 Open Access Article
Open Access ArticleDecoupling the structural origins of strength and toughness in industrially relevant heterogeneous elastomers
Yuri
Ishikura
a,
Erica
Uehara
b and
Yuji
Higuchi
 *c
*c
aR&D Department, Sekisui Chemical Co., Ltd, 2-1 Hyakuyama, Shimamoto-cho, Mishima-gun, Osaka 618-8589, Japan
bGraduate School of Informatics, Kyoto University, Japan
cResearch Institute for Information Technology, Kyushu University, 744 Motooka, Nishi-ku, Fukuoka, 819-0395, Japan. E-mail: higuchi.yuji.474@m.kyushu-u.ac.jp; Fax: +81-92-802-2642; Tel: +81-92-802-2646
First published on 10th March 2026
Abstract
Using coarse-grained molecular dynamics, we investigate the structure–property relationships for industrially relevant heterogeneous elastomer networks formed by random crosslinking. Our simulations reveal a clear decoupling of mechanical properties: strength and elongation are governed by the network's cycle rank (a measure of global topology), while toughness (fracture energy) is governed by the effective chain ratio and strand length uniformity. Notably, network uniformity, which is critical for toughness, is enhanced by using wider reactive site spacing and lower-functionality crosslinkers. This insight provides a rational design guideline to overcome the common strength–toughness trade-off by independently tuning molecular parameters to optimize distinct mechanical properties in these complex yet industrially vital materials.
Introduction
Elastomers are a group of polymeric materials characterized by rubber elasticity, exhibiting significant reversible deformation in response to external forces.1–3 This characteristic rubber elasticity stems from the polymer chains forming a three-dimensional network structure via chemical or physical crosslinking points.4 Because of their outstanding flexibility and toughness, elastomers have established themselves as essential foundational materials not only for traditional industrial products like tires and sealants but also for cutting-edge fields such as soft robotics, wearable devices, and regenerative medical scaffolds, and their importance continues to grow.4–7 The mechanical properties required for these diverse applications, which include elastic modulus, fracture strength, toughness, and energy dissipation capacity, are critically determined by the microscopic crosslinking structure of the polymer network.7–10 Therefore, designing materials to achieve desired functions necessitates understanding the correlation between the network structure and macroscopic properties at the molecular level.Based on this structure–property correlation, attempts have been made to control the network structure to achieve desired properties. For example, tetra-PEG gels11 with structures close to ideal networks, characterized by a uniform molecular weight between crosslinks, were synthesized by reacting tetra-branched prepolymers at their ends, experimentally contributing to the theoretical verification of fracture energy.1 Double-network (DN) gels,12,13 which combine a rigid primary network with a flexible secondary network, have been reported to exhibit extremely high toughness by efficiently dissipating energy through the breaking of sacrificial bonds.5 The concept of a DN structure was also applied to elastomers, resulting in enhanced mechanical properties.14 This demonstrates the importance of network structures for improving the mechanical properties of elastomers. Furthermore, self-healing functionality achieved through hybrid crosslinking methods that introduce multiple crosslinks with different properties, such as covalent and hydrogen bonds, has been actively studied.15–17 However, even in these advanced material systems, completely controlling network heterogeneity remains experimentally challenging. Controlling the network structure is particularly difficult for elastomers formed by random crosslinking reactions, which are widely employed in industry. Thus, clarification of the relationship between the randomly crosslinked network structure and mechanical properties at the molecular level is strongly desired.
Molecular dynamics (MD) simulations provide an effective means to elucidate the relationship between mechanical properties and molecular-level structures. This approach has been applied to elastomers,17,18 polymer glasses,19–21 polymer gels,22,23 polymer composites,24–27 and semicrystalline polymers.28–41 In particular, coarse-grained (CG) MD simulations are an extremely powerful tool for clarifying the relationship between the polymer network structure and mechanical response.2,3,10,11 Previous studies using CGMD to investigate structure–property correlations for crosslinked elastomers have largely focused on end-linked networks such as tetra-PEG gels.7,12,13,42 These systems, with their uniform molecular weights between crosslinks, have served as ideal networks for direct comparison with theoretical models, playing a crucial role in understanding the fundamental principles of elasticity and fracture. However, industrially produced natural rubber and numerous synthetic rubbers are crosslinked via random radical or condensation reactions. Consequently, they form heterogeneous networks with broad distributions of inter-crosslink distances, including dangling chains and loop structures.3,11 Such network heterogeneity is known to generate local variations in stress concentration and energy dissipation that affect the overall mechanical properties of the material.7 Recent work by Mohanty et al. demonstrated that fracture in heterogeneous networks propagates along the “shortest path” where tensile stress concentrates, rather than at the crack tip, highlighting the significance of heterogeneity.43 However, systematic simulation studies that might enable control over this microscopic heterogeneity and prediction of material properties remain scarce.
Therefore, the aim of this study was to elucidate the structure–property correlations of elastomers featuring heterogeneous crosslinked networks using CGMD simulations. Specifically, we focus on two fundamental molecular design parameters that govern network heterogeneity: (1) the spacing between crosslinking sites on the prepolymer chains and (2) the number of functional groups per crosslinker. These are key elements for controlling the network chain-length distribution and the topology of crosslinking points. For example, in their CGMD simulations of DN gels, Higuchi et al. demonstrated that increasing the chain length and the distance between crosslinking points of the soft second network enhances peak stress and ductility relative to the rigid first network.23 Furthermore, Yin et al. experimentally demonstrated that using a bifunctional crosslinker instead of a tetrafunctional one allows chain elongation to dominate, affording both a low elastic modulus and high toughness.44 In this study, we construct a model system by systematically varying these parameters while assuming an identical monomer composition and perform laterally constrained uniaxial extension simulations. This approach elucidates how the microscopic network structure influences the stress–strain curve, elastic modulus, elongation at break, and structural changes leading to failure.
Methods
Simulation model and conditions
CGMD simulations using a bead–spring model with three-dimensional periodic boundary conditions were performed to investigate the tensile characteristics of thermoset elastomers. Polymer chains were modeled as linear polymers of 400 beads, with 100 chains arranged in the simulation box. Beads designated as crosslinking sites were defined as “reactive beads,” while all other beads were “normal beads,” as shown in Fig. 1.The simulation used dimensionless parameters with the energy and length units set to  and σ = 1, respectively. Non-bonding interactions were described using the Lennard-Jones potential:
and σ = 1, respectively. Non-bonding interactions were described using the Lennard-Jones potential:
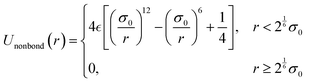 | (1) |
 and σ0 = σ is the bead diameter. Bonding interactions in the main chain were described using a harmonic potential:
and σ0 = σ is the bead diameter. Bonding interactions in the main chain were described using a harmonic potential:| Ubond(r) = K(r − r0)2 | (2) |
 | (3) |
The fraction of reactive beads (ϕreactive) was set to 0.025, 0.05, or 0.1. Reactive beads were arranged regularly with spacings of 10, 20, or 40 beads. Terminal beads were normal beads. Crosslinkers with two, three, or four functional groups were added at 0.5–5 wt% relative to the polymer. A crosslinking reaction occurred whenever a reactive bead and a functional group of a crosslinker were within a distance of 1σ. This reaction was attempted every 100 simulation steps.
Initially, chains were randomly arranged using OCTA,45 and the system was equilibrated using the NPT ensemble without reactions to relax the polymer melt structure at a temperature of T = 1.0T0, where the temperature unit is  (kB is the Boltzmann constant). The equilibrium simulation time was 1.0 × 108τ, where the time unit
(kB is the Boltzmann constant). The equilibrium simulation time was 1.0 × 108τ, where the time unit  (m is the bead mass). Subsequently, the crosslinking reaction was carried out for up to 1.0 × 106τ under a constant volume. After the reaction, another equilibrium calculation was performed for 1.0 × 108τ using the NPT ensemble to relax the crosslinked structure.
(m is the bead mass). Subsequently, the crosslinking reaction was carried out for up to 1.0 × 106τ under a constant volume. After the reaction, another equilibrium calculation was performed for 1.0 × 108τ using the NPT ensemble to relax the crosslinked structure.
Laterally constrained uniaxial extension simulations were performed at T = 1.0T0. The system was elongated in the z direction at a constant engineering strain rate corresponding to 100% strain every 103τ. The elongation ratio is defined as λ = Lz/Lz0, where Lz is the length of the simulation box in the tensile direction and Lz0 is its initial length. The lengths of the x and y axes were fixed. We conducted void detection in the polymer network using a grid-based approach under periodic boundary conditions. Particle positions were wrapped within the simulation box, and the box was voxelized into a grid with spacing (dx = 0.5). Voids were identified as unoccupied voxels, and connected components were labeled to calculate cavity volumes.
The simulation time step was dt = 0.005τ for all simulations. The temperature was controlled using a Langevin thermostat in the tensile simulations, while the pressure was maintained using a Nosé–Hoover barostat in the NPT simulations. The thermostat and barostat relaxation times were set to 0.5τ and 5τ, respectively.
Simulations were performed using OCTA/COGNAC45 (for reactions) and LAMMPS46 (for equilibration and elongation).
Data analysis
The following analyses were performed on the calculated stress–strain curves:Uniaxial extensional modulus C 33: the slope of the stress–strain curve in the linear regime, determined by linear fitting in the small-strain range (εzz ≤ 0.006), corresponds to the C33 component of the stiffness matrix, not Young's modulus. This distinction arises due to the lateral constraints applied in the x- and y-directions, aligning with the generalized Hooke's law under the imposed boundary conditions. For isotropic materials, C33 is greater than Young's modulus (E) when Poisson's ratio (ν) is greater than zero. Therefore, the term ‘uniaxial extensional modulus’ is used here to accurately describe the extracted elastic constant under the specific simulation conditions employed. We have verified that the value of C33 converges within this strain range, and additional benchmarks confirm that C33 is invariant with respect to the imposed strain rate. The SI provides detailed data supporting these findings.
Yield stress: the value of the first peak stress on the stress–strain curve.
Hardness in the high-strain region: the slope of the tangent to the stress–strain curve at a strain of S = 300%, calculated numerically over a small interval of S = 300% ± 2%.
To quantify the microscopic structure of the generated networks, structural analysis was performed using the Python graph theory library NetworkX.47 The bead and bond information was converted into nodes (chain ends and crosslinking points) and edges (connecting two nodes), respectively. The following structural parameters were calculated:
Dangling chain: partial chain from the end of the polymer chain to the crosslinking point or the chain end. Dangling chains are considered network defects because they do not effectively transmit stress. Dangling chains were identified by tracing bond connectivity along the polymer backbone. Terminal beads were defined as beads with only one bonded neighbor (degree = 1). From each terminal bead, bonded neighbors were followed sequentially until either a crosslinking point (degree ≥ 3) or another terminal bead was reached. All beads along this linear segment were classified as belonging to a dangling chain. The dangling chain ratio was calculated as Ndangling/Nall, where Ndangling is the number of beads composing the dangling chains and Nall is the total number of beads in the simulation.
Loop: a closed loop formed by crosslinking different sites on the same prepolymer chain. A loop is elastically ineffective. Loops were identified by tracing bond connectivity starting from polymer beads directly bonded to crosslinking-agent beads. The chain was followed along bonded neighbors until another crosslinking-agent bead was reached. When the path terminated at the same crosslinking-agent bead from which it originated, the segment was classified as a loop. The loop ratio was defined as Nloop/Nall, where Nloop is the number of beads composing the loop.
Effective chain: a polymer chain connecting two distinct crosslinking points. Effective chains were defined as polymer segments connecting two distinct crosslinking-agent beads. These segments were identified by tracing bond connectivity starting from beads adjacent to crosslinking agents and classifying paths that terminated at a different crosslinking agent. The effective chain ratio was calculated as Nactive/Nall, where Nactive is the number of beads composing the effective chains.
Strand length: a length (number of beads) between two adjacent crosslinks. The mean and standard deviation of the strand length were calculated to evaluate the network homogeneity.
Cycle rank: a graph theory indicator quantifying the topological connectivity of a network. It is the minimum number of bonds that must be cut to transform the network into a tree structure. A higher cycle rank indicates a more robustly connected network. It was calculated using the formula cycle rank = n − ne + c, where ne is the total number of edges, n is the total number of nodes, and c is the number of connected components.
The extracted mechanical properties were subsequently used for statistical evaluation. The Pearson correlation coefficients reported in this study were calculated by pooling data from all simulated conditions into a single dataset. Linear regression analysis was then performed on the combined dataset to quantify the strength of the linear relationship between structural parameters and mechanical properties.
Results and discussion
Network structure analysis
To establish the targeted molecular design guidelines, we first quantitatively evaluated the effects of prepolymer reactive site spacing and crosslinker functionality on the resulting network topology. Fig. 2 shows the network structure parameters with respect to the number of crosslinks. Ndangling decreased monotonically as the number of crosslinks increased (Fig. 2a). This trend was independent of crosslinker functionality. Notably, wider spacing between reactive sites on the prepolymer suppressed dangling chain formation. On the other hand, Nloop increased with the number of crosslinks (Fig. 2b). Loop formation was particularly pronounced in the systems with trifunctional and tetrafunctional crosslinkers, whereas it was suppressed with bifunctional crosslinkers. This is likely attributable to multifunctional crosslinkers acting as hubs, increasing the probability of capturing multiple reactive sites from the same chain, whereas bifunctional crosslinkers primarily act as chain extenders. The effective chain ratio initially increased with the number of crosslinks before becoming saturated (Fig. 2c). The change in the effective chain ratio was independent of crosslinker functionality and the reactive site spacing.The average strand length and its standard deviation are plotted in Fig. 2d and e, respectively, which indicate the network uniformity. The average strand length decreased with increasing crosslink density, showing similar behavior across all crosslinker functionalities and reactive-site spacings. While crosslink density was the dominant factor controlling strand-length dispersion, increasing the reactive-site spacing produced a secondary but systematic reduction in the standard deviation. This suggests a modest improvement in structural homogeneity at larger reactive-site spacing. In contrast, shorter spacing promotes successive reactions at nearby sites, leading to greater local heterogeneity. This local chain reaction is thought to generate a heterogeneous network structure containing both short and long strands. To further substantiate this interpretation, we calculated the radial distribution function (RDF) between already crosslinked reactive sites and remaining unreacted reactive sites. The RDF analysis revealed that, for systems with shorter reactive-site spacing, a significant population of unreacted reactive sites remains in close proximity to already crosslinked sites. In contrast, for systems with wider reactive-site spacing, unreacted sites were less concentrated around previously crosslinked points (Fig. 3).
These results support the proposed local chain-reaction mechanism: when reactive sites are closely spaced along the prepolymer backbone, the formation of a crosslink increases the likelihood that neighboring reactive sites will subsequently react within the same local region. This spatial correlation enhances local clustering of crosslinks and contributes to the development of heterogeneous network structures containing both short and long strands. Conversely, wider reactive-site spacing suppresses such local correlation effects, leading to a more spatially homogeneous network formation process. From this structural analysis, we found that wider reactive site spacing on the prepolymer and lower crosslinker functionality (bifunctional) lead to a more uniform network with fewer defects.
Finally, the cycle rank was calculated to evaluate the overall topological connectivity. Fig. 2f shows that the cycle rank increased with the total number of crosslinks. The degree of the increase in the cycle rank was unaffected by the reactive site spacing, although it was affected by the crosslinker functionality. This finding is consistent with previous studies by Masubuchi and co-workers,48,49 who reported that the cycle rank of star-polymer networks depends on both the number of crosslinks and the crosslinker functionality.
Correlation between mechanical properties and structural factors: elasticity and yield behavior
Next, laterally constrained uniaxial extension elongation simulations were performed for each constructed network model. The stress–strain curve shown in Fig. 4a exhibits typical strain-hardening behavior, similar to conventional elastomers with 1000 crosslinks, a reactive site spacing of 10, and a bifunctional crosslinker, taken during elongation at a constant strain rate of 0.001/τ. Fig. 4b shows typical snapshots of the bifunctional network. At strains from 0% to 100%, the elastomer deformed, and void generation was observed (Table 1).| Strain | Volume [σ3] | ||||
|---|---|---|---|---|---|
| Void 1 | Void 2 | Void 3 | Void 4 | Void 5 | |
| 0% | — | — | — | — | — |
| 50% | 9245 | 6524 | 535 | 4 | 3 |
| 100% | 33![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 287 287 |
6949 | — | — | — |
The uniaxial extensional modulus C33 and yield stress were calculated, and for reference, hardness at high strain (S = 300%) was also obtained, acknowledging that this high-strain value serves as an indicative measure rather than a definitive physical property. The correlations between these mechanical properties and the structural factors are presented in Fig. 5.
Both yield stress and hardness at high strain exhibited extremely strong positive correlations with the cycle rank (Fig. 4a and b), where the Pearson correlation coefficients were 0.81 and 0.90, respectively. This indicates that the resistance of the materials to large deformation is governed by the strength of the topological constraints across the entire network rather than by the properties of individual strands. In contrast, the uniaxial extensional modulus C33 did not display a clear correlation with any of the structural factors evaluated in this study (Fig. 5c), where the maximum coefficient was only 0.27. This suggests that the elastic response in the low-strain regime is governed by smaller-scale structures, such as the interparticle potential and local chain conformations. The parameters examined in this study, such as the average strand length and its standard deviation, likely exerted a smaller direct impact on local stiffness during minor deformations.
Fracture behavior and dominant structural factors
To elucidate the fracture behavior, we performed simulations where bonds could sever when their length exceeded a critical value (1.5σ). Fig. 6a presents a stress–strain curve for the network featuring 1000 crosslinks, a reactive site spacing of 10 beads, and bifunctional crosslinkers. Fig. 6b shows typical snapshots of the same system during elongation at a constant strain rate of 0.001/τ. We calculated the fracture stress, fracture strain, and fracture energy (the area under the stress–strain curve up to the fracture point) and analyzed their correlation with structural factors. The fracture point was defined as the point exhibiting the maximum stress in the strain region above 100%. In the present study, toughness was quantified as the tensile toughness, defined as the area under the uniaxial stress–strain curve. We acknowledge that elastomer toughness is often characterized using fracture mechanics-based measures such as tearing energy or fracture toughness derived from stress intensity factors.Although these quantities are not identical, they are all fundamentally associated with the material's ability to dissipate energy prior to catastrophic failure. From this perspective, the structural factors identified in this study are expected to influence alternative toughness measures through their impact on stress distribution and energy dissipation near crack tips.
However, direct evaluation of tearing energy or crack-tip process-zone behavior was beyond the scope of the present simulations. Future work incorporating explicit crack simulations would be necessary to quantitatively establish the relationship between the identified network descriptors and fracture mechanics-based toughness measures.
Fig. 7 shows the fracture stress and strain plotted against the cycle rank. A strong correlation was observed between each pair of parameters. The correlation coefficients between network structure parameters and fracture strain/fracture stress are presented in Fig. 8. In both cases, the cycle rank exhibited the highest correlation coefficients, with values of 0.97 for fracture stress and −0.85 for fracture strain. In contrast, as shown in Fig. 9, the fracture energy, which reflects material toughness, was most strongly correlated not with the cycle rank but with the effective chain ratio and the strand length standard deviation (strand length uniformity), where the correlation coefficients were 0.74 and −0.73, respectively.
 | ||
| Fig. 7 Correlation of fracture properties with the cycle rank. Scatter plots showing (a) fracture stress and (b) fracture strain as a function of the cycle rank for all simulated networks. | ||
These findings are further detailed in Fig. 10, which shows plots of the fracture energy against the two structural parameters. Although the data are more scattered than the correlations with the cycle rank presented in Fig. 8, the underlying trends are clearly evident. Specifically, Fig. 9b and 10a show that the fracture energy increased with increasing effective chain ratio and decreased with increasing standard deviation of the strand length, respectively, indicating that a more uniform network leads to higher toughness. It is worth noting that when the cycle rank of a network is low, its connectivity is close to that of a tree. Such networks often contain numerous articulation points and bridge edges and can be disconnected by a small number of cross-chain breaks.
Our results advance the understanding proposed by Masubuchi et al.,48 who suggested that the cycle rank universally describes fracture properties, and offer the more nuanced view proposed below.
First, this study extends network design principles from a theory based solely on the number of effective chains to one incorporating both their number and their quality. While the cycle rank quantifies the number of independent constraint paths, our results show that a high cycle rank does not guarantee high toughness if the quality—specifically, the uniformity of strand lengths—is low.
Second, it provides higher resolution for the fracture mechanism. In heterogeneous networks, stress concentrates on short strands, likely initiating early crack propagation. This aligns with the description by Mohanty et al. of fracture progressing along the “shortest path” of concentrated tension.43
It should be noted that the present simulations focus on randomly crosslinked networks in which multiple reactive sites are distributed along the prepolymer backbone. Such architectures are representative of many industrially produced elastomers formed via radical or condensation reactions. In contrast, end-linked systems such as tetra-PEG gels and telechelic networks possess only two reactive sites per chain and typically exhibit narrower strand-length distributions with suppressed loop formation.
Previous studies on these end-linked networks have shown that their elastic properties can be quantitatively described in terms of elastically effective chain density and defect statistics, largely independent of the specific crosslinking protocol.
In this context, the present work extends such a descriptor-based framework to heterogeneous randomly crosslinked elastomers. By identifying cycle rank as the dominant factor governing strength-related properties and effective chain ratio and strand-length uniformity as the key determinants of toughness, we provide a structural interpretation that is not restricted to a specific network generation mechanism. While quantitative relationships may differ between random and end-linked architectures due to differences in strand-length distributions and defect populations, the decoupling between global topological connectivity and local strand quality may represent a general principle that could be explored and validated across diverse elastomer systems in future studies.
In a uniform network, however, stress is shared more cooperatively among numerous strands, allowing the material as a whole to absorb more energy before failure. This concept of cooperative stress sharing reveals an essential mechanism for toughness that is not fully captured by the average indicator of cycle rank alone.
Therefore, designing tough polymer networks requires not only increasing the number of effective chains (related to the cycle rank) but also enhancing their quality by improving uniformity. The present study builds upon the framework of Masubuchi et al. by identifying uniformity as a critical new design axis for achieving high toughness.
Our results clearly demonstrate that the strength and elongation of elastomers are governed by different network structural factors than their toughness (fracture energy). Specifically, strength depends on topological constraints (cycle rank), whereas toughness is governed by the quantity and uniformity of the effective chains responsible for energy dissipation (Fig. 11). This insight provides a new material design guideline for simultaneously achieving both strength and toughness, which are often subject to a trade-off relationship, by independently controlling the prepolymer primary structure (reactive site spacing) and the crosslinking conditions (crosslinker functionality and amount).
Conclusions
In this study, we have systematically clarified the effects of prepolymer reactive site spacing and crosslinker functionality on the network structure and mechanical properties of crosslinked elastomers using CGMD simulations. Structural analysis revealed that wider reactive site spacing and lower crosslinker functionality suppress the formation of network defects such as dangling chains and loops, leading to a network composed of longer, more uniform strands.Correlation analysis with mechanical properties revealed the most significant finding of this study: the structural factors governing the strength and elongation of an elastomer are distinct from those determining its toughness (fracture energy). Specifically, properties related to strength and elongation—such as yield stress, hardness at high strain, fracture stress and fracture strain—exhibited extremely strong positive correlations with the cycle rank, an indicator of the network's topological constraints. This suggests that the ability of a material to withstand large deformations depends directly on how robustly the network is connected.
Conversely, fracture energy, which reflects material toughness, was found to be strongly governed not by the cycle rank but by the quantity and quality of the energy-dissipating network chains, namely, the effective chain ratio and strand length uniformity. This implies that even with a high cycle rank, numerous network defects or non-uniform strand lengths can cause stress concentration, leading to fracture before the material as a whole can absorb energy.
These findings suggest that the strength and toughness of elastomers originate from distinct hierarchical network parameters. This provides new guidance for the rational design of high-performance elastomers:
1. To enhance strength and elongation, increasing the cycle rank by adjusting the crosslink density and crosslinker functionality should be effective.
2. To improve toughness, designing the prepolymer with appropriate reactive site spacing to form a high number of uniform effective chains is appropriate.
This distinction suggests that coordinated control of these molecular parameters provides a viable route toward partial and targeted tuning of strength and toughness. It should be noted, however, that completely independent control is inherently limited. Increasing crosslink density not only enhances the cycle rank but can also improve strand-length uniformity, thereby influencing toughness-related parameters as well. Consequently, structural parameters are not strictly orthogonal, and modifications intended to enhance strength may simultaneously affect toughness.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). See DOI: https://doi.org/10.1039/D6SM00007J. The input files and initial data set for the simulations are available at https://nomadlab.eu/prod/v1/gui/user/uploads/upload/id/djDW09gRS5WYYpHbec_rmQ.Acknowledgements
This work used computational resources of SQUID provided by Osaka University through the HPCI System Research Project (Project ID: hp250502).Notes and references
- L. R. G. Treloar, The Physics of Rubber Elasticity, Oxford University Press, 1975 Search PubMed.
- P. Bednarczyk, K. Mozelewska and Z. Czech, Int. J. Adhes. Adhes., 2020, 102, 102652 CrossRef CAS.
- Y. Wang, H. Liu, P. Li and L. Wang, Polymers, 2022, 14, 1308 CrossRef CAS PubMed.
- J. L. Barrat, et al. , JPhys: Mater., 2024, 7, 012501 Search PubMed.
- L. Zhang, S. Chen and Z. You, Acc. Chem. Res., 2023, 56, 2907 Search PubMed.
- D. L. Henann, S. A. Chester and K. Bertoldi, J. Mech. Phys. Solids, 2013, 61, 2047 CrossRef.
- M. Yunwei and A. Lallit, J. Appl. Mech., 2018, 85, 081008 CrossRef.
- M. Pharr, J. Sun and Z. Suo, J. Appl. Phys., 2012, 111, 104114 CrossRef.
- M. Tsige and M. J. Stevens, Macromolecules, 2004, 37, 630 CrossRef CAS.
- M. Solar, Z. Qin and M. J. Buehler, J. Mater. Res., 2022, 14, 1308 Search PubMed.
- T. Sakai, T. Matsunaga, Y. Yamamoto, C. Ito, R. Yhoshida, S. Suzuki, N. Sasaki, M. Shibayama and U. Chung, Macromolecules, 2008, 41, 5379 CrossRef CAS.
- J. P. Gong, Soft Matter, 2010, 6, 2583 RSC.
- J. Gong, Y. Katsuyama, T. Kurokawa and Y. Osada, Adv. Mater., 2003, 15, 1155 CrossRef CAS.
- S. Yasuda, R. Yamamoto and S. Hyodo, Rheol. Acta, 2009, 48, 781 Search PubMed.
- J. Gong, Soft Matter, 2010, 6, 2583 RSC.
- E. Ducrot, Y. Chen, M. Bulters, R. P. Sijbesma and C. Creton, Science, 2014, 344, 186 CrossRef CAS PubMed.
- T. L. Sun, T. Kurokawa, S. Kuroda, A. B. Ihsan, T. Akasaki, K. Sato, M. A. Haque, T. Nakajima and J. P. Gong, Nat. Mater., 2013, 12, 932 CrossRef CAS PubMed.
- A. B. Ihsan, T. L. Sun, S. Kuroda, T. Nakajima and J. P. Gong, J. Mater. Chem. B, 2013, 1, 4555 RSC.
- A. Makke, M. Perez, O. Lame and J. L. Barrat, J. Chem. Phys., 2009, 131, 014904 CrossRef PubMed.
- D. G. Tsalikis, M. Ciobanu, C. S. Patrickios and Y. Higuchi, Macromolecules, 2023, 56, 9299 CrossRef CAS.
- J. Rottler and M. O. Robbins, Phys. Rev. E: Stat., Nonlinear, Soft Matter Phys., 2003, 68, 011801 CrossRef PubMed.
- K. Fujimoto, Z. Tang, W. Shinoda and S. Okazaki, Polymer, 2019, 178, 121570 Search PubMed.
- A. Makke, O. Lame, M. Perez and J. L. Barrat, Macromolecules, 2013, 46, 7853 Search PubMed.
- Y. R. Sliozberg, R. A. Mrozek, J. D. Schieber, M. Kröger, J. L. Lenhart and J. W. Andzelm, Polymer, 2013, 54, 2555 Search PubMed.
- Y. Higuchi, K. Saito, T. Sakai, J. P. Gong and M. Kubo, Macromolecules, 2018, 51, 3075 Search PubMed.
- J. Zhao, L. Wu, C. Zhan, Q. Shao, Z. Guo and L. Zhang, Polymer, 2017, 133, 272 Search PubMed.
- K. Hagita, Polymer, 2019, 174, 218 CrossRef CAS.
- P. Khan, R. Kaushik and A. Jayaraj, ACS Omega, 2022, 7, 47567 CrossRef CAS PubMed.
- Y. Higuchi, Y. Hikima, N. Kamiuchi, E. Uehara, Y. Oya, G. Yamamoto and K. Sakakibara, Polymer, 2025, 333, 128539 CrossRef CAS.
- S. Lee and G. C. Rutledge, Macromolecules, 2011, 44, 3096 CrossRef CAS.
- I.-C. Yeh, J. W. Andzelm and G. C. Rutledge, Macromolecules, 2015, 48, 4228 CrossRef CAS.
- T. Yamamoto, Polymer, 2013, 54, 3086 CrossRef CAS.
- S. Jabbari-Farouji, J. Rottler, O. Lame, A. Makke, M. Perez and J.-L. Barrat, ACS Macro Lett., 2015, 4, 147 CrossRef CAS PubMed.
- Y. Higuchi and M. Kubo, Modell. Simul. Mater. Sci. Eng., 2016, 24, 055006 CrossRef.
- A. Moyassari, T. Gkourmpis, M. S. Hedenqvist and U. W. Gedde, Macromolecules, 2019, 52, 807 CrossRef CAS.
- Y. Higuchi and M. Kubo, Macromolecules, 2017, 50, 3690 CrossRef CAS.
- Y. Higuchi, Polym. J., 2018, 50, 579 CrossRef CAS.
- Y. Higuchi, Macromolecules, 2019, 52, 6201 CrossRef CAS.
- Y. Higuchi, Phys. Rev. E, 2021, 103, 042502 CrossRef CAS PubMed.
- Y. Higuchi and G. Matsuba, Macromol. Chem. Phys., 2024, 225, 2400076 CrossRef CAS.
- W. S. Fall, J. Baschnagel and H. Meyer, npj Soft Matter, 2025, 1, 6 Search PubMed.
- S. Yasuda, R. Yamamoto and S. Hyodo, Rheol. Acta, 2009, 48, 781 Search PubMed.
- S. Mohanty, J. Blanchet, Z. Suo and W. Cai, arXiv, 2025, preprint, arXiv:2502.11339 DOI:10.48550/arXiv.2502.11339.
- L. Yin, Y. Zhao, J. Zhu, M. Yang, H. Zhao, J. Pei, S. Zhong and Z. Dang, Nat. Commun., 2021, 12, 4517 Search PubMed.
- M. Doi, Macromol. Symp., 2003, 195, 101 Search PubMed.
- A. P. Thompson, H. M. Aktulga, R. Berger, D. S. Bolintineanu and W. M. Brown, et al. , Comput. Phys. Commun., 2022, 271, 108171 CrossRef CAS.
- A. A. Hagberg, D. A. Schult and P. J. Swart, in Proceedings of the 7th Python in Science Conference (SciPy2008), ed. G. Varoquaux, T. Vaught and J. Millman, Pasadena, CA, 2008, pp. 11–15 Search PubMed.
- Y. Masubuchi, Polymer, 2024, 297, 126880 CrossRef CAS.
- Y. Masubuchi, Y. Doi, T. Ishida, N. Sakumichi, T. Sakai, K. Mayumi, K. Satoh and T. Uneyama, Macromolecules, 2023, 56, 9359 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |










