 Open Access Article
Open Access ArticleGraph neural network-based multi-objective Bayesian optimization for enhanced screening of metal–organic frameworks with optimal separation performance
Lane E.
Schultz
 a,
Nickolas
Gantzler
b,
N. Scott
Bobbitt
a,
Nickolas
Gantzler
b,
N. Scott
Bobbitt
 b,
Dorina F.
Sava Gallis
b,
Dorina F.
Sava Gallis
 c and
Rémi
Dingreville
c and
Rémi
Dingreville
 *a
*a
aCenter for Integrated Nanotechnologies (CINT), Sandia National Laboratories, Albuquerque, NM 87185, USA. E-mail: rdingre@sandia.gov
bMaterial, Physical, and Chemical Sciences Center, Sandia National Laboratories, Albuquerque, NM 87185, USA
cNanoscale Sciences Department, Sandia National Laboratories, Albuquerque, NM 87185, USA
First published on 9th February 2026
Abstract
Metal–organic frameworks (MOFs) are porous crystalline materials with applications in gas capture, drug delivery, and molecular separations. While high-throughput computational screening has traditionally identified promising MOFs, recent advances increasingly harness machine learning to accelerate discovery and screening. Existing optimization methods such as Bayesian optimization (BO) and genetic algorithms often overlook the detailed structure–property relationships critical to MOF performance. Here, we present an optimization workflow that couples graph neural networks (GNNs) with multi-objective BO to enhance MOF discovery and screening. By representing MOFs as graphs embedding atomic- and structural-level features, GNNs capture intricate structure–property correlations, enabling more accurate property predictions than traditional methods relying solely on macroscopic descriptors. Our integrated framework efficiently identifies Pareto-optimal MOF candidates tailored for improved separation of alkanes, alkenes, alcohols, and aromatics, demonstrating the significant advantage of graph-based models in materials optimization workflows.
1 Introduction
Metal–organic frameworks (MOFs) are hybrid materials composed of two fundamental building blocks: (i) single metal ions or polynuclear metal clusters (secondary building units) and (ii) organic ligands that connect these units to form three-dimensional porous structures.1–3 MOFs demonstrate exceptional utility across diverse applications,2 including gas storage,4 separations,5,6 sensing,7,8 luminescence,9,10 environmental remediation,11 catalysis,12,13 and biomedical applications,14–16 to name a few.8,17 In sensing applications, the ability to selectively distinguish one analyte from others is crucial, and MOFs have demonstrated significant potential for volatile organic compound (VOC) detection through selective adsorption of specific target compounds over competing species.8Due to their versatility, MOFs have garnered significant attention, with researchers actively seeking MOFs with enhanced properties for targeted applications. Given their modular construction and the vast chemical space of different nodes, linkers, and topologies, there are a large number of possible MOFs, meaning they can be chosen or carefully designed for specific applications. Specialized databases containing thousands of MOFs have emerged, including the hMOF database,18 ToBaCCo,19ab initio REPEAT charge MOF (ARC-MOF),20,21 computation-ready experimental MOF (CoRE MOF),22 and MOSAEC MOF database (MOSAEC-DB)23 to name a few. ARC-MOF20 features MOFs with density functional theory (DFT) calculated charges and some experimentally synthesized samples. CoRE MOF22 stores experimentally synthesized MOFs along with some material properties. MOSAEC-DB23 includes curated MOFs that were verified by the physical plausibility of metal oxidation states. These extensive databases provide researchers with opportunities for MOF property optimization on more physically feasible structures. While databases like CoRE MOF and ARC-MOF have aggregated tens of thousands of structures, they represent a vast and noisy chemical space. Brute-force high-throughput computational screening of every candidate for every possible application is computationally intractable. The generation of new MOFs has evolved dramatically from basic combinatorial approaches where ligands and secondary building units are interchanged between known structures,18 to sophisticated machine-learning (ML) techniques such as variational autoencoders that explore latent spaces to produce innovative structures.24 The primary bottleneck in materials discovery is no longer generation but rather the efficient screening and rapid identification of the highest-performing, Pareto-optimal candidates from these massive existing databases. Therefore, we utilize the well-established CoRE MOF22 2019 database for much of our work, which provides experimentally validated structures as a reliable foundation. We also use the ARC-MOF database.20,21
Various computational approaches have been employed to search for MOFs with optimal properties,25 including hierarchical screening with ML predictions26 and genetic algorithms.27 Bayesian optimization (BO) has emerged as a particularly effective method, utilizing surrogate models and acquisition functions that balance exploitation (maximizing target properties if greater is better) and exploration (investigating uncertain areas) to efficiently identify promising candidates. Researchers have demonstrated that BO can rapidly screen optimal materials from large search spaces, sometimes requiring only a few proposed materials to find optimal candidates.28 Advanced implementations incorporate multi-fidelity approaches that balance computational cost with accuracy for a similar class of material called covalent organic frameworks,29 and diversity promoting mechanisms like the Vendi score to ensure broad exploration of a material space.30 Others optimize multiple properties simultaneously like in Comlek et al.,31 where MOFs with favorable CO2 working capacity and CO2 over N2 selectivity were selected. Many approaches to MOF discovery employ BO with MOFs represented in a tabular format (i.e., representations where each row corresponds to a material and each column to a feature or property). Tabular formats emphasize macroscopic properties such as pore size and surface area.32–34 While useful, these simplified descriptors overlook critical structure–property relationships at the atomic scale inherent in graph-based representations. They fail to capture the local chemical environment of the metal node, the precise geometry of the organic linker, or the material's topology. Even strategies utilizing Simplified Molecular Input Line Entry System (SMILES) strings encoded as vectors, such as those discussed in Liu et al.,30 may not inherently capture atomic properties like local chemistry or geometry. In contrast, graph neural networks (GNNs) that integrate both structural and chemical information have demonstrated superior performance in property prediction compared to their tabular counterparts, as highlighted by Luo et al.35 and Li et al.36 Both sources indicate that graph-based models achieve better accuracy in predicting properties than models trained on tabular data. Luo et al.35 focused on atomic charge prediction while Li et al.36 focused on prediction of CH4 over N2 selectivity. The integration of property optimization via BO with a GNN serving as the surrogate model holds significant promise for MOFs, offering a way to exploit complex structural and chemical information.
MOFs have been widely studied for chemical separations.5,6 One of the most difficult challenges in adsorptive separations is to distinguish between molecules with similar chemical or geometric characteristics. When a MOF exhibits strong affinity for a specific molecule, it often shows comparable affinity for structurally or chemically related species, making selective adsorption difficult. Consequently, a tradeoff in selectivity inevitably arises, and this tradeoff becomes increasingly pronounced as molecular similarity increases. In this work, we integrate a GNN-based structural representation of MOFs with a BO workflow to identify and screen MOFs capable of separating chemically similar organic molecules (e.g., alcohols, butenes, benzene/toluene). This integrated approach enables the discovery of Pareto-optimal MOFs across multiple challenging separation tasks, effectively prioritizing candidates that optimally balance selectivity between closely related molecules.
We utilized structures from the ARC-MOF20,21 v6 and CoRE MOF22 2019 Version 1.1.3 databases to search for Pareto-optimal MOFs. Each MOF was uniquely characterized using Voronoi polyhedra (VP), which defines the connectivity employed in a graph representation of the framework. These graphs incorporated atomic-level chemical information, local atomic neighborhood environments, and global MOF properties. Using these graph representations, surrogate GNNs were trained on subsets of MOFs and subsequently applied within a BO framework to explore the remaining search space, assessing whether the GNNs effectively identified Pareto-front materials. We validated this approach by comparing multiple model types with our GNN BO workflow to identify MOFs with optimal CH4/N2 selectivity using the data of Li et al.36 Separations were quantified by maximizing the Henry constant of the target species while minimizing those of competing molecules. The Henry coefficient, which represents the proportionality between gas adsorption and pressure in the low-pressure limit, was used as an indicator of affinity toward specific adsorbates. Optimization tasks included one, two, three, and four simultaneous chemical objectives. The GNN-based candidate selection consistently outperformed random selection across most test cases. Feature ablation analyses provided interpretability, revealing the respective contributions of edge-, node-, and global-level features to model predictions. The present framework not only captures structure-to-property mapping but also enables automated discovery of materials with optimized performance. The current search space, however, remains limited by the availability of structures within existing databases.
2 Computational methods
This section details the integrated computational workflow for screening MOF databases, generating the associated targeted Henry coefficients, and optimizing multi-property adsorption performance. This workflow consists of several elements. After curating structural datasets, Henry coefficients were computed as central descriptors for affinity and selectivity. MOF structures were featurized as graphs with rich node, edge, and global attributes. GNN-based surrogates were constructed and optimized to predict material properties and uncertainties, supported by feature ablation interpretation and benchmarking against Gaussian process (GP) surrogates. These surrogate models were iteratively coupled with BO to efficiently identify MOFs with optimal adsorption properties, using acquisition functions and performance metrics for both single- and multi-objective searches. The sub-sections below describe each of these steps in more detail.2.1 MOF database
We used a subset of 77![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 482 and 18
482 and 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 522 MOF structures from the ARC-MOF20,21 v6 and CoRE MOF 2019 databases,22,37 respectively. The ARC-MOF collection includes both experimentally synthesized and computationally generated frameworks, along with DFT-computed atomic charges and several macroscopic MOF properties. The CoRE MOF database comprises experimentally realized MOFs that have been curated for atomistic modeling by addressing issues such as partial occupancies, symmetry inconsistencies, and the presence of solvent molecules. The structural information for each MOF in both datasets is encoded in a crystallographic information file (CIF) containing the element type, site indices, and three-dimensional atomic spatial coordinates. From CoRE MOF, subsets were further filtered to only retain structures with a pore limiting diameter greater than the largest kinetic diameter of adsorbed VOCs. See the SI for tabulated kinetic diameters. We note that these datasets primarily comprise ‘computationally ready’ versions of known structures. While such readiness increases the likelihood that our ‘optimal’ materials can be synthesized, it also means that the model may have lower predictive confidence in ‘dark’ chemical spaces, such as those involving highly reactive open-metal sites or non-traditional coordination geometries that are underrepresented in the training data.
522 MOF structures from the ARC-MOF20,21 v6 and CoRE MOF 2019 databases,22,37 respectively. The ARC-MOF collection includes both experimentally synthesized and computationally generated frameworks, along with DFT-computed atomic charges and several macroscopic MOF properties. The CoRE MOF database comprises experimentally realized MOFs that have been curated for atomistic modeling by addressing issues such as partial occupancies, symmetry inconsistencies, and the presence of solvent molecules. The structural information for each MOF in both datasets is encoded in a crystallographic information file (CIF) containing the element type, site indices, and three-dimensional atomic spatial coordinates. From CoRE MOF, subsets were further filtered to only retain structures with a pore limiting diameter greater than the largest kinetic diameter of adsorbed VOCs. See the SI for tabulated kinetic diameters. We note that these datasets primarily comprise ‘computationally ready’ versions of known structures. While such readiness increases the likelihood that our ‘optimal’ materials can be synthesized, it also means that the model may have lower predictive confidence in ‘dark’ chemical spaces, such as those involving highly reactive open-metal sites or non-traditional coordination geometries that are underrepresented in the training data.
2.2 Henry constants and the setup for optimization
The adsorption affinity of a gas in the dilute limit can be described by the Henry coefficient which quantifies the proportionality between the amount adsorbed and its partial pressure. This metric offers a convenient and computationally efficient means to evaluate host–guest interactions under infinite dilution, where adsorbate–adsorbate interactions are negligible. Because Henry coefficients capture intrinsic framework–adsorbate affinities, they serve as an effective descriptor for screening materials in early-stage separation studies. Mathematically, Henry's law defines the coefficient as, | (1) |
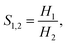 | (2) |
Finding a single material that optimizes multiple selectivities is often infeasible due to inherent trade-offs in material properties. Instead, a set of non-dominated solutions, known as the Pareto front, can be identified for multi-objective optimization. The Pareto front represents solutions that cannot be improved in one objective without degrading another.39 Mathematically, a solution yi belongs to the Pareto front if no other solution yj dominates it:
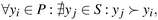 | (3) |
| Dataset | Structures | Objective | Family | Min/Max |
|---|---|---|---|---|
| {1-propanol} | 3634 | 1-Propanol | Alcohol | Min |
| {Butane} | 3653 | Butane | Alkane | Max |
| {Benzene, toluene} | 2125 | Benzene | Aromatic | Max |
| Toluene | Aromatic | Min | ||
| {1-propanol, butane} | 3634 | 1-propanol | Alcohol | Min |
| Butane | Alkane | Max | ||
| {1-propanol, 2-propanol} | 3634 | 1-propanol | Alcohol | Max |
| 2-propanol | Alcohol | Min | ||
| {1-propanol, propane} | 3634 | 1-propanol | Alcohol | Min |
| Propane | Alkane | Max | ||
| {2-cis-butene, 2-trans-butene} | 3260 | 2-cis-butene | Alkene | Max |
| 2-trans-butene | Alkene | Min | ||
| {Butane, propane} | 3653 | Butane | Alkane | Max |
| Propane | Alkane | Min | ||
| {1-propanol, 2-propanol, butane} | 3634 | 1-propanol | Alcohol | Min |
| 2-propanol | Alcohol | Min | ||
| Butane | Alkane | Max | ||
| {1-propanol, 2-propanol, propane} | 3634 | 1-propanol | Alcohol | Min |
| 2-propanol | Alcohol | Min | ||
| Propane | Alkane | Max | ||
| {1-propanol, 2-propanol, butane, propane} | 3634 | 1-propanol | Alcohol | Min |
| 2-propanol | Alcohol | Min | ||
| Butane | Alkane | Max | ||
| Propane | Alkane | Max |
2.3 Data generation for volatile organic compounds Henry coefficients
Henry coefficient data were generated using Widom insertions simulations implemented in RASPA40 v2, closely following the methods described in Bobbitt et al.41 RASPA is an open-source molecular simulation software designed for adsorption and diffusion studies in molecules such as MOFs. Each simulation was performed for 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 insertion cycles per adsorbate. MOF structures were held rigid at their crystallographic minimum positions. Non-bonded interactions were treated using 12–6 Lennard-Jones potentials with parameters taken from the universal force field (UFF)42 for MOFs and TraPPE43–46 for the adsorbates. For most adsorbates, the united atom model was applied, while for benzene and toluene we employed rigid 12-site models with explicit charges. The toluene model was adapted from benzene by replacing a hydrogen atom with an sp3 methyl group. Interaction cross terms were determined using Lorentz–Berthelot mixing rules. Partial charges on the MOFs were computed using the EQeq47 method as implemented in RASPA and treated using Ewald summations. All simulations used a 12.0 Å cutoff radius and 300 K temperature, with simulation boxes duplicated as needed to ensure dimensions at least twice this value in all directions. Ideal gas Rosenbluth weights were pre-computed at 300 K. Given the wide range of values in Henry coefficients, we took their natural logarithms for subsequent analysis. All Lennard-Jones parameters and molecule structures are provided as RASPA definition files in the repository mentioned in the Data availability section at the end of this article.
000 insertion cycles per adsorbate. MOF structures were held rigid at their crystallographic minimum positions. Non-bonded interactions were treated using 12–6 Lennard-Jones potentials with parameters taken from the universal force field (UFF)42 for MOFs and TraPPE43–46 for the adsorbates. For most adsorbates, the united atom model was applied, while for benzene and toluene we employed rigid 12-site models with explicit charges. The toluene model was adapted from benzene by replacing a hydrogen atom with an sp3 methyl group. Interaction cross terms were determined using Lorentz–Berthelot mixing rules. Partial charges on the MOFs were computed using the EQeq47 method as implemented in RASPA and treated using Ewald summations. All simulations used a 12.0 Å cutoff radius and 300 K temperature, with simulation boxes duplicated as needed to ensure dimensions at least twice this value in all directions. Ideal gas Rosenbluth weights were pre-computed at 300 K. Given the wide range of values in Henry coefficients, we took their natural logarithms for subsequent analysis. All Lennard-Jones parameters and molecule structures are provided as RASPA definition files in the repository mentioned in the Data availability section at the end of this article.
Regarding the representativeness of Henry coefficients as indicators of separation performance, we note that Henry's law strictly applies only at low adsorbate concentrations. Nevertheless, Henry coefficients remain informative for low-concentration applications such as VOC capture or chemical sensing.48,49 They are also valuable descriptors for dilute gas separations, consistent with prior studies.50 Moreover, Henry coefficients computed via Widom insertions capture a comprehensive average of adsorbate–framework interactions by sampling the entire pore environment. Accounting for higher-concentration behavior would require explicitly modeling adsorbate–adsorbate interactions, which become non-negligible for species such as alcohols. Incorporating these effects would entail substantial methodological extensions that we identify as an important avenue for future work.
2.4 MOF featurization
 | ||
| Fig. 1 Conceptual framework for graph-based representation of metal–organic frameworks (MOFs). A MOF from CoRE MOF22,37 is shown in panel (a) along with a subset of calculated properties. Panel (b) presents an abstraction of a material structure. Panel (c) illustrates the corresponding graph representation, where atoms are represented as nodes and connections to neighbors as edges. Panel (d) depicts the global features that characterize the entire graph, the node features that encode atomic properties, and the edge features that capture connectivity. | ||
The connections between nodes forming the graph structure were defined using VP, a geometric construction that partitions space around each atom by defining bisecting planes between the central atom and its neighboring atoms. Each MOF structure provided as a CIF was loaded into pymatgen51 as a structural object. The nearest-neighbor network was constructed using pymatgen's IsayevNN class, applying a bond distance tolerance of 0.5 Å without imposing restrictions on the number of Voronoi neighbors.51,52 This strategy, inspired by Luo et al.,35 defines an edge between two atoms if (i) their distance is less than or equal to the sum of their Cordero covalent radii plus a distance tolerance and (ii) they share a common Voronoi facet. The 0.5 Å tolerance was specifically chosen to prevent isolated nodes, as noted in Luo et al.35 Thus each MOF was represented as a graph in which atomic connectivity is encoded through VP-based spatial relationships.
Other featurizations exist. We acknowledge that the smooth overlap of atomic positions (SOAP)53 approach is a powerful method for encoding local atomic information in structures. SOAP representations are differentiable with respect to atomic positions (crucial for molecular dynamics simulations) while maintaining the invariance properties characteristic of graph representations. However, the original SOAP formulation incorporates only positional and element type information and does not incorporate additional chemical descriptors such as electronegativity. In contrast, graph representations offer greater flexibility, allowing the seamless inclusion of diverse chemical and structural features within a unified framework.
Chemical and structural properties of each atomic site were also incorporated as node features. These properties were obtained from two sources: descriptors extracted through pymatgen51 and features adopted from Luo et al.35 From pymatgen, each node was encoded with elemental attributes including Mendeleev number, atomic mass, number of electrons, electron affinity, and the number of electrons in each orbital of the atom up to the 7p orbital (with unoccupied orbitals assigned a value of zero). The VP geometry of each atom was represented using Schläfli notation, which records the specific number of enclosing faces and the number of vertices per face. For example, an atom with nine enclosing faces, where three faces have four vertices and the remaining six have five vertices, is represented as (0, 3, 6), where the index indicates vertex count and is the number of faces. Faces with up to 10 vertices were considered, following the hard-coded constraint in pymatgen's VoronoiAnalyzer. In total, 31 node features were obtained from the properties described above, while additional features were included using modified code and parameterizations provided by Luo et al.35
As in Luo et al.,35 each node was further enriched with atomic properties from Mendeleev,54 including atomic number, atomic weight, van der Waals radius, first ionization energy, Cordero covalent radius, Rahm atomic radius, static dipole polarizability, and Ghosh electronegativity (8 features). Beyond these direct attributes, a set of local environment descriptors were computed to capture geometric and chemical interactions between neighboring atoms.
The property weighted radial distribution functions (pwRDFs)55 quantify how a given atomic property P varies radially around each atom i:
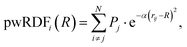 | (4) |
The property weighted angular distribution functions (pwADFs) were computed analogously to describe three-body correlations between atom i connected to atoms j and k:
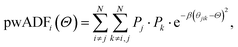 | (5) |
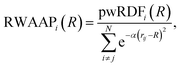 | (6) |
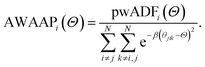 | (7) |
The weighed radial atom centered symmetry function (wRACSF)35,56,57 was also included,
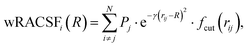 | (8) |
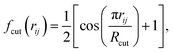 | (9) |
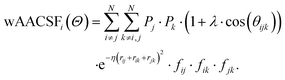 | (10) |
Finally, shell features35,58 characterized property distributions over coordination shells up to the fourth order. We first define what a coordination shell for an atom i represents. The first coordination shell consists of the immediate neighbors surrounding an atom. The second coordination shell encompasses all atoms connected to the first neighbors of atom i. The third coordination shell includes all atoms connected to the second coordination shell atoms, and this pattern continues for higher-order shells. For each atom in a MOF and for each property P and shell n, we computed the average, absolute difference, and standard deviation:
 | (11) |
 | (12) |
 | (13) |
Global features were classified into two categories: material-level and graph-based descriptors. Material-level features were computed from structural and pore geometry data, while graph-based features were derived exclusively from the topological representation of each MOF. Material-level properties, calculated using pymatgen,51 include volume, density, the number of atoms, the number of each element type in a structure normalized by the total number of atoms, the average VP volume for all atoms, and the average number of atom neighbors. Additional geometrical descriptors evaluated with Zeo++,59 include the diameter of the largest included sphere, diameter of largest free sphere, diameter of largest included sphere along the free sphere path, both available and unavailable surface area, and both the available and unavailable void fraction. A spherical probe radius of N2 was used. Graph-based descriptors, computed using NetworkX,60 describe structural connectivity and topology. These include undirected graph density, diameter, degree assortativity coefficient, and average shortest path length. The graph density is defined as  where e and n are the number of edges and nodes respectively and describes the ratio of actual connections to possible connections in the graph. The graph diameter and radius correspond to the maximum and minimum eccentricities across all nodes, representing the farthest and nearest network spans, respectively. Unconnected nodes were excluded from distance calculations. The degree assortativity coefficient is the Pearson correlation coefficient of the degree between pairs of linked nodes, while the average shortest path length is given by
where e and n are the number of edges and nodes respectively and describes the ratio of actual connections to possible connections in the graph. The graph diameter and radius correspond to the maximum and minimum eccentricities across all nodes, representing the farthest and nearest network spans, respectively. Unconnected nodes were excluded from distance calculations. The degree assortativity coefficient is the Pearson correlation coefficient of the degree between pairs of linked nodes, while the average shortest path length is given by
 | (14) |
2.5 Construction, optimization, and feature interpretation of graph neural networks for multi-property regression
Efficient screening of a large set of MOFs with optimized performance involves evaluating thousands of candidates across multiple objectives, which is computationally demanding. BO provides a data-efficient framework for this purpose by iteratively selecting new candidates based on model predictions and associated uncertainties. To perform BO, a surrogate model capable of predicting target properties and quantifying its own uncertainty is essential. While GPs naturally satisfy these requirements, they are computationally expensive and scale poorly with data size. To overcome these limitations, we developed a neural surrogate model based on leveraging the flexible nature of neural networks (NNs), which can learn non-linear structure–property relationships while estimating prediction uncertainties. This framework combines the expressive power of NNs with the uncertainty awareness critical for BO-driven materials screening. Our implementation of GNNs utilizes the PyTorch Geometric (PyG)61 library.The GNN adopted here follows the formulation introduced by Battaglia et al.62 and implemented through the MetaLayer class in PyG. In this architecture, edge, node, and global sub-models learn (from features scaled with scikit-learn's RobustScaler63) complementary hierarchical representations that capture multi-scale structural information within graphs. The latent features extracted from these components are subsequently processed through additional neural layers to infer material properties and their uncertainties. Fig. 2 provides an abstraction of our GNN, showing how multi-level graph features (global, node, and edge) are encoded into latent representations for predicting target properties with uncertainties.
 , connecting nodes
, connecting nodes  and
and  , given a global feature vector,
, given a global feature vector,  , is defined as,
, is defined as, | (15) |
 denotes the new representation of edges. The input layer dimensionality of ϕe is equal to twice the number of node features (comprising features from source and destination nodes), plus the number of edge features and global features. All hidden layers and the output layer maintain consistent width, with rectified linear unit (ReLU) activations and batch normalization between layers. The number of layers and neurons were later optimized, as described in Section 2.5.3.
denotes the new representation of edges. The input layer dimensionality of ϕe is equal to twice the number of node features (comprising features from source and destination nodes), plus the number of edge features and global features. All hidden layers and the output layer maintain consistent width, with rectified linear unit (ReLU) activations and batch normalization between layers. The number of layers and neurons were later optimized, as described in Section 2.5.3.
The node sub-model builds upon latent edge representations to encode neighborhood effects. For each node, latent edge vectors directed toward it are aggregated using the PyG scatter function, combined with its intrinsic node and global features, and passed through a neural network ϕn:
 | (16) |
 is the latent representation of the nodes. This produces latent node features that reflect both local geometry and broader contextual information from neighboring atoms. The architecture of ϕn follows the same design as ϕe.
is the latent representation of the nodes. This produces latent node features that reflect both local geometry and broader contextual information from neighboring atoms. The architecture of ϕn follows the same design as ϕe.
The global sub-model integrates information across the entire structure, combining aggregated node and edge embeddings with global descriptors  . Its update function is given by
. Its update function is given by
 | (17) |
 and ρe→g perform mean aggregations across all nodes and edges, respectively. The network ϕg adheres to the same architectural conventions as ϕe and ϕn sub-models.
and ρe→g perform mean aggregations across all nodes and edges, respectively. The network ϕg adheres to the same architectural conventions as ϕe and ϕn sub-models.
After the latent edge, node, and global representations were established, their outputs were concatenated and passed through a fully connected NN that produces final material-level embeddings (the s neurons in Fig. 2(a)). These embeddings were then connected to separate layers corresponding to each target property (the p neurons in Fig. 2(a)). Each output head contains twice the number of neurons as target properties—one neuron for the predicted value and another for the associated uncertainty—allowing the model to learn both quantities simultaneously.
Model training employed the negative log-likelihood (NLLH) loss function. We assumed Gaussian-distributed prediction errors. For each sample i and target property y, the model generated a mean prediction µi and an associated standard deviation σi. The loss, later averaged over all training instances and target properties, is expressed as
 | (18) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20
20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0 ratio for BO and a 80
0 ratio for BO and a 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10
10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 ratio for parity plots, respectively. Fixed training parameters included a learning rate of 0.001, a batch size of 32, an early stopping criterion of 10 epochs (i.e., the number of epochs with no improvement to terminate learning), and an exponential learning rate scheduler with γ of 0.99. Each of the 100 Optuna trials involved a short 10-epoch training run. Model performance was assessed using the area under the average NLLH loss function versus epoch curve, normalized by the number of epochs to mitigate the effects of early stopping. This approach provided a smoother and more reliable optimization metric than using the minimum or final epoch loss. We selected the optimal architectural parameters from these trials. The final model parameters were taken from the epoch with the lowest observed loss. For BO, this full optimization and training workflow was repeated at each iteration to refine the surrogate model continuously.
10 ratio for parity plots, respectively. Fixed training parameters included a learning rate of 0.001, a batch size of 32, an early stopping criterion of 10 epochs (i.e., the number of epochs with no improvement to terminate learning), and an exponential learning rate scheduler with γ of 0.99. Each of the 100 Optuna trials involved a short 10-epoch training run. Model performance was assessed using the area under the average NLLH loss function versus epoch curve, normalized by the number of epochs to mitigate the effects of early stopping. This approach provided a smoother and more reliable optimization metric than using the minimum or final epoch loss. We selected the optimal architectural parameters from these trials. The final model parameters were taken from the epoch with the lowest observed loss. For BO, this full optimization and training workflow was repeated at each iteration to refine the surrogate model continuously.
2.6 Gaussian process as a surrogate model
We compared our GNN-based surrogate with a commonly used surrogate model: a GP. GP is a non-parametric model that treats a set of points as jointly following a multivariate Gaussian distribution. By conditioning this distribution on observed data, the GP yields predictive means and standard deviations for new inputs. We implemented the SingleTaskGP from BoTorch65 with ExactMarginalLogLikelihood to train surrogate GP models, following the approach used in Deshwal et al.28 Our GP-based BO workflow was validated against their published results, with corresponding verification details provided in the SI. For comparisons with more than one objective, the KroneckerMultiTaskGP model type was used. Because GP models operate on fixed-length tabular inputs, they were trained solely on global-level features extracted from the graph representations of MOFs.2.7 Bayesian optimization
Given that both the GNN- and GP-based surrogates provide mean predictions and associated uncertainties, they can be directly applied within BO. Algorithm 1 outlines the general procedure. The process begins by fitting a surrogate model capable of estimating property values and uncertainties. Candidate materials are evaluated using an acquisition function that ranks them according to their expected improvement or contribution to the Pareto front. The top-ranked candidate is then selected, evaluated for its true properties, and appended to the training dataset. The surrogate is retrained on the expanded dataset, and the cycle repeats until a predetermined budget is reached. The budget may correspond to the number of BO iterations (as used in this study), total computation time, or other practical constraints.We applied BO to explore the space of MOFs with optimal Henry coefficients. Each surrogate was initially trained with 25 data points. For single-objective Bayesian optimization (SOBO), we used the expected improvement (EI) acquisition function. For multi-objective Bayesian optimization (MOBO), we used the expected hypervolume improvement (EHI). Both implementations were from BoTorch.65
The hypervolume improvement identifies the current Pareto front from measured objectives, then quantifies the additional hypervolume contributed by a new candidate relative to a reference point, measuring the improvement across all objectives at each iteration. A detailed formulation is provided by Xu et al.39 For SOBO, where only one objective is optimized, the improvement is computed as the difference between the best achieved value and the reference point, rather than a hypervolume. All reference points were chosen via the infer_reference_point function from BoTorch.65
We chose random selection of candidates from the available search space as a baseline for comparison. This approach ignores prior information and mimics unguided exploration, offering a conservative measure against which BO performance can be evaluated.
To quantitatively compare SOBO and MOBO results across different optimization runs, we introduced a normalized performance metric defined as follows. For each iteration, we computed the difference between GNN performance (e.g., hypervolume improvement) and random selection performance as a function of the number of proposed candidates, normalized by the performance gap between the best possible outcome and random selection. The area under the curve (AUC) of this normalized performance curve provides a single score representing how closely the GNN approaches optimal performance relative to random selection. Positive values indicate that the GNN outperforms random selection, negative values the opposite, and zero corresponds to equivalent performance. This metric was calculated for both hypervolume improvement and Pareto-front coverage to ensure consistency across optimization scenarios.
 | (19) |
3 Results & discussion
3.1 Comparison between surrogate model types
We start by comparing the performance of the GNN-based surrogate against the more classical GP surrogate for selectivity. Adsorption data from Li et al.36 were used as objectives. These three objectives are adsorbed CH4, adsorbed N2, and CH4 over N2 selectivity (SCH4/N2). These published data are from Grand Canonical Monte Carlo (GCMC) simulations for a 50% binary mixture of CH4 and N2 at 1 bar and 298 K (i.e., not Henry coefficients). Adsorptions are in units of mol per kg. Ten independent SOBO runs were conducted using GNN and BO surrogate models, along with random selection to provide a naive baseline selection process (Fig. 3(a)). Each run started from 25 randomly chosen initial samples (Algorithm 1) and proceeded for 850 iterations. Both GNN and GP surrogates significantly outperformed random selection for maximization of SCH4/N2. However, none of the GP models identified the highest selectivity MOF within the allocated budget. From approximately 550 sampled MOFs, the GP surrogate ceased improving, indicated by zero standard deviation in acquisition scores. This stagnation reflects local optima rather than repeated sampling of the same MOF. In contrast, all ten GNNs eventually identified the optimal MOF within the budget, typically evaluating about 1% (i.e., 850 suggested structures) of the 77![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 482 structures. GNNs models experienced a similar temporary stagnation between 200 and 350 iterations but recovered to find superior candidates.
482 structures. GNNs models experienced a similar temporary stagnation between 200 and 350 iterations but recovered to find superior candidates.
We compared the case for optimizing two objectives (Fig. 3(b)) similar to that of the single objective in Fig. 3(a). Specifically, surrogates were programmed to maximize adsorbed CH4 and minimize adsorbed N2. Three independent runs were averaged instead of ten used for the single objective. It is clear that GNNs outperform GPs and do so faster than the previous comparison. For a randomly chosen GNN surrogate, we show how the model explored both the input and objective space of our MOFs. We only include the first 200 samples to avoid visual clutter. The global level features of MOFs were gathered, transformed using scikit-learn's RobustScaler,63 and projected onto a two-dimensional (2D) space with the use of Uniform Manifold Approximation and Projection (UMAP)66 (Fig. 3(c)). The min_dist parameter of UMAP was set to 0.01. As seen in Fig. 3(c), there are sparse and dense regions in the low-dimensional 2D UMAP space. The surrogate samples points from both sparse and dense regions, indicating exploration. Fig. 3(d) shows the adsorption objectives on each axis and the same sampling of MOFs from Fig. 3(c). The GNN surrogate has sampled MOFs with objectives in the direction of higher CH4 adsorption and lower N2 adsorption (i.e., in the direction of the Pareto front). Identifying MOFs that maximize CH4 uptake while minimizing N2 adsorption directly translates to materials that enhance methane recovery and reduce energy costs in pressure-swing adsorption or membrane-based separation processes. Values of 0 on the color maps indicate MOFs that have not been sampled by the GNN. The superior performance of GNN over GP is attributed to its ability to capture complex structure–property relationships beyond the global features used by GPs. Consequently, GNNs surrogates were utilized for all subsequent optimization tasks in this work.
3.2 Correlations across Henry coefficients
This subsection focuses on challenging separations of VOCs using MOFs. As established in Section 2.2, maximizing separation involves increasing the Henry coefficient for the favored component while minimizing it for the excluded one. This task becomes difficult when Henry coefficients for both species are highly correlated (i.e., Pearson correlation factors ranging from 0.98 to effectively 1.0 for our VOCs). Although, no single MOF perfectly discriminates between VOCs, the Pareto front identifies the set of optimal trade-offs offering the best achievable selectivities (ratios of Henry coefficients, eqn (2)). We highlight one such Pareto-optimal MOF for later feature sensitivity analysis (Section 3.3).Strong correlations between Henry coefficients indicate that the adsorption physics are dominated by a scaling relationship where increased binding site density enhances both metrics simultaneously. Successful separation of alkanes from alcohols, for instance, requires minimizing the Henry coefficients of 1-propanol and 2-propanol while maximizing those for butane and propane in the four objective case, representing competing objectives that define a multi-dimensional Pareto front. The Pareto-optimal MOFs identified by our GNN model are those that maximize the ‘residual’ performance, achieving higher selectivity than would be predicted by the adsorption capacity alone. From an industrial perspective, even a small deviation from the dominant trend that would lead to a 2% increase in selectivity can result in significantly lower regeneration costs and higher product purity. Our aim is to develop a surrogate model for BO that can accurately identify the Pareto optimal set and distinguish it from suboptimal solutions.
3.3 Performance of surrogate model across Henry coefficients
This subsection validates the predictive accuracy of the GNN surrogate model on the multi-objective optimization Henry coefficient targets. Utilizing 800 randomly sampled MOFs for training, and 100 each for early stopping validation and testing, the model's generalization was assessed. Parity plots for 1-propanol and butane (Fig. 4) demonstrate strong predictive fidelity, with root mean squared error (RMSE) values of 1.355 and 1.491 ln([mol kg−1 Pa−1]) for 1-propanol and butane, respectively. Considering that the true Henry coefficient values for these VOCs span approximately 11 natural log units, an RMSE of around 1.5 ln([mol kg−1 Pa−1]) corresponds to an error of approximately 14% relative to the data span, acceptable for early-stage candidate screening. Similar accuracy for 2-propanol and propane (shown in the SI) confirms robustness.To evaluate the surrogate's ability to capture meaningful structure–property connections, we measured EHI score for all points. Among the top five scoring candidates, the three highest-scoring were not on the Pareto front, while the remaining two were Pareto-optimal. We selected the highest-scoring of the two Pareto front points (indicated by the star shape in Fig. 4) for the feature importance ablation study. This analysis revealed that node-level features predominantly drive model predictions, with eight of the top ten most important features arising from node descriptors and only two from global features for both 1-propanol and butane. The absence of edge features among the top contributors highlights the dominant role of local atomic environments captured at the node level. This underscores the limitation of surrogate models restricted to tabular global features, which would inherently miss significant predictive information embedded in the graph topology and atomic features. These results validate the surrogate's reliability and justify its use within the BO framework to effectively navigate strongly correlated multi-objective MOF design spaces.
3.4 Comparison on Bayesian optimization and the number of optimized objectives
The results presented in the previous sections demonstrated that the GNN surrogate can reliably predict multiple objectives associated with multiple selectivities in MOFs. Building on these results, we now evaluate the efficacy of our GNN as a surrogate model within BO for one- to four-objective optimization tasks, as summarized in Table 1.Fig. 5 illustrates results from a representative MOBO run optimizing butane selectivity over 1-propanol and 2-propanol. Panel (a) shows hypervolume improvement versus the number of proposed candidates, where the GNN-based surrogate significantly outperforms random selection, achieving near-maximal hypervolume after about 200 candidates, while random selection falls short even after 1000. The random proposals were averaged across 10 runs. Panel (b) shows the fraction of identified Pareto front points instead of hypervolume improvement, with the GNN identifying nearly 60% of points on the Pareto front after 1000 proposed candidates compared to 26% for random search.
Panels (c) and (d) compare the AUC for normalized hypervolume improvement and Pareto front coverage across runs with one to four objectives. In panel (c), the GNN's AUC of normalized hypervolume improvement, relative to random selection, remains largely consistent across runs optimizing one through four objectives. By contrast, panel (d) shows that the AUC of normalized fraction of Pareto-front points identified is highest for single-objective optimization, similar for two- and three-objective cases, and drops sharply in the four-objective scenario. While normalized hypervolume improvement remains stable across objectives, coverage declines for four-objective optimizations due to the exponential growth of the Pareto front dimensionality and the intrinsic difficulty of covering it extensively. For example, the single-objective front contains only one optimal point, the three-objective front (shown in panels (a) and (b)) has 211 points, and the four-objective front comprises 1440 points (approximately 40% of the dataset). Despite this, the GNN identified Pareto-front points at rates similar to random selection in higher dimensions, but the specific points it selected yielded substantially greater hypervolume improvement, maintaining superior optimization.
Finally, Table 2 reports GNN BO performance across various VOC separations within the same functional group family, highlighting generally positive AUC values except for the {benzene, toluene} dataset for the metric of fraction, where performance falls to random baseline, likely due to dataset size and the high similarity of these molecules. Overall, the GNN surrogate consistently enables effective optimization for challenging VOC separations in MOFs.
| Dataset | # Objectives | Metric | AUC |
|---|---|---|---|
| {1-Propanol} | 1 | Hypervolume | 942.5 |
| Fraction | 876.5 | ||
| {Butane} | 1 | Hypervolume | 776.4 |
| Fraction | 681.5 | ||
| {1-propanol, 2-propanol} | 2 | Hypervolume | 801.3 |
| Fraction | 140.0 | ||
| {1-propanol, butane} | 2 | Hypervolume | 772.0 |
| Fraction | 236.1 | ||
| {1-propanol, propane} | 2 | Hypervolume | 893.1 |
| Fraction | 64.8 | ||
| {2-cis-butene, 2-trans-butene} | 2 | Hypervolume | 824.6 |
| Fraction | 107.4 | ||
| {Benzene, toluene} | 2 | Hypervolume | 766.8 |
| Fraction | −7.1 | ||
| {Butane, propane} | 2 | Hypervolume | 852.5 |
| Fraction | 333.8 | ||
| {1-propanol, 2-propanol, butane} | 3 | Hypervolume | 867.8 |
| Fraction | 278.7 | ||
| {1-propanol, 2-propanol, propane} | 3 | Hypervolume | 771.5 |
| Fraction | 112.6 | ||
| {1-propanol, 2-propanol, butane, propane} | 4 | Hypervolume | 817.4 |
| Fraction | 8.4 |
3.5 Experimental feasibility of top-ranked MOFs
To bridge the gap between computational prediction and experimental realization, we evaluated the properties of six MOFs and demonstrated how they qualitatively occupy the space of available structural properties. We selected a subset of global graph features commonly used to interpret MOFs and constructed stacked histograms with 15 bins (Fig. 6(a) and (c)) to illustrate the distribution of these properties. Fig. 6(b) presents the top three MOFs on the Pareto front predicted by the GNN model used to generate Fig. 4. These MOFs exhibit large non-accessible surface areas, meaning that adsorbates are unlikely to find suitable uptake sites. Although the GNN surrogate successfully converges to the Pareto front in the four-objective optimization space, the proposed MOFs show traits that are inconsistent with physically reasonable expectations. Unusual behavior emerges in higher-dimensional spaces where the objectives are strongly correlated and have competing optimization directions. This issue did not arise in the results comparing adsorption values (i.e., not Henry coefficients) from Li et al.36 For the two-objective screening discussed in Section 3.1, the MOFs on the Pareto front selected by the GNN surrogate are shown in Fig. 6(d) and exhibit realistic physical characteristics. For instance, the optimal MOFs possess low non-accessible surface areas and large void spaces (i.e., higher largest free sphere, largest included sphere, and largest included sphere along free path). These trends align with known structure–adsorption relationships, suggesting that the GNN implicitly captures chemically and physically meaningful patterns when identifying optimal materials. The three MOFs in question are DB12-IHUTIQ_clean_repeat.cif (Fig. 6(e)), DB0-m3_o490_o15_f0_fsc.sym.2_repeat.cif (Fig. 6(f)), and DB1-ZnN4-SiF6-irmof6_No2_repeat.cif (Fig. 6(g)). We emphasize that our method screens and finds candidates on the Pareto front, but the Pareto front may lack physically reasonable characteristics which is especially true for higher dimension objective spaces. | ||
| Fig. 6 Selected MOF properties, their span, and values for Pareto optimal metal–organic framework (MOFs). Panel (a) shows a stacked histogram of properties for CoRE MOF shown in the legend. The horizontal axis is the base 10 logarithm of values (with values of zero kept as zero) while the vertical axis represents the counts. Panel (b) shows the property values for MOFs selected by a graph-neural network (GNN). Panels (c) and (d) show the same type of information as panels (a) and (b), respectively, but for the ARC-MOF instead. Panels (e)–(g) show the OVITO67 visualizations of MOFs from panel (d). | ||
Among the three optimal candidates in Fig. 6(d), DB12-IHUTIQ_clean_repeat.cif appears to be the most synthetically feasible. This material likely originates from the CCDC database and has been previously reported in the literature.68 Its organic linker, 1,4-bis(4-pyridyl)-2,3-diaza-1,3-butadiene, is commercially available, overcoming one of the most common limitations in MOF synthesis—linker accessibility. The second candidate, DB0-m3_o490_o15_f0_fsc.sym.2_repeat.cif, also appears synthetically feasible. It consists of a bridging tetratopic carboxylate linker and a pillared N-based linker, both of which are available commercially. Although this structure has not been previously reported, its composition suggests it could be readily synthesized. In contrast, DB1-ZnN4-SiF6-irmof6_No2_repeat.cif is the least feasible of the three. While conceptually synthesizable, the linker has not been previously synthetically realized. Nevertheless, this structure is isoreticular with the SIFSIX family of materials, implying that MOF synthesis would be straightforward if the linker were accessible.69
4 Conclusions
This work demonstrates how detailed structure–property relationships of MOFs can be effectively integrated into a BO workflow using GNNs. By representing MOFs as graphs with multi-scale features encoded at node, edge, and global levels, GNNs-based surrogates surpass traditional models such as GPs limited to global, tabular representations. Utilizing expected improvement and expected hypervolume improvement acquisition functions for single- and multi-objective optimization respectively, the GNN-enabled optimization efficiently navigates complex design spaces, achieving faster hypervolume gains, notably in challenging separations such as CH4 over N2 selectivity, when compared to GP-based approaches.Despite the inherent strong correlations among Henry coefficients of VOCs studied, GNNs identified Pareto-optimal MOFs with superior performance compared to random search. Ablation analyses underscored the critical role of node-level features in capturing local atomic environments, which are largely overlooked by models restricted to macroscopic descriptors. These results highlight the fundamental advantage of graph-based representations in encoding physicochemically relevant information essential for accurate surrogate modeling and effective candidate selection in materials optimization tasks.
Moving forward, integrating dynamic or flexible MOF representations that capture framework flexibility and adsorption site heterogeneity could improve realism and predictive accuracy. Incorporating multi-fidelity simulation data or experimental feedback into the BO loop may accelerate convergence and robustness of predictions. Enhancing uncertainty quantification methods within GNN architectures will better guide exploration–exploitation trade-offs. Lastly, extending this workflow to optimize additional properties such as adsorption kinetics or stability metrics would broaden its applicability to real-world materials design challenges.
Author contributions
R. D. and L. E. S. conceived the idea. L. E. S. featurized MOFs as graphs, trained and optimized GNNs, developed the BO routine, and wrote significant portions of this article. N. G. and N. S. B. performed simulations to measure Henry coefficients for VOCs, wrote the corresponding methods, and reviewed this article. D. F. S. G. and R. D. supervised the project and contributed to the draft of this article.Conflicts of interest
There are no conflicts to declare.Data availability
All raw data, processed data, and computational codes generated in this study are available from the corresponding author, Rémi Dingreville, upon reasonable request at E-mail: rdingre@sandia.gov; . The CoRE MOF 2019 database (Version 1.1.3) is publicly available at https://zenodo.org/record/3677685 and the ARC-MOF (V6) database is accessible at https://doi.org/10.5281/zenodo.13891643. Previously published data used to validate our methods are available at https://github.com/lwx199906/MOF-GRU.git.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ta09133k.
Acknowledgements
This work was supported by the Laboratory Directed Research and Development Program at Sandia National Laboratories. Computational capabilities were supported by the Center for Integrated Nanotechnologies (CINT), an Office of Science User Facility operated for the U.S. Department of Energy (DOE), Office of Science. This article has been authored by an employee of National Technology & Engineering Solutions of Sandia, LLC under Contract No. DE-NA0003525 with the U.S. Department of Energy (DOE). The employee owns all right, title and interest in and to the article and is solely responsible for its contents. The United States Government retains and the publisher, by accepting the article for publication, acknowledges that the United States Government retains a non-exclusive, paid-up, irrevocable, world-wide license to publish or reproduce the published form of this article or allow others to do so, for United States Government purposes. The DOE will provide public access to these results of federally sponsored research in accordance with the DOE Public Access Plan https://www.energy.gov/downloads/doe-public-access-plan.References
- V. F. Yusuf, N. I. Malek and S. K. Kailasa, ACS Omega, 2022, 7, 44507–44531 Search PubMed.
- H. Furukawa, K. E. Cordova, M. O'Keeffe and O. M. Yaghi, Science, 2013, 341, 1230444 Search PubMed.
- H.-C. J. Zhou and S. Kitagawa, Chem. Soc. Rev., 2014, 43, 5415–5418 Search PubMed.
- G. Li, H. Kobayashi, J. M. Taylor, R. Ikeda, Y. Kubota, K. Kato, M. Takata, T. Yamamoto, S. Toh, S. Matsumura and H. Kitagawa, Nat. Mater., 2014, 13, 802–806 Search PubMed.
- J.-R. Li, R. J. Kuppler and H.-C. Zhou, Chem. Soc. Rev., 2009, 38, 1477–1504 Search PubMed.
- S. K. Firooz and D. W. Armstrong, Anal. Chim. Acta, 2022, 1234, 340208 Search PubMed.
- L. E. Kreno, K. Leong, O. K. Farha, M. Allendorf, R. P. Van Duyne and J. T. Hupp, Chem. Rev., 2012, 112, 1105–1125 CrossRef CAS PubMed.
- S. Okur, T. Hashem, E. Bogdanova, P. Hodapp, L. Heinke, S. Bräse and C. Wöll, ACS Sens., 2024, 9, 622–630 CrossRef CAS PubMed.
- J. Rocha, L. D. Carlos, F. A. A. Paz and D. Ananias, Chem. Soc. Rev., 2011, 40, 926–940 RSC.
- J. I. Deneff, L. E. S. Rohwer, K. S. Butler, B. Kaehr, D. J. Vogel, T. S. Luk, R. A. Reyes, A. A. Cruz-Cabrera, J. E. Martin and D. F. Sava Gallis, Nat. Commun., 2023, 14, 981 Search PubMed.
- Y. Hu, Z. Huang, L. Zhou, D. Wang and G. Li, J. Sep. Sci., 2014, 37, 1482–1488 CrossRef CAS PubMed.
- J. Lee, O. K. Farha, J. Roberts, K. A. Scheidt, S. T. Nguyen and J. T. Hupp, Chem. Soc. Rev., 2009, 38, 1450–1459 RSC.
- R. E. Sikma, D. J. Vogel, R. A. Reyes, M. L. Meyerson, P. G. Kotula and D. F. Sava Gallis, Adv. Mater., 2024, 36, 2407435 CrossRef CAS PubMed.
- J. Della Rocca, D. Liu and W. Lin, Acc. Chem. Res., 2011, 44, 957–968 Search PubMed.
- F. Ke, Y.-P. Yuan, L.-G. Qiu, Y.-H. Shen, A.-J. Xie, J.-F. Zhu, X.-Y. Tian and L.-D. Zhang, J. Mater. Chem., 2011, 21, 3843–3848 Search PubMed.
- D. F. Sava Gallis, K. S. Butler, J. O. Agola, C. J. Pearce and A. A. McBride, ACS Appl. Mater. Interfaces, 2019, 11, 7782–7791 Search PubMed.
- P. Falcaro, R. Ricco, A. Yazdi, I. Imaz, S. Furukawa, D. Maspoch, R. Ameloot, J. D. Evans and C. J. Doonan, Coord. Chem. Rev., 2016, 307, 237–254 CrossRef CAS.
- C. E. Wilmer, M. Leaf, C. Y. Lee, O. K. Farha, B. G. Hauser, J. T. Hupp and R. Q. Snurr, Nat. Chem., 2012, 4, 83–89 Search PubMed.
- Y. J. Colón, D. A. Gómez-Gualdrón and R. Q. Snurr, Cryst. Growth Des., 2017, 17, 5801–5810 Search PubMed.
- J. Burner, J. Luo, A. White, A. Mirmiran, O. Kwon, P. G. Boyd, S. Maley, M. Gibaldi, S. Simrod, V. Ogden and T. K. Woo, Chem. Mater., 2023, 35, 900–916 Search PubMed.
- J. Burner, J. Luo, A. White, A. Mirmiran, O. Kwon, P. G. Boyd, S. Maley, M. Gibaldi, S. Simrod and T. K. Woo, ab initio REPEAT Charge MOF Database (ARC-MOF), 2025, DOI:10.5281/zenodo.13891643.
- Y. G. Chung, E. Haldoupis, B. J. Bucior, M. Haranczyk, S. Lee, H. Zhang, K. D. Vogiatzis, M. Milisavljevic, S. Ling, J. S. Camp, B. Slater, J. I. Siepmann, D. S. Sholl and R. Q. Snurr, J. Chem. Eng. Data, 2019, 64, 5985–5998 Search PubMed.
- A. White, M. Gibaldi, J. Burner, R. A. Mayo and T. Woo, Alarming structural error rates in MOF databases used in data driven workflows identified via a novel metal oxidation state-based method, https://chemrxiv.org/engage/chemrxiv/article-details/6706b96312ff75c3a1fb0365 Search PubMed.
- Z. Yao, B. Sánchez-Lengeling, N. S. Bobbitt, B. J. Bucior, S. G. H. Kumar, S. P. Collins, T. Burns, T. K. Woo, O. K. Farha, R. Q. Snurr and A. Aspuru-Guzik, Nat. Mach. Intell., 2021, 3, 76–86 Search PubMed.
- Z. Han, Y. Yang, J. Rushlow, J. Huo, Z. Liu, Y.-C. Hsu, R. Yin, M. Wang, R. Liang, K.-Y. Wang and H.-C. Zhou, Chem. Soc. Rev., 2025, 54, 367–395 Search PubMed.
- R. Wang, Y. Zhong, L. Bi, M. Yang and D. Xu, ACS Appl. Mater. Interfaces, 2020, 12, 52797–52807 CrossRef CAS PubMed.
- T. D. Pham and R. Q. Snurr, Langmuir, 2025, 41, 4585–4593 Search PubMed.
- A. Deshwal, C. M. Simon and J. R. Doppa, Mol. Syst. Des. Eng., 2021, 6, 1066–1086 RSC.
- N. Gantzler, A. Deshwal, J. R. Doppa and C. M. Simon, Digital Discovery, 2023, 2, 1937–1956 RSC.
- T.-W. Liu, Q. Nguyen, A. B. Dieng and D. A. Gómez-Gualdrón, Chem. Sci., 2024, 15, 18903–18919 Search PubMed.
- Y. Comlek, T. D. Pham, R. Q. Snurr and W. Chen, npj Comput. Mater., 2023, 9, 1–14 Search PubMed.
- Y. G. Chung, D. A. Gómez-Gualdrón, P. Li, K. T. Leperi, P. Deria, H. Zhang, N. A. Vermeulen, J. F. Stoddart, F. You, J. T. Hupp, O. K. Farha and R. Q. Snurr, Sci. Adv., 2016, 2, e1600909 CrossRef PubMed.
- A. W. Thornton, C. M. Simon, J. Kim, O. Kwon, K. S. Deeg, K. Konstas, S. J. Pas, M. R. Hill, D. A. Winkler, M. Haranczyk and B. Smit, Chem. Mater., 2017, 29, 2844–2854 CrossRef CAS PubMed.
- M. Pardakhti, E. Moharreri, D. Wanik, S. L. Suib and R. Srivastava, ACS Comb. Sci., 2017, 19, 640–645 Search PubMed.
- J. Luo, O. B. Said, P. Xie, M. Gibaldi, J. Burner, C. Pereira and T. K. Woo, npj Comput. Mater., 2024, 10, 224 Search PubMed.
- W. Li, Y. Situ, L. Ding, Y. Chen and Q. Yang, ACS Appl. Mater. Interfaces, 2023, 15, 59887–59894 CrossRef CAS PubMed.
- Y. G. Chung, E. Haldoupis, B. J. Bucior, M. Haranczyk, S. Lee, K. D. Vogiatzis, S. Ling, M. Milisavljevic, H. Zhang, J. S. Camp, B. Slater, J. Ilja Siepmann, D. S. Sholl and R. Q. Snurr, Computation-Ready Experimental Metal–Organic Framework (CoRE MOF) 2019 Dataset, https://zenodo.org/record/3677685 Search PubMed.
- K. S. Walton and D. S. Sholl, AIChE J., 2015, 61, 2757–2762 CrossRef CAS.
- S. Xu, J. Zhang, H. Chen, Y. Gao, Y. Gao, H. Gao and X. Jia, Appl. Sci., 2022, 13, 352 Search PubMed.
- D. Dubbeldam, S. Calero, D. E. Ellis and R. Q. Snurr, Mol. Simul., 2016, 42, 81–101 CrossRef CAS.
- N. S. Bobbitt, R. E. Sikma, J. P. Sammon, M. Chandross, J. I. Deneff and D. F. Sava Gallis, ACS Sens., 2025, 10, 360–375 Search PubMed.
- C. J. Casewit, K. S. Colwell and A. K. Rappe, J. Am. Chem. Soc., 1992, 114, 10046–10053 Search PubMed.
- B. Chen, J. J. Potoff and J. I. Siepmann, J. Phys. Chem. B, 2001, 105, 3093–3104 CrossRef CAS.
- N. Rai and J. I. Siepmann, J. Phys. Chem. B, 2007, 111, 10790–10799 CrossRef CAS PubMed.
- M. G. Martin and J. I. Siepmann, J. Phys. Chem. B, 1998, 102, 2569–2577 Search PubMed.
- C. D. Wick, M. G. Martin and J. I. Siepmann, J. Phys. Chem. B, 2000, 104, 8008–8016 CrossRef CAS.
- C. E. Wilmer, K. C. Kim and R. Q. Snurr, J. Phys. Chem. Lett., 2012, 3, 2506–2511 Search PubMed.
- N. S. Bobbitt, R. E. Sikma, J. P. Sammon, M. Chandross, J. I. Deneff and D. F. Sava Gallis, ACS Sens., 2025, 10, 360–375 Search PubMed.
- J. Gonzalez, K. Mukherjee and Y. J. Colón, J. Chem. Eng. Data, 2023, 68, 291–302 CrossRef CAS.
- D. Banerjee, C. M. Simon, A. M. Plonka, R. K. Motkuri, J. Liu, X. Chen, B. Smit, J. B. Parise, M. Haranczyk and P. K. Thallapally, Nat. Commun., 2016, 7, ncomms11831 CrossRef CAS PubMed.
- S. P. Ong, W. D. Richards, A. Jain, G. Hautier, M. Kocher, S. Cholia, D. Gunter, V. L. Chevrier, K. A. Persson and G. Ceder, Comput. Mater. Sci., 2013, 68, 314–319 CrossRef CAS.
- O. Isayev, C. Oses, C. Toher, E. Gossett, S. Curtarolo and A. Tropsha, Nat. Commun., 2017, 8, 15679 CrossRef CAS PubMed.
- A. P. Bartók, R. Kondor and G. Csányi, Phys. Rev. B, 2013, 87, 184115 CrossRef.
- Ł. Mentel, mendeleev – A Python resource for properties of chemical elements, ions and isotopes, 2014, https://github.com/lmmentel/mendeleev Search PubMed.
- M. Fernandez, N. R. Trefiak and T. K. Woo, J. Phys. Chem. C, 2013, 117, 14095–14105 Search PubMed.
- J. Behler, J. Chem. Phys., 2011, 134, 074106 CrossRef PubMed.
- M. Gastegger, L. Schwiedrzik, M. Bittermann, F. Berzsenyi and P. Marquetand, J. Chem. Phys., 2018, 148, 241709 CrossRef CAS PubMed.
- S. Kancharlapalli, A. Gopalan, M. Haranczyk and R. Q. Snurr, J. Chem. Theory Comput., 2021, 17, 3052–3064 Search PubMed.
- T. F. Willems, C. H. Rycroft, M. Kazi, J. C. Meza and M. Haranczyk, Microporous Mesoporous Mater., 2012, 149, 134–141 Search PubMed.
- A. A. Hagberg, D. A. Schult and P. J. Swart, Proceedings of the 7th Python in Science Conference, Pasadena, CA USA, 2008, pp. , pp. 11–15 Search PubMed.
- M. Fey and J. E. Lenssen, ICLR Workshop on Representation Learning on Graphs and Manifolds, 2019 Search PubMed.
- P. W. Battaglia, J. B. Hamrick, V. Bapst, A. Sanchez-Gonzalez, V. Zambaldi, M. Malinowski, A. Tacchetti, D. Raposo, A. Santoro, R. Faulkner, C. Gulcehre, F. Song, A. Ballard, J. Gilmer, G. Dahl, A. Vaswani, K. Allen, C. Nash, V. Langston, C. Dyer, N. Heess, D. Wierstra, P. Kohli, M. Botvinick, O. Vinyals, Y. Li and R. Pascanu, Relational inductive biases, deep learning, and graph networks, arXiv, 2018, preprint, arXiv:1806.01261, DOI:10.48550/arXiv.1806.01261, http://arxiv.org/abs/1806.01261.
- F. Pedregosa, G. Varoquaux, A. Gramfort, V. Michel, B. Thirion, O. Grisel, M. Blondel, P. Prettenhofer, R. Weiss, V. Dubourg, J. Vanderplas, A. Passos, D. Cournapeau, M. Brucher, M. Perrot and E. Duchesnay, J. Mach. Learn. Res., 2011, 12, 2825–2830 Search PubMed.
- T. Akiba, S. Sano, T. Yanase, T. Ohta and M. Koyama, Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 2019 Search PubMed.
- M. Balandat, B. Karrer, D. R. Jiang, S. Daulton, B. Letham, A. G. Wilson and E. Bakshy, Adv. Neural Inf. Process. Syst., 2020, 33, 21524–21538 Search PubMed.
- L. McInnes, J. Healy, N. Saul and L. Grossberger, J. Open Source Softw., 2018, 3, 861 CrossRef.
- A. Stukowski, Model. Simulat. Mater. Sci. Eng., 2009, 18, 015012 CrossRef.
- J. Ye, X. Li, Z. liang Xu and H. Xu, J. Mol. Struct., 2015, 1093, 162–165 CrossRef CAS.
- O. Shekhah, Y. Belmabkhout, Z. Chen, V. Guillerm, A. Cairns, K. Adil and M. Eddaoudi, Nat. Commun., 2014, 5, 4228 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |





