 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Impact of protein corona morphology on nanoparticle diffusion in biological fluids: insights from a mesoscale approach
Beatrice
Cipriani
 *ab and
Hender
Lopez
*ab and
Hender
Lopez
 *a
*a
aSchool of Physics, Clinical and Optometric Sciences, Technological University Dublin, Grangegorman D07 ADY7, Ireland. E-mail: hender.lopezsilva@tudublin.ie
bFOCAS Research Institute, TU Dublin, City Campus, Dublin 8, Ireland
First published on 5th February 2026
Abstract
Nanoparticles (NPs) demonstrate considerable potential in medical applications, including targeted drug delivery and diagnostic probes. However, their efficacy depends on their ability to navigate through the complex biological environments inside living organisms. In such environments, NPs interact with a dense mixture of biomolecules, which can reduce their mobility and hinder diffusion. Understanding the factors influencing NP diffusion in these environments is key to improving nanomedicine design and predicting toxicological effects. In this study, we propose a computational approach to model NP diffusion in crowded environments. We introduce a mesoscale model that accounts for the combined effects of the protein corona (PC) and the crowded medium on NP mobility. By including volume-exclusion interactions and modelling the PC both explicitly and implicitly, we identify key macromolecular descriptors that affect NP diffusion. Our results show that the morphology of the PC can significantly affect the diffusion of NPs, and the roles of the occupied volume fraction and the size ratio between tracers and crowders are analysed. The results also show that approximating large macromolecular assemblies with a hydrodynamic single-sphere model leads to inexact diffusion estimates. To overcome the limitations of single-sphere representations, a strategy for an accurate parameterization of NP–PC systems using a single-sphere model is presented and the validity and limitations of the model are discussed.
1 Introduction
The potential use of nanoparticles (NPs) in biological, pharmaceutical, chemical, and medical fields has produced an interdisciplinary research field known as nanomedicine.1 It is worth noting that effective nanomedical applications rely on the NP's ability to target and navigate specific structures within living organisms. In this way, assessing the mobility of NPs is a critical factor that can influence potential applications, requiring the evaluation of several key aspects. Additionally, when NPs interact with biological fluids, they get immediately coated by a biomolecule rich layer known as the protein corona (PC) or biomolecule corona which determines its bioactivity.2 Its composition depends on several factors such as the NP's size, shape, material, and the biological medium content.2–4 However, it is clear that its formation leads to morphological changes that can result in anisotropic structures with patchy patterns on the surface,5–8 although the specific structure and organization of the PC depend on the NP type.9 As a result, NPs with a PC are unlikely to maintain an ideal, smooth spherical shape. This transformation impacts the NP's mobility and transport properties by altering its size, shape, interaction dynamics and overall its biological identity, thus influencing its bio-distribution and bio-availability.10–13 Furthermore, nowadays NPs can be designed with a multitude of shapes, such as spheres, flowers, or stars, and their surface morphology is often engineered to include features like spikes, virus- or raspberry-like patterns14–18 and it has been shown that the shape alone also affects their bioactivity.19 These findings suggest that simple sphere models might not be adequate to understand, predict and finally engineer NPs with a desirable diffusion behaviour for nanomedicine applications.Another factor to consider is that the biological environments where the transport happens introduce additional complexity to the dynamics of the NPs due to macromolecular crowding. Biological fluids, particularly within cells, are densely populated with proteins and other macromolecules that can occupy up to 40% of the total volume. Furthermore, these crowded environments are often polydisperse, complicating the analysis and study of transport properties. To understand the dynamics of NPs under such conditions, colloids theory for spherical particles is commonly used, an approach that is also applied to protein solutions.20 Over the years, this method has provided insights into protein phase diagrams, crystallization, self-assembly, aggregation, and diffusion.21–23 It is well known that colloidal systems at high concentrations display reduced diffusion rates due to particle interactions, including hydrodynamic effects, crowding, and collisions.24,25 Theories describing concentrated monodisperse soft sphere colloids and polymers have effectively captured these phenomena.26–29 However, for non-spherical particles, the application of these theories is by no means straightforward. Surface roughness and particle shape influence diffusion rates, with rough particles diffusing more slowly than smooth ones near walls in pure solvents.30 Two recent computational studies have emphasized the role of morphology in NP diffusion.31,32 The first study examined how the distribution, size, and morphology of functional groups on spherical NPs influence their translational diffusion both at infinite dilution and near rigid walls. The findings revealed that the transport properties of functionalized NPs are notably affected by the morphology of the attached groups. In the second study, it was shown that NPs with different anisotropic shapes display unique self-diffusion and sedimentation behaviours, with a stronger dependence on volume fraction compared to spheres. Moreover, it has been proved how in crowded media, interactions among rough colloids can lead to rotational arrest and phenomena such as second glass transition in both translational and rotational diffusion, which are absent for systems with smooth particles.33 While colloid models have successfully described the dynamics of some proteins under specific crowded conditions,23,24,34 they are typically restricted to spherical and isotropic particles in monodisperse solutions. More complex modelling approaches are often required to address anisotropic objects.20,25,35 For example, it has been demonstrated that anisotropic shapes and interactions significantly influence protein diffusion, particularly at high concentrations.36–38 All of these findings suggest that understanding NP diffusion in biological fluids requires integrating macromolecular crowding and PC-induced morphological features into existing models. NP diffusion plays a crucial role in determining the in situ properties of the PC, particularly its thickness. Hydrodynamic size measurements, often obtained through fluorescence-based techniques,39–43 rely on diffusion models that typically assume spherical particles. This simplification, commonly adopted in both experimental and simulation studies to enable more tractable modelling and analysis, treats complex structures as effective spheres.44 These are typically defined by an equivalent hydrodynamic radius, derived from the diffusion coefficient at infinite dilution of the non-spherical body. This approach is widely used to estimate PC thickness in biological fluids.45,46 Since these methods are grounded in diffusion theory, an accurate understanding of nanoparticle mobility is essential for reliable measurements.
In this context, this work aims to model the diffusion of NPs in polydisperse crowded environments. For this task, we developed a computational model to simulate NP dynamics in protein-rich media, accounting for macromolecular crowding and the presence of a PC on spherical NPs. We adopt a Coarse-Grained Meso-Scale (CG-MS) approach that ensures computational feasibility over the timescales relevant for the long-time diffusion. Here, proteins are modelled as spheres, and NPs with a PC are treated as single rigid objects using a raspberry-like model, a representation commonly used in computational studies of NPs with a PC.47–49 With this model, we then quantify how the PC properties and macromolecular crowding jointly affect NP diffusion. To achieve this, our analysis explores variations in PC morphology, spatial organization, and composition to assess their impact on diffusion. We compare explicit against implicit PC representation, with the aim of determining the conditions under which an equivalent single-sphere approximation is valid. Finally, we evaluated the role of the medium composition (monodisperse versus polydisperse crowders) and of the occupied volume fraction in NP dynamics in biological environments.
2 Methods
NPs when interacting with living organisms typically diffuse through biological fluids under crowded molecular conditions, often coated by a PC layer. Simulating such systems at the atomistic resolution level can be computationally expensive. For instance, considering a 20-nm diameter spherical NP diffusing in a box of 110 × 110 × 110 nm3 with crowders occupying a volume fraction of ϕ = 0.3 (typical of biological fluids) would require simulating between 1000–10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 proteins over time scales relevant to NP diffusion – usually on the order of several microseconds or even milliseconds. Employing full-atomistic or even moderately coarse-grained models to reach these timescales for a few hundred proteins (corresponding to ≈50
000 proteins over time scales relevant to NP diffusion – usually on the order of several microseconds or even milliseconds. Employing full-atomistic or even moderately coarse-grained models to reach these timescales for a few hundred proteins (corresponding to ≈50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–100
000–100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 residues) would result in a massive investment of computational resources. Moreover, when simulating a highly polydisperse system, we must account for multiple replicas to obtain robust statistics. In this work, to address these challenges, we adopted a mesoscale representation of the proteins in solution and within the PC, wherein each protein is represented by a single sphere. We stress that the hydrodynamic radius is an input parameter defining the spherical representation of proteins, and no alternative non-spherical representations are included for medium proteins. This approach preserves the tracer's morphological properties, allowing us to assess their contribution to diffusion, while with spherical crowders we can still evaluate different size contribution and concentration effects through volume exclusion interactions. The size of each sphere is set to the hydrodynamic radius of the protein computed in pure solvent. Using this approach enables efficient exploration of the NP's diffusion within complex biological environments while balancing computational costs and system comparability.
000 residues) would result in a massive investment of computational resources. Moreover, when simulating a highly polydisperse system, we must account for multiple replicas to obtain robust statistics. In this work, to address these challenges, we adopted a mesoscale representation of the proteins in solution and within the PC, wherein each protein is represented by a single sphere. We stress that the hydrodynamic radius is an input parameter defining the spherical representation of proteins, and no alternative non-spherical representations are included for medium proteins. This approach preserves the tracer's morphological properties, allowing us to assess their contribution to diffusion, while with spherical crowders we can still evaluate different size contribution and concentration effects through volume exclusion interactions. The size of each sphere is set to the hydrodynamic radius of the protein computed in pure solvent. Using this approach enables efficient exploration of the NP's diffusion within complex biological environments while balancing computational costs and system comparability.
2.1 Nanoparticle and protein corona (NP–PC) models
In this work, each NP is modelled as a sphere of 20 nm in diameter while the PC is represented by spheres of different sizes (each size representing a protein type) attached to the NP. These sizes were set to the hydrodynamic radius (rH) of the proteins obtained by computing the translational diffusion coefficient (Dt0) of each protein type at infinite dilution (i.e. in pure solvent) using HYDROPRO software (version 10)50 and then using the Stokes–Einstein relationship to calculate rH. The atomic coordinates for the atoms of the proteins used for these calculations were obtained from the Protein Data Bank51 when available, and alternatively from the Alphafold repository.52 The viscosity and density of the solvent were set to 0.01 poise and 1 g ml−1, respectively, while a temperature of 293 K was used in all calculations. The number of binding sites available on the surface of the NP, and therefore the number of proteins that can be forming the PC, was calculated for each protein type using the approach developed by Rouse and Lobaskin.47 This method accounts for the steric occupancy of every protein, and is developed as follows. First, the projected binding area of each protein that constitutes the PC is calculated with the formula,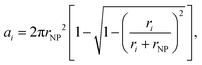 | (1) |
| Ai = [Ci]·4πrNP2, | (2) |
The first two systems modelled are 20-nm NPs with polydispersed plasma PCs. The two proteomic compositions of the PC were selected from experimental data, specifically from 20 nm citrate coated silver and gold NPs (Ag-CIT and Au-CIT NPs) as reported in ref. 56 (see Tables S1–S2 for the composition of the PCs). We will refer to these systems as P1 and P2 NPs. Fig. 1a and b show the relative occupancy of each protein in the corona and a rendering of the final system.
As mentioned above, different configurations of the same PC were generated. For both P1 and P2 NPs, 1000 different layouts of the PC were constructed. For each of these configurations of the NP with a PC, the hydrodynamic radius of the NP–PC was derived from the computed Dt0 using the Stokes–Einstein relationship. To investigate the role of PC's layout on the long-time diffusivity, from this set of 1000 configurations, we selected four for the production BD simulations in crowded medium. The four configurations were selected based on the following criteria: a NP–PC with the modal hydrodynamic size (t1), a NP–PC with the largest hydrodynamic radius (t2), a NP–PC with the smallest hydrodynamic radius (t3), and a randomly selected NP–PC (t4), as shown in Fig. S1 and S2. Additionally, we derived two simplified models, referred to as P1-S and P2-S, from the original P1 and P2 NP–PC systems. In these simplified models, the PC is composed of only three distinct protein sizes (Fig. 1c and d), in contrast to the full distribution of protein sizes present in the original P1 and P2 models (Fig. 1a and b). These three sizes were obtained by discretizing the original protein distributions into groups of proteins with similar hydrodynamic radii. For the P1-S model, the protein sizes were grouped into three representative radii: 2.8 nm, 5.0 nm, and 8.7 nm. The number of size types was chosen to preserve the overall hydrodynamic radius of the NP–PC, such that the resulting rH of the P1-S matches that of the original P1 NP–PC of 18.9 nm. Similarly, for the P2-S model, the simplified composition includes proteins with radii of 3.0 nm, 5.2 nm, and 8.6 nm, also preserving the same rH as the original P2 NP–PC of 22.3 nm. This reduction in complexity was implemented to test whether a simplified representation of the PC, in which the detailed size distribution is reduced but the overall hydrodynamic size and morphology are preserved, has any impact on tracer diffusion.
To assess whether the diffusion of the NPs can be influenced by the polydispersity of the PC, we modelled three more 20 nm NPs with monodisperse PCs, i.e. made only of one kind of protein (Fig. 1e). As protein types we chose three types: small size proteins (spheres of radius 2.6 nm), medium size proteins (spheres radius 5.5 nm) and large proteins (spheres of radius 8.7 nm). These sizes were selected as they are commonly found in both the P1 and P2 coronas (see Fig. 1a and b). We will refer to these systems as monodisperse small (MS), monodisperse medium (MM), and monodisperse large (ML) coronas. For these five systems (P1, P2, MS, MM and ML NPs) we perform BD simulations by treating the NP–PC complex as rigid-bodies (RBs) and also by modelling the NP–PC complex as a single sphere of size determined by the NP–PC hydrodynamic size (equivalent hydrodynamic single sphere, h-SS). In this way, instead of only representing the NP–PC system by the use of a multi-beads RB as shown in Fig. 1, we now also represent the NP–PC complex as a single sphere of which its size is derived using the Stokes–Einstein formula from the Dt0 calculated with HYDRO++ from the RB structure. Thus, we are able to simulate the PC both explicitly and implicitly, evaluating the validity of the h-SS approximation on NP diffusion. Finally, we modelled three more 20 nm NPs with a simplified polydisperse PCs, i.e. using only three protein types. The corona compositions of these three systems were not based on experimental data, but were artificially designed to reproduce three different morphologies – from a more compact homogeneous-shaped NP to a more rough and irregularly-shaped one (Fig. 1f). We will refer to these as P3, P4, and P5. A summary of all NP–PC systems is presented in Table 1.
| System | ID | r H [nm] |
|---|---|---|
| Ag-CIT NP with polydisperse PC | P1 t1* | 18.9 |
| P1 t2 | 19.0 | |
| P1 t3 | 18.7 | |
| P1 t4 | 18.8 | |
| Au-CIT NP with polydisperse PC | P2 t1* | 22.3 |
| P2 t2 | 22.5 | |
| P2 t3 | 22.1 | |
| P2 t4 | 22.3 | |
| P1 NP with simplified PC | P1-S | 18.9 |
| P2 NP with simplified PC | P2-S | 22.3 |
NP with polydisperse PC (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0 protein ratio) 0 protein ratio) |
P3 | 18.0 |
NP with polydisperse PC (2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 protein ratio) 1 protein ratio) |
P4 | 20.5 |
NP with polydisperse PC (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 protein ratio) 2 protein ratio) |
P5 | 22.2 |
| NP with monodisperse PC (small) | MS* | 14.5 |
| NP with monodisperse PC (medium) | MM* | 19.0 |
| NP with monodisperse PC (large) | ML* | 24.3 |
2.2 Medium model
In this study we simulated NPs diffusing in different media at different concentrations. We modelled both polydisperse and monodisperse suspensions. For the polydisperse suspensions, we simulated human plasma, a common incubation medium for in vitro experiments. Human plasma is a complex suspension of biomolecules containing over 3000 identified proteins but the 20 most abundant represent approximately 98% of the total protein mass of plasma.57 To provide a comprehensive representation of polydispersity of the plasma proteome, the plasma model simulated was composed of the 29 most abundant proteins in plasma. The hydrodynamic properties for each protein at infinite dilution were calculated from the full-atomistic protein structures using HYDROPRO, with the same parameters as reported in Section 2.1. After, each protein in the medium was represented by a single bead of the size of its rH. Physical properties of the proteins together with their concentrations in plasma are reported in Table S3. Finally, the polydispersity index of the medium was calculated as α = σ/〈rH〉, where σ is the standard deviation of the hydrodynamic sizes in solution and 〈rH〉 is the mean of the size distribution.In the monodisperse suspensions, a NP–PC was allowed to diffuse within a simulation box containing only one crowder type (corresponding to α = 1). The crowder sizes considered in this study range from 2.1 nm to 25 nm, covering a broad spectrum of possible crowding conditions. The selected sizes include: 2.1 nm, representing the smallest protein in the polydisperse medium; 3.5 nm, corresponding to the most abundant protein in the polydisperse medium; 3.7 nm, which represents the average size of all proteins in the polydisperse medium; 3.9 nm, obtained using the effective radius formula 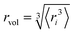 from ref. 34, where the angle brackets denote the average over the entire distribution of proteins in the human plasma model (this value should provide an effective monodisperse equivalent for the polydisperse medium); 8.7 nm, corresponding to the largest protein in the polydisperse medium; 18.9 nm, matching the hydrodynamic size of the tracer in solution (P1 NP); 25 nm, included to extend the range of crowding conditions explored, specifically to scenarios where the crowders are larger than the tracer in solution. Table 2 summarizes all the media tested, detailing their composition, average size, and volume occupancy.
from ref. 34, where the angle brackets denote the average over the entire distribution of proteins in the human plasma model (this value should provide an effective monodisperse equivalent for the polydisperse medium); 8.7 nm, corresponding to the largest protein in the polydisperse medium; 18.9 nm, matching the hydrodynamic size of the tracer in solution (P1 NP); 25 nm, included to extend the range of crowding conditions explored, specifically to scenarios where the crowders are larger than the tracer in solution. Table 2 summarizes all the media tested, detailing their composition, average size, and volume occupancy.
| Composition | 〈rcr〉 [nm] | ϕ tot |
|---|---|---|
| Polydisperse plasma | 3.7 | 0.05, 0.1, 0.2, 0.3, 0.4, 0.5 |
| Mono-crowded | 2.1 | 0.3 |
| Mono-crowded | 3.5 | 0.005, 0.025, 0.05, 0.1, 0.2, 0.3, 0.4, 0.5 |
| Mono-crowded | 3.7 | 0.3 |
| Mono-crowded | 3.9 | 0.3 |
| Mono-crowded | 4.1 | 0.3 |
| Mono-crowded | 4.3 | 0.3 |
| Mono-crowded | 8.7 | 0.05, 0.1, 0.2, 0.3, 0.4, 0.5 |
| Mono-crowded | 18.9 | 0.3 |
| Mono-crowded | 25.0 | 0.3 |
2.3 Brownian dynamics (BD)
To investigate the diffusion of NPs with a PC layer under macromolecular crowding conditions, we performed overdamped BD simulations using the HOOMD-blue software package.55 For the NP–PC RB, an anisotropic integrator was used, ensuring that the correct MOI were defined. More specifically, a central particle located at the centre of mass of the NP–PC RB is defined and then forces, energies and torques are transferred from the constituent particles to the central one, and then are used to integrate the equations of motion of the central particle and update the constituent particles of the RB. BD simulations were performed in a cubic box with periodic boundary conditions. The box size for each system was determined to ensure it was at least three times the size of the tracer, with the NP–PC volume fraction, ϕNP, held constant across simulations within the same medium. For example, in the plasma medium with ϕtot = 0.3, the NP–PC volume fraction was set to ϕNP = 0.015. A total of 1086 proteins were initially modelled for the plasma medium, with their quantities determined based on the experimental molarities provided in Table S3. The number of each protein species was set according to its respective molarity, preserving the experimental concentration ratios in the plasma medium. The volume occupied by this set of proteins was calculated as,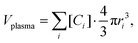 | (3) |
| ϕcrowders = (ϕtot − ϕNP). | (4) |
| Vcrowders = Vbox·ϕcrowders. | (5) |
Formally, the scaling factor is defined as
 | (6) |
| Ni = f·[Ci]. | (7) |
HOOMD-blue employs a self-consistent system of units.55 The system temperature was set to 293 K, with energy units defined such that kBT = 1 where kB is Boltzmann's constant. To establish a realistic time scale, we set the simulation length unit σ = 1 nm and matched the translational diffusion coefficient Dt0 of both the proteins in solution and the tracer under dilute conditions to the values computed using HYDROPRO and HYDRO++, respectively.50,54 In this way, the reduced time unit was defined as τ = 1 µs. For each system, between 5 and 50 independent systems were simulated, followed by production runs lasting up to 1250τ. To avoid particle overlaps in dense systems, equilibration was performed by initially simulating a larger box and gradually compressing it into the target packing fraction. A fixed integration time step of dt = 5 × 10−5τ was used throughout the simulations.
During the BD simulations, one single NP–PC (RB or h-SS representation) is allowed to diffuse through the chosen medium. All the proteins and the tracer in the solution only interact via Weeks–Chandler–Anderson (WCA) potential,58 as per the formula:
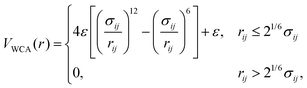 | (8) |
2.4 Calculation of the translational diffusion coefficient
For the NP–PC complexes and proteins in solution we calculated the ensemble averaged mean squared displacement (MSD) of the each particle type over time. For better statistics, we used the so called “window method”, more specifically the implementation available in Freud's library.59 Then, from a linear fit of the MSD vs. time curve, the translational diffusion was obtained from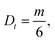 | (9) |
In discussing the results, the diffusion coefficients are reported in normalized form as (Dt/Dt0), where (Dt0) denotes the translational diffusion coefficient of the corresponding particle in the absence of crowders (i.e., at infinite dilution). These reference diffusion coefficients were derived from the crystal structures of proteins in solution using HYDROPRO,50 and from the RB tracer using HYDRO++,54 respectively.
3 Results and discussion
3.1 Effects of the PC
In this section, we focus on the effects of the PC on the long-time diffusion (see Table 1 for details on the systems simulated). In particular, we analysed how various PC properties might impact the diffusivity of the NP–PC complex. Fig. 2a reports the normalised long-time translational diffusion coefficients as a function of the hydrodynamic radius for the crowders in the medium, i.e. plasma proteins for the systems P1t1 and P2t1. These proteins correspond to those reported in Table S3. Fig. 2a also reports the average values of the diffusivities of the proteins in the medium across all 10 simulations (4 variations for each of the P1 and P2 NPs models and their simplified versions), and comparing these values to the ones for P1t1 and P2t1 indicates that there is very small variability between all simulated systems. A similar trend was noted for the remaining eight models. These findings suggest that the diffusion of proteins in the suspension is not significantly affected by the specific type of NP–PC complex, at least under the conditions tested, where only a single NP–PC occupies a small volume fraction of the system. On the other hand, regarding the NPs, the results for the P1 and P2 NPs are depicted in Fig. 2b. For each system, the four layouts (t1, t2, t3, t4) and the simplified PC (P1-S and P2-S) lead to nearly indistinguishable Dt estimates (within the error bars). These results suggest that simulating a single configuration of the PC is sufficient for accurately capturing the dynamics of the NP–PC complex. Also, reducing the number of protein types in the corona does not result in a significant loss of information, as long as the overall morphology is preserved. In this way, we establish that the specific layout of proteins in the corona does not heavily influence their overall mobility. Therefore, to simplify the following analysis we will only present the t1 arrangement, without further discussion of the other cases (t2, t3, t4 and Simplified).Fig. 3 shows the normalized translational long-time diffusion coefficients as a function of rH for 8 RB NP–PCs (see Table 1 for details) and 5 equivalent h-SS (P1, P2, MS, MM, ML). Despite the range of compositions and polydispersities of the PC simulated, all RB systems follow the same monotonic decrease of Dt/Dt0 as a function of rH. Notably, both monodisperse (MS, MM, ML) and polydisperse coronas (P1-P5) align along the same trend line. A similar behaviour is also observed for the h-SS NPs, but the trend is shifted to higher values of Dt/Dt0. This shows that the explicitly modelled RB systems (solid circles) diffuse consistently slower than the equivalent h-SS approximations (open circles). For the RB and the h-SS models with the same rH, the ratio between their Dt decreases linearly with the size of the tracer, as shown in the inset graph of Fig. 3. As our results suggest, the hydrodynamic size, or Stokes radius, alone does not accurately reflect the dynamics of the NP–PC systems, as the type of model employed (RB or h-SS) determines the diffusivity of a NP even if they have the same hydrodynamic size. This discrepancy complicates the effective application of existing theoretical26–29 and computational23,24,34 models, as well as the interpretation of experimental results that derive the hydrodynamic size from the diffusion behaviour.40–43 This issue is particularly relevant for NPs with a PC, as the PC thickness is often derived from the diffusion coefficient.45 Although the h-SS and RB models have the same hydrodynamic size at infinite dilution, they yield different diffusion coefficients under crowded conditions, highlighting that using the computed Dt to infer size can lead to inconsistent results depending on the tracer's morphology (RB or h-SS). It is clear that in order to accurately represent a NP–PC complex by a SS model, the size of the simplified model cannot be based purely on the rH of the NP–PC system, so an alternative descriptor must be derived.
Here, we propose the use of the accessible surface area as a geometric descriptor to model the NP–PC SS size in crowding conditions. Our approach is based on the well-known Solvent Surface Accessible Area (SASA),60 that corresponds to the area accessible to a solvent molecule, usually water. It is normally calculated by implementing the rolling ball algorithm, which consists in simulating a probe rolling over the surface of interest61 to quantify what surface area is accessible to the probe. The size of the probe can be adjusted depending on the level of detail one wants to include in the representation of the surface. In our case, the crowders will “see” the tracer, and depending on their sizes relative to the NP–PC complex, they will perceive more or less details of the surface morphology, affecting the diffusivity of the NP. As in this study the tracer interacts with nanometer-sized crowders, the interest is in deriving a surface accessible to them rather than water molecules as is commonly done. For the specific medium composition (polydisperse plasma) and volume fraction (ϕ = 0.3) used in our simulations, we find that the optimal size of the probe is rprobe ≈ 1 nm. For a more accurate definition and to avoid confusion with the SASA method, we refer to our modified SASA as crowders accessible surface area (CASA), which has been calculated employing the VMD software.62 Using the calculated CASA, we derive an effective size,  for the RB systems, which will then be used as an alternative to the rH. Note that for h-SS models reff = rH + rprobe, corresponding to the surface accessible to the probe, as setting reff = rH would describe the excluded surface area instead of the accessible one. We emphasize that, unlike the tracer (NP–PC), for which both RB and h-SS representations have been tested, proteins in the medium are represented solely as hydrodynamic spheres characterized by the input parameter rH; i.e. different modelling approaches have been applied exclusively to the tracer, not to the crowders. Because no non-spherical representations exist for these proteins, deriving a CASA-based effective radius for them would not be meaningful within this modelling framework, as only the spherical representation is used for all proteins across the investigated systems. Fig. 4 shows the normalized diffusivities plotted against the defined effective radius, reff. With this representation, both the detailed RB models and the simplified h-SS models collapse onto a single trend curve. This indicates that reff serves as a meaningful descriptor capable of unifying the diffusional behaviour of different tracer representations. The P1 RB tracer, for instance, aligns more closely with an h-SS model that has the hydrodynamic radius of the P2 system, rather than that of P1.
for the RB systems, which will then be used as an alternative to the rH. Note that for h-SS models reff = rH + rprobe, corresponding to the surface accessible to the probe, as setting reff = rH would describe the excluded surface area instead of the accessible one. We emphasize that, unlike the tracer (NP–PC), for which both RB and h-SS representations have been tested, proteins in the medium are represented solely as hydrodynamic spheres characterized by the input parameter rH; i.e. different modelling approaches have been applied exclusively to the tracer, not to the crowders. Because no non-spherical representations exist for these proteins, deriving a CASA-based effective radius for them would not be meaningful within this modelling framework, as only the spherical representation is used for all proteins across the investigated systems. Fig. 4 shows the normalized diffusivities plotted against the defined effective radius, reff. With this representation, both the detailed RB models and the simplified h-SS models collapse onto a single trend curve. This indicates that reff serves as a meaningful descriptor capable of unifying the diffusional behaviour of different tracer representations. The P1 RB tracer, for instance, aligns more closely with an h-SS model that has the hydrodynamic radius of the P2 system, rather than that of P1.
The inset of Fig. 4 further supports the relevance of tracer morphology by showing the normalized diffusivity of the RB systems as a function of a roughness parameter ρ, defined here as ρ = CASARB/CASAh-SS. While surface roughness can be quantified using various approaches, such as root-mean-square height fluctuations or curvature-based methods,63 we adopt here a surface-area–based definition. In this approach, roughness is defined as the ratio between the actual surface area of an object and that of an ideal smooth reference, which in our case corresponds to the CASA of the RB and h-SS representations, respectively. Owing to the rolling-ball algorithm, CASA inherently accounts for surface irregularities and protrusions at the length scale relevant to the crowders, thus naturally incorporating surface roughness. The observed decrease in diffusivity with increasing ρ indicates that enhanced surface corrugation increases steric hindrance and effectively enlarges the tracer as perceived by the surrounding crowders.
Using an SS approximation based on hydrodynamic size seemed like an obvious choice for modelling diffusion. However, this approach seems to break down under certain conditions. Previous research for protein suspensions by Balbo et al.36 on self-crowded solutions of bovine serum albumin (BSA) and γ-globulin (IgG) highlighted the significant role of macromolecular shape in influencing translational diffusion. In their study, treating IgG as a h-SS failed to accurately replicate experimental data, while the spherical assumption worked well for the more globular BSA. Our findings suggest a new key mechanism, particularly relevant for solutions where the tracer (NPs with PC in our case) and the crowders are of different nature. Both the RB and h-SS representations in our study fall under the broad definition of “globular” shape, but despite this, their diffusion behaviour differs. We observe that better agreement is achieved when the area of the RB accessible to crowders is taken into account, as systems with similar accessible surface areas (no matter if SS or RB) tend to exhibit comparable diffusion patterns. We now compare our results with those reported by Ando and Skolnick (AS)23 on Escherichia coli cytoplasm. While a direct comparison of data points is not feasible due to different composition of the protein medium and different volume fractions, a similar distinction between the RB and h-SS representations is evident in AS's study. In their work, the long-time diffusion constants of spherical systems are clearly lower than those of molecular-shaped systems, particularly at higher concentrations. However, AS proposed that macromolecular shape has a minimal impact on diffusion in crowded environments, suggesting that an h-SS is a reasonable approximation for in vivo protein diffusion. Specifically, they observed that the difference between molecular-shaped proteins and their spherical approximations was negligible, at least for the experimental diffusion of green fluorescent protein (GFP). Notably, GFP has a rH of approximately 2.4 nm, and among the most abundant macromolecules that compose E. coli is one of the smallest. We can assume that most crowders in the cytoplasm perceive GFP as a small sphere. To motivate this assumption, we must recall the concept of accessible surface area and how it is computed. In the rolling-ball algorithm, the choice of the probe radius influences the measured surface area, as smaller probes are able to capture finer details, resulting in a larger surface area. Conversely, using a very large probe, maybe larger than the size of the main structure, would “smooth out” surface features and approximate the shape as a more uniform rounded form. The extent to which the smoothed-out object approaches a perfect sphere also depends on its original geometry and overall anisotropy. However, we can infer that for a small, globular protein such as GFP, the morphology has little to no impact on its motion in a solution made by much larger crowders. This observation implies that the influence of morphology on diffusion, even for globular objects, becomes significant only when the tracer is larger than the crowders. As the size of the tracer increases relative to the crowders, morphological effects become progressively more important in determining diffusion behaviour. Based on our findings and those of AS, the equivalent hydrodynamic sphere model remains effective for small proteins in solution. This is because when the size of the crowders is equal to or larger than the tracer, the surface area accessible for collisions is reduced compared to smaller crowders. For large macromolecular assemblies like NP–PC, the difference between the h-SS approximation and more detailed representations becomes too significant to ignore, and this discrepancy becomes increasingly relevant as the size of the tracer increases with respect to the size of the crowders. Understanding the influence of morphology on tracer diffusion is essential for a correct interpretation of in vivo conditions, where crowding and heterogeneity are significant.34,36,40 From our data, diffusion appears to be significantly influenced by more sophisticated morphological features such as roughness, not just overall shape and hydrodynamic size, with their impact seemingly dependent on the specific composition of the medium, beyond the simple volume fraction of crowders. This makes its interpretation non-straightforward and challenges the assumptions of the single-sphere model for large bodies in polydisperse solutions. Our analysis highlights the roughness of the surface as an important factor that influences NP–PC mobility under conditions of macromolecular crowding, and we propose the concept of CASA as a relevant geometrical descriptor for a correct interpretation of the diffusion. In this analysis, the medium composition and volume fraction are kept constant across all simulated tracers, which all exhibit a raspberry-like morphology. Consequently, the size of the probe used in the calculation of reff is also held constant. From our analysis and discussion, the appropriate probe size used to compute the effective surface area is expected to vary with medium composition, concentration, and tracer shape. The optimal value of reff therefore should emerge from the interplay of all these factors. Their combined effects on bridging RB and h-SS diffusion behaviours will be explored in more detail in the following section.
3.2 Effects of the medium
In this section, we focus on the role of the medium on NP diffusion. To do that, we simulated only one NP–PC type (P1) in different media at different concentrations. First, it is important to determine whether the polydisperse of the plasma medium (with a polydispersity index of α = 0.26) can be approximated as a monodisperse suspension (α = 1) without affecting the diffusion of the tracer in solution. We derived two effective crowder sizes and we assessed their ability to replicate the behaviour of the polydisperse plasma. To do that, the Dt of the P1 NP in five mono-crowded suspensions was measured, with crowder sizes (rcr) of 3.5, 3.7, 3.9, 4.1, and 4.3 nm, all at a fixed total volume fraction of ϕ = 0.3. Among these, two sizes were derived as effective representations based on specific criteria, while the remaining three were selected to ensure a representative distribution of data points. Specifically, in the medium with rcr = 3.7 nm, the size of the crowders is derived as the mean hydrodynamic radius of proteins in the polydisperse plasma, whereas in the medium with rcr = 3.9 nm, the size of the crowders is derived from colloidal diffusion theory34 (see Section 2.2 for details). Fig. 5 shows the Dt of the P1 RB tracer in polydisperse plasma normalised over the Dt of the same tracer in the five mono-crowded media (rcr of 3.5, 3.7, 3.9, 4.1, and 4.3 nm, as detailed above). | ||
| Fig. 5 Diffusivity of P1 RB in polydisperse plasma medium normalised over the diffusivities of the same tracer in different mono-crowded media as a function of the crowders' size rcr. | ||
Our results show that the diffusivity in rcr = 3.9 nm is the closest match to that of the polydisperse plasma. This suggests that the effective size of crowders, as derived from ref. 34, provides a more accurate representation of the crowding effects seen in polydisperse suspensions. However, this agreement could be coincidental, and other scaling approaches might also yield good agreement under the same conditions investigated here. In contrast, the crowder size rcr = 3.7 nm, which was based on the mean protein size in plasma, resulted in a lower diffusivity. Notably, when the effective size is properly chosen, the sensitivity to polydispersity becomes quite low. However, within the narrow range of approximately 3.7–4.1 nm for the effective crowder radius, the Dpoly/Dmono changes markedly, indicating high sensitivity to rcr in this interval. At the extremes of this range, we observe a plateau-like behaviour, suggesting reduced sensitivity to variations in crowder size. Although this simplification may depend on the specific NP–PC simulated and may not be generalizable without incorporating hydrodynamic interactions (HIs), it constitutes a step toward establishing a scaling theory for polydisperse solutions, which is currently unavailable.
Fig. 6 shows the translational diffusivities of P1 RB normalised over the equivalent h-SS ones (DtRB/DtSS) in three media, two mono-crowded and the polydisperse plasma, at different volume fractions. As the volume fraction of the solution increases, the deviation between the RB representation and the h-SS model becomes more pronounced. We observe a similar trend across all three media. At low volume fractions the two models, as expected, converge but the rate at which this converges happens depends on the crowders' size. For rcr = 8.7 nm (blue circles in the figure), the two models converge at ϕ = 0.05. For rcr = 3.5 nm, the two models converge at even lower packing fractions (0.025 and 0.005). These findings are significant for two key reasons: first, they underscore that the influence of the tracer's morphology on diffusion increases with the packing fraction of the solution; second, they emphasize that the h-SS approximation is only accurate (as compared to the RB model) at very low concentrations, with the upper concentration at which both models predict the same diffusivity depending on the size of the crowders. This last statement is of particular interest, as the hydrodynamic radius is widely adopted in colloid-theory based models for the diffusion of macromolecules even at high packing fractions.23,34,36 These results, together with the ones shown in Fig. 4, would suggest that the size ratio between the tracer and the crowder, together with the tracer's morphology and medium volume fraction, are key factors in the correct interpretation of macromolecular diffusion. When a tracer interacts with a population of much smaller crowders, the surface area available for collisions becomes bigger compared to interactions with larger crowders, as can be depicted from the 2D representation in Fig. 6. This is particularly significant in the case of patchy-like bodies, which may feature concave regions that can only be accessed by smaller crowders in the solution. In this discussion, we assumed that the morphological properties of the analysed systems remain unchanged across different concentrations. It is known that macromolecular crowding induces changes in the folding and compactness of individual proteins,64–67 and large-scale simulations also show that increased protein concentration can induce unfolding,68 providing additional evidence for crowding effects. However, the impact of crowding on more complex structures, such as the PC, remains underexplored. It is unclear whether macromolecular crowding influences morphology, surface roughness, and overall structural integrity of the PC in a way similar to individual proteins. However, a recent study on the biomolecular corona surrounding vesicles hypothesizes that discrete regions of protruding aggregates likely form the corona's architecture, even at high concentrations.8 This finding supports the notion of an anisotropic, patchy layer of biomolecules around the central particle, even under macromolecular crowding conditions, aligning with the model proposed here. Thus, while the patchy, anisotropic nature of the PC has been observed, recent evidence indicates that the validity of the employed description depends on NP chemistry,9 and it may therefore be only valid to particular particle type and under specific conditions.
From the inset in Fig. 3, we already observed that DtRB/DtSS decreases linearly as the size of the tracer rtr increases. Fig. 6 further proves that at the same volume fraction, larger crowders in the medium result in higher values of DtRB/DtSS. These observations would suggest that when to use a h-SS model as an alternative to the RB model will depend on the size ratio between tracer and crowders, and that the h-SS approximation may be valid within certain limits. To explore this hypothesis more systematically, we analyzed the translational diffusivities of the P1 RB and h-SS tracers (both with hydrodynamic radius rtrH = 18.9 nm) in five mono-crowded environments, each with different crowder sizes (rcr = 2.1, 3.5, 8.7, 18.9, 25 nm), while keeping the total volume fraction constant at ϕtot = 0.3. Fig. 7 shows the normalized diffusivity as a function of the crowder-to-tracer size ratio rcr/rtrH. Already within this representation, the diffusivities of the RB and h-SS tracers approach each other as the crowder-to-tracer size ratio increases, indicating that tracer morphology becomes progressively less relevant when crowders are larger than or comparable in size to the size of the tracer. The same data have also been plotted using the CASA-derived effective tracer radius rtreff to define the crowder-to-tracer size ratio rcr/rtreff. These results are reported in Fig. S9. While this mapping slightly reduces the quantitative difference between RB and h-SS diffusivities across size ratios, it does not change the qualitative trend already observed in Fig. 7: morphological effects on tracer diffusion are already strongly suppressed when crowders are of the same hydrodynamic size of the tracer.
Beyond this small methodological difference, the results show that DtRB/DtSS increases with increasing size ratio, meaning that the diffusivities of RB and h-SS tracers converge as crowders become larger compared to the tracer. This indicates that, for the specific raspberry shape analysed in this study, when crowders are larger than the tracer, the tracer morphology has a negligible impact on diffusion. We compare now these findings with the results of Ando and Skolnick23 and Balbo et al.36 In the former study, a h-SS representation was sufficient to capture the diffusion of GFP in a complex, polydisperse cytoplasmic environment, emphasizing the dominant role of HIs, which were treated with a detailed model that included far-field and near-field HIs. In contrast, Balbo et al. found that the same h-SS approximation failed to accurately describe the diffusion of IgG under self-crowding conditions, while it worked well for BSA under identical conditions. They concluded that molecular shape could have a stronger impact on diffusion than other factors. Since Balbo and Ando treat HIs using different approximations, their results are not directly comparable to each other, nor to ours, as we do not include HIs at all in this model. Our focus was on morphological effects, and excluding other interactions from the model allowed us to isolate their influence. In this way, the results presented here suggest that the influence of morphology of the tracer on its diffusion becomes negligible when the tracer is significantly smaller than the surrounding crowders, as in the case of GFP in the cytoplasmic solution. However, when the tracer and crowders are of comparable size, as in the self-crowding conditions for BSA and IgG, shape effects become more pronounced. The fact that the h-SS model succeeded for BSA but failed for IgG under the same size ratio conditions (i.e., rtr/rcr = 1) highlights the role of the tracer intrinsic shape. BSA, being globular, is well approximated by a sphere, while IgG, with its extended Y-shaped structure, deviates substantially from spherical symmetry and shows model-dependent diffusion behaviour at the rtr/rcr = 1. This highlights that the validity of the h-SS model is not governed solely by the overall shape (i.e.: globular, Y-shaped, rod-like), but also by the tracer-to-crowder size ratio. Therefore, the interplay between these two factors must be jointly considered when choosing an appropriate model to describe diffusion in crowded environments.
Fig. 8 shows the radial distribution function (RDF) computed for the same systems as in Fig. 6, using the center-of-mass distance between the NP–PC complex and crowders. The RDFs, resolved at 0.02 with binning adjusted for system size, highlight how crowder size, polydispersity, and tracer geometry shape local structure. All systems exhibit increased structuring with crowding, with the h-SS model showing sharper peaks due to its smooth, spherical geometry, indicating stronger local ordering. In contrast, the RB model, comprising multiple beads, presents broader, less defined peaks, reflecting a more irregular surface and reduced local ordering.
Crowder size further modulates structuring: smaller crowders (Fig. 8a and b) produce broader RDFs, particularly for the RB model, while larger crowders (Fig. 8e and f) yield sharper peaks and reduced sensitivity to tracer geometry. Polydispersity (Fig. 8c and d) smooths RDF profiles, reflecting heterogeneous environments. As crowder size increases relative to the tracer, the RDFs and diffusion coefficients of the two models converge. However, at higher volume fractions (Fig. 8b, d and f), differences between h-SS and RB models grow more pronounced, emphasizing the role of tracer morphology under crowded conditions.
Although translational diffusion analysis showed that the RB model can be approximated by an isotropic representation, provided it accounts for both hydrodynamic size and morphological contributions (dependent on the tracer/crowder size ratio and volume fraction), this simplification sacrifices detail on local structuring. Representing a NP–PC as an equivalent sphere may suffice for translational diffusion analysis, but it limits insight into microscale interactions, a critical factor for future studies aiming to capture both translational dynamics and spatial organization.
4. Conclusions and outlook
In this study, we presented a mesoscale model for NPs covered by a PC and investigated their diffusion in crowded media, specifically the case of the diffusion of NP PC complexes that are larger than the crowding agents. This focus on the diffusion of large tracers in crowded polydisperse environments appears to be under-explored in the current literature, and the findings presented could provide valuable insights, particularly regarding NP diffusion. Here the PC is treated as rigid-body, under the assumption that, once formed, it is steadily adsorbed on the NP within the time scales relevant for the diffusion. Proteins, both in solution and in the PC, have been modelled as soft repulsive spheres of equivalent hydrodynamic radii and interacting via volume exclusion interactions. This approximation allowed us to examine how specific properties of the tracer and of the medium influenced translational diffusion. Different compositions of the PC and different sized crowding agents have been tested, at several volume fractions.In the analysis of the diffusivity in the long time, the combined effects of macromolecular crowding and morphology emerge as critical factors for the derivation of the tracer's effective size and, consequently, of the PC's thickness. We find that as the volume fraction of the crowding agents increases, the diffusion of the molecularly-shaped representations diverges significantly from the equivalent single-sphere ones. This suggests that deriving the overall size/PC thickness from the diffusion coefficient using the Stokes–Einstein relationship at volume fractions >0.05 can lead to incorrect estimations. The fact that globular bodies with the same hydrodynamic radius at infinite dilution exhibit significantly different diffusion behaviours at higher packing fractions challenges the accuracy of size estimations under crowded conditions, calling into question the validity of current approaches for determining true sizes in such environments.
Our results show that the deviation between a rigid-body model and a single-sphere model depends both on the tracer-to-crowder size ratio and the total occupied volume fraction. This scaling reveals that, in polydisperse systems containing macromolecules of widely differing sizes, the hydrodynamic single-sphere approximation fails to reliably describe the diffusion of larger macromolecules, even though it remains adequate for smaller ones. This finding provides a framework for reconciling previous discrepancies between experimental or simulation results and the hydrodynamic single-sphere model, especially for more complex objects in crowded environments. It also underscores that tracer morphology and the relative size ratio between tracer and crowders are key determinants of diffusion.
We therefore propose the accessible surface area as a central parameter in diffusion analysis. The probe size required for accurately calculating this accessible surface area (and for meaningfully applying the single-sphere approximation) appears to be specific to the system under investigation. That different structural models of the same tracer, under identical crowding conditions, yielded significantly different diffusion coefficients directly demonstrates the decisive role of morphology. Our results suggest that this morphological effect arises from two main factors: (1) smaller crowders have access to more surface area of the NP–PC complex; (2) greater volume fractions increase the sensitivity of diffusion to the specific morphology of the tracer. Consequently, the diffusivity of the NP–PC complex is jointly determined by the overall volume occupation of the crowders, their size distribution relative to the tracer, and the tracer's morphology, factors that must be considered when extending single-sphere models to more complex crowding scenarios.
It is important to note that this study does not account for hydrodynamic interactions or the shape of the crowders. Both factors are crucial for fully formulating and deriving a scaling law, as well as for enabling a direct comparison between simulations and experimental data.23,40 Future work should prioritize incorporating these aspects into the proposed model. Furthermore, a detailed assessment of the effects of macromolecular crowding on the features of the PC is needed. It remains unclear whether the raspberry-like rigid model is still valid at high packing fractions and how high concentrations of crowders might affect PC's organization, thickness, softness and morphology. Despite this, our findings provide valuable insights into NP diffusion in polydisperse, concentrated media, improving our understanding of NP mobility in biological environments, that is crucial for predicting the toxicological and pharmacokinetic behaviour of these nanomaterials in therapeutic and nanomedical applications. Furthermore, these results could be extended to other systems, such as large proteins or molecular assemblies, to advance the study of macromolecular diffusion of large, anisotropic bodies in polydisperse solutions.
Conflicts of interest
There are no conflicts to declare.Data availability
Data supporting this article has been included as part of the supplementary information (SI). Supplementary information includes details of the plasma and corona proteins used for modelling the systems, including the atomic coordinates employed to derive their hydrodynamic radii and the translational and rotational diffusion coefficients at infinite dilution for the Brownian Dynamics simulations. It also provides justification for the choice of the Brownian Dynamics approach and the ensemble-averaged center-of-mass mean squared displacement of the systems investigated in this work. See DOI: https://doi.org/10.1039/d5sm01152c.Example scripts to perform Brownian dynamics simulations and to generate the datasets used in this study are available in the Figshare repository (https://doi.org/10.6084/m9.figshare.31123654). The deposit is entitled: Supplementary data and scripts for “Impact of protein corona morphology on nanoparticle diffusion in biological fluids: insights from a mesoscale approach”, a submission to Soft Matter.
Additional input files defining all investigated models and scripts used to generate the simulation trajectories and perform the data analysis are available from the corresponding authors upon reasonable request.
Acknowledgements
The research conducted in this publication was funded by the Irish Research Council under grant number GOIPG/2023/5034 and by the TU Dublin First Time Supervisor Award. The authors wish to acknowledge the Irish Centre for High-End Computing (ICHEC) for the provision of computational facilities and support.References
- K. Riehemann, S. W. Schneider, T. A. Luger, B. Godin, M. Ferrari and H. Fuchs, Angew. Chem., Int. Ed., 2009, 48, 872–897 CrossRef CAS PubMed.
- M. P. Monopoli, F. B. Bombelli and K. A. Dawson, Nat. Nanotechnol., 2011, 6, 11–12 Search PubMed.
- T. Cedervall, I. Lynch, S. Lindman, T. Berggård, E. Thulin, H. Nilsson, K. A. Dawson and S. Linse, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 2050–2055 CrossRef CAS PubMed.
- T. Kopac, Int. J. Biol. Macromol., 2021, 169, 290–301 CrossRef CAS.
- S. Sheibani, K. Basu, A. Farnudi, A. Ashkarran, M. Ichikawa, J. F. Presley, K. H. Bui, M. R. Ejtehadi, H. Vali and M. Mahmoudi, Nat. Commun., 2021, 1–9 Search PubMed.
- A. S. Picco, L. B. Capeletti and M. B. Cardoso, Nano Lett., 2021, 21, 8250–8257 CrossRef.
- C. Cao, L. Zhang, B. Kent, S. Wong, C. J. Garvey and M. H. Stenzel, Angew. Chem., Int. Ed., 2021, 10342–10349 CrossRef CAS.
- A. Musicò, A. Zendrini, S. G. Reyes,, V. Mangolini, L. Paolini, M. Romano, A. Papait, A. R. Silini, P. Di Gianvincenzo, A. Neva, M. Cretich, O. Parolini, C. Almici, S. E. Moya, A. Radeghieri and P. Bergese, Nanoscale Horiz., 2025, 10, 104–112 RSC.
- A. Martinez-Serra, G. Marchetti, F. D’Amico, I. Fenoglio, B. Rossi, M. P. Monopoli and G. Franzese, Int. J. Biol. Macromol., 2025, 330, 148095 CrossRef CAS PubMed.
- M. Qin, J. Zhang, M. Li, D. Yang, D. Liu, S. Song, J. Fu, H. Zhang, W. Dai, X. Wang, Y. Wang, B. He and Q. Zhang, Theranostics, 2020, 10, 1213–1229 CrossRef CAS PubMed.
- S. Tenzer, D. Docter, J. Kuharev, A. Musyanovych, V. Fetz, R. Hecht, F. Schlenk, D. Fischer, K. Kiouptsi, C. Reinhardt, K. Landfester, H. Schild, M. Maskos, S. K. Knauer and R. H. Stauber, Nat. Nanotechnol., 2013, 8, 772–781 Search PubMed.
- F. Etoc, E. Balloul, C. Vicario, D. Normanno, D. Liße, A. Sittner, J. Piehler, M. Dahan and M. Coppey, Nat. Mater., 2018, 17, 740–746 CrossRef CAS PubMed.
- S. Ritz, S. Schöttler, N. Kotman, G. Baier, A. Musyanovych, J. Kuharev, K. Landfester, H. Schild, O. Jahn, S. Tenzer and V. Mailänder, Biomacromolecules, 2015, 16, 1311–1321 Search PubMed.
- Y. Yu, Y. Xie, P. Zeng, D. Zhang, R. Liang and W. Wang, Nanomaterials, 2019, 1–12 Search PubMed.
- Z. Li, Y. Hu, Z. Miao, H. Xu, C. Li, Y. Zhao, Z. Li, M. Chang, Y. Sun, F. Besenbacher, P. Huang and M. Yu, Nano Lett., 2018, 6778–6788 CrossRef CAS.
- L. Tian, X. Li, P. Zhao, X. Chen, Z. Ali, N. Ali, B. Zhang, H. Zhang and Q. Zhang, Macromolecules, 2015, 48, 7592–7603 CrossRef CAS.
- Y. Zhang, H. Zhang, X. Cai, Y. Li, Y. Chen, L. Wang, W. Li, Y. Wang, L. Li and B. Li, ACS Nano, 2022, 16, 1234–1246 Search PubMed.
- Q. Cai, V. Castagnola, L. Boselli, A. Moura, H. Lopez, W. Zhang, J. M. de Araújo and K. A. Dawson, Nanoscale Horiz., 2022, 7, 288–298 RSC.
- F. Zhao, Y. Zhao, Y. Liu, X. Chang, C. Chen and Y. Zhao, Small, 2011, 7, 1322–1337 Search PubMed.
- A. Stradner and P. Schurtenberger, Soft Matter, 2020, 307–323 RSC.
- A. Stradner, H. Sedgwick, F. Cardinaux, W. C. K. Poon, S. U. Egelhaaf and P. Schurtenberger, Lett. Nat., 2004, 432, 492–495 CrossRef CAS PubMed.
- G. Foffi, G. Savin, S. Bucciarelli, N. Dorsaz, G. M. Thurston, A. Stradner and P. Schurtenberger, Proc. Natl. Acad. Sci. U. S. A., 2014, 111, 16748–16753 CrossRef CAS.
- T. Ando and J. Skolnick, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 18457–18462 CrossRef CAS.
- P. M. Blanco, M. Via, J. L. Garcés, S. Madurga and F. Mas, Entropy, 2017, 19, 105 CrossRef.
- F. Roosen-runge, M. Hennig, F. Zhang, R. M. J. Jacobs, M. Sztucki and H. Schober, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 11815–11820 CrossRef CAS PubMed.
- M. Tokuyama and I. Oppenheim, Phys. A, 1995, 216, 85–119 CrossRef CAS.
- G. D. J. Phillies, Macromolecules, 1987, 20, 558–564 CrossRef CAS.
- D. Langevin and F. Rondelez, Polymer, 1978, 19, 875–882 CrossRef CAS.
- L. H. Cai, S. Panyukov and M. Rubinstein, Macromolecules, 2011, 44, 7853–7863 CrossRef CAS PubMed.
- P. R. Lang, J. A. Rivera-mora, Y. Liu, S. Monter and C.-P. Hsu, Soft Matter, 2021, 17, 10301–10311 RSC.
- D. Moreno-Chaparro, N. Moreno, F. B. Usabiaga and M. Ellero, J. Chem. Phys., 2023, 158, 104108 CrossRef CAS.
- Y. M. Wani, G. Kovakas and M. P. Howard, Soft Matter, 2024, 3942–3953 RSC.
- B. Ilhan, F. Mugele and M. H. G. Duits, J. Colloid Interface Sci., 2022, 607, 1709–1716 CrossRef CAS PubMed.
- M. Grimaldo, H. Lopez, C. Beck, F. Roosen-Runge, M. Moulin, J. M. Devos, V. Laux, M. Härtlein, S. Da Vela, R. Schweins, A. Mariani, F. Zhang, J. L. Barrat, M. Oettel, V. T. Forsyth, T. Seydel and F. Schreiber, J. Phys. Chem. Lett., 2019, 10, 1709–1715 CrossRef CAS PubMed.
- F. Hirschmann, H. Lopez, F. Roosen-Runge, T. Seydel, F. Schreiber and M. Oettel, J. Chem. Phys., 2023, 158, 084112 CrossRef CAS PubMed.
- J. Balbo, P. Mereghetti, D. P. Herten and R. C. Wade, Biophys. J., 2013, 104, 1576–1584 CrossRef CAS PubMed.
- A. Gulotta, M. Polimeni, S. Lenton, C. G. Starr, A. Stradner, E. Zaccarelli and P. Schurtenberger, Mol. Pharmaceutics, 2024, 21, 2250–2271 CrossRef CAS.
- S. Bucciarelli, J. S. Myung, B. Farago, S. Das, G. A. Vliegenthart, O. Holderer, R. G. Winkler, P. Schurtenberger, G. Gompper and A. Stradner, Sci. Adv., 2016, 58, 1–8 Search PubMed.
- H. Felix and T. Franosch, Rep. Prog. Phys., 2013, 76, 046602 CrossRef PubMed.
- T. Skóra, F. Vaghefikia, J. Fitter and S. Kondrat, J. Phys. Chem. B, 2020, 124, 7537–7543 CrossRef.
- S. Kwon and S. Lee, J. Korean Phys. Soc., 2020, 77, 700–706 CrossRef.
- T. Kühn, T. O. Ihalainen, J. Hyväluoma, N. Dross, S. F. Willman, J. Langowski, M. Vihinen-Ranta and J. Timonen, PLoS One, 2011, 6, e22962 CrossRef PubMed.
- F. Babayekhorasani, D. E. Dunstan, R. Krishnamoorti and J. C. Conrad, Soft Matter, 2016, 12, 8407–8416 RSC.
- F. Roosen-runge, P. Schurtenberger and A. Stradner, J. Phys.: Condens. Matter, 2021, 33, 154002 CrossRef CAS.
- M. Cui, R. Liu, Z. Deng, G. Ge, Y. Liu and L. Xie, Nano Res., 2014, 7, 345–352 CrossRef CAS.
- A. A. Nikitin, A. Y. Yurenya, R. R. Gabbasov, V. M. Cherepanov, M. A. Polikarpov, M. A. Chuev, A. G. Majouga, V. Y. Panchenko and M. A. Abakumov, J. Phys. Chem. Lett., 2021, 12, 6804–6811 CrossRef CAS PubMed.
- I. Rouse and V. Lobaskin, Biophys. J., 2021, 1–17 Search PubMed.
- F. Tavanti, A. Pedone and M. C. Menziani, Int. J. Mol. Sci., 2019, 20, 3539 CrossRef CAS.
- P. Mosaddeghi Amini, J. Subbotina and V. Lobaskin, Nanomaterials, 2023, 13, 1857 CrossRef.
- A. Ortega, D. Amorós and J. García De La Torre, Biophys. J., 2011, 101, 892–898 CrossRef CAS PubMed.
- RCSB Protein Data Bank, https://www.rcsb.org/.
- AlphaFold, https://alphafold.ebi.ac.uk/.
- S. Milani, F. Baldelli Bombelli, A. S. Pitek, K. A. Dawson and J. Rädler, ACS Nano, 2012, 6, 2532–2541 CrossRef CAS PubMed.
- J. G. De La Torre, G. Del Rio Echenique and A. Ortega, J. Phys. Chem. B, 2007, 111, 955–961 CrossRef PubMed.
- J. A. Anderson, J. Glaser and S. C. Glotzer, Comput. Mater. Sci., 2020, 173, 109363 CrossRef CAS.
- W. Lai, Q. Wang, L. Li, Z. Hu, J. Chen and Q. Fang, Colloids Surf., B, 2017, 152, 317–325 CrossRef CAS PubMed.
- G. L. Hortin, Clin. Chem., 2006, 52, 1223–1237 CrossRef CAS.
- J. D. Weeks, D. Chandler and H. C. Andersen, J. Chem. Phys., 1971, 54, 5237–5247 CrossRef CAS.
- B. Dice, V. Ramasubramani, E. Harper, M. Spellings, J. Anderson and S. Glotzer, Proceedings of the 18th Python in Science Conference, 2019, pp. 27–33 CAS.
- T. J. Richmond, J. Mol. Biol., 1984, 178, 63–89 CrossRef CAS PubMed.
- A. Shrake and J. A. Rupley, J. Mol. Biol., 1973, 79, 361–371 CrossRef PubMed.
- W. Humphrey, A. Dalke and K. Schulten, J. Mol. Graphics, 1996, 14, 33–38 CrossRef CAS.
- S. Trevisani and P. L. Guth, Remote Sens., 2025, 17, 3864 CrossRef.
- C. M. Miller, Y. C. Kim and J. Mittal, Biophys. J., 2016, 111, 28–37 CrossRef PubMed.
- V. Nguemaha, S. Qin and H. X. Zhou, J. Phys. Chem. B, 2018, 122, 11262–11270 CrossRef CAS PubMed.
- L. Haas-Neill, K. Joron, E. Lerner and S. Rauscher, Protein Sci., 2024, 1–57 Search PubMed.
- N. Tokuriki, M. Kinjo, S. Negi, M. Hoshino, Y. Goto, I. Urabe and T. Yomo, Protein Sci., 2004, 125–133 CrossRef CAS PubMed.
- V. Bianco, G. Franzese and I. Coluzza, Chem. Phys. Chem., 2020, 21, 377–384 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |







