 Open Access Article
Open Access ArticleStrengthening biofilms with selective metal ions
Kiera J.
Croland
 a and
R. Kōnane
Bay
a and
R. Kōnane
Bay
 *ab
*ab
aDepartment of Chemical & Biological Engineering, University of Colorado Boulder, Boulder, USA. E-mail: konane.bay@colorado.edu
bMaterials Science and Engineering Program, University of Colorado Boulder, Boulder, USA
First published on 26th March 2026
Abstract
Biofilms are structured microbial communities consisting of bacteria embedded in a self-produced extracellular polymeric substance (EPS) that enables survival in diverse environments. The EPS can integrate materials from the surrounding environment, such as metal ions, which can provide additional mechanical protection to the embedded bacteria from environmental stressors. While previous studies demonstrated that metal ions impact the erosion behavior of biofilms, key quantitative properties, such as failure strain, remain largely undocumented due to difficulties in handling these viscoelastic and soft biomaterials. In this work, we introduce a technique to characterize the impact of metal ions on the uniaxial stress–strain response of bulk bacterial biofilms. Through applying this method to Bacillus subtilis pellicles, we demonstrate that exposure to selective metal ions increases both the low strain elastic modulus and maximum stress, while decreasing failure strain. These effects are consistent with ion-mediated EPS crosslinking and are reversible through the introduction of a strong chelating agent, while variations in pH alone have a negligible impact on measured mechanical properties. We compare our results to previous biofilm erosion studies and provide insights into how metal ion interactions can alter the mechanical behavior of biofilms, which will aid in future biofilm mitigation strategies for biofouling or healthcare applications.
1. Introduction
Biofilm-forming bacteria increase their resilience to the surrounding environment by aggregating together and encasing themselves in an extracellular polymeric substance (EPS). Biofilms are complex structures that readily colonize diverse surfaces, including human tissue,1,2 medical devices,3,4 and water distribution systems.5,6 The protective EPS provides constituent bacteria with structural integrity, surface adhesion, and protection from antimicrobial agents.7 As a result, biofilms pose significant challenges across many industries.5 In clinical settings, biofilm infections resist conventional antibiotic therapy and often require surgical intervention for complete removal.2 Biofilms that form on equipment surfaces in industrial settings, such as tanks and pipes, similarly require costly interventions through physical removal or equipment replacement.5,6 Despite these challenges, biofilms can be engineered to offer significant benefits. In wastewater treatment facilities, biofilms serve as bioreactors that efficiently degrade organic pollutants and remove contaminants from effluent streams.8 Additionally, biofilm properties can be exploited in engineered living materials (ELMs)9,10 to create stimuli-responsive intelligent systems. Across all applications, mechanical properties fundamentally govern biofilm behavior, yet characterization of these fragile biological materials remains technically challenging.11,12Current approaches to biofilm mechanical characterization often yield results that are difficult to compare across systems due to a lack of standardization.11,12 Many mechanical testing methods for biofilms disrupt the native biofilm architecture during sample preparation and mounting (e.g., shear rheometry),1,13–19 do not yield quantitative mechanical parameters (e.g., no decipherable modulus,20 relative detachment values,13–15 and stability21), and do not capture responses under physiologically relevant conditions.22–24 While microscale techniques (e.g., microrheology25 and AFM) preserve the local structure, they do not reflect bulk mechanical behavior, which is essential for understanding the macroscopic behavior of these inherently heterogenous structures. Furthermore, laboratory-grown biofilms often lack the complexity found in natural environments. In situ biofilms routinely incorporate external molecules or components from their surroundings into the EPS, altering material properties. Environmental factors, such as nutrient availability21 and host conditions,1 have been shown to influence the EPS composition and production. Therefore, it is critical to understand how environmental stimuli, such as the exposure to metal ions,13,14 biopolymers,15 or antibiotics,15 impact biofilm mechanics.
Metal ion interactions with the EPS are of particular interest due to their prevalence on colonized surfaces. Metal ions are abundant in biological settings, such as within the human body (e.g., blood and tissues) and on abiotic surfaces (e.g., metal pipes and industrial processing tanks). Additionally, bacteria are able to deteriorate metals through microbially induced corrosion (MIC).26 Although metal ions can exhibit toxicity to planktonic bacteria,13,27 biofilms can actively incorporate ions into the EPS, leading to their long-term survival. Prior studies have demonstrated that treatment with metal ions can significantly influence biofilm mechanical properties, such as modulus (via shear rheology),13,14,19,28 adhesion or failure behavior (via erosion assays),13,14 and surface hydrophobicity.29 However, standard shear rheology techniques for biofilms often require scraping colonies from solid substrates and loading them onto a rheometer, which disrupts the biofilm structure. Erosion studies preserve the biofilm structure but do not yield quantitative mechanical parameters.
In this work, we introduce a new approach to tensile testing bacterial biofilms. Our approach builds on existing rheological, erosion resistance, and tensile testing techniques by quantitatively capturing both high- and low-strain behavior in response to exogenous environmental signals. Tensile stresses are physiologically relevant to biofilms in flow environments, such as pipes or catheters, where shear flow can induce extensional (stretching) deformation of surface-attached biofilms. We employ the soil-dwelling, non-pathogenic bacterium Bacillus subtilis (B. subtilis) as our model organism due to its well-characterized genetic regulation of biofilm formation30 and matrix composition,31 robust biofilm formation capabilities,32 and precedence in biofilm mechanics research.13–15,17,20,22–24,33B. subtilis is also ecologically and practically significant—it colonizes in the human gastrointestinal tract,34 is used as a probiotic,35 and forms root-associated biofilms that promote nutrient uptake.36 Prior work has applied tensile testing to B. subtilis colonies on agar plates20 as well as pellicles,22,23 which are biofilms that grow at the air–liquid interface; however, these studies were limited by their ability to provide quantitative mechanical parameters and long experimental timelines, respectively. The method described here implements a custom-built cantilever-based instrument, The Uniaxial Tensile Tester for UltraThin films (TUTTUT), which has previously been used for tensile testing on synthetic polymer ultrathin films (<200 nm).37–41 Critically, TUTTUT characterization preserves the native pellicle structure, enables the introduction of environmental cues at various points of biofilm development, and provides quantitative measurements of mechanical properties across the complete strain range in one test. Using this method, we find that B. subtilis pellicles display viscoelastic behavior, consistent with previous reports.22,23 To investigate how environmental metal ion incorporation influences biofilm mechanical response, we characterized how the B. subtilis pellicle stress–strain response changes upon exposure to Fe3+, Fe2+, Cu2+, Cu+, Ca2+, and Na+. Our results suggest that certain metal ions can significantly impact low strain elastic modulus (E), maximum stress (σmax), and failure strain (εf), providing quantitative insight into metal ion effects on biofilm mechanics and demonstrating the versatility of our TUTTUT system for studying the impact of diverse environments on the mechanical behavior of biofilms.
2. Results and discussion
Here, we modify TUTTUT to directly measure the uniaxial stress–strain response of B. subtilis pellicles (Fig. 1A). Prior to tensile testing biofilms, TUTTUT was used to characterize the stress–strain response of ultrathin polymer films (<200 nm).37–41 TUTTUT leverages a liquid support bath to manipulate difficult-to-handle materials, making it well-suited for characterizing ultrasoft biofilms (i.e., ∼100 Pa–10 kPa13–15,22,23). TUTTUT is a cantilever-based method, in which force resolution can be tuned by adjusting the material and geometry of the cantilever, enabling precise force measurements across a wide modulus range—from soft biofilms with elastic moduli in the Pa range to stiff polymer films in the GPa range. This makes TUTTUT broadly applicable to materials with widely varying mechanical properties. To the best of our knowledge, only one other research group has tested the tensile properties of pellicles.22,23 Their approach, however, relies on growing the biofilm directly on the liquid surface within their instrument, resulting in natural adhesion onto the force sensors and instrument walls. While this approach preserves in situ biofilm growth, it constrains experimental throughput and parameter space, as evidenced by the authors’ report of approximately 100 biofilms studied over five years.22In this work, B. subtilis pellicles are grown outside of the liquid support bath in TUTTUT (Fig. 1A), allowing successive measurements without culture time restraints. We accomplish this by growing rectangular pellicles in 3D printed molds which generate two rectangular test samples from a connected media source (see the Experimental methods section and Fig. S1). To minimize clamping and end effects, samples were prepared with lengths greater than their widths (aspect ratios 1.85–3.86 (L/W), see the SI). Prior work on rectangular freestanding polymer thin films has shown that apparent geometry dependent differences in tensile response primarily arise from system compliance rather than intrinsic material behavior.42 After growth, additional liquid media are added to the culture container to float the biofilms out of the molds (Movie S1 and Fig. S2). One biofilm is transferred to TUTTUT, and the other is harvested onto a glass slide for determining the biofilm thickness (see the Experimental methods section and Fig. S3). Because tensile testing and thickness measurements are performed on two separate pellicle strips, some variability between films could introduce error into the calculated stress. Films grown within the same culture container exhibited an average thickness difference of 10.6 ± 8.88% (Fig. S4). Additional sources of uncertainty may arise from surface wrinkling or trapped water during optical profilometry measurements of pellicles on glass slides. However, these effects were minimized through consistent handling and imaging conditions and are not expected to substantially alter the reported mechanical trends. After transferring the biofilm to the bath, the biofilm is attached to the bath and the cantilever (Fig. 1A). To perform tensile testing, a linear actuator stretches the biofilm at a constant strain rate (0.0026–0.0045 s−1, previously used in B. subtilis tensile testing22) causing the cantilever to deflect. The cantilever is calibrated for force and displacement, and using biofilm geometry, we calculate stress and strain (see the Experimental methods section).
The stress–strain curves for untreated B. subtilis pellicles are shown in Fig. 1B. Similar to what has previously been observed,22 there is an initial linear-elastic regime at low strains followed by nonlinear behavior. Low strain elastic modulus (E) was determined from the slope of the stress–strain curve between 0 and 5% strain (Fig. S5). To assess the influence of a potential toe region, linear fits were also performed excluding the initial 0.5% strain; however, consistent E trends were obtained in both analyses (see Table S1). In addition, E was also calculated from linear fits over the 0–10% strain range (Table S2). Because some samples exhibited non-linear behavior beginning around 6% strain for some samples (see Fig. S5C), E values reported in the main text are based on the 0–5% strain region. Notably, two populations of E within the stress–strain curves appear to be present for untreated pellicles, despite being prepared and tested under the same conditions; however, we found no clear factors (e.g. sample history or handling) that could account for the variation. We determine the low strain elastic modulus, E ∼ 850 ± 310 Pa, the maximum stress, σmax ∼ 120 ± 62 Pa, and the failure strain, εf ∼ 26 ± 15%. E and σmax are of the same order of magnitude as the literature values (E ∼ 200–400 Pa and σmax ∼ 50–200 Pa).22,23 In prior tensile studies, residual compressive stress is present due to pellicle growth in the testing apparatus.22,23 Here, we release pellicles from their culture molds prior to loading, thereby relieving growth-induced compressive stresses and defining a relaxed reference state for tensile deformation. Variations between our results and previously reported values may be attributed to differences in sample age and thickness measurement protocols. In this study, samples were tested on day 3, whereas prior studies evaluated samples aged for 2 (ref. 22) or 4–7 (ref. 23) days. Moreover, the thickness was measured individually for each culture container using optical profilometry in the present work, while previous studies assumed a constant average thickness of 350 µm across all samples, as measured using a needle plunge technique and microscopy.22–24 Furthermore, whereas previous researchers required five years to collect data on over 100 pellicles,22 TUTTUT enabled the characterization of 82 pellicles within just 26 testing days, corresponding to an average throughput of approximately ∼3.2 samples per day. We note that a small number of tensile tests were excluded from analysis due to pellicle slipping at the cantilever or clamp (12 samples) or due to experimental setup issues (15 samples). However, even with these exclusions, our throughput is still a substantial improvement in experimental efficiency compared to previous tensile testing methods.
Beyond increasing the throughput for measuring the mechanics of untreated pellicles, TUTTUT provides a platform to measure the influence of external environmental factors (e.g. free metal ions) on the mechanical properties of biofilms (Fig. 2). Here, we assess the impact of Cu2+ and Fe3+, which are relevant to biofilms in copper and iron piping systems. We subsequently tested Fe2+ and Cu+ to probe the impact of metal identity and ion valency. Additionally, we probe the influence of Ca2+ and Na+, which are common free metal ions in water systems.43,44 After three days of growth, pellicles were released from their molds and exposed to 50 mM metal ion solutions for one hour by replacing the culture media with metal-supplemented media, such that they are in a free-floating state (see the Experimental methods section and Fig. S2). The metal ion concentration and exposure time were selected to mirror previous studies employing rheology and erosion analyses to probe the influence of metal ion exposure on biofilm mechanics. Additionally, we calculated an estimated diffusion time of ∼36 minutes, assuming the highest biofilm thickness (859 µm), lowest ion diffusivity (0.604 × 10−9 m2 s−1 for Fe3+),45 and ratio of effective diffusivity through biofilms to aqueous diffusivity (0.56).46 Treatment with 50 mM Fe3+ and Cu2+ resulted in a pellicle color change from off-white to rust and blue (Fig. 2A), respectively, indicating the absorption of these ions into the biofilm matrix. We observe no change in lateral dimension; however, we note a reduction in thickness compared to that of untreated biofilms for all metal ion treatments except Fe3+-treated pellicles. This change in thickness is accounted for in stress calculations. For all metal ion treatments, we measure a stress–strain curve with a shape similar to that of the untreated biofilms (Fig. 2A). Metal ion treatment leads to an increase in E and σmax as well as a decrease in εf compared to the untreated biofilms (Fig. 2B–D). To understand the impact of different metal ion treatments on biofilm mechanics, we compare E, σmax, and εf.
Among pellicles treated with the trivalent ion, Fe3+ had the largest increase in the average E—an increase by an order of magnitude—followed by Fe2+-treated pellicles, which resulted in a >3-fold increase in E, and then Cu2+-treated pellicles (∼2.9-fold increase in E, Fig. 2B). Pellicles treated with other bivalent and monovalent ions exhibit a more moderate increase in E, although these changes are not statistically significant (Fig. 2B). It should be noted that Fe2+ and Cu+ solutions likely underwent partial oxidation to Fe3+ and Cu2+, respectively, as suggested by a slight color change in the surrounding media during testing. Thus, the mechanical responses observed for these samples may reflect contributions from both oxidation states rather than a single, well-defined ionic species. Researchers have observed similar increases in the storage modulus (G′) for Pseudomonas aeruginosa (P. aeruginosa),28Escherichia coli,19 and B. subtilis (B-113 and NCIB 361014) colonies with the addition of trivalent metal cations. Increases in E have also been observed in P. aeruginosa biofilms with Ca2+ exposure due to Ca2+-mediated ionic crosslinking of alginate in the P. aeruginosa EPS.47 The increase in E with Fe3+ and Cu2+ treatment seen in this work is consistent with previously reported changes in G′ for ion-treated NCBI 3610 colonies;14 however, while we observe a slight, although not significant, increase in E with Na+ and Ca2+ treatment, previous measurements show a decrease or no change in G′.13,14 This discrepancy may arise from differences in sample preparation or growth method, which are known to influence the measured mechanical properties of biofilms.48 Here, tensile tests are performed on pellicle biofilms, whereas previous work implemented shear rheology and erosion assays on biofilm colonies grown on agar plates.
The increase in E is associated with nonspecific ionic crosslinking between EPS components and metal ions. In B. subtilis, positively charged metal cations interact with negatively charged functional groups on proteins (e.g., carboxylic acid and aspartate), leading to increased crosslinking of the EPS.26,49,50 Additionally, cryo-electron microscopy on the B. subtilis EPS component TasA has suggested that aspartate residues on TasA can coordinate metal cations.51 Furthermore, erosion assays performed on metal ion-treated NCIB 3610 colonies, specifically those performed on mutants lacking key EPS components, demonstrated that enhanced biofilm attachment arises from interactions between the metal ions and the EPS, not specific to any particular EPS component.14 This mechanism parallels that of metal-ion coordinated hydrogels, where reversible coordination between multivalent cations and polymer functional groups enhances mechanical properties, such as stiffness and toughness.52 Biofilms are often compared to colloidal hydrogels, where the cells are the colloids and the EPS is the crosslinked gel.49,53,54 Similar to the structural role that metal ion coordination plays in hydrogels, ion coordination in biofilms may reinforce the EPS through dynamic crosslinking interactions. Although Na+ is monovalent and therefore unlikely to form crosslinks in the biofilm matrix, its addition is associated with a modest, though not statistically significant, increase in E. This effect may arise from nonspecific ionic screening that reduces electrostatic repulsion within the EPS and/or osmotic deswelling of the biofilm matrix.
In further support of our ionic crosslinking hypothesis, we observe a trend of decreasing εf with the addition of the metal ions (Fig. 2D). Although Fe3+ treated biofilms are the only group which displays a statistically significant decrease in εf from the untreated biofilms, a trend of decreasing εf with metal ion treatment is present. We propose that this decrease arises from an increase in network connectivity—specifically, higher crosslink density—which limits the ability of the biofilm to deform before failure. This interpretation aligns with findings from tensile tests on ionically crosslinked hydrogels, where increasing crosslink density, achieved through increased metal cation content, leads to a reduction in εf.55 In further support of ion-mediated stabilization, previous work showed that supplementing B. subtilis NCIB 3610 cultures with Ca2+ limits biofilm dispersal by promoting structural stabilization,56 suggesting that metal ion-induced stabilization may influence biofilm persistence and failure. To our knowledge, this is the first quantification of εf in B. subtilis pellicles. We note that εf has been quantified for the surface of other biofilms using a microcantilever-based technique.57
Two failure modes were commonly observed during tensile testing: either failure initiated at pre-existing cracks introduced during removal from culture molds or new cracks or holes formed and propagated under tension. In most cases, failure occurred near the grip interface, typically within approximately 5 mm from the end of the film. While the presence of pre-existing cracks did not appear to significantly alter the εf, samples with pre-existing cracks tended to exhibit lower maximum stress values at failure (Table S3). However, definitive conclusions cannot be drawn due to a limited number of replicates.
The addition of metal ions results in an increase in σmax, with Fe3+ producing the highest σmax, followed by Cu2+, Fe2+, Cu+, and then Na+ (Fig. 2C). Similar trends are seen in yield stress, σy (see Fig. S6). We attribute these increases to ionic crosslinking; however, we note that the differences in σmax between the untreated group and the Fe2+, Cu+, Na+, and Ca2+ treated groups are not statistically significant. Because σmax reflects the maximum stress a biofilm can sustain prior to failure, it is relevant for understanding biofilm resistance to extreme mechanical perturbations, such as high shear or disruptive forces.
Differences in σmax values between metal ion treatments have several potential sources, including variations in the spatial distribution of the metal ions within the EPS or metal ion electronic structure. X-ray fluorescence (XRF) measurements from prior work support this idea, showing that Ca is distributed uniformly throughout the biofilm matrix, whereas Zn, Mn, and Fe are specifically enriched within biofilm wrinkles.58 The spatial distribution of ionic crosslinks can influence mechanical behavior by creating regions of localized stress concentration that are more susceptible to crack initiation,59 which can impact the measured σmax. In addition to spatial effects, differences in metal identity may also contribute to the observed changes in all properties measured. For example, in alginate hydrogels crosslinked with metal ions, E follows the trend, Fe3+ > Cu2+ > Ca2+, which is attributed to differences in ion charge, size, coordination number, and binding affinity. Trivalent ions, like Fe3+, can coordinate with three carboxylic groups on alginate, forming more crosslinks and generating stiffer networks. Divalent ions such as Fe2+, Cu2+ and Ca2+ typically coordinate with two groups, leading to more compliant structures.60 We observe a similar trend in biofilms, where Fe3+ treatment induces the largest change in measured properties among all ions tested. The difference in E for Cu2+ and Ca2+ crosslinkers in alginate hydrogels was attributed to the affinity of each ion to binding alginate.60 This could play a role in the differences in E observed in biofilms for divalent crosslinkers here: ions may have differing affinities for EPS polymers. Additionally, metal ions exhibit preferred coordination geometries based on their electronic configurations, but the specific binding environments presented by EPS components may constrain ions into less favorable geometries,52 giving rise to metal identity-dependent mechanical properties.
To test our ionic crosslinking hypothesis, we introduced an additional wash step to remove metal ions from pellicles pre-treated with Fe3+. Fe3+ was selected for this study because it caused the most pronounced mechanical differences from untreated biofilms. In this study, following treatment with 50 mM Fe3+, pellicles were subjected to a secondary one hour wash step using LB–glycerol–manganese (LBGM) media or 25 mM ethylenediaminetetraacetic acid disodium salt (EDTA), a strong chelating agent. The LBGM media wash results in slight recovery of the stress–strain response (Fig. 3A) and moderate reduction of the rust color associated with Fe3+ treatment (Fig. 3A). These changes suggest limited removal of Fe3+ ions and partial disruption of the ionic crosslinks within the EPS using LBGM media. In contrast, the EDTA wash produces a more dramatic color change, indicating more effective removal of Fe3+ from the EPS (Fig. 3A). Mechanically, the EDTA wash resulted in pellicle stress–strain curves and mechanical property metrics (E, σmax, or εf) that closely resembled those of EDTA-only controls (i.e., pellicle treated with EDTA but no Fe3+), indicating reversal of Fe3+ induced stiffening (Fig. 3B–D, refer to Fig. S7 for yield stress). The stronger impact of EDTA wash compared to that of the LBGM media wash is due to the high affinity of EDTA for metal ions,61,62 allowing it to more effectively disrupt Fe3+–EPS interactions. While color changes and EDTA reversibility support ionic participation, additional studies would be required to directly quantify metal content or map its spatial distribution within the EPS. Trends seen here agree with erosion studies on B. subtilis B-1 colonies, where an EDTA wash following Fe3+ treatment restored biofilm detachment to ∼100%.13
Notably, treatment with EDTA alone seemed to impact biofilm mechanics. While the Na+ ions present in the EDTA solution could contribute to the observed changes, EDTA-specific interactions are also likely to play a role. In contrast to prior findings showing that EDTA compromises the structural integrity in biofilms formed by pathogenic bacteria,63,64 we see a slight decrease in εf for the EDTA only treatment, potentially indicative of increased network connectivity. This mechanical response could be influenced by interactions between EDTA and extracellular DNA (eDNA) in the EPS65 or pH effects.58,66 Previous studies have suggested that EDTA may enhance single-stranded DNA binding to B. subtilis cells,65 potentially introducing additional crosslinks within the biofilm matrix and contributing to the observed reduction in εf. Additionally, previous work has shown that TasA fibers exhibit polymorphism and undergo configurational changes or aggregation under acidic conditions.58,66 This may result in biofilm matrix stiffening due to EDTA exposure and could impact stress–strain behavior.
Given the acidity of EDTA and that the addition of metal ions to LBGM media causes a reduction in pH (Table S4), we next explored the impact of pH on the biofilm tensile response. Similar to our metal ion studies, we perform TUTTUT testing on pellicles exposed to LBGM media adjusted to pH 2.0, 4.0, 8.0, and 10.0 for one hour (Fig. 4 and Fig. S8 and S9). Treatment with pH adjusted media causes slight changes in measured mechanical properties for a few treatment groups, although much smaller than those caused by the metal ion solution with its corresponding pH. Thus, we attribute the increase in E and σmax and the decrease in εf to the presence of metal ions as opposed to the pH change. However, LBGM media adjusted to pH 4.0, which is closest to the pH of 25 mM EDTA, result in a moderate increase in E and a decrease in εf, suggesting that pH may contribute to the effects observed with EDTA treatment.
3. Conclusions
We introduce a method, TUTTUT, to directly measure the uniaxial stress–strain relationship of bacterial biofilms. Using TUTTUT, we find that the E, σmax, and εf of B. subtilis pellicles are impacted by the absorption of the metal ions Fe3+, Fe2+, Cu2+, Cu+, Na+, and Ca2+ into the biofilm matrix. pH is found to have a minimal influence on the observed changes in mechanical properties. Additionally, the mechanical properties of untreated pellicles can be partially recovered from Fe3+ pre-treated pellicles through the introduction of a chelating agent, EDTA. The role of metal ions in controlling biofilm mechanics highlights the importance of considering the ionic environment when developing strategies for biofilm removal or control. The presence of strongly bound metal ions, such as Fe3+, may confer increased mechanical stability, making biofilms more resistant to mechanical removal. Effective biofilm mitigation strategies may therefore require the identification and targeted removal of ions from the EPS. Furthermore, these results are relevant to a wide range of applications in which biofilms interact with materials—such as medical devices, water infrastructure, and industrial surfaces—where local ion availability may significantly influence the biofilm structure, persistence, and response to treatment.4. Experimental methods
4.1 Bacterial culture
Liquid cultures of B. subtilis NCIB 3610 were generated by inoculating 2% w/v LB broth (Lennox) with a piece of frozen bacterial glycerol stock, incubating overnight at 30 °C and shaking at 240 rpm. Overnight cultures were then diluted 1000× in LB and incubated at 30 °C with shaking at 240 rpm until an optical density (OD) at 600 nm of ∼0.1 was reached, resulting in our starting culture. To achieve the desired pellicle testing geometry, we 3D printed culture molds (i.e., volume-fixed containers) that have two rectangular compartments (25 mm × 7.5 mm × 10 mm deep) and one circular port for adding additional media to the mold after biofilm growth (Fig. S1). Rectangular sample geometries ease the calculation of mechanical properties and allow for comparisons across conditions. Culture molds grow two pellicle strips from a connected media reservoir: one is used for tensile testing and the other for thickness measurements. The molds were secured within sterile magenta plant culture boxes and then filled with LBGM media (2% LB, 0.1 mM MnCl2, and 3% glycerol) such that there was ∼1 cm of media covering the bottom of the culture (7.5–35 mL total volume). Our starting culture was inoculated into the molds, resulting in an OD600 ≈ 0.001. The culture box was covered with parafilm and incubated at room temperature for 3 days prior to mechanical testing.4.2 Biofilm harvesting and testing
After 3 days of growth, a scalpel was used to detach biofilm samples from the edges of the 3D printed molds. Biofilms were floated up out of their culture molds by adding additional media to the container (Movie S1 and Fig. S2). Then, one biofilm was transferred from the culture container to the TUTTUT bath. For metal ion studies, biofilm samples were floated in 50 mM FeCl3, FeCl2, CuSO4, CuCl, CaCl2, or NaCl supplemented media for an exposure time of one hour prior to transferring to TUTTUT. The remaining biofilm was harvested on a glass slide for thickness measurements. Only biofilms in which both pellicle strips displayed a wrinkled morphology were used for measurements. Once in the TUTTUT bath, a clamp fabricated from rectangular coverglass slides (30 mm × 5 mm) with an ∼0.2 mm PDMS (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 cross-linker
10 cross-linker![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) silicone elastomer base) coating on one side was dropped onto one end of the biofilm and was secured into place on the bath using magnets. The free edge of the biofilm was then aligned with the extension piece of the cantilever, and the bath was raised to attach the sample to the cantilever grip, resulting in samples with aspect ratios (L/W) ranging from 1.8 to 3.9 (see the SI for all sample aspect ratios). To perform tensile testing, a linear actuator on TUTTUT caused the clamp to move away from the cantilever at a fixed velocity (vactuator = 0.1 mm s−1), stretching the biofilm at a fixed strain rate (0.00295–0.00446 s−1) and causing the cantilever to deflect (Movie S1). Tensile tests in which the pellicle slipped from either the clamp or cantilever grip were excluded from analysis.
silicone elastomer base) coating on one side was dropped onto one end of the biofilm and was secured into place on the bath using magnets. The free edge of the biofilm was then aligned with the extension piece of the cantilever, and the bath was raised to attach the sample to the cantilever grip, resulting in samples with aspect ratios (L/W) ranging from 1.8 to 3.9 (see the SI for all sample aspect ratios). To perform tensile testing, a linear actuator on TUTTUT caused the clamp to move away from the cantilever at a fixed velocity (vactuator = 0.1 mm s−1), stretching the biofilm at a fixed strain rate (0.00295–0.00446 s−1) and causing the cantilever to deflect (Movie S1). Tensile tests in which the pellicle slipped from either the clamp or cantilever grip were excluded from analysis.
4.3 Thickness measurements
A Keyence VR-3000 3D Optical Profilometer (OP) was used to scan the 3D profiles of biofilms on glass slides. Using instrument analysis software, eight equally sized squares were selected within the sample, returning each square's average thickness with respect to the instrument stage. The thickness of the glass slide was subtracted from measurements, and the eight thickness values were averaged to obtain the average biofilm thickness for that culture container.4.4 Investigation of pH effects
LBGM media (pH 7.0) were prepared as described in the Bacterial culture section and pH was adjusted to 2.0, 4.0, 8.0, or 10.0 using 1 M HCl and 1 M NaOH. To determine the impact of pH on the biofilm stress–strain response, biofilms were cultured for 3 days and then floated in pH adjusted media for an exposure time of one hour prior to tensile testing.4.5 Chelation based analysis of biofilm-Fe3+ affinity
Pellicles were treated with 50 mM FeCl3 for one hour prior to transfer to a wash bath, containing either 25 mM EDTA or LBGM media. After one hour, biofilms were transferred to TUTTUT for mechanical testing as described in the Biofilm harvesting and testing section.4.6 Cantilever fabrication and calibration
Force readings were determined using the stiffness of cantilevers (Sc) fabricated from rectangular polyethylene terephthalate (PET) sheets. The cantilever material was selected to be at least 2× stiffer than biofilm samples. Cantilever calibrations and stress–strain calculations were performed as previously described.37 In brief, Sc was calculated by measuring resonance frequency (f) at varying cantilever lengths (Lc). f was measured using a LK-G5000 Series Laser Displacement Sensor (1000 Hz) to capture the cantilever displacement (δcant) that occurred in response to tapping the cantilever. A Fourier transform was fit to these data to determine f. Three f measurements were taken for five different Lc values approximately 5 mm apart. We calculated the stiffness using f, Lc, total cantilever length (Ltotal), and total cantilever mass (mtotal) by67 | (1) |
To establish a relationship between laser displacement (δlaser) and δcant, a displacement is applied in the x-direction using the linear actuator and the resulting δlaser is measured. A linear fit yields slope m1. δcant is determined by dividing δlaser by m1, and force is then determined by F = Sc × δcant. Then, biofilm displacement (δfilm) is determined by δfilm = vactuator × t − δcant, where t is the time. Strain (ε) is determined by 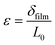 , where L0 is the initial length of the film between grips. Stress (σ) is calculated by
, where L0 is the initial length of the film between grips. Stress (σ) is calculated by 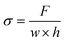 , where w and h are the biofilm width and thickness, respectively.
, where w and h are the biofilm width and thickness, respectively.
4.7 Statistical analysis
All quantitative data are presented as mean ± standard deviation (SD) (n ≥ 3). Statistical significance between sample means was determined using a one-way analysis of variance (ANOVA), followed by a Fisher's least significant difference (LSD) post hoc test for pairwise comparisons in OriginLab. Statistical significance with p < 0.05, p < 0.01, and p < 0.001 is indicated with an *, **, and ***, respectively.Author contributions
Kiera J. Croland: conceptualization, data curation, formal analysis, investigation, methodology, validation, visualization, and writing—original draft. R. Kōnane Bay: conceptualization, funding acquisition, methodology, project administration, resources, software, supervision, visualization, and writing—review and editing.Conflicts of interest
There are no conflicts to declare.Data availability
Supplementary information: all stress–strain responses and the properties extracted from stress–strain curves (low strain elastic moduli, maximum stresses, yield stresses, and failure strains). See DOI: https://doi.org/10.1039/d5sm01096a.Acknowledgements
KC sincerely thanks Ava Crowley for supporting this work through generating diagrams of TUTTUT and for guidance and training in the Huli Materials Lab. We would like to acknowledge the Living Materials Laboratory of Wil Srubar at CU Boulder for providing the B. subtilis NCIB 3610 strain used in this work. We also thank Joselle McCracken and the White laboratory at CU Boulder for supporting this work through providing access to the structured light profilometer for thickness measurements. This project was financially supported by the National Science Foundation Graduate Research Fellowship (NSF DGE # 2040434) and the University of Colorado Boulder Startup Funds.References
- K. Kovach, M. Davis-Fields, Y. Irie, K. Jain, S. Doorwar, K. Vuong, N. Dhamani, K. Mohanty, A. Touhami and V. D. Gordon, npj Biofilms Microbiomes, 2017, 3, 1 CrossRef PubMed.
- D. G. Metcalf and P. G. Bowler, Burns Trauma, 2013, 1, 2321–3868 CrossRef PubMed.
- X. Li, L. Sun, P. Zhang and Y. Wang, Coatings, 2021, 11, 294 CrossRef CAS.
- H. J. Busscher, H. C. van der Mei, G. Subbiahdoss, P. C. Jutte, J. J. A. M. van den Dungen, S. A. J. Zaat, M. J. Schultz and D. W. Grainger, Sci. Transl. Med., 2012, 4, 153rv10 CrossRef PubMed.
- M. Cámara, W. Green, C. E. MacPhee, P. D. Rakowska, R. Raval, M. C. Richardson, J. Slater-Jefferies, K. Steventon and J. S. Webb, npj Biofilms Microbiomes, 2022, 8, 1–8 Search PubMed.
- G. Liu, Y. Zhang, W.-J. Knibbe, C. Feng, W. Liu, G. Medema and W. van der Meer, Water Res., 2017, 116, 135–148 CrossRef CAS PubMed.
- P. S. Stewart and J. W. Costerton, Lancet, 2001, 358, 135–138 CrossRef CAS PubMed.
- S. Saini, S. Tewari, J. Dwivedi and V. Sharma, Mater. Adv., 2023, 4, 1415–1443 RSC.
- J. Huang, S. Liu, C. Zhang, X. Wang, J. Pu, F. Ba, S. Xue, H. Ye, T. Zhao, K. Li, Y. Wang, J. Zhang, L. Wang, C. Fan, T. K. Lu and C. Zhong, Nat. Chem. Biol., 2019, 15, 34–41 CrossRef CAS PubMed.
- L. M. González, N. Mukhitov and C. A. Voigt, Nat. Chem. Biol., 2020, 16, 126–133 CrossRef PubMed.
- T. Guélon, J.-D. Mathias and P. Stoodley, in Biofilm Highlights, ed. H.-C. Flemming, J. Wingender and U. Szewzyk, Springer, Berlin, Heidelberg, 2011, pp. 111–139 Search PubMed.
- H. Boudarel, J.-D. Mathias, B. Blaysat and M. Grédiac, npj Biofilms Microbiomes, 2018, 4, 17 CrossRef PubMed.
- S. Grumbein, M. Opitz and O. Lieleg, Metallomics, 2014, 6, 1441–1450 CrossRef CAS PubMed.
- M. Klotz, M. Kretschmer, A. Goetz, S. Ezendam, O. Lieleg and M. Opitz, RSC Adv., 2019, 9, 11521–11529 RSC.
- E. N. Hayta and O. Lieleg, Biomater. Sci., 2019, 7, 4675–4686 RSC.
- J. Yan, A. Moreau, S. Khodaparast, A. Perazzo, J. Feng, C. Fei, S. Mao, S. Mukherjee, A. Košmrlj, N. S. Wingreen, B. L. Bassler and H. A. Stone, Adv. Mater., 2018, 30, 1804153 CrossRef PubMed.
- S. Liu, J. Huang, C. Zhang, L. Wang, C. Fan and C. Zhong, Synth. Syst. Biotechnol., 2022, 7, 965 CrossRef CAS PubMed.
- L. A. Bakhtiari, M. J. Wells and V. D. Gordon, Biophys. Rev., 2021, 2, 031402 CrossRef CAS PubMed.
- A. Sarlet, V. Ruffine, K. G. Blank and C. M. Bidan, ACS Omega, 2023, 8, 4667–4676 CrossRef CAS PubMed.
- S. Grumbein, M. Werb, M. Opitz and O. Lieleg, J. Rheol., 2016, 60, 1085–1094 CrossRef CAS.
- I. Dogsa, M. Brloznik, D. Stopar and I. Mandic-Mulec, PLoS One, 2013, 8, e62044 CrossRef CAS PubMed.
- E. C. Hollenbeck, C. Douarche, J.-M. Allain, P. Roger, C. Regeard, L. Cegelski, G. G. Fuller and E. Raspaud, J. Phys. Chem. B, 2016, 120, 6080–6088 CrossRef CAS PubMed.
- M. Trejo, C. Douarche, V. Bailleux, C. Poulard, S. Mariot, C. Regeard and E. Raspaud, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 2011–2016 CrossRef CAS PubMed.
- C. Douarche, J.-M. Allain and E. Raspaud, Biophys. J., 2015, 109, 2195–2202 CrossRef CAS PubMed.
- M. U. Rahman, D. F. Fleming, L. Wang, K. P. Rumbaugh, V. D. Gordon and G. F. Christopher, npj Biofilms Microbiomes, 2022, 8, 1–9 Search PubMed.
- I. B. Beech and J. Sunner, Curr. Opin. Biotechnol, 2004, 15, 181–186 CrossRef CAS PubMed.
- J. A. Lemire, J. J. Harrison and R. J. Turner, Nat. Rev. Microbiol., 2013, 11, 371–384 CrossRef CAS PubMed.
- O. Lieleg, M. Caldara, R. Baumgärtel and K. Ribbeck, Soft Matter, 2011, 7, 3307–3314 RSC.
- C. Falcón García, M. Kretschmer, C. N. Lozano-Andrade, M. Schönleitner, A. Dragoŝ, Á. T. Kovács and O. Lieleg, npj Biofilms Microbiomes, 2020, 6, 1 CrossRef PubMed.
- H. Vlamakis, Y. Chai, P. Beauregard, R. Losick and R. Kolter, Nat. Rev. Microbiol., 2013, 11, 157–168 CrossRef CAS PubMed.
- S. S. Branda, F. Chu, D. B. Kearns, R. Losick and R. Kolter, Mol. Microbiol., 2006, 59, 1229–1238 CrossRef CAS PubMed.
- S. Gingichashvili, D. Duanis-Assaf, M. Shemesh, J. D. B. Featherstone, O. Feuerstein and D. Steinberg, Microorganisms, 2020, 8, 62 CrossRef CAS PubMed.
- D. Romero, C. Aguilar, R. Losick and R. Kolter, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 2230–2234 CrossRef CAS PubMed.
- H. A. Hong, R. Khaneja, N. M. K. Tam, A. Cazzato, S. Tan, M. Urdaci, A. Brisson, A. Gasbarrini, I. Barnes and S. M. Cutting, Res. Microbiol., 2009, 160, 134–143 CrossRef CAS PubMed.
- M. Marzorati, P. Van den Abbeele, S. S. Bubeck, T. Bayne, K. Krishnan, A. Young, D. Mehta and A. DeSouza, Microorganisms, 2020, 8, 1028 CrossRef CAS PubMed.
- A. Hashem, B. Tabassum and E. Fathi Abd_Allah, Saudi J. Biol. Sci., 2019, 26, 1291–1297 CrossRef CAS PubMed.
- R. K. Bay, S. Shimomura, Y. Liu, M. Ilton and A. J. Crosby, Macromolecules, 2018, 51, 3647–3653 CrossRef CAS.
- Y. Liu, Y.-C. Chen, S. Hutchens, J. Lawrence, T. Emrick and A. J. Crosby, Macromolecules, 2015, 48, 6534–6540 CrossRef CAS.
- W. J. Choi, R. K. Bay and A. J. Crosby, Macromolecules, 2019, 52, 7489–7494 CrossRef CAS.
- R. K. Bay, K. Zarybnicka, J. Jančář and A. J. Crosby, ACS Appl. Polym. Mater., 2020, 2, 2220–2227 CrossRef CAS.
- R. K. Bay, T. Zhang, S. Shimomura, M. Ilton, K. Tanaka, R. A. Riggleman and A. J. Crosby, Macromolecules, 2022, 55, 8505–8513 CrossRef CAS.
- H. Zhang, K. Fujiki, K. Ido, N. Kusayanagi, D. Sakagami, J. Li and Y. Okamura, Polym. Test., 2020, 91, 106825 CrossRef CAS.
- H. M. Thippeswamy, R. Shanbhog, M. N. Kumar, S. N. Prashanth and P. Smitha, Sci. Rep., 2025, 15, 10689 CrossRef CAS PubMed.
- H. S. Faust, Am. J. Clin. Nutr., 1982, 35, 1459–1467 CrossRef CAS PubMed.
- P. Vanysek, CRC Hand Book of Chemistry and Physics, 1993, pp. 5–92 Search PubMed.
- P. S. Stewart, Biotechnol. Bioeng., 1998, 59, 261–272 CrossRef CAS PubMed.
- V. Körstgens, H.-C. Flemming, J. Wingender and W. Borchard, Water Sci. Technol., 2001, 43, 49–57 CrossRef.
- S. Geisel, E. Secchi and J. Vermant, Interface Focus, 2022, 12, 20220032 CrossRef PubMed.
- J. N. Wilking, T. E. Angelini, A. Seminara, M. P. Brenner and D. A. Weitz, MRS Bull., 2011, 36, 385–391 CrossRef CAS.
- I. W. Sutherland, Microbiology, 2001, 147, 3–9 CrossRef CAS PubMed.
- J. Böhning, M. Ghrayeb, C. Pedebos, D. K. Abbas, S. Khalid, L. Chai and T. A. M. Bharat, Nat. Commun., 2022, 13, 7082 CrossRef PubMed.
- E. Khare, N. Holten-Andersen and M. J. Buehler, Nat. Rev. Mater., 2021, 6, 421–436 CrossRef CAS.
- N. Ido, A. Lybman, S. Hayet, D. N. Azulay, M. Ghrayeb, S. Liddawieh and L. Chai, Soft Matter, 2020, 16, 6180–6190 RSC.
- S. Arnaouteli, N. C. Bamford, N. R. Stanley-Wall and Á. T. Kovács, Nat. Rev. Microbiol., 2021, 19, 600–614 CrossRef CAS PubMed.
- K. J. Henderson, T. C. Zhou, K. J. Otim and K. R. Shull, Macromolecules, 2010, 43, 6193–6201 CrossRef CAS.
- M. Nishikawa and K. Kobayashi, J. Bacteriol., 2021, 203, 10–1128 CrossRef PubMed.
- S. Aggarwal and R. M. Hozalski, Biofouling, 2010, 26, 479–486 CrossRef PubMed.
- D. N. Azulay, O. Spaeker, M. Ghrayeb, M. Wilsch-Bräuninger, E. Scoppola, M. Burghammer, I. Zizak, L. Bertinetti, Y. Politi and L. Chai, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2118107119 CrossRef PubMed.
- N. Jian, J. Wang, L. Zuo and K. Zhang, Mater. Des., 2023, 225, 111522 CrossRef CAS.
- H. Malektaj, A. D. Drozdov and J. deClaville Christiansen, Polymers, 2023, 15, 3012 CrossRef CAS PubMed.
- A. Kovács, D. S. Nemcsok and T. Kocsis, J. Mol. Struct. THEOCHEM, 2010, 950, 93–97 CrossRef.
- R. M. Hutcheson, M. D. Engelmann and I. F. Cheng, Biometals, 2005, 18, 43–51 CrossRef CAS PubMed.
- R. Cavaliere, J. L. Ball, L. Turnbull and C. B. Whitchurch, MicrobiologyOpen, 2014, 3, 557–567 CrossRef CAS PubMed.
- X. Chen and P. S. Stewart, Appl. Microbiol. Biotechnol., 2002, 59, 718–720 CrossRef CAS PubMed.
- M. J. Tevethia and M. Mandel, J. Bacteriol., 1970, 101, 844–850 CrossRef CAS PubMed.
- M. Ghrayeb, S. Hayet, N. Lester-Zer, Y. Levi-Kalisman and L. Chai, Microorganisms, 2021, 9, 529 CrossRef CAS PubMed.
- M. F. Ashby and D. Cebon, J. Phys. IV France, 1993, 03, C7–C9 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |




