Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
ZIF-8-based biocomposites via reactive extrusion: towards industrial-scale manufacturing
Nikita
Gugin
 ab,
Alexander
Schwab
ab,
Francesco
Carraro
ab,
Alexander
Schwab
ab,
Francesco
Carraro
 c,
Isabella
Tavernaro
c,
Isabella
Tavernaro
 a,
Jana
Falkenhagen
a,
Jana
Falkenhagen
 a,
José Antonio
Villajos
a,
José Antonio
Villajos
 ad,
Paolo
Falcaro
ad,
Paolo
Falcaro
 *c and
Franziska
Emmerling
*c and
Franziska
Emmerling
 *ab
*ab
aFederal Institute for Materials Research and Testing (BAM), Richard-Willstätter-Str. 11, D-12489 Berlin, Germany. E-mail: franziska.emmerling@bam.de
bDepartment of Chemistry, Humboldt-Universität zu Berlin, Brook-Taylor-Str. 2, D-12489 Berlin, Germany
cInstitute of Physical and Theoretical Chemistry, Graz University of Technology, Stremayrgasse 9, 8010 Graz, Austria. E-mail: paolo.falcaro@tugraz.at
dDepartment Hydrogen and Power-to-X, Iberic Centre of Research in Energy Storage, Av. Universidad S/N, 10003 Cáceres, Spain
First published on 20th January 2026
Abstract
Mechanochemistry, a sustainable synthetic method that minimizes solvent use, has shown great promise in producing metal–organic framework (MOF)-based biocomposites through ball milling. While ball milling offers fast reaction times, biocompatible conditions, and access to previously unattainable biocomposites, it is a batch-type process typically limited to gram-scale production, which is insufficient to meet commercial capacity. We introduce a scalable approach for the continuous solid-state production of MOF-based biocomposites. Our study commences with model batch reactions to examine the encapsulation of various biomolecules into Zeolitic Imidazolate Framework-8 (ZIF-8) via hand mixing, establishing a foundation for upscaling. Subsequently, the process is scaled up using reactive extrusion, enabling continuous and reproducible kilogram-scale production of bovine serum albumin (BSA)@ZIF-8 with tunable protein loading. Furthermore, we achieve the one-step formation of shaped ZIF-8 extrudates encapsulating clinical therapeutic hyaluronic acid (HA). Upon release of HA from the composite, the molecular weight of HA is preserved, highlighting the industrial potential of reactive extrusion for the cost-effective and reliable manufacturing of biocomposites for drug-delivery applications.
Introduction
The rapid advancement of science in the 20th century has brought biotechnology into key sectors such as pharmaceuticals, agriculture, and food production. Recent developments at the intersection of molecular and biomolecular sciences have led to the creation of advanced technologies, such as genetic vaccines and biosensing devices.4–6 However, the broader commercial use and cost-effectiveness of these new technologies are restricted by the inherent instability of biomolecules outside their native environment.7,8 The need for ‘cold chain’ logistics to store and deliver vaccines9 and the limited reusability of enzymes in biocatalysis10 contribute to these challenges. A promising method for stabilizing bioactive macromolecules involves integrating them into the pores or immobilizing them onto the surfaces of metal–organic frameworks (MOFs).11 The resulting MOF-biocomposites reduce the lability of the guest molecules, such as protein unfolding,12 thereby increasing their stability against physical and chemical stressors, including heat, organic solvents, or proteolytic agents.13 Apart from enhancing thermodynamic stability, a MOF shell provides additional valuable properties when encapsulating enzymes and biotherapeutics, including size-selectivity for catalysis,14 controllable release profile for drug delivery,15 and structural tunability for targeted medicine.16,17 Therefore, the encapsulation of biomolecules in MOFs is promising for biotechnology and biomedicine.Since the conventional solvothermal synthesis of MOFs often requires harsh solvothermal conditions that are detrimental to biomolecules, their immobilization can be achieved through post-synthetic infiltration or surface bioconjugation.13 Alternatively, a single-step encapsulation (e.g., biomimetic mineralization,18 co-precipitation,19 or de novo approach)20 can be utilized: in the presence of some MOF building blocks, MOF particles grow on the biomolecule's surface under biocompatible conditions (Fig. 1a and b). This approach is suitable for the encapsulation of bio-entities with larger dimensions than the MOF pores openings. Successful encapsulation via biomimetic mineralization includes enzymes,21 antibodies,22 viruses,23 and living cells.24 However, we note that spontaneous encapsulation is typically limited to soluble bioentities with a negative surface charge.25–27 Additionally, despite the successful application of flow chemistry for the continuous synthesis of protein@MOF materials,28 the production rate is currently limited to a maximum of a few grams per hour,28 insufficient to fully realize the commercial potential of MOF-biocomposites.
 | ||
| Fig. 1 (a) Encapsulation method for preparing MOF biocomposites. (b) Comparison of encapsulation methods: biomimetic mineralization versus mechanochemistry, focusing on the challenges of upscaling mechanochemistry. (c) The work performed in this study. 1st step: screening of reaction conditions in the solid phase using a simple and fast hand-mixing approach; 2nd step: using the obtained data to scale MOF biocomposites through reactive extrusion. The crystal structures of BSA and GOx are reproduced under the terms of the CC-BY license.2,3 | ||
A promising development in the field of MOF chemistry is the utilisation of mechanical force to perform synthesis with little or no solvent.29 Intensive research on reactions in ball mills has resulted in a large library of mechanochemically available MOFs.30–32 Optimization of synthesis conditions and mechanistic studies, including time-resolved in situ (TRIS) monitoring methods,33–35 improved reaction control and reproducibility – prerequisites for large-scale production.36–38 The latter has been implemented as a continuous twin-screw extrusion approach, with production rates up to 4 kg h−1, and space-time yields (STY) up to 140![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 kg m−3 d−1.39–41
000 kg m−3 d−1.39–41
Given the potential of mechanochemistry for industrial production of materials, it was a natural step to use the mechanochemical approach to synthesize functional MOF-biocomposites.42 The strategy for applying mechanochemistry can vary: inducing amorphization of MOF-biocomposites through ball milling to tailor drug-release kinetics43,44 or incorporating biomolecules into pre-assembled MOFs by grinding.45–48 Alternatively, mechanochemistry can provide biocomposites by assembling MOFs and encapsulating biomaterials in one step, akin to the biomimetic mineralization approach (Fig. 1b). The first reported example is the synthesis of lipase@ZIF-8, achieved by grinding zinc oxide, 2-methylimidazole, and lipase produced from Burkholderia ubonensis with trace ethanol in an agate mortar.49 The general applicability of the mechanochemical method was then demonstrated by Wei et al., who used ball-milling to encapsulate enzymes β-glucosidase, invertase, β-galactosidase, and catalase into ZIF-8, UiO-66-NH2, and Zn-MOF-74.50 Subsequent works have expanded the list of mechanochemically produced biocomposites to include Candida rugosa lipase encapsulated in ZIF-8 and Zn-MOF-74,51 ibuprofen@HKUST-1,52 5-fluorouracil@MIL-100(Fe),53 and ZIF-90 loaded with BSA, catalase, or E. coli bacteria.54 Additionally, photosensitizers (Ru or Cu coordination complexes) and wheat germ lipase have been co-immobilized in UiO-67 for photoenzymatic enantioselective catalysis.55
The mechanochemical approach to biocomposites offers several advantages. Firstly, it does not require bulk solvents, which is beneficial for those biomolecules with enhanced stability in their dry or lyophilized form.56 Secondly, it enables the synthesis of MOF biocomposites that could not be prepared via one-pot syntheses because of the harsh conditions required to grow some MOFs (e.g., UiO-66-NH2)57. Thirdly, it provides quick reaction times, as fast as only 10 seconds for catalase@ZIF-90, having been reported by the Shieh group.54 Finally, in some cases, the biological activity of the biocomposites produced by this method can outperform that of those produced by water-based synthesis.54
However, current research in this field is limited to pioneering syntheses with ball mills, typically at the 100 mg scale.58 Ball milling is a batch process with limited temperature control, which becomes increasingly problematic at larger production scales.38 This makes scaling up MOF biocomposites in ball mills impractical, particularly for thermally sensitive biomolecules.
As a continuous alternative, reactive extrusion offers a promising route for scalable MOF biocomposite synthesis. Although large-scale production of ZIF-8 and ZIF-8/polymer composites via continuous reactive extrusion has already been demonstrated,39,59 these processes rely on hot melt extrusion at 150–200 °C – temperatures incompatible with sensitive biomolecules. In our previous work, we showed that ZIF-8 can be synthesized by reactive extrusion even under mild conditions of 40 °C,40 thereby providing a green and scalable platform for MOF production. The present study advances this methodology to the encapsulation of biomacromolecules, introducing challenges related to preserving stability and function, investigating loading capacity (protein wt%) and encapsulation efficiency (the percentage of the initial biomolecule amount successfully immobilized), determining biomolecule location, ensuring reproducibility during continuous processing, and enabling shaping into defined, processable forms.
Here, we present a sustainable and scalable method for encapsulating biomacromolecules within ZIF-8, the most commonly used MOF in biocomposite synthesis.27 In contrast to traditional solvent-based approaches, we report the solid-state formation of ZIF-8 biocomposites via simple hand-mixing batch reactions (Fig. 1c, 1st step). We demonstrate this approach using four biomacromolecules: two model compounds, bovine serum albumin (BSA) protein and carboxymethyl-dextran (CM-dextran) carbohydrate, and two commercially valuable compounds, glucose oxidase (GOx) enzyme and hyaluronic acid (HA) carbohydrate. The mild synthesis conditions preserve GOx activity, confirming the method's suitability for sensitive biomaterials. We further extend the method to nucleic acids (salmon DNA) and lipids (eicosapentaenoic acid (EPA) ethyl ester). Using BSA@ZIF-8, we demonstrate the transition from small-scale batch reactions to larger-scale reactive extrusion, ensuring consistent product quality during continuous operation (Fig. 1c, 2nd step). To demonstrate the applicability of reactive extrusion for MOF-based drug production, we encapsulated a model therapeutic, CM-dextran, and a clinical therapeutic, HA, within ZIF-8. Finally, we fabricated shaped HA@ZIF-8 extrudates to improve processability and analyzed the molar mass distribution of released HA, confirming that its structural integrity is retained during extrusion-based encapsulation.
Experimental section
Batch hand-mixing reactions
In a typical experiment, as-purchased 2-methylimidazole (130.3 mg, 1.59 mmol) and basic zinc carbonate (69.7 mg, 0.127 mmol) powders were placed together with the desired amount of biomolecule powder in a 2 ml Eppendorf tube. After mixing the powders for 10 minutes, ethanol and deionized water (ethanol/water = 25/75 v/v, η = 0.30 µl mg−1) were added as auxiliary liquids using a micropipette and thoroughly mixed with a spatula for 5 minutes. The resulting material was resuspended in 1.5 ml of ethanol or water for washing, followed by centrifugation for 3 minutes. This washing procedure was repeated three times, discarding the supernatant each time. The recovered material was then dried overnight in a vacuum oven at room temperature. For a more detailed description of the procedure, see SI.Reactive extrusion
BSA@ZIF-8: in a typical experiment, basic zinc carbonate and 2-methylimidazole powders were thoroughly mixed in a molar ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 12.5 along with the desired amount of pre-ground BSA powder in a 500 ml beaker for 5 minutes. The mixture was fed into the first segment of the pre-heated (40 °C) extruder barrel using the automatic volumetric feeder at a speed of 5% (approximately 1.5 g min−1) and processed at a screw speed of 40 rpm. Simultaneously, absolute ethanol was supplied into the second segment of the extruder barrel using a peristaltic pump at a flow rate of 0.25–0.4 ml min−1 (corresponding to η = 0.17–0.27 µl per mg of solid mixture). After about 20 minutes from the start of extrusion, the extrudate was collected. The as-synthesized material was resuspended in absolute ethanol (10 ml of ethanol per 1 g of extrudate), vortexed, and centrifuged. This washing procedure was repeated three times. Subsequently, the extrudate was washed with an aqueous 10 wt% sodium dodecyl sulfate (SDS) solution (10 ml of SDS solution per 1 g of extrudate), vortexed, allowed to rest for 30 minutes, and then centrifuged. This was followed by three more ethanol-washing steps before drying the sample overnight at room temperature under vacuum. For a more detailed description of the procedure, see SI.
12.5 along with the desired amount of pre-ground BSA powder in a 500 ml beaker for 5 minutes. The mixture was fed into the first segment of the pre-heated (40 °C) extruder barrel using the automatic volumetric feeder at a speed of 5% (approximately 1.5 g min−1) and processed at a screw speed of 40 rpm. Simultaneously, absolute ethanol was supplied into the second segment of the extruder barrel using a peristaltic pump at a flow rate of 0.25–0.4 ml min−1 (corresponding to η = 0.17–0.27 µl per mg of solid mixture). After about 20 minutes from the start of extrusion, the extrudate was collected. The as-synthesized material was resuspended in absolute ethanol (10 ml of ethanol per 1 g of extrudate), vortexed, and centrifuged. This washing procedure was repeated three times. Subsequently, the extrudate was washed with an aqueous 10 wt% sodium dodecyl sulfate (SDS) solution (10 ml of SDS solution per 1 g of extrudate), vortexed, allowed to rest for 30 minutes, and then centrifuged. This was followed by three more ethanol-washing steps before drying the sample overnight at room temperature under vacuum. For a more detailed description of the procedure, see SI.
CM-dextran@ZIF-8 was synthesized following a procedure similar to BSA@ZIF-8. A reagent mixture of basic zinc carbonate (27.9 g, 0.051 mol), 2-methylimidazole (52.2 g, 0.636 mol), and CM-dextran (6.43 g) was processed using a water/ethanol mixture (75/25 v/v) as the auxiliary liquid, supplied at a flow rate of 0.25 ml min−1. The as-synthesized extrudate was washed three times with ethanol, once with DMSO, followed by three additional ethanol washing steps, and then dried overnight at room temperature under vacuum. For a more detailed procedure, see the SI.
HA(10%)@ZIF-8 was synthesized following a procedure similar to BSA@ZIF-8. A reagent mixture of basic zinc carbonate (42.8 g, 0.078 mol), 2-methylimidazole (80.0 g, 0.974 mol), and hyaluronic acid sodium salt (9.9 g) was processed using a water/ethanol mixture (75/25 v/v) as the auxiliary liquid, supplied at a flow rate of 0.35 ml min−1. The dense paste was collected over 60 minutes, yielding 58 g of raw extrudate. The extrudate was washed three times with ethanol, once with an aqueous 10 wt% sodium dodecyl sulfate (SDS) solution, followed by three additional ethanol washing steps, and then dried overnight at room temperature under vacuum.
A total of 45 g of the shaped HA(10%)@ZIF-8 extrudate with a nominal diameter of 2 mm was obtained over a 50-minute experiment using a die faceplate. The as-synthesized material retained its shape after three ethanol washes and was subsequently dried overnight under vacuum, reducing to 30 g. In contrast, washing with water led to extrudate disaggregation, likely due to the dissolution of HA, which could function as an internal binder.
Results and discussion
Batch mixing reactions
Reactive extrusion is a large-scale technique that requires significant material input, making it essential to develop a simple model for testing reaction conditions on a smaller scale. Although ball milling has been reported as a route to ZIF-8 synthesis,60–62 and could in principle serve as a preliminary tool for exploring extrusion-related parameters,63,64 the two approaches rely on different mechanical activation modes,33 operate over distinct reaction times, and involve different processing conditions, which limits the direct transferability of results between them.65 Nevertheless, batch mixing can aid in optimizing reactive extrusion.66 This led us to adapt a previously reported simple hand-mixing approach for ZIF-8 (ref. 67) to examine the preparation of ZIF-8-based biocomposites.Screening of reaction conditions
The manual batch mixing approach to ZIF-8 involves mixing powders of 2-methylimidazole (HmeIm, C4H6N2) and basic zinc carbonate (ZnCarb, [ZnCO3]2·[Zn(OH)2]3) for five minutes in the presence of small amounts of ethanol.67 To ensure full conversion of the reagents to ZIF-8, we screened reaction conditions at the 200 mg scale by testing ethanol, water, and an ethanol/water (50/50 v/v) mixture as auxiliary liquids across a range of η values (0.075–0.3 µl mg−1), where η is the volume of solvent divided by the sample weight (see Batch syntheses of biocomposites and Table S1 in the SI).68 Powder X-ray diffraction (PXRD) analysis (Fig. S1) and attenuated total reflectance Fourier transform infrared (ATR-FTIR) spectroscopy (Fig. S2) of the ethanol-washed products confirm the consistent formation of highly crystalline and pure ZIF-8 in all experiments, with only minor leftovers of unreacted ZnCarb detected when using a low η value of 0.075 µl mg−1.After confirming the efficiency of the batch mixing approach for synthesizing pure ZIF-8, we explored the preparation of BSA@ZIF-8 biocomposite. We modified the aforementioned synthesis of pure ZIF-8 by adding BSA directly to the reagent mixture, either as a powder or an aqueous solution. We screened ethanol, water, and ethanol/water mixtures (25/75, 50/50, 75/25 v/v) as auxiliary liquids in a range of η values (Table S2). When using the water-washing procedure following the synthesis, ZIF-8 with sodalite (sod) topology was formed, along with a minor secondary phase, Zn(meIm)2·ZnCO3 or ZIF-C (Fig. S3). According to the ZIF Phase Analysis app,69 this secondary phase constitutes 4% and may result from unreacted zinc carbonate species, amorphous zinc carbonate/ZIF-8 clusters that serve as a source of carbonate for ZIF-C, or from the exposure of the initially formed ZIF-8 to carbon dioxide and moisture, as previously reported.62,70 The ethanol wash produced the pure sod ZIF-8 phase (Fig. S4); the ZIF-C to ZIF-8 conversion has been reported when ethanol is used.15 ATR-FTIR spectroscopy of the water- and ethanol-washed samples revealed the characteristic BSA amide I band at 1600–1700 cm−1 along with the characteristic spectrum of ZIF-8 (Fig. S5 and S6), confirming the formation of BSA@ZIF-8 biocomposites. Thus, the batch screening demonstrates the rapid (5 min) and consistent formation of ZIF-8 biocomposites with the model protein BSA under various reaction conditions.
Biocomposites of diverse biomacromolecules
Based on our screening results, we applied the optimal reaction conditions to synthesize model ZIF-8 biocomposites with BSA and carboxymethyl-dextran (CM-dextran), as well as functional composites with hyaluronic acid (HA) and glucose oxidase (GOx) (Fig. 2a). HA is a glycosaminoglycan (GAG)-based clinical therapeutic, while GOx, an oxidoreductase enzyme, is widely used in large-scale technological applications.71,72 All these biomolecules have previously been shown to form biocomposites with ZIF-8 via a biomimetic mineralization approach.15,21,26,73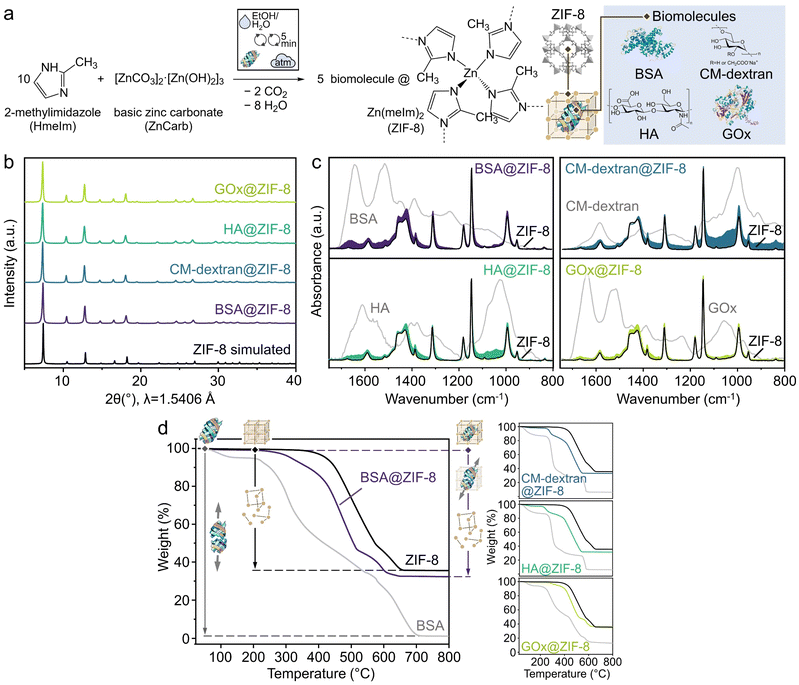 | ||
| Fig. 2 (a) Schematic representation1 of the hand-mixing batch synthesis of ZIF-8 biocomposites with BSA, CM-dextran, HA, or GOx biomolecules, using a water/ethanol mixture as the auxiliary liquid. (b) PXRD patterns of the synthesized materials compared with the reference diffraction pattern of ZIF-8. (c) ATR-FTIR spectra of the resulting materials, highlighting the presence of characteristic biomolecule bands along with the spectrum of ZIF-8. (d) Thermogravimetric analysis curves showing distinct mass losses between 220 and 280 °C, which are absent for reference ZIF-8. | ||
Similar to the BSA@ZIF-8 samples (Fig. S3), the PXRD patterns of the water-washed CM-dextran@ZIF-8, HA@ZIF-8, and GOx@ZIF-8 powders are dominated by the sod ZIF-8 phase, with ZIF-C remaining a secondary phase at up to 17% for CM-dextran@ZIF-8, 13% for HA@ZIF-8, and 22% for GOx@ZIF-8, as determined by the ZIF Phase Analysis app69 (Fig. S7–S9). After three cycles of water washing, residual linker remained in the CM-dextran@ZIF-8 and HA@ZIF-8 samples, as indicated by ATR-FTIR spectroscopy (Fig. S10 and S11). This may be due to linker entrapment within the dense ZIF-C phase and the agglomerated morphology of the extruded biocomposite, combined with the hydrophobicity of ZIF-8, all of which can hinder access to the washing medium. This contrasts with the solution-based syntheses typically yielding well-dispersed ZIF-8 crystals from which water can more readily remove excess linker.28 Ethanol washing, on the other hand, produces phase-pure BSA@ZIF-8, CM-dextran@ZIF-8, and HA@ZIF-8, with only a minor reflection of the secondary ZIF-C phase at 2θ ≈ 11° observed in the diffractogram of GOx@ZIF-8 (∼4%,69Fig. 2b). By examining the washed samples by ATR-FTIR spectroscopy, we observed the characteristic spectrum of ZIF-8 along with broad bands of biomolecules: BSA amide I band at 1600–1700 cm−1; CM-dextran C–OH stretching vibration at ∼1020 cm−1; HA amide I and II bands at 1540–1630 cm−1; and GOx amide I band at 1600–1700 cm−1 (Fig. 2c and S10–S12).74,75 The intensity of CM-dextran and HA bands increases from x = 5 to 15 in carbohydrate(x%)@ZIF-8 (Fig. S10 and S11), where x represents the carbohydrate weight proportion in the ideal end product (see Batch syntheses of biocomposites in SI for details). Thermogravimetric analysis (TGA) of the ethanol-washed samples shows distinct mass losses between 220 and 280 °C for the biocomposites, which are absent in pristine ZIF-8 (Fig. 2d and S13–S15). These mass losses are attributed to the decomposition of encapsulated biomolecules and increase from x = 5 to x = 15 for CM-dextran(x%)@ZIF-8 and HA(x%)@ZIF-8 (Fig. S13 and S14). The loading capacity (LC) and encapsulation efficiency (EE) for CM-dextran and HA at x = 10 are calculated as 5.2 wt% (EE% = 52%) and 6.4 wt% (EE% = 64%), respectively (see Procedures in the SI for details on evaluating biomolecule content from TGA curves). We note that the EE value for HA aligns with the ∼60% reported for the biomimetic mineralization approach.76 The GOx LC determined by BCA assay after biocomposite degradation was similar in the case of ethanol- and water-washed samples (i.e., 2.4 wt% and 2.2 wt%, respectively). Enzyme activity was measured in triplicate using the D-glucose assay (Fig. S16, see GOx catalytic assay in SI), resulting in values of 0.04 and 0.14 U/mgenzyme for ethanol- and water-washed samples, respectively. Control BSA@ZIF-8 samples exhibited no enzymatic activity (Fig. S16).
These data demonstrate that the current hand-mixing synthesis method can produce protein-, carbohydrate-, and enzyme-based biocomposites. Remarkably, GOx@ZIF-8 was enzymatically active. Furthermore, by using different washing procedures, we obtained two distinct phases, suggesting similarity with the water-based biomimetic mineralization approach.15 We note that previous works on ZIF biocomposites showed phase-dependent release profiles, suggesting that our dry method could be relevant for biomedicine. We further extended this approach to nucleic acids and lipids by preparing salmon DNA@ZIF-8 and EPA ethyl ester@ZIF-8 composites (see Batch syntheses of biocomposites and Fig. S17 and S18), confirming compatibility with all four major biomacromolecule classes – proteins, carbohydrates, nucleic acids, and lipids.77
Although a dedicated study would be required to examine in detail the structure–property relationship with a more systematic approach, the structural and catalytic data collected here enable us to preliminarily relate particle characteristics to the observed enzymatic performance of GOx@ZIF-8. Both water- and ethanol-washed GOx@ZIF-8 are predominantly sod ZIF-8 with different fractions of ZIF-C, as revealed by PXRD and quantitative phase analysis (Fig. 2b and SI S9). Together with the comparable GOx loadings determined by BCA assay (see SI), these data indicate that the higher specific activity of the water-washed material (SI Fig. S16) cannot be explained solely by the differences in enzyme content. Instead, we hypothesize that the enhanced activity depends on higher ZIF-C content and the resulting particle microstructure. In our system, ZIF-8 biocomposites form as nanosized crystallites that aggregate into larger particles (up to ∼100 µm), as exemplified by extruded BSA@ZIF-8 (Fig. 3d–f and SI Fig. S25). Such hierarchical particles provide diffusion pathways that couple interparticle voids with the microporous sod ZIF-8 framework. We propose that the increased fraction of ZIF-C and the associated microstructural heterogeneity in the water-washed GOx@ZIF-8 increase defect density and accessible transport into these micrometre-sized particles, enhancing mass-transfer for glucose and O2 and thus access to immobilized GOx. Conversely, the ethanol-washed sample, in which ZIF-C is present only as a minor phase, is likely to yield bulkier particles with a more homogeneous microporous framework.78 As a consequence, a higher effective diffusion resistance is expected, leading to a lower biocomposite specific activity under identical assay conditions.
Despite all these promising results, the hand-mixing method has limited reproducibility and scalability, restricting its potential for industrial production of MOF biocomposites. Nevertheless, our approach provides a basis for developing a continuous, reproducible, and large-scale approach to synthesizing MOF biocomposites via reactive extrusion.
Twin-screw extrusion
Twin-screw extrusion is a continuous and scalable synthetic approach, widely used in industry, where materials are processed by two rotating screws within a closed steel barrel. Its high reproducibility and versatility are ensured by precise temperature control, adjustable feed rates, screw profile design, and rotation speeds. Historically used in industry for shaping polymers, reactive extrusion has recently shown promise in sustainable large-scale chemical production, including MOFs, active pharmaceutical ingredients, and organic compounds,39,65,79–85 but, to our knowledge, its application in synthesizing MOF-based biocomposites has not yet been reported. Nevertheless, the recent implementation of TRIS monitoring techniques, such as Raman spectroscopy and energy-dispersive X-ray diffraction (EDXRD),35,40 has enhanced our understanding and control of extrusion reactions, particularly in MOF synthesis. This advancement facilitates an efficient transition from batch mixing reactions to continuous twin-screw extrusion for ZIF-8 biocomposite production. Building on our previous knowledge of reactive extrusion of ZIF-8, we explored this method for the continuous synthesis of ZIF-8-based biocomposites.Extrusion of ZIF-8
The procedure was first tested with pure ZIF-8 synthesis, without protein addition. In our synthesis, ZnCarb and HmeIm powders were mixed in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
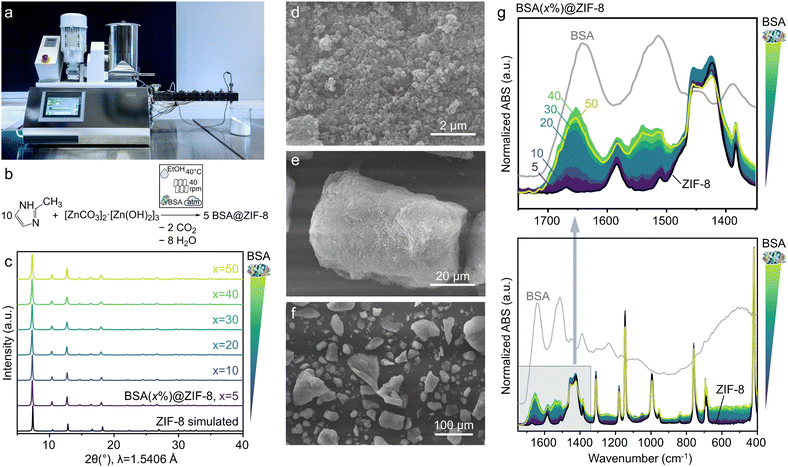
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 12.5 molar ratio and continuously fed into a preheated extruder barrel at 40 °C, along with a small amount of ethanol to initiate the reaction (Fig. 3a). After 20 minutes of extrusion, the white viscous paste was collected in three fractions (ZIF-8(1–3), 15 minutes each). The as-synthesized material was washed three times with ethanol to remove any unreacted linker, then immersed in a 10% sodium dodecyl sulfate (SDS) aqueous solution for 30 minutes, followed by three ethanol washes to remove residual SDS.86 The washed sample was dried overnight at room temperature under vacuum before analysis (see Reactive extrusion of biocomposites in SI for detailed experimental procedures). PXRD and ATR-FTIR analyses confirmed the consistent formation of phase-pure ZIF-8 with negligible unreacted precursors (Fig. S19 and S20). These findings are consistent with the TGA analysis (Fig. S21), which shows a weight loss of 64.5% at 400−650 °C, close to the theoretical weight loss of 64.2% observed in the ZIF-8-to-ZnO decomposition. Analysis of fractions collected at different times gave identical results, confirming consistent qualitative and quantitative yields and products during the continuous synthesis.
12.5 molar ratio and continuously fed into a preheated extruder barrel at 40 °C, along with a small amount of ethanol to initiate the reaction (Fig. 3a). After 20 minutes of extrusion, the white viscous paste was collected in three fractions (ZIF-8(1–3), 15 minutes each). The as-synthesized material was washed three times with ethanol to remove any unreacted linker, then immersed in a 10% sodium dodecyl sulfate (SDS) aqueous solution for 30 minutes, followed by three ethanol washes to remove residual SDS.86 The washed sample was dried overnight at room temperature under vacuum before analysis (see Reactive extrusion of biocomposites in SI for detailed experimental procedures). PXRD and ATR-FTIR analyses confirmed the consistent formation of phase-pure ZIF-8 with negligible unreacted precursors (Fig. S19 and S20). These findings are consistent with the TGA analysis (Fig. S21), which shows a weight loss of 64.5% at 400−650 °C, close to the theoretical weight loss of 64.2% observed in the ZIF-8-to-ZnO decomposition. Analysis of fractions collected at different times gave identical results, confirming consistent qualitative and quantitative yields and products during the continuous synthesis.
Extrusion of BSA@ZIF-8 biocomposites
Next, we investigated the extrusion synthesis of BSA@ZIF-8 by adding varying amounts of BSA powder to the ZIF-8 precursor mixture (Fig. 3b), thus preparing different BSA(x%)@ZIF-8 (x = 5, 10, 20, 30, 40, 50 wt%) samples. PXRD analysis of the washed extrudates confirms the formation of phase-pure ZIF-8 across all investigated x (Fig. 3c). Notably, the absolute intensity of characteristic ZIF-8 reflections decreases with x increasing from 5 to 50 (Fig. S22), indicating a reduction in the relative ZIF-8/BSA content within the ethanol-washed samples. For the sample with x = 20, the quantitative phase analysis (Fig. S23) shows that ethanol washing increases the fraction of the crystalline form of ZIF-8 in the sample from 25 to 43 wt%. Crystallite sizes estimated from PXRD data using the Scherrer equation decrease from ∼75 nm for pure ZIF-8 to ∼50 nm at x ≥ 30, where they reach a plateau (Fig. S24). This trend towards smaller crystal sizes with higher x is also evident in scanning electron microscopy (SEM) images, which show nanoparticles ranging from 50 to 200 nm (Fig. 3d and S25). These nanoparticles form agglomerates up to 100 µm (Fig. 3e, f and S25), distinguishing the morphology of extruded ZIF-8 and BSA@ZIF-8 from the individual cubic and rhombic ZIF-8 crystals, 50–2000 nm in size, typically produced in solution.87,88To confirm the presence of BSA in the washed solid, the samples were analyzed by ATR-FTIR spectroscopy. The spectra show the characteristic features of ZIF-8 together with the amide I (1610–1700 cm−1; primarily C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, with minor contributions from CN stretching, CCN deformation, and NH in-plane bending) and amide II bands (1500–1560 cm−1, mainly N–H in-plane bending and CN stretching) of BSA (Fig. 3g).89,90 This analysis confirms that BSA is either adsorbed on or encapsulated in ZIF-8. The amide I band for the BSA@ZIF-8 composites is shifted by up to 24 cm−1 towards higher wavenumbers compared to pure BSA. This shift is larger than that observed for BSA adsorbed on the surface of ZIF-8 nanoparticles (11 cm−1)6 and contrasts with the report of a redshift of 23 cm−1.91 The observed blueshift may arise from Zn2+ coordination to carbonyl oxygen sites, consistent with the hard and soft acid base (HSAB) theory considerations, accompanied by alterations in secondary structure,6,92 and non-covalent interactions such as van der Waals forces, hydrogen bonding, hydrophobic interactions, or electrostatic forces.6 The intensity of BSA bands increases with higher x, as demonstrated by integrating the amide I band of BSA and the ZIF-8 band at 420 cm−1 (Zn–N stretching) and plotting the BSA/ZIF-8 ratios in Fig. S26. These ratios indicate that the actual BSA loading in the ZIF matrix increases up to x = 40 and reaches a maximum. A slight decrease is observed for x = 50. This trend correlates with the crystallite sizes decreasing as the actual BSA content in the biocomposite increases (Fig. S24).
O stretching, with minor contributions from CN stretching, CCN deformation, and NH in-plane bending) and amide II bands (1500–1560 cm−1, mainly N–H in-plane bending and CN stretching) of BSA (Fig. 3g).89,90 This analysis confirms that BSA is either adsorbed on or encapsulated in ZIF-8. The amide I band for the BSA@ZIF-8 composites is shifted by up to 24 cm−1 towards higher wavenumbers compared to pure BSA. This shift is larger than that observed for BSA adsorbed on the surface of ZIF-8 nanoparticles (11 cm−1)6 and contrasts with the report of a redshift of 23 cm−1.91 The observed blueshift may arise from Zn2+ coordination to carbonyl oxygen sites, consistent with the hard and soft acid base (HSAB) theory considerations, accompanied by alterations in secondary structure,6,92 and non-covalent interactions such as van der Waals forces, hydrogen bonding, hydrophobic interactions, or electrostatic forces.6 The intensity of BSA bands increases with higher x, as demonstrated by integrating the amide I band of BSA and the ZIF-8 band at 420 cm−1 (Zn–N stretching) and plotting the BSA/ZIF-8 ratios in Fig. S26. These ratios indicate that the actual BSA loading in the ZIF matrix increases up to x = 40 and reaches a maximum. A slight decrease is observed for x = 50. This trend correlates with the crystallite sizes decreasing as the actual BSA content in the biocomposite increases (Fig. S24).
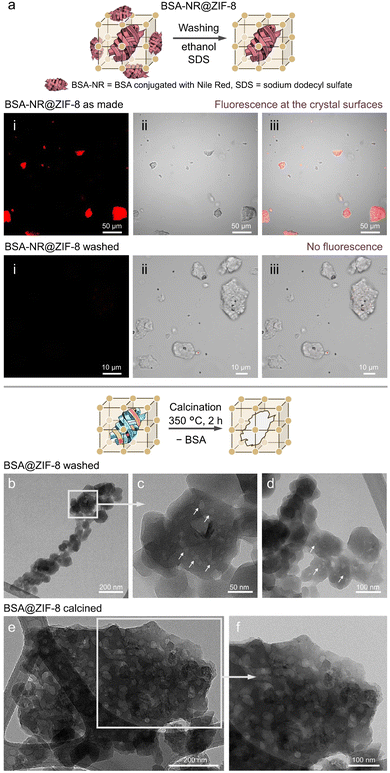
For practical applications of biocomposites, understanding the spatial distribution of biomolecules in/on MOFs is crucial. Specifically, it has been demonstrated that encapsulation is particularly effective for shielding biomolecules against chemical and mechanical stressors and could therefore be an enabling immobilization strategy for large-scale production. To ensure that reactive extrusion produces biocomposites with encapsulated biomolecules, we first examine potential infiltration mechanisms. As the pore window of sod ZIF-8 (0.34 nm)93 is one order of magnitude smaller than the diameter of BSA (6.6 nm),94 the literature dismisses this option. Then, to selectively remove surface adsorbed BSA, we followed a protocol from the literature,86 and the ZIF biocomposite was washed with ethanol and an aqueous 10% SDS surfactant solution. To validate the effectiveness of this washing protocol for our biocomposites, we used BSA tagged with the fluorescent stain Nile Red (NR), and we synthesized BSA-NR(10%)@ZIF-8. Confocal laser scanning microscopy (CLSM) of the as-made material reveals red fluorescence on the surface of biocomposite particles (Fig. 4a). As free Nile Red is non-emissive under the measurement conditions, the signal originates exclusively from BSA-NR. Variations in fluorescence intensity after washing steps reflect changes in adsorbed BSA-NR, providing insight into its retention behavior. Following the standard washing procedure (three ethanol washes, 30 minutes soaking in SDS, and three additional ethanol washes), CLSM images of the biocomposite showed no fluorescence at the crystal surface, indicating complete removal of adsorbed BSA-NR (Fig. 4a). In contrast, ethanol washing without SDS was insufficient to completely remove surface-adsorbed BSA-NR (Fig. S27). Furthermore, washes with pure water were ineffective in removing surface-adsorbed BSA-NR (Fig. S28 and S29). Thus, our data confirmed that the combined ethanol and aqueous SDS washes enable the quantitative removal of surface-adsorbed BSA. Therefore, the protein bands detected by ATR-FTIR in BSA@ZIF-8 samples (Fig. 3g) are due to the protein encapsulated within ZIF-8.
The encapsulation of BSA within ZIF-8 is further confirmed by transmission electron microscopy (TEM) measurements. TEM imaging of BSA(20%)@ZIF-8 reveals agglomerated rhombic dodecahedron crystals with regions of lower electron density, resembling pores (Fig. 4b–d). These regions are absent in pure ZIF-8 crystals (Fig. S30a and b) and range in size from 7 to 14 nm, which can be attributed to protein encapsulation.19 The lower boundary of 7 nm aligns with the radius of gyration of BSA within ZIF-8,18 while the 14 nm regions likely correspond to the inclusion of BSA dimers. TEM images of the biocomposite calcined at 350 °C for 2 hours reveal distinct mesoporous channels within the particle clusters (Fig. 4e and f), which are not present in the calcined pure ZIF-8 (Fig. S31). These mesopores are likely formed as a result of the decomposition of the encapsulated BSA.19 PXRD analysis of the calcined BSA@ZIF-8 and ZIF-8 samples confirms that both retain their crystallinity (Fig. S32). Unlike biocomposites obtained by solution-based approaches,19,95 not all crystals in the bulk BSA@ZIF-8 sample contain such pores, indicating inhomogeneities in the nanometer regime, possibly due to a different mechanism of biocomposite formation. In solution, biomolecules typically serve as nuclei for MOF formation, while in the solid state, the biomolecule is encapsulated due to the reaction of dense ZIF building blocks that yield the porous ZIF-8, resulting in a volumetric expansion of the final product.
Protein loading and encapsulation efficiency are decisive parameters determining the potential applications of MOF-based biocomposites. BSA loading was initially determined using the Bradford assay (see Procedures in SI).96 The results show a gradual increase in BSA loading as x increases, peaking at 27 wt% for the BSA(40%)@ZIF-8 sample (Fig. S33), followed by a decrease to 22 wt% for BSA(50%)@ZIF-8, in agreement with ATR-FTIR results (Fig. S26). Alternatively, TGA analysis provides a more statistically representative BSA loading value, analyzing three 30 mg fractions per sample (90 mg total) compared to the 5 mg used in the Bradford assay. The TGA of the BSA@ZIF-8 samples shows gradual mass losses starting at 170 °C, attributed to the decomposition of encapsulated BSA, followed by the decomposition of ZIF-8 that starts when T > 300 °C (Fig. 5a). After complete decomposition at 700 °C, the residual masses for BSA@ZIF-8 samples are smaller than those for pristine ZIF-8 and decrease with higher x. The calculated BSA loadings (see Procedures in SI) show a maximum of 26 wt% for the BSA(40%)@ZIF-8 sample (Fig. 5b), which is consistent with the 27 wt% obtained from the Bradford assay (Fig. S33). Encapsulation efficiencies, calculated based on the TGA values, show a maximum of 96% for the BSA(5%)@ZIF-8 and decrease with higher x (Fig. 5b).
The decrease in BSA loading observed for BSA(50%)@ZIF-8 can be attributed to complexation of BSA to the surfaces of growing ZIF-8 crystals,6 which sterically limits further crystal growth. The surface-bound protein is removed during ethanol/SDS washing, leading to a reduction in retained BSA at x = 50. This interpretation is consistent with the observed trend toward smaller crystallite sizes, as evidenced by PXRD and SEM data (Fig. S24 and S25).
The N2 adsorption isotherms collected at 77 K were of Type I (Fig. 5c),97 with the shape resembling that of the pristine ZIF-8 (Fig. S34), confirming the microporous nature of all BSA@ZIF-8 samples. The Brunauer−Emmett−Teller (BET) surface areas show a gradual decrease with increasing TGA-calculated BSA loading, from 1748 m2 g−1 for pristine ZIF-8 to 1019 m2 g−1 for BSA(40%)@ZIF-8. This decrease is ascribed to two factors: the addition of non-porous protein mass18 and the increased defectivity of the composite, as indicated by the reduced crystallite size (Fig. S24). At lower BSA loadings, reduced crystallinity is the main contributor to surface area reduction, whereas at higher BSA loadings, the addition of protein mass has a more significant impact (Fig. S35).
To examine the continuous production of the biocomposite with consistent composition, we conducted a 90-minute extrusion of BSA(10%)@ZIF-8. Four fractions were collected every 15 minutes, each for 5 minutes (Fig. S36). TGA analysis shows stable protein content across the fractions 1–4, with BSA loadings of 9.2, 8.7, 9.1, and 9.0 wt%, respectively (Fig. S36). The space-time yield (STY) during continuous operation, calculated from the mass of activated BSA(10%)@ZIF-8, reaches 2.96 × 104 kg m−3 d−1, with a production rate of 1.30 kg d−1 (see STY calculation in Procedures in the SI). This significantly outperforms the production rate of 0.05 kg d−1 previously achieved using a microfluidic reactor.28 As comparative data in Table S3 show, among all scalable continuous methods that could support ZIF-8 biocomposite production, reactive extrusion is the only approach that combines high STY, high production rate, and low solvent consumption at the same time. Since this work did not target maximal productivity, further optimisation of process parameters could yield even greater STY and production rates.
To assess the effect of extrusion temperature, additional experiments were performed at 35 °C, 45 °C, and with a gradient from 35 °C to 45 °C along the barrel. PXRD and FTIR confirmed that the ZIF-8 structure remained intact, while TGA revealed variations in protein loading (Fig. S37). These differences are attributed to temperature-induced changes from a paste-like to a denser extrudate and indicate that, although ZIF-8 formation itself is stable, biomolecule encapsulation can be sensitive even to moderate temperature variations.
Extrusion of ZIF-8-based drugs
Recent studies demonstrating MOF-based drugs as pH-responsive carriers for the encapsulation and controlled release of carbohydrate- and glycosaminoglycan-based therapeutics26,76 prompted us to investigate reactive extrusion for scalable synthesis of ZIF-8 encapsulated with the model carbohydrate-based therapeutic CM-dextran and the clinical drug HA.CM-dextran@ZIF-8 was synthesized following a procedure similar to BSA@ZIF-8, using a 75/25 v/v water/ethanol mixture as the auxiliary liquid (0.25 ml min−1, corresponding to η = 0.17 µl mg−1) instead of pure ethanol. PXRD and ATR-FTIR analyses confirmed the phase purity of the washed sample and the successful encapsulation of CM-dextran within the ZIF-8 framework (Fig. S38 and S39), while TGA indicated a CM-dextran loading of 11 wt% (Fig. S40). SEM images showed nanoparticles up to 200 nm forming agglomerates (Fig. S41), resembling those observed in the extruded BSA@ZIF-8 sample (Fig. S25). The procedure was also effective in producing HA@ZIF-8 with HA loading of 10.0 wt% (Fig. S42–S44). In all cases, the ZIF matrix formed with sod topology. To evaluate the long-term stability, HA@ZIF-8 stored at 4 °C for seven months was analysed by PXRD, ATR-FTIR, and TGA (Fig. S45). The data show no significant changes compared to the freshly prepared material, demonstrating that biocomposite remains stable under these conditions.
Generally, MOFs and MOF-based biocomposites are produced as crystalline powders, which are difficult to process, integrate, and recycle.98 To improve the compatibility of MOFs for industrial use, various shaping protocols are used, like tableting, granulation, or post-synthetic extrusion.79,99,100 However, these approaches often require binders and need precise optimization to balance porosity and mechanical strength.79 Here, we produced shaped HA@ZIF-8 extrudate in a single step directly from the reagent powders without external binders. This was achieved by installing a die faceplate and following the standard synthetic procedure. The resulting extrudate with a nominal diameter of 2 mm (Fig. 6a and b) retains its shape when soaked in ethanol for at least 3 days and, after drying, can be pelletized for better processability (Fig. 6c). PXRD and ATR-FTIR confirm the sod ZIF-8 phase of both ethanol-washed HA@ZIF-8 shaped extrudate and its powder analogue and encapsulation of HA within the ZIF-8 framework (Fig. 6d and e). N2 adsorption experiments on HA@ZIF-8 in powder form and as shaped extrudate (Fig. 6f and S46) show Type I adsorption isotherms with the shape resembling those of the pristine ZIF-8 and BSA@ZIF-8 samples (Fig. S34), indicating microporous materials.97 The HA@ZIF-8 shaped extrudate exhibits slightly higher N2 uptake and BET surface area than its powder counterpart (1482 vs. 1433 m2 g−1, Fig. 6f), consistent with its slightly lower content of non-porous HA, as indicated by TGA (Fig. S47). The shaped extrudate was characterized by SEM (Fig. 6g and h), showing nanoparticles up to 200 nm on both the surface (Fig. S48) and cross-section (Fig. 6g), similar to the extruded BSA@ZIF-8 powder samples (Fig. 3d and S25). However, exfoliation is observed at the extrudate's edges (Fig. 6h), where partial crystallite grain coalescence occurs (Fig. S49), likely induced by shear forces at the die during extrusion.
To determine whether the extrusion protocol, washing procedure, or the chemical environment of ZIF-8 affects the molecular weight of HA, we analyzed the molar mass distribution of HA released from both HA@ZIF-8 powders and shaped extrudates. For this purpose, the ZIF-8 matrix was dissolved under acidic conditions (see Procedures in the ESI), releasing HA, which was subsequently analyzed by size exclusion chromatography (SEC). The resulting molar mass distribution of HA shows only a slight shift toward lower masses on the high-molecular-weight side, compared to HA prior to encapsulation and the HA/ZIF-8 physical mixture references (Fig. 6h). Given the good reproducibility, this shift corresponds to a reduction in Mp and Mw of less than 8% (Table S4). However, considering the SEC method's error margins of approximately 10%, we conclude that the HA@ZIF-8 production process, both for powders and shaped extrudates, does not result in measurable HA degradation. In addition, SEC analysis of HA released from HA@ZIF-8 after 7 months of storage at 4 °C (Fig. S50) revealed molar mass distributions in very good agreement with the initial measurements (Table S4), indicating preserved HA molecular integrity upon long-term storage.
We have successfully upscaled biocomposite synthesis from 200 mg batch reactions to continuous kilogram-scale reactive extrusion. Unlike water-based encapsulation approaches, where negatively charged biomolecules induce MOF formation at their surface,13 the solid-state procedure does not seem to have the same effect. Instead, encapsulation likely results from the 7.6-fold volume expansion during the ZnCarb-to-ZIF-8 transformation, estimated based on the molecular weights and densities of ZnCarb and ZIF-8 using Tanaka et al.‘s formula.101 As ZIF-8 forms and expands, it entraps biomolecules within its structure, facilitating biocomposite formation. This mechanism could enable biomolecule encapsulation regardless of surface charge or solubility and may be compatible with various mechanochemically accessible materials.30,58 While a higher biomolecule content (x = 50 and above) can hinder efficient ZIF-8 formation and reduce biomolecule loading due to increased BSA adsorption on growing crystals, this issue may be mitigated by increasing the linker excess. Additionally, thorough homogenization of the reaction mixture and reduction of biomolecule particle size may enhance uniform biomolecule distribution in the final product.
In view of their scalable production, these ZIF-8 biocomposites are promising for various practical applications, as discussed in recent reviews on MOF biocomposites.13,16,102–104 BSA@ZIF-8, a model protein, exemplifies protein@MOF composites used in drug delivery, biobanking/biopreservation, and biosensing.104,105 GOx@ZIF-8 highlights applications in glucose biosensing and insulin delivery systems.106,107 CM-dextran@ZIF-8 serves as a model system for carbohydrate-based biotherapeutics,26 while HA@ZIF-8 demonstrates the relevance of this approach to commercially important GAGs, with potential in anti-inflammatory and anticoagulant therapies, as well as wound healing and tissue engineering.104,108 Salmon DNA@ZIF-8 represents nucleic acid@MOF composites, with potential applications in gene expression regulation, silencing, and editing, as well as in drug delivery and cell manipulation.104,109–111 Finally, encapsulation of EPA ethyl ester, a derivative of the commercially important omega-3 fatty acid EPA,112 illustrates the potential of lipid@MOF composites as delivery vehicles for hydrophobic drugs.113
Overall, the demonstrated compatibility with diverse biomolecules, the mild synthesis conditions, and the advantages of reactive extrusion – including reduced solvent consumption, continuous processing, and scalability – make this approach particularly attractive for the production of functional biocomposites. In addition, the direct one-step formation of shaped extrudates provides a route toward 3D printing of biocomposites using a twin-screw extruder printing head.104,108,113,114
Conclusions
In conclusion, we have established the first solvent-minimized, room-temperature continuous reactive extrusion process for the one-step production of ZIF-8 biocomposites. This route addresses key challenges such as reproducibility, biomolecule stability, encapsulation efficiency, and the preservation of biological function – limitations that have, until now, hindered the translation of MOF biocomposites from the laboratory to industry.The synthesis involves mixing basic zinc carbonate, 2-methylimidazole, and biomolecule powders with a small amount of liquid, either ethanol or water, corresponding to 0.30 µl per mg of solid (23 wt%). Our small-scale trial experiments using simple hand-mixing batch reactions successfully produced protein-, carbohydrate-, enzyme-, lipid-, and DNA-encapsulated ZIF-8 in 5 minutes. This simple and rapid process preserved the enzymatic activity of the encapsulated glucose oxidase and allowed partial conversion of sodalite ZIF-8 to its carbonate derivative, ZIF-C, by post-synthetic washes.
Next, we adapted the batch procedure to reactive extrusion and tested it with different loadings of bovine serum albumin (BSA). This reaction produced pure, highly crystalline, and porous BSA@ZIF-8 biocomposites, achieving an encapsulation efficiency of up to 96% and a maximum BSA loading of 26 wt%. Our data demonstrate that in continuous operation, the reaction maintains consistent protein content over time and can yield 1.30 kg d−1 of activated product, with a space-time yield of 2.96 × 104 kg m−3 d−1. In addition, reactive extrusion effectively produced ZIF-8-based drugs encapsulating a model carbohydrate-based therapeutic, CM-dextran, and a clinical therapeutic, hyaluronic acid (HA). Finally, to improve the processability of extruded biocomposites, we employed a 2 mm die faceplate to shape HA@ZIF-8 extrudates in a single step, directly from reagent powders. HA released from ZIF-8 retains its initial molar mass, indicating excellent preservation of molecular integrity and highlighting the suitability of reactive extrusion for the large-scale production of biomolecule@ZIF materials.
Our study directly addresses key limitations hindering the biomedical application of MOFs – namely, upscaling, reproducibility, and green synthesis. Given the wide range of porous frameworks available by mechanochemical methods and fewer restrictions on biomolecule selection due to the absence of bulk solvent, we anticipate that this work will encourage the exploration of previously unattainable biocomposites.
Author contributions
CRediT: conceptualization: NG, FC, PF, FE; funding acquisition: NG, JAV, FE; investigation: NG, AS, IT, JF; methodology: NG, FC, PF; project administration: NG; resources: PF, FE; supervision: JAV, PF, FE; visualization: NG; writing – original draft: NG, AS, FC; writing – review & editing: NG, AS, FC, IT, JF, JAV, PF, FE.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: experimental procedures, synthesis protocols for biocomposites, details of reactive extrusion, optimization studies, methods for biomolecule quantification and activity assays, as well as supporting tables and figures. See DOI: https://doi.org/10.1039/d5ta08276e.Acknowledgements
Parts of this work were performed at the electron microscopy center at BAM. The authors acknowledge Ines Feldmann (BAM, Berlin) for SEM measurements, Carsten Prinz (BAM, Berlin) for TEM measurements, Annett Zimathies and Carsten Prinz (BAM, Berlin) for the measurements of N2 adsorption. The authors also thank Dr Kirill Yusenko (Ruhr-Universität Bochum) for the quantitative phase analysis. Furthermore, N. G. thanks Michael Hafner (TU Graz) and Catharina Masser (TU Graz) for their assistance in the laboratory. N. G. acknowledges funding from the School of Analytical Sciences Adlershof (SALSA STF23-02). Jose A. Villajos acknowledges funding from European Union's Next Generation EU funds, coordinated at national level by the Recovery, Transformation and Resilience Plan.Notes and references
- A. A. Michalchuk, E. V. Boldyreva, A. M. Belenguer, F. Emmerling and V. V. Boldyrev, Front. Chem., 2021, 9, 685789 Search PubMed.
- Crystal Structure of Bovine Serum Albumin (Version 2.X), NIH 3D, 2016, DOI:10.60705/3DPX/3447.2.
- Glucose Oxidase from Apergillus Niger (Version 2.X). NIH 3D., 2018, DOI:10.60705/3DPX/9567.2.
- G. Madhariya, S. Diwan, R. Chauhan, N. K. Chandrawanshi and P. K. Mahish, in Handbook of Biomolecules, ed. C. Verma and D. K. Verma, Elsevier, 2023, pp. 397–418, DOI:10.1016/b978-0-323-91684-4.00027-x.
- J. V. Gade, A. Singh and B. Jain, in Handbook of Biomolecules, ed C. Verma and D. K. Verma, Elsevier, 2023, pp. 551–574, DOI:10.1016/b978-0-323-91684-4.00019-0.
- A. Mittal, S. Gandhi and I. Roy, Sci. Rep., 2022, 12, 10331 Search PubMed.
- W. Xu, L. Jiao, Y. Wu, L. Hu, W. Gu and C. Zhu, Adv. Mater., 2021, 33, e2005172 CrossRef PubMed.
- N. Zheng, Y. Cai, Z. Zhang, H. Zhou, Y. Deng, S. Du, M. Tu, W. Fang and X. Xia, Nat. Commun., 2025, 16, 604 CrossRef CAS PubMed.
- R. P. Welch, H. Lee, M. A. Luzuriaga, O. R. Brohlin and J. J. Gassensmith, Bioconjugate Chem., 2018, 29, 2867–2883 CrossRef CAS PubMed.
- A. Madhavan, R. Sindhu, P. Binod, R. K. Sukumaran and A. Pandey, Bioresour. Technol., 2017, 245, 1304–1313 CrossRef CAS PubMed.
- C. Doonan, R. Ricco, K. Liang, D. Bradshaw and P. Falcaro, Acc. Chem. Res., 2017, 50, 1423–1432 CrossRef CAS PubMed.
- R. Murty, M. K. Bera, I. M. Walton, C. Whetzel, M. R. Prausnitz and K. S. Walton, J. Am. Chem. Soc., 2023, 145, 7323–7330 CrossRef CAS PubMed.
- W. Liang, P. Wied, F. Carraro, C. J. Sumby, B. Nidetzky, C. K. Tsung, P. Falcaro and C. J. Doonan, Chem. Rev., 2021, 121, 1077–1129 CrossRef CAS PubMed.
- C. Wu, X. Zhao, D. Wang, X. Si and T. Li, Chem. Sci., 2022, 13, 13338–13346 RSC.
- F. Carraro, M. J. Velasquez-Hernandez, E. Astria, W. Liang, L. Twight, C. Parise, M. Ge, Z. Huang, R. Ricco, X. Zou, L. Villanova, C. O. Kappe, C. Doonan and P. Falcaro, Chem. Sci., 2020, 11, 3397–3404 RSC.
- Y. H. Wijesundara, T. S. Howlett, S. Kumari and J. J. Gassensmith, Chem. Rev., 2024, 124, 3013–3036 CrossRef CAS PubMed.
- A. V. Desai, S. M. Vornholt, L. L. Major, R. Ettlinger, C. Jansen, D. N. Rainer, R. de Rome, V. So, P. S. Wheatley, A. K. Edward, C. G. Elliott, A. Pramanik, A. Karmakar, A. R. Armstrong, C. Janiak, T. K. Smith and R. E. Morris, ACS Appl. Mater. Interfaces, 2023, 15, 9058–9065 CrossRef CAS PubMed.
- K. Liang, R. Ricco, C. M. Doherty, M. J. Styles, S. Bell, N. Kirby, S. Mudie, D. Haylock, A. J. Hill, C. J. Doonan and P. Falcaro, Nat. Commun., 2015, 6, 1–8 Search PubMed.
- F. Lyu, Y. Zhang, R. N. Zare, J. Ge and Z. Liu, Nano Lett., 2014, 14, 5761–5765 CrossRef CAS PubMed.
- F.-K. Shieh, S.-C. Wang, C.-I. Yen, C.-C. Wu, S. Dutta, L.-Y. Chou, J. V. Morabito, P. Hu, M.-H. Hsu and K. C.-W. Wu, J. Am. Chem. Soc., 2015, 137, 4276–4279 CrossRef CAS PubMed.
- K. Y. Wang, J. Zhang, Y. C. Hsu, H. Lin, Z. Han, J. Pang, Z. Yang, R. R. Liang, W. Shi and H. C. Zhou, Chem. Rev., 2023, 123, 5347–5420 CrossRef CAS PubMed.
- C. Wang, S. Tadepalli, J. Luan, K. K. Liu, J. J. Morrissey, E. D. Kharasch, R. R. Naik and S. Singamaneni, Adv. Mater., 2017, 29, 1604433 CrossRef.
- S. Li, M. Dharmarwardana, R. P. Welch, C. E. Benjamin, A. M. Shamir, S. O. Nielsen and J. J. Gassensmith, ACS Appl. Mater. Interfaces, 2018, 10, 18161–18169 CrossRef CAS.
- K. Liang, J. J. Richardson, J. Cui, F. Caruso, C. J. Doonan and P. Falcaro, Adv. Mater., 2016, 28, 7910–7914 CrossRef CAS PubMed.
- N. K. Maddigan, A. Tarzia, D. M. Huang, C. J. Sumby, S. G. Bell, P. Falcaro and C. J. Doonan, Chem. Sci., 2018, 9, 4217–4223 RSC.
- E. Astria, M. Thonhofer, R. Ricco, W. B. Liang, A. Chemelli, A. Tarzia, K. Alt, C. E. Hagemeyer, J. Rattenberger, H. Schroettner, T. Wrodnigg, H. Amenitsch, D. M. Huang, C. J. Doonan and P. Falcaro, Mater. Horiz., 2019, 6, 969–977 RSC.
- S. Kumari, R. N. Ehrman and J. J. Gassensmith, Matter, 2023, 6, 2570–2573 CrossRef CAS.
- F. Carraro, J. D. Williams, M. Linares-Moreau, C. Parise, W. Liang, H. Amenitsch, C. Doonan, C. O. Kappe and P. Falcaro, Angew. Chem., Int. Ed., 2020, 59, 8123–8127 CrossRef CAS PubMed.
- F. Afshariazar and A. Morsali, J. Mater. Chem. A, 2022, 10, 15332–15369 RSC.
- S. Głowniak, B. Szczęśniak, J. Choma and M. Jaroniec, Mater. Today, 2021, 46, 109–124 CrossRef.
- B. Szczęśniak, S. Borysiuk, J. Choma and M. Jaroniec, Mater. Horiz., 2020, 7, 1457–1473 RSC.
- J. M. Marrett, F. Effaty, X. Ottenwaelder and T. Friscic, Adv. Mater., 2025, 37, e2418707 Search PubMed.
- A. A. L. Michalchuk and F. Emmerling, Angew. Chem., Int. Ed., 2022, 61, e202117270 CrossRef CAS PubMed.
- T. Friščić, I. Halasz, P. J. Beldon, A. M. Belenguer, F. Adams, S. A. Kimber, V. Honkimäki and R. E. Dinnebier, Nat. Chem., 2013, 5, 66–73 CrossRef PubMed.
- N. Y. Gugin, K. V. Yusenko, A. King, K. Meyer, D. Al-Sabbagh, J. A. Villajos and F. Emmerling, Chem, 2024, 10, 3459–3473 CAS.
- K. J. Ardila-Fierro and J. G. Hernandez, Chemsuschem, 2021, 14, 2145–2162 CrossRef CAS PubMed.
- E. Colacino, V. Isoni, D. Crawford and F. García, Trends Chem., 2021, 3, 335–339 Search PubMed.
- J. F. Reynes, V. Isoni and F. Garcia, Angew. Chem., Int. Ed., 2023, 62, e202300819 Search PubMed.
- D. Crawford, J. Casaban, R. Haydon, N. Giri, T. McNally and S. L. James, Chem. Sci., 2015, 6, 1645–1649 Search PubMed.
- N. Y. Gugin, J. A. Villajos, O. Dautain, M. Maiwald and F. Emmerling, ACS Sustainable Chem. Eng., 2023, 11, 5175–5183 Search PubMed.
- B. Karadeniz, A. J. Howarth, T. Stolar, T. Islamoglu, I. Dejanovic, M. Tireli, M. C. Wasson, S. Y. Moon, O. K. Farha, T. Friscic and K. Uzarevic, ACS Sustainable Chem. Eng., 2018, 6, 15841–15849 Search PubMed.
- W. Wang, M. Chai, M. Y. B. Zulkifli, K. Xu, Y. Chen, L. Wang, V. Chen and J. Hou, Mol. Syst. Des. Eng., 2023, 8, 560–579 RSC.
- C. Orellana-Tavra, E. F. Baxter, T. Tian, T. D. Bennett, N. K. Slater, A. K. Cheetham and D. Fairen-Jimenez, Chem. Commun., 2015, 51, 13878–13881 Search PubMed.
- C. Orellana-Tavra, R. J. Marshall, E. F. Baxter, I. A. Lazaro, A. Tao, A. K. Cheetham, R. S. Forgan and D. Fairen-Jimenez, J. Mater. Chem. B, 2016, 4, 7697–7707 Search PubMed.
- J. Q. Sha, L. H. Wu, S. X. Li, X. N. Yang, Y. Zhang, Q. N. Zhang and P. P. Zhu, J. Mol. Struct., 2015, 1101, 14–20 CrossRef CAS.
- S. Rojas, F. J. Carmona, C. R. Maldonado, P. Horcajada, T. Hidalgo, C. Serre, J. A. R. Navarro and E. Barea, Inorg. Chem., 2016, 55, 2650–2663 CrossRef CAS.
- S. A. Noorian, N. Hemmatinejad and J. A. R. Navarro, Microporous Mesoporous Mater., 2020, 302, 110199 CrossRef CAS.
- C. Pina-Vidal, V. Berned-Samatan, E. Piera, M. A. Caballero and C. Tellez, Polymers, 2024, 16, 637 CrossRef CAS PubMed.
- H. He, H. Han, H. Shi, Y. Tian, F. Sun, Y. Song, Q. Li and G. Zhu, ACS Appl. Mater. Interfaces, 2016, 8, 24517–24524 CrossRef CAS.
- T.-H. Wei, S.-H. Wu, Y.-D. Huang, W.-S. Lo, B. P. Williams, S.-Y. Chen, H.-C. Yang, Y.-S. Hsu, Z.-Y. Lin and X.-H. Chen, Nat. Commun., 2019, 10, 1–8 Search PubMed.
- E. Ozyilmaz and O. Caglar, Process Biochem., 2023, 134, 276–285 CrossRef CAS.
- J. Nawrocki, D. Prochowicz, A. Wisniewski, I. Justyniak, P. Gos and J. Lewinski, Eur. J. Inorg. Chem., 2020, 2020, 796–800 Search PubMed.
- B. E. Souza and J. C. Tan, Crystengcomm, 2020, 22, 4526–4530 RSC.
- P. K. Lam, T. H. Vo, J. H. Chen, S. W. Lin, C. L. Kuo, J. J. Liao, K. Y. Chen, S. R. Huang, D. Li, Y. H. Chang, H. Y. Chen, H. T. Hsieh, Y. A. Hsu, H. K. Tsao, H. C. Yang and F. K. Shieh, J. Mater. Chem. A, 2023, 11, 24678–24685 RSC.
- S. Wang, W. S. Liu, J. Wang, J. Y. Yu, F. D. Wang, C. A. Jin, K. Y. Wang, P. Cheng, Z. J. Zhang and Y. Chen, ACS Mater. Lett., 2024, 6, 2609–2616 CrossRef CAS.
- M. F. Chaplin and C. Bucke, Enzyme Technology, Cambridge University Press, 1990 Search PubMed.
- C. L. Luu, T. T. V. Nguyen, T. Nguyen and T. C. Hoang, Adv. Nat. Sci.: Nanosci., 2015, 6, 025004 Search PubMed.
- Q. Chen, Z.-W. Li, S. Huang, G. Chen and G. Ouyang, RSC Mechanochem., 2025, 2, 336–350 RSC.
- Y. Quan, R. Shen, R. Ma, Z. Zhang and Q. Wang, ACS Sustainable Chem. Eng., 2022, 10, 7216–7222 Search PubMed.
- S. Tanaka, K. Kida, T. Nagaoka, T. Ota and Y. Miyake, Chem. Commun., 2013, 49, 7884–7886 Search PubMed.
- P. J. Beldon, L. Fabian, R. S. Stein, A. Thirumurugan, A. K. Cheetham and T. Friscic, Angew. Chem., Int. Ed., 2010, 49, 9640–9643 CrossRef CAS PubMed.
- I. Brekalo, W. Yuan, C. Mottillo, Y. Lu, Y. Zhang, J. Casaban, K. T. Holman, S. L. James, F. Duarte, P. A. Williams, K. D. M. Harris and T. Friscic, Chem. Sci., 2020, 11, 2141–2147 Search PubMed.
- J. Andersen, H. Starbuck, T. Current, S. Martin and J. Mack, Green Chem., 2021, 23, 8501–8509 Search PubMed.
- Q. Cao, J. L. Howard, D. E. Crawford, S. L. James and D. L. Browne, Green Chem., 2018, 20, 4443–4447 Search PubMed.
- R. R. A. Bolt, J. A. Leitch, A. C. Jones, W. I. Nicholson and D. L. Browne, Chem. Soc. Rev., 2022, 51, 4243–4260 Search PubMed.
- D. E. Crawford, S. L. James and T. McNally, ACS Sustainable Chem. Eng., 2018, 6, 193–201 Search PubMed.
- N. Gugin, J. A. Villajos, I. Feldmann and F. Emmerling, RSC Adv., 2022, 12, 8940–8944 Search PubMed.
- T. Friščić, S. L. Childs, S. A. Rizvi and W. Jones, Crystengcomm, 2009, 11, 418–426 Search PubMed.
- M. R. Hafner, L. Villanova and F. Carraro, Crystengcomm, 2022, 24, 7266–7271 RSC.
- C. Mottillo and T. Friscic, Angew. Chem., Int. Ed., 2014, 53, 7471–7474 CrossRef CAS PubMed.
- S. B. Bankar, M. V. Bule, R. S. Singhal and L. Ananthanarayan, Biotechnol. Adv., 2009, 27, 489–501 CrossRef CAS PubMed.
- G. Abatangelo, V. Vindigni, G. Avruscio, L. Pandis and P. Brun, Cells, 2020, 9, 1743 Search PubMed.
- M. J. Velasquez-Hernandez, E. Astria, S. Winkler, W. Liang, H. Wiltsche, A. Poddar, R. Shukla, G. Prestwich, J. Paderi, P. Salcedo-Abraira, H. Amenitsch, P. Horcajada, C. J. Doonan and P. Falcaro, Chem. Sci., 2020, 11, 10835–10843 Search PubMed.
- Y. Liu, Z. Li, S. Zou, C. Lu, Y. Xiao, H. Bai, X. Zhang, H. Mu, X. Zhang and J. Duan, Int. J. Biol. Macromol., 2020, 155, 103–109 Search PubMed.
- G. Liu, R. Y. Hong, L. Guo, Y. G. Li and H. Z. Li, Appl. Surf. Sci., 2011, 257, 6711–6717 Search PubMed.
- M. d. J. Velásquez-Hernández, E. Astria, S. Winkler, W. Liang, H. Wiltsche, A. Poddar, R. Shukla, G. Prestwich, J. Paderi, P. Salcedo-Abraira, H. Amenitsch, P. Horcajada, C. J. Doonan and P. Falcaro, Chem. Sci., 2020, 11, 10835–10843 Search PubMed.
- J. B. Reece, L. A. Urry, M. L. Cain, S. A. Wasserman, P. V. Minorsky and R. B. Jackson, Campbell Biology, Pearson Education, 2011 Search PubMed.
- S. A. Polash, A. Poddar, F. Carraro, G. Bryant, P. Falcaro and R. Shukla, RSC Appl. Interfaces, 2026, 3, 176–191 Search PubMed.
- J. Casaban, Y. Zhang, R. Pacheco, C. Coney, C. Holmes, E. Sutherland, C. Hamill, J. Breen, S. L. James, D. Tufano, D. Wong, E. Stavrakakis, H. Annath and A. Moore, Faraday Discuss., 2021, 231, 312–325 RSC.
- B. D. Egleston, M. C. Brand, F. Greenwell, M. E. Briggs, S. L. James, A. I. Cooper, D. E. Crawford and R. L. Greenaway, Chem. Sci., 2020, 11, 6582–6589 Search PubMed.
- O. Galant, G. Cerfeda, A. S. McCalmont, S. L. James, A. Porcheddu, F. Delogu, D. E. Crawford, E. Colacino and S. Spatari, ACS Sustainable Chem. Eng., 2022, 10, 1430–1439 Search PubMed.
- T. Stolar, S. Lukin, M. Tireli, I. Sovic, B. Karadeniz, I. Kerekovic, G. Matijasic, M. Gretic, Z. Katancic, I. Dejanovic, M. di Michiel, I. Halasz and K. Uzarevic, ACS Sustainable Chem. Eng., 2019, 7, 7102–7110 Search PubMed.
- Q. Cao, D. E. Crawford, C. Shi and S. L. James, Angew. Chem., Int. Ed., 2020, 59, 4478–4483 CrossRef CAS PubMed.
- T. Stolar, S. Grubesic, N. Cindro, E. Mestrovic, K. Uzarevic and J. G. Hernandez, Angew. Chem., Int. Ed., 2021, 60, 12727–12731 CrossRef CAS PubMed.
- V. Chantrain, T. Rensch, W. Pickhardt, S. Gratz and L. Borchardt, Chem.–Eur. J., 2024, 30, e202304060 CrossRef CAS PubMed.
- W. Liang, R. Ricco, N. K. Maddigan, R. P. Dickinson, H. Xu, Q. Li, C. J. Sumby, S. G. Bell, P. Falcaro and C. J. Doonan, Chem. Mater., 2018, 30, 1069–1077 CrossRef CAS.
- A. Schejn, L. Balan, V. Falk, L. Aranda, G. Medjahdi and R. Schneider, Crystengcomm, 2014, 16, 4493–4500 Search PubMed.
- N. L. Torad, M. Hu, Y. Kamachi, K. Takai, M. Imura, M. Naito and Y. Yamauchi, Chem. Commun., 2013, 49, 2521–2523 Search PubMed.
- A. Barth, Biochim. Biophys. Acta, 2007, 1767, 1073–1101 CrossRef CAS PubMed.
- W. Liang, H. Xu, F. Carraro, N. K. Maddigan, Q. Li, S. G. Bell, D. M. Huang, A. Tarzia, M. B. Solomon and H. Amenitsch, J. Am. Chem. Soc., 2019, 141, 2348–2355 Search PubMed.
- D. Tocco, D. Chelazzi, R. Mastrangelo, A. Casini, A. Salis, E. Fratini and P. Baglioni, J. Colloid Interface Sci., 2023, 641, 685–694 CrossRef CAS PubMed.
- Y. Feng, A. Schmidt and R. A. Weiss, Macromolecules, 1996, 29, 3909–3917 Search PubMed.
- K. S. Park, Z. Ni, A. P. Cote, J. Y. Choi, R. Huang, F. J. Uribe-Romo, H. K. Chae, M. O'Keeffe and O. M. Yaghi, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 10186–10191 Search PubMed.
- R. G. Queiroz, G. H. Varca, S. Kadlubowski, P. Ulanski and A. B. Lugao, Int. J. Biol. Macromol., 2016, 85, 82–91 Search PubMed.
- H. Zheng, Y. Zhang, L. Liu, W. Wan, P. Guo, A. M. Nystrom and X. Zou, J. Am. Chem. Soc., 2016, 138, 962–968 CrossRef CAS PubMed.
- M. M. Bradford, Anal. Biochem., 1976, 72, 248–254 CrossRef CAS PubMed.
- K. S. W. Sing, D. H. Everett, R. A. W. Haul, L. Moscou, R. A. Pierotti, J. Rouquerol and T. Siemieniewska, Pure Appl. Chem., 1985, 57, 603–619 CrossRef CAS.
- X. Qi, K. Liu and Z. Chang, Chem. Eng. J., 2022, 441, 135953 Search PubMed.
- D. Chakraborty, A. Yurdusen, G. Mouchaham, F. Nouar and C. Serre, Adv. Funct. Mater., 2023, 34, 2309089 Search PubMed.
- J. A. Powell, J. A. D. Marquez, G. A. Johnson, R. O. K. Ozdemir and Q. S. Wang, Ind. Eng. Chem. Res., 2024, 63, 21290–21299 Search PubMed.
- S. Tanaka, K. Sakamoto, H. Inada, M. Kawata, G. Takasaki and K. Imawaka, Langmuir, 2018, 34, 7028–7033 Search PubMed.
- W. He, J. Guo, Y. Qiu, J. Zhang, C. Xue, Z. Wang and J. Zhu, Coord. Chem. Rev., 2026, 546, 217052 CrossRef CAS.
- A. L. Wang, M. Walden, R. Ettlinger, F. Kiessling, J. J. Gassensmith, T. Lammers, S. Wuttke and Q. Peña, Adv. Funct. Mater., 2023, 2308589, DOI:10.1002/adfm.202308589.
- M. d. J. Velásquez-Hernández, M. Linares-Moreau, E. Astria, F. Carraro, M. Z. Alyami, N. M. Khashab, C. J. Sumby, C. J. Doonan and P. Falcaro, Coord. Chem. Rev., 2021, 429, 213651 CrossRef.
- C. Z. Wang, H. C. Sun, J. Y. Luan, Q. S. Jiang, S. Tadepalli, J. J. Morrissey, E. D. Kharasch and S. Singamaneni, Chem. Mater., 2018, 30, 1291–1300 CrossRef CAS.
- M. Mohammad, A. Razmjou, K. Liang, M. Asadnia and V. Chen, ACS Appl. Mater. Interfaces, 2019, 11, 1807–1820 CrossRef CAS.
- C. Zhang, S. Hong, M. D. Liu, W. Y. Yu, M. K. Zhang, L. Zhang, X. Zeng and X. Z. Zhang, J. Controlled Release, 2020, 320, 159–167 CrossRef CAS PubMed.
- A. Bigham, N. Islami, A. Khosravi, A. Zarepour, S. Iravani and A. Zarrabi, Small, 2024, 20, e2311903 CrossRef PubMed.
- S. A. Polash, A. Poddar, S. Pyreddy, F. Carraro, A. M. D'Angelo, G. Bryant, P. Falcaro and R. Shukla, ACS Appl. Mater. Interfaces, 2025, 17, 3002–3012 CrossRef CAS.
- A. Poddar, J. J. Conesa, K. Liang, S. Dhakal, P. Reineck, G. Bryant, E. Pereiro, R. Ricco, H. Amenitsch, C. Doonan, X. Mulet, C. M. Doherty, P. Falcaro and R. Shukla, Small, 2019, 15, e1902268 CrossRef PubMed.
- B. Arora, A. Yoon, M. Sriram, P. Singha and S. S. H. Rizvi, Innovative Food Sci. Emerging Technol., 2020, 64, 102429 CrossRef CAS.
- N. A. Auchterlonie and G. H. Bescoby, in Omega-3 Delivery Systems, ed P. J. García-Moreno, C. Jacobsen, A.-D. Moltke Sørensen and B. Yesiltas, Academic Press, 2021, pp. 79–106, DOI:10.1016/b978-0-12-821391-9.00014-4.
- M. Cano-Sarabia, F. Aydin, L. Meng, M. Gil-Bonillo, J. Fonseca, M. Dietrich, S. Renner, H. Amenitsch, P. Falcaro, I. Imaz and D. Maspoch, Small, 2025, 21, e2407051 Search PubMed.
- P. B. Wang, J. Q. Li, T. Yang, J. W. Hu, M. Edeleva, L. Cardon and J. Zhang, Rapid Prototyp. J., 2025, 31, 492–504 CrossRef.
| This journal is © The Royal Society of Chemistry 2026 |