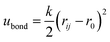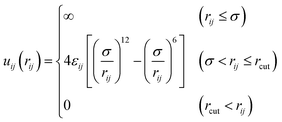Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Following the formation of single-chain nanoparticles generated by interblock crosslinking within diblock copolymers: a Monte Carlo simulation study with adjustable interaction strength between the blocks
Anja
Voigt
a,
Christian
Strauch
 b,
Tom
Höfken
b,
Mirco
Wahab
c,
Wiebke
Hadwich
a,
Christopher
Barner-Kowollik
b,
Tom
Höfken
b,
Mirco
Wahab
c,
Wiebke
Hadwich
a,
Christopher
Barner-Kowollik
 de,
Conrad
Hübler
de,
Conrad
Hübler
 a,
Stefanie
Schneider
a,
Stefanie
Schneider
 *b and
Felix A.
Plamper
*b and
Felix A.
Plamper
 *afg
*afg
aInstitute of Physical Chemistry, TU Bergakademie Freiberg, Lessingstr. 45, 09599 Freiberg, Germany. E-mail: plamper@chemie.tu-freiberg.de
bInstitute of Physical Chemistry II, RWTH Aachen University, Landoltweg 2, 52056 Aachen, Germany. E-mail: schneider@pc.rwth-aachen.de
cInstitute of Glass Technology, TU Bergakademie Freiberg, Leipziger Straße 28, 09599 Freiberg, Germany
dInstitute of Functional Interfaces (IFG), Karlsruhe Institute of Technology (KIT), Hermann-von-Helmholtz-Platz 1, 76344 Eggenstein-Leopoldshafen, Germany
eSchool of Chemistry and Physics, Centre for Materials Science, Queensland University of Technology (QUT), 2 George Street, Brisbane, QLD 4000, Australia
fCenter for Efficient High Temperature Processes and Materials Conversion ZeHS, TU Bergakademie Freiberg, 09599 Freiberg, Germany
gFreiberg Center for Water Research ZeWaF, TU Bergakademie Freiberg, 09599 Freiberg, Germany
First published on 13th April 2026
Abstract
We investigated the folding and crosslinking of diblock copolymers with interblock-crosslinkable units in dilute solution. We used a bead-spring model for the polymer and Monte Carlo simulations for the crosslinking. At no and small interblock attractive interaction, the observed zipping between spatially proximate crosslinkers results in single-chain nanoparticles resembling expanded ladder-type polymers. Stronger attractive interactions between the different blocks lead to an enhanced internal confinement resulting in more compact, randomly crosslinked structures. The structural outcome of the “reactive” Monte Carlo (MC) simulations was also qualitatively recovered by applying “reactive” molecular dynamics (MD) simulations.
1. Introduction
Single-chain nanoparticles (SCNPs), which consist of only one parental polymer chain being internally crosslinked, have emerged as a versatile class of nanostructures inspired by the folding behaviour of naturally occurring macromolecules. Their unique properties have enabled applications in catalysis, sensing, and nanomedicine.1 Furthermore, they can be used for viscosity adjustment2,3 or polymer blending purposes.4–6 Recent advances have highlighted SCNPs as bioinspired nanoreactors for synthesizing nanomaterials and capturing CO2,7 showcasing their potential for sustainable applications.8 For instance, SCNPs have been utilized as nanoreactors for (photo)catalytic processes,9–11 leveraging their confined macromolecular environment12 similar to enzymatic systems.13 Novel pathways have been developed to generate SCNPs14 with enzyme-mimetic activity, demonstrating compartmentalized15 catalytic sites and reductase-like properties.16 Furthermore, SCNPs have been shown to act as multifunctional nanoreactors, with tailored polymeric pockets enhancing substrate recognition and catalytic efficiency.17Additionally, SCNPs have demonstrated catalytic functionality in biological media, enabling spatially controlled reactions for therapeutic applications.18 The versatility of SCNP synthesis has expanded significantly, with studies emphasizing the role of precursor topology,19 crosslinking chemistry in achieving specific morphologies,20 as well as light-induced avenues to control their folding state.21 Computational studies have provided insights into chain folding,22 SCNP formation and SCNP behaviour, though they have been identified as a challenge in terms of being an effective link in SCNP research for guiding the experimental design toward unique topologies.23 Molecular dynamics MD simulations have revealed the impact of orthogonal chemistry on achieving compact and spherical SCNPs,24 and elucidated the exact structure of photocatalytic pocket design within complex SCNPs.25 Furthermore, simulations elucidate the interaction between SCNPs and lipid membranes.26 Molecular confinement effects during synthesis have been shown to influence SCNP morphology, particularly for ring precursors, which adopt crumpled globular conformations.27 Machine learning approaches have further elucidated the relationship between precursor sequence patterning and SCNP morphology, offering a platform for sequence-based design.28 Also the internal motion,29 diffusion30 or effect of shear rate on SCNP formation31 could be captured by simulations.
Early on, a taxonomy emerged that classified the macromolecular construction principles underpinning SCNP design, most notably repeat unit folding (where the folding units are randomly dispersed in the parent polymer chain) and selective point folding (where the folding only occurs between specifically placed folding units at defined positions within the prepolymer chain).32,33 However, the folding of block copolymer systems has neither been systematically classified nor explored in detail, although such systems are offering unique opportunities, including the design of highly bespoke catalytic pockets to progress specific (photo)catalytic reactions or the design of SCNP morphologies – including pancake-like, sandwich-like or pom–pom structures – otherwise not accessible.
Thus, based on the above noted advances in SCNP technology34,35 and the opportunities in block precopolymer design, the folding behaviour of diblock copolymers into SCNPs awaits detailed exploration and exploitation.
As an initial critical step, we investigate the influence of interblock interaction potentials on the intra-molecular crosslinking and resulting SCNP structure using pseudo-kinetic Monte Carlo (MC) simulations.36–38 “Reactive” MC simulations were employed before.39–41 With this strategy, we address gaps in understanding SCNP structure formation and aim to contribute to the rational design of SCNPs for advanced applications, such as effective surfactants.42
Our work was inspired by the observation that block copolymers made of poly(dimethylaminoethyl methacrylate) (PDMAEMA) and poly(propylene oxide) (PPO) show an attractive interaction between the monomers of the different blocks, which is especially visible at interfaces and for advanced macromolecular architectures like branched miktoarm star polymers.43–45 In earlier MC simulation studies, we investigated the conformational changes and intramolecular complexation behaviour of these star polymers, by using a bead spring polymer model and similar generic interaction potentials as employed in this study.46,47 For the sake of simplicity, we assumed good solvent conditions for each block (which is a reasonably good approximation for PPO and PDMAEMA in water at low temperature, where both constituents are soluble, while both polymers phase separate at higher temperatures). While subtle conformational changes occur already at lower attractive interaction strengths, a pronounced collapse of the polymer chain primarily takes place for a larger attractive interaction, and a strong reduction in the radius of gyration and the end-to-end distance is only apparent at ε ≥ 0.6kBT (see also Fig. S1 and S2 in the SI). Of note, the amount of complexed monomer units taken from the simulation and (low temperature) experimental results are in line when assuming an interaction strength ε of approximately 0.6kBT for PDMAEMA and PPO units.48
Here, we focus on the question of whether this special behaviour as seen for PPO and PDMAEMA could be used to generate advanced polymer architectures, which are challenging to obtain by other means. More specifically, we investigate the generation of SCNPs from diblock copolymers, where the number and position of crosslinks and the resulting SCNP topology can be altered by adjusting the attractive interactions between the two blocks.
Experimentally, the crosslinking occurs either thermally or photochemically in the diluted state,49–51 providing control in bridging only between the blocks by advanced photochemistry.52 In addition, tuning of the interaction strength can be achieved by varying the temperature for thermo-responsive polymers like PPO and PDMAEMA (also see our previous paper for a structural picture of the complex;44 alternatively, electrostatic contributions in the case of polyampholyte block copolymers could also lead to tuneable behaviour dependent on pH and salt). Finally, different SCNP structures could result, as also demonstrated experimentally by introducing hydrophobic substituents leading to chain collapse before crosslinking.53
In this study we use a bead spring model for the diblock co-polymer and perform Monte Carlo simulations with the possibility to form additional bonds between crosslinkers during the simulation. With this, we combine aspects of our previous simulation work on block-type copolymers45,46,48,54,55 and of our simulation work on nanogels56–60 as SCNPs can be regarded as the smallest microgels, consisting of just one chain. The results from the MC simulations are compared to MD simulations of the same model (see the SI).
So far, we have focused on a scenario with equidistant locations of the crosslinkable monomer bead (every tenth bead rendered crosslinkable), which represents the average expected distribution of an ideal, azeotropic copolymerization of the main monomer and a crosslinkable monomer. Stochastic sequence variations61,62 will be an issue for future investigations.
The details of the model as well as the parameters for the simulation are given in Section 2 and our results are shown and discussed in Section 3.
In the first part we show the results of the dynamic crosslinking and SCNP formation under various conditions. We compare the resulting distributions of the crosslinks to a perfect ladder-type polymer and to a randomly crosslinked chain, i.e. a polymer, where all crosslinking options are treated equally, irrespective of the location along the chain.
In the second part, we investigate the structures, which we obtained, and compare them again to the structures of perfect ladder-type polymers.
In the last part of the results section, we look at the dynamics of the crosslink-formation under different conditions.
2. Model and simulation parameters
The diblock copolymer was modeled as a bead-spring chain, consisting of 100 beads of type A and 100 beads of type B. To account for excluded volume, all beads interact with a hard-sphere potentialwhere rij is the distance between beads i and j and σ is the bead diameter σ = 0.4 nm.
For chain connectivity, consecutive beads along the polymer chain additionally interact with a harmonic bond potential
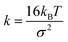 , rij being the actual distance between the two beads and r0 = 1.25 σ = 0.5 nm being the equilibrium distance.
, rij being the actual distance between the two beads and r0 = 1.25 σ = 0.5 nm being the equilibrium distance.
Every tenth bead in the chain, beginning with bead 5, was designated as a crosslinkable bead. Each of these crosslinkable beads can form a new permanent bond with a corresponding crosslinkable bead from the other block during the simulation (see Fig. 1).
 | ||
| Fig. 1 Structure of the polymer investigated with numbered beads for crosslinking, while bridges can only be established between the blue and red block (between bright blue and green beads). | ||
Additionally, we implemented an attractive potential for interactions between beads of type A and beads of type B. The potential depth εij was varied in three increments, ranging from 0.0kBT (indicating no attractive interaction) to 1.2kBT (representing a relatively strong attractive interaction). In this context, we employed the attractive component of a truncated Lennard-Jones potential in conjunction with the hard sphere potential.
A single diblock chain was placed in a simulation box with a length of 300σ (equivalent to 120 nm; with rcut = 150σ, or 60 nm). Monte Carlo simulations were conducted in the canonical (NVT) ensemble, utilizing exclusively single-particle translational moves with a maximum displacement of 0.6 nm. This approach was implemented to prevent artificial movement of larger chain segments and to achieve pseudo-kinetic behavior in the evolution of chain configurations. The simulation temperature was set at T = 298.15 K.
All other simulations were conducted in four distinct phases. In the first phase, the system was equilibrated for 0.5 × 106 Monte Carlo (MC) steps at 0.6kBT and for 106 MC steps at both 0.0kBT and 1.2kBT. The second phase involved setting new crosslinks during a crosslinking run, which consisted of 106 MC steps at 0.0kBT, 0.5 × 106 MC steps at 0.6kBT, and 0.1 × 106 MC steps at 1.2kBT. In the third phase, the resulting SCNP was equilibrated for an additional 106 MC steps. Finally, in the fourth phase, we evaluated structural parameters such as end-to-end distance (Ree) and radius of gyration (Rg) for the equilibrated SCNP over another 106 MC steps.
In the second phase, permanent (chemical) crosslinks were established with a probability of 1.0 if two previously uncrosslinked crosslinkable beads were found within a distance smaller than the critical crosslinking distance (rdistcl = 1.25σ for εij = 1.2kBT, or rdistcl = 1.50σ for all other cases). For the crosslinks, the same harmonic potential is used as for the consecutive beads along the chain.
For each parameter set in the SCNP simulations, we performed 200 simulations using different seeds for the random number generator to ensure robust statistical representation of the formed SCNPs. These simulations were executed with a modified version of the modular simulation software Molsim.63
To facilitate comparison with our SCNP results, we also conducted structural analyses on linear diblock copolymers exhibiting AB attraction and on preformed ladder polymers (with crosslinks established between beads 5 and 195, 15 and 185, etc.). For these polymer systems, we determined the end-to-end distance and radius of gyration following equilibration of the chains. To exclude an influence of the chosen method on the final structure of the SCNP, we performed additional molecular dynamics (MD) simulations for comparison (see the SI).
3. Results and discussion
3.1. Dynamic crosslinking simulations
Crosslinking simulations were conducted with a 100% crosslinking probability upon encountering crosslinkable beads within the critical threshold radius rdistcl. In the following, we focus on simulations with interaction energies ε of 0.0kBT, 0.6kBT and 1.2kBT for the attractive Lennard-Jones potential between beads of different particle type. For each set of parameters, 200 independent dynamic crosslinking procedures were simulated to obtain a statistical distribution of SCNP topologies.The resulting averaged microstructures were analyzed by generating heat maps that depict the frequency with which each single crosslinking option is realized (central column Fig. 2). In addition, the contact frequency of all possible pairs of crosslinkers in the absence of crosslinking (left column Fig. 2) and the initial three crosslinks at the onset of the SCNP formation (right column Fig. 2) were also identified. Our analysis reveals that an increased interaction strength leads to an overall higher contact probability among crosslinkable beads (as shown in the left column of Fig. 2). Notably, a partially antiparallel arrangement of blocks appears to be favored at lower interaction strengths. This preference can be attributed to the inherent monomer sequence along the chain of the copolymers and is translated into the resulting 3D-configuration of the macromolecule. Initial crosslinks are preferentially formed between monomers that are in close proximity because of their arrangement along the chain; thus, crosslinkers located near the block junctions tend to connect first—an observation consistent regardless of interaction strength. However, as the interaction strength increases, the likelihood of establishing one of the first crosslinks far away from the block junction increases. This is evidenced by a pronounced brightness in the bottom right corner of the heat map at ε = 0.0kBT and a homogeneous color of the heat map at ε = 1.2kBT (refer to Fig. 2(c) and (i)). Such behavior can be explained by a more globular structure arising from sufficiently high attraction between blocks, which facilitates closer proximity among all crosslinkable beads.
Interestingly, going from no attraction to weak attraction has only marginal influence on the average structure of the resulting SCNP as the heat maps look rather identical for 0.0kBT and 0.6kBT (Fig. 2(b) and (e)). However, it is worth noting that brighter regions appear in both upper right and lower left corners when attractive interactions are absent, indicating a lower probability of crosslinks between central crosslinker beads and crosslinker beads at the chain end of the other block. Even though a substantial number of initial crosslinks are set “off-center” at intermediate and high interaction energies, these “off-center” crosslinks will also be set at a later stage of the SCNP formation in the case of absent interblock interactions. Nevertheless, distinct internal structural characteristics emerge under conditions with no attractive interaction versus those with high attractive interactions. There is a clear trend when increasing the interaction strength even further. While a more ladder-like structure is observed for ε = 0.0kBT, a more random crosslinking becomes prevalent at ε = 1.2kBT, with nearly identical probabilities for crosslinking for all mutually crosslinkable monomers indicating a more compact structure.
The interaction strength significantly influences the internal structure of SCNPs, as evidenced by the frequency distribution of distances between crosslinks (see Fig. 3). Notably, a distance of 10 monomers between crosslinks is frequently observed, with a local maximum at this value. This preference arises because the most likely crosslink occurs closest to the junction point of the two blocks (specifically, bead 95 linking to bead 105). In contrast, this 10-monomer distance cannot be established with any other crosslinking configuration. Similarly, a distance of 190 monomers can only occur once (between bead 5 and bead 195), resulting in the lowest probability among all measured distances. Intermediate distances reveal several possibilities, with a maximum frequency observed near a distance of 110 monomers. Theoretically, one would expect a most frequent spacing of approximately 100 monomers (as illustrated in Fig. 3 ). This peak becomes more pronounced with increased interaction strengths and shifts closer to the theoretical value of 100. This behavior aligns with an idealized triangular distribution that reflects the number of available crosslinking options for specific spacings.
). This peak becomes more pronounced with increased interaction strengths and shifts closer to the theoretical value of 100. This behavior aligns with an idealized triangular distribution that reflects the number of available crosslinking options for specific spacings.
The average distance between the crosslinked monomers is 84.7 for 0.0kBT and 96.8 for 0.6kBT (the difference to the average 100 can probably be explained by the incomplete crosslinking for some of the structures). It appears that molecules lacking interblock attraction exhibit enhanced crosslinking among monomers positioned at shorter distances within the molecule, whereas those with interblock attraction yield averages closer to theoretical expectations.
Thus, we conclude that higher interaction strengths promote an “ideal” statistical crosslinking scenario where all crosslinkable beads are treated equivalently. This phenomenon can be explained by the more compact conformations observed under stronger interactions; in these cases, the positioning of beads along the chain exerts less influence on their precise locations within the partially collapsed globule surrounded by other beads. Conversely, extended conformations tend to inhibit contacts between beads that are situated far from block junctions.
3.2. Ladder-type polymer and comparison of structures
Despite the increased randomness in crosslinking observed under attractive conditions, the antiparallel diagonal of the heat maps suggests a significant contribution from zipping mechanisms that lead to SCNPs with ladder-type sequences. An extreme case of such SCNPs would resemble a perfect “ladder polymer”,64 where bead 95 links to bead 105, bead 85 to bead 115, bead 75 to bead 125, … until bead 5 links to bead 195. To further explore this phenomenon, we also investigated an ideal ladder-type polymer that does not exhibit any topological through-space connections between the formed rings; specifically, it lacks intramolecularly catenated rings (see Fig. 4).65A comparison of the ideal ladder-type polymer to the dynamically crosslinked SCNPs allowed the evaluation of the impact of stochastic crosslinking versus controlled, predetermined crosslinking strategies. Regarding the end-to-end distance, the ladder polymers show the smallest value, since the two ends are captured by the nearest crosslink (bead 5 with bead 195). Consequently, the end-to-end distance may not serve as an appropriate metric for comparing the compactness of SCNPs with that of ladder polymers. Conversely, attractive interactions are effectively captured in both end-to-end distance and radius of gyration, as the structures appear smaller with increased ε. When considering the radius of gyration as a more reliable indicator of compactness, it becomes evident that ladder polymers are generally less compact than their dynamically crosslinked SCNP counterparts, aside from a few outliers (Fig. 5). This observation can be attributed to the relatively extended conformation of the double strand in ladder polymers, which results in a halving of contour length. Thus, dynamically crosslinked SCNPs demonstrate significantly greater compactness compared to the extreme case of a pure ladder polymer though ladder-type sequences can be found in the SCNPs investigated here (see e.g. “double strand” in Fig. 4(b), left hand side). The observed outliers are often indicative of inadequately crosslinked SCNPs, which predominantly occur at lower interaction strengths. Nevertheless, 80% of the SCNPs exhibit a minimum of 9 crosslinks, even at ε = 0.0kBT. By comparing the SCNPs after their synthesis conditions were released (i.e., by setting the interaction parameter ε to 0.0kBT for analysis following synthesis at ε = 0.6kBT, and so forth), we observe in Fig. 6 that SCNPs synthesized under higher ε values display a more compact structure when evaluated under identical equilibrated conditions (for further details, see the SI). This analysis reveals a significant dependence of the resulting SCNP structures on the synthesis conditions. We conclude that stronger attractions between the blocks lead to more random and compact configurations of the SCNPs, while ladder-type sequences are predominantly associated with lower interaction strengths. For some of the parameter sets, we analyzed the structures also in the inverse space by calculating the average form factors of up to 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 configurations for each of the 200 SCNPs after equilibration at 0.0kBT (averaged over 200 particles for each condition; Fig. 6). The formfactor will be helpful for comparison with corresponding scattering experiments. While the form factor of the uncrosslinked diblock resembles the one of a Gaussian chain (Debye form factor), the ladder-type polymer has signatures of a one-dimensional object with an intensity scaling close to a q−1 dependence (q−1.3). In between, we see a compaction towards a sphere for increasing interaction strength during crosslinking. The ladder-like sequences present in the SCNPs are not so dominant and exposed that a hint of a q−1 scaling would be discernable in the form factor plots. Nevertheless, the heat maps unequivocally demonstrate the presence of such zipped structures. These can only be directly seen in the scattering for higher contents of these structures, though the form factor obtained for structures “synthesized” at low interaction energies are closest to the ladder type polymers.
000 configurations for each of the 200 SCNPs after equilibration at 0.0kBT (averaged over 200 particles for each condition; Fig. 6). The formfactor will be helpful for comparison with corresponding scattering experiments. While the form factor of the uncrosslinked diblock resembles the one of a Gaussian chain (Debye form factor), the ladder-type polymer has signatures of a one-dimensional object with an intensity scaling close to a q−1 dependence (q−1.3). In between, we see a compaction towards a sphere for increasing interaction strength during crosslinking. The ladder-like sequences present in the SCNPs are not so dominant and exposed that a hint of a q−1 scaling would be discernable in the form factor plots. Nevertheless, the heat maps unequivocally demonstrate the presence of such zipped structures. These can only be directly seen in the scattering for higher contents of these structures, though the form factor obtained for structures “synthesized” at low interaction energies are closest to the ladder type polymers.
 | ||
| Fig. 6 Representations of the form factors calculated for the structures obtained for different “synthesis” conditions (uncrosslinked diblock, crosslinked at 0.6 and 1.2kBT, ideal ladder polymer). | ||
Finally, we compared the MC results to “reactive” MD simulations in order to “validate” the independence of the structural outcome of the simulation method used in this paper.66 Both the observed trends in the end-to-end distances and especially in the radii of gyration (see the SI, Fig. S12) and the trends shown in the heat maps are reproduced by the MD approach (Fig. S13).
3.3. Pseudo-kinetics of SCNP formation
Finally, the crosslinking process was monitored as a function of the elapsed Monte Carlo steps (see Fig. 7), providing insights into the pseudo-kinetics of SCNP formation. The distinction between different interaction strengths becomes particularly pronounced, as crosslinking is facilitated in the presence of an attractive potential, even when reducing rdistcl to 5.0 Å at ε = 1.2kBT. This is evident from the plot showing the number of simulation steps required to establish the last crosslink (Fig. 7(a)). Here, an apparent “autocatalytic” behaviour can be observed, as the production of the final SCNPs is reduced and rather unlikely at low step numbers, but the production rate of fully crosslinked SCNPs is increased at intermediate step numbers, (wrongly) suggesting an increasing rate constant during the course of the reaction. In the end, the conformational freedom becomes limited to find one of the few remaining crosslinking partners for some topologically demanding SCNPs, particularly at higher ε. While one might anticipate that initial crosslink formation facilitates further crosslinking within the same molecule due to increased compaction, this “autocatalytic” principle does not hold true in our study, as discussed in the following paragraph. Analyzing the number of steps between crosslinks, the short step numbers become increasingly prevalent with increasing interaction strengths (Fig. 7(b)). These short step counts between crosslinks originate most likely from the small to intermediate formation times, before existing crosslinks impede further crosslinking because of steric restriction caused by the crosslinking.An examination of the Monte Carlo steps taken to establish each n-th crosslink (Fig. 7(c)) produced a diagram indicating nearly linear dependence for molecules without interblock attraction and a nonlinear curve for those with interblock attraction, which exhibits a deceleration toward the end of SCNP formation (refer to the SI for a non-logarithmic representation in Fig. S8). Hence, a real acceleration in crosslinking is not seen during the course of the simulation toward single SCNPs, however, an impression of pseudo-autocatalytic behavior regarding overall SCNP generation can be inferred from Fig. 7(a). This reduction during final SCNP formation occurs despite each crosslinking event resulting in a more compact structure with closer proximity among other crosslinkable monomers. Concurrently, reduced conformational freedom counterbalances this effect, as illustrated in Fig. 7(c). Notably, in cases involving attractive interactions, crosslinking slows down during the final phase of SCNP generation despite overall acceleration attributed to attractive interactions between both blocks.
Conclusions
This study investigates the folding and crosslinking behaviour of dilute, coarse-grained diblock copolymers that carry crosslinkable units along the chain, which can only be crosslinked between the two blocks (interblock crosslinks). This crosslinking leads to SCNPs with an optional amphiphilic and Janus-like nature. The influence of interblock interaction strength and dynamic crosslinking on chain conformation was explored. Simulations revealed that an increasing interaction strength leads to more compact structures, quantified by a decrease in the end-to-end distance and radius of gyration. Analysis of contact frequencies between crosslinker pairs identified preferential crosslinking between spatially proximate crosslinkers, particularly in an antiparallel ladder-type arrangement. However, dynamically crosslinked polymers exhibited a wider range of conformations compared to predefined “ladder” polymers, highlighting the impact of the stochastic crosslinking processes. An increase in interaction energy between both blocks leads to an accelerated and a more random crosslinking, where also beads far away from the block junction have a certain probability to crosslink at the beginning of the SCNP formation. This observation provides valuable information for designing specific crosslinking patterns for tailored nanostructures from diblock copolymers. We propose that imperfect ladder polymers can be obtained by the dynamic crosslinking process described here, in analogy to the hyperbranched polymers being the imperfect version of dendrimers. This study also emphasizes the ability of Monte Carlo simulations to determine the probability of folded polymeric structures by comparison between theoretically possible (ladder-like) polymers and those formed statistically. Finally, our study could guide synthesis, where negligible attraction between the blocks or even repulsion between the blocks could pave the way to ladder-type SCNPs, while stronger attraction under synthesis conditions would result in more compact SCNPs. In future, we plan to experimentally investigate the PPO-b-PDMAEMA system in more detail. Simulations using effective interaction potentials could further improve the comparison with the experimental findings. We plan to estimate noncovalent interaction parameters for pairs of different polymer building blocks. The parametrization will be performed using computationally efficient yet accurate quantum-chemical methods, such as r2SCAN-3c67 or double-hybrid density functionals.68 Solvation effects will be implicitly included in the interaction parameters through the use of appropriate implicit solvation models during the quantum-chemical calculations. This approach will allow us to replace the generic potentials used in our simulations, with effective potentials from atomistic models while maintaining computational efficiency. Furthermore, statistical variation of the monomer sequence during simulations will be a major advance, though one needs to select a representative set of copolymers to subsequently form the SCNP formation therefrom (in our case, there are ∼2 × 1013 different monomer sequences for just one block!). This task will be the subject of our further research.Author contributions
Anja Voigt: investigation, data curation, formal analysis, software, visualization, writing – original draft. Christian Strauch: conceptualization, data curation, methodology, software, supervision. Tom Höfkens: investigation, data curation, methodology, software. Mirco Wahab: resources, software, supervision. Wiebke Hadwich: resources, visualization. Christopher Barner-Kowollik: conceptualization, project administration, writing – review and editing. Conrad Hübler: resources, data curation, methodology, software, validation, supervision, writing – review and editing. Stefanie Schneider: conceptualization, supervision, writing – review and editing. Felix A. Plamper: conceptualization, project administration, supervision, writing – original draft, writing – review and editing.Conflicts of interest
There are no conflicts of interest to declare.Data availability
Data for this article, including input files and files with extracted data used for plots, are included in the supplementary information (SI) as a zip-file. Supplementary information: snapshots on chain conformation, contact frequencies, different representations of figures in the main part, and pseudo-kinetic plots. Zip-file with input data and extracted data. See DOI: https://doi.org/10.1039/d5sm01153a.Acknowledgements
The authors acknowledge computing time on the compute cluster of the Faculty of Mathematics and Computer Science of Technische Universität Bergakademie Freiberg, operated by the computing center (URZ) and funded by the German Research Foundation (Deutsche Forschungsgemeinschaft, DFG) under DFG grant number 397252409. Furthermore, the authors thank the DFG for funding in the frame of the experimental project 524806266 (PL 571/14-1), which gave the incentive for the current work. C.B.-K. acknowledges the Alexander-von-Humboldt Foundation for a Professorial Fellowship. The introduction was partly written with help of scienceOS (March 2025 version, https://www.scienceOS.ai) and https://deepL.com was sometimes used to translate from a German report.References
- D. Arena, C. Nguyen, L. M. A. Ali, E. Verde-Sesto, A. Iturrospe, A. Arbe, U. İşci, Z. Şahin, F. Dumoulin, M. Gary-Bobo and J. A. Pomposo, Amphiphilic Single-Chain Polymer Nanoparticles as Imaging and Far-Red Photokilling Agents for Photodynamic Therapy in Zebrafish Embryo Xenografts, Adv. Healthcare Mater., 2024, 13, e2401683 CrossRef PubMed.
- Y. Ruan, Q. Zou, H. Zhang, Y. Zhang, H. Zhang, W. Wu, H. Liu, J. Yan and G. Liu, The Extreme of Disentanglement: 100-fold Viscosity Reduction in Model Single-Chain Nanoparticles, Macromolecules, 2024, 57, 6583–6592 CrossRef CAS.
- C. Pyromali, N. Patelis, M. Cutrano, M. Gosika, E. Glynos, A. J. Moreno, G. Sakellariou, J. Smrek and D. Vlassopoulos, Nonmonotonic Composition Dependence of Viscosity upon Adding Single-Chain Nanoparticles to Entangled Polymers, Macromolecules, 2024, 57, 4826–4832 CrossRef CAS PubMed.
- Y. Chen, Z. Hu and H. Pu, Microscopic studies on remarkable rheological behavior of single-chain nanoparticles and linear polymer blends, Eur. Polym. J., 2025, 224, 113687 CrossRef.
- P. Zhao, Z. Yan, Y. Liang and R. Zhang, Structures and Segmental Dynamics in Single-Chain Polymer Nanoparticles-Based All-Polymer Nanocomposites, Macromolecules, 2025, 58, 3478–3487 CrossRef CAS.
- Y. Fang, J. Liu, Z. Zhu, Z. Lv and Y. Lin, Crystallization Kinetics of Poly(ethylene oxide) in All-Polymer Nanocomposites Based on Poly(methyl methacrylate) Single-Chain Nanoparticles, ACS Appl. Polym. Mater., 2024, 6, 12217–12227 CrossRef CAS.
- W. Fan, X. Tong, F. Farnia, B. Yu and Y. Zhao, CO 2 -Responsive Polymer Single-Chain Nanoparticles and Self-Assembly for Gas-Tunable Nanoreactors, Chem. Mater., 2017, 29, 5693–5701 CrossRef CAS.
- J. Rubio-Cervilla, E. González and J. A. Pomposo, Advances in Single-Chain Nanoparticles for Catalysis Applications, Nanomaterials, 2017, 7, 341 CrossRef PubMed.
- J. Pinacho-Olaciregui, E. Verde-Sesto, D. Taton and J. A. Pomposo, Gold Nanoclusters Synthesized within Single-Chain Nanoparticles as Catalytic Nanoreactors in Water, Polymers, 2024, 16, 378 CrossRef CAS PubMed.
- K. Mundsinger, A. Izuagbe, B. T. Tuten, P. W. Roesky and C. Barner-Kowollik, Single Chain Nanoparticles in Catalysis, Angew. Chem., Int. Ed., 2024, 63, e202311734 CrossRef CAS PubMed.
- M. Nagao, K. Mundsinger and C. Barner-Kowollik, Photoinduced Energy/Electron Transfer within Single-Chain Nanoparticles, Angew. Chem., Int. Ed., 2025, 64, e202419205 CrossRef CAS PubMed.
- H. Frisch, J. P. Menzel, F. R. Bloesser, D. E. Marschner, K. Mundsinger and C. Barner-Kowollik, Photochemistry in Confined Environments for Single-Chain Nanoparticle Design, J. Am. Chem. Soc., 2018, 140, 9551–9557 CrossRef CAS PubMed.
- C. K. Lyon, A. Prasher, A. M. Hanlon, B. T. Tuten, C. A. Tooley, P. G. Frank and E. B. Berda, A brief user's guide to single-chain nanoparticles, Polym. Chem., 2015, 6, 181–197 RSC.
- J. He, L. Tremblay, S. Lacelle and Y. Zhao, Preparation of polymer single chain nanoparticles using intramolecular photodimerization of coumarin, Soft Matter, 2011, 7, 2380–2386 RSC.
- J. F. Thümmler and W. H. Binder, Compartmentalised single-chain nanoparticles and their function, Chem. Commun., 2024, 60, 14332–14345 RSC.
- I. Perez-Baena, F. Barroso-Bujans, U. Gasser, A. Arbe, A. J. Moreno, J. Colmenero and J. A. Pomposo, Endowing Single-Chain Polymer Nanoparticles with Enzyme-Mimetic Activity, ACS Macro Lett., 2013, 2, 775–779 CrossRef CAS PubMed.
- H. Rothfuss, N. D. Knöfel, P. W. Roesky and C. Barner-Kowollik, Single-Chain Nanoparticles as Catalytic Nanoreactors, J. Am. Chem. Soc., 2018, 140, 5875–5881 CrossRef CAS PubMed.
- Y. Liu, S. Pujals, P. J. M. Stals, T. Paulöhrl, S. I. Presolski, E. W. Meijer, L. Albertazzi and A. R. A. Palmans, Catalytically Active Single-Chain Polymeric Nanoparticles: Exploring Their Functions in Complex Biological Media, J. Am. Chem. Soc., 2018, 140, 3423–3433 CrossRef CAS PubMed.
- L. Zhang, X.-Z. Zhang, J.-T. Lyu, L.-X. Yu, C.-Y. Wang, Z.-Y. Sun, Z.-Y. Lu and H.-J. Qian, Surface-Cross-linked Protein-like Single-Chain Nanoparticle Globules Unexpectedly Stabilized with a Low Cross-linking Degree, Macromolecules, 2024, 57, 858–868 CrossRef CAS.
- R. Chen and E. B. Berda, 100th Anniversary of Macromolecular Science Viewpoint: Re-examining Single-Chain Nanoparticles, ACS Macro Lett., 2020, 9, 1836–1843 CrossRef CAS PubMed.
- P. H. Maag, F. Feist, V. X. Truong, H. Frisch, P. W. Roesky and C. Barner-Kowollik, Visible-Light-Induced Control over Reversible Single-Chain Nanoparticle Folding, Angew. Chem., Int. Ed., 2023, 62, e202309259 CrossRef CAS PubMed.
- S. Wijker, D. Dellemme, L. Deng, B. Fehér, I. K. Voets, M. Surin and A. R. A. Palmans, Revealing the Folding of Single-Chain Polymeric Nanoparticles at the Atomistic Scale by Combining Computational Modeling and X-ray Scattering, ACS Macro Lett., 2025, 14, 428–433 CrossRef CAS PubMed.
- A. M. Hanlon, C. K. Lyon and E. B. Berda, What Is Next in Single-Chain Nanoparticles?, Macromolecules, 2016, 49, 2–14 CrossRef CAS.
- F. Lo Verso, J. A. Pomposo, J. Colmenero and A. J. Moreno, Multi-orthogonal folding of single polymer chains into soft nanoparticles, Soft Matter, 2014, 10, 4813–4821 RSC.
- K. Mundsinger, B. T. Tuten, L. Wang, K. Neubauer, C. Kropf, M. L. O'Mara and C. Barner-Kowollik, Visible-Light-Reactive Single-Chain Nanoparticles, Angew. Chem., Int. Ed., 2023, 62, e202302995 CrossRef CAS PubMed.
- Y. Guo, M. Werner, W. Li, J.-U. Sommer and V. A. Baulin, Shape-Adaptive Single-Chain Nanoparticles Interacting with Lipid Membranes, Macromolecules, 2019, 52, 9578–9584 CrossRef CAS.
- M. Formanek and A. J. Moreno, Effects of precursor topology and synthesis under crowding conditions on the structure of single-chain polymer nanoparticles, Soft Matter, 2017, 13, 6430–6438 RSC.
- R. A. Patel, S. Colmenares and M. A. Webb, Sequence Patterning, Morphology, and Dispersity in Single-Chain Nanoparticles: Insights from Simulation and Machine Learning, ACS Polym. Au, 2023, 3, 284–294 CrossRef CAS PubMed.
- M. J. A. Hore, Analysis of the internal motions of thermoresponsive polymers and single chain nanoparticles, Soft Matter, 2025, 21, 770–780 RSC.
- B. Mei, A. J. Moreno and K. S. Schweizer, Unified Understanding of the Structure, Thermodynamics, and Diffusion of Single-Chain Nanoparticle Fluids, ACS Nano, 2024, 18, 15529–15544 CrossRef CAS PubMed.
- M. D. Chertok, H. A. Stone and M. A. Webb, Effect of shear flow and precursor polymer design on single-chain nanoparticle formation, Soft Matter, 2025, 21, 8265–8278 RSC.
- O. Altintas and C. Barner-Kowollik, Single chain folding of synthetic polymers by covalent and non-covalent interactions: current status and future perspectives, Macromol. Rapid Commun., 2012, 33, 958–971 CrossRef CAS PubMed.
- O. Altintas and C. Barner-Kowollik, Single-Chain Folding of Synthetic Polymers, Macromol. Rapid Commun., 2016, 37, 29–46 CrossRef CAS PubMed.
- E. Guazzelli, E. Martinelli, G. Galli, L. Cupellini, S. Jurinovich and B. Mennucci, Single-chain self-folding in an amphiphilic copolymer: An integrated experimental and computational study, Polymer, 2019, 161, 33–40 CrossRef CAS.
- H. Rabbel, P. Breier and J.-U. Sommer, Swelling Behavior of Single-Chain Polymer Nanoparticles: Theory and Simulation, Macromolecules, 2017, 50, 7410–7418 CrossRef CAS.
- F. A. Detcheverry, D. Q. Pike, P. F. Nealey, M. Müller and J. J. de Pablo, Monte carlo simulation of coarse grain polymeric systems, Phys. Rev. Lett., 2009, 102, 197801 CrossRef PubMed.
- K. Binder and W. Paul, Monte Carlo simulations of polymer dynamics: Recent advances, J. Polym. Sci., Part B:Polym. Phys., 1997, 35, 1–31 CrossRef CAS.
- K. Binder and W. Paul, Recent Developments in Monte Carlo Simulations of Lattice Models for Polymer Systems, Macromolecules, 2008, 41, 4537–4550 CrossRef CAS.
- E. Norouzi Farahani, S. Arzemanzadeh, M. Mahnama and E. Hosseinian, Atomistic insights into the mechanical properties of cross-linked Poly(N-isopropylacrylamide) hydrogel, Polymer, 2024, 297, 126798 CrossRef CAS.
- A. Malani, S. M. Auerbach and P. A. Monson, Monte Carlo Simulations of Silica Polymerization and Network Formation, J. Phys. Chem. C, 2011, 115, 15988–16000 CrossRef CAS.
- P. M. Blanco and P. Košovan, The explicit bonding reaction ensemble Monte Carlo method, J. Chem. Phys., 2024, 161, 094906 CrossRef CAS PubMed.
- S. Li, S. Sun, J. Luo, Z. Xia, Y. Wang, T. P. Russell, S. Shi and Z. Yang, Polymer Single-Chain Nanoparticles: Shaping Solid Surfactants, Macromol. Rapid Commun., 2024, 45, e2400393 CrossRef PubMed.
- A. A. Steinschulte, B. Schulte, M. Erberich, O. V. Borisov and F. A. Plamper, Unimolecular Janus Micelles by Microenvironment-Induced, Internal Complexation, ACS Macro Lett., 2012, 1, 504–507 CrossRef CAS PubMed.
- A. A. Steinschulte, B. Schulte, S. Rütten, T. Eckert, J. Okuda, M. Möller, S. Schneider, O. V. Borisov and F. A. Plamper, Effects of Architecture on the Stability of Thermosensitive Unimolecular Micelles, Phys. Chem. Chem. Phys., 2014, 16, 4917–4932 RSC.
- A. Steinschulte, W. Xu, F. Draber, P. Hebbeker, A. Jung, D. Bogdanovski, S. Schneider, V. V. Tsukruk and F. A. Plamper, Interface-Enforced Complexation between Copolymer Blocks, Soft Matter, 2015, 11, 3559–3565 RSC.
- P. Hebbeker, F. A. Plamper and S. Schneider, Effect of the Molecular Architecture on the Internal Complexation Behavior of Linear Copolymers and Miktoarm Star Polymers, Macromol. Theory Simul., 2015, 24, 110–116 CrossRef CAS.
- P. Hebbeker, A. A. Steinschulte, S. Schneider and F. A. Plamper, Balancing Segregation and Complexation in Amphiphilic Copolymers by Architecture and Confinement, Langmuir, 2017, 33, 4091–4106 CrossRef CAS PubMed.
- P. Hebbeker, A. A. Steinschulte, S. Schneider, J. Okuda, M. Möller, F. A. Plamper and S. Schneider, Complexation in Weakly Attractive Copolymers with Varying Composition and Topology, Macromolecules, 2016, 49, 8748–8757 CrossRef.
- T. Ma, B. Mu and W. Tian, Concurrent Folding and Fluorescent Functionalization of Single-Chain Nanoparticles via Heterogeneous Photochemistry, Angew. Chem., Int. Ed., 2025, 64, e202500354 CrossRef CAS PubMed.
- Y. Vo, R. Raveendran, C. Cao, R. Y. Lai, M. Lossa, H. Foster and M. H. Stenzel, Solvent Choice during Flow Assembly of Photocross-Linked Single-Chain Nanoparticles and Micelles Affects Cellular Uptake, ACS Appl. Mater. Interfaces, 2024, 16, 59833–59848 CrossRef CAS PubMed.
- Y. Chen, Z. Hu, Z. Shen, X. Xue and H. Pu, Preparation of superstructured comb polymers based on tadpole-shaped single-chain nanoparticles, Chem. Sci., 2024, 15, 17590–17599 RSC.
- F. Feist, J. P. Menzel, T. Weil, J. P. Blinco and C. Barner-Kowollik, Visible Light-Induced Ligation via o -Quinodimethane Thioethers, J. Am. Chem. Soc., 2018, 140, 11848–11854 CrossRef CAS PubMed.
- M. Alqaisi, J. F. Thümmler, F. Lehmann, F.-J. Schmitt, L. Lentz, F. Rieder, D. Hinderberger and W. H. Binder, Tuning nanoparticles’ internal structure: fluorinated single-chain nanoparticles (SCNPs) generated by chain collapse of random copolymers, Polym. Chem., 2024, 15, 2949–2958 RSC.
- P. Hebbeker, T. G. Langen, F. A. Plamper and S. Schneider, Spacer Chains Prevent the Intramolecular Complexation in Miktoarm Star Polymers, J. Phys. Chem. B, 2018, 122, 4729–4736 CrossRef CAS PubMed.
- P. Hebbeker, F. A. Plamper and S. Schneider, Aggregation of Star Polymers, Macromol. Theory Simul., 2018, 27, 1800033 CrossRef.
- C. Strauch and S. Schneider, Ionisation and swelling behaviour of weak polyampholyte core–shell networks – a Monte Carlo study, Soft Matter, 2023, 19, 938–950 RSC.
- C. Strauch and S. Schneider, Monte Carlo simulation of the ionization and uptake behavior of cationic oligomers into pH-responsive polyelectrolyte microgels of opposite charge – a model for oligopeptide uptake and release, Soft Matter, 2024, 20, 1263–1274 RSC.
- C. Strauch, L. Roß and S. Schneider, Exploring guest molecule uptake in pH-responsive polyelectrolyte microgels via Monte Carlo simulations, Soft Matter, 2024, 20, 9664–9672 RSC.
- C. Hofzumahaus, P. Hebbeker and S. Schneider, Monte Carlo simulations of weak polyelectrolyte microgels, Soft Matter, 2018, 14, 4087–4100 RSC.
- C. Hofzumahaus, C. Strauch and S. Schneider, Monte Carlo simulations of weak polyampholyte microgels: pH-dependence of conformation and ionization, Soft Matter, 2021, 17, 6029–6043 RSC.
- K. A. Curtis, A. Statt and W. F. Reinhart, Predicting self-assembly of sequence-controlled copolymers with stochastic sequence variation, Soft Matter, 2025, 21, 2143–2151 RSC.
- A. A. Gavrilov and A. V. Chertovich, Copolymerization of Partly Incompatible Monomers: An Insight from Computer Simulations, Macromolecules, 2017, 50, 4677–4685 CrossRef CAS.
- J. Reščič and P. Linse, MOLSIM: A modular molecular simulation software, J. Comput. Chem., 2015, 36, 1259–1274 CrossRef PubMed.
- H. Balan, G. Sadasivan, E. Paul and K. M. Sureshan, Single-Crystal-to-Single-Crystal Synthesis of a Rope-Ladder Polymer, Angew. Chem., Int. Ed., 2025, 64, e202506699 CrossRef CAS PubMed.
- Y. Tezuka, Supramol. Polym. Chem., 2012, 293–303 CAS.
- W. Shinoda, M. Shiga and M. Mikami, Rapid estimation of elastic constants by molecular dynamics simulation under constant stress, Phys. Rev. B:Condens. Matter Mater. Phys., 2004, 69, 134103 CrossRef.
- S. Grimme, A. Hansen, S. Ehlert and J.-M. Mewes, r2SCAN-3c: A “Swiss army knife” composite electronic-structure method, J. Chem. Phys., 2021, 154, 64103 CrossRef CAS PubMed.
- S. Kozuch and J. M. L. Martin, Spin-component-scaled double hybrids: An extensive search for the best fifth-rung functionals blending DFT and perturbation theory, J. Comput. Chem., 2013, 34, 2327–2344 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |