 Open Access Article
Open Access ArticleViscoelastic properties of tumor spheroids revealed by a microfluidic compression device and a modified power law model
Mrinal
Pandey
 a,
Bangguo
Zhu
a,
Bangguo
Zhu
 c,
Kaitlyn
Roach
c,
Kaitlyn
Roach
 b,
Young Joon
Suh
b,
Young Joon
Suh
 a,
Jeffrey E.
Segall
d,
Chung-Yuen
Hui
a,
Jeffrey E.
Segall
d,
Chung-Yuen
Hui
 c and
Mingming
Wu
c and
Mingming
Wu
 *a
*a
aBiological and Environmental Engineering Department, Cornell University, Ithaca, NY, USA. E-mail: mw272@cornell.edu
bMeinig School of Biomedical Engineering, Cornell University, Ithaca, NY, USA
cSibley School of Mechanical and Aerospace Engineering, Cornell University, Ithaca, NY, USA
dDepartment of Pathology, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA
First published on 5th February 2026
Abstract
Clinically, palpation is one of the important diagnostic methods to assess tumor malignancy. In laboratory research, it is well accepted that the bulk stiffness of the tumor and the surrounding tissue is closely correlated with the malignant state of the tumor. Here, we postulate that, in addition to tumor stiffness, tumor viscoelasticity – the fact that tumor tissue takes time to bounce back after compression, can also be used to evaluate the tumor malignancy state. In this work, we characterized the viscoelastic properties of tumor spheroids using a recently developed microfluidic compression device by quantifying their relaxation dynamics upon load removal. Tumor spheroids were made using breast tumor cells spanning various malignancy levels; non-tumorigenic epithelial (MCF10A), moderately malignant tumor (MCF7) and triple negative metastatic tumor (MDA-MB-231) cell line. Spheroids embedded within a 3D extracellular matrix were periodically compressed, and their strain responses were recorded using microscopic imaging. Our results revealed that the measured strain relaxation dynamics can be successfully described by a modified power law model, demonstrated that non-tumorigenic tumor spheroids were more elastic, exhibited shorter relaxation time and less plasticity than those of tumorigenic spheroids. This work highlights that viscoelastic properties in addition to bulk stiffness of the tumor spheroids can serve as a complementary mechanical biomarker of tumor malignancy and demonstrate the validity of a modified power law model for the mechanical characterization of a living tissue.
Introduction
Tumor mechanics emerges as a key biomarker of tumor malignancy state, with extensive studies demonstrating that increased tissue stiffness promotes tumor progression.1–3 This elevated stiffness arises from the uncontrolled proliferation of tumor cells4 and the stiffening of the surrounding extracellular matrix.5,6 Paradoxically, multiple studies have shown that, at the single cell level, malignant cells are often softer than their non-malignant counterpart.7–11 We know that tumors, or living tissues in general, are inherently complex and highly adaptive, capable of dynamically remodelling the mechanical properties of their surrounding matrices in response to changes in the microenvironment.12,13 Tumor cells/tissues are nonlinear elastic, and are very sensitive to the surrounding mechanical environment.14 As a result, the reported values for tumor cells/tissues cover a wide range.9–11,15 It is thus important to develop strategies to investigate tumor mechanics using physiologically realistic experimental conditions and at multiple time and length scales.Tumor mechanics studies today have largely focused on the bulk stiffness of the tumor cell/tissue or surrounding ECMs. It is only recently that the viscoelastic properties of ECM, instead of stiffness alone, have been found to play important roles in cell mechano-transduction16–18 and tumor invasion.19–22 This is not a surprise since tumor invasion is a dynamic process.12 In a landmark study, Chaudhuri et al.21 highlighted the importance of examining viscoelastic properties of extracellular matrices, emphasizing their critical roles in regulating cell migration behavior. In contrast, to date, very little is learned about the viscoelasticity of tumor tissue, and how it correlates with tumor malignant state.23 Examining the viscoelastic properties of tumors is also crucial from a basic science viewpoint. We ask the basic question: how can we theoretically model the mechanics of living materials? The simplest class of solid materials is the Hookean solid where the stress is proportional to the strain. In contrast, biological materials are complex in their behavior; they often deform and adapt when subject to external stresses. Type I collagen, a primary protein that provides architectural support to mammalian cells, is an example.21,24 Collagen consists of a cross-linked fibre network that aligns when stretched and buckles when compressed. This leads to the complex nonlinear elastic and anisotropic material properties of collagen.25,26 Living materials such as tissues (e.g. cell embedded collagen) enhance the complexity of the materials further. Here, the basic constituents, cells, can sense, move, as well as modify the extracellular matrix (ECM) architecture. For example, tumor cells are mechano-sensors; they move and re-arrange the surrounding ECM spatially when a tissue is subjected to an external force.27 In this context, it adds one more dimension to the material mechanics.
Beyond stiffness and viscoelasticity, cells and tissues also undergo irreversible plastic deformation under load. These plastic deformations are emerging as another important biomarker for cancer invasiveness, and single-cell studies have begun to quantify this plastic component of deformation, showing that cells can undergo permanent shape changes after large or repeated mechanical perturbations.28 Plasticity of the tissue remains to be explored in detail.
Commercially available parallel rheometers are the current workhorse for characterizing the mechanics of biological materials today. In parallel rheometers, biological samples are sheared or compressed between two parallel plates, and a stress–strain relationship is obtained. While straightforward, this setup is not easily compatible with living tissues or optical imaging, limiting its use for living materials. AFM addressed some of these issues by enabling force measurements in 3D environments,29–31 and optical compatibility but its fine tip restricts analysis to localized single-cell mechanics. Micro-pipette aspiration32–34 has been used to contribute to valuable insights of tissue mechanics although the data analysis and experimentation are challenging.
Microfluidic platforms have emerged as promising tools for studying cell and tissue mechanics in biologically relevant settings, with better visualization and fine control over bulk cell and tissue mechanics. Examples include PDMS micro-piston systems,35–37 rheo-microscopy based tools,38–40 and constriction based devices that probe mechanical resistance by forcing aggregates through narrow channels.19 Despite their innovations, each of these methods has inherent limitations in terms of throughput, physiological relevance, or spatial resolution and optical compatibility. To complement these approaches, in this manuscript we use an integrated method that combines a microfluidic compression device, which provides a physiologically realistic 3D environment for tumor spheroids, with a modified power-law model to investigate their viscoelastic mechanical properties (and associated plastic deformations) over time. The advantage of our device is the ability to embed the tumor tissue within a 3D extracellular matrix providing a physiologically realistic environment for the tumor spheroids.
Results and discussion
Microfluidic compression device setup and calibration
In this work, we use a previously developed micro-rheometer in our lab,41 which is a three-layer PDMS based microfluidic device. The key capability of the microfluidic device is to provide a well-controlled compression to the tumor spheroids embedded in a physiologically realistic 3D ECM and compatible with microscopic imaging (Fig. 1). Briefly, the device consists of 12 functional units, of which 6 are control units and the other 6 are compression units (Fig. 1A). Each functional unit is built from three PDMS layers bonded together. The cross-sectional view of each unit is shown in Fig. 1B. The main features of the compression unit are the cell chamber layer (L1) that holds the spheroid embedded ECMs and the pressure control unit (L2 + L3) that provides well defined compression to the spheroid. The layer 2 is a thin flexible PDMS membrane with a circular piston. The layer 3 contains pressure chambers and a microchannel. This microchannel connects the pressure chambers of the six compression units (not the control units) and is connected to an external pressure source. In the compression units, when pressure is applied, the piston layer L2 bends downward, compressing the spheroid in the well beneath it (see Fig. 1B, and Movie S5). To calibrate piston displacement under varying pressure levels, we measured piston displacement by locating the focal point of 1 µm green, fluorescent beads on the piston surface as a function of the pressure applied (Fig. S1A). Briefly, beads were placed at the bottom of the piston and upon applying pressure, piston displacement caused beads to shift vertically and defocus. The displacements were measured by refocusing the beads using a motorized stage.42 The piston displacement versus pressure curve was validated against the COMSOL computation as shown in Fig. S1A, where the red line is the result of COMSOL simulation and dots are experimental measurements. The actual device geometry (Fig. S2A) is chosen for COMSOL simulation. The simulation was performed using an axisymmetric model and the PDMS was modelled as a linear elastic material with modulus of 1.33 MPa and Poisson's ratio of 0.49. In the simulation, the bottom glass was modelled as a rigid material with fixed constraints, the sides of the device were fixed, and the pressure was applied to the top surface Fig. S2B and Movie S5. An extra fine mesh was applied across the model to ensure numerical accuracy.For a typical experiment, the cell chamber layer (L1) was first placed on a standard glass slide (75 mm × 25 mm × 1 mm) facilitating microscopic imaging of the embedded tumor spheroids. Each cell chamber is 3 mm in diameter, and 500 µm in depth. The rim of each cell chamber contains 4 slits, each 0.5 mm wide, to allow media transport during compression. For more details on device design and development please refer to.43 Second, tumor spheroids were embedded in 1.5 mg mL−1 type I collagen at a concentration of 1296 spheroids per mL and were introduced into the cell chamber. Note that on average 2–3 collagen spheroids were embedded per cell chamber to avoid potential spheroid–spheroid interaction during the experimental run. In our current work we use a 1.5 mg mL−1 collagen matrix to provide anchorage and to prevent bulk movement of spheroids during cyclic compression. A 1.5 mg mL−1 collagen is a soft matrix that minimally interferes with the measurement of the mechanical properties of the spheroids and provides more stability and repeatability of experiments. Third, the pressure control unit (L2 + L3) was carefully aligned and placed atop L1, ensuring that each piston was centered precisely over the corresponding cell chamber. This alignment is important for uniform compressive forces across each individual spheroid. Lastly, the mid-z plane of the spheroid is imaged using a bright field microscope. A convolutional neural network, U-Net44 segmentation algorithm, was used to trace the spheroid boundaries. The bright yellow boundaries are obtained by using U-Net algorithm (see Fig. 1C–E). The top row (Fig. 1C–E) shows the MDA-MB-231, MCF7 and MCF10A spheroids before any load was applied on them. The bottom panel (Fig. 1F–H) shows the same spheroids after compressive load was applied on them. The red/green outlines in Fig. S1B are also traced spheroid outlines with/without compression respectively. These outlines of the spheroids are used to compute the spheroid area A in the mid-z plane, and the spheroid diameter is computed using  . We see that the spheroid increases in its diameter upon compression (Fig. S1C). Uniform spheroids were generated using a standardized agarose microwell array technique previously developed in our labs.45 Fig. S1D shows that there are no significant differences in initial spheroid sizes among the different cell lines, ensuring a fair basis for comparative mechanical testing.
. We see that the spheroid increases in its diameter upon compression (Fig. S1C). Uniform spheroids were generated using a standardized agarose microwell array technique previously developed in our labs.45 Fig. S1D shows that there are no significant differences in initial spheroid sizes among the different cell lines, ensuring a fair basis for comparative mechanical testing.
Spheroid size response under cyclic compression exhibits viscoelastic material behavior
To study the viscoelastic properties of tumor spheroids, we observed the tumor spheroid size response under a controlled dynamic compression (Fig. 2). A square pressure wave with a max pressure of 14 kPa and a period of 40 seconds (20 s compression followed by 20 s relaxation) is applied to the spheroid through the pressure control unit (Fig. 2A, and Movies S1 and S2). The spheroid is imaged at mid-z plane optically, and its size responses are shown in Fig. 2B. Using the spheroid images, we calculated the normalized area change of the spheroid, (A − A0)/A0, where A is the spheroid area in the mid-z plane at any time point, and A0 is the initial spheroid area before compression. Upon compression, the spheroid area increases rapidly and reaches a near stable value. Once the load is removed (20 s relaxation phase), the spheroid area decreases and moves towards its initial area. After load removal, a rapid initial area recovery is observed, followed by a progressively slower recovery (see Fig. 2B). A similar pattern of area change is observed for next two cycles of compression (and relaxation). This rapid area relaxation (elastic response) followed by a gradual area relaxation over time (viscous response) was observed in all cell types.We computed strain response curves using the data in Fig. 2B. First, we converted the normalized area change into height change (Δh/h) using a geometric relationship assuming that the spheroid is incompressible at the time scale of our interest and a neo-Hookean material for the spheroid. The detailed computation is in Supplementary theoretical model Section 3 and Fig. S5 and S6. Δh/h here is defined as the linear spheroid strain in our measurements.
To evaluate the viscoelasticity of the tumor spheroid, we focus on the relaxation phase of the strain response curve, more specifically, the first cycle of strain relaxation (see the dashed box in Fig. 2B). An important feature of a linear viscoelastic material with a single time scale is that the strain response to a sudden release of a constant compression decays exponentially.46,47 Here, we found that the relaxation response of tumor spheroids exhibited a clear power-law relationship when plotted on a log–log scale (see Fig. 2C–E and S3) across all three cell lines. This indicated that multi-time scales are involved in the relaxation dynamics of tumor spheroids. To estimate the timeit takes for spheroids to bounce back after compression, we analysed the absolute slope values of the relaxation curves across all spheroid types and found that the non-tumorigenic MCF10A spheroids exhibited the fastest recovery (see Fig. 2F) while the metastatic MDA-MB-231 spheroids were the slowest, and MCF7 spheroids in the middle. Interestingly, while the recovery slopes of MCF7 and MCF10A spheroids were not significantly different, the slopes of MDA-MB-231 and MCF10A spheroids differed significantly. The faster recovery of MCF10A spheroids indicates an overall more elastic behavior, whereas the slower recovery of malignant MDA-MB-231 spheroids is indicative of a more fluid like behavior.
Overall, the differential strain relaxation curves for MDA-MB-231 spheroids versus MCF10A spheroids may come from the differential cell–cell adhesion of these two types of spheroids. In MCF10A spheroids, cells adhere to each other via a direct cell–cell adhesion molecule, E-cadherin, while the MDA-MB-231 cells adhere to each other through a secondary adhesion, cell-ECM, via β1-integrin.48,49 The tightly cohesive cell–cell adhesion in MCF10A spheroids may lead to a more solid like behavior than the MDA-MB-231 cell spheroids where cell–cell adhesion is weak. This is consistent with the fast bounce back time for MCF10A spheroids. In addition, single cell stiffness of MCF10A is stiffer than MDA-MB-231 cells, which could also lead to a high elasticity which may contribute to the faster rebound time seen in Fig. 2F. Our observations are also consistent with a previous study19 reporting that non-tumorigenic MCF10A spheroids recover fastest, indicative of higher elasticity, and attributing this behavior to a well-defined cortical actin rim. Within the context of our power-law rheological model, we therefore speculate that a more cohesive cortical actin cortex in MCF10A (and MCF7) supports faster and more reversible stress relaxation, whereas the more diffuse actin organization in MDA-MB-231 is associated with slower relaxation. In future work we plan to directly investigate the role of the actin cortex in governing the viscoelastic response of tumor spheroids.
Importantly, all spheroid types, regardless of malignancy status, exhibited power-law relaxation behavior, reinforcing that living biological materials display complex rheological behavior50 rather than simple linear viscoelasticity. This characteristic may reflect dynamic cytoskeletal remodelling, cell-ECM interactions, and fluid redistribution occurring over a broad range of timescales. More studies matching the timescales (in minutes to hours) of these cellular phenomena can give deeper insight into cellular mechanics and invasion characteristics.
Viscoelastic properties of tumor spheroids revealed by a modified power law model
The power law behavior observed in the spheroid size relaxation curves (Fig. 2C–E) suggest that the material model for tumor spheroids requires a multi-time scale model beyond the simple Maxwell or Kelvin–Voigt models. Here, we propose a modified power law model that has been successful in describing viscoelastic properties of soft materials previously.51,52 This model, originally described in ref. 53 has a stress relaxation function of: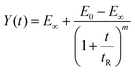 | (1) |
 | ||
| Fig. 3 Viscoelastic properties of tumor spheroids revealed by a modified power law model. (A) The experimental strain response curve for malignant MDA-MB-231, MCF7 and non-tumorigenic MCF10A spheroids. Circles are from experiments, and the lines are fits to the modified power law model. Here the strain is defined as Δh/h, where h is the vertical height of the spheroid, and is computed using the area change data shown in Fig. 2B. (B) Stress relaxation curve computed using the strain response curve in (A) and the modified power-law model for all three types of spheroids. Dashed lines indicate the location of half time when the stress decreases to half its original value. The fit data shown in 3A and the stress relaxation curve in 3B is from a single spheroid. (C) Half relaxation time for malignant for spheroids of three types. The average value for MDA-MB-231 spheroids is 5.571 ± 1.34 s, MCF7 is 3.63 ± 0.633 s and MCF10A is 1.488 ± 0.488 s. (D) Computed effective viscosity of the spheroids as a function of frequency. Each dotted line is data from one spheroid. A total of 18 spheroids, 6 for each cell type were analysed. The stars were obtained using a nonparametric t-test (Mann–Whitney test with **: P < 0.01). | ||
Traditional spring-dashpot rheology models like Maxwell, Kelvin–Voigt and standard linear solids capture relaxation through discrete timescale. However, many soft and living materials exhibit a continuous distribution of relaxation times. Thus, viscoelastic models with broad relaxation spectra, including power-law, modified power-law, and fractional viscoelastic models, have been widely used to describe soft biological materials. Fractional viscoelastic models provide a compact framework for such responses,51,53 and recent studies demonstrate that even single cells can be accurately modelled using fractional Kelvin–Voigt elements.55 The modified power-law model we adopt here is mathematically similar to fractional models in the low-frequency regime and shares their ability to represent broad relaxation spectra while remaining simple, with fewer parameters and an intuitive analytical form, compared to the non-trivial fractional operators in frictional models.51,53 Because our data consist solely of strain versus time responses without direct force measurement, we focus on extracting the relaxation time tR and power-law exponent m, which are robustly identifiable from our experiments. Future work integrating calibrated force measurement, as in Ferraro56et al. or our device,41 will enable determination of the modulus E0 and E∞ and separation of normalized effective viscosity, introduced later, into its dimensional components. To characterize the stress response of the tissue, we computed stress relaxation curves using the fitted parameter m and tR and eqn (1) for the three spheroid types (see Fig. 3B). Here, the half time for stress to relax to its final value is computed. We find that the non-tumorigenic tumor spheroid (MCF10A) relaxes much faster than the malignant MDA-MB-231 spheroids, an average value of 1.27 ± 0.57 s versus 5.57 ± 1.34 s (Fig. 3C). Our results reveal clear differences between malignant and non-malignant spheroids in its half time, which is consistent with our estimate about the bouncing back time in strain (Fig. 2F). This provides a foundation to distinguish differential malignant states of the spheroids based on viscoelastic properties rather than relying solely on elastic measurements alone.
To further quantify the frequency-dependent dissipation, we calculated the normalized effective viscosity η(ω)/E0, derived from the imaginary component of the complex modulus associated with our stress relaxation function and divided by the instantaneous modulus (see Theoretical models’ Section 2 for more details and ref. 57).
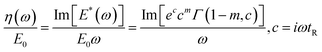 | (2) |
Interestingly, the effective viscosity profiles of the malignant (MDA-MB-231, MCF7) and non-tumorigenic (MCF10A) spheroids revealed a frequency dependent behavior and a crossover at log frequency of −1 corresponding to period of ∼10 s (Fig. 3D). At lower frequencies within our computed window (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) f ≈ −1 to −3.8, corresponding to timescales of ∼10 s–100 min), MDA-MB-231 spheroids exhibited higher effective viscosity than MCF10A, indicating a more viscous-like mechanical behavior in malignant spheroids. This trend reversed at higher frequencies (log
f ≈ −1 to −3.8, corresponding to timescales of ∼10 s–100 min), MDA-MB-231 spheroids exhibited higher effective viscosity than MCF10A, indicating a more viscous-like mechanical behavior in malignant spheroids. This trend reversed at higher frequencies (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) f ≈ −0.9–1.2, corresponding to timescales of ∼8 s to −0.06 s), where malignant spheroids appeared less viscous than their non-tumorigenic counterparts. The observed frequency dependence underscores that the apparent mechanical properties of tumors depend on probing frequency. Thus, a comprehensive evaluation of tumor mechanics requires measurements across a broader frequency spectrum. In addition to these two cell lines, we also looked at MCF 7 cells and consistent to half relaxation data, effective viscosities of MCF 7 were in between MDA-MB-231 and MCF10A.
f ≈ −0.9–1.2, corresponding to timescales of ∼8 s to −0.06 s), where malignant spheroids appeared less viscous than their non-tumorigenic counterparts. The observed frequency dependence underscores that the apparent mechanical properties of tumors depend on probing frequency. Thus, a comprehensive evaluation of tumor mechanics requires measurements across a broader frequency spectrum. In addition to these two cell lines, we also looked at MCF 7 cells and consistent to half relaxation data, effective viscosities of MCF 7 were in between MDA-MB-231 and MCF10A.
Plasticity and spheroid expansion rate under long time compression
Plasticity, a permanent deformation when subjected to an external force, is an important characteristic of biological materials. In tumors, tissue plasticity is often correlated with tumor malignancy progression and invasion.58–60 To quantify plasticity, we monitored spheroid strain relaxation under prolonged periodic compressions using a 40-min square pressure wave (see Movies S3 and S4). Across all three cell lines, spheroid strain increased continuously during compression, and upon release, the strain did not return to its original value (Fig. 4A–C). To quantify this, we measured the residual strain by calculating the difference of the strain at the end of two successive cycles (illustration shown in Fig. 4A).Residual strain provides a measure of irreversible deformation accumulated within the spheroids after each cycle of compression. We found there to be a significant difference in residual strain between malignant and non-malignant cell lines. The malignant MDA-MB-231 spheroid has an average residual strain of 0.166 ± 0.038, and in contrast, the non-tumorigenic MCF10A spheroid has an average residual strain of 0.038 ± 0.011 (see Fig. 4G). Similar residual strain values for cycle 3 and cycle 2 were quantified (see Fig. 4H). The elevated residual strain in MDA-MB-231 spheroids reflects a higher degree of plasticity and potentially structural remodelling in comparison to non-malignant spheroids. In addition, we converted this area deformation to height deformation using FEM analysis discussed in Supplementary theoretical model Section 3 and quantified plasticity using the linear strain values (see Fig. S7).
We observed a continuous increase in spheroid area during compression phase as seen in Fig. 4A–C and Fig. S8A for all types of spheroids. The gradual increase of spheroid area under compression signifies an expansion of the spheroid into the surrounding matrix. We quantified this expansion using a linear fit to the spheroid area change as a function of time (see Fig. 4D–F). For all spheroids, we observed a near linear expansion in area with time. The linear fit to the data showed that the metastatic MDA-MB-231 spheroids exhibited the steepest slopes with an average value of (4.45 ± 0.89) × 10−3 min−1. In contrast, MCF10A spheroids displayed significantly lower slopes, with an average value of (1.18 ± 0.32) × 10−3 min−1, see (Fig. 4I). This is consistent with the understanding that normal epithelial cell line MCF10A has more stable cytoskeleton and stronger cell–cell adhesion that leads to slower expansion than malignant MDA-MB-231.61,62
Together, these findings revealed that tumor spheroids exhibit plasticity under periodic compression, with malignant spheroids demonstrating enhanced plasticity than the non-tumorigenic spheroids. These results complement our short-timescale observations and underscore the importance of investigating tumor mechanics across multiple timescales to fully capture the dynamics of living materials.
Conclusion and future perspectives
In this work, we investigated the viscoelastic properties of breast tumor spheroids using an integrated microfluidic compression device and theoretical modelling. Our results showed that breast tumor spheroids at three different malignancy states followed a modified power law model. Notably, their mechanical phenotypes varied significantly with malignancy state: upon release of a constant compressive strain, non-tumorigenic spheroids recovered rapidly and behaved more elastically on a short time scales (seconds) and possessed less of a permanent deformation (plasticity) than those of malignant tumors at longer time scale (minutes). These observations are consistent with previous work where stable cytoskeleton and stronger e-cadherin mediated cell–cell adhesion has been correlated with less invasion.61,62The modified power law model used here provided a simple and generic method for modelling living materials where multiple time scales are involved. We note that the observed power-law behavior is limited to short-time relaxation, whereas at longer time scales (minutes) plasticity behavior becomes increasingly dominant. In the future, we plan to interrogate biological activities at various time scales individually including cytoskeletal molecule dynamics, ECM remodelling and cell migration to understand the nature of multi-time scale dynamics presented here. We also hope that the modified power law model can be used for characterizing mechanics of other tissue types to further understand the biological implications of viscoelastic properties of living tissue.
Materials and methods
Cells, spheroids, and 3D spheroid culture preparation
Microfluid compression device fabrication
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 polydimethylsiloxane (PDMS) mixture, degassed, and poured into a 5.3 mm thick polycarbonate frame placed around the feature on the silicon wafer. A sheet of laser printer transparency film was placed on top to create a smooth surface, followed by a glass panel with a weight to ensure even thickness. The PDMS was then cured at 60 °C overnight. The cured PDMS was cut, removed from the silicon master, plasma cleaned for 1 minute, and treated with in-house FOTS setup overnight. This PDMS mold was then used for second casting to get actual L1 on a glass slide using 10 g of 10
1 polydimethylsiloxane (PDMS) mixture, degassed, and poured into a 5.3 mm thick polycarbonate frame placed around the feature on the silicon wafer. A sheet of laser printer transparency film was placed on top to create a smooth surface, followed by a glass panel with a weight to ensure even thickness. The PDMS was then cured at 60 °C overnight. The cured PDMS was cut, removed from the silicon master, plasma cleaned for 1 minute, and treated with in-house FOTS setup overnight. This PDMS mold was then used for second casting to get actual L1 on a glass slide using 10 g of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. A clean 25 mm × 75 mm × 1 mm glass slide was taped on all four sides onto the bottom of a Petri dish, and PDMS was poured on top of the glass slide and the feature side of the PDMS mold. After removing all bubbles in a vacuum chamber, the PDMS mold was flipped onto the slide, secured with tape, weighted down flush such that the height of the device is defined by the PDMS mold. The assembly was cured overnight at 60 °C and then carefully peeled away, leaving the L1 securely attached to the glass slide.
1. A clean 25 mm × 75 mm × 1 mm glass slide was taped on all four sides onto the bottom of a Petri dish, and PDMS was poured on top of the glass slide and the feature side of the PDMS mold. After removing all bubbles in a vacuum chamber, the PDMS mold was flipped onto the slide, secured with tape, weighted down flush such that the height of the device is defined by the PDMS mold. The assembly was cured overnight at 60 °C and then carefully peeled away, leaving the L1 securely attached to the glass slide.
To fabricate the piston membrane layer L2, 10 g of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 PDMS was prepared and degassed. A 340 µm thick rectangular frame was placed around the feature on the silicon wafer, and PDMS was poured into the frame. A sheet of transparency film was carefully placed on top without introducing bubbles, followed by a glass slide and a weight. The PDMS was cured at 60 °C overnight. After curing, the frame and excess PDMS were carefully removed, leaving only the center PDMS piece on the wafer ready to be bonded with the L3 later.
1 PDMS was prepared and degassed. A 340 µm thick rectangular frame was placed around the feature on the silicon wafer, and PDMS was poured into the frame. A sheet of transparency film was carefully placed on top without introducing bubbles, followed by a glass slide and a weight. The PDMS was cured at 60 °C overnight. After curing, the frame and excess PDMS were carefully removed, leaving only the center PDMS piece on the wafer ready to be bonded with the L3 later.
To fabricate the pressure chamber layer L3, 25 g of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 PDMS was prepared and degassed. A 5.3 mm thick rectangular frame was positioned on the silicon wafer, and PDMS was poured into it. A sheet of transparency film was placed on top without introducing bubbles, followed by a glass panel and a weight. The PDMS was cured at 60 °C overnight. After curing, the PDMS was cut and removed from the wafer, and a 1.5 mm hole was punched at the inlet using a biopsy punch for pressure delivery.
1 PDMS was prepared and degassed. A 5.3 mm thick rectangular frame was positioned on the silicon wafer, and PDMS was poured into it. A sheet of transparency film was placed on top without introducing bubbles, followed by a glass panel and a weight. The PDMS was cured at 60 °C overnight. After curing, the PDMS was cut and removed from the wafer, and a 1.5 mm hole was punched at the inlet using a biopsy punch for pressure delivery.
Microfluidic compression device assembly and operation
The pressure control unit is made by bonding L2 and L3. Both L2 (attached to the wafer) and L3 PDMS were plasma cleaned for 1 minute in a plasma cleaner (PDC-001-HP, Harrick Plasma, Ithaca, NY, USA). The features were then carefully aligned under a microscope and bonded. The bonded layers were placed in a 90 °C oven for 15 minutes to strengthen the bond, then cooled slowly to room temperature. Finally, the PDMS was carefully detached from the wafer and cut to match the size of L2. Then, 4 access ports were made using 2 mm biopsy punches to allow media replenishment and gas exchange.Prior to experiment, the pressure control chambers were filled with water, and the surface of the sample loading chamber was activated for better adhesion with type 1 collagen gel. For the pressure control unit preparation, the pressure control unit was plasma cleaned for 30 seconds and submerged under water overnight to fill the pressure control chamber with water. Then, a tubing (OD and ID of 1/16″ and 1/32″, respectively) pre-filled with water from the pressure controller (Elveflow OB1 MK4, Paris, France) was connected to the 1.5 mm inlet on the device. To prepare the sample loading chamber layer L1, the well surface was treated such that collagen matrices can be bonded to the surface. Here, each well in the L1 was selectively plasma cleaned for 20 seconds with a mask, treated with 1% polyethylenimine (PEI) for 10 minutes, washed with PBS once, and treated with 0.5% glutaraldehyde for 30 minutes. After three washes with PBS, the L1 was ready for use in the experiment.
On the day of the experiment, 8 µL of 1.5 mg mL−1 collagen embedded tumor spheroids were loaded to the sample chamber L1. The pressure control unit was then aligned under a microscope and lowered onto the L1. The three layers were then sandwiched between a metal frame and a Plexiglas window, secured with screws to ensure uniform pressure around the entire assembly as shown in Fig. 1A. The assembled device was then placed in a 37 °C and 5% CO2 incubator for 45 minutes for collagen polymerization. After the polymerization, the device was carefully filled with cell media through the four access ports. Lastly, the device was connected to the Elveflow pressure controller and placed on a temperature, humidity, and CO2 controlled microscope stage for imaging.
The pressure in the pressure control unit was regulated using a commercially available pressure controller (OB1 MK4, Elveflow Inc., Paris, France), operated via Elvesys software. The software allows users to generate pressure waveforms with customizable profiles. The piston's stability and responsiveness under pressure inputs were validated, for details refer.43 An external pressure source (Model 3 Compressor, Jun-Air, MI, USA) supplied pressure to the OB1 MK4 controller.
Imaging and data analysis
All images were taken using an inverted epi-fluorescent microscope (IX81, Olympus America, Center Valley, PA, USA) with a CCD camera (ORCA-R2, Hamamatsu Photonics, Bridgewater, NJ, USA). In all the experiments the middle z-plane of the spheroids was captured using a 20× objective (Olympus, NA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) =
= ![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) in bright field. The scope has a stage incubator (Precision Plastics Inc., Beltsville, MD, USA) that maintained a temperature of 37 °C, humidity of ∼70%, and 5% CO2 level. The setup was placed on the automated X–Y microscope stage (MS-2000, Applied Scientific Instrumentation, Eugene, OR), and images were taken at ∼16 frames per second for 40 s square wave spheroid mechanics data and at 1 frame per minute rate for 40-minute tumor mechanics data using CellSens software (Olympus America, Center Valley, PA, USA). The 16 frames per second frame rate allows us to observe the compression and relaxation dynamics of the spheroid on a timescale of ∼60 ms.
1) in bright field. The scope has a stage incubator (Precision Plastics Inc., Beltsville, MD, USA) that maintained a temperature of 37 °C, humidity of ∼70%, and 5% CO2 level. The setup was placed on the automated X–Y microscope stage (MS-2000, Applied Scientific Instrumentation, Eugene, OR), and images were taken at ∼16 frames per second for 40 s square wave spheroid mechanics data and at 1 frame per minute rate for 40-minute tumor mechanics data using CellSens software (Olympus America, Center Valley, PA, USA). The 16 frames per second frame rate allows us to observe the compression and relaxation dynamics of the spheroid on a timescale of ∼60 ms.
Author contributions
MP and MW created the research project. MP, YJS and MW designed the project. MP carried out the experiments. MP and KR performed data analysis. BZ, and CYH carried out the theoretical modelling work shown in the SI. The manuscript was collaboratively written by MP, JES, and MW, with contributions from all authors.Conflicts of interest
There are no conflicts to declare.Data availability
Supplementary information is available. See DOI: https://doi.org/10.1039/d5sm00949a.Data sets generated during the study are available from the corresponding author on reasonable request.
Acknowledgements
This work was supported by a grant from the National Institute of Health (Grant No. R01CA221346). The microfluidic device used in this work was developed at the Cornell NanoScale Facility, a member of the National Nanotechnology Coordinated Infrastructure (NNCI), which is supported by the National Science Foundation (Grant NNCI2025233). JES is the Betty and Sheldon Feinberg Senior Faculty Scholar in Cancer Research.References
- D. T. Butcher, T. Alliston and V. M. Weaver, A tense situation: forcing tumour progression, Nat. Rev. Cancer, 2009, 9(2), 108–122, DOI:10.1038/nrc2544.
- F. Bordeleau, B. N. Mason, E. M. Lollis, M. Mazzola, M. R. Zanotelli, S. Somasegar, J. P. Califano, C. Montague, D. J. LaValley and J. Huynh, et al., Matrix stiffening promotes a tumor vasculature phenotype, Proc. Natl. Acad. Sci. U. S. A., 2017, 114(3), 492–497, DOI:10.1073/pnas.1613855114.
- P. P. Provenzano, K. W. Eliceiri, J. M. Campbell, D. R. Inman, J. G. White and P. J. Keely, Collagen reorganization at the tumor-stromal interface facilitates local invasion, BMC Med., 2006, 4(1), 38, DOI:10.1186/1741-7015-4-38 (acccessed 2022-12-14T17:06:35).
- R. K. Jain, J. D. Martin and T. Stylianopoulos, The Role of Mechanical Forces in Tumor Growth and Therapy, Annu. Rev. Biomed. Eng., 2014, 16(1), 321–346, DOI:10.1146/annurev-bioeng-071813-105259 (acccessed 2023-06-07T21:09:07).
- K. R. Levental, H. Yu, L. Kass, J. N. Lakins, M. Egeblad, J. T. Erler, S. F. T. Fong, K. Csiszar, A. Giaccia and W. Weninger, et al., Matrix Crosslinking Forces Tumor Progression by Enhancing Integrin Signaling, Cell, 2009, 139(5), 891–906, DOI:10.1016/j.cell.2009.10.027.
- J. A. Linke, L. L. Munn and R. K. Jain, Compressive stresses in cancer: characterization and implications for tumour progression and treatment, Nat. Rev. Cancer, 2024, 24(11), 768–791 Search PubMed.
- C. Alibert, B. Goud and J. B. Manneville, Are cancer cells really softer than normal cells?, Biol. Cell, 2017, 109(5), 167–189 CrossRef PubMed.
- M. Plodinec, M. Loparic, C. A. Monnier, E. C. Obermann, R. Zanetti-Dallenbach, P. Oertle, J. T. Hyotyla, U. Aebi, M. Bentires-Alj and R. Y. Lim, The nanomechanical signature of breast cancer, Nat. Nanotechnol., 2012, 7(11), 757–765 CrossRef CAS PubMed.
- B. Zbiral, A. Weber, M. D. Vivanco and J. L. Toca-Herrera, Characterization of breast cancer aggressiveness by cell mechanics, Int. J. Mol. Sci., 2023, 24(15), 12208 CrossRef CAS PubMed.
- S. Kwon, W. Yang, D. Moon and K. S. Kim, Comparison of cancer cell elasticity by cell type, J. Cancer, 2020, 11(18), 5403 Search PubMed.
- N. Schierbaum, J. Rheinlaender and T. E. Schäffer, Viscoelastic properties of normal and cancerous human breast cells are affected differently by contact to adjacent cells, Acta Biomater., 2017, 55, 239–248 Search PubMed.
- V. Gensbittel, M. Kräter, S. Harlepp, I. Busnelli, J. Guck and J. G. Goetz, Mechanical adaptability of tumor cells in metastasis, Dev. Cell, 2021, 56(2), 164–179 Search PubMed.
- A. S. G. van Oosten, X. Y. Chen, L. K. Chin, K. Cruz, A. E. Patteson, K. Pogoda, V. B. Shenoy and P. A. Janmey, Emergence of tissue-like mechanics from fibrous networks confined by close-packed cells, Nature, 2019, 573(7772), 96–101, DOI:10.1038/s41586-019-1516-5.
- A. V. Taubenberger, S. Girardo, N. Träber, E. Fischer-Friedrich, M. Kräter, K. Wagner, T. Kurth, I. Richter, B. Haller and M. Binner, 3D microenvironment stiffness regulates tumor spheroid growth and mechanics via p21 and ROCK, Adv. Biosyst., 2019, 3(9), 1900128 CrossRef PubMed.
- T. Fuhs, F. Wetzel, A. W. Fritsch, X. Li, R. Stange, S. Pawlizak, T. R. Kießling, E. Morawetz, S. Grosser and F. Sauer, Rigid tumours contain soft cancer cells, Nat. Phys., 2022, 18(12), 1510–1519 Search PubMed.
- M. Cantini, H. Donnelly, M. J. Dalby and M. Salmeron-Sanchez, The plot thickens: the emerging role of matrix viscosity in cell mechanotransduction, Adv. Healthcare Mater., 2020, 9(8), 1901259 Search PubMed.
- C. T. Mierke, Viscoelasticity, like forces, plays a role in mechanotransduction, Front. Cell Dev. Biol., 2022, 10, 789841 Search PubMed.
- A. Saraswathibhatla, D. Indana and O. Chaudhuri, Cell–extracellular matrix mechanotransduction in 3D, Nat. Rev. Mol. Cell Biol., 2023, 24(7), 495–516 CrossRef CAS PubMed.
- M. Tavasso, A. D. Bordoloi, E. Tanré, S. A. Dekker, V. Garbin and P. E. Boukany, Linking metastatic potential and viscoelastic properties of breast cancer spheroids via dynamic compression and relaxation in microfluidics, Adv. Healthcare Mater., 2025, 14(6), 2402715 Search PubMed.
- A. Zubiarrain-Laserna, D. Martínez-Moreno, J. L. de Andrés, L. de Lara-Peña, O. Guaresti, A. M. Zaldua, G. Jiménez and J. A. Marchal, Beyond stiffness: deciphering the role of viscoelasticity in cancer evolution and treatment response, Biofabrication, 2024, 16(4), 042002 CrossRef CAS PubMed.
- O. Chaudhuri, J. Cooper-White, P. A. Janmey, D. J. Mooney and V. B. Shenoy, Effects of extracellular matrix viscoelasticity on cellular behaviour, Nature, 2020, 584(7822), 535–546 CrossRef CAS PubMed.
- Z. Gong, S. E. Szczesny, S. R. Caliari, E. E. Charrier, O. Chaudhuri, X. Cao, Y. Lin, R. L. Mauck, P. A. Janmey and J. A. Burdick, Matching material and cellular timescales maximizes cell spreading on viscoelastic substrates, Proc. Natl. Acad. Sci. U. S. A., 2018, 115(12), E2686–E2695 Search PubMed.
- M. Hadzipasic, S. Zhang, Z. Huang, R. Passaro, M. S. Sten, G. M. Shankar and H. T. Nia, Emergence of nanoscale viscoelasticity from single cancer cells to established tumors, Biomaterials, 2024, 305, 122431 CrossRef CAS PubMed.
- B. Buchmann, P. Fernández and A. R. Bausch, The role of nonlinear mechanical properties of biomimetic hydrogels for organoid growth, Biophys. Rev., 2021, 2(2), 021401 CrossRef PubMed.
- B. Sun, The mechanics of fibrillar collagen extracellular matrix, Cell Rep. Phys. Sci., 2021, 2(8), 100515 Search PubMed.
- S. Muenster, L. M. Jawerth, B. A. Leslie, J. I. Weitz, B. Fabry and D. A. Weitz, Strain history dependence of the nonlinear stress response of fibrin and collagen networks, Proc. Natl. Acad. Sci. U. S. A., 2013, 110(30), 12197–12202, DOI:10.1073/pnas.1222787110.
- C. M. Nelson, M. M. VanDuijn, J. L. Inman, D. A. Fletcher and M. J. Bissell, Tissue geometry determines sites of mammary branching morphogenesis in organotypic cultures, Science, 2006, 314(5797), 298–300 CrossRef CAS PubMed.
- Z. Yan, X. Xia, W. C. Cho, D. W. Au, X. Shao, C. Fang, Y. Tian and Y. Lin, Rapid plastic deformation of cancer cells correlates with high metastatic potential, Adv. Healthcare Mater., 2022, 11(8), 2101657 CrossRef CAS PubMed.
- Y. Abidine, A. Giannetti, J. Revilloud, V. M. Laurent and C. Verdier, Viscoelastic properties in cancer: From cells to spheroids, Cells, 2021, 10(7), 1704 CrossRef CAS PubMed.
- M. Krieg, G. Fläschner, D. Alsteens, B. M. Gaub, W. H. Roos, G. J. Wuite, H. E. Gaub, C. Gerber, Y. F. Dufrêne and D. J. Müller, Atomic force microscopy-based mechanobiology, Nat. Rev. Phys., 2019, 1(1), 41–57 Search PubMed.
- Y. M. Efremov, W.-H. Wang, S. D. Hardy, R. L. Geahlen and A. Raman, Measuring nanoscale viscoelastic parameters of cells directly from AFM force-displacement curves, Sci. Rep., 2017, 7(1), 1541 CrossRef PubMed.
- R. C. Boot, A. Roscani, L. van Buren, S. Maity, G. H. Koenderink and P. E. Boukany, High-throughput mechanophenotyping of multicellular spheroids using a microfluidic micropipette aspiration chip, Lab Chip, 2023, 23(7), 1768–1778 Search PubMed.
- K. Guevorkian and J.-L. Maître, Micropipette aspiration: A unique tool for exploring cell and tissue mechanics in vivo. Methods in cell biology, Elsevier, 2017, vol. 139, pp. 187–201 Search PubMed.
- E. Zhou, S. Quek and C. Lim, Power-law rheology analysis of cells undergoing micropipette aspiration, Biomech. Model. Mechanobiol., 2010, 9(5), 563–572 CrossRef CAS PubMed.
- S. Onal, M. M. Alkaisi and V. Nock, A flexible microdevice for mechanical cell stimulation and compression in microfluidic settings, Front. Phys., 2021, 9, 654918 Search PubMed.
- D. N. Hohne, J. G. Younger and M. J. Solomon, Flexible microfluidic device for mechanical property characterization of soft viscoelastic solids such as bacterial biofilms, Langmuir, 2009, 25(13), 7743–7751 CrossRef CAS PubMed.
- S. Onal, M. M. Alkaisi and V. Nock, Application of sequential cyclic compression on cancer cells in a flexible microdevice, PLoS One, 2023, 18(1), e0279896 Search PubMed.
- R. Ferraro, S. Guido, S. Caserta and M. Tassieri, i-Rheo-optical assay: Measuring the viscoelastic properties of multicellular spheroids, Mater. Today Bio, 2024, 26, 101066 Search PubMed.
- S. Villa, P. Edera, M. Brizioli, V. Trappe, F. Giavazzi and R. Cerbino, Quantitative rheo-microscopy of soft matter, Front. Phys., 2022, 10, 1013805 CrossRef.
- I. Nagle, M. Tavasso, A. D. Bordoloi, I. A. Muntz, G. H. Koenderink and P. E. Boukany, Invasive cancer cells soften collagen networks and disrupt stress-stiffening via volume exclusion, contractility and adhesion, bioRxiv, 2025, preprint DOI:10.1101/2025.04.11.648338.
- Y. J. Suh, M. Liu, B. Zhu, M. Pandey, B. C. Cheung, J. Kim, N. Bouklas, C. Roh, J. E. Segall and C.-Y. Hui, A microfluidic rheometer for tumor mechanics and invasion studies, Lab Chip, 2025, 25(22), 6018–6032 Search PubMed.
- M. M. Wu, J. W. Roberts and M. Buckley, Three-dimensional fluorescent particle tracking at micron-scale using a single camera, Exp. Fluids, 2005, 38(4), 461–465, DOI:10.1007/s00348-004-0925-9.
- Y. J. Suh, M. Liu, B. Zhu, M. Pandey, B. Cheung, J. Kim, N. Bouklas, C. Roh, J. E. Segall, C.-Y. Hui and M. Wu, A microfluidic rheometer for tumor mechanics and invasion studies. Lab on a Chip, submitted Search PubMed.
- O. Ronneberger; P. Fischer and T. Brox, U-net: Convolutional networks for biomedical image segmentation. In Medical image computing and computer-assisted intervention–MICCAI 2015: 18th international conference, Munich, Germany, October 5-9, 2015, proceedings, part III 18, 2015, Springer, pp. 234–241.
- W. Song, C.-K. Tung, Y.-C. Lu, Y. Pardo, M. Wu, M. Das, D.-I. Kao, S. Chen and M. Ma, Dynamic self-organization of microwell-aggregated cellular mixtures, Soft Matter, 2016, 12(26), 5739–5746 RSC.
- Y.-C. Fung, Biomechanics: mechanical properties of living tissues, Springer Science & Business Media, 2013 Search PubMed.
- J. Song, N. Holten-Andersen and G. H. McKinley, Non-Maxwellian viscoelastic stress relaxations in soft matter, Soft Matter, 2023, 19(41), 7885–7906 Search PubMed.
- A. Ivascu and M. Kubbies, Diversity of cell-mediated adhesions in breast cancer spheroids, Int. J. Oncol., 2007, 31(6), 1403–1413 CAS.
- Y. L. Huang, Y. Ma, C. Wu, C. Shiau, J. E. Segall and M. Wu, Tumor spheroids under perfusion within a 3D microfluidic platform reveal critical roles of cell-cell adhesion in tumor invasion, Sci. Rep., 2020, 10(1), 9648, DOI:10.1038/s41598-020-66528-2.
- J. De Sousa, R. Freire, F. Sousa, M. Radmacher, A. Silva, M. Ramos, A. Monteiro-Moreira, F. Mesquita, M. Moraes and R. Montenegro, Double power-law viscoelastic relaxation of living cells encodes motility trends, Sci. Rep., 2020, 10(1), 4749 CrossRef CAS PubMed.
- A. Bonfanti, J. L. Kaplan, G. Charras and A. Kabla, Fractional viscoelastic models for power-law materials, Soft Matter, 2020, 16(26), 6002–6020 Search PubMed.
- S. Aime, L. Cipelletti and L. Ramos, Power law viscoelasticity of a fractal colloidal gel, J. Rheol., 2018, 62(6), 1429–1441 CrossRef CAS.
- R. L. Bagley, Power law and fractional calculus model of viscoelasticity, AIAA J., 1989, 27(10), 1412–1417 CrossRef CAS.
- R. M. Christensen, Theory of viscoelasticity, Courier Corporation, 2013 Search PubMed.
- M. Das, J. L. Waeterloos, C. Clasen and G. H. McKinley, Single cells are compactly and accurately described as fractional Kelvin-Voigt materials, Rheol. Acta, 2025, 1–15 Search PubMed.
- R. Ferraro, S. Caserta and S. Guido, A Low-Cost, User-Friendly Rheo-Optical Compression Assay to Measure Mechanical Properties of Cell Spheroids in Standard Cell Culture Plates, Adv. Mater. Technol., 2024, 9(4), 2301890 CrossRef CAS.
- J. Vincent, Structural biomaterials, Structural Biomaterials, Princeton University Press, 2012 Search PubMed.
- S. R. Torborg, Z. Li, J. E. Chan and T. Tammela, Cellular and molecular mechanisms of plasticity in cancer, Trends Cancer, 2022, 8(9), 735–746 CrossRef CAS PubMed.
- J. Gotzmann, M. Mikula, A. Eger, R. Schulte-Hermann, R. Foisner, H. Beug and W. Mikulits, Molecular aspects of epithelial cell plasticity: implications for local tumor invasion and metastasis, Mutat. Res./Rev. Mutat. Res., 2004, 566(1), 9–20 Search PubMed.
- P. Friedl and S. Alexander, Cancer invasion and the microenvironment: plasticity and reciprocity, Cell, 2011, 147(5), 992–1009 Search PubMed.
- The Physical Sciences – Oncology Centers Network, A physical sciences network characterization of non-tumorigenic and metastatic cells, Sci. Rep., 2013, 3(1), 1449 Search PubMed.
- C. Alibert, B. Goud and J.-B. Manneville, Are cancer cells really softer than normal cells, Biol. Cell, 2017, 109(5), 167–189, DOI:10.1111/boc.201600078 (acccessed 2022-12-04T21:48:33).
- J. D. Shields, M. E. Fleury, C. Yong, A. A. Tomei, G. J. Randolph and M. A. Swartz, Autologous chemotaxis as a mechanism of tumor cell homing to lymphatics via interstitial flow and autocrine CCR7 signaling, Cancer Cell, 2007, 11(6), 526–538, DOI:10.1016/j.ccr.2007.04.020.
- M. Pandey, Y. J. Suh, M. Kim, H. J. Davis, J. E. Segall and M. Wu, Mechanical compression regulates tumor spheroid invasion into a 3D collagen matrix, Phys. Biol., 2024, 21(3), 036003 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |



