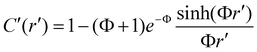Open Access Article
Open Access ArticleMolecular concentration field design using closed-form steady-state solutions
Dong Woo
Kim
 a,
Alison
Grinthal
a and
Rebecca
Schulman
a,
Alison
Grinthal
a and
Rebecca
Schulman
 *abc
*abc
aDepartment of Chemical and Biomolecular Engineering, Johns Hopkins University, 3400 N. Charles Street, Baltimore, MD 21218, USA. E-mail: rschulm3@jhu.edu
bDepartment of Computer Science, Johns Hopkins University, 3400 N. Charles Street, Baltimore, MD 21218, USA
cDepartment of Chemistry, Johns Hopkins University, 3400 N. Charles Street, Baltimore, MD 21218, USA
First published on 11th February 2026
Abstract
Control over spatial concentration fields represents a fundamental challenge in designing synthetic biological systems and programmable soft materials. While nature creates morphogen gradients that orchestrate complex developmental processes, synthetic approaches have largely relied on empirical optimization and computationally intensive simulations. Here, we present an analytical framework for steady-state concentration fields generated by finite-sized localized sources in diffusion–degradation systems and derive closed-form solutions for one-, two-, and three-dimensional geometries. By expressing these solutions in dimensionless form, we show that gradient steepness and spatial structure are organized by the Thiele modulus, which captures the competition between diffusion and degradation length scales. The analysis reveals distinct design regimes: in degradation-dominated systems, gradient shape is governed by exponential decay and becomes dimension-independent, whereas in diffusion-dominated systems, gradient magnitude and extent follow dimension-dependent power-law scaling. Building on these results, we introduce a quantitative design strategy that uses threshold-based criteria to program concentration ranges by tuning physically accessible parameters, most directly the production rate, while holding transport and degradation properties fixed. Comparisons with numerical solutions and reported experimental systems demonstrate consistency with the predicted scaling behavior. Together, this work provides a generalizable and physically transparent framework for designing steady-state concentration fields in synthetic biological and soft matter systems, enabling predictive control of gradient-mediated organization without reliance on extensive numerical optimization.
Introduction
Chemical concentration fields in living cells and tissues guide biological processes across scales, from individual cellular behavior to complex multicellular patterning. Cells respond to diverse molecular gradients—for instance, bacteria follow nutrient concentrations,1,2 immune cells migrate along chemokine gradients,3 neuronal axons grow in response to spatial distributions of guidance molecules,4 and morphogen distributions direct embryonic pattern formation.5 Similarly, synthetic systems have been developed in which concentration variations tune colloidal phase transitions,6,7 control filament growth dynamics,8,9 orchestrate droplet motility,10,11 direct condensate formation,12,13 regulate biochemical oscillations,14,15 and establish spatially controlled patterning in synthetic biological circuits.16–24 For realizing the full potential of these systems, a theoretical framework capable of predicting and controlling concentration fields is required for the rational design of chemical gradient landscapes tailored to applications ranging from tissue engineering to dissipative materials.25These chemical gradients arise from the interplay of localized reactions and molecular transport and are commonly described by reaction–diffusion dynamics, in which local production, degradation, and diffusion of molecules collectively shape spatial concentration profiles.26,27 To capture and predict these behaviors, theoretical frameworks based on reaction-diffusion equations have been developed using a range of mathematical and computational approaches, each with distinct advantages and limitations. Partial differential equation (PDE) formulations with numerical solutions provide detailed spatiotemporal dynamics for complex multi-component systems, but the high computational cost often makes it challenging to explore large parameter spaces or apply them across many system variants.28–34 To balance accuracy and computational efficiency, semi-analytical models that combine closed-form solutions with numerical evaluation steps have been developed.35–37 These approaches are particularly useful for exploring parameter spaces and optimizing system designs in specific reaction-diffusion scenarios with the numerical evaluation of integrals or special functions, which can still impose computational overhead and limit their general applicability for different dimensionality, or complex boundary conditions.
Alternatively, analytical approaches have been developed through local accumulation time, which provide morphogen gradient dynamics before reaching steady state,38 while their later work extended their model to multi-dimensional systems.39 Other reaction–diffusion models describing diffusion and first-order morphogen degradation have been studied, in which steady-state solutions under idealized boundary conditions yield exponential concentration profiles characterized by a decay length 1/λ = (D/k)0.5, where k is the degradation rate constant and D is the diffusion constant.40–43 In these frameworks, the ratio λ/L—where L is the system size—emerges as a key dimensionless parameter governing pattern scaling and robustness, and closed-form solutions are available primarily for one-dimensional continuous domains with prescribed Dirichlet or Neumann boundary conditions. However, existing closed-form solutions do not directly address systems with finite-sized localized sources, nor do they provide a unified description across dimensionality for the predictive design of steady-state concentration gradients in systems such as dissipative materials that require continuous energy input to maintain far-from-equilibrium organization.
Here, we address this challenge by developing an analytically tractable framework for predicting steady-state concentration fields generated by localized-source reaction–diffusion systems. We derive closed-form steady-state solutions for one-, two-, and three-dimensional geometries in which continuous molecular production at finite-sized sources is balanced by diffusion and first-order degradation. By expressing these solutions in dimensionless form, we show that gradient steepness and spatial structure are organized by the Thiele modulus, which captures the competition between diffusion and degradation length scales and delineates diffusion- and degradation-dominated regimes. Building on these results, we introduce a transferable, threshold-based design strategy that leverages analytically derived scaling relationships to connect application-level requirements—such as target concentration and target active range—to experimentally tunable parameters, particularly the production rate, across dimensionalities. Together, this framework provides a quantitative and transferable foundation for designing biochemical gradients in synthetic materials, tissue engineering platforms, and autonomous soft matter systems that respond to long-term chemical cues.
Results and discussion
Analytical framework
To establish a tractable and physically grounded model of the concentration field, we consider a system composed of three essential components: a localized production source, diffusing molecules, and degradation enzymes or other catalysts (Fig. 1a). The source with diameter 2R is modeled as a highly porous region that is symmetric about its center in one-, two-, or three-dimensional space (Fig. 1b–d). Within the confined region of the source, molecules are homogeneously produced at a constant rate rp, while simultaneously diffusing and undergoing degradation via a first-order reaction rc with rate constant k via catalyzed degradation. The interplay of production, diffusion, and degradation generates a concentration gradient, described by the concentration field C(r,t), where r is the spatial coordinate and symmetric from the center of the source and t is time. Concentration fields can be calculated by solving piecewise partial differential equations in the separate regions. For the source region (0 < r < R), the system has production, degradation, and diffusion: | (1) |
Beyond the source (r > R), the system has only diffusion and degradation without the production reaction:
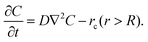 | (2) |
At steady state, where the concentration field no longer varies with time (∂C/∂t = 0), four boundary conditions—symmetry, two continuity conditions, and a dilution condition (see SI for details)—allow us to derive exact solutions across 1D, 2D, and 3D systems (Table S1). The full derivation is provided in the SI.
To generalize the analytical solutions, we used a dimensionless framework defined by three quantities: C′ = C/(rp/k), r′ = r/R, Φ = R(k/D)0.5. Here, concentration C is scaled by the ratio of production to degradation rates, the spatial coordinate r is normalized by the source radius R, and the Thiele modulus Φ quantifies the relative strength of degradation versus diffusion.44 Specifically, Φ characterizes the relative contributions of diffusion and degradation to concentration decay: when Φ > 1, degradation dominates, whereas when Φ < 1, diffusion is dominant. The resulting solutions with and without dimensionless quantities are summarized in Table 1 and Table S1. In 1D systems, the steady-state concentration profiles are well studied and are governed by hyperbolic functions within the source region and exponential decay outside the source: cosh(Φr′) within the source region r' < 1 and exp(−Φr′) in the outer region for r′ > 1.45 However, few solutions in 2D and 3D have been reported. In 2D systems, the solutions involve modified Bessel functions of the first and second kind (I1 and K0), capturing the logarithmic behavior. For 3D systems, the solutions are described by sinh(Φr′)/r′ within the source and exp(−Φr′)/r′ outside the source, reflecting the radial decay of concentration in spherical symmetry. The analytical solutions closely match independent numerical simulations across 1D, 2D, and 3D system configurations and a range of parameters (Φ = 0.3, 1, 3), with total deviations below 10−6 (Fig. 2a–c). While this dimensionless formulation organizes gradient behavior into diffusion- and degradation-dominated regimes, practical gradient design requires additional tuning of physically accessible parameters such as the production rate, as discussed in Design strategy section.
 | ||
| Fig. 2 Validation of analytical concentration field solutions in 1D, 2D, and 3D systems through comparison with numerical simulations, and their semi-logarithmic plots. Dimensionless concentration profiles C′ =C/(rp/k) as a function of dimensionless radial distance r/R demonstrate the precision of our closed-form analytical solutions (solid lines) against independent numerical calculations31 (circles) for three distinct system dimensions in Fig. 1b–d: (a) 1D, (b) 2D, and (c) 3D systems, and their semi-logarithmic plots (d)–(f). The systematic variation of the dimensionless parameter Φ = R(k/D)0.5 from 0.3 to 3 reveals how the characteristic length scale controls gradient steepness with larger Φ values producing sharper concentration drops near the source boundary. The dashed lines in d–f represent analytical extrapolations that extend from the far from the source to near the source. The shaded gray region indicates the source. For numerical calculation, we solved eqn (1) and (2) by formulating them as boundary-value problems and using MATLAB's bvp4c, a fourth-order collocation solver with adaptive mesh refinement. | ||
Spatial decay behavior near and far from the source
We investigate the scaling principles that govern how concentration profiles decay as a function of r/R. To analyze this behavior, we used semi-logarithmic plots for 1D, 2D, and 3D systems across different parameter ranges (Φ = 0.3, 1, 3) (Fig. 2d–f). In the region far from the source (r/R ≫ 1), the profiles for 1D, 2D, and 3D systems each become linear in the semi-log plots. This behavior is consistent with the analytical solutions, which show that the dimensionless concentration derivative (dC'/dr) asymptotically converges to an exponential form proportional to ∼exp(−Φ) (Table 1). This behavior reflects an exponential decay, arising from diffusion and first-order degradation. Importantly, the linear relationship can be used for controlling the gradient steepness by manipulating Φ (= R(k/D)0.5). Consistent with this interpretation, increasing either the source radius or the degradation rate produces steeper gradient profiles.For the 1D system, the linear trend appears both near and far from the source boundaries, consistent with classical diffusion theory46 (Fig. 2d). In contrast, the 2D and 3D systems exhibit nonlinear transitions near the source boundaries (r/R∼1), where the profiles deviate from linearity (Fig. 2e and f). As dimensionality increases, the available volume or area for diffusion grows, allowing molecules to disperse more rapidly away from the source—e.g., higher diffusion flux in 3D system compared to 1D system where molecules are restricted to a line. These deviations are more pronounced at low Φ, where diffusion dominates over degradation, producing a more gradual initial decay of concentration and corresponding nonlinear behavior near the source. Mathematically, this is reflected in the dimensionless concentration derivatives near the source: in 2D, dC′/dr ∼ exp(−Φ)/r′, while in 3D, dC′/dr ∼ exp(−Φ)/r', up to dimension-dependent prefactors. This trend illustrates how diffusion becomes increasingly sensitive to spatial dimensionality, leading to faster concentration decay near the source in higher-dimensional systems.
Shape of concentration gradients: steepness and magnitude
To characterize the steepness and magnitude of concentration gradients, the simple linear relationship valid in the far-field cannot be applied near the source, where 2D and 3D exhibit nonlinear transitions. To capture these features, we quantified gradient magnitude using the concentration at the source surface and gradient steepness using the half-width at half-maximum (HWHM). The surface concentration at r/R = 1 was chosen, rather than the center concentration, because the gradient outside the source functions as the relevant morphogen signal, and the maximum external concentration occurs at the surface.
and gradient steepness using the half-width at half-maximum (HWHM). The surface concentration at r/R = 1 was chosen, rather than the center concentration, because the gradient outside the source functions as the relevant morphogen signal, and the maximum external concentration occurs at the surface.
The surface concentration was calculated as a function of Φ (Fig. 3a). In the diffusion -dominated regime (Φ < 1),  (summarized in Table S2) exhibits distinct power-law scaling behaviors with Φ, where the slope of the log–log plot corresponds to the scaling exponent that characterizes how the gradient peak varies with Φ. Specifically, 1D systems show linear scaling (slope = 1), 2D systems display intermediate behavior (slope ≈ 1.5), and 3D system exhibits quadratic scaling (slope = 2). These different slopes indicate how the radius of the source and degradation rate constant as Φ = R(k/D)0.5 becomes effective in higher dimensions. By contrast, in the degradation-dominated regime (Φ > 1),
(summarized in Table S2) exhibits distinct power-law scaling behaviors with Φ, where the slope of the log–log plot corresponds to the scaling exponent that characterizes how the gradient peak varies with Φ. Specifically, 1D systems show linear scaling (slope = 1), 2D systems display intermediate behavior (slope ≈ 1.5), and 3D system exhibits quadratic scaling (slope = 2). These different slopes indicate how the radius of the source and degradation rate constant as Φ = R(k/D)0.5 becomes effective in higher dimensions. By contrast, in the degradation-dominated regime (Φ > 1),  remains constant with respect to Φ.
remains constant with respect to Φ.
 | ||
Fig. 3 Dimensional scaling laws for concentration field optimization and design. (a) Surface concentration  as a function of dimensionless parameter Φ = R(k/D)0.5 demonstrates distinct scaling behaviors across dimensional configurations, with 1D system exhibiting linear scaling (slope = 1), 2D system showing intermediate behavior (slope = 1.5), and 3D system displaying quadratic scaling (slope = 2), reflecting how source-to-volume ratios fundamentally determine achievable surface concentrations. (b) Half-width at half-maximum (HWHM) normalized by source radius R reveals the spatial extent characteristics of concentration fields, where 1D system exhibits the strongest spatial confinement effects (slope = −1), 2D system shows moderate spatial expansion (slope = −0.5), and 3D system maintains constant width (slope = 0). HWHM/R values from experimental data were compared. Zadorin et al.43 (△) used morphogen gradients in 1D microfluidic devices to direct particle organization. The others (Kim et al.47 (○), Gines et al.48 (◊), Karzbrun et al.14 (□)) built concentration fields in 2D microfluidic devices. Detailed values are summarized in Table S4. as a function of dimensionless parameter Φ = R(k/D)0.5 demonstrates distinct scaling behaviors across dimensional configurations, with 1D system exhibiting linear scaling (slope = 1), 2D system showing intermediate behavior (slope = 1.5), and 3D system displaying quadratic scaling (slope = 2), reflecting how source-to-volume ratios fundamentally determine achievable surface concentrations. (b) Half-width at half-maximum (HWHM) normalized by source radius R reveals the spatial extent characteristics of concentration fields, where 1D system exhibits the strongest spatial confinement effects (slope = −1), 2D system shows moderate spatial expansion (slope = −0.5), and 3D system maintains constant width (slope = 0). HWHM/R values from experimental data were compared. Zadorin et al.43 (△) used morphogen gradients in 1D microfluidic devices to direct particle organization. The others (Kim et al.47 (○), Gines et al.48 (◊), Karzbrun et al.14 (□)) built concentration fields in 2D microfluidic devices. Detailed values are summarized in Table S4. | ||
The HWHM, normalized by the source radius R (summarized in Table S3) was defined as the distance from the center of the source at which the concentration reaches half of the surface concentration  and was calculated as a function of Φ (Fig. 3b). In the diffusion-dominated regime (Φ < 1), the HWHM/R exhibits power-law scaling with Φ, where the slope indicates how strongly gradient steepness depends on Φ. The 1D system exhibits strongest dependence (slope = −1), indicating that gradients become sharper as Φ increases. The 2D system displays moderate dependence (slope = −0.5), whereas the three-dimensional system maintains a nearly constant HWHM/R ≈ 2. This result means that the concentration falls to half of the surface value at approximately twice the source radius, regardless of Φ (Fig. 3c). Thus, the dependence on Φ weakens with increasing dimensionality. In the degradation-dominated regime, HWHM/R is independent of Φ across 1D, 2D, and 3D systems.
and was calculated as a function of Φ (Fig. 3b). In the diffusion-dominated regime (Φ < 1), the HWHM/R exhibits power-law scaling with Φ, where the slope indicates how strongly gradient steepness depends on Φ. The 1D system exhibits strongest dependence (slope = −1), indicating that gradients become sharper as Φ increases. The 2D system displays moderate dependence (slope = −0.5), whereas the three-dimensional system maintains a nearly constant HWHM/R ≈ 2. This result means that the concentration falls to half of the surface value at approximately twice the source radius, regardless of Φ (Fig. 3c). Thus, the dependence on Φ weakens with increasing dimensionality. In the degradation-dominated regime, HWHM/R is independent of Φ across 1D, 2D, and 3D systems.
The physical origin of the slope in the low Φ regime likely arises from the enhanced diffusive flux in higher dimensions, as discussed in the previous section, which enables molecules to disperse into larger available volume, leading to stronger dilution near the source and corresponding changes in scaling exponents. For quasi-one-dimensional or quasi-two-dimensional systems, where confinement in width or height is finite but not negligible, the scaling exponents are expected to interpolate between those of the lower- and higher-dimensional systems. For example, a quasi-2D system with finite height but sources confined to the bottom boundary (e.g., sedimented biomolecular condensates13) allows diffusion away from the substrate into the bulk, and is therefore better approximated by a hemispherical or effectively three-dimensional geometry, leading to slope values intermediate between the 2D and 3D limits.
For comparison with experimental observations, we compiled experimental values of HWHM/R, from previously reported morphogen and reaction–diffusion systems in 1D and 2D microfluidic configurations. These data points are overlaid with our analytical scaling predictions in Fig. 3b, and the corresponding physical parameters (diffusion coefficients, source sizes, degradation rates, and calculated Thiele moduli) are summarized in Table S4. As shown, the experimental measurements are consistent with the dimension-dependent scaling trends predicted by the analytical solutions, supporting the applicability of the theoretical framework to experimentally realizable reaction–diffusion systems.
Design strategy for concentration fields
Based on these insights, we propose a practical strategy for designing steady-state concentration fields using threshold-based criteria (Fig. 4a). In both synthetic reaction–diffusion systems and biological morphogen frameworks, spatial organization is often governed by whether local concentrations exceed a critical threshold, beyond which specific structures, patterns, or functional responses are activated. This threshold-based interpretation is widely employed in synthetic systems to direct material assembly and particle organization, as well as in biological systems to regulate tissue patterning and morphogenesis.5,26,42,43,49 Motivated by this common principle, we introduce a design guideline in which concentration fields are engineered to achieve a target threshold concentration C* over a prescribed spatial extent by tuning physical parameters.The first design step is to determine whether the target concentration field is steep or extended by assessing whether the half-width at half-maximum (HWHM) is smaller or larger than twice the source radius (2R). When HWHM < 2R—corresponding to the degradation-dominated regime (Φ > 1)— the concentration profile decays steeply, with gradient shape governed by simple exponential decay (C′ ∼ e−Φr′), and the surface concentration approaches a constant value (C′surface ≈ 0.5) independent of system dimensionality. In contrast, when HWHM > 2R —the diffusion-dominated regime (Φ < 1)— the concentration field extends over longer distances, and both gradient steepness and magnitude follow dimension-dependent power-law scaling with Φn, where the exponent reflects the effective dimensionality of molecular transport. Once the appropriate regime is determined, the physical parameters R, k, and D can be selected to achieve the desired gradient profile; for instance, operating in the diffusion-dominated regime is advantageous for generating long-range concentration fields in localized RNA transcription systems using DNA-immobilized objects, where diffusion coefficients, source sizes, and degradation rates are often constrained to narrow experimental ranges (Fig. 4b).13,47
In the second step, the threshold concentration (C*) and target active range (L) are specified based on upstream biochemical or material constraints (Fig. 4c). For example, a threshold concentration of C* = 1 nM and a target active range of L = 100 µm are representative of systems such as protein-aptamer–based biosensors, where dissociation constants typically span sub-nanomolar to micromolar concentrations,50 or linker-mediated particle assembly processes that operate in the nanomolar to micromolar range.6,43 By tuning the production rate while holding degradation and transport parameters fixed, we determine production rates of 28, 150, and 850 nM min−1 that yield a concentration of 1 nM at a distance of 100 µm for the 1D, 2D, and 3D cases, respectively.
Finally, to extend the target active range to longer distances, the concentration field can in principle be tuned by adjusting the production rate, degradation rate constant, or source radius. Among these parameters, we focus on the production rate as the primary control variable because it modulates the overall magnitude of the concentration field without altering gradient steepness, whereas changes in source size or degradation rate affect both magnitude and spatial decay (Tables S2 and S3). In practical implementations, the production rate can be directly controlled through experimentally accessible parameters; for example, in localized transcription reactions within hydrogels or biomolecular condensates, it can be tuned by varying the concentration of immobilized DNA templates or the activity of the transcription machinery.13,47
By rearranging the analytical solutions in Table S1, we find that the production rate required to maintain a fixed threshold concentration at distance L scales as exp(λL), 1/K0(λL), L·exp(λL), for 1, 2, 3D systems, respectively (Fig. 4d). These relationships provide direct guidelines for selecting production rates that extend the target active range, for example to 2L or 3L, without modifying degradation kinetics or source geometry. We note that while analogous design rules based on varying the degradation rate or source size are less analytically tractable, they can be obtained numerically. Using the production rates determined from Fig. 4d, the resulting concentration profiles in one-, two-, and three-dimensional systems exhibit threshold crossing at 1L, 2L or 3L, respectively, as shown in Fig. 4e.
Conclusions
In this work, we developed an analytically tractable framework for predicting and designing steady-state concentration fields generated by localized reaction–diffusion sources in one-, two-, and three-dimensional systems. By deriving closed-form solutions for finite-sized sources and expressing them in dimensionless form, we identified how gradient steepness and spatial structure are organized by the Thiele modulus, which delineates diffusion- and degradation-dominated regimes. These results clarify how dimensionality fundamentally shapes concentration decay and spatial extent, and they establish scaling relationships that are consistent with numerical simulations and reported experimental systems.14,43,47,48Beyond analytical description, this framework enables a practical, threshold-based strategy for concentration field design. Rather than relying on empirical tuning or extensive numerical optimization, target concentration ranges can be programmed by mapping application-specific requirements—such as threshold concentration and spatial reach—onto physically accessible parameters. In particular, we show that tuning the production rate provides a direct and experimentally viable means to extend or contract target active ranges while preserving gradient steepness and degradation kinetics, offering a robust design handle across dimensions.
Together, these results provide a quantitative and generalizable foundation for engineering reaction–diffusion concentration fields in synthetic and dissipative materials,25,51,52 where precise spatial control under nonequilibrium conditions is essential. The framework is broadly applicable to systems ranging from programmable biosensors31,53 and gradient-guided assembly to fuel-driven soft matter architectures that sustain autonomous spatial organization.54–56 Future extensions incorporating time-dependent dynamics,25,52 nonlinear reaction kinetics,57 or spatial heterogeneity51 could further expand the applicability of this approach, positioning analytical gradient design as a key tool for the emerging field of programmable soft matter and synthetic morphogen systems, with direct relevance to tissue engineering and regenerative medicine.58–62
Author contributions
Conceptualization, R. S. and D. W. K.; funding acquisition, R. S.; investigation, D. W. K.; project administration, R. S.; software, D. W. K.; resources, R. S.; supervision, R. S.; validation, D. W. K.; visualization, D. W. K.; writing – original draft, R. S., D. W. K. and A. G.; writing – review & editing, R. S., D. W. K. and A. G.Conflicts of interest
There are no conflicts to declare.Data availability
Data and code associated with the publication are available at both https://github.com/DongWoo94/ConcentrationFieldModels and https://doi.org/10.7281/T1MLLJ33, Johns Hopkins Research Data Repository. Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5sm01121c.Acknowledgements
We thank Marc Kamionkowski for insightful discussions. R. S. and A. G. acknowledge support from a Vannevar Bush Faculty Fellowship, ONR N00014-23-1-2868. D. W. K. and therefore the analytical and simulation work, were supported by the U.S. Department of Energy through DE-SC0010426.References
- J. H. Wheeler, K. R. Foster and W. M. Durham, Nat. Microbiol., 2024, 9, 2308–2322 CrossRef CAS PubMed.
- V. Sourjik and N. S. Wingreen, Curr. Opin. Cell Biol., 2012, 24, 262–268 CrossRef CAS PubMed.
- J. W. Griffith, C. L. Sokol and A. D. Luster, Annu. Rev. Immunol., 2014, 32, 659–702 CrossRef CAS PubMed.
- A. L. Kolodkin and M. Tessier-Lavigne, Cold Spring Harbor Perspect. Biol., 2011, 3, a001727 Search PubMed.
- K. W. Rogers and A. F. Schier, Annu. Rev. Cell Dev. Biol., 2011, 27, 377–407 CrossRef CAS PubMed.
- J. Lowensohn, B. Oyarzún, G. Narváez Paliza, B. M. Mognetti and W. B. Rogers, Phys. Rev. X, 2019, 9, 041054 CAS.
- W. B. Rogers and V. N. Manoharan, Science, 2015, 347, 639–642 Search PubMed.
- R. Schulman, B. Yurke and E. Winfree, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 6405–6410 CrossRef CAS PubMed.
- D. Krishnamurthy and M. Prakash, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2304981120 CrossRef CAS PubMed.
- Y. Zhang, A. McMullen, L.-L. Pontani, X. He, R. Sha, N. C. Seeman, J. Brujic and P. M. Chaikin, Nat. Commun., 2017, 8, 21 CrossRef PubMed.
- N. J. Cira, A. Benusiglio and M. Prakash, Nature, 2015, 519, 446–450 Search PubMed.
- M. Morita, T. Sakamoto, S. I. M. Nomura, S. Murata, M. Yanagisawa and M. Takinoue, Adv. Mater. Interfaces, 2024, 11, 2300898 Search PubMed.
- E. Kengmana, E. Ornelas-Gatdula, K.-L. Chen and R. Schulman, J. Am. Chem. Soc., 2024, 146, 32942–32952 Search PubMed.
- E. Karzbrun, A. M. Tayar, V. Noireaux and R. H. Bar-Ziv, Science, 2014, 345, 829–832 Search PubMed.
- M. Weitz, J. Kim, K. Kapsner, E. Winfree, E. Franco and F. C. Simmel, Nat. Chem., 2014, 6, 295–302 CrossRef CAS PubMed.
- Y. Fang, V. V. Yashin, S. J. Dickerson and A. C. Balazs, J. Appl. Phys., 2018, 123, 215107 CrossRef.
- A. Manfrin, Y. Tabata, E. R. Paquet, A. R. Vuaridel, F. R. Rivest, F. Naef and M. P. Lutolf, Nat. Methods, 2019, 16, 640–648 CrossRef CAS PubMed.
- S. Kondo and T. Miura, Science, 2010, 329, 1616–1620 Search PubMed.
- B. A. Grzybowski and W. T. Huck, Nat. Nanotechnol., 2016, 11, 585–592 CrossRef CAS PubMed.
- S. Biswas, V. V. Yashin and A. C. Balazs, Soft Matter, 2018, 14, 3361–3371 Search PubMed.
- M. Moradi, O. E. Shklyaev and A. C. Balazs, Proc. Natl. Acad. Sci. U. S. A., 2024, 121, e2319777121 Search PubMed.
- K. Abe, I. Kawamata, M. N. Shin-ichiro and S. Murata, Mol. Syst. Des. Eng., 2019, 4, 639–643 RSC.
- I. Kawamata, T. Hosoya, F. Takabatake, K. Sugawara, S.-I. Nomura, T. Isokawa, F. Peper, M. Hagiya and S. Murata, 2016.
- I. Kawamata, S. Yoshizawa, F. Takabatake, K. Sugawara and S. Murata, 2016.
- E. Del Grosso, E. Franco, L. J. Prins and F. Ricci, Nat. Chem., 2022, 14, 600–613 CrossRef CAS PubMed.
- T. D. Banerjee and A. Monteiro, Commun. Biol., 2025, 9, 87 Search PubMed.
- H. Connahs, S. Tlili, J. van Creij, T. Y. Loo, T. D. Banerjee, T. E. Saunders and A. Monteiro, Development, 2019, 146, dev169367 CrossRef CAS PubMed.
- A. N. Ford Versypt, P. D. Arendt, D. W. Pack and R. D. Braatz, PLoS One, 2015, 10, e0135506 Search PubMed.
- M. S. Johnson, J. N. Mueller, C. Daniels, H. N. Najm and J. Zádor, J. Phys. Chem. A, 2024, 128, 3685–3702 CrossRef CAS PubMed.
- D. Scalise and R. Schulman, Technology, 2014, 2, 55–66 CrossRef.
- P. J. Dorsey, D. Scalise and R. Schulman, R. Soc. Open Sci., 2022, 9, 220200 Search PubMed.
- D. Scalise and R. Schulman, Nat. Comput., 2016, 15, 197–214 Search PubMed.
- K. Abe, S. Murata and I. Kawamata, Soft Matter, 2021, 17, 6160–6167 Search PubMed.
- S. Chen and G. Seelig, Soft Matter, 2020, 16, 3555–3563 RSC.
- K. Al Noufaey, Results Phys., 2018, 9, 609–614 CrossRef.
- H. Y. Alfifi, ANZIAM J., 2017, 59, 167–182 Search PubMed.
- H. Alfifi, T. R. Marchant and M. Nelson, J. Math. Chem., 2016, 54, 1632–1657 Search PubMed.
- P. V. Gordon, C. Sample, A. M. Berezhkovskii, C. B. Muratov and S. Y. Shvartsman, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 6157–6162 Search PubMed.
- P. V. Gordon, C. B. Muratov and S. Y. Shvartsman, J. Chem. Phys., 2013, 138, 104121 CrossRef PubMed.
- B. Vasiev, J. Theor. Biol., 2016, 404, 109–119 CrossRef PubMed.
- H.-W. Kang, L. Zheng and H. G. Othmer, Interface Focus, 2012, 2, 465–486 Search PubMed.
- P. C. Bressloff, Phys. Biol., 2022, 19, 066005 Search PubMed.
- A. S. Zadorin, Y. Rondelez, G. Gines, V. Dilhas, G. Urtel, A. Zambrano, J.-C. Galas and A. Estevez-Torres, Nat. Chem., 2017, 9, 990–996 CrossRef CAS PubMed.
- E. W. Thiele, Ind. Eng. Chem., 1939, 31, 916–920 CrossRef CAS.
- B. Cushman-Roisin, Thayer School of Engineering Dartmouth College, University Lecture, 2012 Search PubMed.
- P. J. Dorsey, M. Rubanov, W. Wang and R. Schulman, ACS Macro Lett., 2019, 8, 1133–1140 CrossRef CAS PubMed.
- D. W. Kim, M. Rubanov, A. Grinthal, P. Moerman and R. Schulman, Matter, 2025, 8(7), 102208 CrossRef CAS.
- G. Gines, A. Zadorin, J.-C. Galas, T. Fujii, A. Estevez-Torres and Y. Rondelez, Nat. Nanotechnol., 2017, 12, 351–359 CrossRef CAS PubMed.
- H. Inomata, Dev., Growth Differ., 2017, 59, 41–51 CrossRef CAS PubMed.
- H. Lee, T. Xie, B. Kang, X. Yu, S. W. Schaffter and R. Schulman, Nat. Commun., 2024, 15, 7973 Search PubMed.
- B. Dúzs and I. Szalai, Commun. Chem., 2020, 3, 168 CrossRef PubMed.
- J. Boekhoven, W. E. Hendriksen, G. J. Koper, R. Eelkema and J. H. van Esch, Science, 2015, 349, 1075–1079 Search PubMed.
- S. Jia, S. C. Phua, Y. Nihongaki, Y. Li, M. Pacella, Y. Li, A. M. Mohammed, S. Sun, T. Inoue and R. Schulman, Nat. Commun., 2021, 12, 5729 Search PubMed.
- M. Lovrak, W. E. Hendriksen, M. T. Kreutzer, V. van Steijn, R. Eelkema and J. H. van Esch, Soft Matter, 2019, 15, 4276–4283 Search PubMed.
- J. Zenk, D. Scalise, K. Wang, P. Dorsey, J. Fern, A. Cruz and R. Schulman, RSC Adv., 2017, 7, 18032–18040 RSC.
- P. J. Dorsey, D. Scalise and R. Schulman, Angew. Chem., 2021, 133, 342–348 CrossRef.
- K. L. Narayanan, V. Meena, L. Rajendran, J. Gao and S. P. Subbiah, Appl. Comput. Math., 2017, 6, 143–160 CrossRef.
- Z. Zeravcic, V. N. Manoharan and M. P. Brenner, Rev. Mod. Phys., 2017, 89, 031001 CrossRef.
- Z. Zeravcic and M. P. Brenner, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 4342–4347 Search PubMed.
- E. W. Gehrels, W. B. Rogers, Z. Zeravcic and V. N. Manoharan, ACS Nano, 2022, 16, 9195–9202 CrossRef CAS PubMed.
- N. Tompkins, N. Li, C. Girabawe, M. Heymann, G. B. Ermentrout, I. R. Epstein and S. Fraden, Proc. Natl. Acad. Sci. U. S. A., 2014, 111, 4397–4402 CrossRef CAS PubMed.
- I. Hunter, M. M. Norton, B. Chen, C. Simonetti, M. E. Moustaka, J. Touboul and S. Fraden, Phys. Rev. E, 2022, 105, 024310 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |