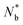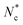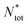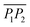Open Access Article
Open Access ArticleMultiscale reorganisation of colloidal aggregation by percolating bacterial networks
Laura Stricker†
*a,
Samuel G. V. Charlton†*b and
Eleonora Secchi *b
*b
aInstitute of Process Engineering, Otto von Guericke University Magdeburg, 39106 Magdeburg, Germany. E-mail: laura.stricker@ovgu.de
bETH Zurich, Department of Civil, Environmental and Geomatic Engineering, Zurich, 8093, Switzerland. E-mail: charlton@ifu.baug.ethz.ch; secchi@ifu.baug.ethz.ch
First published on 22nd April 2026
Abstract
Self-assembly in colloidal suspensions emerges from the interplay of local ordering and constraints imposed by the surrounding medium. While motile bacteria are known to alter colloidal dynamics, the influence of non-motile species remains largely unexplored. Here, we study suspensions of colloids and non-motile Comamonas denitrificans sedimenting near the wall and forming percolating networks. Using multiscale structural descriptors, we show that bacteria enhance colloidal aggregation into branch-like clusters. In turn, colloids reinforce bacterial networks by extending their elastic backbone. The analysis of mid-range ordering reveals that, unlike purely colloidal suspensions where ordering propagates across 3–4 neighbour shells, bacterial scaffolds suppress this propagation beyond the first shell. These findings highlight how non-motile bacteria reshape colloidal self-assembly across scales, while providing a quantitative framework for studying complex particle–network interactions. This approach opens pathways to understanding analogous processes in natural systems, including those involving microplastic contaminants.
Bacterial sedimentation is central to geochemical cycling, contributing to pollutant sequestration and carbon storage in sediments.1 Across terrestrial and marine environments, bacteria coexist with particulate matter, such as mineral grains, organic-rich soil aggregates, and particulate organic carbon.2–4 However, the dynamics of bacteria–particle interactions remain poorly understood. Most existing work has focused on motile species. Motile bacteria can hinder particle settling, drive the propagation of topological defects in dense colloidal mixtures, and are themselves accelerated when swimming in proximity to colloids.5–9 These phenomena are exploited in applications such as bacteria-driven mixing and interface stabilisation.10–12 In non-motile systems, studies have largely focused on monospecies suspensions where passive interactions such as depletion attraction and polymer bridging establish aggregation, and phenotypic traits such as filamentation control the topology and mechanics of the resulting networks.13 Less understood is the effect of non-motile bacteria on the formation of colloidal aggregates and networks, as well as the influence of colloidal–bacterial interactions on the development and settling of bacterial flocs.
Here, we study mixed suspensions of colloids and non-motile Comamonas denitrificans bacteria with similar sedimentation velocities. We focus on how the percolating bacterial network affects the near-wall layer colloidal structuring at different length scales; local (nearest neighbours), mid-range, and global (system-spanning) and how colloids in turn influence the bacterial network. Our analysis is based on two-dimensional projections of the sedimented near-wall layer and focuses on relative structural differences between bacteria–colloid mixtures and colloid-only suspensions. Using a combination of structural analysis and multiscale metrics, we characterise the self-assembly mechanisms that emerge in such mixed suspensions. Though multiscale analysis has been previously proposed to study hierarchical clustering and network formation gelation in soft particulate systems,14,15 to the best of our knowledge, this approach has not yet been used to investigate bacterial suspensions. We interpret our results within the framework of self-assembly, which arises from the minimisation of free energy – either through internal energy reduction, associated with short-range ordering via ‘locally favoured structures’ (LFS),16,17 or through entropy maximisation, which is conversely related to ordering at a larger scale.18,19 Geometric confinement strongly modulates colloidal clustering,20–23 and in our system, the near-wall bacterial network imposes such constraints. Short-range ordering remains unaffected, but pronounced changes occur at the mid-range and global scales. We explore this decoupling between length scales using tools from information theory, specifically mutual information,24 and from graph theory, through clique identification methods.25 While previous studies have hinted at a connection between local entropy, which is related to mid-range ordering, and locally favoured structures,26 our analysis offers a broader, multiscale perspective.
1 Materials and methods
1.1 Experimental system
Experiments were conducted by preparing liquid mixtures of colloidal particles and Comamonas denitrificans (referred to as ‘mixed suspensions’) and imaging their behaviour via optical microscopy. These were compared against two control conditions: suspensions containing only colloids (‘colloids-only’) and suspensions containing only bacteria (‘bacteria-only’). All suspensions were loaded into rectangular glass capillaries (0.25 × 10 × 40 mm3; Vitrocom, USA), with approximately ≈300 µL of sample per experiment. The ends of the capillaries were sealed with Vaseline to prevent evaporation, and the vessels were immediately transferred to the microscope stage. Structures formed at the bottom of the chamber were analysed after 140 minutes, once sedimentation was complete.1.2 Bacterial culture
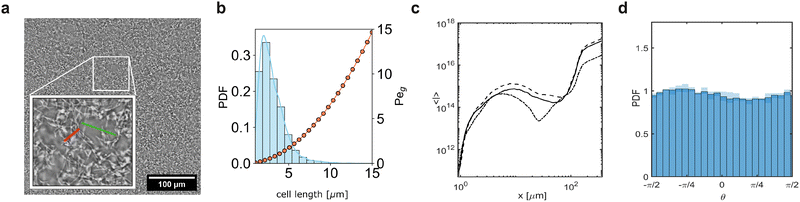
We perform the experiments using Comamonas denitrificans 123 (ATCC 700936), a rod-shaped, Gram-negative bacterium, cultured under conditions optimised to produce a narrow cell length distribution, with an average length of 4.16 µm and a standard deviation of 1.25 µm (see Fig. 2b). Glycerol cryostocks were routinely revived by inoculating 5 mL of Tryptic Soy Broth (TSB; Sigma-Aldrich) and incubating overnight at 30 °C with shaking at 180 rpm in 50 mL Falcon centrifuge tubes. The following day, subcultures were prepared by diluting the overnight culture 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 (v/v) into fresh 5 mL TSB and incubating for 6 h at 30 °C under the same shaking conditions, until cultures reached an optical density (OD600) of 1.0. Bacterial cell suspensions were aliquoted into 1.5 mL centrifuge tubes and washed three times with fresh TSB by centrifugation at 2700 rcf for 90 seconds. After the final wash, the cell pellets were resuspended in TSB containing the colloidal suspension for downstream assays.
100 (v/v) into fresh 5 mL TSB and incubating for 6 h at 30 °C under the same shaking conditions, until cultures reached an optical density (OD600) of 1.0. Bacterial cell suspensions were aliquoted into 1.5 mL centrifuge tubes and washed three times with fresh TSB by centrifugation at 2700 rcf for 90 seconds. After the final wash, the cell pellets were resuspended in TSB containing the colloidal suspension for downstream assays.
1.3 Colloidal particle selection and preparation
For the mixed and the colloids-only suspensions, we use FluoRed polystyrene colloids (2.5% w/v) with a diameter of 3.0 µm (microParticles GmbH, Germany), suspended in conditioned bacterial culture medium. The conditioned medium was obtained by filtering a bacterial culture with OD600 = 1 through a 0.2 µm filter (Filtropur S 0.2, Sarstedt, Germany). The colloids were washed three times by centrifugation at 2700 rcf for 90 seconds, each time replacing the supernatant with conditioned culture medium. After the final wash, the colloids were sonicated for 10 minutes to disrupt any aggregates. TSB is a high–ionic strength medium (≈130 mM salt equivalent, pH 7.0–7.2) in which electrostatic interactions are strongly screened (Debye length <1 nm); the physicochemical environment was identical in suspensions with and without bacteria, ensuring that effective interactions are short-ranged and dominated by steric exclusion. Note that the selected colloid size was chosen to match the sedimentation velocity of C. denitrificans, ensuring comparable gravitational Péclet numbers between bacteria and colloids (see Fig. 2b). Toward this aim, the gravitational Péclet number was estimated as Peg = V0(ρp − ρ)ga/(kBT), where ρp and ρ are the density of the particles and the liquid respectively, Vp is the volume of a particle, T is the temperature, g is the gravitational acceleration constant and kB is the Boltzmann constant.27 Colloids of the selected diameter have Peg ∼ 2.53, which falls within the range estimated for C. denitrificans bacterial cells (Peg = 1.7–4.4), assuming a cell radius of approximately ∼0.5 µm, a length within the 3–5 µm range, and a density ρp = 1.0825 g cm−3.1.4 Bacteria/colloid suspensions
We prepared three types of mixed bacterial–colloidal suspensions, each with a different concentration of colloidal particles (hereafter referred to as ‘colloidal densities’). To this end, a bacterial suspension was divided into three 1 mL aliquots and centrifuged at 2700 rcf for 5 minutes to sediment the cells. Subsequently, 100 µL, 50 µL or 25 µL of supernatant was replaced with an equal volume of colloidal suspension, yielding final volumetric colloid-to-bacteria ratios of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10, 1
10, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20, and 1
20, and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40, respectively. These mixtures correspond to final colloidal densities of 1.68 × 108 particles per mL (‘1
40, respectively. These mixtures correspond to final colloidal densities of 1.68 × 108 particles per mL (‘1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10’, high density), 8.4 × 107 particles per mL (‘1
10’, high density), 8.4 × 107 particles per mL (‘1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20’, medium density), and 4.2 × 107 particles per mL (‘1
20’, medium density), and 4.2 × 107 particles per mL (‘1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40’, low density). Each suspension was vortexed for 90 seconds to ensure appropriate resuspension of the solid phase.
40’, low density). Each suspension was vortexed for 90 seconds to ensure appropriate resuspension of the solid phase.
1.5 Bacteria-only and colloid-only suspensions
To assess the reciprocal influence between colloids and the bacterial network, we performed two sets of control experiments: one with bacteria-only and one with colloids-only suspensions. In the colloids-only controls, we used the same particle concentrations as in the corresponding mixed suspensions. Each sample was prepared by following the same liquid exchange procedure, starting from 1 mL of conditioned culture medium per mixture (see Fig. 3a, c and e). For the bacteria-only control experiments, we used the same bacterial cultures used as in the mixed suspensions, thereby maintaining identical bacterial concentrations across conditions.1.6 Microscopy and imaging conditions
Imaging was performed in both fluorescence and brightfield modes using an inverted microscope (Nikon Ti, Nikon, Japan). Images were acquired at 20× magnification (NA 0.5) with a 1.5× zoom (equivalent to a final 30× magnification), using an sCMOS camera (Photometrics Prime BSI, Teledyne, USA; pixel size 6.5 µm). Bacteria were imaged in brightfield mode with the condenser aperture set to 50% closure. Fluorescent imaging was used to visualise the colloidal particles, with excitation at 555 nm and emission at 570 nm (Lumencor Spectra, Lumencor, USA). The depth of focus of the optical configuration at 550 nm was approximately 2.85 µm, as estimated from the theoretical axial resolution of the imaging system using the standard wave-optical formulation for depth of field (DOF ≈ λn/NA2, where λ is the wavelength, n is the refractive index of the immersion medium, and NA is the numerical aperture). This value should be interpreted as an estimate of the axial extent contributing to the image, i.e. an upper bound for the optical section thickness. We acquired images immediately above the capillary floor, 1.5 µm into the sample. For each field of view, the optimal focus was re-established using the glass surface as a reference, ensuring that the near-wall bacterial layer was consistently imaged within the central, highest-resolution portion of the optical section despite minor local variations across the sample. Since the colloids have a diameter of 3.0 µm and the average bacterial width is approximately 1 µm, one optical section contains at most a single colloid layer and the portions of the bacterial branches that intersect the plane. All samples were imaged 140 minutes after capillary placement on the microscope stage, to ensure complete sedimentation. For each condition, three biological replicates from independent bacterial cultures were prepared to ensure reproducibility. At least four fields of view were acquired per sample to provide statistical robustness.1.7 Structural characterisation of the bacterial network
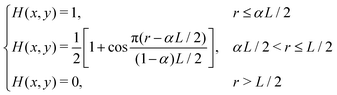 | (1) |
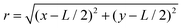 the radial position of the point (x, y) with respect to the centre of the image and α = 0.5 a scaling factor. We then computed the frequency-domain representation of the windowed image by applying a fast Fourier transform (FFT). The resulting 2D power spectrum was integrated over the angular coordinate from 0 to π to obtain a radially averaged profile. Finally, the resulting curves were transformed back from Fourier space to real space to extract the radial profile of the integrated power spectrum, denoted as 〈I(r)〉. The analysis was performed with an in-house code written in ImageJ-macro language, based on the native ImageJ FFT plugin, the Windowing plugin30 and Radial Profile Extended plugin,31 combined with an in-house Matlab code. Note that we chose to extract the characteristic mesoscopic length scale using the FFT approach rather than the ‘chord-length’ method proposed by other authors,32 as the latter requires binarising the microscopy images and therefore makes the results sensitive to the choice of threshold. Conversely, the FFT can be applied directly to grayscale images and is a well-established methodology in the analysis of isotropic, disordered, network-like structures with a characteristic domain size.
the radial position of the point (x, y) with respect to the centre of the image and α = 0.5 a scaling factor. We then computed the frequency-domain representation of the windowed image by applying a fast Fourier transform (FFT). The resulting 2D power spectrum was integrated over the angular coordinate from 0 to π to obtain a radially averaged profile. Finally, the resulting curves were transformed back from Fourier space to real space to extract the radial profile of the integrated power spectrum, denoted as 〈I(r)〉. The analysis was performed with an in-house code written in ImageJ-macro language, based on the native ImageJ FFT plugin, the Windowing plugin30 and Radial Profile Extended plugin,31 combined with an in-house Matlab code. Note that we chose to extract the characteristic mesoscopic length scale using the FFT approach rather than the ‘chord-length’ method proposed by other authors,32 as the latter requires binarising the microscopy images and therefore makes the results sensitive to the choice of threshold. Conversely, the FFT can be applied directly to grayscale images and is a well-established methodology in the analysis of isotropic, disordered, network-like structures with a characteristic domain size.1.8 Structural characterisation of colloidal patterns
1.8.1.1 Coordination number. The coordination number nc of a particle is defined as the number of its nearest neighbours. To calculate it, we counted, for each colloid, the number of particles located within a distance rn = 2ar0 from its centre, where r0 is the average colloid radius and a is a dimensionless constant. In our analysis, we set a = 1.25, and verified that the results remain qualitatively unchanged when a is varied by up to 10%. This choice of rn and its variation range corresponds to the position of the first minimum in the radial distribution function g(r), which identifies the first nearest-neighbours shell.
1.8.1.2 Centrosymmetry parameter. We further evaluated short-range ordering by calculating the centrosymmetry parameter of the colloids, based on their N = 6 nearest neighbours, following the method introduced in ref. 36:
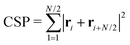 | (2) |
1.8.2.1 Mid-range order parameter. To characterize mid-range ordering, for each particle we calculated the mid-range order parameter si2, defined in ref. 26, 37, 38, and 39 as the ‘local entropy’ and approximated it as the two-body excess entropy.26,37 This kind of parameter is always negative, with more ordered structures corresponding to lower (i.e. more negative) values. For each particle i, the mid-range order parameter calculated over a neighbourhood of radius rm is
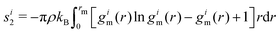 | (3) |
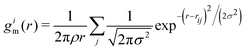 is a modified version of the radial distribution function centered around the ith particle, with rij the distance between particles i and j, and σ a broadening parameter (here σ = 0.05). To mitigate the effects of fluctuations, we calculated the average local entropy mid-range order parameter
is a modified version of the radial distribution function centered around the ith particle, with rij the distance between particles i and j, and σ a broadening parameter (here σ = 0.05). To mitigate the effects of fluctuations, we calculated the average local entropy mid-range order parameter
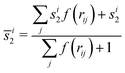 | (4) |
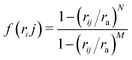 is a switching function with a cutoff ra = rm, N = 6 and M = 12. The averaged local entropy mid-range order parameter provides a stronger footprint of local ordering than the non-averaged one.26,37–39 Note that the absolute value of
is a switching function with a cutoff ra = rm, N = 6 and M = 12. The averaged local entropy mid-range order parameter provides a stronger footprint of local ordering than the non-averaged one.26,37–39 Note that the absolute value of ![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) 2 depends on the details of the estimation of the radial distribution function in Eqn (3), but its comparative value across different systems is still significant, as long as such details are kept constant.40 Since the particle positions are extracted from a single near-wall optical section, gim(r) should be understood here as an effective in-plane local pair-correlation function of the imaged layer, rather than as the bulk 3D radial distribution function of the full suspension. We use si2 for our analysis instead of gim(r) because the latter, though also localized at the particle i, is still a function of distance, whereas s2 provides a single scalar summary of the local correlation profile over the range 0 < r < rm. Accordingly, Eqn (3) is not a thermodynamic entropy, but is a scalar structural functional of the measured local pair correlation, useful for comparative analysis across samples.
2 depends on the details of the estimation of the radial distribution function in Eqn (3), but its comparative value across different systems is still significant, as long as such details are kept constant.40 Since the particle positions are extracted from a single near-wall optical section, gim(r) should be understood here as an effective in-plane local pair-correlation function of the imaged layer, rather than as the bulk 3D radial distribution function of the full suspension. We use si2 for our analysis instead of gim(r) because the latter, though also localized at the particle i, is still a function of distance, whereas s2 provides a single scalar summary of the local correlation profile over the range 0 < r < rm. Accordingly, Eqn (3) is not a thermodynamic entropy, but is a scalar structural functional of the measured local pair correlation, useful for comparative analysis across samples.
1.8.2.2 Relative mutual information. To quantify the relationship between any two properties associated with each colloidal particle, we computed the relative mutual information between them, in line with the approach proposed in ref. 41 for soft matter systems.42 In particular, we evaluated the relative mutual information M(NC, S)/I(Nc), between the coordination number and the average mid-range order parameter
![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) is calculated at a given length scale rm. For a given cutoff rm, the mutual information M(NC, S) is calculated as24
is calculated at a given length scale rm. For a given cutoff rm, the mutual information M(NC, S) is calculated as24
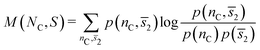 | (5) |
![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) i2(rm) ∈ S, i = 1,…N} and {NC: niC ∈ NC, i = 1, …, N} respectively; p(nC) and p(
i2(rm) ∈ S, i = 1,…N} and {NC: niC ∈ NC, i = 1, …, N} respectively; p(nC) and p(![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) 2) are the probabilities that, for a particle i, the coordination number is niC = nC and the average mid-range order parameter is
2) are the probabilities that, for a particle i, the coordination number is niC = nC and the average mid-range order parameter is ![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) i2 =
i2 = ![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) 2, while p(nC,
2, while p(nC, ![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) 2) indicates a joint probability. The self-information of NC is calculated as
2) indicates a joint probability. The self-information of NC is calculated as
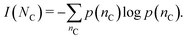 | (6) |
1.8.2.3 Cluster analysis. We identified the clusters as ensembles of particles within a distance rc = 2ar0 from each other. For each cluster, the gyration radius was calculated as
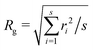 , where ri is the distance between the ith particle of the cluster and the cluster's centre of mass and s is the cluster's size, namely the number of colloids belonging to the cluster. Furthermore, we assessed the structure of the clusters, as emerging from the juxtaposition of uniform building units – i.e. similar local environments – by determining their tiling structure by triangular ‘cliques’ (i.e. triplets of adjacent colloids) and their network topology. To this aim, we first filtered out singlets and doublets, namely particles with less than two neighbours, by applying a rolling ball search algorithm43 with a radius rc around each particle. We then created an undirected graph with the (x, y) coordinates of the remaining colloids as nodes. The edges were delimited by pairs of neighbouring colloids, i.e. colloids within a distance rc. The connected components of the graph, detected through the algorithm described in ref. 25 and 44, corresponded to the colloidal clusters. Hence, their tessellation in triangular tiles was retrieved by detecting the completely connected 3-node subgraphs, i.e. the 3-cycles. Nodes that belonged to such triangular tiles were called 'clique-like' nodes. Nodes that did not belong to such triangular tiles and hence featured two neighbours unconnected to each other, were called ‘branch-like nodes’. Then, we evaluated the structural compactness of each cluster through the parameter ab = sb/s, where sb is the number of branch-like nodes in the cluster itself. Low values of ab indicate a more compact cluster structure, hence we refer to ab as the ‘airiness parameter’. Similarly, for the whole system, we evaluated the 'global airiness parameter' Ab = Nb/Ntot, where Nb and Ntot denote the total number of branch-like colloids and the total number of colloids (with the exclusion of singlets and doublets) respectively. Ab indicates the probability that neighbouring nodes are also connected, and it is therefore a measure of graph connectivity. The analysis was performed with an in-house Python code leveraging the NetworkX45 package.
, where ri is the distance between the ith particle of the cluster and the cluster's centre of mass and s is the cluster's size, namely the number of colloids belonging to the cluster. Furthermore, we assessed the structure of the clusters, as emerging from the juxtaposition of uniform building units – i.e. similar local environments – by determining their tiling structure by triangular ‘cliques’ (i.e. triplets of adjacent colloids) and their network topology. To this aim, we first filtered out singlets and doublets, namely particles with less than two neighbours, by applying a rolling ball search algorithm43 with a radius rc around each particle. We then created an undirected graph with the (x, y) coordinates of the remaining colloids as nodes. The edges were delimited by pairs of neighbouring colloids, i.e. colloids within a distance rc. The connected components of the graph, detected through the algorithm described in ref. 25 and 44, corresponded to the colloidal clusters. Hence, their tessellation in triangular tiles was retrieved by detecting the completely connected 3-node subgraphs, i.e. the 3-cycles. Nodes that belonged to such triangular tiles were called 'clique-like' nodes. Nodes that did not belong to such triangular tiles and hence featured two neighbours unconnected to each other, were called ‘branch-like nodes’. Then, we evaluated the structural compactness of each cluster through the parameter ab = sb/s, where sb is the number of branch-like nodes in the cluster itself. Low values of ab indicate a more compact cluster structure, hence we refer to ab as the ‘airiness parameter’. Similarly, for the whole system, we evaluated the 'global airiness parameter' Ab = Nb/Ntot, where Nb and Ntot denote the total number of branch-like colloids and the total number of colloids (with the exclusion of singlets and doublets) respectively. Ab indicates the probability that neighbouring nodes are also connected, and it is therefore a measure of graph connectivity. The analysis was performed with an in-house Python code leveraging the NetworkX45 package.
1.8.3.1 Lacunarity. The lacunarity λ of the colloidal pattern was calculated based on the detected positions of the colloidal centroids as
 | (7) |
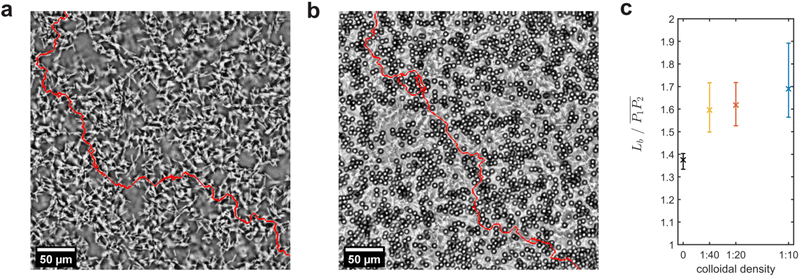
1.8.3.2 Elastic backbone. The elastic backbone of the percolating structure was defined as the union of all the shortest paths connecting two representative points, P1 and P2.46 The length of each one of such paths was denoted as Lb. In order to extract the elastic backbone, we processed the images collected by phase-contrast microscopy as described below. First, we performed a contrast-limited adaptive histogram equalisation (CLAHE).47 We then binarised the resulting images via thresholding, and we despeckled them with a median filter based on a 3 × 3 pixel square. In the resulting black and white images, both bacteria and particles were depicted in white, while the background was black. Such images were skeletonised by applying the thinning algorithm described in ref. 48. The extracted skeleton underwent an additional dilatation to match the thickness of the bacteria. Hence, we were able to extract the binarised connected bacterial structure. For the mixed suspensions, additional steps were required to guarantee that the colloids incorporated inside the network were correctly represented. To this aim, we separately reconstructed the binarised representation of the colloids, and we subsequently added it to the bacterial network. The binarised representation of the colloids was reconstructed by using the positions of their centroids as detected by TrackMate35 from the fluorescence images. Using such positions, we constructed a binary mask of single pixels, and we then applied a morphological dilatation filter based on a disk structure matching the colloidal diameter. Hence, we summed this image to the previously extracted binary bacterial network to reconstruct a binarised version of the complete percolating structure. These operations were performed by using an in-house code written in ImageJ macro language based on the FiJi native plugins ‘CLAHE’, ‘Threshold’, ‘Remove outliers’ and ‘Skeletonize’, as well as the ‘Morphological filters’ from the MorphoLibJ toolbox.49 Finally, we used the binarised image representing the percolating structure to extract the elastic backbone by means of the burning algorithm.46 To this end, in each image we identified the largest connected area and selected its two most distant points, P1 and P2, corresponding as closely as possible to the extremes of one of the image diagonals. The burning algorithm was then applied using P1 as the starting point and P2 as the endpoint, employing an 8-connected neighbourhood scheme to identify connected pixels.
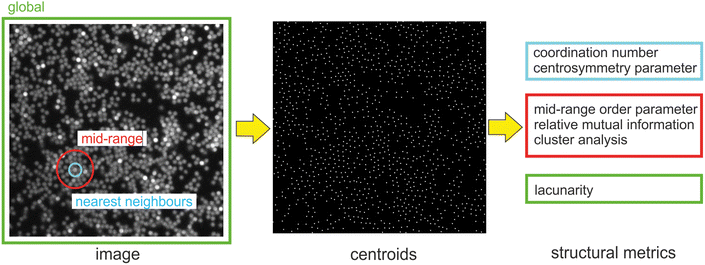
A list of the main symbols used throughout the manuscript is provided in Table 1. A sketch of the workflow adopted to characterize the patterns formed by the colloids is displayed in Fig. 1.
2 Results and discussion
2.1 Bacterial network
After sedimentation, Comamonas denitrificans formed a percolating network that spanned the entire field of view, composed of branched, interconnected structures distributed across the capillary floor, consistent with previous observations for C. denitrificans50 (Fig. 2a). Cells were visualised in a focal plane located 1.5 µm above the capillary floor, within a single optical section of thickness 2.85 µm; therefore, the observations reported here specifically probe the near-wall bacterial layer. The experiments were performed using stationary-phase cells, for which growth during the experimental time frame was negligible; therefore, the interactions within this network are governed by geometric confinement and the bacterial cell surface properties.We performed Fourier analysis on bacteria-only suspensions to access the characteristic length scales. Peaks in the power spectrum 〈I(r)〉 revealed a dominant spatial frequency corresponding to a characteristic length of approximately 7.5 ± 1 µm in the bacterial network (Fig. 2c). Visual inspection confirmed this length matched the width of bacterial branches and the gaps between them (Fig. 2a). Additionally, we investigated the presence of preferential directions by determining the orientation of the local gradient of the image intensity at each point, as detected by Sobel filtering.33 The resulting probability distribution (PDF) of orientations θ indicated no dominant direction (Fig. 2d). We thus conclude that the bacterial network is isotropic.
2.2 Mixed bacterial/colloidal suspensions
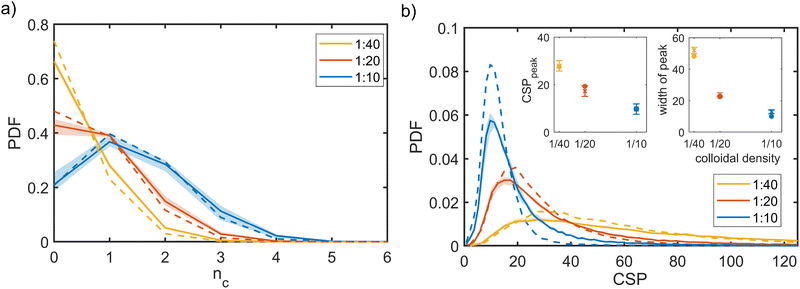
In the mixed suspensions, colloids were embedded within the bacterial network. Bacteria altered colloidal clustering, increasing heterogeneity in the resulting clusters (Fig. 3b–h) compared to the colloids-only suspensions (Fig. 3a–g). To quantify this effect, we performed a structural analysis of the colloidal aggregates in the mixed and colloids-only suspensions. Our analysis spanned multiple spatial scales: local (nearest neighbours), mid-range, and global (system-spanning). We used probability distribution functions (PDF) of different structural parameters.In mixed suspensions, the peaks of the CSP PDFs were shifted towards lower values compared to their colloids-only counterparts, indicating increased local symmetry, consistent with enhanced clustering (Fig. 4b). The shift, however, remained lower than 10%. Thus, bacteria did not significantly affect nearest-neighbour ordering.
![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) 2, computed for each particle based on the positions of its neighbours within a circular region of radius rm (see the Materials and Methods Section 1.8.2). Though this kind of parameter was originally defined in the context of equilibrium systems,53 it has subsequently been extensively employed with the name of ‘two-body excess entropy’ also for non-equilibrium systems,40,54,55 including 2D and quasi-2D particle systems.56,57 We note, however, that for such systems,
2, computed for each particle based on the positions of its neighbours within a circular region of radius rm (see the Materials and Methods Section 1.8.2). Though this kind of parameter was originally defined in the context of equilibrium systems,53 it has subsequently been extensively employed with the name of ‘two-body excess entropy’ also for non-equilibrium systems,40,54,55 including 2D and quasi-2D particle systems.56,57 We note, however, that for such systems, ![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) i2 should not be considered as thermodynamic entropy in the equilibrium sense. Instead, it is a structural metric reflecting ordering on the length-scale rm that has been proven particularly suitable to identify clustering and nucleated states.40 We calculated the probability distribution function of
i2 should not be considered as thermodynamic entropy in the equilibrium sense. Instead, it is a structural metric reflecting ordering on the length-scale rm that has been proven particularly suitable to identify clustering and nucleated states.40 We calculated the probability distribution function of ![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) 2 using rm equal to 4ar0 and 8ar0, corresponding to approximately two and four coordination shells, respectively (Fig. 5a and b).
2 using rm equal to 4ar0 and 8ar0, corresponding to approximately two and four coordination shells, respectively (Fig. 5a and b).
In each case, the ![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) 2 distributions were broader for mixed suspensions than for colloids-only suspensions (insets Fig. 5a and b), indicating increased structural heterogeneity in the presence of bacteria. For mid-range order parameters calculated with a cutoff rm = 4ar0, the peak of the distribution shifted to the left in mixed suspensions with middle (red curve) and low particle concentrations (yellow curve). This indicates that ordering on the length scale of the second neighbouring shell is more pronounced when bacteria are present. Conversely, for high colloidal concentrations (blue curve), the presence of bacteria did not affect the position of the peak. Interestingly, when the mid-range order parameter was calculated with a cutoff rm = 8ar0, the situation was reversed: for middle and low particle concentrations, the position of the peak was not influenced by the presence of bacteria, while for high particle concentrations the peak of the mixed suspensions had a shift towards the right with respect to their colloids-only counterpart. This suggests that at high colloidal concentrations, ordering at the scale of the fourth neighbour shell is more pronounced in the absence of bacteria.
2 distributions were broader for mixed suspensions than for colloids-only suspensions (insets Fig. 5a and b), indicating increased structural heterogeneity in the presence of bacteria. For mid-range order parameters calculated with a cutoff rm = 4ar0, the peak of the distribution shifted to the left in mixed suspensions with middle (red curve) and low particle concentrations (yellow curve). This indicates that ordering on the length scale of the second neighbouring shell is more pronounced when bacteria are present. Conversely, for high colloidal concentrations (blue curve), the presence of bacteria did not affect the position of the peak. Interestingly, when the mid-range order parameter was calculated with a cutoff rm = 8ar0, the situation was reversed: for middle and low particle concentrations, the position of the peak was not influenced by the presence of bacteria, while for high particle concentrations the peak of the mixed suspensions had a shift towards the right with respect to their colloids-only counterpart. This suggests that at high colloidal concentrations, ordering at the scale of the fourth neighbour shell is more pronounced in the absence of bacteria.
2.3 Ordering propagation
To gain further insights into the different behaviours of the examined systems on different length scales, we investigated how local nearest-neighbours ordering influenced mid-range ordering. We evaluated the relative mutual information M(NC, S)/I(Nc), between the coordination number and the average mid-range order parameter calculated for a given cutoff distance rm, where M(NC, S) denotes the mutual information and I(Nc) the self-information of the coordination number (Fig. 6). The relative mutual information M(NC, S)/I(Nc) can be interpreted as a correlation, namely a number between 0 and 1, quantifying how much information one has on the mid-range order parameter of a particle i, if its coordination number niC is known. The higher the relative mutual information, the stronger the correlation. Since![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) i2 quantifies ordering around a particle i over a distance rm, by varying rm we could establish the spatial influence of the coordination number (reflecting the nearest-neighbour arrangement) on the ordering degree over a length scale rm.
i2 quantifies ordering around a particle i over a distance rm, by varying rm we could establish the spatial influence of the coordination number (reflecting the nearest-neighbour arrangement) on the ordering degree over a length scale rm.
The relative mutual information trends were similar for mixed (solid lines) and colloids-only suspensions (dashed lines) for low and middle particle concentrations, but at high particle concentrations, the colloids-only suspension displayed a peak near 7 µm (corresponding to 3–4 neighbouring shells), absent in mixed suspensions. The presence of a peak at a given distance 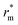 indicates a stronger interdependence between coordination number and mid-range order parameter at this distance. We interpret
indicates a stronger interdependence between coordination number and mid-range order parameter at this distance. We interpret 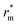 as a “propagation distance” of mid-range ordering. Conversely, no peak indicates a more heterogeneous, disordered structure. This was confirmed by the observation that, for high particle concentrations, on the length scale of two neighbouring shells (rm = 4ar0) the PDFs of mid-range order parameters had a peak at the same location regardless of the presence of bacteria (Fig. 5a), while on a length scale of four neighbouring shells (rm = 8ar0) colloids-only suspensions had a lower
as a “propagation distance” of mid-range ordering. Conversely, no peak indicates a more heterogeneous, disordered structure. This was confirmed by the observation that, for high particle concentrations, on the length scale of two neighbouring shells (rm = 4ar0) the PDFs of mid-range order parameters had a peak at the same location regardless of the presence of bacteria (Fig. 5a), while on a length scale of four neighbouring shells (rm = 8ar0) colloids-only suspensions had a lower ![[s with combining macron]](https://www.rsc.org/images/entities/i_char_0073_0304.gif) 2, i.e. a higher ordering degree (Fig. 5b).
2, i.e. a higher ordering degree (Fig. 5b).
We conclude that in colloidal-only suspensions, the local ordering of the nearest neighbours propagates up to 3–4 shells around a particle, whereas for mixed suspensions it stops after the first. We attribute this to the isotropic orientation of the bacterial network. Hence, it acts as a filter by mechanically obstructing the propagation of configurational ordering. Interestingly, the bacterial characteristic length of the bacterial network alone (∼7.5 ± 1 µm) was consistent with the suppression of mid-range order propagation beyond 3–4 shells in mixed suspensions.
2.4 Cluster analysis
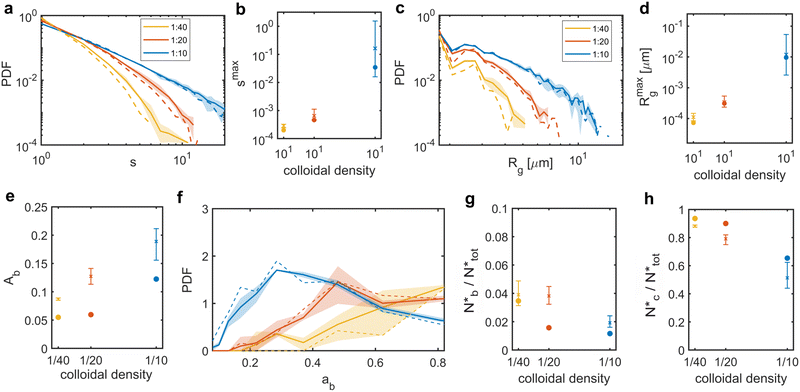
To investigate how systems that are indistinguishable at the nearest-neighbours level can differ at the mid-range, we analysed the distributions and structure of the colloidal clusters, i.e. agglomerates of colloids with an inter-particle distance within rm = 2ar0. Mixed suspensions at low and medium colloidal densities (red and yellow solid lines) contained more large clusters than their colloids-only counterparts, consistent with the observation that the presence of bacteria promotes colloidal clustering (Fig. 7a and c). However, at high colloidal densities, the differences between the distributions of the colloids-only and mixed suspensions were not significant. Similarly, for all colloidal densities, the maximum cluster size smax and the maximum gyration radius Rmaxg were not substantially affected by the presence of bacteria (see the insets in Fig. 7b and d).We further analysed the cluster structure by identifying repeating building blocks using a graph theory approach, where colloids are treated as nodes. Nodes with less than two neighbours were excluded. In particular, we investigated the topology of triangular ‘cliques’ i.e. triplets of adjacent colloids. The nodes were classified as ‘clique-like’ (belonging to triangular cliques, compact), or ‘branch-like’ (non-clique, open). The global airiness parameter Ab = Nb/N, where Nb and N are the number of branch-like nodes and the total number of connected nodes, respectively was 40–50% higher in the mixed suspensions, indicating a more open structure (Fig. 7e), consistent with the global lacunarity analysis (see Section 2.3). To examine the distribution of branch-like nodes, we distinguished three types of clusters: mixed (clique cores with branches), purely clique-like, and purely branch-like. Their structure was quantified using an airiness parameter ab = sb/s, where sb is the number of branch-like nodes in the cluster (Fig. 7f). The distribution of the airiness parameter showed little difference between systems at low and medium densities, but at high density, the mixed suspensions contained fewer clusters with low ab. In contrast, purely clique-like and purely branch-like clusters differed between the colloids-only and the mixed suspensions. Bacteria increased the fraction of branch-like clusters 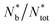 while decreasing clique-like clusters
while decreasing clique-like clusters 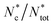 across all densities (Fig. 7g and h). We conclude that differences in global airiness between systems with and without bacteria mainly arise from shifts in the numbers of purely branch-like and purely clique-like clusters, while mixed clusters remain largely unchanged.
across all densities (Fig. 7g and h). We conclude that differences in global airiness between systems with and without bacteria mainly arise from shifts in the numbers of purely branch-like and purely clique-like clusters, while mixed clusters remain largely unchanged.
2.5 Lacunarity
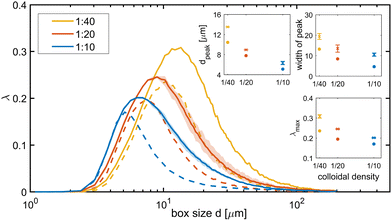
We assessed ordering on a global level and the emergence of characteristic length scales by calculating the lacunarity λ of the colloidal pattern, for different system-spanning tesselations with square tiles (‘boxes’) of side d.58 The lacunarity measures the heterogeneity and gappiness of a pattern on the scale d, with higher λ values indicating a higher heterogeneity on the considered length scale.59,60 Mixed suspensions had higher (∼30%) and wider (∼30–50%) lacunarity peaks, shifted towards larger values of d (∼25–40%), compared to colloids-only suspensions (Fig. 8). These changes reflect greater heterogeneity, a broader gap-size distribution, and the presence of larger gaps in the mixed suspensions. This is consistent with the clique analysis, which showed that the bacteria promote a branch-like airier colloidal packing.2.6 Elastic backbone
Lastly, we evaluated the effect of the colloids on the bacterial network by comparing the percolating structures in bacteria-only and mixed suspensions. We assessed the length of the elastic backbone of the percolating structures, defined as the union of all the shortest paths connecting two representative points P1 and P2,46 and we further normalised it by the distance between two such points. The elastic backbone represents the stress-bearing structure of the network.61 Mixed suspensions exhibited longer elastic backbones than bacteria-only suspensions, and backbone length increased with colloidal density (Fig. 9a–c). As the length of the elastic backbone is related to the resistance to stresses, we speculate that the presence of particles reinforces the bacterial network, making it more resistant to disruption.3. Conclusions
In this work, we studied how the presence of sedimenting colloids influences the near-wall layer structure of the percolating network formed by bacteria during sedimentation and how the network, in turn, influences colloidal aggregation. We find that the presence of bacteria favours colloidal clustering. The microscopic origin of the observation was investigated by characterizing the structures emerging in mixed and colloids-only suspensions at different length scales (local, mid-range, global). At the local level, the colloidal arrangement is similar with and without bacteria, as shown by the coordination number and the centrosymmetry parameter. However, at the mid-range scale, we observed enhanced clustering in the presence of bacteria, as evidenced by the increased number of larger clusters in the mixed suspensions. Such clusters were more branch-like and less compact than those in colloids-only suspensions. These observations explain why at the global scale the presence of bacteria induces a greater heterogeneity and gappiness in the colloidal surface-layer pattern, as shown by higher lacunarity values. We attribute this to the randomly orientated bacterial network acting as a barrier to local ordering propagation. This suppression of ordering propagation is not specific to the biological nature of the bacteria, but arises from the presence of a disordered, percolating fibrous scaffold that geometrically constrains colloidal rearrangements. We therefore expect similar behaviour for non-biological rod-like or fibrous networks in the isotropic regime, while different responses may emerge if the scaffold develops orientational (e.g. nematic) order. The conclusion was supported by analysis of the mid-range order parameter local entropy, which quantifies the mid-range ordering. We found that in purely colloidal suspensions ordering was stronger and propagated further (up to 3–4 particle shells) than in mixed suspensions.Finally, we assessed the effect of the colloids on the bacterial near-wall layer network by comparing mixed colloids-bacteria suspensions to the bacteria-only suspensions. In particular, we evaluated the length of the elastic backbone, the load-bearing structure of a percolating system. We found that the presence of colloids resulted in longer elastic backbones, possibly indicating a structure with greater stress resistance, than in the bacteria-only case. Further investigation is needed to confirm this hypothesis, e.g. by evaluating the rheological properties of such suspensions.
While previous efforts to investigate and control near-wall colloidal patterning have typically relied on synthetic depletants or engineered surface geometries, our study is the first to demonstrate that living biological matter can act as a self-assembling scaffold to drive these near-wall microstructural transitions.62,63 Overall, our study reveals that the presence of sedimenting non-motile bacteria with colloidal particles gives rise to collective structures distinct from those in purely colloidal and purely bacterial suspensions. These findings provide a basis for using bacterial scaffolds to modulate colloidal assembly and offer a framework for studying particle–network interactions across scales. While primarily fundamental, this approach may also help to rationalise unwanted bacterial aggregation in environmental systems, including those influenced by microplastic contaminants.
Author contributions
LS and ES developed the original idea. SC, ES and LS designed the experiments. SC and LS performed the experiments. LS and SC performed the data analysis. LS wrote the first draft of the manuscript, and all authors contributed to subsequent versions.Conflicts of interest
There are no conflicts to declare.Data availability
All codes used for the analysis are available from the authors upon reasonable request. The centroid position data used in this study are provided as supplementary information (SI) in the form of two.xyz files corresponding to high-density suspensions with and without bacteria, which allow reproduction of the structural analyses shown in Fig. 3–8. See DOI: https://doi.org/10.1039/d5sm01062d.Acknowledgements
The authors acknowledge financial support from SNSF PRIMA Grant No. 179834 (to E.S.), and Marie Skłodowska-Curie Actions Individual Fellowship Grant No. 101033169 (to S.C.).Notes and references
- A. Mandal, A. Dutta, R. Das and J. Mukherjee, Marine Poll. Bull., 2021, 170, 112626 CrossRef CAS PubMed.
- L. Krause, D. Biesgen, A. Treder, S. A. Schweizer, E. Klumpp, C. Knief and N. Siebers, Geoderma, 2019, 351, 250–260 CrossRef CAS.
- L. Zhang, D. Tu, X. Li, W. Lu and J. Li, BMC Microbiol., 2020, 20, 254 CrossRef CAS PubMed.
- A. Roberto, J. Van Gray and L. Leff, Water Res., 2018, 134, 353–369 CrossRef CAS PubMed.
- C. Valeriani, M. Li, J. Novosel, J. Arlt and D. Marenduzzo, Soft Matter, 2011, 7, 5228–5238 RSC.
- J. Singh, A. E. Patteson, B. O. Torres Maldonado, P. K. Purohit and P. E. Arratia, Soft Matter, 2021, 17, 4151–4160 RSC.
- B. O. Torres Maldonado, R. Ran, K. L. Galloway, Q. Brosseau, S. Pradeep and P. E. Arratia, Phys. Fluids, 2022, 34, 113305 CrossRef CAS.
- T. Yao, V. Kos, Q. Zhang, Y. Luo, E. Steager, M. Ravnik and K. Stebe, Sci. Adv., 2022, 8, eabn8176 CrossRef PubMed.
- S. Kamdar, S. Shin, P. Leishangthem, L. Francis, X. Xu and X. Cheng, Nature, 2022, 603, 819–823 CrossRef CAS PubMed.
- L. Vaccari, M. Molaei, T. H. Niepa, D. Lee, R. L. Leheny and K. J. Stebe, Adv. Colloid Interface Sci., 2017, 247, 561–572 CrossRef CAS PubMed.
- H. Firoozmand and D. Rousseau, Food Hydrocolloids, 2014, 42, 204–214 CrossRef CAS.
- H. Firoozmand and D. Rousseau, Food Res. Int., 2016, 81, 66–73 CrossRef CAS.
- S. Charlton, G. Melaugh, D. Marenduzzo, C. MacPhee and E. Secchi, Phys. Rev. E, 2025, 111, 024410 CrossRef CAS PubMed.
- M. Bantawa, B. Keshavarz, M. Geri, B. Mehdi, D. Thibaut, G. H. McKinley and E. Del Gado, Nat. Phys., 2023, 19, 1178–1184 Search PubMed.
- M. Nabizadeha, F. Nasirianb, X. Li, Y. Saraswatd, R. Waheibid, L. C. Hsiaod, D. Bi, B. Ravandi and S. Jamali, Proc. Natl. Acad. Sci. U. S. A., 2024, 121, e2316394121 CrossRef PubMed.
- G. Tarjus, S. Kivelson, Z. Nussinov and P. Viot, J. Phys.: Condens. Matter, 2005, 17, R1143–R1182 CrossRef CAS.
- C. P. Royall, S. R. Williams, T. Ohtsuka and H. Tanaka, Nat. Mater., 2008, 7, 556–561 CrossRef CAS PubMed.
- B. de Nijs, S. Dussi, F. Smallenburg, J. Meeldijk, D. Groenendijk, L. Filion, A. Imhof, A. van Blaaderen and M. Dijkstra, Nat. Mater., 2015, 14, 56–60 CrossRef CAS PubMed.
- D. Frenkel, Phys. A, 1999, 263, 26–38 CrossRef CAS.
- O. Velev, A. Lenhoff and E. Kaler, Science, 2000, 287, 2240–2243 CrossRef CAS PubMed.
- V. Manoharan, M. Elsesser and D. Pine, Science, 2003, 301, 483–487 CrossRef CAS PubMed.
- F. Bai, D. Wang, Z. Huo, W. Chen, L. Liu, X. Liang, C. Chen, X. Wang, Q. Peng and Y. Li, Angew. Chem., 2007, 46, 6650–6653 CrossRef CAS PubMed.
- S. Wintzheimer, T. Granath, M. Oppmann, T. Kister, T. Thai, T. Kraus, N. Vogel and K. Mandel, ACS Nano, 2018, 12, 5093–5120 CrossRef CAS PubMed.
- T. Cover and J. Thomas, Elements of information theory, John Wiley & Sons, New Jersey, 2005 Search PubMed.
- C. Bron and J. Kerbosch, Commun. ACM, 1973, 16, 575–577 CrossRef.
- J. E. Hallet, F. Turci and P. C. Royall, Nat. Commun., 2018, 9, 1–10 Search PubMed.
- A. E. González, J. Phys.: Condens. Matter, 2002, 14, 2335–2345 CrossRef.
- L. J. Teece, M. A. Faersb and P. Bartlett, Soft Matter, 2011, 7, 1341–1351 RSC.
- W. K. Pratt, Digital Image Processing: PIKS Scientific Inside, John Wiley & Sons Inc, New York, 2007 Search PubMed.
- B. H. Glünder, https://www.gluender.de/Miscellanea/MiscTexts/MiscExcerpts/ReadMe_W.html, 2020.
- P. Carl, https://questpharma.u-strasbg.fr/html/radial-profile-ext.html, 2020.
- L. Testard, V. Berthier and K. Walter, Phys. Rev. Lett., 2011, 106, 125702 CrossRef PubMed.
- R. C. Gonzalez and R. E. Woods, Digital image processing, Pearson, New York, 2014, pp. 186–188 Search PubMed.
- J.-Y. Tinivez, J. Schindelin, J. Eglinger and M. Hiner, Directionality plugin for ImageJ/Fiji, software, https://imagej.net/plugins/directionality (accessed 2018).
- J.-Y. Tinevez, N. Perry, J. Schindelin, G. Hoopes, G. Reynolds, E. Laplantine, S. Bednarek, S. Shorte and K. Eliceiri, Methods, 2017, 115, 80–90 CrossRef CAS PubMed.
- C. K. Kelchner, S. J. Plimpton and J. C. Hamilton, Phys. Rev. B: Condens. Matter Mater. Phys., 1998, 58, 11085–11088 CrossRef CAS.
- P. M. Piaggi and M. Parrinello, J. Chem. Phys., 2017, 147, 114112 CrossRef PubMed.
- H. Tanaka, T. Kawasaki, H. Shintani and K. Watanabe, Nat. Mater., 2010, 9, 324–331 CrossRef CAS PubMed.
- M. Leocmach, J. Russo and H. Tanaka, J. Chem. Phys., 2013, 138, 12A536 CrossRef PubMed.
- P. Piaggi, O. Valsson and M. Parrinello, Phys. Rev. Lett., 2017, 119, 015701 CrossRef PubMed.
- R. L. Jack, A. J. Dunleavy and C. P. Royall, Phys. Rev. Lett., 2014, 113, 095703 CrossRef CAS PubMed.
- E. Stanifer and M. L. Manning, Soft Matter, 2022, 12, 2394–2406 RSC.
- S. R. Sternberg, Computer, 1983, 16, 22–34 Search PubMed.
- E. Tomita, A. Tanaka and H. Takahashi, Theor. Comput. Sci., 2006, 363, 28–42 CrossRef.
- A. A. Hagberg, D. A. Schult and P. J. Swart, Proceedings of the 7th Python in Science Conference, Pasadena, CA USA, 2008, pp. 11–15 Search PubMed.
- H. J. Hermann, D. C. Hong and H. E. Stanley, J. Phys. A: Math. Gen., 1984, 17, L261–L266 CrossRef.
- K. Zuidervel, Graphics gems IV, Academic Press Professional, San Diego, 1994, pp. 474–485 Search PubMed.
- T. Y. Zhang and C. Y. Suen, Commun. ACM, 1984, 27, 235–239 CrossRef.
- D. Legland, I. Arganda-Carreras and P. Andrey, Bioinformatics, 2016, 32, 3532–3534 CrossRef CAS PubMed.
- S. Charlton1, G. Melaugh, D. Marenduzzo, C. MacPhee and E. Secchi, Phys. Rev. E, 2025, 024410, 024410 CrossRef PubMed.
- W. C. K. Poon, J. Phys.: Condens. Matter, 2002, 14, 7581–7597 CrossRef.
- W. K. Kegel and A. van Blaaderen, Science, 2000, 287, 290–293 CrossRef CAS PubMed.
- J. C. Dyre, J. Chem. Phys., 2018, 149, 210901 CrossRef PubMed.
- T. S. Ingebrigtsen and H. Tanaka, Proc. Natl. Acad. Sci. U. S. A., 2018, 115, 87–92 CrossRef CAS PubMed.
- S. A. Ghaffarizadeh and G. J. Wang, J. Phys. Chem. Lett., 2022, 13, 4949–4954 CrossRef CAS PubMed.
- Z. Wang, W. Qi, Y. Peng, A. M. Alsayed, Y. Chen, P. Tong and Y. Han, J. Chem. Phys., 2011, 134, 034506 CrossRef PubMed.
- X. Ma, J. Liu, Y. Zhang, P. Habdas and A. G. Yodh, J. Chem. Phys., 2019, 150, 144907 CrossRef PubMed.
- A. Alicke, L. Stricker and J. Vermant, J. Colloid Interface Sci., 2023, 652, 317–328 CrossRef CAS PubMed.
- P. Meakin, Fractals, scaling and growth far from equilibrium, Cambridge University Press, Cambridge, 2017 Search PubMed.
- T. Smith, G. Lange and W. Marks, J. Neurosci. Methods, 1996, 69, 123–136 CrossRef PubMed.
- C. I. N. Sampaio Filho, J. S. J. Andrade, H. J. Hermann and A. A. Moreira, Phys. Rev. Lett., 2018, 20, 175701 CrossRef PubMed.
- N. Nakamura, Y. Sakamoto and H. Ogi, Sci. Rep., 2021, 11, 8929 CrossRef CAS PubMed.
- B. Sprinkle, S. Wilken, S. Karapetyan, M. Tanaka, Z. Chen, J. R. Cruise, B. Delmotte, M. M. Driscoll, P. Chaikin and A. Donev, Phys. Rev. Fluids, 2021, 6, 034202 CrossRef.
Footnote |
| † These authors contributed equally to the present work. |
| This journal is © The Royal Society of Chemistry 2026 |