Open Access Article
Open Access ArticleLow-cost printed circuit board (PCB) electrochemical biosensors for rapid and label-free detection of Streptococcus pneumoniae
Vincent Vezzaa,
Veerappan Mani a,
Niamh Docherty
a,
Niamh Docherty a,
Adrian Butterwortha,
David Alcornb,
Paul A. Hoskisson
a,
Adrian Butterwortha,
David Alcornb,
Paul A. Hoskisson c and
Damion Corrigan
c and
Damion Corrigan *a
*a
aUniversity of Strathclyde, Centre for Advanced Measurement Science and Health Translation, Pure and Applied Chemistry, Thomas Graham Building, 295 Cathedral St, Glasgow G1 1XL, UK. E-mail: damion.corrigan@strath.ac.uk
bDepartment of Anaesthesia, Royal Alexandra Hospital, Corsebar Rd, Paisley, PA2 9PN, UK
cStrathclyde Institute of Pharmacy and Biomedical Sciences, 161 Cathedral St, Glasgow G4 0RE, UK
First published on 4th February 2026
Abstract
Severe sepsis presents a critical healthcare challenge where rapid pathogen identification is vital for timely intervention. Current diagnostic methods, however, remain inadequate, often delaying targeted treatment. Using readily available printed circuit board (PCB) electrodes, we address this need by developing a low-cost electrochemical DNA biosensor for rapid detection of Streptococcus pneumoniae using the lytA gene as a biomarker. Through systematic evaluation of commercial and custom PCB designs (P1–P4), gold-plated PCB P4 was found as the optimal platform, demonstrating sensitive detection of lytA sequences (20 bp at 4.50 pM limit of detection in buffer) and clinically relevant 235 bp polymerase chain reaction (PCR) amplicons in 100% human serum (1.0–100 pM) within 15 min at room temperature using electrochemical impedance spectroscopy. The performance of the biosensor originates from the optimized electrode geometry, surface properties, and robust self-assembled monolayer functionalization, enabling specific recognition of bacterial DNA without sample pretreatment. This work establishes PCB-based biosensors as a promising solution for point-of-care sepsis diagnostics, offering significant advantages in speed, cost, and operational simplicity compared to conventional methods.
1. Introduction
Severe sepsis is a life-threatening medical emergency that provides a significant challenge to clinicians and health care systems.1 The UK sepsis trust database indicates that sepsis is responsible for 11 million deaths globally each year and 48![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 sepsis related deaths each year in the UK.2 Sepsis is an extremely challenging illness to diagnose as there is often a multitude of signs, symptoms and measurements that can occur and can be attributed to many other serious or benign medical conditions.3 Among the various pathogens responsible for sepsis, Streptococcus pneumoniae infection is commonly fatal particularly in cases of community-acquired respiratory infections, highlighting the urgent need for rapid and specific diagnostic tools.4–6 There is a growing demand for a low-cost, rapid point-of-care (PoC) diagnostic device capable of detecting Streptococcus pneumoniae, not only in hospitals but also in community screening, primary care, and home healthcare settings.7
000 sepsis related deaths each year in the UK.2 Sepsis is an extremely challenging illness to diagnose as there is often a multitude of signs, symptoms and measurements that can occur and can be attributed to many other serious or benign medical conditions.3 Among the various pathogens responsible for sepsis, Streptococcus pneumoniae infection is commonly fatal particularly in cases of community-acquired respiratory infections, highlighting the urgent need for rapid and specific diagnostic tools.4–6 There is a growing demand for a low-cost, rapid point-of-care (PoC) diagnostic device capable of detecting Streptococcus pneumoniae, not only in hospitals but also in community screening, primary care, and home healthcare settings.7
Phenotypic identification, genotypic identification, and mass spectrometry are the three key methods used for this purpose; the main issue with such methodologies is that they can take hours to days to result.8–10 Phenotypic identification, the current gold standard, relies on culturing blood samples to observe bacterial growth, followed by biochemical testing which typically requires more than 72 hours. Genotypic identification relies on DNA or RNA analysis through techniques like 16S/23S ribosomal DNA sequencing and amplified ribosomal DNA restriction analysis (ARDRA), but they are limited by time consumption, the need for specialized equipment and expertise, and reduced effectiveness in distinguishing closely related strains. Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) identifies pathogens by analysing their protein mass spectra through laser desorption and time-of-flight measurement, offering rapid, cost-effective detection but is limited by the need for prior colony cultivation (24–48 hours), dependence on reference spectra, and database availability.10 Notably, all these methods are lab-centric and not suitable for PoC use. The pneumococcal urinary antigen test is the only currently available PoC device that can provide a time to result upon sample application in 15 min; however, it requires temperature equilibration of samples and reagents and involves multiple manual steps, and its generalizability can be uncertain.11,12 These drawbacks highlight the need for the development of better PoC devices.
Electrochemical biosensors are a promising solution, offering potential for miniaturization, low cost, and rapid results.13,14 Specifically, printed circuit board (PCB) electrodes are a compelling platform because they have significant advantages, including high customizability through modern design software, established manufacturing processes for rapid, precise, and low-cost mass production, and a variety of available metal plating finishes.15,16 Increased electrode numbers can be manufactured on smaller surfaces resulting in higher sample sizes within smaller time intervals. Due to the maturation of PCB systems and their extensive incorporation in modern electronic devices, they can be easily integrated into electrochemical readers. They can have additional electronics easily incorporated onto or conjoined to the electrode system of the end goal device. PCB biosensors have been reported in the literature with a wide range of research applications undertaken.17 However, a label-free PCB-based DNA sensor for the direct detection of S. pneumoniae in clinically relevant matrices like human serum has not been demonstrated.
The main objective of this work is to develop PCB-based low-cost, rapid diagnostic test strips for rapid detection of Streptococcus pneumoniae, to tackle existing limitations in sepsis diagnostics. Four different PCBs (P1–P4) have been demonstrated in this work, each having different electrode configurations, sizes, geometries, surface materials, and manufacturing methods (Fig. 1). The four PCB electrodes were not selected to enable direct comparison between identical surfaces, but rather to represent distinct and practically relevant PCB manufacturing routes: PCB P1: fully in-house fabricated prototype, PCB P2: commercially fabricated board with in-house gold plating, PCB P3: standard commercial fabrication (bare copper), and PCB P4: commercially fabricated and optimized for biosensing (gold-plated). The aim of the study was to assess how these different fabrication pathways commonly encountered during biosensor development and scale-up influence electrode surface characteristics and biosensing performance. By functionalising PCB electrodes with a DNA probe sequence that specifically binds to a truncated sequence derived from the S. pneumoniae lytA gene which encodes the major autolysin lytA, a label-free S. pneumoniae biosensor platform was developed.18 Amplification of a 235 bp from the lytA gene has been shown to electrochemically distinguish S. pneumoniae from similar streptococci.19 Our investigation shows that PCB P4 biosensor can detect 10 pM levels of lytA in buffer. The amplified 235 bp lytA was detectable at 1.0 pM in complex human serum with a 15 min time to result at room temperature.
2. Experimental
2.1. Reagents
Deionized (DI) water (resistivity ≥18 MΩ cm), hydrogen peroxide (H2O2), phosphate buffered saline tablets (PBS), potassium chloride (KCl), potassium ferricyanide (K3[Fe(CN)6]), potassium ferrocyanide (K4[Fe(CN)6]), sulfuric acid (H2SO4), tris(2-carboxyethyl)phosphine (TCEP), and 3-mercapto-1-propanol (MCP) were purchased from Sigma. Acid piranha solution was prepared by mixing 18.0 M H2SO4 and 30% H2O2 in a volume ratio of 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v). PBS (pH 7.40) was the buffer used throughout unless otherwise specified. All oligonucleotides (probe, target, and primers) were obtained from Sigma, and their sequences are provided in Table 1.
1 (v/v). PBS (pH 7.40) was the buffer used throughout unless otherwise specified. All oligonucleotides (probe, target, and primers) were obtained from Sigma, and their sequences are provided in Table 1.
| Oligonucleotide | Sequence (5′ - 3′) |
|---|---|
| lytA probe | [ThilC6] [SP18] TGCCGAAAACGCTTGATACA |
| lytA target | TGTATCAAGCGTTTTCGGCA |
| lytA forward primer | TTGGGAACGGTTGCATCATG |
| lytA reverse primer | TCGTGCGTTTTAATTCCAGCT |
| blaOXA-1 forward primer | AACAGAAGCATGGCTCGAAA |
| blaOXA-1 reverse primer | TGGTGTTTTCTATGGCTGAGTT |
2.2. Custom PCB electrodes
Four PCB electrodes featuring 4 different designs and geometries are named as PCB P1, PCB P2, PCB P3, and PCB P4. Detailed specifications of all the electrodes are provided in Table S1. All PCB boards used the same electrochemical cell setup for the plating process. Once each board had obtained a gold surface finish, it was cleaned to remove any impurities and contaminants. The details about the cell configuration for electrode plating and cleaning, the plating procedure, and the cleaning procedure are given in the SI (sections S1–S3, Tables S1 & S2, and Fig. S1 & S2). The reference and counter electrode geometries differ between PCB designs as part of an exploratory study into layout efficiency. While these geometric variations can influence absolute electrochemical parameters, all designs were controlled to maintain stable reference potentials and adequate counter electrode competence. The comparative performance assessment focuses on the normalized biosensing response (i.e., the change in impedance or current, not the absolute values) measured under identical assay conditions, ensuring that observed differences are attributable to the overall sensing platform rather than to the reference/counter electrode geometries.2.3. Fabrication of a DNA biosensor
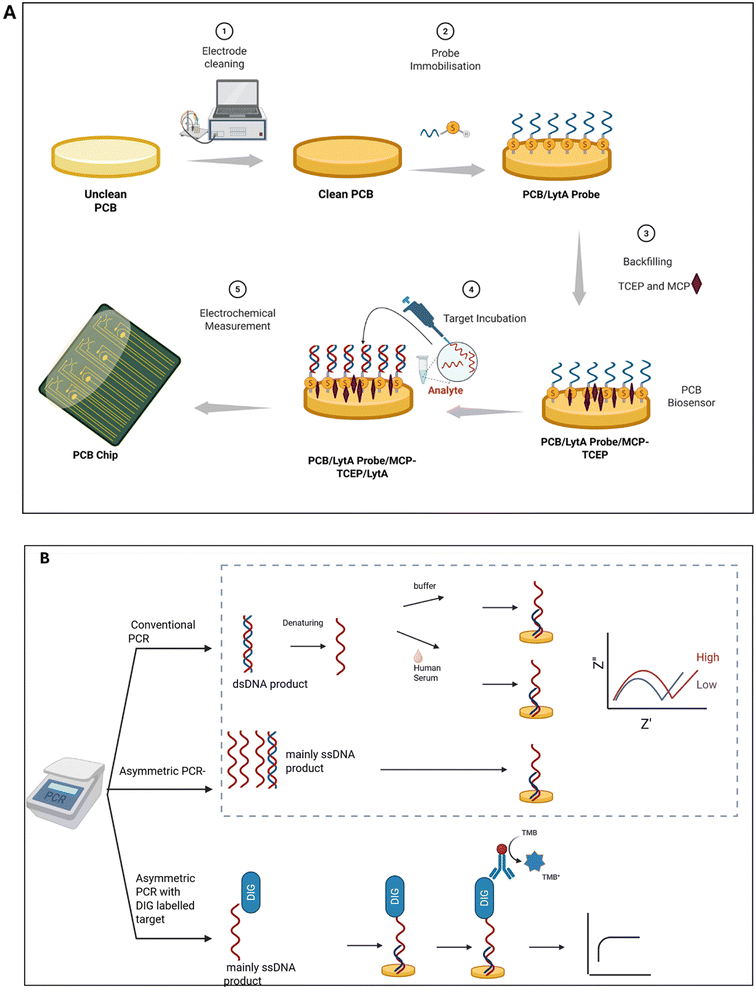
All experiments used the same protocol for probe functionalisation and a scheme is provided in Fig. 2A. A solution mixture containing 3.0 μM lytA probe solution and 15 μM TCEP solution was first prepared. This mixture was applied to the PCB electrode cell surface and incubated overnight at room temperature. PBS washing was then performed to remove any non-specifically bound DNA. The electrode surface was subsequently backfilled with 1.0 mM MCP for 1 h at room temperature, followed by another PBS washing step to eliminate excess materials. This process yielded the final lytA PCB biosensors. For a typical electrochemical measurement of lytA detection, target concentrations were sequentially incubated on the biosensor electrode surface for 30 min at room temperature. After each incubation, the electrodes were washed with PBS to remove unbound materials before measurement. This cycle was repeated for each concentration. The sequences of the lytA primers and probes were adopted from the previous literature.19 The blaOXA-1 (oxa) primers were designed in-house using the E. coli plasmid pEK499 genome sequence, based on methods described in previous studies.202.4. Polymerase chain reaction (PCR) amplification
PCR amplifications were carried out using a miniPCR mini8 thermal cycler, and the resulting products were analysed using a blueGel electrophoresis system (miniPCR Bio, Cambridge, MA, USA) (Fig. 2B). Quantification of PCR products was performed using a Qubit 4 fluorometer (Thermo Fisher Scientific). The 235 bp lytA amplicons were amplified from S. pneumoniae genomic DNA TIGR4 [ATCC BAA-334] (LGC Standards, Middlesex, UK), while the oxa amplicons were amplified from E. coli plasmid pEK499. Asymmetric PCR was achieved by lowering the concentration of the forward primer, resulting in an unequal amount of forward and reverse segments to hybridize, and thereby producing a higher concentration of single-stranded DNA (ssDNA) target amplicons. Asymmetric digoxigenin (DIG)-labelled amplification followed the same approach but used a DIG PCR mix (Sigma-Aldrich) in place of the standard mix, allowing incorporation of DIG into the amplified sequence. The DIG-labelled target was subsequently detected using a horseradish peroxidase (HRP)-conjugated anti-digoxigenin antibody [HRP.21H8] (Abcam), which binds specifically to the DIG moiety. The bound HRP facilitates the oxidation of 3,3′,5,5′-tetramethylbenzidine (TMB), which can then be electrochemically reduced and quantified via chronoamperometry.2.5. PCR amplicons in PBS
Various types of 235 bp lytA PCR amplicons (positive control) and 115 bp oxa PCR amplicons (derived from OXA-48-type ß-lactamase gene; negative control) were utilized in the experiments. The used amplicon types included (1) standard PCR amplicons for lytA and oxa, which required preheating at 95 °C for 5 min to denature the double-stranded DNA, (2) asymmetric lytA and oxa PCR amplicons, which did not require heat treatment, and (3) asymmetric DIG-labelled lytA and oxa PCR amplicons for electrochemical detection via antibody recognition. All amplicons were diluted in 1× PBS, and a range of target concentrations was tested. Target incubation was performed for 15 min at room temperature, followed by PBS and 0.05% Tween washing steps prior to measurement. For DIG-labelled PCR amplicons, chronoamperometric detection was carried out by incubating the electrodes with anti-DIG antibodies (diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 from stock) for 30 min at room temperature. This was followed by a brief 20 s 1× PBS washing step, after which 10 μL of TMB substrate was added and allowed to react for 20 min. Chronoamperometry was then performed with an applied potential (Edc) of −0.15 V for 2 s.
1000 from stock) for 30 min at room temperature. This was followed by a brief 20 s 1× PBS washing step, after which 10 μL of TMB substrate was added and allowed to react for 20 min. Chronoamperometry was then performed with an applied potential (Edc) of −0.15 V for 2 s.
2.6. PCR amplicons in human serum
PCB P4 was used for all experiments given in this section. Standard lytA and oxa PCR amplicons were first denatured by heating at 95 °C for 5 min to convert the double-stranded DNA into single strands. The denatured amplicons were then spiked into 100% human serum (origin: human male AB plasma, USA; sterile filtered, Sigma-Aldrich) to make various target concentrations. Each concentration was sequentially incubated on the electrodes for 15 min at room temperature. After each incubation, the electrodes were washed with 0.05% Tween solution, tested, and then rinsed with PBS before proceeding to the next target concentration. This process was repeated until all concentrations had been tested.2.7. Electrochemical measurements
Electrochemical measurements were performed using a PalmSens4 potentiostat running with PSTrace software (Alvatek Electrochemical Solutions, Houten, The Netherlands). All measurements in this work were performed sequentially on individual working electrodes using an MUX8-R2 multiplexer. Measurements were performed in at least triplicate, and the averaged results were used for data analysis and calculations. Each PCB electrode utilised its inbuilt Au counter and reference electrodes. Unless otherwise specified in the figure captions, the redox buffer used for all measurements consisted of 2.0 mM K3Fe(CN)6 and 2.0 mM K4Fe(CN)6 prepared in 0.5× PBS. The measurement parameters for chronoamperometry (TMB-based): applied potential (Edc) = −0.15 V; duration = 2 s. Electrochemical impedance spectroscopy (EIS) parameters: DC bias potential = 0.0 V (vs. open-circuit potential); frequency range = 1 Hz to 50 kHz.3. Results and discussion
3.1. Designing PCBs
The first PCB designed in this work was designated PCB P1. It consisted of eight working electrodes of various diameters (Table S1), arranged around a central circular reference electrode and an annular counter electrode (Fig. 1A), and was manufactured in-house. The objectives for this prototype were (1) to assess whether the copper pads and traces could be successfully plated with gold, (2) to evaluate the ease and speed of interfacing the PCB with a potentiostat measurement system, and (3) to determine whether the Au surface is capable of providing suitable electrochemical responses that would justify further development for biosensing applications.21,22 PCB P1 incorporates multiple working electrodes with different diameters as part of a geometry exploration study. All working electrodes were used, they were just normalised by their area to make them comparable, and no mixing of data from different diameters occurred within a given analysis. When comparisons were made across different working electrode geometries, current density and area-normalised impedance were used to ensure that the results were normalised to electrode area and therefore directly comparable despite geometric differences.Next, commercially manufactured PCBs were purchased. The design was changed to include eight working electrodes of equal diameter and rectangular reference and counter electrodes, generating a more spatially efficient layout (Table S1). Two versions of this updated design were obtained: (1) PCB P2, which was Au plated by the manufacturer, and (2) PCB P3, which was left non-plated to allow for in-lab testing of various Au plating techniques. Utilising a third-party manufacturer enabled the production of a larger number of devices and better quality of fabrication. Both PCBs P2 and P3 exhibited electrochemical responses characteristic of Au surfaces. Notably, PCB P3 demonstrated superior electrochemical behaviour, although with limited reproducibility. Drawing from the insights gained through the development and testing of PCBs P1 to P3, a refined version was designed and commercially produced. This final electrode, offering optimal and reproducible performance, was designated as PCB P4.
Thus, the four PCB electrodes were selected to represent a progression of fabrication pathways relevant to biosensor development and scale-up: from in-house prototyping (P1) to commercial manufacturing with varying levels of optimization (P2–P4). Their surfaces are inherently dissimilar due to differences in manufacturing processes, which directly influence surface properties such as roughness and active area. This variability allows for a practical comparison of how common fabrication choices impact electrochemical biosensing performance when tested under identical assay conditions. The goal of this comparison was to identify the manufacturing route that provides the optimal balance of sensitivity, reproducibility, and scalability for a low-cost diagnostic platform.
3.2. lytA detection using custom PCB electrodes
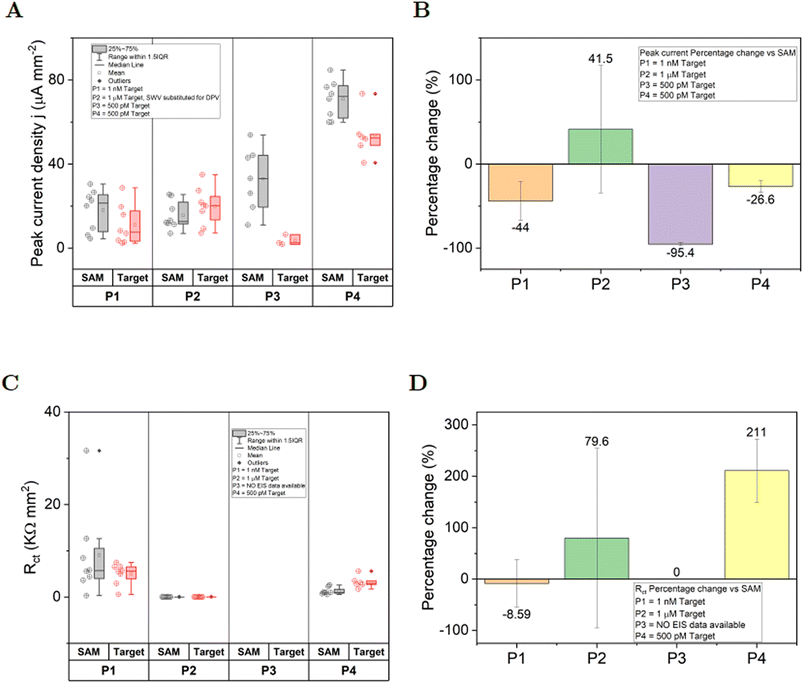
The lytA DNA detection performance of the four PCB designs (P1–P4) was evaluated using differential pulse voltammetry (DPV) and EIS. All electrodes underwent identical self-assembled monolayer (SAM) functionalization, with measurements conducted in 5.0 mM redox buffer. DPV measurements were performed on all four PCB designs before and after incubation with 500 pM lytA target DNA. Fig. 3A presents the DPV current densities, while Fig. 3B shows the corresponding baseline-subtracted currents. PCB P1 exhibited a moderate current decrease (−44%) upon 1.0 nM lytA addition, though the target and control groups overlapped, indicating no significant differentiation. PCB P2 showed a 41.5% current increase at 1.0 μM lytA but similarly lacked statistical distinction from the SAM group. In contrast, PCB P3 demonstrated a pronounced current reduction (≈95%) at 500 pM lytA, with clear separation between target and control data without any overlap. However, it suffered from electrode loss (4 electrodes) and inconsistent performance. PCB P4 also displayed significant current decreases (≈26%) at 500 pM, with compacted distributions and no data overlap between groups. Two outliers were observed, but their inclusion did not alter the conclusion of distinct target/SAM responses. Notably, P4 initially lost two electrodes due to a connector issue (unrelated to the electrodes themselves), which was resolved in subsequent experiments. Fig. 3C presents the charge transfer resistance (Rct) values from EIS measurements before and after incubation with 500 pM lytA target DNA addition for each PCB electrode, and Fig. 3D presents the corresponding percentage changes. PCB P1 showed a minor Rct decrease reduction (−8.6%), with overlapping distributions indicating no significant difference. PCB P2 exhibited minimal impedance values and a 76.6% increase, though not discernible from DPV data due to axis scaling; no statistical distinction was found. P3 produced unreliable EIS fits, precluding analysis. In contrast, PCB P4 demonstrated a clear 211% increase in impedance, with non-overlapping data ranges suggesting reliable target detection.Overall, PCBs P1 and P2 failed to demonstrate reliable lytA detection. PCB P3 showed promising DPV performance but suffered from inconsistent EIS results and electrode loss. PCB P4 outperformed all other designs, offering robust and reproducible detection of 500 pM lytA using both DPV and EIS, with improved electrode consistency following connector repair. Based on its superior sensing performance, stability, and reliability, PCB P4 was selected for all subsequent biosensing experiments.
3.3. Detailed investigation of lytA detection using PCB P4
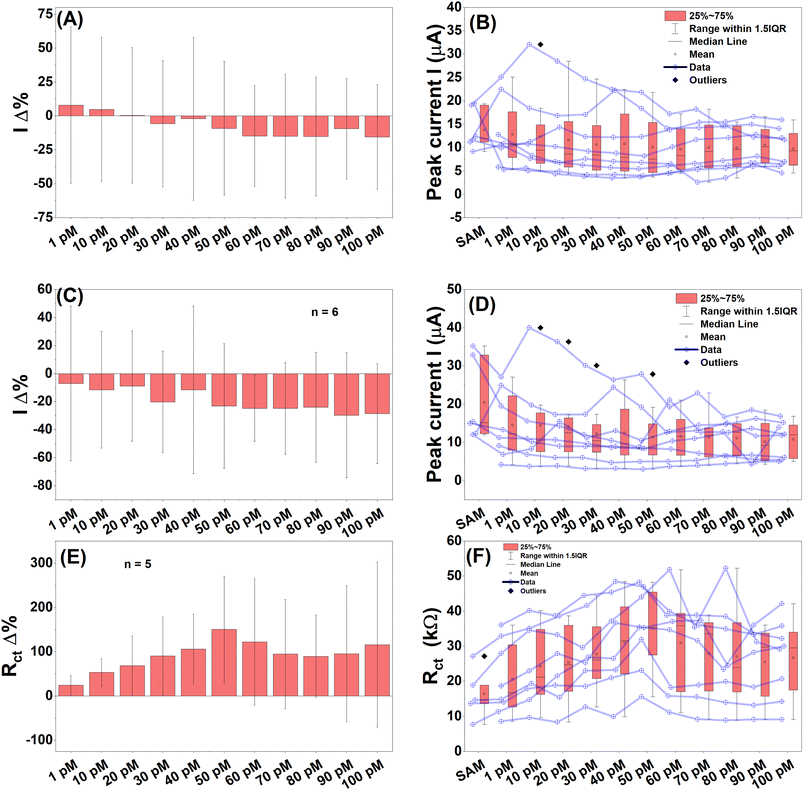
The performance of PCB P4 for detecting various concentrations of lytA target DNA was evaluated, and the results are presented in Fig. 4. The DPV response current linearly decreased and showed a dose-dependent fashion for different target concentration ranges from 500 fM to 500 nM. The mean percentage change in DPV peak current demonstrated a generally linear and dose-dependent decrease in response to increasing lytA concentrations. However, the 5.0 pM concentration slightly deviated from this trend, and a trend reversal was observed at 50 pM, after which the signal continued to decrease with higher concentrations (Fig. 4A). This unexpected trend reversal most likely occurred due to the multiple incubation, washing and testing processes, destabilising the probe DNA. The corresponding raw DPV peak current data showed considerable spread, with significant differences likely occurring between the SAM control and the 30 pM to 50 nM target concentrations; the remaining concentrations appeared statistically similar (Fig. 4B). Square wave voltammetry (SWV) results exhibited comparable percentage change magnitudes and dose–response trends across a range of 500 fM to 500 nM (Fig. 4C), with the raw data reflecting similar behaviour to DPV but with fewer outliers (Fig. 4D). Likely significant differences between the SAM control and target samples were again observed in the 30 pM to 50 nM range. The mean Rct percentage change from the EIS results also showed a dose-dependent increase, with the 5.0 pM concentration again deviating from the overall trend and with a noticeable reversal at 50 pM, consistent with the DPV and SWV results (Fig. 4E). Notably, the magnitude of the EIS response was significantly larger, about 250% compared to the ∼30% observed with DPV and SWV. The raw Rct data showed several outliers; exclusion of these would have reduced the magnitude of the percentage change but improved statistical errors (Fig. 4F). Significant differences between the SAM control and the target DNA were evident from 10 pM up to 50 nM. Overall, EIS was found to be the best performing detection technique among the CV, SWV and EIS, demonstrating the highest magnitude of percentage changes, clearer distinctions between control and target stages, and smaller data spreads for each concentration of lytA.Further analysis of lytA detection performance on PCB P4 was conducted by performing linear regression within the linear response region between 10 pM and 50 pM. The LoD was also calculated. The LoD for lytA was determined to be 10.4 pM, 12.39 pM, and 4.5 pM using DPV, SWV, and EIS, respectively. A second linear response region was observed in all three techniques, between 0 and 2.0 pM, yielding LOD values ranging from 251 to 433 fM. This suggests that the PCB P4 biosensor is capable of detecting femtomolar concentrations of 20 bp complementary S. pneumoniae DNA, indicating potential for ultra-sensitive detection. Notably, a previously reported peptide nucleic acid (PNA)-based PCB device for rapid and high-sensitivity DNA quantification reported an LoD of 57 fM.17 The biosensor described in this work demonstrated comparable LoDs (251–433 fM) without requiring expensive PNA reagents, instead relying on a straightforward SAM modification strategy.
3.4. PCR amplification of genetic sequences at PCB P4
With a suitable platform established and successful detection of the 20 bp complementary lytA DNA sequences in PBS, the performance of the probe was next subjected to longer DNA target fragments. This aimed to more precisely replicate the DNA lengths that a sensor would be required to deal with in PoC settings. For this purpose, a 235 bp sequence (amplified from S. pneumoniae genomic DNA) that contained 20 bp lytA complementary sequence in the middle of the larger sequence was used. The reason for it being in the middle and not at the ends of the sequence was due to the primer design from the previous literature.19 To further validate the specificity of the lytA probe, negative controls were included using 115 bp oxa amplicons.| S. no. | Amplifying target | Length (bp) | Qubit quantification (μg mL−1) |
|---|---|---|---|
| 1. | lytA (S. pneumoniae) | ||
| Positive | 235 | 14.1 | |
| Negative | NA | 0.9 | |
| 2. | lytA (S. pneumoniae) DIG | ||
| Positive | 235 | 3.29 | |
| Negative | NA | 2.57 | |
| 3. | lytA (S. pneumoniae) asymmetric | ||
| Positive | 235 | 6.73 | |
| Negative | NA | 0.399 | |
| 4. | lytA (S. pneumoniae) asymmetric DIG | ||
| Positive | 235 | 8.73 | |
| Negative | NA | 1.11 | |
| 5. | Oxa (E. coli) | ||
| Positive | 115 | 11.3 | |
| Negative | NA | 2.78 | |
| 6. | Oxa (E. coli) DIG | ||
| Positive | 115 | 4.89 | |
| Negative | NA | 2.75 | |
| 7. | Oxa (E. coli) asymmetric | ||
| Positive | 115 | 5.13 | |
| Negative | NA | 2.16 | |
| 8. | Oxa (E. coli) asymmetric DIG | ||
| Positive | 115 | 46.4 | |
| Negative | NA | 2.23 | |
Similarly, for the oxa amplifications, the positive PCR mixture showed a concentration of 11.3 μg mL−1, while the negative control measured 2.78 μg mL−1 (Table 2). The significantly higher concentration in the positive sample confirmed successful nucleic acid amplification. The gel electrophoresis analysis revealed no visible band in the negative lane, while the positive lane showed a clear band in the lower end in the range of 100–200 bp (Fig. S3B), consistent with the desired sequence of 115 bp long amplicon. The distorted appearance of the gel was noted, which could be attributed to water vapour condensation on the plastic cover. These results confirmed successful amplification of the oxa amplicons, enabling the subsequent specificity testing.
3.5. lytA genetic amplicons in PBS using PCB P4
These results can likely be attributed to non-specific biofouling of the electrode surface. As shown in Table 2, both asymmetric lytA and oxa reactions produced similar levels of amplified materials (6.73 and 5.13 μg mL−1, respectively). However, the gel electrophoresis analysis for both showed no indication of bands in their lanes which suggested that the specific DNA material was not present in the sample. If this were true, then both PCBs exposed to the lytA positive and oxa negative were in reality receiving various concentrations of non-specific proteins and nucleotide mixtures that composed the PCR solutions. The responses seen here would then easily be explained as a biofouling dose response of these protein/nucleotide contaminants with none or insufficient quantities of amplicons present.
The PCBs were washed after the previous measurements and TMB was applied for 20 min. The TMB solution turned blue confirming the presence of HRP. Chronoamperometry was performed producing a current decrease for both positive and negative controls (Fig. 6 and S4E, test 1). There were no differences between the controls. To ensure that the unspecifically bound HRP was properly removed, a second washing step incorporating 20 s of 1× PBS, 20 s of 0.05% Tween, and then 20 s of 0.1× PBS was performed and reapplication of TMB for 20 min was undertaken. The response again showed a current decrease for both controls with the negative decreasing more than the positive (Fig. S4E and test 2) producing no differences between the groups. This chronoamperometry method produced TMB colour reactions for both control group solutions when only the positive control should. This meant that the HRP anti DIG antibodies were present on the negative control and were unspecifically bound, with multiple washing attempts unable to successfully remove them.
3.6. lytA genetic amplicon in human serum
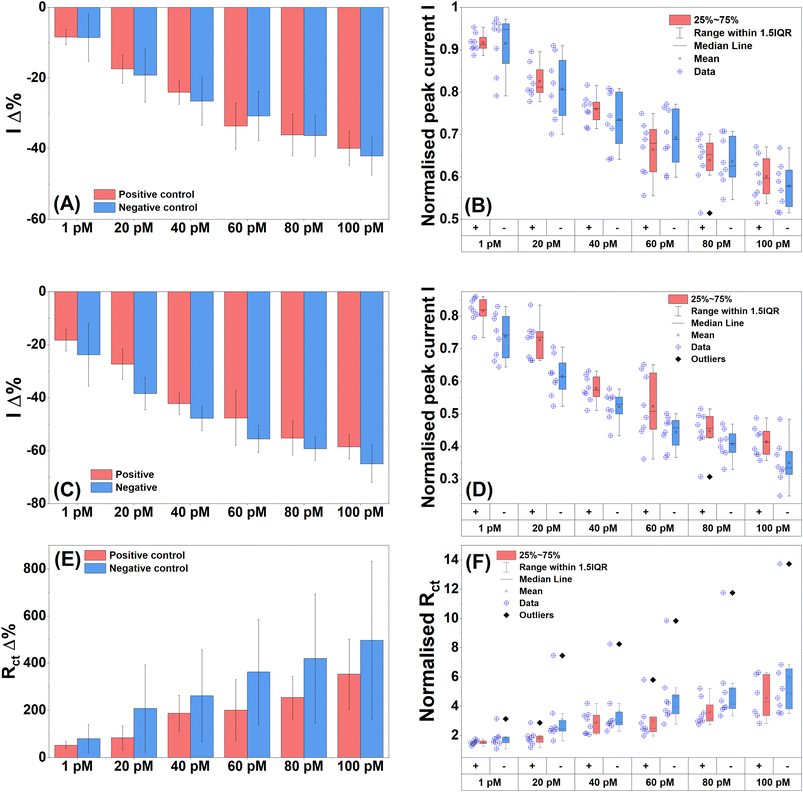
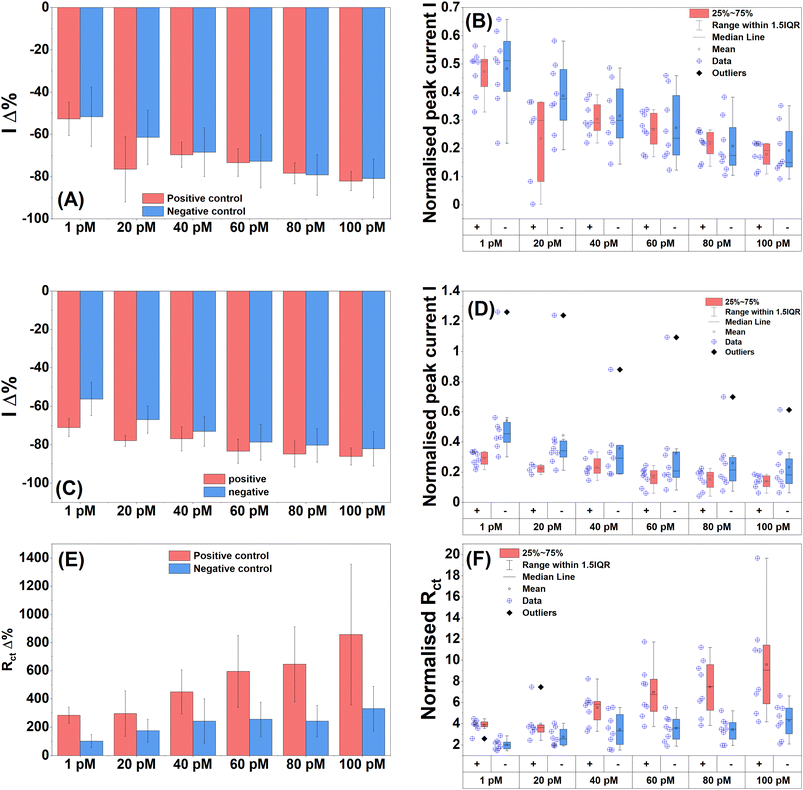
With successful specific detection of full complementary 20 bp and 235 bp normal PCR and asymmetric DIG-labelled amplicons in PBS, the next step was to attempt detection of the 235 bp amplicon (amplified from S. pneumoniae genomic DNA) in a clinically relevant matrix. Normal 235 bp lytA and 115 bp oxa PCR products were heated to 95 °C for 5 min and then spiked into 100% human serum. The serum samples were incubated on the electrode surface for 15 min, Tween washed and then tested. The target concentration range was 1.0 pM to 100 pM. The mean DPV peak current percentage change displayed a dose-dependent response to increasing target with 20 pM failing to follow the trend (Fig. 7A). The negative control also showed a dose-dependent response. The normalised DPV data confirmed the percentage changes with the 20 pM verging on a likely difference between the two controls (Fig. 7B). The other target concentrations showed no differences. The mean SWV peak current percentage changes showed a slight improvement over DPV with more pronounced differences between the control groups (Fig. 7C). Dose dependence was again present for both control groups. The normalised data showed that 1.0 and 20 pM control groups were likely different and 60, 80 and 100 pM were close to being different (Fig. 7D).EIS again proved to be the best performing detection technique with the PCB P4 biosensor. The mean Rct percentage change showed increasing dose dependence for the positive control (Fig. 7E). The negative control increased until 40 pM and then remained constant until 100 pM where it then increased slightly. Therefore, the negative control did not produce a strong dose response, and the sensor demonstrated anti-biofouling properties. The normalised mean Rct data showed likely differences between positive and negative groups for 1.0, 60, 80 and 100 pM (Fig. 7F). Some positive target stages were likely different from each other indicating a strong signal response to target DNA while no differences were found within the negative control groups, meaning that they were all likely equivalent further reinforcing the anti-biofouling performance of the sensor. The LoD values for the various measurement techniques were calculated, which were 23 pM for DPV, 30 pM for SWV, and 18 pM for EIS. The LoD values correlated with the concentrations demonstrated to have likely differences from the box plot data, with the exception of EIS which was able to successfully detect target DNA at 1.0 pM which was significantly lower than the calculated LoD.
Table 3 summarises the comparisons with sensors from the literature. Compared to several previous sensors, the developed sensor demonstrated excellent responses for such a simple device and would hopefully continue showing such responses with testing of reduced concentrations and incubation times. This is a vast improvement on current culture methods which regularly return no culture positive after multiple hours of incubation. This sensor also outperforms the pneumococcal urinary antigen test in terms of time to result, expense and complexity. The time to result values are both 15 min but this should be easily reduced on this sensor system. The sensor does not require to be at specific temperature nor do the sample and reagents.11 This sensor comprises sample addition, washing, redox addition and then results, reducing the complexity of the assay. The antigen test requires samples and multiple reagents. Antigen tests are relatively expensive approximately in £3–£5, whereas this sensor would likely be in the region of pence if it aligns its manufacture protocol with that of glucose strip production which is demonstrated in the next section for a Coronavirus Disease 2019 (COVID-19) sensor. Other tests on the market include a magnetogenosensor assay for lytA that was able to obtain a LoD of 1100 pM.19 Other examples of reported S. pneumoniae sensors include a DNA–antibody nanostructure assay (≈3.26 pM),23 a polymer-based genosensor (≈1 × 106 pM),24 a synergism strategy for gene sequence sensing (0.0005 pM),25 and versatile loop-mediated isothermal amplification with a sensitivity of 0.173 pM.26 The sensor developed in this work demonstrated similar or improved sensitivity to the devices reported. It was also able to achieve this in complex human serum with reduced time to result utilising less complicated functionalisation strategies on an easily manufacturable device. The sensors above that demonstrate low pM detection used complex devices, materials and methods to achieve this. The PCB P4 design, which provides a dedicated counter and reference electrode for each working electrode, creates an array of independent electrochemical cells. This design inherently minimizes the risk of crosstalk, a significant advantage for future parallelized operation in a point-of-care device.
| Sensor/technology | Label free? | Approx. cost per test | Key complexity | LOD (pM) | Time to result | Real sample |
|---|---|---|---|---|---|---|
| Abbreviations: UAT: urinary antigen test; LAMP: loop-mediated isothermal amplification; PNA: peptide nucleic acid; MB = magnetic beads. | ||||||
| Cell culture | Yes | Low | Simple (but slow) | NA | 72 h | Blood |
| PCR | Yes | High | Moderate | NA | Hours | Extracted DNA |
| UAT30 | Yes | ≈£5 | Moderate | NA | 15 min | Urine |
| Magnetosensor19 | No | High (∼£4.65) | High (MB modification) | 1100 | 5 h | Buffer |
| DNA antibody23 | No | Very high | Very high (DNA tetrahedron) | 3.26 | >30 min | Serum |
| Polymer genosensor24 | Yes | High | Moderate | 1 × 106 | 20 min @ 55 °C | Buffer |
| Synergism gene sequence25 | No | Very high | High (multiple probes) | 0.0005 | 20–60 min | 50% serum |
| LAMP26 | No | Moderate (≈£2.80) | High (microfluidics) | 0.173 | ≈2 h | Clinical samples |
| PNA-based lab-on-PCB17 | Yes | Moderate (PNA costly) | High (microfluidics) | 0.057 | 5 min (flow) | Buffer |
| PCB electrodes (this work) | Yes | Very low (≈£0.2) | Low (standard SAM) | 1.0 | 15 min | 100% serum |
For deployment as a practical PoC device, this PCB electrode can be integrated into a single-use, disposable cartridge designed for ‘plug-and-play’ operation. Our group recently demonstrated a similar plug-and-play, easy-to-manufacture PoC device for diagnostic applications.27 Such cartridge houses can protect the delicate electrodes and electrical contacts, incorporates a defined sample chamber with a simple injection port for regulated sample introduction, and ensures robust fluidic containment to prevent user exposure to the sample or reagents. The user would simply inject the prepared sample, insert the cartridge into a portable, handheld reader, and receive a digital readout. This approach, common in commercial glucose meters and other POC diagnostics, effectively decouples the complex microfluidics and electrochemistry from the end-user, ensuring reliability, safety, and ease of use. Future work will focus on prototyping such cartridges and integrating them with low-cost, portable readers.
For translation to a field-deployable POC device, power consumption is a key practical consideration. The electrochemical techniques employed (EIS and DPV) require very low operating currents (<100 μA). Commercial mini-potentiostats demonstrate that such systems can run for extended periods on small batteries or a smartphone connection.28,29 Therefore, a dedicated reader for this PCB platform would have minimal power demands, easily supporting a full day of operation on a compact, rechargeable battery, aligning with the requirements for use in resource-limited settings.
These results demonstrated that SWV could partially, and EIS could fully successfully and specifically detect an unlabelled 235 bp lytA amplicon amplified from S. pneumoniae full genomic DNA over a range of concentrations from 1.0–100 pM in 15 min at room temperature. Clinically relevant blood levels that can result in life-threatening infections can be as low as 1–10 colony-forming units per millilitre of blood (cfu mL−1).31 This equates to approximately attomolar (aM) levels of genetic material for detection. The sensor has shown possible femtomolar LoDs and should comfortably be able to detect these femtomolar concentrations. Attomolar ranges would be a challenge to achieve, nevertheless until the sensor is tested with these concentrations, no feasibility conclusions can be made. If the sensor was unable demonstrate detection at these small concentrations, then PCR amplification would have to be employed to amplify the DNA to detectable levels. This would be easily accomplished as PCR amplification was already employed for target sequence production for these experiments. Therefore, the infrastructure already existed and has been successfully demonstrated. Two hours of amplification was required to produce nanomolar levels of target DNA with the sensor demonstrating picomolar capabilities. Consequently, the attomolar DNA would only require amplification to picomolar levels shortening the current amplification times used. This would result in the successful detection of clinically relevant levels of S. pneumoniae in under 2 h.
4. Conclusions
This study systematically investigated PCB-based electrodes for rapid sepsis diagnostics through S. pneumoniae lytA DNA detection. While prototype PCBs (P1–P3) showed progressive improvements in electrochemical performance, they faced limitations in reliability and consistency. The commercially fabricated PCB P4 emerged as the optimal platform, demonstrating superior and reproducible DNA detection capabilities, making it ideal for sepsis biosensor development. Using a 20 bp lytA sequence, we established a PCB P4-based electrochemical detection system with a 4.5 pM LoD in PBS. The system was further validated with PCR-amplified longer target sequences and successfully detected normal double-stranded DNA amplicons, asymmetric (single-stranded DNA) and asymmetric DIG-labelled amplicons, and 235 bp lytA sequences spiked in 100% human serum (1–100 pM detection via EIS in 15 min at room temperature). While some asymmetric PCR products showed detection limitations due to potential quantity/contamination issues, the platform demonstrated clinically relevant sensitivity. This work presents a functional biosensor capable of detecting 1.0 pM lytA amplicons in complex matrices within 15 minutes at ambient temperature. With further optimization, this system shows strong potential for reduced detection times, enhanced sensitivity, and direct clinical sample analysis. The developed PCB platform establishes a foundation for rapid, point-of-care sepsis diagnostics with significant translational promise.Conflicts of interest
The authors declare no conflicts of interest.Data availability
Key data that support the findings of this study have been included in the manuscript. Additional data that support this article have been included as part of the supplementary information (SI). Supplementary information: electrochemical set-up for gold plating and cleaning, procedure for gold plating on PCB electrodes, cleaning of gold PCB electrodes, table for PCB specifications, optimized chemical cleaning and electrochemical procedure, and additional figures. See DOI: https://doi.org/10.1039/d5sd00210a.Acknowledgements
This work was supported by the Centre for Advanced Measurement Science and Health Translation, University of Strathclyde. PAH would like to acknowledge the Royal Academy of Engineering Research Chair Scheme for long term personal research support (RCSRF2021\11\15). DA would like to acknowledge educational support from Baxter Pharmaceuticals and MSD (Merck Sharp and Dohme). DC would like to thank the Dowager Countess Eleanor Peel Trust and Tenovus Scotland for seed grants in the area of sepsis monitoring. VV's EngD studentship was funded by the EPSRC CDT in Biomedical Devices and Health Technologies (EP/L015595/1).References
- J.-L. Vincent, EBioMedicine, 2022, 86, 104318 CrossRef CAS PubMed.
- T. K. Burki, Lancet Respir. Med., 2018, 6, 826 CrossRef PubMed.
- C. F. Duncan, T. Youngstein, M. D. Kirrane and D. O. Lonsdale, Curr. Infect. Dis. Rep., 2021, 23, 1–14 CrossRef PubMed.
- I. Martin-Loeches, L. F. Reyes and A. Rodriguez, Thorax, 2025, 80, 565–575 CrossRef PubMed.
- N. Gopal, N. Chauhan, U. Jain, S. K. Dass, H. S. Sharma and R. Chandra, Artif. Cells, Nanomed., Biotechnol., 2023, 51, 476–490 CrossRef CAS PubMed.
- A. S. Tanak, A. Sardesai, S. Muthukumar and S. Prasad, Bioeng. Transl. Med., 2022, 7, e10310 CrossRef CAS PubMed.
- S. Balayan, N. Chauhan, R. Chandra, N. K. Kuchhal and U. Jain, Biosens. Bioelectron., 2020, 169, 112552 Search PubMed.
- R. Varghese, R. Jayaraman and B. Veeraraghavan, J. Microbiol. Methods, 2017, 141, 48–54 CrossRef CAS PubMed.
- S. L. Goh, B. P. Kee, K. Abdul Jabar, K. H. Chua, A. M. Nathan, J. Bruyne, S. T. Ngoi and C. S. J. Teh, Pathog. Global Health, 2020, 114, 46–54 CrossRef CAS PubMed.
- W.-T. Fan, T.-T. Qin, R.-R. Bi, H.-Q. Kang, P. Ma and B. Gu, Eur. J. Clin. Microbiol. Infect. Dis., 2017, 36, 1005–1012 CrossRef CAS.
- P. Kim, A. Deshpande and M. B. Rothberg, Infect. Drug Resist., 2022, 2219–2228 CrossRef CAS PubMed.
- B. Kim, J. Kim, Y. H. Jo, J. H. Lee, J. E. Hwang, M. J. Park and S. Lee, PLoS One, 2018, 13, e0200620 CrossRef PubMed.
- P. Lasserre, B. Balansethupathy, V. J. Vezza, A. Butterworth, A. Macdonald, E. O. Blair, L. McAteer, S. Hannah, A. C. Ward and P. A. Hoskisson, Anal. Chem., 2022, 94, 2126–2133 CrossRef CAS PubMed.
- S. Kumar, S. Tripathy, A. Jyoti and S. G. Singh, Biosens. Bioelectron., 2019, 124, 205–215 CrossRef PubMed.
- S. Mahapatra, R. Kumari and P. Chandra, Trends Biotechnol., 2024, 42, 591–611 Search PubMed.
- H. Shamkhalichenar, C. J. Bueche and J.-W. Choi, Biosensors, 2020, 10, 159 CrossRef CAS PubMed.
- P. Jolly, J. Rainbow, A. Regoutz, P. Estrela and D. Moschou, Biosens. Bioelectron., 2019, 123, 244–250 CrossRef CAS PubMed.
- D. Llull, R. Lpez and E. Garca, J. Clin. Microbiol., 2006, 44, 1250–1256 CrossRef CAS PubMed.
- S. Campuzano, M. Pedrero, J. L. Garca, E. Garca, P. Garca and J. M. Pingarrn, Anal. Bioanal. Chem., 2011, 399, 2413–2420 CrossRef CAS PubMed.
- N. Woodford, A. Carattoli, E. Karisik, A. Underwood, M. J. Ellington and D. M. Livermore, Antimicrob. Agents Chemother., 2009, 53, 4472–4482 CrossRef CAS PubMed.
- D. Pletcher, A first course in electrode processes, Royal Society of Chemistry, 2009 Search PubMed.
- R. G. Compton and C. E. Banks, Understanding voltammetry, World Scientific, 2007 Search PubMed.
- J. Wang, M. C. Leong, E. Z. W. Leong, W. S. Kuan and D. T. Leong, Anal. Chem., 2017, 89, 6900–6906 CrossRef CAS PubMed.
- F. P. Ferreira, A. C. Honorato-Castro, J. V. da Silva, S.-C. Orellana, G. C. Oliveira, J. M. Madurro and A. G. Brito-Madurro, Polym. Eng. Sci., 2018, 58, 1308–1314 CrossRef CAS.
- F. Li, Z. Yu, Y. Xu, H. Ma, G. Zhang, Y. Song, H. Yan and X. He, Anal. Chim. Acta, 2015, 886, 175–181 CrossRef CAS PubMed.
- H. Wang, Z. Ma, J. Qin, Z. Shen, Q. Liu, X. Chen, H. Wang, Z. An, W. Liu and M. Li, Biosens. Bioelectron., 2019, 126, 373–380 CrossRef CAS PubMed.
- A. Dobrea, N. Hall, S. Milne, D. K. Corrigan and M. Jimenez, Sci. Rep., 2024, 14, 14154 CrossRef CAS PubMed.
- A. Ainla, M. P. S. Mousavi, M.-N. Tsaloglou, J. Redston, J. G. Bell, M. T. Fernández-Abedul and G. M. Whitesides, Anal. Chem., 2018, 90, 6240–6246 CrossRef CAS PubMed.
- K. A. Bautista, E. Madsen, S. D. Riegle and J. C. Linnes, ACS Electrochem., 2025, 1, 386–394 CrossRef CAS.
- F. Gutirrez, M. Mar, J. C. Rodrguez, A. Ayelo, B. Soldn, L. Cebrin, C. Mirete, G. Royo and A. M. Hidalgo, Clin. Infect. Dis., 2003, 36, 286–292 CrossRef PubMed.
- S. O. Kelley, ACS Sens., 2017, 2, 193–197 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |