yang, J. Y. Park, Q. Hu, K. Ma, C. Zhu, Y. S. Zhao, L. Dou and K. Wang, Chem. Commun., 2025, 61, 7644–7647 RSC.
- P. Du, Y. Zhao, P. Song, S. Yang, S. Liu, J. Zhang, H. Cai, J. Ni and J. Li, J. Mater. Sci.: Mater. Electron., 2024, 35, 1870 CrossRef CAS.
- Z. Lu, X. Xu, Y. Gao, Z. Wu, A. Li, Z. Zhan, Y. Qu, Y. Cai, X. Huang, J. Huang, Z. Zhang, T. Luo, L. Peng, P. Liu, T. Shi and W. Xie, Surf. Interfaces, 2022, 34, 102343 CrossRef CAS.
- R. J. D. Tilley, MRS Bull., 2017, 42, 325 CrossRef.
- D. B. Mitzi, J. Chem. Soc., Dalton Trans., 2001, 1–12, 10.1039/b007070j.
- M. A. Green, A. Ho-Baillie and H. J. Snaith, Nat. Photonics, 2014, 8, 506–514 CrossRef CAS.
- S. Rajukkannu, W. Bunpheng, R. Dhairiyasamy and V. Gopinath, Sci. Rep., 2025, 15, 833 CrossRef CAS PubMed.
- N. K. Elangovan, R. Kannadasan, B. B. Beenarani, M. H. Alsharif, M.-K. Kim and Z. Hasan Inamul, Energy Rep., 2024, 11, 1171–1190 CrossRef.
- S. Khatoon, S. Kumar Yadav, V. Chakravorty, J. Singh, R. Bahadur Singh, M. S. Hasnain and S. M. M. Hasnain, Mater. Sci. Energy Technol., 2023, 6, 437–459 CAS.
- A. Younis, C.-H. Lin, X. Guan, S. Shahrokhi, C. Y. Huang, W. Yutao, T. He, S. Singh, L. Hu, J. Retamal, J.-H. He and T. Wu, Adv. Mater., 2021, 33, 2005000 CrossRef CAS PubMed.
- Y. Gao, Z. Song, Q. Fu, Y. Chen, L. Yang, Z. Hu, Y. Chen and Y. Liu, Adv. Mater., 2024, 36, 2405921 CrossRef CAS PubMed.
- A. Mahapatra, V. Anilkumar, A. Scarperi, D. J. Kubicki, P. Yadav, M. Mączka and D. Prochowicz, ACS Photonics, 2024, 11, 5091–5099 CrossRef CAS.
- X. Li, S. Aftab, S. Hussain, F. Kabir, A. M. A. Henaish, A. G. Al-Sehemi, M. R. Pallavolu and G. Koyyada, J. Mater. Chem. A, 2024, 12, 4421–4440 RSC.
- K. R. Hansen, C. Y. Wong, C. E. McClure, B. Romrell, L. Flannery, D. Powell, K. Garden, A. Berzansky, M. Eggleston, D. J. King, C. M. Shirley, M. C. Beard, W. Nie, A. Schleife, J. S. Colton and L. Whittaker-Brooks, Matter, 2023, 6, 3463–3482 CrossRef CAS.
- Y. Shen, S. Hu, Y. Meng, S. Yip and J. C. Ho, Mater. Today Electron., 2024, 8, 100100 CrossRef.
- X. Li, J. M. Hoffman and M. G. Kanatzidis, Chem. Rev., 2021, 121, 2230–2291 CrossRef CAS PubMed.
- E. Fransson, J. Wiktor and P. Erhart, ACS Energy Lett., 2024, 9, 3947–3954 CrossRef CAS.
- R. Lyu, C. E. Moore, T. Liu, Y. Yu and Y. Wu, J. Am. Chem. Soc., 2021, 143, 12766–12776 CrossRef CAS PubMed.
- Y. Mai, J. Tang, H. Meng, X. Li, M. Liu, Z. Chen, P. Zhang and S. Li, Adv. Compos. Hybrid Mater., 2024, 7, 104 CrossRef CAS.
- E. I. Marchenko, S. A. Fateev, A. A. Petrov, V. V. Korolev, A. Mitrofanov, A. V. Petrov, E. A. Goodilin and A. B. Tarasov, Chem. Mater., 2020, 32, 7383–7388 CrossRef CAS.
- W. Zhang, J. Zhang, C. Zhang, R. Dong, Y. Xu, Z. Yang, J. Zheng, D. Chen, Q. Liu, W. Yang and M.-H. Shang, J. Mater. Chem. A, 2025, 13, 19670–19681 RSC.
- H. A. Chipman, E. I. George and R. E. McCulloch, arXiv, 2010, preprint, arXiv:0806.3286, DOI:10.48550/arXiv.0806.3286.
- F. Tavazza, B. DeCost and K. Choudhary, ACS Omega, 2021, 6, 32431–32440 CrossRef CAS PubMed.
- Y. Wang and D. L. McDowell, in Uncertainty Quantification in Multiscale Materials Modeling, ed. Y. Wang and D. L. McDowell, Woodhead Publishing, 2020, pp. 1–40, DOI:10.1016/B978-0-08-102941-1.00001-8.
- G. Tom, R. J. Hickman, A. Zinzuwadia, A. Mohajeri, B. Sanchez-Lengeling and A. Aspuru-Guzik, Digital Discovery, 2023, 2, 759–774 RSC.
- S.-Y. Ding and W. Wang, Chem. Soc. Rev., 2013, 42, 548–568 RSC.
- K. S. Novoselov, A. Mishchenko, A. Carvalho and A. H. Castro Neto, Science, 2016, 353, aac9439 CrossRef CAS PubMed.
- P. Xu, X. Ji, M. Li and W. Lu, npj Comput. Mater., 2023, 9, 42 CrossRef.
- R. Chang, Y.-X. Wang and E. Ertekin, npj Comput. Mater., 2022, 8, 242 CrossRef.
- Y. Wang, H. Huang, C. Rudin and Y. Shaposhnik, J. Mach. Learn. Res., 2021, 22, 1–73 Search PubMed.
- J. Choi, J. Kim, M. Jeong, B. Park, S. Kim, J. Park and K. Cho, Small, 2024, 20, 2405598 CrossRef CAS PubMed.
- P. Liu, X. Li, T. Cai, W. Xing, N. Yang, H. Arandiyan, Z. Shao, S. Wang and S. Liu, Nano–Micro Lett., 2024, 17, 35 CrossRef PubMed.
- S. Bhattacharya and A. Roy, Comput. Mater. Sci., 2024, 231, 112581 CrossRef CAS.
- L. Lin, T. W. Jones, T. C.-J. Yang, X. Li, C. Wu, Z. Xiao, H. Li, J. Li, J. Qian, L. Lin, J. Q. Shi, S. D. Stranks, G. J. Wilson and X. Wang, Matter, 2024, 7, 38–58 CrossRef CAS.
- J. Li and P. Rinke, Phys. Rev. B, 2016, 94, 045201 CrossRef.
- A. Dunn, Q. Wang, A. Ganose, D. Dopp and A. Jain, npj Comput. Mater., 2020, 6, 138 CrossRef.
- L. Ward, A. Agrawal, A. Choudhary and C. Wolverton, npj Comput. Mater., 2016, 2, 16028 CrossRef.
- P. Reiser, M. Neubert, A. Eberhard, L. Torresi, C. Zhou, C. Shao, H. Metni, C. van Hoesel, H. Schopmans, T. Sommer and P. Friederich, Commun. Mater., 2022, 3, 93 CrossRef CAS PubMed.
- L. Breiman, Mach. Learn., 2001, 45, 5–32 CrossRef.
- J. A. Hanley and B. J. McNeil, Radiology, 1982, 143, 29–36 CrossRef CAS PubMed.
- D. Babaian, D. Hill, P. Yu and S. Guha, J. Mater. Chem. C, 2025, 13, 193–202 RSC.
- X. Gao, X. Zhang, W. Yin, H. Wang, Y. Hu, Q. Zhang, Z. Shi, V. L. Colvin, W. W. Yu and Y. Zhang, Adv. Sci., 2019, 6, 1900941 CrossRef CAS PubMed.
- K. Fedoruk, S. J. Zelewski, J. K. Zaręba, M. Ptak, M. Mączka and A. Sieradzki, J. Mater. Chem. C, 2022, 10, 10519–10529 RSC.
- Z. Yuan, C. Zhou, Y. Tian, Y. Shu, J. Messier, J. C. Wang, L. J. van de Burgt, K. Kountouriotis, Y. Xin, E. Holt, K. Schanze, R. Clark, T. Siegrist and B. Ma, Nat. Commun., 2017, 8, 14051 CrossRef CAS PubMed.
- Y.-Y. Guo, L.-J. Yang and P. Lightfoot, Crystals, 2019, 9, 616 CrossRef.
- S. M. Lundberg and S.-I. Lee, Adv. Neural Inf. Process. Syst., 2017, vol. 30 Search PubMed.
- S. Guo, W. Mihalyi-Koch, Y. Mao, X. Li, K. Bu, H. Hong, M. P. Hautzinger, H. Luo, D. Wang, J. Gu, Y. Zhang, D. Zhang, Q. Hu, Y. Ding, W. Yang, Y. Fu, S. Jin and X. Lü, Nat. Commun., 2024, 15, 3001 CrossRef CAS PubMed.
- J. Hu, I. W. H. Oswald, S. J. Stuard, M. M. Nahid, N. Zhou, O. F. Williams, Z. Guo, L. Yan, H. Hu, Z. Chen, X. Xiao, Y. Lin, Z. Yang, J. Huang, A. M. Moran, H. Ade, J. R. Neilson and W. You, Nat. Commun., 2019, 10, 1276 CrossRef PubMed.
- S. Wang, S. Kalyanasundaram, L. Gao, Z. Ling, Z. Zhou, M. Bonn, P. W. M. Blom, H. I. Wang, W. Pisula and T. Marszalek, Mater. Horiz., 2024, 11, 1177–1187 RSC.
- K. Fedoruk-Piskorska, J. K. Zaręba, S. J. Zelewski, A. Gągor, M. Mączka, S. Drobczyński and A. Sieradzki, ACS Appl. Mater. Interfaces, 2024, 16, 28829–28837 CrossRef CAS PubMed.
- W. Li, J. He, Y. Zhang, L. Ye, G. Yao, Z. Zhang, S. Li, X. Lu, H. Lu, T. Zeng and Z. Yang, Cell Rep. Phys. Sci., 2025, 6, 102509 CrossRef CAS.
- B. Settles, Active Learning Literature Survey, Technical Report #1648, 2009 Search PubMed.
- N. Houlsby, F. Huszár, Z. Ghahramani and M. Lengyel, arXiv, 2011, preprint, arXiv:1112.5745, DOI:10.48550/arXiv.1112.5745.
- (a) CCDC 2490415: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2plgzr; (b) CCDC 2490430: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2plhg8; (c) CCDC 2490431: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2plhh9.
| This journal is © The Royal Society of Chemistry 2026 |






