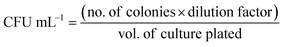A sustainable bioinspired nano-assembly of live marine bacteria for mineralization of phenothiazine dye
Neha
Redkar
 a,
Jyotsna
Mishra
a,
Jyotsna
Mishra
 b,
Rahul Kumar
Das
ac,
Dharmveer
Yadav
b,
Rahul Kumar
Das
ac,
Dharmveer
Yadav
 b,
Cathrine
Manohar
de,
Sumit
Saxena
b,
Cathrine
Manohar
de,
Sumit
Saxena
 abc and
Shobha
Shukla
abc and
Shobha
Shukla
 *abc
*abc
aNanostructures Engineering and Modelling Laboratory, Department of Metallurgical Engineering and Materials Science, Indian Institute of Technology Bombay, Mumbai, MH 400076, India. E-mail: sshukla@iitb.ac.in
bCentre for Research in Nanotechnology and Science, Indian Institute of Technology Bombay, Mumbai, MH 400076, India
cWater Innovation Centre: Technology, Research & Education (WICTRE), Indian Institute of Technology Bombay, Mumbai, MH 400076, India
dBiological Oceanography Department, CSIR-National Institute of Oceanography, Dona Paula, Goa 403004, India
eAcademy of Scientific and Innovative Research (AcSIR), Ghaziabad 201002, India
First published on 13th January 2026
Abstract
Bioinspired materials mimic the remarkable biological properties of natural systems, which promote microbial adherence and enhance the binding of pollutants, thereby improving the efficiency of water remediation processes. In this study, we report the assembly of a bio–nano interface using graphene oxide (GO) and live marine facultative anaerobic bacteria, Bacillus subtilis NAG1. Herein, we systematically investigate the interaction of NAG1 with a broad range of GO concentrations, ranging from 20 to 150 µg mL−1, using cell viability assays and further optimize the biocompatibility across a pH range of 4 to 14. Our findings indicate that GO, at 50 µg mL−1, showed biocompatibility and supported cell proliferation. This optimized GO–NAG1 nano-assembly was employed to efficiently degrade phenothiazine dyes – Azure A (Az-A) and Azure B (Az-B). Additionally, GO enhanced the production of ligninolytic enzymes including laccase (Lac), lignin peroxidase (LiP) and manganese peroxidase (MnP) leading to complete dye breakdown as confirmed by LC-MS analysis. Overall, our study provides strong evidence of the mineralization of phenothiazine dyes using a live anaerobic marine epiphytic bacterial system with GO.
1. Introduction
Nature provides an extensive repository of structural and functional blueprints that have emerged to sustain diverse biological systems.1 Designing biomimetic surfaces has emerged as an effective technique for developing innovative functional materials.2 These surfaces exhibit unique properties such as improved adhesion, multispectral response, hydrophobicity, and self-healing.3 Alteration of such surfaces facilitates microbial growth for various applications, such as pollutant degradation, oil–water separation, and water and air purification.4 Microbes, in particular bacteria, degrade organic pollutants via aerobic and anaerobic processes in which pollutants frequently bind to the bacterial surface via electrostatic interactions, following which enzymes such as laccase, azoreductase, and peroxidase assist in the breakdown of aromatic amines and azo linkages.5 Yet, as free-living organisms, they face limitations such as (1) cell wash-out and low stability, (2) the presence of high concentrations of pollutants limiting viability, (3) survival and cell proliferation, (4) competition with indigenous microbiota, and (5) fluctuating nutrients and deficiencies in the surroundings.6,7 This is in particular true for marine bacterial strains, which adjust to constantly changing habitats.Although biological methods have been widely employed, the existing technologies for complete dye degradation encounter significant operational and process-related limitations. Most physicochemical approaches including photocatalysis, adsorption, membrane filtration and coagulation–flocculation leave the pollutants isolated but not essentially degraded.8 In addition, these removal methods require high energy consumption, high cost and maintenance, and specific pH and temperature conditions and generate toxic secondary pollutants.9 To leverage complete degradation of toxic dyes and meet discharge standards of dye wastewater treatment, multistage or hybrid processes are implemented over conventional one-step removal methods.10,11 These hybrid methods take advantage of an initial step for pollutant enrichment by adsorbents followed by enhanced degradation by biological systems. To our advantage, using live cells rather than immobilized enzymes, can provide a self-sustaining platform for bioremediation. In contrast to passive systems, they adapt easily to external factors, maintain enzymatic activity, and aid in long-term pollutant degradation12 in an environmentally benign manner. Together with biomimetic surfaces, they form synergistic bio–nano interfaces that facilitate cell proliferation, mechanical anchorage, and a favorable microenvironment for enhanced bioremediation13
With the advancement of nanotechnology, biomimetic concepts are increasingly being used at the nanoscale to develop materials that interact with bacterial strains for environmental cleanup.14 Transition metal oxide-based nanomaterials such as zinc oxide, titanium oxide, and copper oxide offer promising remediation options but face limitations due to agglomeration, high production cost, poor scalability and dependency on specific conditions such as pH and UV exposure.15 However, carbon-based nanomaterials such as fullerenes, carbon nanotubes, quantum dots, graphene, and their derivatives have been extensively explored due to their high surface area, tunable physicochemical properties, biocompatibility, versatility, structural diversity, and low cost.16,17 Graphene oxide (GO), a two-dimensional oxidized form of graphene, is a suitable candidate for biomimetics due to its hydrophilicity, high surface area, surface functional groups (hydroxyl, epoxy, and carboxyl groups), and ease of dispersion in aqueous environments.18
Although well reported as an antibacterial material, studies reveal that GO nanosheets serve as an excellent nanoarchitecture for biomimetic scaffolds. Their interaction with cells is influenced by surface area, concentration, sheet size, pH, degree of defects, and the number of functional groups.19,20 GO has been used to enhance the growth of Escherichia coli by promoting cell attachment and biofilm formation.21 Typically, bacterial surfaces contain a high content of hydrophobic amino acids, main constituents of outer membrane proteins and lipopolysaccharides, which influence adhesiveness.22 The flat surface-functionalized GO and bacteria mediate attachment to hydrophobic surfaces via hydrophobic interactions, whereas the edge-tethered GO sheets weaken the adhesion forces.23 Xu and group reported that the biocompatibility of GO with live Shewanella depends on the degree of oxidation, carbonyl groups, defects, and sheet size, which are responsible for hydrophobic attraction, electrostatic repulsion, and adhesion force.24 On the other hand, GO is also reported as a bactericidal agent at high concentrations, causing disruption of the cell membrane by acting as “nano knives”, and through a “wrapping or trapping” effect causing oxidative stress and generation of reactive oxygen species (ROS) leading to cell lysis.25,26 It is therefore critical to develop an effective biomimetic scaffold before its application for bioremediation.
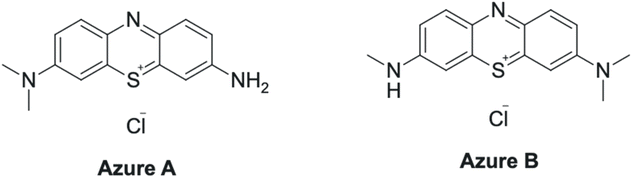
In this study, we investigated the use of live marine epiphytic facultative anaerobic bacteria (a macroalgae-associated strain isolated from mangrove sediments), NAG1, belonging to Bacillus sp. with GO. Systematic studies of the interaction of NAG1 with a broad range of GO concentrations, ranging from 20 to 150 µg mL−1, using cell viability assays, and further optimization of biocompatibility across a range of pH from acidic to alkaline (4 to 14) were performed. The optimum GO–NAG1 nano-assembly was used for the degradation of phenothiazine dyes – Azure A (Az-A) and Azure B (Az-B). Phenothiazine dyes are recalcitrant and have a rigid, planar tricyclic aromatic structure with inherent cationic charge and redox stability, making them widely used in textile industries and biomedical applications.27 As shown in Scheme 1, Az-A and Az-B are derived from oxidative demethylation of methylene blue, and are redox-active, highly stable, and intrinsically toxic due to their intercalation into DNA and partitioning into the lipid membrane of living cells.28 According to studies, they possess a heteroatomic tricyclic backbone that contains sulfur and nitrogen, which gives them resistance to mineralization.29 Our study provides strong evidence of mineralization of phenothiazine dyes using a live anaerobic marine epiphytic bacterial system with GO.
2. Experimental section
2.1. Bacterial strain and culture conditions
NAG1, an epiphytic facultative anaerobic bacterial strain belonging to Bacillus sp., (accession number: MW487389) was provided by CSIR-National Institute of Oceanography (NIO), Goa, India. This strain was isolated from red macroalgae (Gracilaria corticata) found in the marine environment of Dona Paula, Goa (15.4547°N, 73.8045°E). NAG1 was routinely subcultured on Zobell Marine broth (HiMedia, India) supplemented with 1.5% agar powder (bacteriological grade, HiMedia, India). For experiments, a single colony was inoculated in 30 mL Boyd & Kohlmeyer (BK) broth, denoted as BK (2% glucose, 0.1% yeast extract, and 0.2% peptone) and incubated in a shaker incubator at 37 °C (pH 7), 120 rpm. The cells were harvested in the mid-exponential growth phase (OD600 = 0.3–0.4).2.2. Synthesis of GO
An improved Hummer's method was used to synthesize GO nanosheets by chemical exfoliation of graphite powder.30 The synthesized GO suspension solution was washed thoroughly using distilled water until a pH value of 7 was achieved and then sonicated in an ultrasonication bath at 15 kHz frequency (Vibronics Ultrasonic cleaner, 250 W) for 30 minutes (without heating).2.3. Bacterial cell compatibility with GO nanosheets
An aliquot of 1% (v/v) of the total volume of bacterial culture (106 CFU mL−1) was incubated with GO nanosheets at varying concentrations (20 to 150 µg mL−1) at 37 °C with agitation (120 rpm). These samples were denoted as G20, G40, G60, G80, G100, and G150 in further experiments. A freshly prepared BK broth was used in all experiments. OD600 was obtained after every 3 h using a UV-vis spectrophotometer (Shimadzu UV 2600) until the growth reached a stationary phase. The uninoculated media were maintained as a control. Furthermore, NAG1 cell growth was studied over 12 h of incubation across a broad pH spectrum, including pH 4, 7, 11, and 14.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000, v/v) and PI (1
1000, v/v) and PI (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000, v/v) were added. After 20 minutes of incubation at 37 °C, cells were centrifuged at 5000 rpm and washed several times with distilled water. The calcein fluorescence (λex/em = 488/528 nm) and PI fluorescence (λex/em = 561/647 nm) were measured. The cells were visualized using a confocal laser scanning microscope (LSM 780; Carl Zeiss, CZ microscopy, Germany).
1000, v/v) were added. After 20 minutes of incubation at 37 °C, cells were centrifuged at 5000 rpm and washed several times with distilled water. The calcein fluorescence (λex/em = 488/528 nm) and PI fluorescence (λex/em = 561/647 nm) were measured. The cells were visualized using a confocal laser scanning microscope (LSM 780; Carl Zeiss, CZ microscopy, Germany).
2.4. Biodegradation of Az-A and Az-B dye
NAG1 cells were grown in G50 for 12 h, harvested, centrifuged, and washed with water. A working stock of 100 ppm Az-A and Az-B dyes was prepared and filter-sterilized using a 0.22 µm sterile filter. For dye degradation studies, 1% (v/v) bacterial suspension was added as inoculum to BK media supplemented with 50 ppm as the final concentration of each dye and incubated at 37 °C under agitation (120 rpm). Uninoculated dye was maintained as a control. Absorbance was recorded at 620 and 650 nm for Az-A and Az-B dyes, respectively, for 24 h. All experiments were carried out in triplicate, and the removal efficiency was calculated using the following formula:| Dye removal rate (%) = ((Co − Ce)/Co) × 100 |
Furthermore, for the identification of dye degradation products, LC-MS was performed in ESI ionization mode, using a HypersilGold C18 100 × 2.1 mm, 3 µm column at a Fragmentor voltage of 175.0 V.
2.5. Screening of ligninolytic enzymes for dye degradation
2.6. Characterization
X-ray diffraction (XRD) patterns were obtained using a PANalyticalXpert-Pro equipped with a Cu target source operating at 45 kV & 40 mA. The functional groups at the surface were investigated using Fourier transform infrared spectroscopy (FT-IR; Bruker 3000 Hyperion). Raman spectra (Reinshaw, Invia Reflex) were recorded at room temperature using a 100× objective under laser excitation at a wavelength of 532 nm. X-ray photoelectron spectroscopy (XPS) was performed using a Kratos AXIS Supra. The morphology and size were analysed using field emission gun-based high resolution transmission electron microscopy (HR-TEM; FEI Tecnai G2, F30 300 kV) and environmental scanning electron microscopy (ESEM; FEI QUANTA 200). The surface topography was mapped using an atomic force microscope (AFM; MFP-3D BIO Asylum Research, USA).2.7. Statistics
All the experiments were performed in triplicate, and data were represented as mean ± standard deviation. The statistical analysis of differences was carried out using a two-way ANOVA test and a t-test. Data with P < 0.05 was considered to be significant.3. Results and discussion
3.1. Characterization of GO nanosheets
GO was synthesized using the improved method. As seen in Fig. S1a, the TEM micrograph showed large sheets with an increase in transparent regions, indicating few-layered GO. A selected area electron diffraction (SAED) pattern (Fig. S1b) revealed consistent, clear diffraction spots in a hexagonal lattice, with interplanar spacings of 2.3 Å, 1.4 Å, 1.1 Å, and 0.97 Å. The SEM micrograph (Fig. S1c) displayed smooth surfaces consisting of one or more layers with wrinkles at the edges of the sheet, while AFM analysis (Fig. S2a) revealed that the sheet contained multiple folds and soft wrinkles.34 The roughness profile and three-dimensional topography plot are shown in Fig. S2b and c. The average roughness and maximum height of the GO nanosheet were 1.5 nm and 13.1 nm, respectively, as illustrated in Table ST1.The UV-visible spectrum of GO showed a maximum absorption peak at 226 nm (π–π* transition of the aromatic C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds), and a weak shoulder at 292 nm (n–π* transition of C
C bonds), and a weak shoulder at 292 nm (n–π* transition of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bonds) (Fig. S3a). The XRD spectrum revealed a prominent diffraction peak at 2θ = 10.23° corresponding to the (001) plane of GO, suggesting complete oxidation and exfoliation (Fig. S3b). An increased interlayer d-spacing, or basal spacing, of 0.87 nm was observed, due to van der Waals force interactions between GO layers, and aligns with the reported values.35 The FTIR spectrum (Fig. S3c) confirmed the presence of functional groups in GO, with signature peaks at 3395 cm−1 (–OH), 1647 cm−1 (C
O bonds) (Fig. S3a). The XRD spectrum revealed a prominent diffraction peak at 2θ = 10.23° corresponding to the (001) plane of GO, suggesting complete oxidation and exfoliation (Fig. S3b). An increased interlayer d-spacing, or basal spacing, of 0.87 nm was observed, due to van der Waals force interactions between GO layers, and aligns with the reported values.35 The FTIR spectrum (Fig. S3c) confirmed the presence of functional groups in GO, with signature peaks at 3395 cm−1 (–OH), 1647 cm−1 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 1221 and 1077 cm−1 (C–OH and C–O–C), and 2934/2859 cm−1 for CH2 stretching.36 The Raman spectrum (Fig. S3d) showed the D band at 1350 cm−1 (defects), G band at 1596 cm−1 (sp2 carbon), and 2D band at 2695 cm−1 (layered structure). The ID/IG ratio of 1.00 indicated a high degree of disorder due to oxidation and exfoliation. XPS analysis was used to probe valence states of carbon and oxygen in GO. The deconvoluted XPS spectra of C1s (Fig. S3e) revealed three peaks with binding energies of 284.6 eV (graphitic C–C bonds), 286.8 eV (C–O from epoxy and hydroxyl), and 287.9 eV (C
C), 1221 and 1077 cm−1 (C–OH and C–O–C), and 2934/2859 cm−1 for CH2 stretching.36 The Raman spectrum (Fig. S3d) showed the D band at 1350 cm−1 (defects), G band at 1596 cm−1 (sp2 carbon), and 2D band at 2695 cm−1 (layered structure). The ID/IG ratio of 1.00 indicated a high degree of disorder due to oxidation and exfoliation. XPS analysis was used to probe valence states of carbon and oxygen in GO. The deconvoluted XPS spectra of C1s (Fig. S3e) revealed three peaks with binding energies of 284.6 eV (graphitic C–C bonds), 286.8 eV (C–O from epoxy and hydroxyl), and 287.9 eV (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). The O1s spectrum (Fig. S3f) displayed peaks at 531.2 eV (C
O). The O1s spectrum (Fig. S3f) displayed peaks at 531.2 eV (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 532.5 eV (–COOH), and 533.1 eV (–OH).37 The survey spectrum of GO is provided in Fig. S3g.
O), 532.5 eV (–COOH), and 533.1 eV (–OH).37 The survey spectrum of GO is provided in Fig. S3g.
3.2. Bacterial cell viability and optimization of GO–NAG1 nano-assembly
The synthesized GO was incubated with the marine bacterial strain NAG1 belonging to Bacillus sp. to evaluate the biocompatibility at optimal concentration. Fig. 1a illustrates the current work in which NAG1 cells were incubated with different concentrations of GO ranging from 20 to 150 µg mL−1 and optical density (OD) was recorded at 600 nm as presented in Fig. S4. After 12 h of incubation, the media flasks supplemented with GO (G20 to G60) showed turbidity due to an increase in cell density; however, at higher concentrations (G80 to G150), visible growth interference was observed due to dark coloration of GO (Fig. S4a and b). The growth curve of GO-incubated cells demonstrated prominent lag, log, and stationary phases. Cells incubated with G20–G60 showed a similar growth pattern as compared to untreated NAG1 cells (control); however, at 80–150 µg mL−1, GO displayed an inhibitory effect with a decrease in cell density as shown in Fig. S4c. | ||
Fig. 1 Bacterial cell viability at varied GO concentrations. (a) Schematic of the present study, (b) bar graph showing cell viability expressed as colony forming units per mL (CFU mL−1) and ROS generation using the DCFH-DA assay expressed as relative fluorescence units, (c) images of bacterial colonies plated on NA after 12 h of exposure, (d) optimization of GO–NAG1 on 40, 50 and 60 µg mL−1 at pH 4, 7, 11, and 14, (e) % live cells using the live/dead cell assay and (f) ESEM micrograph of G50–NAG1 (Mag: 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 00×; scale bar: 10 µm). Pseudo-coloring of SEM images was done using GIMP software. The original micrograph is provided in Fig. 2d (data were analyzed by one and two-way ANOVA; at all-time points, the results were significant in comparison with the control, p < 0.05. Statistical data are provided in Fig. S5 and Table ST2). 00×; scale bar: 10 µm). Pseudo-coloring of SEM images was done using GIMP software. The original micrograph is provided in Fig. 2d (data were analyzed by one and two-way ANOVA; at all-time points, the results were significant in comparison with the control, p < 0.05. Statistical data are provided in Fig. S5 and Table ST2). | ||
The bacterial viability was assessed by plating each treated sample on NA plates, and the total viable count was represented as CFU mL−1. As shown in Fig. 1b (green bar graph), the CFU mL−1 count increased from 2.7 × 107 (G20) to 4.8 × 107 (G60), indicating that cell viability was preserved at lower concentrations. In contrast, the CFU mL−1 count declined from 2.8 × 107 (G80) to 2 × 106 (G150), indicating suppression of growth due to the presence of a high concentration of GO. Under shaking conditions, the oxygenated functional groups at the basal planes of GO help adsorb nutrients from the supplemented media, making them easily accessible for metabolic processes at lower concentrations (G20 to G60). Additionally, bacterial cells were able to adhere to the nanosheet by secreting extracellular polymeric substances and proliferate.38 As shown in Fig. 1c, the NA agar plate showed isolated dense colonies (G20–G60); however, the number of colonies decreased, indicating a decrease in cell viability. The agar plate of untreated NAG1 cells is displayed in Fig. S4d.
To better understand the mechanism underlying the observed growth patterns, intracellular ROS levels were measured using the DCFH-DA assay. DCFH-DA is a general oxidative species indicator sensitive to a wide range of reactive oxygen species, to evaluate ROS production.39 Our findings demonstrated that ROS generation increased with GO concentration (red bar graph), and this increase was inversely proportional to the bacterial cell viability (green bar graph) as shown in Fig. 1b. At G20 and G40, a gradual rise in fluorescence, indicating minor oxidative stress was observed. This suggests that low concentrations of GO might function as a stress-priming agent without causing damage25 and therefore did not inhibit cell viability and metabolic activity. However, at G60–G100, ROS levels increased significantly by approximately three to six-fold (compared to the control), indicating that GO induces oxidative stress. The oxidative stress mainly occurs due to the formation of reactive species such as superoxide ions (O2−), hydroxyl ions (OH−), and hydrogen peroxide (H2O2), which damage the cellular components such as lipid, protein, and nucleic acids.40 When there is a rise in ROS levels, cell membrane integrity is damaged, triggering cytoplasmic leakage and causing cell death,41 which correlates with reduced cell colonies and cell viability, as observed in our study. The same was further validated by the fluorescence spectra at 522 nm (Fig. S6a) and LSM images of GO-incubated cells (Fig. S6b) which showed an increase in signal intensity confirming the presence of reactive species.
Furthermore, NAG1 cells were incubated with different concentrations of GO that supported viability (G40, G50, and G60), and were cultured under pH conditions including: 4, 7, 11, and 14 (Fig. 1d). A broad pH range was selected, considering the ability of marine bacteria to adapt to extreme environmental conditions for metabolic activity.40 The bacterial growth was varied under different pH conditions, for instance, under acidic conditions (pH 4), no cell growth was observed. Under acidic conditions, GO becomes protonated and less hydrophilic, resulting in larger aggregates (Fig. S7a). This is due to stronger π–π interactions and less electrostatic repulsion.42 Additionally, acidic pH also suppresses key metabolic processes, making cell survival more difficult43 (Fig. S7a and e). Under neutral conditions (pH 7), the CFU mL−1 value increased to 2.8 × 106 (G40), 3.8 × 106 (G50), and 2.5 × 106 (G60), indicating cell viability. At pH 7, GO shows excellent dispersibility, and –COOH groups are deprotonated to –COO− ions, creating a more hydrophilic surface. These charged surfaces also enhance the adsorption of nutrients from the media onto cells, thereby supporting metabolic activity and cell growth44,45(Fig. S7b and f). Under alkaline conditions (pH 11 and 14), the CFU mL−1 value further decreased by approximately two to three-fold, respectively (Fig. S7c, d and g, h). Under such conditions, GO undergoes deprotonation, which increases electronegativity46 and reduces nutrient adsorption. However under extreme alkaline conditions, GO aggregates and settles down, entrapping cells and reducing their viability.47 Overall, NAG1 cells exhibit the highest CFU mL−1 value across pH conditions of 7, 11, and 14. Maximum cell activity was exhibited in the presence of G50 at pH 7; this optimized combination was used for subsequent experiments.
The live and dead bacterial staining assay of G50+NAG1 was performed to further confirm the CFU mL−1 method. The viability of the G50+NAG1 cells was 91.8% compared to the control (untreated NAG1) (Fig. 1e). The LSM images of the assay are provided in Fig. S8, indicating calcein-AM (green) stained live cells. A representative ESEM image depicting viable NAG1 cells beneath the GO nanosheet (G50) is shown in Fig. 1f.
3.3. Spectroscopic analysis of the interaction of GO with NAG1 cells
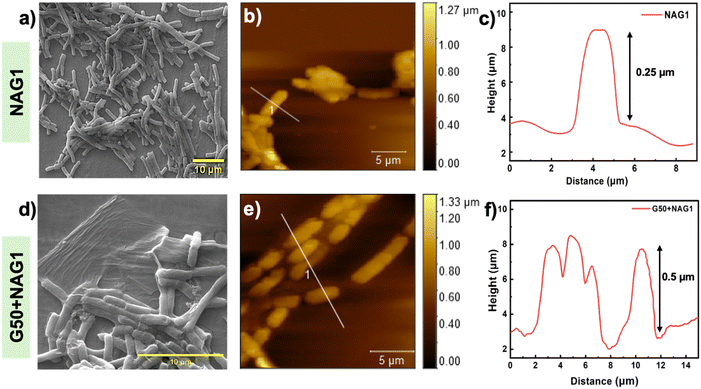
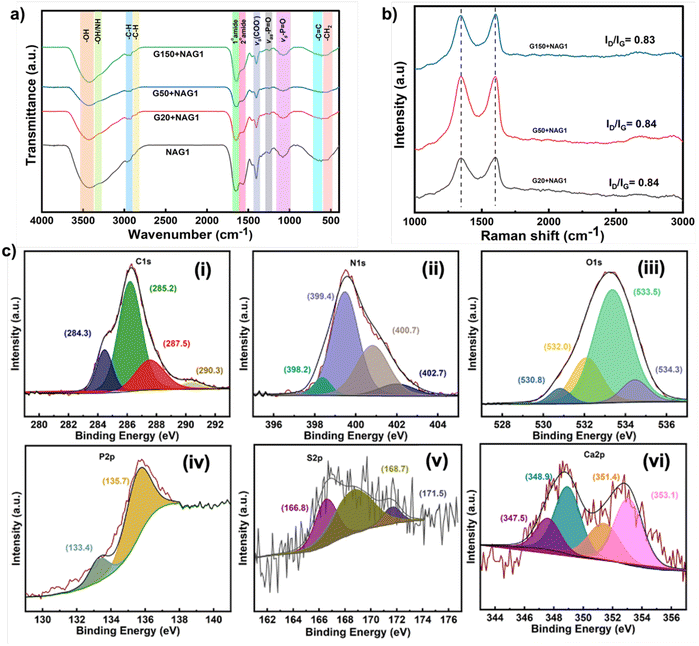
The ESEM micrograph showed that G50 formed increased wrinkles, which acted as a scaffold, resulting in extensive valley formation, allowing cells to adhere, proliferate and aggregate (Fig. 2d). The enclosed cells showed no mechanical damage due to the smooth edges of GO nanosheets and retained the same rod-like structure as the untreated NAG1 (Fig. 2a). The AFM analysis displayed surface topography and mechanical interaction between G50 and NAG1. The GO nanosheets in G50+NAG1, being a few nanometers thin, covered the cells uniformly, causing a reduction in the roughness. The cells showed a length of 4.8 µm and a height of 0.25 µm with a diameter of approximately 2.2 µm (Fig. 2b and c), whereas the untreated NAG1 was 5.5 µm in length and 0.5 µm in height with a diameter of approximately 2 µm (Fig. 2e and f). It was observed that the difference in height between the cells was approximately 250 nm. The detailed parameters of roughness and height are provided in Table ST3. At a higher concentration of GO, NAG1 cells were engulfed, resulting in a 3-fold reduction in roughness (Fig. S9a–c). The ESEM images of higher concentrations of GO (G100 and G150) showed irregular cells trapped under wrinkled nanosheets (Fig. S10).To examine the surface chemistry and functional groups of NAG1, FTIR analysis was performed (Fig. 3a). The G50+NAG1 spectrum was compared to those of untreated and treated NAG1 with the least (G20) and highest (G150) GO concentrations to gain a better understanding of the interactions. The untreated NAG1 a showed a broad peak at 3419 cm−1 (OH/NH stretching from sugars and surface proteins), 2964 cm−1 (C–H stretching from proteins, lipids, and carbohydrates), and 1647 cm−1 (amide I and carbonyl stretching from phospholipids). The peaks at 1235 cm−1 and 617 cm−1, respectively, represent C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bending in alkenes and PO2− vibrations in phospholipids of the cell wall.41
C bending in alkenes and PO2− vibrations in phospholipids of the cell wall.41
After 12 h of incubation with GO, the band intensities of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and N–H decreased, indicating reduced protein and amide content. In G20+NAG1 and G50+NAG1, a weak peak shift at 1248 cm−1 (P
O and N–H decreased, indicating reduced protein and amide content. In G20+NAG1 and G50+NAG1, a weak peak shift at 1248 cm−1 (P![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O asymmetric stretch) and reduced intensity at 1060 cm−1 (PO2− symmetric stretch) suggested nucleic acid and phospholipid interactions with GO. The peaks at 1649 and 1566 cm−1 (amide I and II) were weaker in GO-incubated cells, indicating structural changes. The intensity drop at 2938 cm−1 and 2873–2880 cm−1 reflected alterations in –CH2– groups and lipopolysaccharide/peptidoglycan structures. A notable decrease at 1409 cm−1 suggested C–H deformation and C–O stretch from deprotonated carboxylate groups, pointing to cell membrane modification by GO.48 The Raman spectra of NAG1 incubated with G20, G50, and G150 showed ID/IG ratios of 0.84, 0.84, and 0.83, respectively (Fig. 3b). This suggested that minimal defects were caused by NAG1 across different concentrations. In the case of G150+NAG1, the ID/IG was found to be 0.83, indicating the possibility of partial bacterial reduction of GO or restoration of graphitic domains.49
O asymmetric stretch) and reduced intensity at 1060 cm−1 (PO2− symmetric stretch) suggested nucleic acid and phospholipid interactions with GO. The peaks at 1649 and 1566 cm−1 (amide I and II) were weaker in GO-incubated cells, indicating structural changes. The intensity drop at 2938 cm−1 and 2873–2880 cm−1 reflected alterations in –CH2– groups and lipopolysaccharide/peptidoglycan structures. A notable decrease at 1409 cm−1 suggested C–H deformation and C–O stretch from deprotonated carboxylate groups, pointing to cell membrane modification by GO.48 The Raman spectra of NAG1 incubated with G20, G50, and G150 showed ID/IG ratios of 0.84, 0.84, and 0.83, respectively (Fig. 3b). This suggested that minimal defects were caused by NAG1 across different concentrations. In the case of G150+NAG1, the ID/IG was found to be 0.83, indicating the possibility of partial bacterial reduction of GO or restoration of graphitic domains.49
The chemical interactions between NAG1 and GO were investigated using XPS analysis (Fig. 3c). The high-resolution scans of C1s, N1s, O1s, P2p, S2p, and Ca2p were compared between untreated NAG1 (Fig. S11) and G50+NAG1. The wide spectra of NAG1 and G50+NAG1 validated the presence of key elements (Fig. S12). The C1s region of G50+NAG1 showed varying relative intensities of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C sp2 (284.5 eV), C–N/C–O sp3 (285.6 eV), C–O (286.2 eV), C
C sp2 (284.5 eV), C–N/C–O sp3 (285.6 eV), C–O (286.2 eV), C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (287.3 eV), and O–C
O (287.3 eV), and O–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (290.3 eV) (Fig. 3c-i). The changes in the C
O (290.3 eV) (Fig. 3c-i). The changes in the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C
C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, and O–C
O, and O–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O peaks point to intermolecular hydrogen bonding through aldehyde and carboxylic groups as well as π–π interactions. The sp3 carbon peak represents functional groups found in biomolecules such as N-acetylglucosamine, N-acetylmuramic acid, and amino acids.50 The N1s peaks at 400.8, 398.4, and 399.4 eV denote amide, imine, and –NH2 groups. The presence of meso-diaminopimelic acid and peptide secondary amines is suggested by changes in amine and amide ratios following G50 incubation (Fig. 3c-ii).
O peaks point to intermolecular hydrogen bonding through aldehyde and carboxylic groups as well as π–π interactions. The sp3 carbon peak represents functional groups found in biomolecules such as N-acetylglucosamine, N-acetylmuramic acid, and amino acids.50 The N1s peaks at 400.8, 398.4, and 399.4 eV denote amide, imine, and –NH2 groups. The presence of meso-diaminopimelic acid and peptide secondary amines is suggested by changes in amine and amide ratios following G50 incubation (Fig. 3c-ii).
The O1s spectra show that peaks at 531.4 (O–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 532.5 (C
O), 532.5 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 533.5 (C–O–H), and 534 eV (–O–C–O–C– from the C1–C6 sugar backbone ether linkage) were present on the NAG1 surface, which after binding with G50 showed major changes in aldehyde- and hydroxyl-group oxygen51 (Fig. 3c-iii). A strong satellite peak at 135.7 eV following GO binding was observed in the P2p signal, which is ascribed to surface phospholipids and indicates 2p1/2 participation52 (Fig. 3c-iv). The Ca2p peaks shifted from 351.1/347.3 eV (2p3/2/2p1/2) to 353.1/348.9 eV, indicating that exposure to G50 altered the oxidation state of calcium (Fig. 3c-vi). Overall, the XPS analysis supports the conclusions derived from FTIR (–OH, –NH,2 amide, and C–O) and Raman (change in the sp3/sp2 nature of carbon) analysis.
O), 533.5 (C–O–H), and 534 eV (–O–C–O–C– from the C1–C6 sugar backbone ether linkage) were present on the NAG1 surface, which after binding with G50 showed major changes in aldehyde- and hydroxyl-group oxygen51 (Fig. 3c-iii). A strong satellite peak at 135.7 eV following GO binding was observed in the P2p signal, which is ascribed to surface phospholipids and indicates 2p1/2 participation52 (Fig. 3c-iv). The Ca2p peaks shifted from 351.1/347.3 eV (2p3/2/2p1/2) to 353.1/348.9 eV, indicating that exposure to G50 altered the oxidation state of calcium (Fig. 3c-vi). Overall, the XPS analysis supports the conclusions derived from FTIR (–OH, –NH,2 amide, and C–O) and Raman (change in the sp3/sp2 nature of carbon) analysis.
3.4. Evaluation of Az-A and Az-B dye removal using the GO–NAG1 nano-assembly
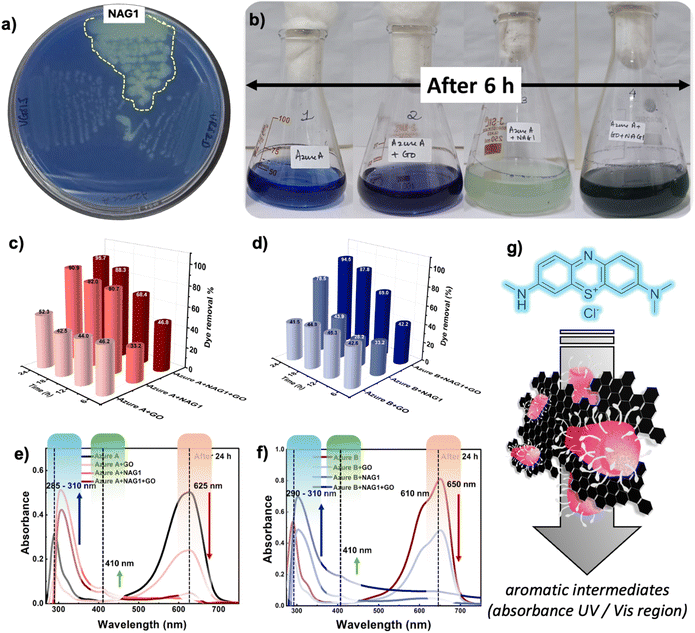
As a qualitative assay, a loopful of NAG1 cells was inoculated onto an Az-A dye-based agar plate, resulting in a visible decolorization zone after 24 h of incubation (Fig. 4a). A final concentration of 50 ppm of Az-A and Az-B dye was treated with GO, NAG1, and GO+NAG1 nano-assembly for 24 h at 120 rpm (under shaking conditions) (Fig. 4b). The concentration of GO used was 50 µg mL−1, which was optimized in the previous section. The absorbance was recorded at 625 and 650 nm for Az-A and Az-B, respectively, at 6, 12, 18, and 24 h.It is noteworthy that the GO+NAG1 nano-assembly outperformed the Azure dyes treated individually with GO and NAG1. Specifically, the GO–NAG1 nano-assembly achieved 95.7% and 94.5% removal efficiencies of Az-A and Az-B, respectively, within 24 h (Fig. 4c and d). In the initial 6 h, GO+NAG1 and GO exhibited a removal rate of 46.8% (Az-A) and 42.6% (Az-B), indicating that the removal was primarily due to adsorption by GO. Az-A and Az-B are cationic dyes that create a strong electrostatic interaction between the N+H group (positive dipole from dye molecules) and oxygen-containing functional groups present on the GO surface. Moreover, Az-A and Az-B contain aromatic thiazine ring structures that interact with the graphitic domains of GO through π–π stacking to form stable adsorption.53 In the case of only NAG1, the dark blue color gradually changed to viridescent (Fig. 4b) within the initial 6 h, indicating the onset of biodegradation. After complete incubation, higher dye removal % was exhibited, 90.9% (Az-A) and 78.5% (Az-B). Notably, the supernatant of the treated dye was clear and transparent, and the absence of coloration in bacterial cells indicates biodegradation of dye rather than cellular adsorption (Fig. S13a).
Both dyes revealed a decrease in characteristic adsorption peaks in the UV-visible absorption spectra (Fig. 4e and f). Following, a 24 h treatment by utilizing NAG1–GO and NAG1, the deep blue color of dye faded, indicating a significant decrease in the peak at 625 nm in Az-A, which corresponds to the π → π* transition of the chromophore area. Similarly, the peak at 650 nm in Az-B, indicative of π–π* transitions in the phenothiazine ring, along with a shoulder peak at 610 nm (monomer–dimer transition), disappeared entirely in GO–NAG1.54 Additionally, a new peak appeared at the absorption wavelengths of about 290 and 400 nm in both NAG1 and GO+NAG1 treated dyes. This might be the adsorption peaks produced by the metabolites formed after biodegradation of dyes,55 which supported our hypothesis. Table 1 shows a comparative report of dye degradation efficiencies exhibited mainly by GO and live bacteria-based nano-assembly. Previous studies have attributed synergistic adsorption and degradation of textile dyes and effluents but require longer incubation time.20,56–58 In many such systems, GO undergoes reduction to rGO,59,60 which is toxic to cells.25 In comparison, the GO–NAG1 nano-assembly reported in this work, achieved efficient degradation efficiency within 24 h with no reduction of GO.
| Graphene and its derivatives | Bacterial strain and source | Target pollutant | Conc. (µg mL−1) | Degradation efficiency (%) | Degradation time (h) | Ref. |
|---|---|---|---|---|---|---|
| GO-based porous hydrogel | Shewanella xiamenensis BC01 (facultative wild-type) | Congo Red | 50 | 99.00 | 20 | 56 |
| rGO-based porous hydrogel | Shewanella xiamenensis BC01 (facultative wild-type) | Congo Red | 100 | 99.80 | 54 | 59 |
| Methylene Blue | 100 | 95.90 | 54 | |||
| Graphene-based porous hydrogel | Shewanella putrefaciens CN32 (facultative wild-type) | Methyl Orange | 20 | 80.60 | 24 | 60 |
| Methylene Blue | 20 | 81.50 | 24 | |||
| GO–gelatin–polyacrylic acid (GO–G–PAA) | Bacterial consortium of Dietzia sp., Bacillus sp. and Pseudomonas mendocina (water bodies near the textile dyeing unit) | Textile effluents | — | 99.47 | 12 | 57 |
| Graphene oxide–calcium alginate hydrogel beads (KG–GO–CA) | Klebsiella grimontii (isolated from soil contaminated with textile effluents) | Acid Blue 113 | 50 | 94.60 | 360 | 58 |
| GO nanosheets | Marine Bacillus sp. NAG1 (mangrove sediments) | Methylene Blue | 50 | 34.70 | 24 | 20 |
| GO nanosheets | Marine Bacillus sp. NAG1 (mangrove sediments) | Azure A | 50 | 95.7 | 24 | This study |
| Azure B | 50 | 94.5 | 24 |
3.5. Synergistic adsorption–biodegradation of dyes by GO–NAG1 nano-assembly
The untreated Az-A showed a peak at 3433 cm−1 that can be attributed to the N–H stretching vibration of the primary amine group. Bands at 2954, 2923, and 2850 cm−1 arise due to C–H bond stretching vibrations, while the vibrations corresponding to 1647 cm−1 can be attributed to stretching vibrations of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C of the aromatic ring (Fig. 5a). The peaks at 1461 and 1407 cm−1 correspond to the asymmetrical and symmetrical bending of N–Me1 and N–Me2 stretches. The two peaks at 1346 and 1240 cm−1 are attributed to in-plane C–H bending vibrations for aromatic rings. The lower frequency bands between 690 and 850 cm−1 correspond to C–S stretching vibrations of the phenothiazine ring.61 Upon Az-A dye degradation, new peaks were observed. In the case of GO+NAG1 and NAG1, peaks at 3304 and 3132 cm−1 can be associated with the asymmetric and symmetric stretching vibrations of N–H bonds, particularly in the aromatic amine/amide region, indicating the breakdown of azo bonds (–N
C of the aromatic ring (Fig. 5a). The peaks at 1461 and 1407 cm−1 correspond to the asymmetrical and symmetrical bending of N–Me1 and N–Me2 stretches. The two peaks at 1346 and 1240 cm−1 are attributed to in-plane C–H bending vibrations for aromatic rings. The lower frequency bands between 690 and 850 cm−1 correspond to C–S stretching vibrations of the phenothiazine ring.61 Upon Az-A dye degradation, new peaks were observed. In the case of GO+NAG1 and NAG1, peaks at 3304 and 3132 cm−1 can be associated with the asymmetric and symmetric stretching vibrations of N–H bonds, particularly in the aromatic amine/amide region, indicating the breakdown of azo bonds (–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–). The peaks at 2923 and 2848 cm−1 are attributed to –CH2– stretching vibrations. In addition, the presence of a new peak and a peak shift from 1577 to 1560 cm−1, along with a prominent peak at 1398 cm−1 is attributed to aromatic/azo stretching due to modification of aromatic structures by enzymatic attack. The distinct peaks in the 1186 to 969 cm−1 region correspond to C–N stretching, S
N–). The peaks at 2923 and 2848 cm−1 are attributed to –CH2– stretching vibrations. In addition, the presence of a new peak and a peak shift from 1577 to 1560 cm−1, along with a prominent peak at 1398 cm−1 is attributed to aromatic/azo stretching due to modification of aromatic structures by enzymatic attack. The distinct peaks in the 1186 to 969 cm−1 region correspond to C–N stretching, S![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O vibrations, and C–O–C linkages, indicating probable breakdown of thiazine rings.
O vibrations, and C–O–C linkages, indicating probable breakdown of thiazine rings.
The FTIR spectra of untreated Az-B (Fig. 5b) showed a peak at 3410 cm−1 attributed to N–H stretching vibration due to the presence of a primary amine group. Broadening in the 3500–3200 cm−1 region is observed due to –OH stretching, showing the presence of water molecules in the sample. The peaks at 2925 cm−1 and 2850 cm−1 correspond to the asymmetric and symmetric C–H stretching vibrations due to the presence of aromatic and methyl groups in the dimethylamino substituent of Az-B. Furthermore, a sharp peak at 1650 cm−1 could arise from C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching in the aromatic ring or N–H bending vibration. The peaks at 1465 cm−1 and 1407 cm−1 indicate C–H stretching vibrations of the aromatic ring, whereas the peak at 1037 cm−1 corresponds to the –C–OH stretching vibrations. The peak at 1238 cm−1 is attributed to C–N stretching associated with aromatic amine and heterocyclic nitrogen compounds. The weak C–S stretching observed at 626 cm−1 indicates the aromatic framework of Az-B dye.54 A significant increase in the relative intensity at 1650 cm−1, which corresponds to C
O stretching in the aromatic ring or N–H bending vibration. The peaks at 1465 cm−1 and 1407 cm−1 indicate C–H stretching vibrations of the aromatic ring, whereas the peak at 1037 cm−1 corresponds to the –C–OH stretching vibrations. The peak at 1238 cm−1 is attributed to C–N stretching associated with aromatic amine and heterocyclic nitrogen compounds. The weak C–S stretching observed at 626 cm−1 indicates the aromatic framework of Az-B dye.54 A significant increase in the relative intensity at 1650 cm−1, which corresponds to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (aromatic ring)/C
O (aromatic ring)/C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds (carbonyl group) present in GO, was observed in GO-treated Az-B, which is attributed to the interaction of GO with the Az-B dye molecule.
C bonds (carbonyl group) present in GO, was observed in GO-treated Az-B, which is attributed to the interaction of GO with the Az-B dye molecule.
Upon 24 h treatment of Az-B dye with NAG1 and GO–NAG1, a minor shift and increase in relative intensity of the peak at 1400 cm−1 were observed, indicating deformation in the C–H group from the methyl group or C–N bending vibration, suggesting partial breakdown of aromatic ring structures into –COO− derivatives during enzymatic attack. The appearance of a peak at 1562 cm−1 in NAG1 and GO+NAG1 indicates N–H bending of the amide II band from bacterial enzymes and suggests partial breakdown of aromatic fragments. The peak at 3301 cm−1 indicates O–H and N–H stretching vibrations due to hydroxylated or amine-containing degraded products. Additionally, the peak at 3118 cm−1 is attributed to aromatic C–H stretching, indicating bacterial protein or enzymatic action.
3.6. Probable degradation mechanism of phenothiazine dyes
To investigate the degradation mechanism of Azure dyes by GO–NAG1 nano-assembly, LC-MS was performed to identify the various degradation intermediates. The results revealed that distinct peaks at retention time of 5.6, 4.7, 2.4, 1.9, and 1.7 minutes for the intermediates and by-products of Azure dyes after treatment (Fig. 6a and b). At 14.5 minutes, m/z 270 (C15H16ClN3S) was found to be higher in abundance in the untreated dye sample. In the first pathway (Fig. 6c), m/z 284 (C16H18N3S) is converted into m/z 274 (C14H16N3OS), which indicates oxidation of thioester (sulfide) to sulfoxide.62 Furthermore, the distinct by-products with m/z 155 at 2.4 minutes, m/z 137 at 1.9 minutes, and m/z 187 at 1.7 minutes denote C7H7O2S+, C9H12N2O, and C7H9NO3S, respectively (Table ST4). | ||
| Fig. 6 LC-MS mass spectra of untreated and treated (a) Az-A and (b) Az-B, and (c) intermediates formed after degradation of phenothiazine dyes. | ||
An upfold in the abundance of m/z 100 at 3.2 minutes in the treated sample showed the formation of a linear aldehyde (C6H12O), which further undergoes mineralization.63–65 In addition, an alternate reduction pathway was observed (Fig. 6c) wherein the dye molecule was broken down into fragments having m/z 224 (C12H20N2S) and m/z 235 (C8H15N2O4S), in accordance with the amine functionality observed in the FTIR data, and further formed a linear short chain of linear aldehyde with m/z 100 (C6H12O). The tandem MS analysis (Fig. S14) provided supportive evidence, with fragmentation of precursor ions with m/z 105, 120, 166, 188, 197 and 211 confirming the identity of each chemical species discussed in the proposed mechanism for Azure dye degradation. Based on this, the +ESI product ions reveal derivative structures such as o-tolualdehyde, phthalic acid and 2,5 dinitrobenzoic acid supporting the formation of each species leading to linear aldehydes (m/z 100) in the mass +ESI scan described in Fig. 6.
Marine anaerobic bacteria possess several classes of ligninolytic enzymes which degrade aromatic compounds present in pollutants.66 As shown in Fig. S15, NAG1 showed the presence of ligninolytic enzymes including Lac, LiP and MnP. In the presence of GO, there was enhanced production of ligninolytic enzymes. This may be attributed to GO acting as a nano-catalyst for cell proliferation67 or GO inducing stress leading to overexpression of lignocellulolytic enzyme synthesis-related genes.68 Lac-mediated one-electron oxidation processes, cleave the azo bond (–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–) present within the chromophore,69 which is consistent with azo bond cleavage and demethylation as shown in the FTIR data (Fig. 5). Additionally, the oxidative breakage (by ˙OH) of the central rings results in the formation of m/z 185 C7H7O2S+, which undergoes hydroxylation reactions releasing NH4+ and SO42−, followed by ring cleavage yielding a short chain of linear aldehyde.70 Previous studies have also reported the formation of a CoA thioester bond and the involvement of benzoyl-CoA-reductase in sequential opening of the aromatic ring structure,71 which is further reduced to linear aldehyde derivatives, correlating with the presence of m/z 100 (C6H12O) in the LC-MS analysis. These aldehyde derivatives are postulated to be mineralized, indicating complete degradation of phenothiazine dyes by the live NAG1–GO nano-assembly.
N–) present within the chromophore,69 which is consistent with azo bond cleavage and demethylation as shown in the FTIR data (Fig. 5). Additionally, the oxidative breakage (by ˙OH) of the central rings results in the formation of m/z 185 C7H7O2S+, which undergoes hydroxylation reactions releasing NH4+ and SO42−, followed by ring cleavage yielding a short chain of linear aldehyde.70 Previous studies have also reported the formation of a CoA thioester bond and the involvement of benzoyl-CoA-reductase in sequential opening of the aromatic ring structure,71 which is further reduced to linear aldehyde derivatives, correlating with the presence of m/z 100 (C6H12O) in the LC-MS analysis. These aldehyde derivatives are postulated to be mineralized, indicating complete degradation of phenothiazine dyes by the live NAG1–GO nano-assembly.
3.7. Significance of the study and its environmental implications
Marine bacteria belonging to genera such as Bacillus, Shewanella, Vibrio, and Halomonas possess molecular mechanisms that enable them to thrive in extreme environmental conditions, including high salinity, extreme pH levels, high temperatures, limited oxygen, and nutrient scarcity. These species inherently possess ligninolytic enzymes such as Lac, LiP, MnP and azo reductase which enable them cope with nutrient limitation and utilize organic pollutants as alternative nutrient sources. These extracellular ligninolytic enzymes achieve C–C bond cleavage, demethylation, hydroxylation, benzyl alcoholization and breaking of aromatic rings. However, when present in free or suspended form in water, they exhibit poor degradation efficiency due to washout, low survival rate and limited contact time with pollutants. Therefore, an immobilized carrier such as GO can provide a high specific area and rich oxygenated functional groups that enable cell attachment. GO also acts as a nano-shield, preventing dyes from being directly exposed to the cells. In addition, its high dye adsorbing capacity, accelerates degradation efficiency compared to free bacteria as demonstrated in our study.The nature and fate of the intermediates and by-products formed play an important role in ecological safety. Several dye degradation processes generate toxic and persistent aromatic structures which tend to bioaccumulate in the system. In contrast, the current findings highlight the formation of linear aldehydes and their derivatives as end-products, which are more likely to undergo mineralization. The GO–NAG1 nano-assembly offers a sustainable and efficient approach for the removal of phenothiazine dyes as it provides faster degradation rates and most importantly, no generation of secondary pollutants. For practical scalability, the GO–bacteria systems can be further immobilized into beads, membranes or bioreactors for the treatment of mixed dyes and textile effluents.
4. Conclusion
In the current study, we demonstrated the nano-assembly of live NAG1 (Bacillus sp.) cells onto biomimetic GO nanosheets for sustainable degradation of phenothiazine dyes – Azure A and Azure B. The GO–NAG1 nano-assembly was systematically optimized across varying concentrations of GO (20 to 150 µg mL−1) and pH values from 4 to 14. According to the cell viability results, GO promotes NAG1 cell proliferation at low concentrations and acts as a bactericidal agent at higher concentrations due to oxidative stress. The morphological analysis exhibited that wrinkles on GO act as a scaffold that promotes a favorable microenvironment, allowing cells to adhere, proliferate, and aggregate NAG1 cells (without mechanical damage). The optimized GO–NAG1 nano-assembly was further employed for the degradation of Azure A and Azure B dyes. Within 24 hours, GO–NAG1 showed 95.7 and 94.5% dye removal efficiencies, respectively, outperforming NAG1 and GO individually. LC-MS studies confirmed the proposed pathways, indicating the mineralization of dyes via two pathways – first by subsequent demethylation and second by enzymatic attack of ligninolytic enzymes such as Lac, LiP and MnP followed by the formation of linear aldehyde derivatives. This work highlights the synergistic effect of live NAG1 cells and GO nanosheets as a scaffold in facilitating the enhanced dye degradation without the generation of secondary pollutants.Author contributions
Neha Redkar: methodology, investigation, data curation, formal analysis and writing – original draft; Jyotsna Mishra: investigation, data curation, formal analysis, writing – review & editing; Rahul Kumar Das: formal analysis, writing – original draft, review & editing; Dharmveer Yadav: formal analysis, writing – review & editing, Cathrine Manohar Sumathi: writing – review & editing; Sumit Saxena: validation, supervision, funding acquisition, resources and writing – review & editing; Shobha Shukla: conceptualization, methodology, validation, supervision, project administration, resources, funding acquisition and writing – review & editing.Conflicts of interest
The authors have no relevant financial or non-financial interests to disclose.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: figures, tables, and other information. See DOI: https://doi.org/10.1039/d5ta06267e.Acknowledgements
This work was financially supported by the Department of Science and Technology (DST), Ministry of Science and Technology, Technology Mission Division, through grant no. DST/TM/WTI/WIC/2K17/100(C) and the Science and Engineering Research Board (SERB) through grant no. EMR/2017/005144. N. R. would like to thank the Council of Scientific & Industrial Research for the financial assistance provided through grant no. 09/087(1025)/2019-EMR-I. The authors would like to acknowledge the Sophisticated Analytical Instrument Facilities (SAIF), Central Facility, along with the Industrial Research and Consultancy Center (IRCC), of the Indian Institute of Technology Bombay, for providing research facilities. We acknowledge the Biological Oceanography Department, CSIR-NIO, Goa, for providing us with marine bacterial isolates.References
- M. A. H. Eid and G. Al-Abdallah, Sustainable Dev., 2024, 32, 6045–6056 CrossRef.
- E. Stratakis, J. Bonse, J. Heitz, J. Siegel, G. D. Tsibidis, E. Skoulas, A. Papadopoulos, A. Mimidis, A. C. Joel, P. Comanns and J. Krüger, Mater. Sci. Eng. R Rep., 2020, 141, 100562 Search PubMed.
- Y. Zhou, H. Lu, J. Wang, J. Zhou, X. Leng and G. Liu, J. Hazard. Mater., 2018, 356, 82–90 Search PubMed.
- G. Wen, Z. Guo and W. Liu, Nanoscale, 2017, 9, 3338–3366 Search PubMed.
- S. Varjani, P. Rakholiya, H. Y. Ng, S. You and J. A. Teixeira, Bioresour. Technol., 2020, 314, 123728 CrossRef CAS PubMed.
- B. Biswas, L. N. Warr, E. F. Hilder, N. Goswami, M. M. Rahman, J. G. Churchman, K. Vasilev, G. Pan and R. Naidu, Chem. Soc. Rev., 2019, 48, 3740–3770 RSC.
- M. A. Ruiz-Fresneda, E. González-Morales, C. Gila-Vilchez, A. Leon-Cecilla, M. L. Merroun, A. L. Medina-Castillo and M. T. Lopez-Lopez, Mater. Horiz., 2024, 11, 5533–5549 RSC.
- N. Donkadokula, N. Yadav, A. Kishore, K. Kola, I. Naz and D. Saroj, Rev. Environ. Sci. Biotechnol., 2020, 19, 543–560 CrossRef CAS.
- Y. Song, L. Wang, X. Qiang, W. Gu, Z. Ma and G. Wang, J. Water Process Eng., 2023, 55, 104242 Search PubMed.
- C. X.-H. Su, P. C. Y. Ling, S. W. Lim and P. E. Poh, J. Environ. Chem. Eng., 2016, 4, 3618–3631 CrossRef CAS.
- J. Fan, Y. Luo, X. Chen, L. Li and J. Li, Chemosphere, 2018, 191, 315–323 CrossRef CAS PubMed.
- J. Wachtmeister and D. Rother, Curr. Opin. Biotechnol., 2016, 42, 169–177 CrossRef CAS PubMed.
- A. Dsouza, C. Constantinidou, T. N. Arvanitis, D. M. Haddleton, J. Charmet and R. A. Hand, ACS Appl. Mater. Interfaces, 2022, 14, 47323–47344 Search PubMed.
- A. M. Brozell, J. de Grooth and E. M. V. Hoek, Nat. Water, 2024, 2, 392–396 CrossRef.
- Y. Yoon, P. L. Truong, D. Lee and S. H. Ko, ACS Nanosci. Au, 2022, 2, 64–92 CrossRef CAS PubMed.
- A. Pandey, R. K. Das, A. K. Chakraborty, S. Shukla and S. Saxena, J. Clean. Prod., 2022, 362, 132257 CrossRef CAS.
- J. Mishra, T. Suryawanshi, N. Redkar, R. Kumar Das, S. Saxena, A. Majumder, K. Kondabagil and S. Shukla, ChemSusChem, 2025, 18, e202402056 CrossRef CAS PubMed.
- X. Sun, C. Huang, L. Wang, L. Liang, Y. Cheng, W. Fei and Y. Li, Adv. Mater., 2021, 33, 2001105 CrossRef CAS PubMed.
- A. C. Barrios, Y. Wang, L. M. Gilbertson and F. Perreault, Environ. Sci. Technol., 2019, 53, 14679–14687 CrossRef CAS PubMed.
- M. Deb, N. Redkar, C. S. Manohar, A. S. Jagtap, S. Saxena and S. Shukla, Environ. Pollut., 2023, 326, 121490 CrossRef CAS PubMed.
- O. N. Ruiz, K. A. S. Fernando, B. Wang, N. A. Brown, P. G. Luo, N. D. McNamara, M. Vangsness, Y. P. Sun and C. E. Bunker, ACS Nano, 2011, 5, 8100–8107 CrossRef CAS PubMed.
- P. C. Henriques, A. T. Pereira, A. L. Pires, A. M. Pereira, F. D. Magalhaes and I. C. Goncalves, ACS Appl. Mater. Interfaces, 2020, 12, 21020–21035 Search PubMed.
- J. Xue, S. BinAhmed, Z. Wang, N. G. Karp, B. L. Stottrup and S. Romero-Vargas Castrillón, Environ. Sci. Technol. Lett., 2018, 5, 14–19 CrossRef CAS.
- W. Xu, Z. Jin, X. Pang, Y. Zeng, X. Jiang, Y. Lu and L. Shen, ACS Appl. Bio Mater., 2020, 3, 4263–4272 CrossRef CAS PubMed.
- S. Liu, T. H. Zeng, M. Hofmann, E. Burcombe, J. Wei, R. Jiang, J. Kong and Y. Chen, ACS Nano, 2011, 5, 6971–6980 Search PubMed.
- O. Akhavan, E. Ghaderi and A. Esfandiar, J. Phys. Chem. B, 2011, 115, 6279–6288 Search PubMed.
- S. Revoju, A. Matuhina, L. Canil, H. Salonen, A. Hiltunen, A. Abate and P. Vivo, J. Mater. Chem. C, 2020, 8, 15486–15506 Search PubMed.
- H. Zhang, X. Liu, B. Liu, F. Sun, L. Jing, L. Shao, Y. Cui, Q. Yao, M. Wang, C. Meng and Z. Gao, J. Hazard. Mater., 2023, 458, 131939 Search PubMed.
- J. C. Bollinger, E. C. Lima, L. Mouni, S. Salvestrini and H. N. Tran, Environ. Chem. Lett., 2023, 21, 2525–2552 Search PubMed.
- S. Saxena, T. A. Tyson and E. Negusse, J. Phys. Chem. Lett., 2010, 1, 3433 Search PubMed.
- J. Karmacharya, S. Lee, J. Kim, H. Park, S. Lee and S. Kim, Chemosphere, 2025, 372, 144137 CrossRef CAS PubMed.
- Y. Ding, L. Zhang, X. Li, H. Wang, Z. Li and Q. Zhou, J. Hazard. Mater., 2022, 431, 128544 CrossRef CAS PubMed.
- S. Kudo, T. Kataoka, S. Sugiura, H. Hirakawa and Y. Tanaka, ACS Omega, 2017, 2, 7329–7333 Search PubMed.
- K. Haddad, A. Abokifa, S. An, J. Lee, B. Raman, P. Biswas and J. D. Fortner, J. Mater. Chem. A, 2023, 11, 447–459 RSC.
- S. Saxena, T. A. Tyson, S. Shukla, E. Negusse, H. Chen and J. Bai, Appl. Phys. Lett., 2011, 99, 013104 Search PubMed.
- V. Brusko, A. Khannanov, A. Rakhmatullin and A. M. Dimiev, Carbon, 2024, 229, 119507 CrossRef CAS.
- D. Yadav, R. K. Das, S. Saxena and S. Shukla, J. Clean. Prod., 2023, 411, 137266 CrossRef CAS.
- P. Braylé, E. Pinelli, B. Schoefs, E. Flahaut, J. Silvestre, L. Gauthier and M. Barret, Carbon, 2024, 226, 119224 CrossRef.
- Q. Jiang, F. E, J. Tian, J. Yang, J. Zhang and Y. Cheng, ACS Appl. Mater. Interfaces, 2020, 12, 16150–16158 CrossRef CAS PubMed.
- A. Mougi, npj Biodiversity, 2024, 3, 32 CrossRef PubMed.
- M. Wang, Z. Li, Y. Zhang, Y. Li, N. Li, D. Huang and B. Xu, J. Hazard. Mater., 2021, 412, 125333 CrossRef CAS PubMed.
- R. Castellanos-Espinoza, S. Fernández-Tavizón, U. Sierra-Gómez, E. A. Elizalde-Peña, G. Luna-Bárcenas, L. A. Baldenegro-Pérez, L. I. Olvera, L. V. González-Gutiérrez, C. M. Ramos-Castillo, N. Arjona and B. L. España-Sánchez, Appl. Surf. Sci., 2023, 623, 156953 CrossRef CAS.
- C. Ratzke and J. Gore, PLoS Biol., 2018, 16, e2004248 CrossRef PubMed.
- I. Sengupta, P. Bhattacharya, M. Talukdar, S. Neogi, S. K. Pal and S. Chakraborty, Colloid Interface Sci. Commun., 2019, 28, 60–68 CrossRef CAS.
- S. Romero-Vargas Castrillón, F. Perreault, A. F. De Faria and M. Elimelech, Environ. Sci. Technol. Lett., 2015, 2, 112–117 CrossRef.
- H. Tang, S. Zhang, T. Huang, J. Zhang and B. Xing, Environ. Sci. Technol., 2021, 55, 14639–14648 CrossRef CAS PubMed.
- V. Palmieri, F. Bugli, M. C. Lauriola, M. Cacaci, R. Torelli, G. Ciasca, C. Conti, M. Sanguinetti, M. Papi and M. De Spirito, ACS Biomater. Sci. Eng., 2017, 3, 619–627 CrossRef CAS PubMed.
- T. Pulingam, K. L. Thong, M. E. Ali, J. N. Appaturi, I. J. Dinshaw, Z. Y. Ong and B. F. Leo, Colloids Surf., B, 2019, 181, 6–15 CrossRef CAS PubMed.
- Q. Xu, X. Lin, L. Gan, G. Owens and Z. Chen, J. Colloid Interface Sci., 2022, 605, 881–887 CrossRef CAS PubMed.
- K. Meyer, C. Addy, S. Akashi, D. I. Roper and J. R. Tame, Biochem. Biophys. Res. Commun., 2018, 499, 594–599 CrossRef CAS PubMed.
- Y. Yang, J. Liu, B. R. Clarke, L. Seidel, J. R. Bolla, P. N. Ward, P. Zhang, C. V. Robinson, C. Whitfield and J. H. Naismith, Nat. Commun., 2021, 12, 4349 CrossRef CAS PubMed.
- M. Kjærvik, M. Ramstedt, K. Schwibbert, P. M. Dietrich and W. E. S. Unger, Front. Chem., 2021, 9, 666161 CrossRef PubMed.
- J. Xiao, W. Lv, Z. Xie, Y. Tan, Y. Song and Q. Zheng, J. Mater. Chem. A, 2016, 4, 12126–12135 RSC.
- I. Haq and A. Raj, Chemosphere, 2018, 196, 58–68 CrossRef CAS PubMed.
- Y. Tian, K. Wu, S. Lin, M. Shi, Y. Liu, X. Su and R. Islam, ACS Omega, 2024, 9, 7668–7678 CAS.
- L. Shen, Z. Jin, D. Wang, Y. Wang and Y. Lu, Chemosphere, 2018, 190, 201–210 CrossRef CAS PubMed.
- P. Banerjee, S. Mishra, S. Mondal, A. Saha and S. Chattopadhyay, Clean Technol. Environ. Policy, 2018, 20, 2287–2298 CrossRef CAS.
- H. Tiwari, P. Tripathi, R. K. Sonwani and R. S. Singh, Bioresour. Technol., 2023, 387, 129614 CrossRef CAS PubMed.
- L. Shen, Y. Wang, Y. Liu, M. Zhang, J. Yu and H. Yang, Ind. Eng. Chem. Res., 2019, 58, 7817–7824 CrossRef CAS.
- Z. Hua, Y. Li, X. Wang, J. Chen, L. Zhao and Y. Zhang, Environ. Res., 2023, 236, 116739 CrossRef CAS PubMed.
- E. Martínez-Periñán, A. Domínguez-Saldaña, A. M. Villa-Manso, C. Gutiérrez-Sánchez, M. Revenga-Parra, E. Mateo-Martí, F. Pariente and E. Lorenzo, Sens. Actuators, B, 2022, 370, 132761 Search PubMed.
- J. Wang, D. Li, W. Tao, Y. Lu, X. Yang and J. Wang, Biomacromolecules, 2019, 20(7), 1740–1747 CrossRef CAS PubMed.
- N. Gautam, K. B. Singh, D. D. Upadhyay and G. Pandey, RSC Adv., 2023, 13, 23181–23196 Search PubMed.
- P. Jia, H. Tan, K. Liu and W. Gao, Mater. Res. Bull., 2018, 102, 45–50 CrossRef CAS.
- S. Gangola, G. Bhandari, S. Joshi, A. Sharma, H. Simsek and P. Bhatt, Environ. Res., 2023, 232, 116332 Search PubMed.
- L. F. Muriel-Millán, S. Millán-López and L. Pardo-López, Appl. Microbiol. Biotechnol., 2021, 105, 7171–7185 CrossRef PubMed.
- S. Noreen, M. Asgher, S. Hussain, R. A. Haroon, M. Bilal and H. M. N. Iqbal, Environ. Technol. Innov., 2021, 24, 101884 CrossRef CAS.
- S. Liu, Y. Zhang, X. Wang, L. Chen, H. Li and J. Zhao, Appl. Environ. Microbiol., 2022, 88, e00960–22 Search PubMed.
- G. Mustafa, S. Khan, R. Ahmad, M. Ali and F. Shah, J. Environ. Chem. Eng., 2025, 13, 115605 CrossRef CAS.
- J. Van Buren, M. Smith, L. Johnson and K. Thompson, Environ. Sci. Technol., 2020, 54, 8352–8361 CrossRef CAS PubMed.
- O. Tiedt, J. Fuchs, W. Eisenreich and M. Boll, J. Biol. Chem., 2018, 293, 10264–10274 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |