 Open Access Article
Open Access ArticleSpatially resolved fast dynamics reveal the aggregation mechanism in two-dimensions
Tamoghna
Das
 *a and
Mahesh M.
Bandi
b
*a and
Mahesh M.
Bandi
b
aWPI Nano Life Science Institute (WPI-NanoLSI), Kanazawa University, Kakuma-machi, Kanazawa, 920-1192, Japan. E-mail: tamoghna@staff.kanazawa-u.ac.jp
bNonlinear and Non-equilibrium Physics Unit, Okinawa Institute of Science and Technology, 1919-1 Tancha, Onna, Okinawa, 904-0495, Japan. E-mail: bandi@oist.jp
First published on 23rd January 2026
Abstract
Two typical morphologies of two-dimensional aggregates are considered: compact crystalline clusters and string-like non-compact conformations. Simulated trajectories of both types of aggregates are analysed with fine spatial resolution. While the long-time geometry of such trajectories appears to be statistically identical for the two conformations, the self-overlap statistics reveal two distinct short-time pre-caging mechanisms. While the length-scale is directly proportional to the time-scale for particles in compact aggregates, a non-monotonic relationship holds for non-compact clusters. The relationship between short length-scale and fast time-scale for particle localization might hold the key to the structure–function relationship of aggregate forming systems and other non-equilibrium soft materials.
Introduction
Competition between the length and time scales lies at the heart of the formation of self-assembled structures.1–3 This competition happens most naturally in the presence of a short-range attraction and a long-range repulsion as it appears in systems ranging from astronomical length scale, such as the pasta phase in neutron stars4 to micron-sized colloidal systems,5–9 and nano-scale metallic, organometallic and polymeric self-assemblies10–12 to biologically relevant macro-molecule formations.13–15 While the exact nature of the interactions varies widely depending on the systems, the geometric frustration originating from the competing interactions leads to the localization or caging of the constituent units. The emergent finite length and time scales of the cage then govern the global structure and dynamics of the resulting self-assembled structures. Over the past decades, this picture has been actively constructed and revised through numerous theoretical and experimental efforts.16–19The caging of a diffusive particle by its neighbors is a purely dynamic process. Once caged, the particle rattles within a local energy minimum until a collective, cooperative rearrangement of the neighbors provides an escape route. Through these dynamic rearrangement events, the particle continues its motion via a series of intermittent cage-free ballistic motions punctuated by occasional within-cage rattling motions. This intuitive picture has been instrumental in providing a microscopic basis for the observation of co-existing spatially distinct regions of different mobilities aka dynamical heterogeneity, a central and common feature of non-equilibrium particulate systems.20 However, the identification of a cage has always been system dependent.
The energy landscape21 of an interacting many-body system is a multi-dimensional object, difficult to visualize, but a useful theoretical construct necessary to understand the system and its underlying dynamics. The rattling motion of a caged particle gives us access to a local basin of this landscape. A huge body of work is devoted to characterizing the spatio-temporal organization of such basins in terms of particle motion statistics.22–30 The list of references in this regard is only a representative selection. In contrast, the intermittent cage-hopping motion of particles, which can be informative about the evolution of local basins, has received relatively less attention.31–33 In this work, we attempt detailed characterization of the delocalization dynamics of a particle in a generic self-assembly system with an aim to understand the role of geometric frustration in this process.
Specifically, we have analysed the simulated trajectories of an aggregate forming system in two dimensions. Two limiting aggregate conformations are chosen for comparison: compact clusters with a local crystalline arrangement of particles and string-like non-compact aggregates. Although these are the steady state structures, i.e., no coarsening is observed, they are highly dynamic. Particles detach from their parent clusters and move freely before they reattach to the same or a neighboring cluster. We refer to this intermediate excursion of particles, in between their cages, as the pre-caging dynamics. Examining the self-overlap of a particle as a function of a certain bounding length scale provides us with information about this entire dynamic behaviour. It turns out that this intermediate cage-free motion is not ideally free as it must be under the influence of the underlying potential energy landscape. In fact, the nature of pre-caging in two different aggregation environments is found to be quite opposite which reveals certain intriguing relations between the fast dynamics and the energy landscape as explained later. In what follows, we first describe the features of our model system and the simulation protocol briefly. The main findings are presented next. Following this, we conclude our study with a detailed discussion and mention of possible future directions.
Method and model
A two-dimensional system of about 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 monodisperse particles that interact pair-wise via Φ(r) is considered within a periodic box. The density of the system is set at ρ = 0.4 by choosing a box of suitable size. The trajectory of the system is simulated in a canonical ensemble using molecular dynamics. The motion of individual particles is assumed to follow Langevin dynamics where thermal fluctuations are modeled in a spatially uncorrelated and temporally delta-correlated fashion. The temperature T of the system is expressed in ε units assuming the Boltzmann constant kB = 1. The equations of motion are integrated using the velocity Verlet algorithm as implemented within LAMMPS.34 Setting the integration time steps δt = 10−5, the time is expressed as
000 monodisperse particles that interact pair-wise via Φ(r) is considered within a periodic box. The density of the system is set at ρ = 0.4 by choosing a box of suitable size. The trajectory of the system is simulated in a canonical ensemble using molecular dynamics. The motion of individual particles is assumed to follow Langevin dynamics where thermal fluctuations are modeled in a spatially uncorrelated and temporally delta-correlated fashion. The temperature T of the system is expressed in ε units assuming the Boltzmann constant kB = 1. The equations of motion are integrated using the velocity Verlet algorithm as implemented within LAMMPS.34 Setting the integration time steps δt = 10−5, the time is expressed as 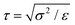 . The mass of the particles is set to unity.
. The mass of the particles is set to unity.
The particular realization of competing interactions,35Φ(r) = ΦSA + ΦLR, used in this study is a superposition of a short-range attraction (SA), ΦSA and a long-range repulsion (LR), ΦLR, henceforth jointly referred to as SALR to represent Φ(r). The two potentials have the following forms: ΦSA = 4ε[(σ/r)2α − (σ/r)α] and ΦLR = (Aσ/r)exp(−r/ξ) where σ is the particle diameter. Setting the energy scale by ε, the strength of attraction is matched with the repulsion strength A = 4ε. The range of attraction is fixed at 0.2σ by setting α = 18 and the range of repulsion ξ is tuned to obtain different morphologies. This simple realization of SALR interaction is motivated by a more detailed model for polymer-grafted nanoparticle systems.36,37 To model SALR type interactions, attempts have been made from a very simple square-well type potential38 to much more sophisticated potentials with many parameters to mimic the exact experimental conditions. See ref. 39 for a comprehensive list. However, for any system with competing interactions, it is possible to define a single control parameter as the ratio of relevant length scales.40 Hence, the results presented here are in general transferable to other systems with similar SALR interactions.
We have chosen two specific values of ξ = 0.5 and 0.8 for this study. For each ξ, we begin our simulation by preparing a high temperature equilibrium configuration Ti = ε. An ensemble of 100 equilibrium configurations is saved as the initial condition. Each equilibrium liquid configuration is then cooled very slowly to a final temperature Tf = 0.05ε over a period of time of 104τ. This slow cooling protocol is chosen to minimize the preparation rate dependence commonly reported in particulate glassy systems.41 After reaching Tf, the simulation runs for a period of 103τ, and the trajectories during this time are saved for further analysis. Temperature fluctuation during this production run is monitored and found to be around 10−3ε ensuring a thermodynamic steady state. A detailed discussion about the energy fluctuation in this system can be found in our previous study.42
Note the presence of the energy barrier in the effective potential, Φ(r), as a result of the competition between the two length scales of the two contributing potentials (Fig. 1A). This is the key to the geometric frustration experienced by a particle in a many-body configuration. By tuning ξ, the height of this barrier and the nature of the minimum behind this barrier can be controlled. At ξ = 0.5, there is a negative global minimum in Φ(r) for small pair-wise distance, r ∼ σ, behind a modest positive energy barrier that decays to zero asymptotically at large r values (≫3σ). The particles, under the dominant close-range attraction, aggregate into compact, locally crystalline arrangements of finite size (Fig. 1B). As the repulsive length scale increases to ξ = 0.8, the global Φ(r) minimum becomes local and is now protected by an even stronger positive energy barrier. The system can only achieve a global minimum state if the particles are far enough from each other pointing to an infinite dilution. For any modest density of interacting particles, the system would then be perpetually frustrated and metastable. In such a scenario, the particles prefer to arrange themselves in locally stable ramified structures of nearly string-like conformations (Fig. 1C).
The overall shapes of representative trajectories from two different conformations appear to be very different. Particles in compact clusters appear to explore the space in such a way that over long time the trajectory fills a nearly circular region (Fig. 1B, inset). In comparison, the trajectory geometry of particles in string-like aggregates appears to be more jagged in space as the particle moves within a narrow restricted space (Fig. 1C, inset). The spatial localization and escape from that is more evident though, in this latter case of string-like aggregates. The displacement statistics, as reported earlier,42 for both aggregates is non-Gaussian but with different features. The self-displacement fluctuations are dominantly Gaussian with a small exponential tail for the compact clusters. The Gaussian nature is recessive and is buried within the long exponential tail for string-like clusters. The residence time of a particle in a compact cluster is also significantly larger than that in a string-like structure. It was also shown that the interrelation between the mobility of an individual particle and its neighborhood fluctuation is dictated by the fractal dimension of the cluster it forms.43 This remarkable structure–dynamics correspondence prompted us to examine the particle dynamics rigorously.
Results
The two aggregate morphologies exhibit markedly different long-time dynamics, as evidenced by their mean-squared displacement (MSD) 〈r2(t)〉 (Fig. 2). The compact cluster system shows a ballistic (〈r2(t)〉 ∼ t2) to diffusive (〈r2(t)〉 ∼ t) crossover through a strong intermediate time relaxation (∼t0.4) spanning over almost two decades of time scale. This intermediate relaxation is much more gentle for the string-like aggregates (〈r2(t)〉 ∼ t0.8) before it turns to diffusive behaviour. However, the diffusive dynamics does not persist in the long time; instead, it turns to a typical sub-diffusion (∼t0.5) within our observation time. We speculate that the ramification of available space due to the specific aggregation morphology in this system is responsible for such behaviour. However, we leave this aspect for future exploration and focus on the individual trajectories, instead, to understand the dynamics of the departure from ballistic behaviour.Recalling the visual difference in the representative trajectories, we first investigate the statistical properties of these trajectories imagining them as growing polymers. Each monomer in these polymers would represent the position of a particle at a certain time. The total number of monomers ![[scr N, script letter N]](https://www.rsc.org/images/entities/char_e52d.gif) would then be exactly equal to the total time t over which the trajectories are recorded. Unlike the usual polymers, polymers with such a trajectory would not be self-avoiding and would be highly entangled in space. Yet, we find that both end-to-end distance 〈
would then be exactly equal to the total time t over which the trajectories are recorded. Unlike the usual polymers, polymers with such a trajectory would not be self-avoiding and would be highly entangled in space. Yet, we find that both end-to-end distance 〈![[script L]](https://www.rsc.org/images/entities/char_e144.gif) 〉 and the radius of gyration
〉 and the radius of gyration ![[scr R, script letter R]](https://www.rsc.org/images/entities/char_e531.gif) g of these polymers (trajectories) are unable to pick up any such signature even after a long time. Since the MSDs for different aggregates show distinct features after the initial ballistic regime, only the part of the trajectories after this regime is considered for comparing their shapes. 〈
g of these polymers (trajectories) are unable to pick up any such signature even after a long time. Since the MSDs for different aggregates show distinct features after the initial ballistic regime, only the part of the trajectories after this regime is considered for comparing their shapes. 〈![[script L]](https://www.rsc.org/images/entities/char_e144.gif) 〉 grows linearly with
〉 grows linearly with ![[scr N, script letter N]](https://www.rsc.org/images/entities/char_e52d.gif) for trajectories in both of the conformations (Fig. 3A). The shape of the polymer, quantified by
for trajectories in both of the conformations (Fig. 3A). The shape of the polymer, quantified by ![[scr R, script letter R]](https://www.rsc.org/images/entities/char_e531.gif) g is related to the number of monomers,
g is related to the number of monomers, 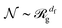 , where df is the fractal dimension of the polymer. For the trajectory polymers, a crossover from nearly linear (df ∼ 1) to almost circular shape (df ∼ 2) is observed as a function of increasing
, where df is the fractal dimension of the polymer. For the trajectory polymers, a crossover from nearly linear (df ∼ 1) to almost circular shape (df ∼ 2) is observed as a function of increasing ![[scr N, script letter N]](https://www.rsc.org/images/entities/char_e52d.gif) (time) (Fig. 3B). We mention that the large-
(time) (Fig. 3B). We mention that the large-![[scr N, script letter N]](https://www.rsc.org/images/entities/char_e52d.gif) behaviour is reported for localization of DNA into knots under strong adsorption.44 Though the shape crossover corresponds to the departure from ballistic dynamics, these measures turn out to be inadequate to provide any further detailed information about the process and also unable to distinguish between the trajectories from two different morphologies. This prompts us to examine the finer details of trajectories with the aid of a self-overlap function.
behaviour is reported for localization of DNA into knots under strong adsorption.44 Though the shape crossover corresponds to the departure from ballistic dynamics, these measures turn out to be inadequate to provide any further detailed information about the process and also unable to distinguish between the trajectories from two different morphologies. This prompts us to examine the finer details of trajectories with the aid of a self-overlap function.
How long a particle stays at a specific position is quantified using the self-overlap function by computing the total number of occurrences of a particle within a small cut-off radius σR around its initial position, over time t. According to this measure, a diffusing particle is expected to be found exponentially far away from its original position at a later time. In contrast, a localized particle is expected to be found within a region of size σR over a characteristic time τR. Examining this behavior over a range of σR will then be informative about spatio-temporal statistics of the localization–delocalization process. Formally, the measure reads as 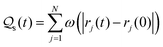 , with ω = 1 when |rj(t) − rj(0)| < σR and ω = 0 otherwise. Typically, σR is interpreted as the caging length and there is active debate about how to assign a reasonable value to it in the context of glassy materials.45–48 Here, as we are only interested in the pre-caging dynamics, we adopt a more straightforward definition. We choose the upper bound of
, with ω = 1 when |rj(t) − rj(0)| < σR and ω = 0 otherwise. Typically, σR is interpreted as the caging length and there is active debate about how to assign a reasonable value to it in the context of glassy materials.45–48 Here, as we are only interested in the pre-caging dynamics, we adopt a more straightforward definition. We choose the upper bound of 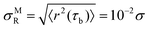 where τb is the ballistic timescale determined from MSD and study the behaviour of
where τb is the ballistic timescale determined from MSD and study the behaviour of ![[scr Q, script letter Q]](https://www.rsc.org/images/entities/char_e530.gif) s(t) for a set of σR ≤ σMR.
s(t) for a set of σR ≤ σMR.
We notice that for both of the aggregate conformations, the early time decay of ![[scr Q, script letter Q]](https://www.rsc.org/images/entities/char_e530.gif) s(t) is Gaussian but their long time decay behaviour is quite different. For compact clusters, this long time decay is best described as a stretched exponential, (Fig. 4A), whereas for string-like clusters, it decays algebraically (Fig. 4B). While the width of the early time peak of Qs(t) would provide us information about the localization time within the specified σR, the departure of the particle from this localization zone and its occasional revisit would contribute to the long time decay of
s(t) is Gaussian but their long time decay behaviour is quite different. For compact clusters, this long time decay is best described as a stretched exponential, (Fig. 4A), whereas for string-like clusters, it decays algebraically (Fig. 4B). While the width of the early time peak of Qs(t) would provide us information about the localization time within the specified σR, the departure of the particle from this localization zone and its occasional revisit would contribute to the long time decay of ![[scr Q, script letter Q]](https://www.rsc.org/images/entities/char_e530.gif) s(t). We could then estimate the residency time τR of a particle within σR in a statistical sense by fitting this long time tail of
s(t). We could then estimate the residency time τR of a particle within σR in a statistical sense by fitting this long time tail of ![[scr Q, script letter Q]](https://www.rsc.org/images/entities/char_e530.gif) s(t). We find that this space–time (σR–τR) relationship is opposite within the two aggregation environments (Fig. 5).
s(t). We find that this space–time (σR–τR) relationship is opposite within the two aggregation environments (Fig. 5).
For the smallest σR considered, the typical residence time τR for particles in string-like clusters is almost equal to one unit of time, τ, which is surprisingly large. As we increase σR, up to a certain value, τR decreases monotonically by almost 4 orders of magnitude before it jumps directly to a much larger value and slowly approaches τb. We relate this curious localization behaviour with the specific form of pair potential for ξ = 0.8 which has a positive local minimum behind a strong positive energy barrier. While the barrier strongly protects any localization at the local minimum, any deviation from that is very unfavorable for the barrier and the discrete jump is indicative of the escape from the barrier. Particles in a compact cluster (ξ = 0.5) are, in comparison, much more mobile as it has very small τR at a very small σR. In this case, the global negative minimum of the potential might be inaccessible due to the positive energy barrier. As with increasing σR, more of the accessible region can be probed and τR is found to increase monotonically. However, note the sharp rise in τR occurring at the same σR as for the ξ = 0.8 case. This supports our previous intuition that this sharp change is related to the crossing of the energy barrier. Note that the behavior of both morphologies at large σR and long τR are very similar. This explains why the statistics of their trajectory geometry, as characterized using 〈![[script L]](https://www.rsc.org/images/entities/char_e144.gif) 〉 and
〉 and ![[scr R, script letter R]](https://www.rsc.org/images/entities/char_e531.gif) g, show no difference. Yet,
g, show no difference. Yet, ![[scr Q, script letter Q]](https://www.rsc.org/images/entities/char_e530.gif) s(t) computed with finer resolution has identified a fundamental difference in the particle dynamics for different morphologies. Putting this together, we argue that σR, probed by
s(t) computed with finer resolution has identified a fundamental difference in the particle dynamics for different morphologies. Putting this together, we argue that σR, probed by ![[scr Q, script letter Q]](https://www.rsc.org/images/entities/char_e530.gif) s(t), reveals an effective reaction coordinate to study the spatio-temporal features of a vast class of aggregate-forming systems.
s(t), reveals an effective reaction coordinate to study the spatio-temporal features of a vast class of aggregate-forming systems.
Conclusions
Overall, this study provides us with a clear correspondence between the observed macroscopic structure, the microscopic dynamics governing it, and most importantly, how the underlying geometric frustration orchestrates this structure–dynamics correspondence. We have demonstrated the role of frustration, arising from competing interaction, by choosing two different realisations of competition between short-range attraction and long-range repulsion. In one case, there is a global minimum at short interparticle distance and a local minimum at long-distance separated by a modest energy barrier, resulting in modest frustration, promoting spatially symmetric particle aggregation of a compact shape. This conformation allows the particles to rattle within the cage of their neighbours, and occasionally jump between the cages, a scenario well-known for glassy systems. This motion is reflected by the intermediate-time slow-relaxation due to frustration. However, the system recovers from this frustration in the long-time showing diffusive motion. In the second, a different dynamical scenario emerges upon spatial inversion of the positions of minima, i.e., the short-distance positive local minimum and long-distance global minimum separated by a strong energy barrier. In this case of pronounced frustration, strong-localisation at short distances is energetically favored, resulting in asymmetric string-like clusters, from which particle detachment is unfavorable and abrupt. Due to the inherent metastable nature of the potential, the system, in this case, continues to be sub-diffusive even in the long time. We emphasise that this case of strong frustration is fundamentally different from the traditional glassy materials where the frustration usually originates from a combination of polydispersity and thermal quenching. Both the causes are cautiously avoided in our study. Noting that the cluster shape in the strong frustration case can be tuned by changing the range of attraction, it might be worth studying the spatio-temporal dynamics in detail for such purely non-equilibrium by design systems.Identification of the remarkable role of particle-interaction level frustration in dictating the structure and dynamics at finite scales was possible by extending the definition of the self-overlap function for finer spatial resolution. This function has been routinely used to elucidate the nature of caged particle dynamics in the context of glassy systems. In fact, recently, it has been concluded that the behaviour of self-overlap doesn't strongly depend on the chosen length-scale of overlap from a detailed examination of three different glass-forming systems.49 However, we note that the reported trend resembles well the large overlap-length behaviour where length- and time-scales are found to have a linear relationship. Exploring the fast-time, short-length nature of the trajectories has revealed new frustration-dependent features. Based on these findings, we argue that such frustration-centric space–time relationships can provide a new generic framework for understanding the dynamical systems far from equilibrium.
On a related note, we discuss the scope of our findings in the context of another commonly used measure of glassy systems, namely, dynamic entropy. Dynamic entropy50 is a thoughtful simplification of Kolmogorov–Sinai entropy that estimates the rate of information growth needed to describe the evolution of a dynamical system.51,52 This is the inverse of the mean first passage time needed by a particle to escape a certain predefined length-scale. Noting the computational similarity with the self-overlap function, the inverse of the time-scales extracted from the self-overlap distributions can be treated as reasonable proxies for dynamic entropy. While establishing a formal relation between this quantity and the bulk response dynamics would require further studies, this view can be useful to understand material information processing. Notably, the self-overlap statistics, being ideally accessible by high-resolution dynamic microscopy, can be informative about the role of frustration in governing the structure–dynamics–function relationship in myriad non-equilibrium situations.
Finally, we conclude by mentioning the immediate applicability of our findings. The fundamental difference in the aggregation process of different morphologies, as revealed by our study, is important for the following two reasons. First, the analysis presented here will significantly decrease the waiting time required for studying the long-time behavior of non-equilibrium systems in order to characterize them. In particular, with the advent of high-throughput high-resolution dynamic microscopy, quantification of appropriate reaction coordinates is now within the reach of direct experimental validation. Furthermore, a deeper understanding of how these different modes of localization are manifested in more dense systems would be beneficial for material design, since different aggregate morphologies are known to have different material response properties. For example, bio-condensates in a healthy cell have a compact spherical shape and are soft and viscoelastic. In contrast, hard, needle-shaped amyloid structures are found in degrading cells, sick with neuro-degenerative diseases.53 This brings us to the second important point. Our approach is aligned with the basic principles of high-resolution dynamic microscopy methods. Adapting our approach to high-speed microscopy techniques would provide us with useful information about the basic principles of shape transformation in biological condensates.54,55 This knowledge would be pivotal in the design of novel therapeutic strategies.56
Conflicts of interest
There are no conflicts to declare.Data availability
All data presented are contained within the manuscript. The codes used for simulation and analysis are available from the author upon reasonable request.Acknowledgements
The authors acknowledge the use of high-performance computing at the OIST Graduate University for this study. M. M. B. was supported by the Nonlinear and Non-equilibrium Physics Unit, OIST Graduate University through subsidy funding from the Cabinet Office of the Prime Minister of Japan. T. D. was supported by the World Premier International Research Center Initiative (WPI), MEXT, Japan and by JSPS Grant Number KAKENHI 24K23179.References
- J. L. Lebowitz and O. Penrose, J. Math. Phys., 1966, 7, 98 CrossRef CAS.
- P. C. Hemmer and G. Stell, Phys. Rev. Lett., 1970, 24, 1284 CrossRef.
- P. Fratzl, J. L. Lebowitz, O. Penrose and J. Amar, Phys. Rev. B: Condens. Matter Mater. Phys., 1991, 44, 4794 CrossRef PubMed.
- C. J. Horowitz, M. A. Perez-Garcia and J. Piekarewicz, Phys. Rev. C: Nucl. Phys., 2004, 69, 045804 CrossRef.
- A. I. Campbell, V. J. Anderson, J. S. van Duijneveldt and P. Bartlett, Phys. Rev. Lett., 2005, 94, 208301 CrossRef PubMed.
- E. Zaccarelli, S. Andreev, F. Sciortino and D. R. Reichman, Phys. Rev. Lett., 2008, 100, 195701 CrossRef PubMed.
- J.-X. Chen, J.-W. Mao, S. Thakur, J.-R. Xu and F.-Y. Liu, J. Chem. Phys., 2011, 135, 094504 Search PubMed.
- D. McDermott, C. J. O. Reichhardt and C. Reichhardt, Soft Matter, 2014, 10, 6332–6338 RSC.
- X. Xu, Z. Wang, X. Xu, G. Fang and M. Gu, J. Chem. Phys., 2020, 152, 054906 CrossRef CAS PubMed.
- A. Jayaraman and K. S. Schweizer, Macromolecules, 2009, 42, 8423–8434 CrossRef CAS.
- J. L. Nugent, S. S. Moganty and L. A. Archer, Adv. Mater., 2010, 3677 Search PubMed.
- J. Fonseca, L. Meng, I. Imaz and D. Maspoch, Chem. Soc. Rev., 2023, 52, 2528–2543 RSC.
- A. Stradner, H. Sedgwick, F. Cardinaux, W. C. Poon, S. U. Egelhaaf and P. Schurtenberger, Nature, 2004, 432, 492–495 Search PubMed.
- T. P. Knowles, M. Vendruscolo and C. M. Dobson, Nat. Rev. Mol. Cell Biol., 2014, 15, 384–396 Search PubMed.
- J. Rouwhorst, C. van Baalen, K. Velikov, M. Habibi, E. van der Linden and P. Schall, npj Sci. Food, 2021, 5, 32 CrossRef PubMed.
- V. J. Anderson and H. N. Lekkerkerker, Nature, 2002, 416, 811–815 CrossRef CAS PubMed.
- K. Kroy, M. Cates and W. Poon, Phys. Rev. Lett., 2004, 92, 148302 CrossRef CAS PubMed.
- P. J. Lu, E. Zaccarelli, F. Ciulla, A. B. Schofield, F. Sciortino and D. A. Weitz, Nature, 2008, 453, 499–503 CrossRef CAS PubMed.
- J. Rouwhorst, C. Ness, S. Stoyanov, A. Zaccone and P. Schall, Nat. Commun., 2020, 11, 3558 CrossRef CAS PubMed.
- M. D. Ediger, Annu. Rev. Phys. Chem., 2000, 51, 99–128 CrossRef CAS PubMed.
- D. J. Wales, Annu. Rev. Phys. Chem., 2018, 69, 401–425 CrossRef CAS PubMed.
- A. Widmer-Cooper, P. Harrowell and H. Fynewever, Phys. Rev. Lett., 2004, 93, 135701 CrossRef PubMed.
- P. M. Reis, R. A. Ingale and M. D. Shattuck, Phys. Rev. Lett., 2007, 98, 188301 CrossRef CAS PubMed.
- E. Zaccarelli and W. C. Poon, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 15203–15208 Search PubMed.
- S. Mazoyer, L. Cipelletti and L. Ramos, Phys. Rev. E: Stat., Nonlinear, Soft Matter Phys., 2009, 79, 011501 CrossRef PubMed.
- D. S. Simmons, M. T. Cicerone, Q. Zhong, M. Tyagi and J. F. Douglas, Soft Matter, 2012, 8, 11455–11461 Search PubMed.
- B. Li, K. Lou, W. Kob and S. Granick, Nature, 2020, 587, 225–229 CrossRef CAS PubMed.
- G. Krishnan and U. Harbola, Int. J. Mod. Phys. B, 2022, 36, 2250065 CrossRef CAS.
- M. Ruiz-Ruiz, A. Vila-Costa, T. Bar, C. Rodrguez-Tinoco, M. Gonzalez-Silveira, J. A. Plaza, J. Alcalá, J. Fraxedas and J. Rodriguez-Viejo, Nat. Phys., 2023, 19, 1509–1515 Search PubMed.
- R. M. Alkemade, F. Smallenburg and L. Filion, J. Chem. Phys., 2023, 158, 134512 Search PubMed.
- A. Widmer-Cooper and P. Harrowell, Phys. Rev. Lett., 2006, 96, 185701 CrossRef PubMed.
- R. Zhang and K. S. Schweizer, J. Chem. Phys., 2010, 133, 104902 Search PubMed.
- D. S. Simmons and J. F. Douglas, Soft Matter, 2011, 7, 11010–11020 RSC.
- A. P. Thompson, H. M. Aktulga, R. Berger, D. S. Bolintineanu, W. M. Brown, P. S. Crozier, P. J. In't Veld, A. Kohlmeyer, S. G. Moore and T. D. Nguyen, et al. , Comput. Phys. Commun., 2022, 271, 108171 CrossRef CAS.
- F. Sciortino, S. Mossa, E. Zaccarelli and P. Tartaglia, Phys. Rev. Lett., 2004, 93, 055701 Search PubMed.
- D. Meng, S. K. Kumar, J. M. D. Lane and G. S. Grest, Soft Matter, 2012, 8, 5002–5010 Search PubMed.
- X. Liu, Y. Ni and L. He, J. Phys.: Condens. Matter, 2016, 28, 445201 CrossRef CAS PubMed.
- A. Denton and H. Löwen, J. Phys.: Condens. Matter, 1997, 9, 8907 Search PubMed.
- Y. Liu and Y. Xi, Curr. Opin. Colloid Interface Sci., 2019, 39, 123–136 Search PubMed.
- T. Das, T. Lookman and M. M. Bandi, Soft Matter, 2015, 11, 6740 Search PubMed.
- F. Sciortino and P. Tartaglia, Adv. Phys., 2005, 54, 471–524 Search PubMed.
- T. Das, T. Lookman and M. M. Bandi, Soft Matter, 2016, 12, 9674 Search PubMed.
- T. Das and M. Bandi, J. Phys.: Condens. Matter, 2020, 32, 214004 Search PubMed.
- E. Ercolini, F. Valle, J. Adamcik, G. Witz, R. Metzler, P. D. L. Rios, J. Roca and G. Dietler, Phys. Rev. Lett., 2007, 98, 058102 CrossRef PubMed.
- R. A. Laviolette and F. H. Stillinger, J. Chem. Phys., 1995, 83, 4079 CrossRef.
- Z. H. Jin, P. Gumbsch, K. Lu and E. Ma, Phys. Rev. Lett., 2001, 87, 055703 Search PubMed.
- N. Lacevic, F. W. Starr, T. B. Schroder and S. C. Glotzer, J. Chem. Phys., 2003, 119, 7372 Search PubMed.
- X. Wang, W.-S. Xu, H. Zhang and J. F. Douglas, J. Chem. Phys., 2019, 151, 184503 CrossRef PubMed.
- P. Pareek, M. Adhikari, C. Dasgupta and S. K. Nandi, J. Chem. Phys., 2023, 159, 174503 Search PubMed.
- P. Allegrini, J. F. Douglas and S. C. Glotzer, Phys. Rev. E: Stat. Phys., Plasmas, Fluids, Relat. Interdiscip. Top., 1999, 60, 5714 CrossRef CAS PubMed.
- K. Koss, O. Petrov, M. Myasnikov, K. Statsenko and M. Vasiliev, J. Exp. Theor. Phys., 2016, 123, 98–107 Search PubMed.
- J. F. Douglas, Q.-L. Yuan, J. Zhang, H. Zhang and W.-S. Xu, Soft Matter, 2024, 20, 9140–9160 Search PubMed.
- T. W. Han, B. Portz, R. A. Young, A. Boija and I. A. Klein, Cell Chem. Biol., 2024, 31, 1593–1609 Search PubMed.
- M. V. Garabedian, W. Wang, J. B. Dabdoub, M. Tong, R. M. Caldwell, W. Benman, B. S. Schuster, A. Deiters and M. C. Good, Nat. Chem. Biol., 2021, 17, 998–1007 CrossRef CAS PubMed.
- Z.-G. Qian, S.-C. Huang and X.-X. Xia, Nat. Chem. Biol., 2022, 18, 1330–1340 Search PubMed.
- Y. Li and S. Sun, EMBO J., 2025, 44, 613–638 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |





