Open Access Article
Open Access ArticleBiophysical characterization of spCas9 binding and cleavage using real-time electronic biosensors
Deependra Kumar
Ban†
a,
Kshama
Parate†
 a,
Deepta
Bharadwaj
a,
Austin
Wong
a,
Lorelai
Schoch
a,
Deepta
Bharadwaj
a,
Austin
Wong
a,
Lorelai
Schoch
 a,
Kenneth
Visk
a and
Kiana
Aran
a,
Kenneth
Visk
a and
Kiana
Aran
 *ab
*ab
aShu Chien-Gene Lay Department of Bioengineering, Jacobs School of Engineering Department of Bioengineering, University of California, San Diego, CA, USA 92093. E-mail: karan@ucsd.edu
bDepartment of Medicine, Division of Geriatrics, Gerontology & Palliative Care, School of Medicine, University of California, San Diego, CA, USA 92093
First published on 9th March 2026
Abstract
CRISPR–Cas9 enables curative genome editing but requires precise control of target recognition, particularly when single-nucleotide polymorphisms (SNPs) influence specificity. Conventional biochemical and optical assays often rely on endpoint or ensemble-averaged measurements and therefore fail to resolve the real-time binding dynamics underlying off-target interactions. Here, we report a label-free, non-faradaic electrochemical impedance spectroscopy (nfEIS) platform that directly monitors spCas9–gRNA interactions on gold microelectrodes with single-base resolution at the sickle cell disease (SCD) locus. A guide RNA was designed to perfectly match the SCD mutation (A to T) while introducing a single PAM-proximal mismatch with the wild-type DNA (WD) sequence. Using 63-nucleotide synthetic DNA substrates representing SCD and WD targets, concentration-dependent binding assays were performed to extract equilibrium parameters. Hill-model analysis revealed higher affinity for the SCD target (kD = 0.09 nM) relative to WD (kD = 0.3 nM), confirming strong on-target binding and weakened interaction at the mismatch site. Magnesium dependence evaluation showed that 5 mM Mg2+ enhanced discrimination by stabilizing on-target complexes while destabilizing mismatched binding, whereas at 1 mM Mg2+ this selectivity was lost. Time-resolved kinetic measurements using 1 nM spCas9 and exponential fitting of the curve revealed rapid association (t1/2 = 1.85 min) and dissociation rates (t1/2 = 5.24 min) for SCD, consistent with efficient R-loop formation. In contrast, the WD target exhibited slower association (t1/2 = 2.68 min) and recurring transient binding with delayed dissociation (t1/2 = 34.38 min), corroborated by endpoint gel assays. Cas9 lacking gRNA showed only weak, unstable interactions. Overall, these results demonstrate that Cas9 specificity arises from both affinity differences and binding-residence dynamics. nfEIS thus provides a real-time, label-free platform for probing Cas9 fidelity, Mg2+-dependent activation, and gRNA design for therapeutic genome editing and diagnostics.
Introduction
Since the advent of CRISPR–Cas9, a Nobel Prize winning technology, genome editing has shifted from concept to clinic, offering realistic paths to treat previously intractable genetic diseases.1,2 The precision of this system, however, depends on a finely coordinated sequence of molecular events at the DNA target site, including PAM engagement, seed-proximal R-loop nucleation, stepwise RNA:DNA hybrid propagation, and Mg2+-dependent activation of the HNH and RuvC nuclease domains. Because each of these steps is energetically coupled, even a single-nucleotide polymorphism (SNP) within the target sequence can perturb the overall Cas9 energy landscape by altering its binding residence time and the likelihood of productive cleavage.3–6 Therefore, a mechanistic understanding of these energy barriers is essential for optimizing on-target efficiency while minimizing off-target effects. Moreover, recent studies have identified the residence time of the Cas9–DNA complex as a critical kinetic determinant that dictates whether a binding event proceeds to cleavage or results in premature dissociation, thereby governing both the fidelity and efficiency of genome editing.7,8 Biochemical and single-molecule analyses have revealed that Cas9 interrogates its target through a PAM-first recognition mechanism, followed by directional R-loop propagation along the protospacer sequence. During this process, Mg2+ ions play a dual role: stabilizing the catalytically competent conformation and gating the transition from target binding to cleavage.4,9 Despite these advances, it remains unclear how these sequential molecular events manifest as time-resolved binding signatures after binding with the DNA and how cofactors such as Mg2+ influence the discrimination between perfectly matched and mismatched targets after binding with a specific DNA sequence. Bridging this gap requires quantitative approaches that can directly capture the dynamics of Cas9–gRNA–DNA interactions in real time and under native electrochemical environments.In this regard, optical and structural methods such as fluorescence labeling, Förster resonance energy transfer (FRET), and cryo-EM have mapped key conformational states of Cas9 and elucidated aspects of its target search and cleavage cycle. However, these approaches typically rely on fluorescent tags, specialized instrumentation, or endpoint imaging, which complicate the direct observation of real-time, surface-bound kinetics10–12 (Table S1). By contrast, electronic biosensing provides a means to circumvent these limitations by converting molecular binding events into electrical signals without the need for optical labels or signal amplification. For example, graphene field-effect transistor (gFET) platforms, including CRISPR-chip, have established label-free, amplification-free detection of genomic targets and even single-nucleotide polymorphisms (SNPs).13,14 Yet, most of these demonstrations focused on specific sequence (i.e., target vs. non-target and SNP) detection, offering limited information about the continuous dynamics of biomolecular interaction at the interface. Therefore, there remains a critical need to develop label-free, real-time methodologies capable of quantitatively resolving the binding dynamics and conformational transitions of spCas9 and its cofactors.
To address these challenges, we utilized mediator-free, non-faradaic electrochemical impedance spectroscopy (nfEIS) as a real-time, label-free analytical technique to resolve the binding dynamics (e.g., binding affinity, effect of Mg2+, and sequence specific binding kinetics) of spCas9–gRNA complexes on DNA-functionalized gold microelectrodes. In this method, frequency-resolved impedance spectra capture subtle variations in the electrical double layer and in the dielectric and ionic properties of the interfacial biolayer, thereby encoding signatures of binding, conformational transitions, and molecular reorganization.15 These interfacial processes were modeled using minimal equivalent circuits (e.g., R1 − (CPEdl//Rp)), which allowed a quantitative extraction of apparent affinity, cooperativity, and relaxation kinetics directly from electrical data.15,16 Because nfEIS operates at low excitation potentials over a broad frequency range, it minimally perturbs the intrinsic spCas9–DNA interactions while sensitively detecting charge redistribution and biolayer evolution at the electrode interface.17 Due to its advantages, nfEIS has been widely applied to interrogate biological interactions at electrode interfaces, owing to its sensitivity to changes in interfacial capacitance, dielectric properties, and charge distribution arising from biomolecular binding. Prior studies have employed nfEIS to investigate protein–DNA interactions, including transcription factor binding and sequence-specific nucleic acid recognition, as well as antibody–antigen interactions and enzymatic processes occurring at functionalized electrode surfaces.18–22 nfEIS has been used to quantify binding affinities and kinetic parameters in DNA hybridization assays, to resolve antibody–antigen recognition without labels, and to monitor conformational rearrangements associated with enzyme–substrate complex formation.19–23 In addition, impedance-based approaches have been applied to study cellular adhesion, membrane remodeling, and protein adsorption dynamics, demonstrating the technique's versatility in probing interfacial biological phenomena under physiologically relevant conditions.24–26 Despite these advances, to the best of our knowledge, nfEIS has not previously been applied to study CRISPR–Cas systems or to resolve the real-time binding dynamics and sequence discrimination behavior of Cas nucleases. The present work therefore extends the application space of nfEIS to CRISPR-based genome editing, enabling direct, label-free interrogation of Cas9–DNA interactions and their modulation by cofactors such as Mg2+. Using specific DNA sequence corresponding to the sickle-cell mutation and its wild-type counterpart, which differ by a single base in the PAM-proximal seed region,13 we systematically examined spCas9 concentration-dependent and time-resolved binding behavior under varying Mg2+ concentrations. This unified framework enabled the direct derivation of affinity constants, cooperativity factors, and kinetic parameters, linking molecular-scale binding dynamics to electrical readouts in real time. Collectively, these results establish nfEIS as a mechanistically important platform for analyzing residence-time dependent target sequence recognition and guiding the design of high-fidelity CRISPR systems for genome-editing and diagnostic applications.
Experimental
Materials
The acrylic substrate (0.7 mm thickness) for gold electrode fabrication was purchased from Olycraft. All the reagents were prepared in UltraPure™ DNase/RNase free distilled water (Invitrogen, Waltham, MA, USA). Custom dsDNA oligonucleotides with 5′ disulfide modification (ThiolMC6-D) were synthesized by IDT technologies (Coralville, IA, USA). The 63-nucleotide length dsDNA includes a sickle cell disease specific single point mutation sequence (Hereafter called SCD) and wild type DNA sequence (WD) (see SI Table S2).13 SCD specific guide RNA for Streptococcus pyogenes-CRISPR associated protein (spCas9) was custom synthesized from Synthego. spCas9 was purchased from Aldevron (9212-0.25MG). DNA immobilization buffer was prepared with a final concentration of 5.05 mM Tris-HCl, 0.505 M KH2PO4 and 0.25 mM TCEP. TCEP was used for reduction of the disulfide bond on the custom oligos. spCas9 reaction buffer was prepared with 200 μM Tris-HCl, 50 μM MgCl2 (pH 7.5).Sensor fabrication
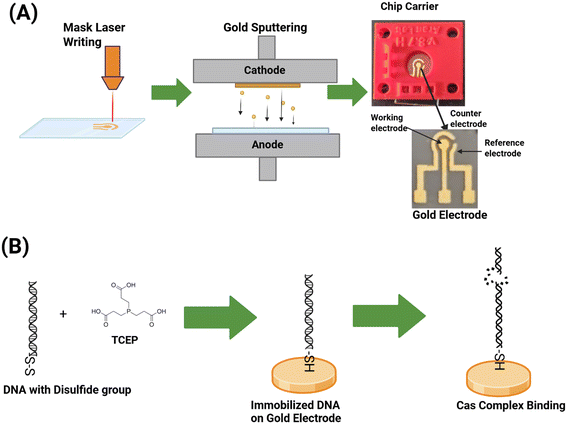
First, masking tape was fixed to an acrylic sheet and air bubbles were removed by squeezing. A three electrode impedimetric sensor was fabricated using Glowforge Plus (40 W CO2 laser cutter and engraver) with a speed setting of 450 (∼42.5 mm per second) and a power setting of 15 (∼6 Watts).After laser cutting and peeling the tape mask to expose the inner pattern, a 10 nm chromium metal glue layer and 100 nm gold metal were sputter coated27 using a Denton Discovery 635 DC sputter coater in an Ar gas environment at the process power of 200 W and the process pressure of 2.5 mTorr. The remaining masking tape was peeled off to reveal the electrode pattern. Individual chips were cut from the acrylic sheet using a Glowforge Pro laser cutter (speed: 300, power: 100) and cleaned for EIS measurement.
Surface cleaning and baseline characterization of the sensor
All chips were rinsed five times using ultrapure water, sonicated in ethanol (5 min), rinsed again (3×), and dried for further use. Initial EIS spectra were recorded in ultrapure water and then in 0.2 mM Tris-HCl (0.01×) to verify baseline stability during medium exchange. Only devices exhibiting stable baselines and intact continuity proceeded to functionalization (Fig. S1).dsDNA immobilization and surface passivation
The dsDNA (10 μM) (target, SCD or control, WD) was incubated with 0.25 mM tris(2-carboxyethyl)phosphine (TCEP) for 30 min at room temperature to reduce disulfide bonds to produce the thiol group (Fig. 2B), as custom synthesized dsDNA from IDT contains a disulfide bond (i.e., S–S) to protect the free thiol group. A 35 μL droplet of the dsDNA with the freshly generated thiol group was placed on the top of the gold working electrode and incubated for 1 h to immobilize the DNA. To confirm dsDNA layer formation, EIS measurements were performed by applying 10 mV of DC voltage between the working electrode (WE) and reference electrode (RE) with an AC frequency range of 100 Hz to 200 kHz while collecting current between the WE and counter electrode (CE) using 0.01× Tris-HCl buffer. Following DNA immobilization, L-cysteine (1 mM) was used to passivate exposed gold working electrodes, excess analyte was rinsed using 0.01× Tris-HCl buffer and passivation mediated non-faradaic impedance was measured.spCas9 and gRNA complex preparation and activity assay
To evaluate the effect of varying spCas9–gRNA concentrations (0, 0.1, 1, and 10 nM) on binding dynamics, the spCas9–gRNA complex was prepared by mixing Cas9 enzyme and guide RNA in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio using 0.01× Tris-HCl buffer containing 5 mM MgCl2. The selected concentration of spCas9 ensures that spCas9 remains in the limiting concentration with respect to dsDNA while allowing sufficient surface occupancy to generate a reproducible nfEIS signal. Moreover, higher spCas9 concentrations (micromolar regime) led to protein crowding and biofouling effects at the electrode surface, producing impedance changes unrelated to diffusion-mediated interfacial interactions with the substrate even after surface passivation. The selected concentration range therefore represents an experimentally optimized compromise that preserves interfacial specificity while maintaining measurement fidelity. The mixture was incubated at 37 °C for 30 minutes to ensure complete ribonucleoprotein complex formation. Following incubation, 35 μL of the spCas9–gRNA complex solution was dispensed onto the DNA-functionalized gold microelectrode and continuous measurement was performed for 45 minutes at room temperature to measure association dynamics of spCas9. After this binding step, the excess solution was gently removed and washed five times, and an equal volume (35 μL) of fresh 0.01× Tris-HCl buffer with 5 mM MgCl2 was added to monitor spCas9 interfacial interaction mediated change in polarization resistance. For kinetic analysis, 1 nM of spCas9–RNA and spCas9 only were exposed to SCD and WD immobilized sensors and continuous non-faradaic impedance measurement was performed for 30 min for association response. After 30 minutes of association measurement, the excess sample was removed to prevent contributions from bulk solution and loosely bound interfacial interactions. Following removal of bulk spCas9, dissociation of spCas9 from the dsDNA–gold electrode interface was continuously monitored for 1 hour using measurement buffer (Fig. S3). All impedance measurements were conducted using a Hioki impedance analyzer under an applied DC bias of 10 mV and a frequency sweep from 100 Hz to 200 kHz. Real impedance (Z), imaginary impedance (Z″), parallel capacitance, and phase angle values were continuously recorded using the instrument's software. Nyquist plots were generated by plotting the imaginary impedance (Z″) against the real impedance (Z) to evaluate changes in polarization characteristics. The interfacial polarization resistance (Rp) in non-faradaic EIS arising from ion interactions, biomolecular binding, and charge realignment at the working electrode is calculated by subtracting the solution resistance (Rsol) from the measured interfacial resistance (Rint).17,28 This difference reflects the net change in interfacial polarization resistance (Rp), which was used to analyze the spCas9 binding, Mg ion effect, and spCas9 binding kinetics. Furthermore, we have also assessed the influence of magnesium ion concentration, and parallel experiments were performed under identical conditions using 1 mM MgCl2, and the resulting impedance spectra were analyzed following the same procedure.
1 molar ratio using 0.01× Tris-HCl buffer containing 5 mM MgCl2. The selected concentration of spCas9 ensures that spCas9 remains in the limiting concentration with respect to dsDNA while allowing sufficient surface occupancy to generate a reproducible nfEIS signal. Moreover, higher spCas9 concentrations (micromolar regime) led to protein crowding and biofouling effects at the electrode surface, producing impedance changes unrelated to diffusion-mediated interfacial interactions with the substrate even after surface passivation. The selected concentration range therefore represents an experimentally optimized compromise that preserves interfacial specificity while maintaining measurement fidelity. The mixture was incubated at 37 °C for 30 minutes to ensure complete ribonucleoprotein complex formation. Following incubation, 35 μL of the spCas9–gRNA complex solution was dispensed onto the DNA-functionalized gold microelectrode and continuous measurement was performed for 45 minutes at room temperature to measure association dynamics of spCas9. After this binding step, the excess solution was gently removed and washed five times, and an equal volume (35 μL) of fresh 0.01× Tris-HCl buffer with 5 mM MgCl2 was added to monitor spCas9 interfacial interaction mediated change in polarization resistance. For kinetic analysis, 1 nM of spCas9–RNA and spCas9 only were exposed to SCD and WD immobilized sensors and continuous non-faradaic impedance measurement was performed for 30 min for association response. After 30 minutes of association measurement, the excess sample was removed to prevent contributions from bulk solution and loosely bound interfacial interactions. Following removal of bulk spCas9, dissociation of spCas9 from the dsDNA–gold electrode interface was continuously monitored for 1 hour using measurement buffer (Fig. S3). All impedance measurements were conducted using a Hioki impedance analyzer under an applied DC bias of 10 mV and a frequency sweep from 100 Hz to 200 kHz. Real impedance (Z), imaginary impedance (Z″), parallel capacitance, and phase angle values were continuously recorded using the instrument's software. Nyquist plots were generated by plotting the imaginary impedance (Z″) against the real impedance (Z) to evaluate changes in polarization characteristics. The interfacial polarization resistance (Rp) in non-faradaic EIS arising from ion interactions, biomolecular binding, and charge realignment at the working electrode is calculated by subtracting the solution resistance (Rsol) from the measured interfacial resistance (Rint).17,28 This difference reflects the net change in interfacial polarization resistance (Rp), which was used to analyze the spCas9 binding, Mg ion effect, and spCas9 binding kinetics. Furthermore, we have also assessed the influence of magnesium ion concentration, and parallel experiments were performed under identical conditions using 1 mM MgCl2, and the resulting impedance spectra were analyzed following the same procedure.
Quality control of the chip
All chips were subjected to an initial quality-control (QC) screening prior to surface functionalization. QC was based on the impedance response during buffer exchange from deionized water to 0.01× Tris buffer, which probes the cleanliness, conductivity, and electrochemical stability of the gold working electrodes. Chips exhibiting at least an 80% decrease in interfacial polarization resistance (Rp) under these conditions were considered to have clean and conductive electrodes and were retained for subsequent surface modification steps (see representative response in Fig. S1). Following dsDNA immobilization and cysteine-based surface passivation, chips were further validated by comparing their Rp values to buffer-only control chips processed in parallel. Only chips showing a statistically significant change in polarization resistance relative to controls were included in downstream binding, kinetic, and Mg2+-dependence measurements (Fig. 3B and C). For all experimental conditions, data were collected from a minimum of three independently fabricated and QC-qualified chips, and results are reported as mean ± standard deviation to capture inter-chip variability. Moreover, inter-chip variability in sensor response was quantified by calculating the coefficient of variation (CV) (n = 58), defined as the standard deviation of ΔRp divided by the mean ΔRp. Here, ΔRp corresponds to the change in polarization resistance measured during buffer exchange from deionized water to 0.01× Tris-HCl buffer. These criteria ensured consistent baseline electrochemical behavior across devices while minimizing variability introduced by fabrication defects.Interaction Analysis
Nyquist plots for the spCas reaction and wash steps were used to extract parameters to analyze the data, such as the polarization resistance (Rp) deduced by fitting the data to a Randles circuit using a custom code developed in Python (SI). To compare the different treatment mediated changes in Rp value, the relative shift in Rp value was plotted against the concentration and time. This relative change analysis led to a positive shift (i.e. Rp increasing with respect to baseline) and a negative shift (i.e. decrease in Rp value but not representing negative resistance) with treatment. These values were then plotted together with the on-target, DNA sequences with SNP, and spCas9 only control for comparison. Time-dependent Rp values were obtained for spCas9-gRNA complexes with SCD and WD targets, as well as for spCas9 alone (without gRNA) with SCD. The data were fitted using the Hill equation (see below) to determine the dissociation constant (kD) and the Hill coefficient (h). A value of h > 1 indicates cooperative binding, whereas h < 1 suggests non-cooperative or unstable interactions. This analysis was used to evaluate the specificity of spCas9 binding to the target DNA.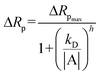 | (1) |
Furthermore, we have used GraphPad prism to fit time dependent change in mean values of Rp resistance using exponential function and separately fitted the curve for the association and dissociation phase to calculate the association rate constant (K (min−1)), time constant (τ = 1/K), half-life (t1/2 = ln(2)/K) for initial spCas9 or spCas9–gRNA complex binding and subsequent dissociation phase to further investigate the binding mechanism:
| Rp(t) = AeKt + C | (2) |
Results and discussion
Sensor quality control and surface functionalization
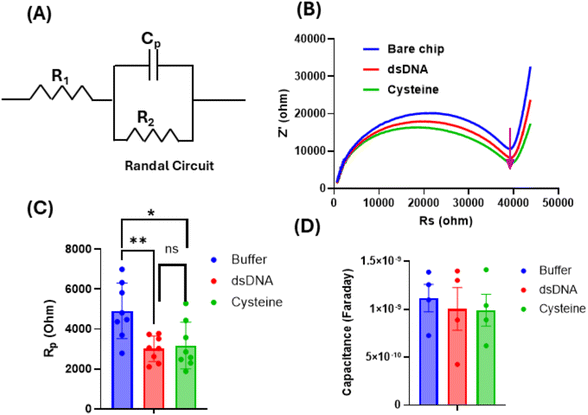
All sensors underwent quality control prior to surface functionalization. Each chip was visually inspected for defects, verified for electrode continuity, and assessed for baseline stability during the medium exchange from deionized water to the measurement buffer. Only devices exhibiting a robust, reproducible decrease of at least 80% in polarization resistance (Rp) during DI to 0.01× Tris buffer transition were selected for dsDNA and cysteine conjugation steps (Fig. 3). Furthermore, inter-chip variability was quantified by calculating the coefficient of variation (CV) from the change in polarization resistance (ΔRp) measured during the deionized water to 0.01× Tris-HCl buffer exchange. Across the qualified chips, this analysis yielded a CV of ∼26% (n = 58), reflecting moderate fabrication-dependent variability typical of microfabricated, surface-sensitive electrochemical sensors. Importantly, the applied quality-control and selection criteria ensured that this variability did not compromise the reproducibility of downstream nfEIS measurements. Furthermore, chips were also measured for double layer/biolayer polarization mediated capacitance by fitting a Randel circuit (Fig. 3A) using Hioki commercial software. The polarization resistance R2 (i.e., Rp) decreased significantly after dsDNA attachment and remained lower after cysteine blocking (n = 3), visible as a smaller semicircle in the Nyquist plot (Fig. 3B and C). This reduction is attributed to electrostatic interactions of the polyanionic phosphate backbone of DNA and carboxylic acid/amine group of the cysteine, which enhances local ionic polarization/screening at the interface and thereby facilitates interfacial charge accommodation, even in the absence of a redox mediator.In contrast, parameter trends show that dsDNA immobilization followed by L-cysteine passivation did not show a measurable change in Cp relative to the bare-chip buffer baseline, consistent with a thin, compact molecular layer formation that minimally perturbs the effective double layer (Fig. 3D).
Effect of MgCl2 and Cas9 concentration on the Cas9 binding with the target
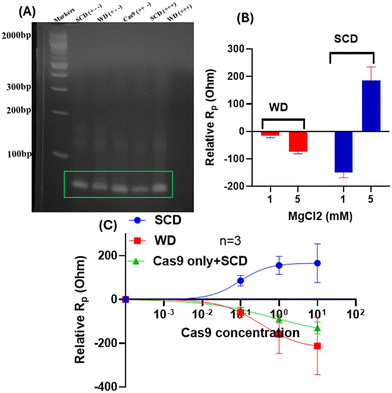
Prior to impedance measurements, we verified sequence selectivity with a gel-shift assay (Fig. 4A and S2). Ethidium-bromide-stained gels imaged under UV illumination showed efficient cleavage of the perfectly matched sickle-cell disease (SCD) substrate by the spCas9–gRNA ribonucleoprotein (RNP), as evidenced by a pronounced reduction in the full-length double-stranded DNA band relative to control lanes (Fig. S2). In contrast, reactions containing the seed-mismatched wild-type (WD) and non-target dsDNA (RD) showed little to no cleavage under identical conditions. Similarly, the SCD substrate incubated with spCas9 complexed with non-specific gRNA (RgRNA) samples also retained intense, well-defined full-length bands (Fig. 4A and S2). These results provide direct biochemical evidence that the spCas9–gRNA RNP selectively recognizes and cleaves the SCD target while sparing the WD sequence, demonstrating single-nucleotide discrimination under the experimental conditions. Semi-quantitative densitometric analysis performed using Fiji (ImageJ) gel analysis macros revealed that the band intensity of the SCD substrate was reduced by approximately 33–77% relative to the WD substrate across three independent agarose gel experiments (Fig. S2A–C). These gel-based measurements consistently confirm the high specificity of spCas9 when paired with a perfectly matched guide RNA. However, gel-shift or cleavage assays offer only limited insight into the biophysical characteristics of Cas9–gRNA–DNA interactions in practice. Specifically, such assays do not resolve crucial kinetic parameters such as the binding affinity (kD) of the spCas9–gRNA complex for its target sequence, the rate of association/cleavage, or whether binding proceeds via a cooperative mechanism. Likewise, gel-electrophoresis lacks the temporal resolution to detect subtle but functionally important events such as spCas9 binding mediated changes in charge redistribution that may occur before cleavage. In real-world genome-editing applications, these kinetic and biophysical features are critical because a complex that binds strongly but dissociates slowly, even without cleavage, may hinder enzyme turnover, reduce editing efficiency, or elevate off-target risk. Previous studies have shown that tight binding and extended residence times of spCas9 on DNA can interfere with repair processes and significantly influence editing outcomes.1–3,30 Therefore, relying solely on gel-based readouts may overestimate or underestimate true editing efficiency or mismatch discrimination by spCas9, especially within the more complex milieu of a living cell. It follows that for practical therapeutic genome editing and specific diagnostics in biological fluids, it is ideal to select spCas9–gRNA complexes that either do not bind non-target sequences at all, or if they do bind, have very short residence times to minimize unwanted engagements and improve fidelity.To overcome these limitations and gain a deeper understanding of the binding affinity, cooperativity coefficient, and the Mg2+-dependent stabilization of the spCas9–gRNA–DNA complex, we employed non-faradaic electrochemical impedance spectroscopy (EIS), a highly sensitive, label-free technique capable of probing molecular interactions at the sensor–electrolyte interface in real time. This mediator-free approach allows direct observation of dielectric and polarization resistance changes associated with complex formation, charge redistribution, and conformational transitions, offering a powerful means to evaluate the biophysical properties of Cas9-mediated target recognition.
It has been reported that magnesium ions and their concentration play a critical role in stabilizing CRISPR–Cas9–DNA complexes and facilitating target recognition.31 Therefore, intracellular free Mg2+ concentrations, depending on cell type and physiological conditions, become an important factor for gene editing applications.32,33 Accordingly, we first examined target recognition at 1 mM of Mg2+, a concentration that lies within the upper range of physiologically relevant levels (e.g., typical range 0.2 1 mM in eukaryotes), using mediator-free, non-faradaic electrochemical impedance spectroscopy (nfEIS) on electrodes functionalized with SCD (specific complementary DNA) and WD (wobble/mismatch DNA) sequences.13 With the RNP complex (spCas9–gRNA) held constant at 1 nM, increasing the MgCl2 concentration from 1 mM to 5 mM resulted in markedly different electrochemical responses for the two target types (Fig. 4B). On the SCD-functionalized electrodes, elevated Mg2+ concentration induced a relative positive shift in polarization resistance (Rp), consistent with enhanced stabilization and compaction of the spCas9–gRNA–DNA interfacial complex. In contrast, WD-functionalized electrodes exhibited a distinct and opposing impedance response, indicating that Mg2+ does not equivalently stabilize mismatched R-loop configurations. These divergent behaviors are attributed to mismatch-induced differences in R-loop stability and interfacial interaction dynamics. The reduced discrimination observed at 1 mM Mg2+ in our nfEIS measurements therefore likely reflects a biologically relevant trade-off between binding efficiency and target specificity under mammalian intracellular conditions rather than a limitation of the sensing platform itself. Indeed, a prior study also demonstrated that decreasing Mg2+ from 5 mM to ≥1 mM led to an approximately 17-fold reduction in Cas9 cleavage activity.34 Taken together, our results suggest that nfEIS is sensitive to precisely those ionic effects that govern spCas9 specificity in vivo. Intermediate Mg2+ concentrations (e.g., 0.5–2 mM), which bridge mammalian and bacterial regimes, are therefore expected to produce a gradual transition between high-specificity/low-activity and high-activity/low-specificity states, consistent with established spCas9 kinetic models11,35 (Fig. 4B).
Collectively, these observations suggest that Mg2+ modulates the electrostatic and structural environment of the spCas9–gRNA–DNA complex in a manner that enhances on-target stabilization while limits nonspecific or mismatched binding. Such a finding is particularly significant for eukaryotic gene-editing applications, where the intracellular free Mg2+ ion concentration is often lower than the optimal range (5 mM or more) required for maximal spCas9 activity, in contrast to the higher Mg2+ ion levels present in prokaryotic systems, where spCas9 originally evolved.34 This difference in ionic environment could therefore influence spCas9 target binding efficiency and specificity in mammalian cells. Consistent with the interpretation by Egger et al.34 demonstrating that reduced Mg2+ concentration from 5 mM to ≥1 mM led to an approximately 17-fold decrease in Cas9 (e.g., Geobacillus stearothermophilus Cas9 (GeoCas9)) activity, showing the ion's dual role in catalysis and target discrimination.
Furthermore, we investigated the role of varying concentrations of the spCas9–gRNA complex in the presence of 5 mM MgCl2 to understand the concentration-dependent binding dynamics on the electrode surface. The relative shift in non-faradaic polarization resistance (ΔRp) was measured after incubating the sensor with 0–10 nM spCas9–gRNA complex for 45 minutes (Fig. 4C). After incubation, the electrode surface was rinsed with a measurement buffer to remove loosely bound analytes and then subjected to impedance analysis. The resulting nfEIS spectra revealed a clear concentration-dependent increase in polarization resistance for the SCD-functionalized surface, indicating progressive stabilization of the Cas9–gRNA–DNA complex with increasing RNP concentration. Fitting the ΔRp data to the Hill equation yielded an apparent binding affinity (kD) of 0.09 nM and a Hill coefficient (h) of 1.14, suggesting strong binding and slightly positive cooperativity. In contrast, measurements on WD (mismatch DNA) and the Cas9-only (no gRNA) controls with SCD showed substantially weaker and less specific interactions, with an apparent kD more than three times larger than spCas9–gRNA-SCD and h < 1, indicative of non-cooperative or negatively cooperative binding behavior (Fig. 4C).
The observed sub-nanomolar affinity (kD ≈ 0.09 nM) for the spCas9–gRNA complex on the SCD surface underscores the tight and specific association between the ribonucleoprotein and its perfectly matched target sequence. The sub-nanomolar kD observed for SCD agrees with prior biophysical studies reporting tight Cas9–gRNA binding in the 0.1–1 nM range under similar buffer conditions.1,36,37 Moreover, the affinity values and cooperative behavior reflect the multi-step nature of Cas9 target engagement, which involves coordinated transitions such as PAM recognition, DNA bending, and R-loop propagation rather than a single-step interaction. The slightly positive cooperativity observed here is consistent with models in which early conformational transitions facilitate subsequent stabilization of the Cas9–gRNA–DNA complex.38,39
In contrast, we observed weaker binding and non-cooperative behavior (h < 1) on the WD conjugated gold surface and with spCas9-only-SCD, which contains a single mismatch in the seed region and no gRNA, indicating impaired R-loop propagation and unstable ternary complex formation, hindering the formation of the RNA–DNA hybrid and altering conformational activation pathways.11,36 The decrease in cooperativity (h < 1) suggests that once partial or mismatched binding occurs, subsequent molecular rearrangements are less favorable, potentially leading to non-productive binding.
Overall, the non-faradaic impedance approach provided an electrochemical fingerprint of these molecular interactions, reflecting changes in interfacial charge distribution that accompany Cas9 binding, R-loop stabilization, or mismatch-induced dissociation. The increase in Rp observed for the SCD-modified surface reflects the formation of a more compact, tightly bound, and less interfacial charge layer, consistent with stable Cas9–gRNA–DNA complex formation. In contrast, the smaller Rp shift measured for the WD surface suggests higher ionic mobility and a more permeable interface due to weaker and less stable binding. These findings are in agreement with the model proposed by Aldag et al., which indicates that R-loop stability determines the number and persistence of Cas9–DNA complexes on fully matched targets, thereby facilitating efficient spCas9 cleavage through stronger interactions with the DNA substrate.40 To further validate this interpretation, DNase I was included as a positive control for nonspecific DNA-binding and enzymatic cleavage activity. In contrast to the weak and transient responses observed under spCas9-only conditions, DNase I produced a pronounced nfEIS signal, with approximately a two-fold decrease in polarization resistance (Rp) observed at enzyme concentrations of 0.01 U mL−1 and 1 U mL−1 (n = 2). This distinct response reflects DNA binding protein interaction and clearly differentiates nonspecific protein–DNA binding, thereby confirming the platform's ability to resolve non-specific versus sequence specific spCas9 interfacial dynamics (Fig. S1C and D).
Analyzing kinetics of Cas9 activity
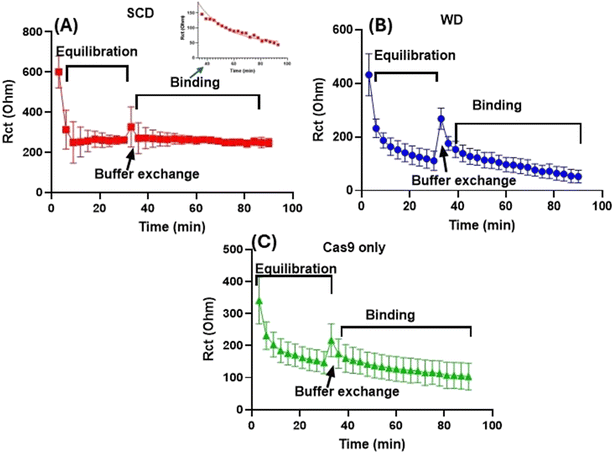
We next analyzed the dynamic behavior of spCas9 on surface-immobilized DNA using nfEIS by separating the association and dissociation rate and fitting each independently with a single-exponential model. The association phase was recorded during the first 30 minutes after introducing 1 nM spCas9–gRNA or spCas9-only. Following the initial incubation period, unbound or weakly adsorbed protein (i.e., spCas9–gRNA/spCas9) was removed to minimize nonspecific contributions. The subsequent evolution of interfacial reorganization and stabilization of protein was monitored for 60 minutes (Fig. 5, Table 1). This two-step approach allowed us to independently quantify the rate of target recognition and early complex formation and the stability or residence time of the bound spCas9 complexes after removal of free protein (see the Experimental section).| S. No. | spCas9–gRNA-SCD | spCas9–gRNA-WD | spCas9-SCD | |||
|---|---|---|---|---|---|---|
| Association | Dissociation | Association | Dissociation | Association | Dissociation | |
| K (min−1) | 0.374 | 0.132 | 0.25 | 0.02 | 0.19 | 0.005 |
| τ (min) | 2.67 | 7.56 | 3.85 | 49.66 | 5.25 | 18.84 |
| Half-life (t1/2) | 1.85 | 5.24 | 2.68 | 34.38 | 3.64 | Unstable |
For the perfect-match SCD sequence, the association phase displayed the fastest kinetics among all tested conditions (K1 = 0.374 min−1, time constant (τ) = 2.67 min, half-life = 1.85 min), indicating rapid on-target recognition due to perfect match with gRNA. This rapid change in interfacial impedance indicates efficient PAM recognition and R-loop initiation on a fully matched target. In the interfacial interaction rate analysis (i.e., dissociation/desorption), the SCD complex relaxed with a moderate rate (K2 = 0.132 min−1, time constant (τ) = 7.56 min, half-life = 5.24 min), suggesting that the spCas9–gRNA–DNA complex persists long enough to undergo catalytic activation and change in structured R-loop but does not remain bound for extended periods on the surface bound DNA. When comparing nfEIS methods with single-molecule studies such as smFRET and surface based methods such as surface plasmon resonance (SPR), it was revealed that the residence time for Cas9 ranges from min to hours depending on complementarity, DNA length, and interaction forces.4,8,11,36,41–43 Moreover, the residence times measured for SCD in this study were shorter than those observed in optical-tweezers, SPR and smFRET studies. However, this difference is expected because these single-molecule and surface based assays typically used long DNA substrates (hundreds to thousands of base pairs), which allow Cas9 to undergo target search, 1D sliding, hopping, and repeated rebinding processes known to significantly extend observable dwell times.36,44,45 In contrast, our sensor uses a short (63-nt), immobilized duplex, which eliminates search dynamics and isolates spCas9 activity to only local binding, R-loop formation, and dissociation. Therefore, the apparent shorter residence times measured here more accurately reflect the true catalytic lifetime of spCas9 once it has already located its target DNA sequence.
In contrast, the single-mismatch WD sequence showed a slower association rate (K1 = 0.25 min−1, time constant (τ) = 3.85 min, half-life = 2.68 min), indicating that the mismatch reduces the efficiency of early R-loop formation (Fig. 5B, Table 1). More importantly, the cleavage rate analysis revealed a very prolonged relaxation (K2 = 0.02 min−1, time constant (τ) = 49.66 min, and half-life = 34.38 min). This extended residence time is consistent with a mismatch-stabilized Cas9–DNA complex that binds tightly but fails to activate the HNH nuclease domain. In this regard, previous smFRET, cryo-EM, and biochemical studies have shown that PAM-proximal mismatches can trap Cas9 in a linear or partially zipped R-loop that prevents catalytic alignment while still maintaining strong DNA binding.4,45,46 Therefore, the WD sequence acts as a kinetic trap, producing a transient recurring binding but non-productive complex that remains on the surface more than six times longer than the perfectly matched SCD target.3
Moreover, spCas9 without gRNA showed the weakest and least stable interactions across both phases. The association rate (K1 = 0.19 min−1, time constant (τ) = 5.25 min, half-life = 3.64 min) was slower than both SCD and WD, and the effective lifetime of interfacial interaction rate (K1 = 0.005 min−1, time constant (τ) = 18.84 min) indicated an intermediate lifetime. These values suggest that apo-Cas9 binds largely through nonspecific electrostatic interactions with surface DNA rather than structured R-loop formation. Therefore, Cas9 lacking gRNA might adopt a conformation that is incompatible with productive PAM recognition and unable to form stable R-loops,47 Therefore, spCas9-only serves as a baseline for nonspecific protein–DNA contacts (Fig. 5C, Table 1).
These results show that fitting the association and dissociation phases separately with exponential models provides a clear and intuitive view of spCas9 binding dynamics. The perfect-match SCD target, which has the strongest equilibrium affinity (kD = 0.09 nM), exhibits fast recognition and a moderate residence time, consistent with efficient R-loop formation and dissociation. In contrast, the WD mismatch, despite having only a modestly weaker affinity (kD = 0.3 nM), shows slower initial binding and a dramatically longer residence time. This behavior indicates that the mismatch traps spCas9 in a recurring transient and cleavage-incompetent state, where the enzyme binds but cannot complete HNH activation or catalysis. spCas9-only conditions show weak and short-lived interactions, confirming that gRNA is required for specific and stable target engagement. Therefore, these kinetic and affinity measurements together reveal that Cas9 able to discriminate between the SCD and WD sequences is governed not only by binding strength (i.e., 3-fold smaller kD for SCD, and differential binding at 5 mM Mg2+) but also by large differences in binding dynamics (Fig. 5), with mismatch sites producing clear difference in Rp values collected using nfEIS.
Overall, nfEIS-based analysis of binding kinetics, residence time, and Mg2+-dependent behavior provided a quantitative framework for understanding Cas9 specificity and sequence discrimination mechanisms. The prolonged residence times observed for mismatched targets are consistent with prior studies showing that PAM-proximal mismatches can kinetically trap Cas9 in cleavage-incompetent conformational states associated with incomplete R-loop formation.11,30,35,36,48 Mismatch position relative to the PAM is therefore a critical determinant of Cas9 behavior: PAM-proximal mismatches, particularly within the seed region, strongly perturb R-loop initiation and complex stabilization, whereas PAM-distal mismatches are often better tolerated and primarily affect cleavage rather than binding affinity.36,49–51 As a result, Cas9 exhibits distinct kinetic and conformational responses depending on both mismatch position and multiplicity, with kinetic trapping near the PAM representing a major contributor to off-target binding in cellular systems.
By enabling direct, quantitative comparison of residence times across different gRNA-target combinations, nfEIS provides an experimental basis for identifying gRNA designs or chemical modifications that selectively destabilize mismatch-bound complexes while preserving rapid and productive on-target binding. In this regard, the present results align closely with earlier findings showing that prolonged off-target residence without cleavage has been shown to contribute to cellular toxicity and unintended regulatory effects, underscoring the importance of minimizing such interactions.11,52 Accordingly, gRNA modifications such as truncation of the guide sequence (17–18 nt instead of 20 nt), introduction of destabilizing nucleotide substitutions, or chemical modifications have been shown to selectively reduce off-target binding and residence times while preserving strong on-target interactions.51,53,54
Moreover, these findings provide a foundation for integrating nfEIS into high-throughput CRISPR guide RNA screening pipelines. In a standard genome-editing workflow, gRNAs are designed and synthesized prior to cellular delivery, while functional evaluation typically occurs only after electroporation and subsequent pool or clonal analysis. We envision incorporating nfEIS immediately following guide synthesis and prior to cell-based editing steps (Fig. S4). In this format, arrays of independently addressable electrodes functionalized with target and predicted off-target DNA sequences would enable parallel measurement of binding affinity, association kinetics, dissociation behavior, and residence time across large gRNA libraries. Similar multiplexed electrochemical platforms have been demonstrated for nucleic acid detection and protein–DNA interaction studies, supporting the feasibility of scaling nfEIS to higher-throughput formats.18–20 In this context, nfEIS could function as a rapid, label-free prescreening tool to rank candidate guides based on kinetic discrimination metrics before advancing to electroporation, pool analysis, clonal isolation, and downstream quality control. Such integration would complement, rather than replace, established cellular validation workflows in therapeutic genome editing and diagnostic development.
Collectively, these results position nfEIS as both a mechanistic tool for probing Cas9–DNA interactions and a complementary platform for guiding rational gRNA design. By linking electrical readouts to binding dynamics and residence-time-dependent specificity, nfEIS extends existing sequencing- and fluorescence-based approaches and provides a direct, label-free strategy for the development of safer and more effective genome-editing systems.54–56
Conclusion
Overall, our nfEIS measurements, combined with equilibrium binding analysis and time-dependent single-exponential kinetic fitting, revealed clear and informative differences in how spCas9 interacts with perfectly matched versus mismatched DNA targets and how these interactions influence spCas9 binding and dissociation. We found that the perfect-match SCD sequence, which exhibits the strongest affinity (kD = 0.09 nM), supports rapid target recognition and a moderate residence time consistent with efficient R-loop formation and productive activity. In contrast, the WD mismatch, despite having only a slightly weaker affinity (kD = 0.3 nM), shows slower initial binding and a markedly prolonged residence time, indicating formation of a transient but recurring binding of an incompetent complex that acts as a kinetic trap. Cas9 alone displays only weak, short-lived interactions, confirming the essential role of gRNA in establishing specific and stable DNA binding. Importantly, the concentration of MgCl2 played a critical role in tuning spCas9's ability to discriminate between SCD and WD. At 5 mM MgCl2, on-target SCD complexes were stabilized while mismatched WD interactions remained unstable, enabling clear single-nucleotide discrimination. However, at 1 mM MgCl2, this discrimination was lost, demonstrating that Mg2+ strongly influences both catalytic activation and selectivity of spCas9–gRNA complexes. Together, these findings show that spCas9 specificity is governed not only by affinity but also by distinct differences in binding kinetics and Mg2+-dependent stabilization of R-loop states. Therefore, nfEIS provides a powerful, label-free platform for resolving productive versus nonproductive Cas9–DNA interactions and holds significant potential for evaluating gRNA specificity, off-target susceptibility, diagnostics and kinetic mechanisms in CRISPR-based genome-editing applications.Author contributions
Conceptualization by KA and DKB. Experimental design by DKB with KA. Experiment performed by KP under the supervision of DKB and KA. KV and LS helped in chip fabrication, chip holder printing and data acquisition. Data analysis by DKB, DB, and AW. All the authors edited the manuscript and provided feedback.Conflicts of interest
There is no conflict of interest to declare.Data availability
The data supporting this article have been included as part of the Supplementary information.Supplementary information (SI): SI file contains Tables S1: comparison of Cas9-based methods and Table S2: nucleic acid sequences used in this study; Fig. S1–S4 (Nyquist plots showing interfacial resistance Rp and DNase I interaction, agarose gel analysis of spCas9 activity, nfEIS kinetic analysis of spCas9-dsDNA interactions, and the proposed nfEIS-based CRISPR prescreening workflow). The code used for data analysis is also provided in the SI. See DOI: https://doi.org/10.1039/d5sd00227c.
Acknowledgements
The authors are grateful to NIH RO1 HL139605, NSF EAGER 316843-00001 at UCSD, EFMA 2526846 at NSF, and UCSD: Aims Project 2045293 for financial support for this research. This work is supported by the U.S. National Science Foundation Center for Pandemic Insights (NSF CPI) award number 2412522. The schematics in Fig. 1 and 2 were created with http://BioRender.com.References
- M. Jinek, K. Chylinski, I. Fonfara, M. Hauer, J. A. Doudna and E. Charpentier, Science, 2012, 337, 816–821 CrossRef CAS PubMed.
- P. Mali, L. Yang, K. M. Esvelt, J. Aach, M. Guell, J. E. DiCarlo, J. E. Norville and G. M. Church, Science, 2013, 339, 823–826 CrossRef CAS PubMed.
- J. P. K. Bravo, M.-S. Liu, G. N. Hibshman, T. L. Dangerfield, K. Jung, R. S. McCool, K. A. Johnson and D. W. Taylor, Nature, 2022, 603, 343–347 CrossRef CAS PubMed.
- M. Pacesa, L. Loeff, I. Querques, L. M. Muckenfuss, M. Sawicka and M. Jinek, Nature, 2022, 609, 191–196 CrossRef CAS PubMed.
- K. Babu, V. Kathiresan, P. Kumari, S. Newsom, H. P. Parameshwaran, X. Chen, J. Liu, P. Z. Qin and R. Rajan, Biochemistry, 2021, 60, 3783–3800 CrossRef CAS.
- K. A. Hossain, L. Nierzwicki, M. Orozco, J. Czub and G. Palermo, eLife, 2025, 13, RP102538 CrossRef PubMed.
- H. Ma, L.-C. Tu, A. Naseri, M. Huisman, S. Zhang, D. Grunwald and T. Pederson, J. Cell Biol., 2016, 214, 529–537 CrossRef CAS PubMed.
- M.-S. Liu, S. Gong, H.-H. Yu, D. W. Taylor and K. A. Johnson, in Methods in Enzymology, ed. S. Bailey, Academic Press, 2019, vol. 616, pp. 289–311 Search PubMed.
- J. Lee and C. Jeong, BMB Rep., 2025, 58, 8 CrossRef CAS PubMed.
- Y. Wang, J. Mallon, H. Wang, D. Singh, M. Hyun Jo, B. Hua, S. Bailey and T. Ha, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2010650118 CrossRef CAS.
- Y. S. Dagdas, J. S. Chen, S. H. Sternberg, J. A. Doudna and A. Yildiz, Sci. Adv., 2017, 3, eaao0027 CrossRef PubMed.
- X. Zhu, R. Clarke, A. K. Puppala, S. Chittori, A. Merk, B. J. Merrill, M. Simonović and S. Subramaniam, Nat. Struct. Mol. Biol., 2019, 26, 679–685 CrossRef CAS PubMed.
- S. Balderston, J. J. Taulbee, E. Celaya, K. Fung, A. Jiao, K. Smith, R. Hajian, G. Gasiunas, S. Kutanovas, D. Kim, J. Parkinson, K. Dickerson, J.-J. Ripoll, R. Peytavi, H.-W. Lu, F. Barron, B. R. Goldsmith, P. G. Collins, I. M. Conboy, V. Siksnys and K. Aran, Nat. Biomed. Eng., 2021, 5, 713–725 CrossRef CAS PubMed.
- R. Hajian, S. Balderston, T. Tran, T. deBoer, J. Etienne, M. Sandhu, N. A. Wauford, J.-Y. Chung, J. Nokes, M. Athaiya, J. Paredes, R. Peytavi, B. Goldsmith, N. Murthy, I. M. Conboy and K. Aran, Nat. Biomed. Eng., 2019, 3, 427–437 CrossRef CAS PubMed.
- C. Robinson, V. B. Juska and A. O'Riordan, Environ. Res., 2023, 237, 116877 CrossRef CAS PubMed.
- M. A. MacDonald and H. A. Andreas, Electrochim. Acta, 2014, 129, 290–299 CrossRef CAS.
- H. Stevenson, N. Radha Shanmugam, A. Paneer Selvam and S. Prasad, SLAS Technol., 2018, 23, 5–15 CrossRef CAS.
- F. Lisdat and D. Schäfer, Anal. Bioanal. Chem., 2008, 391, 1555–1567 CrossRef CAS PubMed.
- J. S. Daniels and N. Pourmand, Electroanalysis, 2007, 19, 1239–1257 CrossRef CAS PubMed.
- C. Berggren, B. Bjarnason and G. Johansson, Electroanalysis, 2001, 13, 173–180 CrossRef CAS.
- E. Katz and I. Willner, Electroanalysis, 2003, 15, 913–947 CrossRef CAS.
- M. Labib, E. H. Sargent and S. O. Kelley, Chem. Rev., 2016, 116, 9001–9090 CrossRef CAS PubMed.
- I. E. Barsoukov, Impedance spectroscopy theory, experiment, and applications, Wiley Online Library, 2005 Search PubMed.
- I. Giaever and C. R. Keese, Proc. Natl. Acad. Sci. U. S. A., 1991, 88, 7896–7900 CrossRef CAS PubMed.
- J. Wegener, C. R. Keese and I. Giaever, Exp. Cell Res., 2000, 259, 158–166 CrossRef CAS PubMed.
- N. Meini, M. Ripert, C. Chaix, C. Farre, G. De Crozals, R. Kherrat and N. Jaffrezic-Renault, Mater. Sci. Eng., C, 2014, 38, 286–291 CrossRef CAS PubMed.
- C. W. Tan and J. Miao, Thin Solid Films, 2009, 517, 4921–4925 CrossRef CAS.
- A. S. Tanak, B. Jagannath, Y. Tamrakar, S. Muthukumar and S. Prasad, Anal. Chim. Acta: X, 2019, 3, 100029 CAS.
- A. Kakatkar, T. S. Abhilash, R. D. Alba, J. M. Parpia and H. G. Craighead, Nanotechnology, 2015, 26, 125502 CrossRef PubMed.
- E. A. Boyle, J. O. L. Andreasson, L. M. Chircus, S. H. Sternberg, M. J. Wu, C. K. Guegler, J. A. Doudna and W. J. Greenleaf, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 5461–5466 CrossRef CAS.
- Ł. Nierzwicki, K. W. East, J. M. Binz, R. V. Hsu, M. Ahsan, P. R. Arantes, E. Skeens, M. Pacesa, M. Jinek, G. P. Lisi and G. Palermo, Nat. Catal., 2022, 5, 912–922 CrossRef.
- A. M. Romani, Arch. Biochem. Biophys., 2011, 512, 1–23 CrossRef CAS PubMed.
- F. J. Arjona and J. H. F. de Baaij, J. Physiol., 2018, 596, 747–750 CrossRef CAS PubMed.
- A. R. Eggers, K. Chen, K. M. Soczek, O. T. Tuck, E. E. Doherty, B. Xu, M. I. Trinidad, B. W. Thornton, P. H. Yoon and J. A. Doudna, Cell, 2024, 187, 3249–3261.e14 CrossRef CAS PubMed.
- M. Klein, B. Eslami-Mossallam, D. G. Arroyo and M. Depken, Cell Rep., 2018, 22, 1413–1423 CrossRef CAS.
- S. H. Sternberg, S. Redding, M. Jinek, E. C. Greene and J. A. Doudna, Nature, 2014, 507, 62–67 CrossRef CAS PubMed.
- C. D. Richardson, G. J. Ray, M. A. DeWitt, G. L. Curie and J. E. Corn, Nat. Biotechnol., 2016, 34, 339–344 CrossRef CAS.
- V. Mekler, L. Minakhin, E. Semenova, K. Kuznedelov and K. Severinov, Nucleic Acids Res., 2016, 44, 2837–2845 CrossRef PubMed.
- J. C. Cofsky, D. Karandur, C. J. Huang, I. P. Witte, J. Kuriyan and J. A. Doudna, eLife, 2020, 9, e55143 CrossRef CAS PubMed.
- P. Aldag, M. Rutkauskas, J. Madariaga-Marcos, I. Songailiene, T. Sinkunas, F. Kemmerich, D. Kauert, V. Siksnys and R. Seidel, Nat. Commun., 2023, 14, 3654 CrossRef CAS PubMed.
- C. R. Cromwell, K. Sung, J. Park, A. R. Krysler, J. Jovel, S. K. Kim and B. P. Hubbard, Nat. Commun., 2018, 9, 1448 CrossRef PubMed.
- A. T. Raper, A. A. Stephenson and Z. Suo, J. Am. Chem. Soc., 2018, 140, 2971–2984 CrossRef CAS PubMed.
- F. Zheng, Z. Chen, J. Li, R. Wu, B. Zhang, G. Nie, Z. Xie and H. Zhang, Adv. Sci., 2022, 9, 2105231 CrossRef CAS PubMed.
- M. D. Newton, B. J. Taylor, R. P. C. Driessen, L. Roos, N. Cvetesic, S. Allyjaun, B. Lenhard, M. E. Cuomo and D. S. Rueda, Nat. Struct. Mol. Biol., 2019, 26, 185–192 CrossRef CAS PubMed.
- D. Singh, S. H. Sternberg, J. Fei, J. A. Doudna and T. Ha, Nat. Commun., 2016, 7, 12778 CrossRef CAS PubMed.
- H. Feng, J. Guo, T. Wang, C. Zhang and X. Xing, Nucleic Acids Res., 2021, 49, 1263–1277 CrossRef CAS.
- R. Sundaresan, H. P. Parameshwaran, S. D. Yogesha, M. W. Keilbarth and R. Rajan, Cell Rep., 2017, 21, 3728–3739 CrossRef CAS PubMed.
- D. Singh, Y. Wang, J. Mallon, O. Yang, J. Fei, A. Poddar, D. Ceylan, S. Bailey and T. Ha, Nat. Struct. Mol. Biol., 2018, 25, 347–354 CrossRef CAS PubMed.
- M. Jinek, A. East, A. Cheng, S. Lin, E. Ma and J. Doudna, eLife, 2013, 2, e00471 CrossRef PubMed.
- F. Jiang, D. W. Taylor, J. S. Chen, J. E. Kornfeld, K. Zhou, A. J. Thompson, E. Nogales and J. A. Doudna, Science, 2016, 351, 867–871 CrossRef CAS PubMed.
- P. D. Hsu, D. A. Scott, J. A. Weinstein, F. A. Ran, S. Konermann, V. Agarwala, Y. Li, E. J. Fine, X. Wu, O. Shalem, T. J. Cradick, L. A. Marraffini, G. Bao and F. Zhang, Nat. Biotechnol., 2013, 31, 827–832 CrossRef CAS PubMed.
- S. H. Sternberg, B. LaFrance, M. Kaplan and J. A. Doudna, Nature, 2015, 527, 110–113 CrossRef CAS PubMed.
- Y. Fu, J. D. Sander, D. Reyon, V. M. Cascio and J. K. Joung, Nat. Biotechnol., 2014, 32, 279–284 CrossRef CAS PubMed.
- I. M. Slaymaker, L. Gao, B. Zetsche, D. A. Scott, W. X. Yan and F. Zhang, Science, 2016, 351, 84–88 CrossRef CAS PubMed.
- B. P. Kleinstiver, V. Pattanayak, M. S. Prew, S. Q. Tsai, N. T. Nguyen, Z. Zheng and J. K. Joung, Nature, 2016, 529, 490–495 CrossRef CAS PubMed.
- C. A. Vakulskas, D. P. Dever, G. R. Rettig, R. Turk, A. M. Jacobi, M. A. Collingwood, N. M. Bode, M. S. McNeill, S. Yan, J. Camarena, C. M. Lee, S. H. Park, V. Wiebking, R. O. Bak, N. Gomez-Ospina, M. Pavel-Dinu, W. Sun, G. Bao, M. H. Porteus and M. A. Behlke, Nat. Med., 2018, 24, 1216–1224 CrossRef CAS PubMed.
Footnote |
| † Equal first author. |
| This journal is © The Royal Society of Chemistry 2026 |