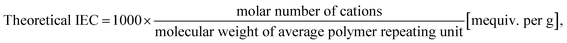Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Orchestrating explainable AI, ChatGPT, and human expertise: a framework for extracting polymer design guidelines
Yin Kan Phua a,
Nana Terasobab,
Manabu Tanaka
a,
Nana Terasobab,
Manabu Tanaka bc,
Tsuyohiko Fujigaya
bc,
Tsuyohiko Fujigaya *ade and
Koichiro Kato
*ade and
Koichiro Kato *aef
*aef
aDepartment of Applied Chemistry, Graduate School of Engineering, Kyushu University, 744 Motooka, Nishi-ku, Fukuoka 819-0395, Japan. E-mail: fujigaya.tsuyohiko.948@m.kyushu-u.ac.jp; kato.koichiro.957@m.kyushu-u.ac.jp
bDepartment of Applied Chemistry, Tokyo Metropolitan University, 1-1 Minami-osawa, Hachioji, Tokyo 192-0397, Japan
cResearch Center for Hydrogen Energy-based Society (ReHES), Tokyo Metropolitan University, 1-1 Minami-osawa, Hachioji, Tokyo 192-0397, Japan
dInternational Institute for Carbon Neutral Energy Research, Kyushu University, 744 Motooka, Nishi-ku, Fukuoka 819-0395, Japan
eCenter for Molecular Systems, Kyushu University, 744 Motooka, Nishi-ku, Fukuoka 819-0395, Japan
fResearch Institute for Information Technology, Kyushu University, 744 Motooka, Nishi-ku, Fukuoka 819-0395, Japan
First published on 14th April 2026
Abstract
To accelerate the rational design of high-performance functional polymers, such as anion exchange membranes (AEMs), the establishment of chemically meaningful and actionable design guidelines is essential. Machine learning (ML) models, particularly artificial neural networks (ANNs), offer high predictive accuracy for such materials but suffer from limited interpretability due to their black-box nature. Although explainable artificial intelligence (XAI) methods such as SHapley Additive exPlanations (SHAP) provide a unified framework for model explanation, their application to ANN models is hindered by the expensive computation cost associated with the high dimensionality of molecular descriptors commonly used to represent polymer structures. In this study, a framework that combines statistical (minimum redundancy maximum relevance) and explainable ANN-based (permutation importance via ELI5) feature selection was developed, reducing the input space to 67 key descriptors. This dimensionality reduction enabled computationally feasible SHAP analysis while enhancing the predictive accuracy of the ANN by 40.87%. However, the resulting key descriptors were often difficult to interpret in physicochemical terms. To address this, large language models (LLMs) such as ChatGPT were employed to analyze descriptor source code and assist human experts in deriving chemically intuitive insights. By orchestrating XAI, LLM assistance, and expert knowledge, the framework successfully extracted design guidelines for AEMs. Based on these insights, two candidate AEM polymers with predicted anion conductivities ≥0.1 S cm−1 at 80 °C were proposed, exceeding typical commercialization thresholds. This study illustrates a generalizable, explainable, and efficient pathway for integrating ML, XAI, and LLMs in polymer informatics, with broad applicability across descriptor-based materials research.
Introduction
Recent advances in machine learning (ML) and artificial intelligence (AI) have fundamentally reshaped scientific methodologies, enabling the accelerated discovery of novel materials with tailored functionalities across diverse material classes.1–28 These techniques showed effectiveness in predicting target properties, enabling high-throughput virtual screening, and substantially reducing experimental workloads and resource consumption during materials development.29–31 Among the various ML models, artificial neural networks (ANNs)32,33 have received particular attention for their high predictive ability,4,6,7,9,12,13,15,18,19,23–28 owing to their capacity to capture complex, nonlinear relationships between explanatory and target variables,34–37 as opposed to tree-based models, which are weak in extrapolation.38–40 Despite the high predictive performances of ANNs, they are commonly regarded as black-box models41 due to the difficulties associated with interpreting their internal decision-making processes.42 During materials research, it is necessary to design new structures while optimizing and balancing various interdependent physical and chemical factors. Explainable ML/AI (XAI) can address this challenge by clarifying how input variables contribute to predictions.XAI can be achieved through post-hoc explanation methods, which can be broadly classified into two types, namely model-dependent and model-agnostic (model-independent). Model-dependent methods, such as feature importance in random forests,43 rely on the model's internal structure and parameters. Such mechanisms are typically absent in ANNs, rendering model-agnostic methods, such as local interpretable model-agnostic explanations (LIME),44 SHapley Additive exPlanations (SHAP),45 and permutation importance,46 more suitable. LIME offers local explanations by analyzing how input variables influence individual predictions, while permutation importance provides global insights by assessing the overall feature importance. While each can be useful independently,3,5,6,14,47–52 materials science applications benefit from both global explanations to uncover general structure–property relationships and local explanations to understand the behavior of individual materials or data points.53–55
SHAP fulfills this requirement by offering a unified framework capable of delivering both global and local explanations with internal consistency,56 making it well-suited for materials science applications.9,22,53,54,57–60 Previous studies primarily applied SHAP to non-ANN-based models,9,22,57,58,60 while SHAP analyses of ANNs remain limited to low-dimensional datasets (typically <100 features).53,54,59 This is primarily due to the high computational cost associated with calculating SHAP values,56,61–64 hindering its application towards high-dimensional datasets.
Representing organic molecules and polymers in a machine-readable numerical form typically involves molecular descriptors and/or fingerprints derived from tools such as the Mordred library65 and RDkit package,66 all of which employ feature vectors with several hundreds to <2000 dimensions. This high dimensionality limits the practical application of SHAP in the ANN-based modeling of organic and polymeric materials. This is where dimensionality reduction (feature selection) can be useful due to its ability to decrease the number of variables in a high-dimensional dataset while preserving meaningful characteristics of the original data. Various dimensionality reduction techniques exist, including manual feature selection based on expert knowledge (feature elimination),47,48 wrapper-based feature selection methods such as recursive feature elimination (RFE),67–69 embedded feature selection using model-internal metrics (feature importance in RF), feature extraction methods such as principal component analysis (PCA),70,71 and filter-based methods leveraging statistical or information-theoretic metrics. Manual selection of relevant descriptors requires expertise in molecular descriptors, whereas feature extraction methods (e.g., PCA) compress several descriptors into components at the cost of interpretability. RFE is effective for reducing the dimensions of molecular descriptors,3,6,72–78 but it suffers from extensive computational costs due to its iterative retraining strategy, which is especially problematic for datasets with high dimensionalities.69,74 Additionally, embedded feature selection works by simultaneously performing model training and feature selection by internally evaluating feature relevance through criteria inherent to the learning algorithm itself.79–81 Since an ANN does not possess any inherent feature importance measures, it is generally incompatible with embedded feature selection. Moreover, filter-based feature selection involves selecting features based on statistical or information-theoretic criteria independent of predictive models. This requires a low computational cost but typically evaluates features independently, thereby missing inter-variable relationships.82,83 Despite such variety, no established method exists that is simultaneously explainable, computationally feasible, and broadly applicable for reducing the dimensionality of molecular descriptors in SHAP-based ANN analysis. In addition, previous materials science reports that have applied SHAP to explain ML/AI have mainly considered explanations based on individual variables.84–89 However, reports have been published in the contexts of genetic analysis,90 aviation safety,91 and financial fraud,92 analyzing important variables obtained from feature importance or SHAP in groups formed through knowledge-driven methods, thereby showcasing the ability to obtain a simplified explanation and a more general scope. These considerations motivate the development of new approaches tailored for high-dimensional datasets, along with validation of the effectiveness of group-wise SHAP important variable analysis in materials science. Additionally, molecular descriptors are often mathematically complex, making their interpretation inherently challenging; as a result, descriptors frequently remain poorly understood despite effectively encoding structural information that enables ML/AI models to capture trends and make accurate predictions.71,93,94 Consequently, even when important descriptors are identified via explainable ANN methods, an interpretative framework is essential to translate them into meaningful, actionable chemical insights and practical experimental guidelines. This interpretative step is crucial for orchestrating effective collaboration between explainable AI and human expert knowledge, enhancing the practical utility and scientific value of ML-driven materials discovery.
In this study, a generalizable model-agnostic framework that integrates SHAP-based XAI methods with ANN models is proposed, aiming to systematically extract chemically intuitive and chemically interpretable design guidelines from high-dimensional molecular descriptor datasets for polymeric materials. The effectiveness of this framework is validated using a polymer dataset of anion exchange membranes (AEMs), a class of functional polymeric materials that play a central role in electrochemical energy conversion technologies relevant to the hydrogen economy.95 AEMs serve as solid polymer electrolytes in anion exchange membrane fuel cells (AEMFCs) and anion exchange membrane water electrolyzers (AEMWEs), where they conduct hydroxide or other anions between electrodes while providing electronic insulation and gas separation.96,97 Compared to proton exchange membrane-based systems, AEM-based devices can operate under alkaline conditions that permit the use of non-precious metal catalysts, offering potential advantages in cost reduction and resource sustainability.98
Despite these advantages, the molecular design of high-performance AEM polymers remains challenging due to inherent trade-offs between achieving high anion conductivity and maintaining long-term alkaline stability.97,99–103 High anion conductivity typically requires elevated ion-exchange capacity (IEC) through dense incorporation of cationic functional groups along the polymer backbone or side-chains.104 However, increased IEC is often accompanied by higher water uptake and swelling, which can compromise mechanical and dimensional stability and accelerate chemical degradation under alkaline operating conditions.104 As a result, AEM polymer design often involves navigating competing and interdependent molecular requirements related to backbone chemistry, side-chain architecture, and ionic group distribution. This complex, multidimensional design space motivates the use of data-driven and interpretable modeling approaches capable of disentangling structure–property relationships and identifying molecular features that simultaneously favor conductivity and stability. In this study, both anion conductivity and alkaline stability are targeted using a unified predictive model, thereby providing a directly relevant materials context for demonstrating the proposed explainable ML framework.
To overcome the dimensionality barrier that limits the use of SHAP in ANN-based modeling, statistical and explainable ANN-based feature selection techniques are combined to reduce the descriptor dimensionality in two steps, preserving the explainability during the latter process (Scheme 1). The ability of this feature selection strategy to enable computationally feasible SHAP analysis is evaluated, along with its model performance. Initially, conventional SHAP analysis is applied to identify crucial individual molecular descriptors, offering insight into variable-specific structure–property relationships. Subsequently, a novel pairwise SHAP interpretation method is proposed to analyze the interrelationships between top-ranked explanatory variables, potentially revealing chemically meaningful interactions between descriptors that may not be evident through individual descriptor analysis. This could facilitate the extraction of actionable materials design guidelines and eventually accelerate the rational design of high-performance polymers. As a demonstration of orchestrating ANN models, XAI-driven feature interpretation, and domain expert knowledge, the proposed framework is applied to the top influential descriptors, whose physicochemical meanings are further elucidated through a collaborative analysis involving ChatGPT and human experts, thereby highlighting the practicality, transparency, and interpretability of this integrated methodology.
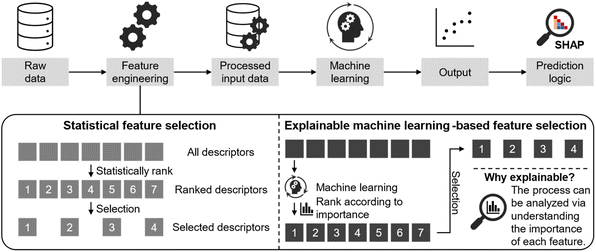 | ||
| Scheme 1 Schematic illustration outlining the systematic incorporation of SHAP into an ANN when working with a high-dimensional database. | ||
Experimental
Data curation
This study employed an in-house database curated using our previously reported method, which is now publicly available on GitHub.71,94 The data included in the database include the chemical structures of AEM polymers, experimental conditions (measurement temperature for anion conductivity, concentration of alkaline solution, temperature, and duration for alkaline stability tests), the ion-exchange capacity (IEC), the anion conductivities of pristine AEM polymers (pristine anion conductivity), and the conductivities measured during the alkaline stability test (degraded anion conductivity).The chemical structures were handled according to the method described in our previous reports.71,94 Initially, the anion-conducting polymers were separated into either homo- or copolymers, wherein the homopolymers contain one type of repeating structure, whereas the copolymers had two. Subsequently, the chemical structures were separated into two blocks, namely blocks A and B, based on the number of repeating units in the copolymer. Block A comprised repeating units in the AEM material containing an anion-conducting moiety, while block B constituted another repeating unit that did not contain an anion-conducting moiety (Fig. 1). Anion conductivity measures the ability of an AEM to conduct anions, with a unit of Siemens per centimeter (S cm−1), and is temperature- and humidity-dependent. Generally, anion conductivity increases as temperature and/or humidity increases. Ideally, the anion conductivity of an AEM polymer should be ≥0.1 S cm−1 at its operating temperature.105 The IEC was calculated theoretically as follows:
Descriptor conversion of the anion-conducting polymer structures
The chemical structures were converted into machine-readable numerical forms using the Mordred molecular descriptor library,65 following our previous reports.71,94 Mordred is an open-source library that calculates a comprehensive set of molecular descriptors and fingerprints, yielding 1825 features per structure. The resulting high-dimensional dataset was used for feature selection and ANN modeling in subsequent steps.Pre-processing and statistical-based feature selection
The high-dimensional database described above was initially scaled using StandardScaler from scikit-learn106 to ensure that each feature possessed zero mean and unit variance, which is beneficial for stabilizing ANN training. Features exhibiting an extremely low variance were removed by applying a variance threshold (threshold = 0), eliminating predictors that contributed negligibly to the model. Subsequently, minimum redundancy maximum relevance (mRMR) feature selection107,108 was performed to reduce the remaining feature set to 1000 dimensions. mRMR is a feature selection technique that calculates the mutual information between each feature and the target variable (maximizing relevance), while penalizing excessive overlap among the features themselves (minimizing redundancy). Through iterative scoring and selection, mRMR ensures that the final subset has the greatest predictive power with the least redundancy among the variables.Construction of the ANN and explainable ML-based feature selection
The ANN was constructed using the MLPRegressor module from scikit-learn.106 The explanatory variables included variables that survived the mRMR filtering, while the target variable was the anion conductivity (as measured in the pristine state and during the alkaline stability test). Cross-validation was performed using a group k-fold (n_split = 5), and the hyperparameters were tuned using Optuna.109 Specifically, the number of neurons in each of three or five hidden layers (ranging from 2 to 600, depending on the model), the activation function (Rectified Linear Unit (ReLU), tanh, or logistic), the solver (stochastic gradient descent (SGD) or Adam), the initial learning rate (0.00001–0.01, logarithmic scale), the regularization parameter α (0.1–5, logarithmic scale), and the batch size (64, 128, 256, 512, or 1024) were automatically optimized based on the trial suggestions. After identifying the best-performing model hyperparameters from cross-validation, the model was retrained using the best hyperparameters and the combined cross-validation dataset (combined training–validation dataset), yielding the optimal model for evaluation. Following this tuning step, an explainable ML-based feature selection procedure was performed on the best performing model using ELI5, a model-agnostic tool that computes global permutation importance with negligible computational cost. ELI5 derives a global explanation by estimating the significance of each feature measured through the overall deterioration in the predictive performance when that feature is shuffled across all samples in the validation set. Based on the importance of each explanatory variable obtained from the global explanation, the 64 top-ranked features were retained, and the final predictive ANN model was trained again using these reduced features, with the types of hyperparameters tuned to be the same as in the first model.ANN model performance metrics
The ANN model performance was evaluated using the root mean squared error (RMSE) and the mean absolute error (MAE). Both metrics quantify the prediction error between the actual and predicted values, enabling direct comparison across models. RMSE emphasizes large errors due to the squaring of residuals and is therefore more sensitive to outliers. In contrast, MAE calculates the average absolute difference and provides a more balanced view by reducing the influence of outliers. Using both metrics therefore offers complementary perspectives related to model accuracy and robustness. The equation used to calculate the RMSE is as follows:
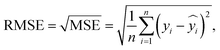 | (1) |
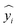 is the predicted value of yi. Additionally, the MAE was calculated as follows:
is the predicted value of yi. Additionally, the MAE was calculated as follows:
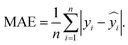 | (2) |
Prediction logic evaluations for the models
The prediction logic for each model was evaluated using SHAP,45 which quantifies the contribution of each explanatory variable to the predicted output based on cooperative game theory. More specifically, the KernelExplainer module in SHAP was employed due to the fact that it is model-agnostic,45 rendering it suitable for ANNs that typically lack specialized explainer variants available for tree-based or gradient-boosted models. The top 20 explanatory variables, ranked by their average absolute SHAP values, were extracted, revealing the most influential predictors driving the ANN's predicted outputs.Results and discussion
Pre-processing and statistical-based feature selection
The AEM database used in this study was reported in our previous studies71,94 and is openly available on GitHub. The database covers 2452 data points corresponding to anion conductivity, which were collected from 78 papers published between 2007 and 2024 and which belong to 323 copolymers and 23 homopolymers. The anion conductivity data originated from both temperature dependence anion conductivities of pristine AEMs and AEMs that were undergoing degradation tests. All anion conductivity data were recorded under fully hydrated conditions. The 346 AEM polymer structures were incorporated into the database by categorizing the repeating units into blocks A and B using the method described in our previous reports.71,94 Both blocks were converted to a numerical form using Mordred, producing a database containing a total of 3226 descriptors. Among the descriptors obtained, those containing non-numerical values or empty rows were removed, leaving 2238 molecular descriptors (1210 for block A and 1028 for block B) in the database. Seven descriptors with low variance were removed, yielding 2231 descriptors (1204 belonging to block A and 1027 belonging to block B). The statistical-based feature selection method mRMR was then employed to further reduce the descriptors to <1000, yielding a total of 998 descriptors (749 for block A and 249 for block B). To ensure the retention of experimentally important variables, certain parameters (including the molar ratio between blocks A and B, information related to the presence or absence of polymer cross-linking, the alkaline stability test temperature, the concentration of the alkaline solution employed during the alkaline stability test, the duration of the alkaline stability test, the anion conductivity measurement temperature, and the IEC) were intentionally reintroduced into the final dataset if they had been excluded during the pre-processing steps. After this targeted reincorporation and statistical feature selection, the final dataset comprised 1007 dimensions. Such reincorporation is particularly important in the context of AEM polymers, where the database contains temperature-dependent measurements related to anion conductivity. When applying this workflow to other materials systems, the choice of experimentally important variables can be adapted based on domain-specific knowledge.ANN construction and explainable ML-based feature selection
The ANN was constructed using the MLPRegressor module of the scikit-learn library in Python. Two ANNs with different numbers of hidden layers were tested in this study, namely three hidden layers (named Model I) and five hidden layers (named Model II). The ANN exhibiting the greatest test predictive accuracy was used for the remainder of the study. The hyperparameters of both models were optimized using Optuna (number of trials = 2000) followed by manual fine-tuning (the optimized hyperparameters of Models I and II are shown in Tables S1 and S2, respectively), and the final models with optimized hyperparameters were used to predict the 14 AEM polymers present in the test database (separated from the training–validation database) to evaluate their generalization performance on unseen data. The RMSE and MAE values from combined training and validation indicated distinct differences in terms of the prediction performances between Models I and II, with Model II outperforming Model I by 35.65% (RMSE) and 34.57% (MAE), corresponding to absolute differences of 0.0082 and 0.0056 S cm−1, respectively (Table 1, Fig. S1a, b and Scheme S1a, b). Test predictions also revealed substantial differences in accuracy between the two models, with Model II showing a 0.0119 S cm−1 reduction in the RMSE (34.10% improvement) and a 0.0112 S cm−1 reduction in the MAE (39.02% improvement) compared to Model I (Table 1 and Fig. 2a). The greater disparity observed in the MAE compared to the RMSE suggests that Model II consistently made predictions closer to the actual values across most test samples, whereas Model I may have produced fewer extreme outliers but a higher overall error (Fig. 2a). Regardless, both models significantly outperformed previously reported MLPRegressor-based results (MLPRegressor, 2 hidden layers, RMSE = 0.344 S cm−1),94 likely due to the increased model depth enabling the ANNs to capture more complex structure–property relationships, as well as the use of a larger dataset. However, due to the high dimensionality (>1000 features), direct SHAP analysis at this stage was computationally prohibitive (Fig. S2). Exact Shapley value computation scales exponentially with the number of features (O(2D)), and while KernelSHAP provides a sampling-based approximation, its computational cost (and associated memory footprint) still increases rapidly with dimensionality in practice due to the need for more coalition evaluations and larger internal allocations as the feature space grows.110 This effect is quantified in Table S6 and Fig. S2b, c, which benchmark KernelSHAP across D = 25–1000 under controlled conditions (Table S6 Panel A) and under manuscript-style full-training conditions at D = 67 (Table S6 Panel B). Even after dimensionality reduction to D = 67, full-training KernelSHAP remains computationally intensive (Table S6 Panel B), underscoring the necessity of the staged dimensionality-reduction workflow adopted in this study (Fig. S2). Considering the consistently superior predictive performance and generalization capability of Model II, it was selected as the baseline model for subsequent feature selection and explainability analyses.| Combined training–validation accuracy | Test predictive accuracy | |||
|---|---|---|---|---|
| RMSE/S cm−1 | MAE/S cm−1 | RMSE/S cm−1 | MAE/S cm−1 | |
| Model I | 0.0230 | 0.0162 | 0.0349 | 0.0287 |
| Model II | 0.0148 | 0.0106 | 0.0230 | 0.0175 |
| Model II-mini | 0.0098 | 0.0071 | 0.0136 | 0.0089 |
Subsequently, ELI5 was applied to Model II to extract the top 64 important explanatory variables (Fig. 2b and Table S3). Model II identified the anion conductivity measurement temperature and the IEC as the two most important variables, aligning with chemical intuition for AEM polymers. Further inspection revealed that most top-ranked descriptors belonged to the Moran autocorrelation (MATS) and Moreau–Broto autocorrelation (AATS and ATSC) families (Fig. 2b), primarily representing topological and electronic descriptors. Sigma electrons (MATS5d_B, AATSC5d_B, AATS6d_A, and Xc-5d_B) and ionization potential-related descriptors (MATS1i_B and AATSC1i_A) were particularly prioritized (Table S4). Furthermore, Model II identified seven block B-related descriptors, implying that the values of these descriptors (including the ionization potential-related MATS1i_B) affect the anion conductivity, although it was unclear whether a high value led to a higher anion conductivity purely due to the variable importance (Fig. 2b). Notably, MATS1i_B is a descriptor that assesses the uniformity of the ionization potential or charge distribution across adjacent atoms, such as in the case where the adjacent atoms have similar chemical properties (e.g., the six carbons in a benzene ring). Possessing higher uniformity is known to enhance the alkaline stability of the AEM, with the highly uniform backbones of aryl-ether-free poly(aryl piperidinium)-based AEMs representing a prominent example.111 The focus on underlying electronic structure-related descriptors exhibited by Model II correctly reflects the chemical properties of AEM polymers, as anion conductivity is influenced by the stability and ease of transportation of anions by cationic functional groups (e.g., quaternary ammonium ions). These results demonstrate that utilizing global permutation importance for explainable feature selection enhances the transparency of the modeling process, providing clear justifications for the inclusion of chemically meaningful descriptors in the succeeding model trained using a database with a reduced feature set.
After obtaining the top 64 explanatory variables ranked by ELI5 for Model II, a new dimensionally reduced database was constructed accordingly. In line with the earlier statistical pre-processing step, experimentally important variables including the molar ratio between blocks A and B, cross-linking information, the alkaline stability test temperature, the concentration of the alkaline solution, the duration of the alkaline stability test, the anion conductivity measurement temperature, and the IEC were reincorporated into the final database if they had not been selected by ELI5 as one of the top 64 variables. This step does not contradict the logic of model-driven feature selection but rather complements it by ensuring that variables critical to the experimental context, particularly those known to influence the temperature-dependent conductivity behavior in AEM polymers, are preserved for model explainability and scientific validity. As a result, the final input database consisted of 67 variables. Based on Model II, a new ANN model (named Model II-mini) was constructed using the dimensionally reduced dataset (final input database) obtained through ELI5 feature selection. Model II-mini was trained based on the final output database and optimized with Optuna, with a significantly lower number of nodes in each hidden layer (∼1/10) compared to those of Models I and II (Table S5).
Model II-mini demonstrated a significantly improved predictive accuracy compared to Models I and II (Table 1 and Fig. 3a, b). It should be noted that all three models were trained and evaluated using the same underlying experimental database; the distinction lies in the input feature dimensionality, with Models I and II utilizing 1007 descriptors obtained after statistical feature selection and Model II-mini utilizing the 67 top-ranked descriptors identified through explainable ML-based feature selection using ELI5 permutation importance. More specifically, the RMSE for the combined training–validation dataset decreased by 0.0132 S cm−1 (57.39% improvement) and 0.0050 S cm−1 (33.78% improvement) relative to Models I and II, respectively, while the test dataset RMSE dropped by 0.0213 S cm−1 (61.03% improvement) and 0.0094 S cm−1 (40.87% improvement), respectively. These improvements indicate that Model II-mini effectively mitigated overfitting, a problem commonly encountered in high-dimensional ANNs, by removing noisy or redundant variables while retaining essential chemical information. The reduced input dimensionality also enabled a corresponding reduction in model complexity, with Model II-mini employing approximately one-tenth the number of neurons per hidden layer compared to Models I and II (Tables S1, S2 and S5). This architectural simplification improved the bias-variance tradeoff by reducing variance-related generalization error while maintaining sufficient model capacity to capture the underlying structure–property relationships, in accordance with the bias-variance tradeoff principle.112 As such, a dual-purpose highly predictive model capable of accurately estimating the anion conductivities of AEM polymers, both in their freshly synthesized state and throughout the course of alkaline stability testing, was obtained. Overall, the feature selection workflow presented herein not only sharpened the model focus on chemically meaningful trends, but also improved its generalization performance, capturing the behavior of AEM polymers.
 | ||
| Fig. 3 Predicted vs. actual anion conductivity plots from the (a) training–validation and (b) test prediction processes of Model II-mini. | ||
It is important to note that the model operates in descriptor space rather than structural space. Mordred descriptors provide an algorithmic mapping from molecular structure to a vector of physicochemical and topological features, enabling evaluation of candidates whose exact monomer identities were not observed during training. Prediction reliability is governed by the applicability domain in the 67-dimensional descriptor space: candidates whose descriptor vectors lie within or near the training distribution are expected to yield more reliable predictions, while those substantially outside the training distribution should be interpreted with appropriate caution. To characterize generalization behavior in the model feature space, the relationship between k-nearest-neighbor (k-NN) distance (standardized 67D space, k = 5 (default), Euclidean) and prediction error was evaluated for the held-out test polymers (Fig. S3 and Table S7). Following classical k-NN applicability-domain practice, high-quantile distance references (e.g., the 95th percentile of the training-set internal k-NN distance distribution) are often used as conservative indicators of sparse regions in descriptor space.113,114 The analysis indicates that the percentile rank is conservative for this dataset, where the majority of test data points exceed the 95th percentile while the model retains strong predictive accuracy (RMSE = 0.0136 S cm−1), whereas the absolute k-NN distance provides a more informative continuous indicator of prediction reliability (Fig. S3).
Prediction logic evaluations using SHAP
SHAP was successfully applied to Model II-mini (Fig. 4a). The resulting SHAP summary plot ranks the input variables by their overall impact, wherein each dot represents the SHAP value of an explanatory variable for a specific sample, and the dot color reflects the original explanatory variable (feature) value. Among the top 20 most influential variables identified by SHAP, two were experimental in origin: the anion conductivity measurement temperature and IEC. Both are well-established factors influencing conductivity, with higher temperatures generally enhancing ion mobility and increased IEC improving conductivity up to a point, beyond which excessive swelling may compromise AEM membrane stability. The prominence of these variables indicates that Model II-mini effectively captured key chemical principles. The remaining top variables were molecular descriptors, and the impacts of the top three important variables (AATSC5i_A, AATS8v_A, and SRW06_A) on the predicted anion conductivity were examined (Tables 2 and S8). AATSC5i_A is a descriptor that captures the electron distribution by averaging ionization potentials between atoms that are five bonds apart in the block A monomer. A high AATSC5i_A value indicates aromatic-rich structures containing groups with a high ionization potential or electron-withdrawing substituents,115 as seen in partially fluorinated poly(arylene ether)s (e.g., QPE-PYR-X5Y3,116 highest AATSC5i_A = 0.3724, anion conductivity 0.00126 S cm−1 at 80 °C). AATS8v_A reflects the bulkiness by averaging the van der Waals volumes of the atoms that are eight bonds apart. High AATS8v_A values are typically observed in polymers with bulky substituents that are consistently spaced by eight-bond intervals, such as poly(arylene ether sulfone) (e.g., PAES-Q-80,117 highest AATS8v_A = 216.7770, anion conductivity 0.0679 S cm−1 at 80 °C). SRW06_A captures the cyclicity and graph complexity by counting six-step closed walks within block A. High SRW06_A values originate from block A monomers rich in carbocyclic rings, such as fluorene-based poly(arylene ether sulfone) (e.g., TrimPES-0.4,118 highest SRW06_A = 8.6665, anion conductivity 0.121 S cm−1 at 80 °C). Higher values of these descriptors have larger impacts on the predicted values but not necessarily higher predicted anion conductivities. For example, the highest-performing AEM polymer (C-QPx1y0, conductivity 0.319 S cm−1 at 80 °C)119 exhibits AATSC5i_A, AATS8v_A, and SRW06_A values of 0.004205, 117.2667, and 6.4281, respectively, falling between the second and third quartiles, the first and second quartiles, and below the first quartile, respectively (Table 3). In contrast, PPO-7Q-0.8,120 the lowest-performing AEM polymer (conductivity 0.00115 S cm−1 at 80 °C), shows values below the first quartile (AATSC5i_A = −0.3141 and SRW06_A = 6.4938) and between the first and second quartiles (AATS8v_A = 108.0371) (Table 3). These observations suggest that high anion conductivity is not simply associated with high values of all important descriptors, highlighting the complex interplay between molecular descriptors and anion conducting properties. Across the dataset (Fig. S4a and c), high conductivity was generally associated with a balanced combination of these descriptors rather than extreme values in any single descriptor, as signified by C-QPx1y0, which exhibits a balance between the top three descriptors. This highlights the complex connection between structural features and ion transport, along with the importance of analyzing inter-descriptor relationships to guide AEM design, favoring balance over the maximization of individual descriptors.| Descriptor | Type | Property | Lag | Chemical meaning | High value trend | Low value trend |
|---|---|---|---|---|---|---|
| AATSC5i_A | Averaged & centered autocorrelation; block A | Ionization potential | 5 | Electron distribution across 5-bond distances | Multiple atoms with higher ionization potentials than carbon | Mostly constituted of carbon atoms |
| AATS8v_A | Averaged autocorrelation; block A | van der Waals volume | 8 | Long-range atomic size spread | Bulky, branched, and multiple heteroatom chains | Compact, limited branching, homogeneous chains |
| SRW06_A | Self-returning walk count; block A | Topological structure | 6 | Cyclicity and local structural complexity | Rigid, cyclic motifs | Simple linear chains |
| Descriptor | First quartile | Second quartile | Third quartile | Minimum value | Maximum value |
|---|---|---|---|---|---|
| AATSC5i_A | −0.2474 | −0.02510 | 0.03686 | −0.9884 | 0.3724 |
| AATS8v_A | 106.8337 | 133.4384 | 168.0280 | 31.0837 | 216.7770 |
| SRW06_A | 6.5236 | 7.0639 | 7.6625 | 5.7137 | 8.6665 |
Three pairs were formed from the top three descriptors, namely AATSC5i_A/AATS8v_A, AATSC5i_A/SRW06_A, and AATS8v_A/SRW06_A. To unravel the correlations between the descriptors in each pair, the SHAP value of each descriptor was used to calculate the Pearson correlation coefficient,121 r (Table 4), via the following equation:121
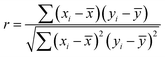 | (3) |
![[x with combining macron]](https://www.rsc.org/images/entities/i_char_0078_0304.gif) and ȳ are the means of the x and y values, respectively. Although the use of the SHAP value to calculate the correlation coefficient has been reported in the field of soil salinity analysis,122 it was used solely for feature selection by utilizing the target–property relationship between the model prediction and the input variables formed by SHAP. Here, the correlation coefficients were used to guide whether the pairs formed are of chemical significance, wherein values >0.7 reflect a high correlation, 0.5–0.69 indicate a moderate correlation, and <0.49 indicate a low (or zero) correlation.123 Accordingly, the AATS8v_A/SRW06_A pair represents the most correlated pair (correlation coefficient = 0.7688; Table 4), whereas the AATSC5i_A/SRW06_A pair is the least correlated pair (correlation coefficient = 0.2340); the AATSC5i_A/AATS8v_A pair is moderately correlated (correlation coefficient = 0.5329) (Fig. S4d–f). Considering the individual meanings of the descriptors, the moderate AATSC5i_A/AATS8v_A correlation occurs because both components consider the heteroatoms present in the polymer structure, which either lead to a higher ionization potential or a larger van der Waals volume. These descriptors are not highly correlated because it is possible to increase the ionization energy without significantly affecting the van der Waals volume, such as through adding heteroatoms to aliphatic chains.124 The low correlation observed for AATSC5i_A/SRW06_A can be attributed to the fact that AATSC5i_A focuses on heteroatoms, while SRW06_A only concentrates on six-step closed walks, thereby ignoring the heterogeneity within the polymer structure.125 This correlation decreased further to 0.1766 when recalculated using only AEM polymers with anion conductivities ≥0.1 S cm−1 at 80 °C (Table 4), and the AATSC5i_A/SRW06_A pair was therefore excluded from subsequent analysis. The high correlation for AATS8v_A/SRW06_A occurs since both descriptors evaluate the polymer bulkiness, i.e., AATS8v_A through the atomic van der Waals volume, and SRW06_A through cyclic structures. Their correlation is strong but remains below 0.8, likely because SRW06_A focuses solely on cyclicity and does not explicitly account for heteroatoms, a factor that influences AATS8v_A. Interestingly, correlation trends from the SHAP values mirrored those from the descriptor values, although the descriptor-based correlations were slightly higher. This difference arises because descriptor-based correlations solely reflect direct structural similarity, whereas the SHAP values capture additional nonlinear and context-dependent interactions within the model. For instance, vertically aligned clusters in the SHAP scatter plots (Fig. 4b–d), where multiple data points share identical descriptor values (x-axis) yet show differing SHAP values (y-axis), illustrate how various experimental conditions can modulate the contribution of a descriptor. Thus, SHAP-based analysis provides deeper insights into the combined influence of structural features and experimental conditions on the predicted conductivity, surpassing the interpretability provided by descriptor correlations alone. Overall, these pairwise descriptor correlations highlight complex underlying structural relationships affecting AEM conductivity, emphasizing the requirement for deeper analysis. To accurately interpret how these descriptors and descriptor pairs are influenced by polymer structures, it is crucial to further investigate their calculation mechanisms through examination of their computational algorithms, which are available in the Mordred library.65
and ȳ are the means of the x and y values, respectively. Although the use of the SHAP value to calculate the correlation coefficient has been reported in the field of soil salinity analysis,122 it was used solely for feature selection by utilizing the target–property relationship between the model prediction and the input variables formed by SHAP. Here, the correlation coefficients were used to guide whether the pairs formed are of chemical significance, wherein values >0.7 reflect a high correlation, 0.5–0.69 indicate a moderate correlation, and <0.49 indicate a low (or zero) correlation.123 Accordingly, the AATS8v_A/SRW06_A pair represents the most correlated pair (correlation coefficient = 0.7688; Table 4), whereas the AATSC5i_A/SRW06_A pair is the least correlated pair (correlation coefficient = 0.2340); the AATSC5i_A/AATS8v_A pair is moderately correlated (correlation coefficient = 0.5329) (Fig. S4d–f). Considering the individual meanings of the descriptors, the moderate AATSC5i_A/AATS8v_A correlation occurs because both components consider the heteroatoms present in the polymer structure, which either lead to a higher ionization potential or a larger van der Waals volume. These descriptors are not highly correlated because it is possible to increase the ionization energy without significantly affecting the van der Waals volume, such as through adding heteroatoms to aliphatic chains.124 The low correlation observed for AATSC5i_A/SRW06_A can be attributed to the fact that AATSC5i_A focuses on heteroatoms, while SRW06_A only concentrates on six-step closed walks, thereby ignoring the heterogeneity within the polymer structure.125 This correlation decreased further to 0.1766 when recalculated using only AEM polymers with anion conductivities ≥0.1 S cm−1 at 80 °C (Table 4), and the AATSC5i_A/SRW06_A pair was therefore excluded from subsequent analysis. The high correlation for AATS8v_A/SRW06_A occurs since both descriptors evaluate the polymer bulkiness, i.e., AATS8v_A through the atomic van der Waals volume, and SRW06_A through cyclic structures. Their correlation is strong but remains below 0.8, likely because SRW06_A focuses solely on cyclicity and does not explicitly account for heteroatoms, a factor that influences AATS8v_A. Interestingly, correlation trends from the SHAP values mirrored those from the descriptor values, although the descriptor-based correlations were slightly higher. This difference arises because descriptor-based correlations solely reflect direct structural similarity, whereas the SHAP values capture additional nonlinear and context-dependent interactions within the model. For instance, vertically aligned clusters in the SHAP scatter plots (Fig. 4b–d), where multiple data points share identical descriptor values (x-axis) yet show differing SHAP values (y-axis), illustrate how various experimental conditions can modulate the contribution of a descriptor. Thus, SHAP-based analysis provides deeper insights into the combined influence of structural features and experimental conditions on the predicted conductivity, surpassing the interpretability provided by descriptor correlations alone. Overall, these pairwise descriptor correlations highlight complex underlying structural relationships affecting AEM conductivity, emphasizing the requirement for deeper analysis. To accurately interpret how these descriptors and descriptor pairs are influenced by polymer structures, it is crucial to further investigate their calculation mechanisms through examination of their computational algorithms, which are available in the Mordred library.65
| Descriptor 1 | Descriptor 2 | Correlation coefficient | |
|---|---|---|---|
| SHAP value-based | Descriptor value-based | ||
| All data | |||
| AATSC5i_A | AATS8v_A | 0.5329 | 0.5894 |
| AATSC5i_A | SRW06_A | 0.2340 | 0.2378 |
| AATS8v_A | SRW06_A | 0.7688 | 0.7928 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||
| Anion conductivity measured at 80 °C ≥ 0.1 S cm−1 | |||
| AATSC5i_A | AATS8v_A | 0.5417 | 0.4858 |
| AATSC5i_A | SRW06_A | 0.1766 | 0.2157 |
| AATS8v_A | SRW06_A | 0.6071 | 0.6999 |
Molecular descriptor explanation using ChatGPT
In-depth examinations of the AATSC5i_A, AATS8v_A, and SRW06_A source codes were performed using ChatGPT (versions 4o and o3; accessed May 13, 2025), a chatbot program developed by OpenAI and based on the large language model (LLM) technology.126 Such LLMs provide coding assistance that is also beneficial for the chemistry field,127–129 and it would be desirable for the workflow incorporating the descriptor explanations to be constructed with minimal prior domain knowledge regarding the descriptor or the descriptor source code. It should be emphasized that ChatGPT was not employed as an autonomous source of chemical knowledge or as a decision-making agent in this study; rather, it was deliberately positioned as an assistive interpretative tool within a human-in-the-loop framework, where all generated explanations were critically evaluated, corrected where necessary, and validated by domain experts. The role of ChatGPT was specifically limited to (i) parsing and summarizing descriptor source code obtained from the Mordred library and (ii) facilitating the translation of mathematically abstract descriptor definitions into chemically intuitive language, thereby accelerating expert interpretation rather than replacing it. While LLMs such as ChatGPT are trained on broad corpora that incidentally include chemistry and materials science literature, their domain-specific reliability remains limited, and they may generate incomplete or misleading explanations if used without expert oversight.130–132 To mitigate this risk, descriptor source code, formal definitions, and domain-specific background information were explicitly provided as inputs, and all generated interpretations were cross-checked against established physicochemical principles and experimental knowledge. Using ChatGPT, the workflow is as follows:(1) Obtain source code for AATSC5i_A, AATS8v_A, and SRW06_A from Mordred (https://mordred-descriptor.github.io/documentation/master/descriptors.html).
(2) Ask ChatGPT to explain AATSC5i_A (for example) based on (i) the meaning of the descriptor from Mordred and (ii) the source code of the descriptor.
(3) Analyze the answer provided by ChatGPT based on chemical domain knowledge.
During this workflow, four representative limitations that further motivate the human-in-the-loop safeguards described above were observed. First, ChatGPT occasionally produced mechanistic-sounding narratives directly from correlational patterns (e.g., inferring specific ion-transport mechanisms from descriptor–conductivity correlations); such statements were rewritten using hypothesis-level language and cross-checked against established polymer physics principles. Second, initial outputs emphasized simple screening-threshold heuristics suggesting monotonic improvement, whereas expert analysis indicated non-monotonic tradeoffs that require a balanced optimization perspective. Third, architecturally meaningful insights, such as the structural significance of certain lag values (e.g., lag-5 and lag-8) as characteristic bond distances in AEM polymer architectures, often emerged only after explicit expert prompting, highlighting that domain knowledge is essential to formulate scientifically meaningful questions. Fourth, design suggestions required expert filtering for synthetic feasibility and chemical stability. Importantly, all quantitative results reported herein were computed and verified independently by the authors; ChatGPT outputs were used only to assist with drafting qualitative interpretations. Consistent with this cautious framing, evaluations of GPT models on chemistry tasks report improved accuracy in newer versions but persistent computational and interpretive errors, indicating that careful human oversight remains necessary even as model versions improve.133 Table S9 provides concrete examples of this refinement process. When used within this validated workflow, providing descriptor source code, definitions, and domain-specific background (Fig. 5) enabled ChatGPT to generate more detailed draft explanations for key individual descriptors (Table 5) and descriptor pairs (Table 6), which facilitated clearer visualization and discussion of AATSC5i_A and AATS8v_A. In contrast, for SRW06_A and its pairs, the added value was limited, consistent with SRW06_A being a comparatively straightforward ring-count/topology descriptor largely reflecting the presence and number of six-membered rings. These results indicate that, under expert oversight, ChatGPT can accelerate descriptor interpretation by improving the clarity and completeness of draft explanations. ChatGPT stated that the AATSC5i_A/AATS8v_A pair describes how regularly high-ionization potentials and bulky atoms coexist, and how they are periodically spaced along the main/side-chain; however, be aware that the presence of long, bulky alkyl chains does not affect the ionization potential (AATSC5i_A) but increases the bulk (AATS8v_A). In particular, the five- and eight-bond distances of the AATSC5i_A/AATS8v_A pair were identified by Model II-mini because five represents the shortest graph distance that spans an entire para-substituted phenyl ring, while eight is the major tether length from the main-chain structure to the quaternary ammonium ion, as well as the para-to-para distance between cationic groups on adjacent phenyl groups in a biphenyl or poly(aryl ether ketone)-like main-chain. This explanation is coherent with the database content, as both of these constituted the majority of the AEM polymers in the database.71
 | ||
| Fig. 5 Example for the prompt used in this study and the actual answer provided by ChatGPT-o3. The actual answer consists of six sections; only the first section is shown here. | ||
| Descriptor | Factors increasing the descriptor value | Factors decreasing the descriptor value |
|---|---|---|
| AATSC5i_A | Majority of atoms lie on the same side of the ionization-potential mean, giving a positive covariance | Alternation of high- and low-ionization potential atoms every five bonds, yielding negative covariance |
| Similar high-ionization potential atoms (O, F, and quaternary-N) arranged exactly five bonds apart | A high-ionization-potential heteroatom opposite a low-ionization-potential carbon at a five-bond distance | |
| Electron-withdrawing groups (F, CF3, SO2, and quaternary ammonium) installed so their centers are separated by five σ-bonds | Donor/acceptor alternation or long aliphatic tethers that bring dissimilar ionization potential atoms together at a five-bond distance | |
| Long, aromatic-rich A block that provides numerous five-bond pathways between like heteroatoms | Very short or highly branched blocks that limit or disrupt regular five-bond pairings | |
| AATS8v_A | Many bulky atoms (Cl, Br, P, quaternary-N substituents, long alkyls, and aromatic rings) | Dominance of small atoms (H, light heteroatoms) in the graph |
| Regular spacing of those bulky atoms so that the graph distance between the atoms is exactly 8 bonds (e.g. para–para substituted aromatics; tether lengths of 7 σ-bonds) | Bulky atoms clustered closer than 8 bonds or significantly further apart; few 8-bond paths | |
| A long linear or lightly branched backbone that preserves many distinct 8-bond paths | Very short, highly cross-linked or heavily branched repeating units that shorten most path lengths | |
| Repetition of side-chains of similar size yielding numerous equivalent paths | Alternation between large and very small atoms (large–small pairs give moderate products) | |
| SRW06_A | Six-membered rings (benzene, piperidine, and cyclohexane): each ring supplies many 6-step closed walks (each way round the ring plus retraced variants) | Strictly linear chains with no rings or long branches have very few closed walks |
| Fused aromatic systems or multiple rings in one repeating unit: walk counts combine additively and multiplicatively | Very short monomers (≤5 heavy atoms) cannot support 6-step cycles at all, giving SRW06 = ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 = 0 1 = 0 |
|
| Dense branching that brings atoms back to the origin in six steps (e.g., 1,3,5-trisubstituted phenyl) | Extensive spacer segments between functional moieties push closed circuits beyond 6 steps, and so they do not contribute here | |
| Heteroatom substitutions inside the rings (N, O, and S) do not change the walk count itself but may appear more than once because each heteroatom is a graph node |
| Pair | Factors increasing both descriptor values | Factors decreasing both descriptor values | Factors promoting the opposite effect (one up/one down) |
|---|---|---|---|
| AATSC5i_A & AATS8v_A | Two quaternary-N or O/F atoms separated by exactly 5 bonds and again encountered (through backbone repetition) at an 8-bond spacing | Aliphatic chains with mixed C and heteroatoms → low IP covariance and small van der Waals products | Add bulky tert-butyl carbons (↑ AATS8v_A) without heteroatoms (↔ AATSC5i_A) |
| Para-di-substituted phenyl where ammonium head groups such as [N(CH3)4]+ sit 5 bonds apart (around the ring) and 8 bonds apart (across adjacent rings) | Very short/branched units that break 8-bond paths | Introduce alternating donor/acceptor heteroatoms (↓ AATSC5i_A vs. ↔/↑ AATS8v_A) | |
| AATS8v_A & SRW06_A | Fused or multi-ring systems with bulky substituents (biphenyl, naphthalene, and fluorene) | Linear aliphatics with few side-groups | Add branching bulky tert-butyls along a flexible chain (↑ AATS8v_A) but no rings (SRW06_A low) |
From the above findings, AEM polymer design guidelines can be derived: (i) including aromaticity, preferably benzene with its hydrogen substituted with other atoms/groups that are more electron-withdrawing (AATSC5i_A) and/or as biphenyl (AATS8v_A), (ii) having a side-chain length of eight bonds from the connecting point between main and side-chain to the ammonium site (AATS8v_A), and (iii) avoiding pure alkyl chains of more than 4 units by introducing electron-withdrawing heteroatoms every 4 carbons would help to inflate AATSC5i_A, positively affecting ionization potential. These guidelines conform with the latest understandings that biphenyl moieties or long side-chains enhance the durability (by avoiding aryl-ether cleavage reactions) and phase separation,99 while also providing a more quantitative guideline on the bond interval that is required along a long alkyl side-chain to maximize water and ion transport.104,134,135
While several extracted motifs align with the established AEM literature, the novelty of this work lies in the traceable, descriptor-to-guideline workflow, including the pairwise correlation analysis of SHAP-attributed contributions, that quantitatively recovers and refines these principles from experimental datasets without hard-coding material-specific design rules. Based on these design guidelines, a conceptual block A monomer structure was designed for an AEM polymer, since both AATSC5i_A and AATS8v_A are descriptors of block A (Fig. 6, blue background). It should be noted that the polymer structures created by experts and shown in Fig. 6 are presented as conceptual design examples intended to demonstrate how descriptor-level insights can be translated into chemically plausible polymer architectures, rather than as claims of optimized or synthesis-ready materials. To assess whether the conceptual designs fall within the applicability domain of Model II-mini, k-NN distances were computed in the standardized 67-dimensional descriptor space used by Model II-mini (k = 5, Euclidean; Table S7). Although percentile ranking relative to the training-dataset internal distance distribution places these designs in the extreme tail (≥99.49th percentile), percentile references are conservative for this dataset, and the absolute k-NN distance provides a more informative continuous novelty indicator (Fig. S3 and Table S7). The conceptual designs exhibit k-NN distances of 4.71–6.90, placing them in the same near-neighbor distance band associated with strong predictive performance (RMSE = 0.01427–0.00192 S cm−1) for multiple held-out test polymers with comparable distances. For example, AEM polymer 3 (k-NN distance = 4.71) is comparable to test polymer H22C9N136 (4.62, RMSE = 0.00357 S cm−1), while AEM polymers 1 and 4 (k-NN distance ≈ 6.8–6.9) are comparable to PmTDMP137 (6.86, RMSE = 0.00499 S cm−1). This analysis suggests that the predictions are reasonably supported within the learned descriptor space. Nevertheless, such predictions remain extrapolative to a certain degree and require experimental validation to confirm predicted properties, particularly given that outliers such as PPO-PipOH138 (RMSE = 0.02940 S cm−1 despite moderate k-NN distance) demonstrate that proximity to training data is necessary but not sufficient for accurate prediction. Experimentalists can apply this approach by calculating Mordred descriptors for candidate monomers and assessing their alignment with the identified optimal ranges (Tables 2 and 3) and applicability domain of Model II-mini, thereby using the framework as a quantitative screening filter prior to synthesis.
Importantly, this descriptor-guided framework can also be applied in the inverse direction: given a specific monomer or set of candidate monomers, their descriptor values can be evaluated against the SHAP analysis-identified optimal ranges to guide decisions on side-chain modification, block composition, and functionalization strategies, enabling rational polymer design under realistic synthetic constraints. Without specific recommendations regarding heteroatom spacers or anion-conducting moieties, ether linkages and trimethylammonium ions were selected, since the hydrophilic ether linkage can potentially mitigate Hofmann elimination at the C3 methylene group adjacent to the trimethylammonium site by contributing to the conservation of high OH− hydration.139,140 These functional groups have well-established synthetic accessibility in polymer and/or AEM research.101,141,142 Although a block A homopolymer may appear promising in terms of its anion conductivity, a fully functionalized block A-only AEM polymer would possess an excessively high IEC, raising concerns regarding water solubility and membrane swelling. To address potential over-swelling, a B-block segment that shares the main-chain structure of block A but lacks the side-chain was introduced, thereby converting the homopolymer into a block copolymer with a reduced functionalization density (denoted as AEM polymer 1). The block A/block B molar ratio was set at 8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 as a representative example; alternative ratios can be explored using the same framework to optimize monomer distribution for specific application requirements. AEM polymer 1 exhibited AATSC5i_A and AATS8v_A values surpassing the third quartile and falling between the second and third quartiles, respectively. Subsequently, the performance of AEM polymer 1 was evaluated using Model II-mini. The anion conductivity was predicted at 80 °C, while the alkaline stability was assessed under 1 M NaOH conditions at 60 °C after 100 days. The block A/block B molar ratio was set at 8
2 as a representative example; alternative ratios can be explored using the same framework to optimize monomer distribution for specific application requirements. AEM polymer 1 exhibited AATSC5i_A and AATS8v_A values surpassing the third quartile and falling between the second and third quartiles, respectively. Subsequently, the performance of AEM polymer 1 was evaluated using Model II-mini. The anion conductivity was predicted at 80 °C, while the alkaline stability was assessed under 1 M NaOH conditions at 60 °C after 100 days. The block A/block B molar ratio was set at 8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2. AEM polymer 1 exhibited a predicted anion conductivity exceeding 0.1 S cm−1 at 80 °C and retained 67.94% of that conductivity (0.07639 S cm−1) after 100 days of alkaline exposure (1 M NaOH, 60 °C) (Table 7 and Fig. 6, purple background). Detailed predicted conductivity profiles for AEM polymers 1 and 3, including temperature-dependent conductivity and time-resolved degradation trajectories over 0–100 days of alkaline exposure, are provided in Table S10. Both polymers exhibit a gradual decline in predicted conductivity with increasing exposure time, with AEM polymer 3 retaining 71.33% of its initial conductivity at 80 °C after 100 days, comparable to AEM polymer 1 (67.94%). To contextualize this prediction, the retention is moderate compared with some highly durable AEMs reported under comparable alkaline-aging protocols, while remaining chemically plausible given the functional motifs adopted in the conceptual design.99,101,103 Importantly, these values are model predictions intended to guide prioritization rather than to replace experimental validation, and they suggest that the extracted design guidelines can yield realistic candidates for subsequent synthesis-focused studies. Three additional block A variants were designed, with their block B structures set according to their main-chain structures but without a side-chain. Their block A/block B ratio was defined as 8
2. AEM polymer 1 exhibited a predicted anion conductivity exceeding 0.1 S cm−1 at 80 °C and retained 67.94% of that conductivity (0.07639 S cm−1) after 100 days of alkaline exposure (1 M NaOH, 60 °C) (Table 7 and Fig. 6, purple background). Detailed predicted conductivity profiles for AEM polymers 1 and 3, including temperature-dependent conductivity and time-resolved degradation trajectories over 0–100 days of alkaline exposure, are provided in Table S10. Both polymers exhibit a gradual decline in predicted conductivity with increasing exposure time, with AEM polymer 3 retaining 71.33% of its initial conductivity at 80 °C after 100 days, comparable to AEM polymer 1 (67.94%). To contextualize this prediction, the retention is moderate compared with some highly durable AEMs reported under comparable alkaline-aging protocols, while remaining chemically plausible given the functional motifs adopted in the conceptual design.99,101,103 Importantly, these values are model predictions intended to guide prioritization rather than to replace experimental validation, and they suggest that the extracted design guidelines can yield realistic candidates for subsequent synthesis-focused studies. Three additional block A variants were designed, with their block B structures set according to their main-chain structures but without a side-chain. Their block A/block B ratio was defined as 8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, and the resulting polymers were named AEM polymers 2–4 (Fig. 6, gray background). The same Model II-mini was used to predict the anion conductivities at 80 °C of AEM polymers 2–4. Among them, the side-chain of AEM polymer 2 was maintained at five bond lengths to fulfill the criteria of AATSC5i_A, but due to the difference in ionization potentials between carbon and ammonium, the obtained AATSC5i_A value was the lowest amongst all four conceptual structures (Table 7 and Fig. 6, gray background). None surpassed the performance of the initial concept, with the highest conductivity (0.1024 S cm−1, AEM polymer 3) being 0.01 S cm−1 lower than the initial design (Table 7 and Fig. 6, gray background). To benchmark these predictions against the broader experimental landscape, it is noted that among the 326 AEM polymers in the training database with conductivity values reported at 80 °C, only 47 (14.42%) exhibit anion conductivities ≥0.1 S cm−1, a target often used as a practical performance criterion for AEMFC/AEMWE-relevant conditions.143 Within the four illustrative conceptual designs proposed using the extracted guidelines, two exceed this criterion (AEM polymers 1 and 3). While this outcome is not presented as a statistical claim given the small sample size, it illustrates how the framework can enrich promising candidates prior to committing synthetic resources, thereby improving the efficiency of early-stage AEM polymer development.
2, and the resulting polymers were named AEM polymers 2–4 (Fig. 6, gray background). The same Model II-mini was used to predict the anion conductivities at 80 °C of AEM polymers 2–4. Among them, the side-chain of AEM polymer 2 was maintained at five bond lengths to fulfill the criteria of AATSC5i_A, but due to the difference in ionization potentials between carbon and ammonium, the obtained AATSC5i_A value was the lowest amongst all four conceptual structures (Table 7 and Fig. 6, gray background). None surpassed the performance of the initial concept, with the highest conductivity (0.1024 S cm−1, AEM polymer 3) being 0.01 S cm−1 lower than the initial design (Table 7 and Fig. 6, gray background). To benchmark these predictions against the broader experimental landscape, it is noted that among the 326 AEM polymers in the training database with conductivity values reported at 80 °C, only 47 (14.42%) exhibit anion conductivities ≥0.1 S cm−1, a target often used as a practical performance criterion for AEMFC/AEMWE-relevant conditions.143 Within the four illustrative conceptual designs proposed using the extracted guidelines, two exceed this criterion (AEM polymers 1 and 3). While this outcome is not presented as a statistical claim given the small sample size, it illustrates how the framework can enrich promising candidates prior to committing synthetic resources, thereby improving the efficiency of early-stage AEM polymer development.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 as an example
2 as an example
| AEM polymer no. | AATSC5i_A | AATS8v_A | Predicted anion conductivity/S cm−1 | Predicted alkaline stability/S cm−1 |
|---|---|---|---|---|
| 1 | 0.07185 | 139.1117 | 0.1124 | 0.07639 (67.94%) |
| 2 | −0.06804 | 127.0347 | 0.0741 | — |
| 3 | 0.09846 | 161.8823 | 0.1024 | — |
| 4 | 0.1087 | 162.3189 | 0.0727 | — |
Overall, these results demonstrated the potential of systematically feeding back interpretations from combined SHAP-based, ChatGPT-assisted descriptor analysis into the design–predict–select loop of AEM polymer development. By referring to the guidelines obtained through ChatGPT assisted interpretation of important molecular descriptors, candidate structures can be rationally designed and then screened in silico using the trained ANN model to predict both initial anion conductivity and long-term alkaline stability. Only those candidates falling within the high-performance descriptor range and meeting predictive performance criteria, such as ≥0.1 S cm−1 conductivity at 80 °C, would be prioritized for synthesis. This design-to-prediction workflow supports efficient narrowing of the experimental search space, accelerates identification of promising AEM polymers, and reduces resource consumption in early-stage polymer development. Though applied only to AEM polymers here, this workflow should be applicable to any material wherein molecular descriptors are used to represent the chemical structure. Such generalizability positions this workflow as a useful tool for expanding XAI applications across diverse materials systems.
Conclusions
This study demonstrates a systematic and transparent ML/AI framework that combines statistical (mRMR) and explainable ML (permutation importance) feature selection techniques for analyzing high-dimensional polymer datasets within an ANN framework. The two-step dimensionality reduction, from thousands of descriptors down to 67 key features, enabled efficient SHAP-based model explanation while significantly enhancing the predictive accuracy of the ANN for unseen AEM polymer anion conductivity (+40.87%). SHAP analysis effectively identified the most impactful descriptors, allowing both manual and AI-assisted interpretation (via ChatGPT) to reveal chemically meaningful structure–property relationships. Moreover, this novel pairwise correlation analysis, together with the utilization of ChatGPT, uncovered previously hidden descriptor interactions, providing richer insights and clearer guidelines for polymer design than traditional single-variable analysis alone. Four illustrative conceptual AEM polymer designs were proposed based on the extracted guidelines, and two were predicted to exhibit anion conductivities ≥0.1 S cm−1 at 80 °C, highlighting how the framework can prioritize promising candidates for subsequent experimental validation. Through the proposed method, this study successfully demonstrated the achievement of orchestration between XAI, ChatGPT-assisted molecular descriptor interpretation, and human expertise, transforming complex model outputs into chemically interpretable design insights that can guide rational monomer and polymer design prior to synthesis, whether starting from novel conceptual structures or from experimentally accessible monomers. By enabling descriptor-guided screening and prioritization under realistic synthetic constraints, the framework helps experimentalists narrow the candidate space and allocate synthesis or testing resources more efficiently, complementing, rather than replacing, experimental validation. Furthermore, because the model operates in descriptor space rather than structural space, the framework supports evaluation of near-neighbor chemical space, such as new combinations or modifications of established motifs, while transparently signaling via applicability-domain metrics when predictions extend beyond the training distribution and should be interpreted with appropriate caution. While this method proved effective for AEM polymers and ANN architectures, limitations remain. The current analysis was restricted to pairwise interactions and three descriptors; expanding this to include higher-order interactions and broader chemical classes could further improve insight and generalizability, while simultaneously increasing the number of relationships to be analyzed. Future work may explore integrating this framework with generative models or transfer learning to enhance its applicability to unexplored chemical spaces. More broadly, the present workflow demonstrates how experimentally derived polymer datasets can be translated into quantitative descriptor–property relationships, enabling materials scientists to examine, refine, and reproduce design heuristics within a data-driven framework. Overall, the proposed framework serves as a robust template for extending XAI to complex materials systems, as demonstrated in its application to AEM polymers, ultimately bridging predictive modeling with scientific understanding to accelerate material discovery.Author contributions
The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript. Yin Kan Phua: conceptualization, investigation, data curation, and writing – original draft. Nana Terasoba: data curation. Manabu Tanaka: data curation, writing – review, and editing. Tsuyohiko Fujigaya: supervision, writing – review, and editing. Koichiro Kato: conceptualization, supervision, writing – review, and editing.Conflicts of interest
There are no conflicts to declare.Data availability
The data underlying this study, with the representative snapshot is provided in Table S11, are openly available in the GitHub repository of our previous study (https://doi.org/10.1002/celc.202400252) at https://github.com/KatoGroup-AppChem-KU/AEM_Unsupervised-ML.71 The Mordred descriptor library can be found on GitHub at https://github.com/mordred-descriptor/mordred. The scikit-learn library can be found at https://github.com/scikit-learn/scikit-learn. The codes used to create Models I, II, and II-mini are openly available in GitHub at https://github.com/KatoGroup-AppChem-KU/AEM_Explainable-ML_Workflow.Supplementary information (SI): additional model construction and analysis workflow details; optimized hyperparameter values; complete tables with brief explanations of top 64 important variables from ELI5 and top 20 important variables from SHAP; figure of descriptor value vs. anion conductivity and descriptor value vs. descriptor value, for the top 3 important variables; code availability for this workflow. See DOI: https://doi.org/10.1039/d5ta06120b.
Acknowledgements
This study was supported by the Japan Science and Technology Agency (JST) (ACT-X, Grant No. JPMJAX22AF), the Establishment of University Fellowships towards the Creation of Science Technology Innovation (grant no. JPMJFS2132), JST SPRING (grant no. JPMJSP2136), an “Engineering Research for Pioneering of a New Field” grant provided by the Faculty of Engineering, Kyushu University, JSPS KAKENHI (grant no. JP23H02027), the Data Creation and Utilization-Type Material Research and Development Project (Grant Number JPMXP1122714694) granted by the Ministry of Education, Culture, Sports, Science, and Technology (MEXT), and the “TMU Research Fund for Young Scientists” provided by Tokyo Metropolitan University. The authors thank the Robert T. Huang Entrepreneurship Center of Kyushu University (QREC) for supporting the project via an “Academic Challenge 2021” grant.References
- I. H. Sarker, SN Comput. Sci., 2021, 2, 160 CrossRef PubMed.
- H. Wang, T. Fu, Y. Du, W. Gao, K. Huang, Z. Liu, P. Chandak, S. Liu, P. Van Katwyk, A. Deac, A. Anandkumar, K. Bergen, C. P. Gomes, S. Ho, P. Kohli, J. Lasenby, J. Leskovec, T. Y. Liu, A. Manrai, D. Marks, B. Ramsundar, L. Song, J. Sun, J. Tang, P. Veličković, M. Welling, L. Zhang, C. W. Coley, Y. Bengio and M. Zitnik, Nature, 2023, 620, 47–60 CrossRef CAS PubMed.
- A. E. Comesana, T. T. Huntington, C. D. Scown, K. E. Niemeyer and V. H. Rapp, Fuel, 2022, 321, 123836 CrossRef CAS.
- M. R. S. A. Janjua, A. Irfan, M. Hussien, M. Ali, M. Saqib and M. Sulaman, Energy Technol., 2022, 10, 2200019 CrossRef CAS.
- G. Armeli, J. H. Peters and T. Koop, ACS Omega, 2023, 8, 12298–12309 CrossRef CAS PubMed.
- Z. Chen, B. Yang, N. Song, T. Chen, Q. Zhang, C. Li, J. Jiang, T. Chen, Y. Yu and L. X. Liu, Chem. Eng. J., 2023, 455, 140547 CrossRef CAS.
- A. Mahmood, Y. Sandali and J. L. Wang, Phys. Chem. Chem. Phys., 2023, 25, 10417–10426 RSC.
- V. Besel, M. Todorović, T. Kurtén, H. Vehkamäki and P. Rinke, J. Aerosol Sci., 2024, 179, 106375 CrossRef CAS.
- M. Li, C. R. Zhang, M. L. Zhang, J. J. Gong, X. M. Liu, Y. H. Chen, Z. J. Liu, Y. Z. Wu and H. S. Chen, Phys. Status Solidi A, 2024, 221, 2400008 CrossRef CAS.
- P. Gao, D. Kochan, Y.-H. Tang, X. Yang and E. G. Saldanha, J. Power Sources, 2025, 629, 236035 CrossRef CAS.
- R. Santonocito, A. Cavallaro, A. Pappalardo, R. Puglisi, A. Marano, M. Andolina, N. Tuccitto and G. Trusso Sfrazzetto, Biosens. Bioelectron., 2025, 270, 116986 CrossRef CAS PubMed.
- C. Kuenneth, W. Schertzer and R. Ramprasad, Macromolecules, 2021, 54, 5957–5961 CrossRef CAS.
- M. Aldeghi and C. W. Coley, Chem. Sci., 2022, 13, 10486–10498 RSC.
- Z. Zhang, Z. Jiao, R. Shen, P. Song and Q. Wang, ACS Appl. Eng. Mater., 2022, 1, 596–605 CrossRef.
- S. S. Shukla, C. Kuenneth and R. Ramprasad, MRS Bull., 2023, 49, 17–24 CrossRef.
- Y. Zhang, S. Shi, Y. Lu, R. Qin, X. Zhang, J. Xu and B. Chen, Polym. Compos., 2024, 45, 6914–6932 CrossRef CAS.
- R. Pétuya, S. Durdy, D. Antypov, M. W. Gaultois, N. G. Berry, G. R. Darling, A. P. Katsoulidis, M. S. Dyer and M. J. Rosseinsky, Angew Chem. Int. Ed. Engl., 2022, 61, e202114573 CrossRef PubMed.
- X. Bai, Z. Shi, H. Xia, S. Li, Z. Liu, H. Liang, Z. Liu, B. Wang and Z. Qiao, Chem. Eng. J., 2022, 446, 136783 CrossRef CAS.
- A. Nandy, S. Yue, C. Oh, C. Duan, G. G. Terrones, Y. G. Chung and H. J. Kulik, Matter, 2023, 6, 1585–1603 CrossRef CAS.
- Y. Wang, Z. J. Jiang, D. R. Wang, W. Lu and D. Li, J. Am. Chem. Soc., 2024, 146, 6955–6961 CrossRef CAS PubMed.
- Y. Zhuo, A. Mansouri Tehrani and J. Brgoch, J. Phys. Chem. Lett., 2018, 9, 1668–1673 CrossRef CAS PubMed.
- S. M. Mastelini, D. R. Cassar, E. Alcobaça, T. Botari, A. C.P.L.F. de Carvalho and E. D. Zanotto, Acta Mater., 2022, 240, 118302 CrossRef CAS.
- Y. Yan, Z. Pei, M. C. Gao, S. Misture and K. Wang, Acta Mater., 2023, 253, 118955 CrossRef CAS.
- X. Wan, W. Yu, A. Wang, X. Wang, J. Robertson, Z. Zhang and Y. Guo, ACS Sens., 2023, 8, 2319–2330 CrossRef CAS PubMed.
- N. J. Szymanski, S. Fu, E. Persson and G. Ceder, npj Comput. Mater., 2024, 10, 1–9 Search PubMed.
- Y. Hirabayashi, H. Iga, H. Ogawa, S. Tokuta, Y. Shimada and A. Yamamoto, npj Comput. Mater., 2024, 10, 1–10 Search PubMed.
- J. Kim, J. Jung, S. Kim and S. Han, Comput. Mater. Sci., 2024, 234, 112783 CrossRef CAS.
- K. Yamakoshi, Y. Ohno, K. Kutsukake, T. Kojima, T. Yokoi, H. Yoshida, H. Tanaka, X. Liu, H. Kudo and N. Usami, Adv. Mater., 2024, 36, e2308599 CrossRef PubMed.
- R. Gómez-Bombarelli, J. Aguilera-Iparraguirre, T. D. Hirzel, D. Duvenaud, D. Maclaurin, M. A. Blood-Forsythe, H. S. Chae, M. Einzinger, D. G. Ha, T. Wu, G. Markopoulos, S. Jeon, H. Kang, H. Miyazaki, M. Numata, S. Kim, W. Huang, S. I. Hong, M. Baldo, R. P. Adams and A. Aspuru-Guzik, Nat. Mater., 2016, 15, 1120–1127 CrossRef PubMed.
- E. Kim, K. Huang, S. Jegelka and E. Olivetti, npj Comput. Mater., 2017, 3, 1–9 CAS.
- A. Merchant, S. Batzner, S. S. Schoenholz, M. Aykol, G. Cheon and E. D. Cubuk, Nature, 2023, 624, 80–85 CrossRef CAS PubMed.
- J. J. Hopfield, Proc. Natl. Acad. Sci. U. S. A., 1982, 79, 2554–2558 CrossRef CAS PubMed.
- D. Ackley, G. Hinton and T. Sejnowski, Cogn. Sci., 1985, 9, 147–169 Search PubMed.
- N. Kriegeskorte and T. Golan, Curr. Biol., 2019, 29, R231–R236 CrossRef PubMed.
- J. Park and I. W. Sandberg, Neural Comput., 1991, 3, 246–257 CrossRef CAS PubMed.
- J. Park and I. W. Sandberg, Neural Comput., 1993, 5, 305–316 CrossRef.
- A. Chung Tsoi and F. Scarselli, Neural Netw., 1998, 11, 15–37 CrossRef PubMed.
- T. Hengl, M. Nussbaum, M. N. Wright, G. B. M. Heuvelink and B. Gräler, PeerJ, 2018, 6, e5518 CrossRef PubMed.
- B. Takoutsing and G. B. M. Heuvelink, Geoderma, 2022, 428, 116192 CrossRef.
- G. Buriticá and S. Engelke, arXiv, 2024, preprint, arXiv.2410.23246, DOI:10.48550/arXiv.2410.23246.
- G. Montavon, W. Samek and K.-R. Müller, Digit. Signal Process., 2018, 73, 1–15 CrossRef.
- S. Tan, K. C. Sim and M. Gales, presented in part at the 2015 IEEE Workshop on Automatic Speech Recognition and Understanding, Scottsdale, AZ, 2015 Search PubMed.
- L. Breiman, Mach. Learn., 2001, 45, 5–32 CrossRef.
- M. T. Ribeiro, S. Singh and C. Guestrin, presented in part at the Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, 2016, pp. 1135–1144 Search PubMed.
- S. M. Lundberg and S.-I. Lee, presented in part at the Proceedings of the 31st International Conference on Neural Information Processing Systems, Long Beach, CA, 2017, pp. 4768–4777 Search PubMed.
- A. Altmann, L. Toloşi, O. Sander and T. Lengauer, Bioinformatics, 2010, 26, 1340–1347 CrossRef CAS PubMed.
- X. Zou, J. Pan, Z. Sun, B. Wang, Z. Jin, G. Xu and F. Yan, Energy Environ. Sci., 2021, 14, 3965–3975 RSC.
- Q. Zhang, Y. Yuan, J. Zhang, P. Fang, J. Pan, H. Zhang, T. Zhou, Q. Yu, X. Zou, Z. Sun and F. Yan, Adv. Mater., 2024, 36, e2404981 CrossRef PubMed.
- K. Wang, V. Gupta, C. S. Lee, Y. Mao, M. N. T. Kilic, Y. Li, Z. Huang, W. K. Liao, A. Choudhary and A. Agrawal, Sci. Rep., 2024, 14, 25178 CrossRef PubMed.
- H. Kaneko, ACS Omega, 2023, 8, 23218–23225 CrossRef CAS PubMed.
- A. Masarkar, A. K. Maparu, Y. S. Nukavarapu and B. Rai, ACS Appl. Nano Mater., 2024, 7, 19991–20002 CrossRef CAS.
- K. I. Nassar, T. L. P. Galvão, J. D. Gouveia and J. R. B. Gomes, J. Phys. Chem. C, 2025, 129, 2512–2524 CrossRef CAS.
- A. S. Barnard, Cell Rep. Phys. Sci., 2022, 3, 100696 CrossRef CAS.
- A. S. Barnard and B. L. Fox, Chem. Mater., 2023, 35, 8840–8856 CrossRef CAS.
- T. Liu and A. S. Barnard, Cell Rep. Phys. Sci., 2023, 4, 101630 CrossRef.
- S. M. Lundberg, G. Erion, H. Chen, A. DeGrave, J. M. Prutkin, B. Nair, R. Katz, J. Himmelfarb, N. Bansal and S. I. Lee, Nat. Mach. Intell., 2020, 2, 56–67 CrossRef PubMed.
- L. Liu, Y. Li, J. Zheng and H. Li, J. Membr. Sci., 2024, 693, 122327 CrossRef CAS.
- Y. Yang, Z. Yu and D. S. Sholl, Chem. Mater., 2023, 35, 10156–10168 CrossRef CAS PubMed.
- F. Zheng, Z. Zhu, J. Lu, Y. Yan, H. Jiang and Q. Sun, Chem. Phys. Lett., 2023, 814, 140358 CrossRef CAS.
- C. Frie, A. R. Durmaz and C. Eberl, Fatigue Fract. Eng. Mater. Struct., 2024, 47, 2752–2773 CrossRef.
- Y. Matsui and T. Matsui, Theor. Comput. Sci., 2001, 263, 305–310 CrossRef.
- R. Rodríguez-Pérez and J. Bajorath, J. Comput.-Aided Mol. Des., 2020, 34, 1013–1026 CrossRef.
- K. Aas, M. Jullum and A. Løland, Artif. Intell., 2021, 298, 103502 CrossRef.
- J. Roder, L. Maguire, R. Georgantas III and H. Roder, BMC Med. Inf. Decis. Making, 2021, 21, 211 CrossRef PubMed.
- H. Moriwaki, Y. S. Tian, N. Kawashita and T. Takagi, J. Cheminf., 2018, 10, 4 Search PubMed.
- G. Landrum, P. Tosco, B. Kelley, R. Rodriguez, D. Cosgrove, R. Vianello, sriniker, P. Gedeck, G. Jones, E. Kawashima, N. Schneider, D. Nealschneider, A. Dalke, M. Swain, B. Cole, tadhurst-cdd, S. Turk, A. Savelev, A. Vaucher, M. Wójcikowski, I. Take, R. Walker, V. F. Scalfani, H. Faara, K. Ujihara, D. Probst, N. Maeder, J. Monat, J. Lehtivarjo and G. Godin, rdkit/rdkit: 2025_03_3 (Q1 2025) Release, 2025 Search PubMed.
- I. Guyon, J. Weston, S. Barnhill and V. Vapnik, Mach. Learn., 2002, 46, 389–422 CrossRef.
- X. w. Chen and J. C. Jeong, presented in part at the Sixth International Conference on Machine Learning and Applications, Cincinnati, OH, 2007 Search PubMed.
- B. F. Darst, K. C. Malecki and C. D. Engelman, BMC Genet., 2018, 19(Supplement 1), 1–6 Search PubMed.
- K. Pearson, London, Edinburgh Dublin Philos. Mag. J. Sci., 1901, 2, 559–572 CrossRef.
- Y. K. Phua, N. Terasoba, M. Tanaka, T. Fujigaya and K. Kato, ChemElectroChem, 2024, 11, e202400252 CrossRef CAS.
- Y. Xue, Z. R. Li, C. W. Yap, L. Z. Sun, X. Chen and Y. Z. Chen, J. Chem. Inf. Comput. Sci., 2004, 44, 1630–1638 CrossRef CAS PubMed.
- H. Li, C. W. Yap, C. Y. Ung, Y. Xue, Z. W. Cao and Y. Z. Chen, J. Chem. Inf. Model., 2005, 45, 1376–1384 CrossRef CAS PubMed.
- E. Goldberg, M. Scheringer, T. D. Bucheli and K. Hungerbühler, Environ. Sci. Nano, 2015, 2, 352–360 RSC.
- M. R. Findlay, D. N. Freitas, M. Mobed-Miremadi and K. E. Wheeler, Environ. Sci. Nano, 2018, 5, 64–71 RSC.
- A. Bahl, B. Hellack, M. Balas, A. Dinischiotu, M. Wiemann, J. Brinkmann, A. Luch, B. Y. Renard and A. Haase, NanoImpact, 2019, 15, 100179 CrossRef.
- K. L. Goh, A. Goto and Y. Lu, ACS Omega, 2022, 7, 29787–29793 CrossRef CAS PubMed.
- L. E. Castro-Anaya, E. Marese, J. A. Lozano, G. F. Peixer, J. R. Barbosa Jr and S. Y. Gómez González, J. Chem. Inf. Model., 2025, 65, 1812–1825 CrossRef CAS PubMed.
- Y. Saeys, I. Inza and P. Larrañaga, Bioinformatics, 2007, 23, 2507–2517 CrossRef CAS PubMed.
- D. Newby, A. A. Freitas and T. Ghafourian, J. Chem. Inf. Model., 2013, 53, 2730–2742 CrossRef CAS.
- A. Jović, K. Brkić and N. Bogunović, presented in part at the 38th International Convention on Information and Communication Technology, Electronics and Microelectronics, Opatija, Croatia, 2015, pp. 1200–1205 Search PubMed.
- R. Sheikhpour, M. A. Sarram, S. Gharaghani and M. A. Z. Chahooki, Pattern Recogn., 2017, 64, 141–158 CrossRef.
- M. Nssibi, G. Manita and O. Korbaa, Comput. Sci. Res., 2023, 49, 100559 Search PubMed.
- N. T. P. Hartono, J. Thapa, A. Tiihonen, F. Oviedo, C. Batali, J. J. Yoo, Z. Liu, R. Li, D. F. Marrón, M. G. Bawendi, T. Buonassisi and S. Sun, Nat. Commun., 2020, 11, 4172 CrossRef CAS.
- N. Jeong, S. Park, S. Mahajan, J. Zhou, J. Blotevogel, Y. Li, T. Tong and Y. Chen, Nat. Commun., 2024, 15, 10918 CrossRef CAS PubMed.
- X. Liu, X. Zhang, Y. Sheng, Z. Zhang, P. Xiong, X. Ju, J. Zhu and C. Ye, npj Comput. Mater., 2025, 11, 1–14 Search PubMed.
- C. Wang, Y. J. Kim, A. Vriza, R. Batra, A. Baskaran, N. Shan, N. Li, P. Darancet, L. Ward, Y. Liu, M. K. Y. Chan, S. K. R. S. Sankaranarayanan, H. C. Fry, C. S. Miller, H. Chan and J. Xu, Nat. Commun., 2025, 16, 1498 CrossRef CAS PubMed.
- C. Wang, B. Wang, C. Wang, A. Li, Z. Chang and R. Wang, npj Comput. Mater., 2025, 11, 1–13 Search PubMed.
- T. Zhang, Q. Ye, Y. Liu, Q. Liu, Z. Han, D. Wu, Z. Chen, Y. Li and H. J. Fan, Nat. Commun., 2025, 16, 3644 CrossRef CAS PubMed.
- H. Chen, S. M. Lundberg and S. I. Lee, Nat. Commun., 2022, 13, 4512 CrossRef CAS PubMed.
- B. Gregorutti, B. Michel and P. Saint-Pierre, Comput. Stat. Data Anal., 2015, 90, 15–35 CrossRef.
- K. Lin and Y. Gao, Expert Syst. Appl., 2022, 210, 118354 CrossRef.
- R. Todeschini, V. Consonni, R. Mannhold, H. Kubinyi and G. Folkers, Molecular Descriptors for Chemoinformatics: Volume I: Alphabetical Listing/Volume II: Appendices, References, 2 Volume Set, 2nd, Revised and Enlarged Edition, Wiley, New York, 2009 Search PubMed.
- Y. K. Phua, T. Fujigaya and K. Kato, Sci. Technol. Adv. Mater., 2023, 24, 2261833 CrossRef.
- J. Hyun and H.-T. Kim, Energy Environ. Sci., 2023, 16, 5633–5662 RSC.
- J. Cheng, G. He and F. Zhang, Int. J. Hydrogen Energy, 2015, 40, 7348–7360 CrossRef CAS.
- D. Li, A. R. Motz, C. Bae, C. Fujimoto, G. Yang, F.-Y. Zhang, K. E. Ayers and Y. S. Kim, Energy Environ. Sci., 2021, 14, 3393–3419 RSC.
- Z. F. Pan, L. An, T. S. Zhao and Z. K. Tang, Prog. Energy Combust. Sci., 2018, 66, 141–175 CrossRef.
- H. Lei, X. Yang, Z. Chen, D. Rawach, L. Du, Z. Liang, D. S. Li, G. Zhang, A. C. Tavares and S. Sun, Adv. Mater., 2025, 37, e2410106 CrossRef PubMed.
- G. Sriram, K. Dhanabalan, K. V. Ajeya, K. Aruchamy, Y. C. Ching, T. H. Oh, H.-Y. Jung and M. Kurkuri, J. Mater. Chem. A, 2023, 11, 20886–21008 RSC.
- F. Xu, Y. Li, J. Ding and B. Lin, ChemElectroChem, 2023, 10, e202300445 CrossRef CAS.
- A. Sahul Hameed, S. Munusamy, R. Gokulapriyan and D. J. Yoo, ACS Appl. Polym. Mater., 2024, 6, 12341–12361 CrossRef CAS.
- E. J. Park, P. Jannasch, K. Miyatake, C. Bae, K. Noonan, C. Fujimoto, S. Holdcroft, J. R. Varcoe, D. Henkensmeier, M. D. Guiver and Y. S. Kim, Chem. Soc. Rev., 2024, 53, 5704–5780 RSC.
- X. Chu, J. Liu, S. Miao, L. Liu, Y. Huang, E. Tang, S. Liu, X. Xing and N. Li, J. Membr. Sci., 2021, 625, 119172 CrossRef CAS.
- G. H. A. Wijaya, K. S. Im and S. Y. Nam, Desalin. Water Treat., 2024, 320, 100605 CrossRef CAS.
- F. Pedregosa, G. Varoquaux, A. Gramfort, V. Michel, B. Thirion, O. Grisel, M. Blondel, P. Prettenhofer, R. Weiss, V. Dubourg, J. Vanderplas, A. Passos, D. Cournapeau, M. Brucher, M. Perrot and É. Duchesnay, J. Mach. Learn. Res., 2011, 12, 2825–2830 Search PubMed.
- C. Ding and H. Peng, J. Bioinf. Comput. Biol., 2005, 3, 185–205 CrossRef CAS.
- M. Radovic, M. Ghalwash, N. Filipovic and Z. Obradovic, BMC Bioinf., 2017, 18, 9 CrossRef PubMed.
- T. Akiba, S. Sano, T. Yanase, T. Ohta and M. Koyama, presented in part at the Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, 2019, pp. 2623–2631 Search PubMed.
- L. H. B. Olsen and M. Jullum, in Explainable Artificial Intelligence, ed. R. Guidotti, U. Schmid and L. Longo, Springer, Cham, 2026, pp. 194–218 Search PubMed.
- J. Wang, Y. Zhao, B. P. Setzler, S. Rojas-Carbonell, C. Ben Yehuda, A. Amel, M. Page, L. Wang, K. Hu, L. Shi, S. Gottesfeld, B. Xu and Y. Yan, Nat. Energy, 2019, 4, 392–398 CrossRef CAS.
- T. Hastie, R. Tibshirani and J. Friedman, in The Elements of Statistical Learning, ed. T. Hastie, R. Tibshirani and J. Friedman, Springer, New York, New York, 2009, pp. 9–41 Search PubMed.
- R. P. Sheridan, B. P. Feuston, V. N. Maiorov and S. K. Kearsley, J. Chem. Inf. Comput. Sci., 2004, 44, 1912–1928 CrossRef CAS.
- F. Sahigara, D. Ballabio, R. Todeschini and V. Consonni, J. Cheminf., 2013, 5, 27 CAS.
- H. Liu, Y. Ding, L. A. Walker and R. J. Doerksen, Chem. Res. Toxicol., 2015, 28, 169–174 Search PubMed.
- J. Miyake, K. Fukasawa, M. Watanabe and K. Miyatake, J. Polym. Sci., Part A:Polym. Chem., 2014, 52, 383–389 CrossRef CAS.
- X. Li, G. Nie, J. Tao, W. Wu, L. Wang and S. Liao, ACS Appl. Mater. Interfaces, 2014, 6, 7585–7595 CrossRef CAS PubMed.
- D. Guo, C. X. Lin, E. N. Hu, L. Shi, F. Soyekwo, Q. G. Zhang, A. M. Zhu and Q. L. Liu, J. Membr. Sci., 2017, 541, 214–223 CrossRef CAS.
- A. N. Lai, K. Zhou, Y. Z. Zhuo, Q. G. Zhang, A. M. Zhu, M. L. Ye and Q. L. Liu, J. Membr. Sci., 2016, 497, 99–107 CrossRef CAS.
- H.-S. Dang, E. A. Weiber and P. Jannasch, J. Mater. Chem. A, 2015, 3, 5280–5284 RSC.
- K. Pearson, Science, 1909, 30, 23–25 CrossRef CAS PubMed.
- R. I. Mukhamediev, T. Merembayev, Y. Kuchin, D. Malakhov, E. Zaitseva, V. Levashenko, Y. Popova, A. Symagulov, G. Sagatdinova and Y. Amirgaliyev, Remote Sens., 2023, 15, 4269 CrossRef.
- A. G. Asuero, A. Sayago and A. G. González, Crit. Rev. Anal. Chem., 2007, 36, 41–59 CrossRef.
- Y. H. Zhao, M. H. Abraham and A. M. Zissimos, J. Org. Chem., 2003, 68, 7368–7373 CrossRef CAS PubMed.
- I. Gutman, C. Rücker and G. Rücker, J. Chem. Inf. Comput. Sci., 2001, 41, 739–745 CrossRef CAS PubMed.
- OpenAI, arXiv, 2024, preprint, arXiv:2303.08774, DOI:10.48550/arXiv.2303.08774.
- R. Wang, H. Feng and G. W. Wei, J. Chem. Inf. Model., 2023, 63, 7189–7209 CrossRef CAS.
- Z. Zheng, O. Zhang, H. L. Nguyen, N. Rampal, A. H. Alawadhi, Z. Rong, T. Head-Gordon, C. Borgs, J. T. Chayes and O. M. Yaghi, ACS Cent. Sci., 2023, 9, 2161–2170 CrossRef CAS PubMed.
- C. Wei, Y. Shi, W. Mu, H. Zhang, R. Qin, Y. Yin, G. Yu and T. Mu, ACS Sustainable Chem. Eng., 2025, 13, 5368–5380 CrossRef CAS.
- S. Fergus, M. Botha and M. Ostovar, J. Chem. Educ., 2023, 100, 1672–1675 CrossRef CAS.
- A. D. White, G. M. Hocky, H. A. Gandhi, M. Ansari, S. Cox, G. P. Wellawatte, S. Sasmal, Z. Yang, K. Liu, Y. Singh and W. J. Pena Ccoa, Digital Discovery, 2023, 2, 368–376 RSC.
- S.-Ş. Uçar, I. Lopez-Gazpio and J. Lopez-Gazpio, Educ. Inf. Technol., 2025, 30, 11463–11482 CrossRef.
- C. Morbidoni, A. Mascitti, A. Sarra, L. Tonucci and F. Coccia, J. Chem. Educ., 2026, 103, 286–297 CrossRef CAS.
- J. Pan, J. Han, L. Zhu and M. A. Hickner, Chem. Mater., 2017, 29, 5321–5330 CrossRef CAS.
- Y. Zhu, L. Ding, X. Liang, M. A. Shehzad, L. Wang, X. Ge, Y. He, L. Wu, J. R. Varcoe and T. Xu, Energy Environ. Sci., 2018, 11, 3472–3479 RSC.
- L. Zhu, X. Peng, S. L. Shang, M. T. Kwasny, T. J. Zimudzi, X. Yu, N. Saikia, J. Pan, Z. K. Liu, G. N. Tew, W. E. Mustain, M. Yandrasits and M. A. Hickner, Adv. Funct. Mater., 2019, 29, 1902059 CrossRef.
- D. Pan, P. M. Bakvand, T. H. Pham and P. Jannasch, J. Mater. Chem. A, 2022, 10, 16478–16489 RSC.
- J. Chen, M. Zhang, C. Shen and S. Gao, Ind. Eng. Chem. Res., 2022, 61, 1715–1724 CrossRef CAS.
- S. Willdorf-Cohen, A. N. Mondal, D. R. Dekel and C. E. Diesendruck, J. Mater. Chem. A, 2018, 6, 22234–22239 RSC.
- S. Willdorf-Cohen, A. Zhegur-Khais, J. Ponce-Gonzalez, S. Bsoul-Haj, J. R. Varcoe, C. E. Diesendruck and D. R. Dekel, ACS Appl. Energy Mater., 2023, 6, 1085–1092 CrossRef CAS PubMed.
- J. W. Onorato, Z. Wang, Y. Sun, C. Nowak, L. Q. Flagg, R. Li, B. X. Dong, L. J. Richter, F. A. Escobedo, P. F. Nealey, S. N. Patel and C. K. Luscombe, J. Mater. Chem. A, 2021, 9, 21410–21423 RSC.
- P. Durand, H. Zeng, T. Biskup, V. Vijayakumar, V. Untilova, C. Kiefer, B. Heinrich, L. Herrmann, M. Brinkmann and N. Leclerc, Adv. Energy Mater., 2022, 12, 2103049 CrossRef CAS.
- B. Yang and Z. Cunman, Chem. Eng. J., 2023, 457, 141094 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |