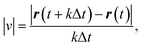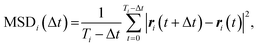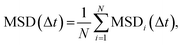Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Nudging cell migration from within through microrod-induced morphological deformation
Masayuki
Hayakawa
 *a,
Tatsuya
Tanaka
b and
Hiroaki
Suzuki
*a,
Tatsuya
Tanaka
b and
Hiroaki
Suzuki
 b
b
aDepartment of Mechanical Engineering, Kyoto Institute of Technology, Goshokaido-cho, Matsugasaki, Sakyo-ku, Kyoto 606-8585, Japan. E-mail: hayakawa@kit.ac.jp
bGraduate School of Science and Engineering, Chuo University, Kasuga 1-13-27, Bunkyo-ku, Tokyo 112-8551, Japan
First published on 10th March 2026
Abstract
Cell migration plays a central role in various biological processes, including development, wound healing, and cancer metastasis, and represents a fundamental form of self-organized motion at the cellular scale. These self-propelled cells can serve as microscale agents with potential applications in bioengineering and microsystem design. To realize such possibilities, it is essential to establish effective methods for controlling their migration. Conventional approaches, such as chemotactic, optogenetic, and substrate-based guidance, depend on external interventions that influence only a limited number of cells. Here, we present a proof-of-concept in Dictyostelium discoideum to bias cell migration by inducing deformation from within the cell. We demonstrate that glass microrods are internalized and that these internalized rods elongate the cells along their own axis. The elongated cells tend to migrate in the direction of their long axis, resulting in enhanced directional persistence. Unlike conventional methods requiring external deterministic cues or patterned environments, our approach enables cells to autonomously and persistently alter their migration behavior through internal morphological deformation. This study introduces a new framework for modulating cell migration and establishes a foundation for developing biohybrid systems that utilize living cells as self-propelled carriers.
Introduction
Cell migration plays an essential role in various biological processes, including development,1–3 wound healing,4–6 and cancer metastasis.3,7,8 It is also a highly efficient and dynamic process driven by the cell's intrinsic energy sources. Cells possessing such migratory capabilities are not merely biological entities but can also be harnessed as mobile components exhibiting advanced locomotion for engineering applications. In particular, their autonomous migratory behavior offers distinct advantages over conventional artificial actuators and has attracted attention for transport in both in vivo and in vitro settings.9–12 Because cell migration is inherently stochastic, a major goal in advancing such technologies is to develop methodologies that enable cells to maintain persistent migration over extended timescales.Currently, strategies for controlling cell migration can be broadly classified into several major types. The first involves chemotactic guidance, which relies on chemotaxis—the ability of cells to migrate in response to chemical concentration gradients.12–15 In this approach, the spatial distribution of specific extracellular chemical cues is used to direct cell movement. Although relatively easy to implement, this method faces limitations because concentration gradients naturally dissipate over time through diffusion, making it difficult to sustain or dynamically adjust stable directional guidance over extended periods.
Another approach involves optogenetic control, which manipulates intracellular signaling pathways using light. A well-established example is the optogenetic activation of Rac1, a small GTPase that promotes actin polymerization and induces lamellipodia formation, thereby determining cell polarity and migration direction.16–18 A photoactivatable form of Rac1 (PA-Rac1) enables spatiotemporal regulation of Rac1 activity through light stimulation. In cells expressing PA-Rac1, localized illumination induces lamellipodia formation at defined sites, allowing precise control over the direction of migration. Other small GTPases, including Cdc42 and RhoA, can also be optogenetically regulated to control distinct aspects of cell migration.19,20 It should be noted that these approaches are not primarily designed to control cell migration but are mainly employed as experimental tools to elucidate the underlying molecular mechanisms. Optogenetic techniques have the distinct advantage of enabling directional control without altering the surrounding microenvironment. However, they require genetic modification and continuous micrometer-scale light stimulation under a microscope, increasing technical complexity and limiting experimental accessibility.
The third approach is substrate-based guidance, in which cell adhesion to engineered substrates promotes lamellipodia formation, polarity establishment, and subsequent migration. Micropatterning serves as a representative example of this technique.21–24 For instance, cells adhering to an asymmetric teardrop-shaped micropattern on a substrate establish a defined polarity and, once released, tend to migrate toward the blunt end of the pattern.23 Furthermore, another study demonstrated that continuously arranging asymmetric triangular patterns on a substrate can effectively rectify cell migration, guiding cells in a unidirectional and long-range manner.24 Although substrate-based guidance enables highly reproducible control of cell migration, it requires cells to be placed on predefined micropatterns, which restricts the range of applicable environments. Beyond chemotaxis, optogenetics, and substrate micropatterning, directed migration can also be guided by light and electric fields (phototaxis25 and electrotaxis26). In addition, substrate mechanics and geometry such as stiffness gradients (durotaxis27) and curvature (curvotaxis28,29) can bias polarity and migration direction. Overall, existing strategies exhibit distinct limitations, including unstable chemical gradients, genetic and technical complexity, and dependence on predefined substrates or applied physical fields.
Among these approaches, substrate-based guidance fundamentally depends on the deformation of cell morphology, thereby regulating lamellipodia formation and the establishment of cell polarity. In other words, if cell morphology could be controlled without relying on externally implemented structures, it may be possible to regulate cell migration more independently of predefined environmental features, similar to genetic approaches. Inspired by this concept, we aimed to guide cell migration from within by internally deforming cellular morphology rather than using external cues. To this end, we examined how the internalization of micrometer-sized structures induces cell deformation and influences migratory behavior. Dictyostelium discoideum was selected as a model system because it is a highly motile amoeboid cell capable of robust phagocytosis and spontaneous migration, making it well suited for investigating motility control. Specifically, we utilized glass microrods and a non-chemotactic mutant of Dictyostelium discoideum known as KI cells.30–32 In this study, we demonstrate that the microrods are internalized through phagocytic activity, forming cell–rod composites that result in an elongated morphology of KI cells (Fig. 1). Furthermore, the velocity vectors of the cells tend to align with the orientation of the rods, accompanied by enhanced directional persistence. Unlike conventional approaches that externally control cell migration, our strategy gently nudges cells to reorganize their migration behavior from within. This approach enhances the directional persistence of cell migration without the need for external structures or genetic modification, providing a simple route toward autonomous modulation.
Methods
Formation of cell–rod composites
In this study, a non-chemotactic mutant strain of Dictyostelium discoideum, referred to as KI cells, was used. KI cells were prepared as follows. One milliliter of Klebsiella aerogenes suspended in 5LP medium (0.5% lactose, 0.5% bactopeptone 211677, OD = 0.1) was spread onto a 9 cm 5LP agar plate (0.5% lactose, 0.5% bactopeptone 211677, 1.5% agar). After the medium had dried, KI cells were inoculated onto the plate and incubated for approximately 5 days at 21 °C. After cultivation, the KI cells and Klebsiella on the plate were collected using phosphate buffer (PB). To remove Klebsiella, the suspension was centrifuged, the supernatant was carefully aspirated, and fresh PB was added. This washing procedure was repeated twice. The number of cells was then counted using a hemocytometer.A cell–rod composite refers to a unified structure in which a KI cell internalizes a glass microrod through phagocytosis. Glass microrods were obtained from a commercial supplier (Nippon Electric Glass Co., Ltd, Japan) with an average length of 15 µm and an average diameter of 3 µm. To form the composites, KI cells (1.0 × 105 cells per mL) and glass microrods (0.6 × 10−5 g mL−1) were co-incubated in PB in a glass-bottom dish at 21 °C overnight. The cell density on the substrate was 1.0 × 104 cells per cm2. During incubation, the microrods were internalized into the cell bodies. The cell–rod composites were observed using an inverted bright-field microscope (Axio Observer, Carl Zeiss AG, Germany).
Classification of cell–rod interactions
Bright-field (BF) images were used to classify cell-rod interactions and to estimate the internalization fraction. First, we defined rod-associated cells as those in which the rod overlapped with the cell region (the area enclosed by the cell outline) in projection; this population was used as the denominator for calculating the internalization fraction. We then classified a rod as internalized when the entire rod was contained within the cell region, based on visual inspection of the BF images. If any portion of the rod extended beyond the cell region, it was categorized as partial attachment and was not counted as internalized.Cell fixation and staining
Coverslips on which the cell–rod composites had been incubated were immersed in 4% paraformaldehyde and kept at room temperature for 15 min. After fixation, the coverslips were gently rinsed by transferring them into PB. Following removal from PB, a permeabilization buffer (0.5% Triton X-100 in PB) was applied dropwise onto the coverslips, which were then incubated at room temperature for 5 min. The coverslips were subsequently rinsed again with PB for approximately 30 s. Thereafter, 100 nM Acti-stain 488 phalloidin (Cytoskeleton, Inc., USA) was applied to the coverslips, which were then incubated for 30 min at room temperature in the dark. After staining, the coverslips were washed twice with PB for 30 s each. Finally, a drop of ProLongTM Glass Antifade Mountant (Thermo Fisher Scientific, Inc., USA) was placed on a glass slide, and the stained coverslip was mounted onto it. The mounted samples were incubated at room temperature in the dark and subsequently observed using a laser-scanning confocal microscope (LSM 700, Carl Zeiss AG, Germany).Cell contour analysis
Time-lapse images of cell–rod composites were acquired at 10 s intervals for 120 s, and each frame was cropped such that the cell centroid was positioned at the center. From the resulting time-lapse images, cell contours were extracted using Cellpose,33 a deep learning-based segmentation tool. In this study, custom models were trained using microscope images obtained with the same equipment as the time-lapse images to enable segmentation of both cell contours and internalized microrods. Contour annotations were performed via the Python API, and both model training and segmentation were carried out using GPU-accelerated computation. The extracted contour data were further analyzed in MATLAB (MathWorks, Inc., MA, USA), primarily using the regionprops function. Specifically, regionprops was applied to the segmented regions (filled contours) obtained by Cellpose to compute the centroid, major/minor axis lengths, and orientation for both the cell and the internalized rod. These parameters define the moment-based equivalent ellipse for each segmented region. Eleven cell–rod composites were analyzed.Circular mapping and correlation analysis
To quantify the spatiotemporal dynamics of cell morphology, a polar coordinate system centered on the cell centroid was introduced. At each time point, the distance from the centroid to the cell boundary was measured for each angular position ϕ, and defined as Amp(ϕ, t). The angular direction ϕ was sampled at 1° intervals (360 divisions), with ϕ = 0° defined as the rightward direction along the horizontal axis and increasing clockwise. The dynamic protrusion and retraction of the membrane were thus represented as temporal variations in Amp(ϕ, t). The autocorrelation function of Amp(ϕ, t) was calculated as follows:where
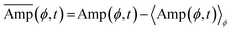 . Angular and temporal averages are denoted by 〈〉ϕ,t. The analyses were conducted using the time-lapse images and cell contour data described in the preceding section (Cell Contour Analysis).
. Angular and temporal averages are denoted by 〈〉ϕ,t. The analyses were conducted using the time-lapse images and cell contour data described in the preceding section (Cell Contour Analysis).
Morphological dynamics were then classified into three patterns: elongation, rotation, and oscillation, based on the CAmp(Δϕ, Δt). If CAmp(Δϕ, Δt) showed no clearly identifiable stripe pattern or instead exhibited a checkerboard-like or mottled appearance, the cell was classified as oscillation. When a stripe pattern was clearly observed, we quantified the angular speed from the stripe slope as |Δθ/Δt|. We then classified these cases as elongation (|Δθ/Δt| ≤ 0.65 deg s−1) or rotation (|Δθ/Δt| > 0.65 deg s−1).
Cell migration analysis
Time-lapse images of cell migration (time interval = 20 s; total duration = 30 min) were analyzed using TrackMate, a plugin for ImageJ (National Institutes of Health, MD, USA), to determine the position of cells r(t) at each time point t. To eliminate the influence of cell–cell contact on migration, only segments in which cells migrated individually were analyzed. Consequently, the tracking durations of the trajectories used for analysis ranged from 480 s to 1800 s. Custom-trained Cellpose models were used as the detector in TrackMate. For the control data (cells without microrods), 32 trajectories of cell centroids were tracked using a model trained to detect cell contours, whereas for the cell–rod composites, 30 trajectories of rod centroids were tracked using a model trained to detect rod contours. In particular, for the microrod, the ellipse parameters (major axis length and orientation) provided by TrackMate's ellipse fit were extracted and used to define the rod major axis. From the obtained trajectories, apparent migration speed was calculated as follows:where Δt is the imaging time interval (20 s) and k ≥ 1 is an integer.
Further, for each tracked cell i, the time-averaged mean square displacement (MSD) was obtained as
where N is the number of tracked cells. The persistence length Lp was also extracted by fitting MSD(Δt) to a persistent random-walk (ballistic-to-diffusive) model:34
where D is the effective diffusion coefficient, v is the characteristic speed, and τ is the persistence time. The persistence length was then given as Lp = vτ. To estimate the uncertainty of Lp, we used bootstrap resampling over cells. For each bootstrap replicate b = 1,…,B, N cells were sampled with replacement, the averaged MSD was calculated, and the model fit was performed to obtain L(b)p. We report the median of L(b)p as the point estimate and the 2.5th–97.5th percentiles as the 95% confidence interval. The relative uncertainty was defined as the standard deviation divided by the mean of the L(b)p distribution.
Results and discussion
Following overnight incubation, three types of interactions between KI cells and glass microrods were observed: no interaction (control; Fig. 2a-i and Movie S1), partial attachment (Fig. 2a-ii and Movie S2), and internalization (Fig. 2a-iii and Movie S3). In the case of partial attachment (Fig. 2a-ii), one end of the microrod adhered to the cell surface without being completely engulfed. In this configuration, although the cell and the rod moved together, the distal end of the rod extended beyond the cell contour. In contrast, in composites where the rod was internalized (Fig. 2a-iii), the microrod remained entirely within the cell contour while moving together with the cell. Because no external delivery methods, such as lipofection, were employed, the internalization of microrods can be attributed to spontaneous phagocytic activity, resulting in the formation of cell–rod composites. Although such large structures might appear unlikely to be internalized, this phenomenon is consistent with previous findings showing that particles comparable in size to cells—such as elliptical disks35 and asterisk-shaped microdevices36—can be engulfed via phagocytosis. As shown in Fig. 2b, among cells that interacted with microrods, the incidence of internalization was approximately one-fourth that of partial attachment. Although partial attachment seems to represent an intermediate state preceding internalization, not all attached rods necessarily progress to that stage. This is likely because, when the rod orientation or contact area is suboptimal, phagocytosis is not triggered and the rods remain partially attached.35 The long-term stability of rod internalization remains to be fully elucidated; however, in time-lapse imaging over 3 h in a PDMS chamber (Fig. S1), the microrod remained fully internalized with no evidence of rod ejection over the tested timescale. This result suggests that, despite their size, internalized microrods remain relatively stable once engulfed. We also performed a trypan blue exclusion assay on cell–rod composites, indicating no obvious plasma-membrane permeabilization (Fig. S2).Fig. 2c–e show maximum-intensity projection images of representative fixed cell–rod composites. F-Actin and nuclei were visualized using phalloidin (green) and Hoechst (blue), respectively. An actin-rich region was observed at one end of the cell, where multiple filopodia-like protrusions extended outward. In contrast, the rear side lacked such actin enrichment. This polarized distribution of actin indicates the establishment of distinct front–rear polarity. Fig. 2f presents cross-sectional views of a representative cell–rod composite, showing ZY sections at three different X positions (top panel), an XY section (middle panel), and an XZ section (bottom panel). In the ZY sections, the actin cytoskeleton appeared as ring-like cross-sectional structures with diameters comparable to that of the microrod (3 µm). Moreover, both the XY and XZ sections displayed rectangular dark regions with dimensions similar to those of the rod, providing strong evidence for microrod internalization. In the ZY section at x = x3, deformation of the nucleus caused by the presence of the microrod was observed. This deformation became more apparent when compared with nuclei of cells without microrods (Fig. S3).
We next focused on the morphology of cells that internalize glass microrods. Fig. 3a and b show time-lapse bright-field and segmented images of a representative cell–rod composite, respectively. To characterize the geometric features of the cell–rod composite, several parameters were defined, as illustrated in Fig. 3c. The centroids of the cell and the rod were denoted as Gcell and Grod, respectively. The cell contour was approximated by an ellipse, with its major and minor axes defined as acell and bcell, respectively. The rod was also approximated by an ellipse, and its major axis was defined as arod. We first calculated the elongation index, R = acell/bcell, for both cell–rod composites and cells without microrod interaction. The time-averaged value of R was calculated for each cell, and the mean across cells (one value per cell) was higher for cell–rod composites than for cells without microrods (Fig. 3d). Specifically, R was 1.49 ± 0.04 for control cells (mean ± SE, N = 22) and 1.65 ± 0.05 for cell–rod composites (mean ± SE, N = 11), indicating that microrod internalization induced cell elongation. We then examined the relationship between R and the length of the internalized rod, arod. The plot in Fig. 3e shows that increases in R were associated with increases in arod. This is likely because longer microrods deform the cell more extensively along their axis, resulting in greater elongation. Since rod-induced polarization in our system may be mediated by such morphological constraints, we speculate that excessively reducing rod length may weaken this effect, although it may facilitate internalization. Furthermore, the absolute value of the angle between acell and arod, denoted as θ (Fig. 3c), had a mean of 15.1°. Fig. 3f shows a histogram of θ for all cell–rod composites (one time-averaged value per cell), indicating that the elongation axis of the cells was well aligned with the microrod axis. In addition, the distance between Gcell and Grod, defined as d, was approximately 2 µm —comparable to the rod radius (1.5 µm)—suggesting that Gcell and Grod were nearly coincident (Fig. 3g).
To investigate how microrod internalization influences the spatiotemporal dynamics of cell morphology, we conducted a contour-based analysis following a method established in previous studies37,38 Specifically, we defined a radial amplitude function, Amp(ϕ, t), representing the distance from the cell centroid to the membrane edge as a function of the angular coordinate ϕ and time t. Previous work demonstrated that the autocorrelation of Amp(ϕ, t) quantitatively evaluates the presence of ordered morphological patterns such as elongation, rotation, and oscillation. These patterns are classified according to characteristic spatiotemporal correlations in cell edge dynamics: elongation corresponds to a sustained extension along a fixed axis, rotation involves the lateral propagation of protrusions around the cell periphery, and oscillation is characterized by periodic deformation, typically with protrusions alternating across the cell axis.
Fig. 4a shows the mapping of Amp(ϕ, t) for a cell without microrod internalization, and Fig. 4b presents its cell contours at t = 20 s and t = 100 s. Points on the contour corresponding to the top 20% of Amp(ϕ, t) values at each time point are marked in green, illustrating that the protrusions change direction over time (Fig. 4b). Although the morphological dynamics are not immediately apparent in Amp(ϕ, t), the corresponding autocorrelation function, CAmp(Δϕ, Δt) exhibits a checkerboard-like or mottled pattern (Fig. 4c), indicating periodic protrusive activity occurring at distinct angular positions (oscillation). This suggests that the cell extends protrusions in one direction and subsequently re-extends them at shifted angles, resulting in alternating cycles of protrusion and retraction that modulate its morphology. Additional examples of CAmp(Δϕ, Δt) from other cells are provided in Fig. S4. The same analysis was next applied to cell–rod composites (Fig. 4d–f). The mapping of Amp(ϕ, t) and the corresponding cell contours at t = 20 s and t = 100 s are shown in Fig. 4d and e, respectively. Over time, peaks of Amp(ϕ, t) appeared around ϕ = 90° and 270°, indicating that the cell maintained a relatively elongated shape (Fig. 4e). Moreover, morphological fluctuations were relatively small around ϕ = 90° (dashed line in Fig. 4d), which corresponds to the rear side of the cell–rod composite (Movie S4). These observations support the notion that pseudopodial activity is spatially biased toward the front, while the rear remains comparatively static,37 thereby suggesting that this front–rear asymmetry is preserved even in the presence of an internalized microrod. The autocorrelation function (Fig. 4f) revealed a distinct elongation pattern, indicating that the internalized microrod stabilized the cell shape along a fixed axis over an extended period. A similar trend was observed in additional cell–rod composites, as presented in Fig. S5. As shown in Fig. 4g, when the cell–rod composites were classified according to their morphological dynamics, the average length of the internalized microrods was clearly greater in the elongation group than in the rotation and oscillation (Rot. & Osc.) group. This finding suggests that cells internalizing longer microrods tend to exhibit elongation patterns.
Thus far, we have demonstrated the effect of microrod internalization on cell morphology. It is well established that cell morphology and motility are closely interconnected.39 To clarify how microrod internalization influences migratory behavior, we conducted tracking analysis of cell migration (Movies S5 and S6). Trajectories of migrating cells without and with microrods are shown in Fig. 5a and Fig. S6, where the tracking duration ranged from 480 s to 1800 s. All trajectories start at (x, y) = (0, 0). For control cells (without microrods), trajectories were obtained by tracking cell centroids, whereas for cell–rod composites, trajectories were determined by tracking microrod centroids. As shown in Fig. 3g, the distance between the cell and microrod centroids was less than 2 µm, indicating that the microrod trajectories could be considered equivalent to the cell trajectories. Cells without microrods exhibited directional changes and tended to follow less linear paths (Fig. 5a, left), whereas cell–rod composites displayed relatively straight trajectories with minimal directional changes (Fig. 5a, right). Next, we compared the MSD between cells without microrods and cell–rod composites (Fig. 5b). The composites exhibited a steeper slope, with the difference from the control becoming increasingly evident as the lag time increased. To quantify the difference in migration persistence, we fitted the MSD to a persistent random-walk model and extracted the persistence length Lp. This analysis yielded Lp = 15.6 µm (95% CI: [8.93, 33.6], rel.unc. = 0.4) for control cells and Lp = 102 µm (95% CI: [50.3, 177], rel.unc. = 0.4) for cell–rod composites, indicating a pronounced increase in persistence upon rod internalization (Table S1). The characteristic speed parameter v obtained from the same fits was comparable within uncertainty between the two conditions (control: v = 0.19 µm s−1; cell–rod composites: v = 0.18 µm s−1; Table S1). The apparent migration speed |v| showed a k dependence, especially for control cells (Fig. S7), indicating that this finite-interval estimate can be influenced by directional persistence when time interval is not sufficiently small. Therefore, we use the fitted characteristic speed v as the estimate of the intrinsic migration speed, rather than the finite-interval apparent speed. We then analyzed the angle α between the velocity vector v and the axis of the internalized microrod (Fig. 5c). A histogram of the time-averaged absolute values of α for each trajectory is shown in Fig. 5d, revealing a peak around 25°. This deviation reflects temporal fluctuations in the migration direction, indicating that the cell moves approximately along the microrod axis but with angular fluctuations of about ±25°. The characteristic scale of ∼25° could reflect intracellular constraints, potentially including steric interactions between the microrod and the cell nucleus (Fig. 2f), although this contribution was not directly quantified in the present study. Such fluctuations are also likely promoted by variations in protrusion formation at the leading edge (Fig. 4d). Overall, cells tend not to move perpendicular to the microrod axis but rather migrate along it. Furthermore, the histogram of the time-averaged α for each trajectory, shown in the inset of Fig. 5d, is centered near 0°, suggesting that there is no significant bias in left–right orientation relative to the microrod axis. In other words, the migration direction and the microrod axis are aligned in cell–rod composites. Although this analysis does not establish the direction of causality, it demonstrates tight coupling between the two.
Conclusion
In this study, we investigated the interactions between KI cells and glass microrods, encompassing both partial attachment and internalization. Notably, internalization occurred without any artificial introduction methods, indicating that microrods were taken up through spontaneous phagocytic activity. Fixed-cell imaging confirmed the presence of microrods within the cells, demonstrating the successful formation of cell–rod composites. Moreover, internalized microrods induced marked morphological changes, promoting elongation of the cells along the microrod axis. Quantitative analysis using the elongation index R revealed that cell–rod composites exhibited higher R values than cells without microrods, particularly when longer microrods were internalized. Contour-based analysis of cell edge dynamics showed that microrod internalization altered cellular deformation patterns, enhancing the tendency toward elongation. Furthermore, cell–rod composites exhibited more persistent migration, and their movement directions tended to align with the orientation of the internalized microrod. The persistent random-walk analysis indicates that the dominant effect is an increase in persistence time and persistence length, while the characteristic speed is comparable between conditions. These findings show that directional persistence of cell migration can be enhanced from within, not through deterministic, externally imposed cues, but by subtly nudging the cell's spontaneous motion via internal morphological deformation.Whether other eukaryotic cells exhibiting similarly active phagocytosis and motility can form cell–rod composites and display comparable migratory behavior remains to be tested. For instance, macrophages, a type of immune cell, could be considered as candidates. Extending the present approach beyond Dictyostelium will be an important direction for future work.9–12
In this study, we consistently employed glass microrods, which are symmetric along both the longitudinal and radial axes. An important next step is to investigate how more complex, asymmetric microstructures influence cellular deformation and migration upon internalization. In particular, curved (“banana-shaped”) microparticles (e.g., SU-8-based structures)40,41 provide a practical symmetry-breaking geometry and may induce a systematic turning bias (non-zero mean angular velocity) during migration. Another promising avenue of research involves modifying the rod material to impart additional functionalities. For instance, using magnetic materials would allow microrod orientation to be controlled by an external magnetic field, providing a well-defined handle to systematically probe rod–migration coupling. In parallel, live imaging of PIP3 and PTEN42,43 under controlled microrod orientation would enable a direct test of whether rod-induced deformation biases and stabilizes the front–rear polarity axis. This could be quantified by measuring the alignment of the front–rear polarity axis (read out from PIP3/PTEN localization) and its temporal fluctuations relative to the rod axis and migration direction. The cell–rod composite may therefore serve as a powerful experimental platform for elucidating the coupling between cell morphology and migration. Alternatively, thermoresponsive hydrogel44 microrods could provide deformable shape cues. In a bilayer design comprising a responsive and a non-responsive layer, differential swelling under temperature changes would induce bending, enabling tunable curvature that could in turn modulate migration patterns.
Collectively, such approaches may provide a useful starting point for developing micrometer-scale motile carriers and related biohybrid systems. We hope that this study will contribute to advancing fundamental strategies for controlling cell migration without reliance on external cues or genetic modification.
Author contributions
M. H.: conceptualization, data curation, formal analysis, investigation, and writing. T. T.: investigation. H. S.: conceptualization, resources, and writing.Conflicts of interest
The authors declare no conflicts of interest.Data availability
The data supporting this study are available in the supplementary information (SI), which includes captions for the supplementary movies, supporting figures and tables, and supplementary analyses. See DOI: https://doi.org/10.1039/d5sm01149c.Acknowledgements
The authors thank Prof. H. Miyazako (The University of Tokyo) for valuable discussions. This work was supported by JSPS KAKENHI (Grant Number 19K12770) and JGC-S (Nikki-Saneyoshi) Scholarship Foundation.References
- E. Scarpa and R. Mayor, J. Cell Biol., 2016, 212, 143–155 CrossRef CAS.
- E. Theveneau, L. Marchant, S. Kuriyama, M. Gull, B. Moepps, M. Parsons and R. Mayor, Dev. Cell, 2010, 19, 39–53 Search PubMed.
- P. Friedl and D. Gilmour, Nat. Rev. Mol. Cell Biol., 2009, 10, 445–457 CrossRef CAS.
- M. Poujade, E. Grasland-Mongrain, A. Hertzog, J. Jouanneau, P. Chavrier, B. Ladoux, A. Buguin and P. Silberzan, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 15988–15993 CrossRef CAS PubMed.
- A. Grada, M. Otero-Vinas, F. Prieto-Castrillo, Z. Obagi and V. Falanga, J. Invest. Dermatol., 2017, 137, e11–e16 CrossRef CAS.
- Y. Xiao, R. Riahi, P. Torab, D. D. Zhang and P. K. Wong, ACS Nano, 2019, 13, 1204–1212 CAS.
- C. D. Paul, P. Mistriotis and K. Konstantopoulos, Nat. Rev. Cancer, 2017, 17, 131–140 CrossRef CAS PubMed.
- P. Mehta, Z. Rahman, P. ten Dijke and P. E. Boukany, Trends Cancer, 2022, 8, 683–697 Search PubMed.
- W. Zhang, M. Wang, W. Tang, R. Wen, S. Zhou, C. Lee, H. Wang, W. Jiang, I. M. Delahunty, Z. Zhen, H. Chen, M. Chapman, Z. Wu, E. W. Howerth, H. Cai, Z. Li and J. Xie, Adv. Mater., 2018, 30, e1805557 CrossRef.
- C. Hermosillo-Abundis, J. A. Flood-Garibay, V. Ayala-Nunez and M. A. Mendez-Rojas, ACS Mater. Lett., 2025, 7, 156–171 CrossRef CAS.
- Y. Qi, X. Yan, T. Xia and S. Liu, Mater. Des., 2021, 198, 109388 CrossRef CAS.
- O. Nagel, M. Frey, M. Gerhardt and C. Beta, Adv. Sci., 2019, 6, 1801242 CrossRef PubMed.
- K. F. Swaney, C.-H. Huang and P. N. Devreotes, Annu. Rev. Biophys., 2010, 39, 265–289 CrossRef CAS.
- P. J. M. V. Haastert and P. N. Devreotes, Nat. Rev. Mol. Cell Biol., 2004, 5, 626–634 CrossRef PubMed.
- M. Skoge, E. Wong, B. Hamza, A. Bae, J. Martel, R. Kataria, I. Keizer-Gunnink, A. Kortholt, P. J. M. V. Haastert, G. Charras, C. Janetopoulos and D. Irimia, PLoS One, 2016, 11, e0154491 CrossRef.
- Y. I. Wu, D. Frey, O. I. Lungu, A. Jaehrig, I. Schlichting, B. Kuhlman and K. M. Hahn, Nature, 2009, 461, 104–108 CrossRef CAS PubMed.
- M. Weitzman and K. M. Hahn, Curr. Opin. Cell Biol., 2014, 30, 112–120 CrossRef CAS PubMed.
- T. Kato, K. Kawai, Y. Egami, Y. Kakehi and N. Araki, PLoS One, 2014, 9, e97749 CrossRef PubMed.
- P. W. Oakes, E. Wagner, C. A. Brand, D. Probst, M. Linke, U. S. Schwarz, M. Glotzer and M. L. Gardel, Nat. Commun., 2017, 8, 15817 CrossRef CAS PubMed.
- P. R. O’Neill, V. Kalyanaraman and N. Gautam, Mol. Biol. Cell, 2016, 27, 1442–1450 CrossRef PubMed.
- K. K. Parker, A. L. Brock, C. Brangwynne, R. J. Mannix, N. Wang, E. Ostuni, N. A. Geisse, J. C. Adams, G. M. Whitesides and D. E. Ingber, FASEB J., 2002, 16, 1195–1204 CrossRef CAS.
- B. Chen, G. Kumar, C. C. Co and C.-C. Ho, Sci. Rep., 2013, 3, 2827 CrossRef.
- X. Jiang, D. A. Bruzewicz, A. P. Wong, M. Piel and G. M. Whitesides, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 975–978 CrossRef CAS PubMed.
- G. Mahmud, C. J. Campbell, K. J. M. Bishop, Y. A. Komarova, O. Chaga, S. Soh, S. Huda, K. Kandere-Grzybowska and B. A. Grzybowski, Nat. Phys., 2009, 5, 606–612 Search PubMed.
- J.-L. Xiao, D.-H. Lu and C.-H. Lee, Appl. Phys. Lett., 2013, 102, 123703 CrossRef.
- B. Cortese, I. E. Palamà, S. D’Amone and G. Gigli, Integr. Biol., 2014, 6, 817–830 CrossRef CAS.
- R. Sunyer and X. Trepat, Curr. Biol., 2020, 30, R383–R387 CrossRef CAS PubMed.
- R. K. Sadhu, M. Luciano, W. Xi, C. Martinez-Torres, M. Schröder, C. Blum, M. Tarantola, S. Villa, S. Penič, A. Iglič, C. Beta, O. Steinbock, E. Bodenschatz, B. Ladoux, S. Gabriele and N. S. Gov, Proc. Natl. Acad. Sci. U. S. A., 2024, 121, e2306818121 CrossRef CAS.
- L. Pieuchot, J. Marteau, A. Guignandon, T. D. Santos, I. Brigaud, P.-F. Chauvy, T. Cloatre, A. Ponche, T. Petithory, P. Rougerie, M. Vassaux, J.-L. Milan, N. T. Wakhloo, A. Spangenberg, M. Bigerelle and K. Anselme, Nat. Commun., 2018, 9, 3995 CrossRef PubMed.
- H. Kuwayama, S. Ishida and P. J. V. Haastert, J. Cell Biol., 1993, 123, 1453–1462 CrossRef CAS.
- H. Kuwayama and S. Ishida, Sci. Rep., 2013, 3, 2272 CrossRef PubMed.
- M. Hayakawa, T. Hiraiwa, Y. Wada, H. Kuwayama and T. Shibata, eLife, 2020, 9, e53609 CrossRef PubMed.
- M. Pachitariu and C. Stringer, Nat. Methods, 2022, 19, 1634–1641 CrossRef CAS.
- C. Bechinger, R. D. Leonardo, H. Löwen, C. Reichhardt, G. Volpe and G. Volpe, Rev. Mod. Phys., 2016, 88, 045006 CrossRef.
- J. A. Champion and S. Mitragotri, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 4930–4934 CrossRef CAS.
- M. I. Arjona, M. Duch, A. Hernández-Pinto, P. Vázquez, J. P. Agusil, R. Gómez-Martínez, M. Redondo-Horcajo, E. Amirthalingam, L. Pérez-García, T. Suárez and J. A. Plaza, Adv. Mater., 2022, 34, e2109581 CrossRef.
- Y. T. Maeda, J. Inose, M. Y. Matsuo, S. Iwaya and M. Sano, PLoS One, 2008, 3, e3734 CrossRef.
- H.-G. Döbereiner, B. J. Dubin-Thaler, J. M. Hofman, H. S. Xenias, T. N. Sims, G. Giannone, M. L. Dustin, C. H. Wiggins and M. P. Sheetz, Phys. Rev. Lett., 2006, 97, 038102 CrossRef PubMed.
- D. L. Bodor, W. Pönisch, R. G. Endres and E. K. Paluch, Dev. Cell, 2020, 52, 550–562 CrossRef CAS.
- C. Fernández-Rico, M. Chiappini, T. Yanagishima, H. de Sousa, D. G. A. L. Aarts, M. Dijkstra and R. P. A. Dullens, Science, 2020, 369, 950–955 CrossRef.
- C. Fernández-Rico, J. S. Urbach and R. P. A. Dullens, Langmuir, 2021, 37, 2900–2906 CrossRef PubMed.
- Y. Arai, T. Shibata, S. Matsuoka, M. J. Sato, T. Yanagida and M. Ueda, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 12399–12404 CrossRef CAS.
- S. Matsuoka and M. Ueda, Nat. Commun., 2018, 9, 4481 CrossRef PubMed.
- A. Das, A. Babu, S. Chakraborty, J. F. R. V. Guyse, R. Hoogenboom and S. Maji, Adv. Funct. Mater., 2024, 34, 2402432 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |