Open Access Article
Open Access ArticleControlling the signal through interfacial design in aptamer-based electrochemical sensors
Ashkan
Koushanpour†
a,
Yara
Raphael†
c,
Edward J.
Harvey
b and
Geraldine E.
Merle
 *bc
*bc
aExperimental Surgery, Faculty of Medicine, McGill University, Montreal, H3A 0C5, Canada
bDepartment of Surgery, Faculty of Medicine, McGill University, Montreal, H3A 0C5, Canada
cDepartment of Chemical Engineering, Polytechnique, Montreal, H3T 1J4, Canada. E-mail: geraldine.merle@polymtl.ca
First published on 12th January 2026
Abstract
Electrochemical aptamer-based (E-AB) sensors represent a distinct class of biosensing platforms that convert target recognition events into faradaic signals through conformational changes at electrode interfaces. Unlike conventional chemical- or enzyme-dependent systems, E-AB sensors operate via purely physical transduction mechanisms, minimizing susceptibility to pH fluctuations, enzymatic degradation, or interfering side reactions. These attributes enable robust operation in complex physiological environments and have been demonstrated in clinically relevant contexts, including intraoperative cancer diagnostics. Nevertheless, broad implementation has been limited by challenges in interfacial chemistry, aptamer stability, electrode nanostructuring, and signal reproducibility. This review critically examines intrinsic factors governing E-AB performance, with emphasis on interfacial engineering, aptamer selection and modification, and nanostructured electrode architectures. Advances in molecular design and materials integration are highlighted, alongside emerging fabrication strategies that enhance sensitivity, dynamic range, and operational stability. By synthesizing recent progress and identifying persistent bottlenecks, this work outlines pathways toward realizing the clinical and technological potential of E-AB sensors.
1. Introduction
In recent years, the demand for rapid, accurate, and cost-effective diagnostic tools has intensified, especially for point-of-care (POC) applications. Biosensors have emerged as a promising solution due to their low fabrication cost, ease of use, and minimal sample volume requirements.1 Among the various biosensing platforms under investigation, electrochemical biosensors have gained prominence. Their appeal lies in their exceptional sensitivity, often reaching picomolar (pM) detection limits and their modular compatibility with a broad spectrum of biorecognition elements. These advantages contribute to the fact that over half of the biosensors reported in the literature utilize electrochemical transducers.The only reagentless and fully quantitative biosensor that has garnered vast commercial success so far, has been the household, and portable glucose biosensors.2 Given that the field of biosensors is still in its early stages most concepts have failed translational from in vitro laboratory studies to in vivo preclinical research and face important challenges before being ready for widespread adoption and clinical application. Therefore, it is encouraging to briefly examine the principles underlying the success of glucose biosensors. The most important key to their success is the production of a readily measurable by-product that is detected unambiguously against the background. In addition, the reaction is enzymatically catalyzed and thus naturally associated with signal amplification. Furthermore, that reaction relies on target-binding-induced chemical evolution instead of specific chemical reactivity, which accounts for a key element that makes them well-suited in contaminated samples e.g., blood, as the enzyme–target complex is less likely to be disrupted. In nature, the chemosensing phenomena rely on biomolecular switches, biomolecular equilibrium between two structural conformations, a process in which a special biomolecular output (biological signal) is brought about by binding-induced changes in conformation or oligomerization (be it a protein or nucleic acid). These natural biosensors enable real-time, continuous target monitoring in highly complex environments. Motivated by such phenomena, significant efforts have been invested in adoption of such switches into the construction of artificial biosensors, for which DNA/RNA aptamers have shown the capability to partially fulfil this goal.
1.1 Emergence of electrochemical biosensors for point-of-care diagnostics
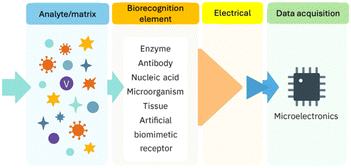
Electrochemical biosensors operate by converting molecular recognition events into measurable electrical signals through a transducing electrode. Electrochemical biosensors are generally composed of several critical components: a substrate for immobilization, a linker, a biorecognition element, a signal receptor, a signal amplification module, and a signal transducer.Among these, the two most essential elements are the biorecognition element and the signal transducer (Fig. 1). The biorecognition element is responsible for conferring high selectivity by specifically binding the target analyte.
The signal transducer, typically an electrode, converts the molecular recognition event into a quantifiable electrical signal.
In this setup, electrical changes induced by analyte binding are monitored and controlled at the electrode surface, allowing for precise, real-time measurement of biological interactions.
1.2 Aptamers as biorecognition element for electrochemical sensor
Among the diverse biorecognition elements, aptamers, short, single-stranded oligonucleotides, have emerged as attractive alternatives to conventional molecules such as antibodies and enzymes.3 Their synthetic accessibility, high stability, and ease of chemical modification offer significant advantages. A comparative overview of the key attributes of antibodies and aptamers is provided in Table 1.| Properties | Antibody | Aptamers |
|---|---|---|
| Generation and synthesis | In vivo selection | In vitro selection |
| Selection process cannot be tailored on demand | Selection process can be tailored as per need | |
| Produced in animals or by recombinant technology | Chemically synthesized in vitro conditions | |
| Difficult to raise antibodies against non-immunogenic entities | Can be developed against non-immunogenic entities | |
| High cost of production | Economical cost of production | |
| Batch-to-batch variation | Negligible batch-to-batch variation | |
| Stability | Requires stringent storage conditions (cold storage) | Can withstand a range of storage conditions |
| Low shelf life | Higher shelf life | |
| High susceptibility to change in pH, temperature, and ionic concentrations | Relatively immune to changes in pH, temperature and ionic concentrations | |
| Stability cannot be increased | Stability can be improved | |
| Modification, specificity and affinity | Comparable specificity and affinity with aptamers | Comparable specificity and affinity with antibodies |
| Affinity and specificity can be tailored | Affinity and specificity can be tailored on demand | |
| Modification is challenging. Or not possible | Amenable to modifications | |
| Difficulties in immobilization | Immobilization is comparatively easier | |
| Structural switching | On binding to its target, the antibody does not undergo target-induced structural change | Aptamers can easily undergo a target-induced structural change |
1.3 Electrochemical aptamer-based sensor
The first E-AB sensor for thrombin detection was developed by Xiao and Plaxco,5 in which they leveraged the reversible binding capabilities of nucleic acid aptamers, which are capable of folding into specific three-dimensional structures upon target binding, thereby enabling both molecular recognition and signal transduction.6 This concept drew inspiration from natural biological systems, where biomolecules often undergo conformational changes upon interaction with their targets. Here, single-stranded DNA (ssDNA), inherently flexible and disordered in the absence of a complementary target, folds into a rigid, well-defined double helix upon hybridization.7 In the foundational E-AB sensor design, thiol-functionalized aptamers were immobilized onto electrode surfaces via self-assembled monolayers (SAMs) and covalently tagged with redox-active molecules, given their lack of intrinsic capability to produce an electrochemical signal. Target binding induced conformational changes in the aptamer structure, thereby modulating the electron transfer between the redox tag and the electrode, generating a quantifiable electrochemical signal.8 This mechanism mimics the behaviour of naturally occurring chemoreceptors in the human body.9 What set E-AB sensors apart from previous electroanalytical methods, such as potentiometry or direct biocatalytic conversion, was their ability to detect a broad range of analytes, not limited to chemically reactive species, simply by designing or selecting the appropriate aptamer.9a This flexibility enabled specific, real-time measurements even in complex physiological environments. Other emerging biosensing technologies, such as CRISPR-based electrochemical biosensors,10 organic electrochemical transistors11 (OECTs), microfluidic-integrated optical biosensors,12 and field-effect transistor (FET)-based biosensors,13 are being actively investigated for their high sensitivity and multiplexing capabilities. FET-based sensors, including graphene and silicon nanowire devices, can achieve extremely low LODs due to direct transduction of binding events into changes in channel conductance; however, their performance is often compromised in physiological fluids by Debye screening effects, frequently necessitating sample dilution or low-ionic-strength buffers. Similarly, CRISPR-based platforms typically rely on enzymatic signal amplification and endpoint detection, while OECTs and optical systems often require more complex device architectures or instrumentation and can be sensitive to environmental drift in complex matrices. In comparison, electrochemical aptamer-based (E-AB) sensors offer reversible, label-free, and reagent-free real-time monitoring directly in complex biological fluids, positioning them as a complementary platform rather than a replacement especially for POC and applications.Recent literature has established important foundations in E-AB sensor fabrication and performance optimization. Schoukroun-Barnes et al.14 provided a comprehensive overview of fabrication strategies for E-AB sensors, demonstrating how surface chemistry and assembly parameters influence signal output and reproducibility. Yoo et al.15 extended this discussion further by evaluating aptamer functionalization approaches that improve real-time detection performance, including modular multifunctionalization strategies for next-generation sensors. Fontaine et al.16 investigated confounding interfacial effects associated with alkanethiol-based self-assembled monolayers, redox reporters, and aptamer configuration, offering insights into sources of signal variability. Finally, Arroyo-Currás et al.17 addressed the translational challenges of adapting E-AB sensors from controlled laboratory settings to complex in vivo diagnostic environments. In contrast, the present review integrates interfacial engineering with electrochemical and operational parameters such as voltammetric interrogation and frequency optimization that collectively govern E-AB sensor performance. In addition, we highlight recent advances in interfacial stabilization and antifouling monolayers, including developments reported after 2023, which have significantly improved long-term operation in complex biological matrices. By examining how interfacial modularity and operational control shape E-AB sensor behaviour, this review aims to identify emerging design strategies that improve analytical performance and operational reliability in real-world diagnostic settings.
2. Biosensor architecture and basic principles
2.1 Mechanisms of signal transduction
In DNA/RNA-based sensors, the interaction between the target and its specific aptamer results in a three-dimensional conformational change induced by specific atomic/molecular interaction. This recognition strategy places significant design demands on interfacial engineering and molecular architecture, both of which are critical for optimizing sensor performance. The relative spatial arrangement in the target/aptamer complex determines the class of targets, which are broadly categorized into two groups: the embedded group and the outside-binding group. Most often, targets are found buried in a specific pocket formed by a specialized oligonucleotide sequence of aptamers, this is characteristic of the embedded group, such as ATP,18 cocaine,19 K+ and, theophylline.20 Here, the design strategies are mostly based on target-induced conformational changes of surface-bound aptamers and are generally less complex. In the case of larger and more complex molecules, such as proteins with multiple binding sites (e.g., thrombin,5 and platelet-derived growth factor-BB (PDGF-BB)21), targets fall into the outside-binding group, where design strategies must be more diverse and interfacial engineering becomes significantly more challenging.2.2 Interfacial recognition modes
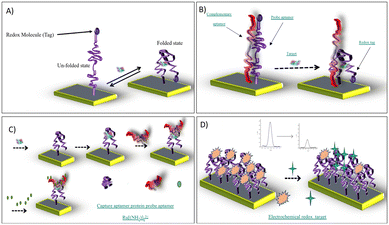
Electrochemical aptamer-based (E-AB) sensors exploit several distinct interfacial signal-transduction strategies, which can be broadly categorized into four major modes (Fig. 2): (a) target-induced structure-switching; (b) target-induced dissociation or displacement; (c) sandwich structure formation; and (d) competitive replacement.In the structure-switching mode (Fig. 2A), target recognition induces conformational rearrangements of the surface-bound aptamer, modulating electron transfer efficiency between a redox tag and the electrode. This strategy represents the most established and widely adopted approach for E-AB sensor construction; however, its utility is often limited by modest signal gain and a relatively poor signal-to-noise ratio, particularly in biological matrices.
The target-induced dissociation/displacement strategy (Fig. 2B) enhances signal fidelity by releasing or displacing a complementary strand upon target binding. By minimizing stochastic strand fluctuations, this mode provides superior baseline stability and clearer electrochemical responses.22 The sandwich structure mode (Fig. 2C), analogous to ELISA, relies on dual recognition of the target by a capture aptamer immobilized on the electrode and a secondary reporter-labelled probe. This dual-binding strategy significantly improves selectivity and quantitative accuracy in complex biological samples.23 Finally, the competitive replacement mode (Fig. 2D) exploits competition between the native target and labelled analogues for aptamer binding sites at the electrode interface. This simple yet powerful format enables highly sensitive readouts using standard electrochemical reporters and is particularly attractive for point-of-care applications.24 These interfacial recognition strategies have also been implemented in clinically relevant sensing platforms, as illustrated by the following patient-based example. A liquid biopsy platform employing quantum dots functionalized with EpCAM and vimentin aptamers was developed for the selective capture of circulating tumour cells (CTCs).25 Following capture, PD-L1 expression on the isolated CTCs was quantified using an electrochemical aptamer-based sensor fabricated on a Fe3O4-coated screen-printed carbon electrode, achieving a limit of detection of 2 ng mL−1. Clinical validation in 41 patients with non-small cell lung cancer demonstrated that increased CTC counts and reduced PD-L1 expression were associated with disease progression. Importantly, the use of an electrochemical aptamer-based sensing interface enabled rapid, reagent-free electrochemical readout immediately after cell capture, illustrating how interfacial sensor design supports near-real-time biomarker detection in complex clinical samples and demonstrates the translational potential of E-AB platforms for molecular monitoring in oncology diagnostics.25
2.3 Role of interfacial engineering in sensor functionality
The analytical performance of electrochemical aptamer-based sensors is fundamentally governed by the nature and quality of the molecular interface. Critical factors such as aptamer orientation, surface density, and spatial distance from the electrode surface directly influence signal generation, sensitivity, and noise suppression.9b At the heart of the signal transduction mechanism is the conformational change that aptamers undergo upon target binding. This structural rearrangement modulates the efficiency of electron transfer between a redox-active reporter and the electrode, resulting in a detectable faradaic current.26 Given this mechanism, the rational design of the aptamer–electrode interface through precise control of surface chemistry, molecular configuration, and immobilization strategy is crucial for high-performance sensing, particularly in complex biological fluids or POC environments. The interfacial region, where the electrode meets the biorecognition element, serves as the origin of all signal transduction events, making it a key determinant of assay sensitivity and specificity. Successful biosensor design, therefore, requires strategic engineering of the molecular architecture at this interface. This includes tuning the composition and packing of SAMs, optimizing aptamer density to balance accessibility and signalling efficiency, and selecting redox tags with favourable electron transfer properties.27 Such interfacial engineering is particularly vital for enabling robust sensor performance in challenging applications like liquid-biopsy-based diagnostics.3. Electrochemical aptamer sensor performance
The analytical performance is governed by multiple interdependent parameters, including sensitivity, selectivity, operational stability, and signal reproducibility. These factors are particularly influenced by the chemical and structural attributes of the sensor interface. Parameters such as probe packing density, redox reporter, backfilling agents, and the aptamer itself collectively define the balance between signal gain and stability of the sensor. For example, a higher probe density will enhance the signal, but often at the expense of stability, while the choice of the backfilling agent will determine electron transfer efficiency and nonspecific adsorption on the sensor surface. Central to E-AB signalling is the modulation of electron transfer efficiency, mediated by conformational changes in the aptamer upon target binding, which governs the proximity and interaction of the redox tag with the electrode surface. Understanding how these interfacial factors shape performance is essential for rational sensor design and provides the foundation for the optimization strategies discussed in the following sections.3.1 Signal gain
Among the various performance metrics, the signal gain is one of the most widely adopted, offering a normalized evaluation of sensor response across different architectures and experimental conditions. Defined as the relative change in signal upon target introduction,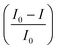 , signal gain enables cross-comparison regardless of absolute current magnitudes.28
, signal gain enables cross-comparison regardless of absolute current magnitudes.28
The signal gain is strongly influenced by the structural and physicochemical properties of the interface. Factors such as the molecular organization of the monolayer, the folding dynamics of the aptamer, and the electron transfer kinetics of the redox label, all modulate signal responsiveness. These interactions often affect the time constants for electron transfer between bound and unbound states, necessitating electrochemical interrogation techniques, like square wave voltammetry or electrochemical impedance spectroscopy, matched to those dynamic timescales. In this section, parameters controlling the chemical and physical nature of the sensor interface, which in turn, determine the performance of the E-AB sensor will be thoroughly discussed.
3.2 Probe packing density
Probe packing density accounts for the most readily observable variable during optimization process toward highest gain and/or best performance. It significantly influences the spatial arrangement of surface-bound aptamers and, consequently, the efficiency of target binding and signal generation. This parameter governs the extent of steric hindrance and electrostatic repulsion between probes, both of which can modulate aptamer folding dynamics and target accessibility.Several studies have been performed to elucidate the effect of packing density on E-AB sensor performance. In 2001, Georgiadis and his co-workers,29 explored the DNA surface density based on the kinetics of target capturing using surface plasmon resonance (SPR). They studied the role of electrostatic forces among double-stranded DNA (dsDNA), and ssDNA as a function of the probe density and the kinetics of monolayer film formation.29 In this work, the kinetics of monolayer film formation showed to have a more pronounced effect on packing density and that ssDNA exhibited greater kinetics and higher density in surface coverage. They demonstrated that the probe density strongly affects target hybridization efficiencies with a higher density value leading to lower hybridization efficiency.30 This was later confirmed by Benight and co-workers32 using cocaine and thrombin, to show the effects of surface density in the signalling gain. In the case of cocaine, low probe density exhibited the highest gain, while an intermediate density for thrombin achieved the best signal gain. This difference was attributed to the inherent structural and spatial characteristics of the aptamers used, with larger aptamers requiring more inter-probe spacing to avoid steric clashes and preserve binding site accessibility.33
Importantly, the target size also contributes: cocaine is a small molecule (∼303 Da), where the dominant factor is the conformational freedom of its relatively compact aptamer, while thrombin is a much larger protein (∼37 kDa), where both the larger aptamer and the bulky target require sufficient spacing to minimize steric hindrance and allow proper binding.28 Thus, for cocaine, reduced steric hindrance at lower densities allows greater conformational freedom and signal amplification, whereas for thrombin, high packing densities are detrimental due to overcrowding and decreased target accessibility. In conclusion, the effect of probe packing density on E-AB sensor performance is dictated by a combined interplay between aptamer structure and target size, with smaller targets (like cocaine) favouring low probe density for optimal signalling, and larger targets (like thrombin) requiring intermediate densities to balance accessibility and signal gain. It was, however, confirmed that controlling the probe aptamer concentration during sensor fabrication successfully modulates the surface density of DNA molecules across an order of magnitude.28 Such control allows the fine-tuning of probe/probe interactions, optimizing folding and signal transduction for each specific aptamer/target pair. Based on these principles, Barton and co-workers23 demonstrated a totally new approach in controlling the probe packing density via in situ electrochemical activation of copper(II) catalyst for Huisgen 1,3-dipolar coupling between the aptamer and the backfilling agents.34 This click-chemistry based approach currently represents one of the most precise methods to regulate surface probe spacing and improve E-AB sensor consistency.
Given the negative charge of the DNA phosphate backbone, the ionic strength also plays a major role in probe behaviour at the electrode interface. Sykes and his co-workers,35 recently reported that ionic strength, or more precisely dielectric permittivity of the solution heavily influences the spatial conformation surface bound DNA sequence. Lower permittivity increased the distance between the redox label and the electrode, reducing the electron transfer rate. Similarly, decreasing ionic strength led to increased electrostatic interactions between DNA and the electrode, thus altering the redox tag's positioning and transfer kinetics.36
The matrix composition, including the presence of stabilizing cations such as K+ and Mg2+, also influences aptamer structure and signal behaviour. Xiao et al.37 investigated the dependence of signal change of thrombin E-AB sensors based on the ionic strength and composition. They found that at a high ionic strength (e.g. 300 mM Tris base, 420 mM NaCl, 60 mM KCl, and 60 mM MgCl2), the apparent binding affinity of an E-AB sensor for thrombin is 50 nM, whereas at lower ionic strength without potassium (100 mM Tris), the affinity improved to 21 nM. This was attributed to the unfolded aptamer structure in low-potassium environments, which undergoes greater conformational rearrangement upon target binding, resulting in improved signal response.37 Overall, packing density is a critical parameter that governs the three-dimensional organization of the probe monolayer and the aptamer–target complex. Its optimization must be tailored to each target for balancing structural flexibility, target accessibility, and signal transduction efficiency, thereby ensuring maximum sensor performance.
3.3 Backfilling agents and surface homogeneity
The structural integrity and homogeneity of the self-assembled monolayer (SAM) formed during E-AB sensor fabrication are key factors influencing probe accessibility, surface stability, and signal reproducibility. Because thiol chemisorption onto gold is inherently oxidative, controlling the applied potential during assembly is a critical parameter in achieving high-quality, defect-free monolayers. Applying a positive potential during assembly has been shown to enhance monolayer quality, promoting improved packing density and reduced defects.38 Ma and Lennox reported a faster kinetics while featuring far fewer defects for surface coverage in preparing a mixed-composition SAM via alkanethiol deposition on gold by the application of constant positive potentials versus the assembly resulted from open circuit potential (OCP).39 Another study showed the influence of the applied potential on the gain with a mixed SAM of fluorophore-labelled DNA and alkylthiol. They showed that a positive applied potential (>0 SCE−1) resulted in ten times higher density compared to deposition at the open circuit potential (OCP) over the same 60 min time-period (Fig. 3).31 Pulsed potential strategies have been shown to accelerate DNA monolayer formation while preserving SAM quality, outperforming constant potential approaches in both speed and uniformity.40 These potential-assisted and pulsed electrochemical assembly techniques clearly improve SAM compactness, reduce defects, and enhance signal reproducibility. However, further refinement is needed to balance assembly speed with long-term electrochemical stability, especially under physiological conditions.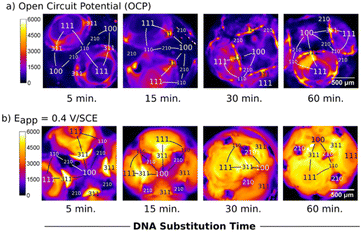 | ||
| Fig. 3 Fluorescence images taken of MCH/DNA layers prepared a) at OCP (no applied potential) and b) at Eapp 0.40 V SCE−1. Images from left to right correspond to increasing time in the deposition solution. Each image is from a different electrode resulting in a different orientation. The stereographic triangle and crystallographic regions analysed are shown on the images. All images are false coloured to represent intensity. Reproduced from ref. 31 with permission from Elsevier,31 copyright 2017. | ||
An alternative approach involves co-adsorption of thiol-modified single-stranded DNA (HS-ssDNA) with short-chain alkanethiol diluents such as mercaptohexanol (MCH).41
The effect of diluent combined with passivation time on surface composition, density, and orientation of HS-ssDNA oligomers was studied by utilizing X-ray photoelectron spectroscopy (XPS), near-edge absorption X-ray absorption fine structure spectroscopy (NEXAFS), and the fluorescence intensity measurements. It was concluded that longer diluent exposure times (>2 h) promote reorganization and vertical orientation of DNA strands within the SAM.42 These findings suggest that probe orientation and density can be finely tuned via co-adsorbate concentration and exposure time, enhancing hybridization accessibility and signal output. This co-adsorption approach offers a controllable and reproducible route to tune DNA packing and accessibility. Nonetheless, the choice of diluent and exposure time must be carefully optimized to prevent unwanted probe desorption and ensure uniform electron-transfer behaviour.
3.4 The length and the chemistry of the backfilling agents
The physicochemical properties of alkanethiol backfilling agents, particularly their chain length and terminal functional groups, have a direct effect on signal transduction in E-AB sensors (Fig. 4). Ricci et al.43 conducted one of the first studies to explore the impact of the alkanethiol length and charge of the SAM passivation layer on the E-AB sensor performance. They observed that the best signalling gain was obtained for an intermediate length of C6-OH compared to short (C2-OH) and long (C11-OH). Shorter co-adsorbates (e.g., C2-OH) allowed for increased probe flexibility and collision frequency but also led to greater baseline noise.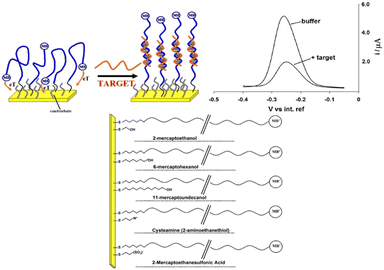 | ||
| Fig. 4 E-AB sensor fabrication and varying lengths of co-adsorbates: here the authors have employed linear probe (top left) as a test bed with which to characterize the effects of surface chemistry on the properties of an E-AB sensor. Because hybridization reduces the rate with which the terminal redox tag collides with the electrode surface and transfers electrons the faradaic current arising from such linear probes is significantly reduced in the presence of a complementary target sequence (top, right). It is thus likely that this suppression and the motion of the unbound and bound probe will be linked to the nature, steric hindrance and charge of the co-adsorbate used for sensor fabrication. They have tested the effects of a range of thiol co-adsorbates (bottom) differing in their length and/or terminal functional groups and chosen to cover a range of charges and steric effects (bottom). Reproduced from ref. 43 with permission from Elsevier,43 copyright 2009. | ||
Conversely, longer chains (e.g., C11-OH) introduced steric hindrance, limiting the redox tag's access to the electrode and thereby reducing signal intensity.
In addition to length, the terminal chemical functionality of the SAM affects both performance and stability. This effect was studied using different terminal functionalities – e.g. hydroxyl, amine, and sulfonate groups (Fig. 5).43 They reported that both positively charged amine and negatively charged sulfonate groups led to improved signal gain, likely due to favorable electrostatic interactions between the SAM and the DNA phosphate backbone. Notably, amine-terminated monolayers enhanced surface stability in static conditions—attributed to their electrostatic binding to DNA.
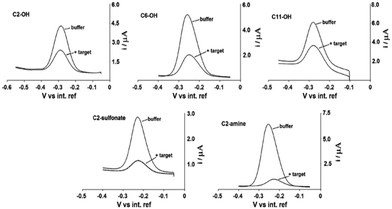 | ||
| Fig. 5 Co-adsorbate effect on E-AB sensor signalling behaviour. Because E-DNA signalling is linked to a binding-specific change in the collision efficiency of the probe-bound redox tag with the electrode surface, the nature (i.e., length and charge) of the co-adsorbate used for sensor production is a determining factor in the performance of E-DNA sensors. Shown are SW voltammograms of sensors fabricated with each of five co-adsorbates before and after the addition of the relevant 17-base target. Here, they find that, among the thiols tested, the positively charged C2-amine (cysteamine) gives rise to the largest and most rapid response to target. Reproduced from ref. 43 with permission from Elsevier,43 copyright 2009. | ||
A major limitation for E-AB sensors deployed in physiological fluids is biofouling, which reduces sensitivity and reliability. Whitesides et al.44 have extensively studied the impact of different alkylthiol monolayer on surface bio-fouling against fibrinogen and lysozyme proteins via surface plasmon resonance spectroscopy (SPR). Here, a single component SAM alkanethiol layer carrying single charge (either positive or negative) resulted in a full monolayer formation of surface fouling proteins, whereas a monolayer constituted from a mixture of two opposite charges acquired less than 1% monolayer of such proteins.44 This result opened the door to the use of zwitterionic monolayers of phosphatidylcholine terminal groups for the fabrication of E-AB sensors44,45 and led to improved signalling in blood despite a strong sensitivity to pH and ionic strength variations. However, zwitterionic SAMs have demonstrated high sensitivity to pH and ionic fluctuations, and their long-term electrochemical stability under continuous operation remains unverified. Progressive desorption of charged monolayers during repeated scans limits their current applicability, highlighting the need for more stable antifouling chemistries.46 Despite ongoing research, 6-mercaptohexanol (C6-OH) remains the most widely used co-adsorbate, balancing adequate passivation, moderate steric hindrance, and acceptable biocompatibility.
3.5 The length of the aptamer
DNA length is another fundamental variable that will impact the organization of surface bound DNA layers. In 2000 Steel et al., examined the influence of oligonucleotide's length on the surface coverage and the corresponding desorption capacity.47 Short and long ssDNAs assembly models on the surface were represented in Fig. 6. | ||
| Fig. 6 Cartoon of two packing configurations for ssDNA probes at a surface with a sticky end-group for specific immobilization. Short probes are envisioned to pack in extended configurations. Longer probes are expected to exist in more flexible, polymeric-like configurations. Reproduced from ref. 47 with permission from Elsevier,47 copyright 2000. | ||
They found that shorter DNA strands produced denser and more stable monolayers due to their rigid, rod-like geometry, whereas longer strands exhibited flatter configurations with multiple substrate contacts, reducing surface coverage.48 Similarly, the desorption phenomenon is more pronounced as the DNA length increases. It was further concluded that oligonucleotides shorter than 24 bases maintain a nearly constant surface density, as their orientation and packing remain consistent within the rod-like regime. In contrast, longer strands increasingly behave like flexible polymers, significantly reducing surface density and increasing desorption potential. Thus, optimizing aptamer length for each target is critical to balance surface coverage, probe flexibility, and structural response.
3.6 The redox reporters and stability
Given the small electrochemical window available on a gold electrode, the number of functional redox labels to be utilized in E-AB sensors is limited. Electrochemical behaviour of most relevant redox molecules that can be used as a label candidate has been illustrated in Fig. 7.49 To date according to the literature, widely used reporters are methylene blue49 (E°′ = −260 mV vs. Ag|AgCl), and anthraquinone (E°′ = −440 mV vs. Ag|AgCl).50 These two molecules can undergo a reversible two-electron and one proton electrochemical reactions. One limitation, however, is that both exhibit pH-dependent redox behaviour, as the proton-coupled nature of their electron transfer causes shifts in potential under varying pH conditions.51 For example, their application in biological fluids with fluctuating pH, like sweat, will influence their electrochemistry and thus affecting E-AB signalling current. Ferrocene (E°′ = +220 mV vs. Ag|AgCl) in contrast exhibits a pH-insensitive one-electron transfer reaction but can be chemically affected by pH.52 More importantly, it has been shown that the oxidized ferrocenium form is susceptible for nucleophilic reaction with even weak nucleophilic agents like chlorides which are normally abundant in biological fluids.52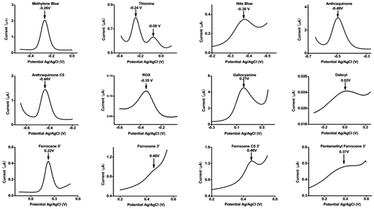 | ||
| Fig. 7 Electrochemical behaviour of different redox candidate. As one can see, dabcyl and ROX, for example, fail to produce clear oxidation and reduction peaks when conjugated to DNA and interrogated using standard square wave voltammetry parameters, and thionine exhibits two peaks in the relevant potential window. In case of ferrocene, three ferrocene-containing constructs were employed: one in which the ferrocene is conjugated directly on to an amine appended to the 5′ end of the DNA, a second in which the ferrocene is conjugated directly on to an amine appended to the 3′ end of the DNA, and a third, ferrocene C5, in which there is an additional spacer between ferrocene and the amide linkage to the DNA. The highly sloping baselines observed at potentials below −0.5 V and above 0.5 V (vs. Ag/AgCl) are due to the reduction of oxygen and the subsequent generation of reactive oxygen species (at low potentials) and the oxidation of gold (at high potentials). These same effects cause significant degradation of the thiol-on-gold SAM; that is, some redox reporters fail because they, themselves fail, and others fail because they report at potentials at which SAM stability is poor. Reproduced from ref. 49 with permission from American Chemical Society,49 copyright 2016. | ||
Because the oxidized form of ferrocene is positively charged, applying a positive potential may promote nonspecific electrostatic interactions and, in certain conditions, contribute to gold surface oxidation or etching. These potential risks caution in using ferrocene as a redox reporter in E-AB sensor fabrication. Anthraquinone provides good chemical stability particularly in chloride media, but has a reduction potential that overlaps with the onset of oxygen reduction on gold.49 Methylene blue, unlike ferrocene and anthraquinone, undergoes an electrochemically stable electron transfer reaction and has a reduction potential distinct from background electrochemical processes.49 The only compromise that needs to be taken into account is the pH variation control to ensure reproducibility in its electrochemical behaviour. Signal drifting is one of the most common characteristics of the redox molecules used as reporters regardless of the matrices whether it is simple buffer or human serum. Recently, a survey on a large set of potential redox reporters (more than a dozens) was conducted jointly by Ricci and Plaxco49 in order to find out which one demonstrates long-duration stability. Their work demonstrates that the performance of methylene blue-based E-AB sensor is unmatched where the sensor's stability against repeated scanning even in complex environments was significantly superior to its alternatives (Fig. 8), and so, has proven to be the most commonly used reporter in the fabrication of E-AB sensors.49
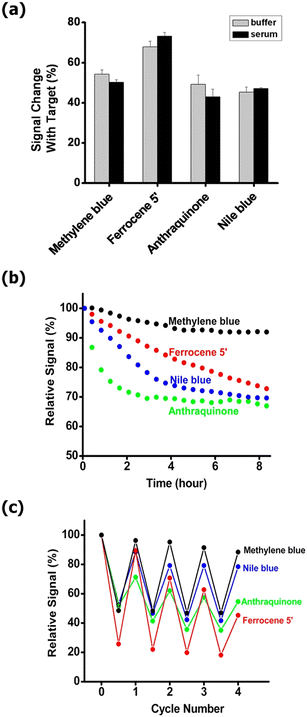 | ||
| Fig. 8 (A) Sensors fabricated with methylene blue, ferrocene, anthraquinone, or Nile blue exhibit similar signal gain in response to target binding whether deployed in simple buffer solutions or in 20% blood serum. (B) They all drift significantly, however, when repeatedly scanned in 20% serum over the course of hours, with methylene blue exhibiting the least drift. (C) Methylene blue-based sensors are likewise the most stable when the sensors are challenged with multiple cycles of hybridization (with saturating target) and regeneration (via di-water wash) in 20% blood serum. Reproduced from ref. 49 with permission from American Chemical Society,49 copyright 2016. | ||
The mechanism of target-induced signalling in E-AB sensors is based on the collision frequency of the redox tag and the electrode surface,53 consequently it seems to be reasonable to argue that the spatial position of the redox reporter within the DNA strand should be critical in its signalling behaviour. To elucidate this, Mayer et al.,54 carried out and detailed quantitative comparative experiment based on three aptamer probes each labelled with methylene blue in the distal end, middle, and proximal end (Fig. 9).
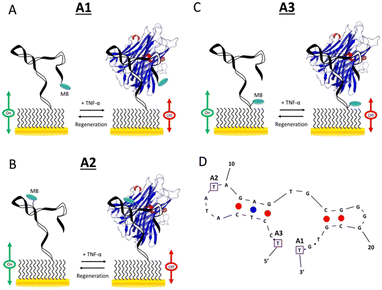 | ||
| Fig. 9 (A–C) Schematic illustrations of the three E-AB sensors. (D) The optimal (minimum energy) structure of the aptamer predicted by Mfold under the following conditions: 0.1 M Na(I), 22 °C. Also included are the locations of the MB redox label in the three different sensor architectures. Reproduced from ref. 54 with permission from Elsevier,54 copyright 2018. | ||
3.7 Stability and analytical performance
Optimized E-AB interface design has been shown to directly influence both analytical performance and operational stability. Early structure-switching electrochemical aptamer sensors reported detection limits in the low-nanomolar regime (e.g., thrombin detection at ∼3 nM (ref. 33)) and paper-based implementations have reported LODs on the order of tens of nanomolar (e.g., 16 nM thrombin; 30 nM DNA).55 In contrast, optimized E-AB interfaces have enabled detection limits in the picomolar range for protein targets directly in serum,56 showing a shift of ∼102–103× in sensitivity depending on target, format, and matrix.Beyond sensitivity, advances in interfacial stabilization have significantly extended operational lifetimes. Early E-AB sensors often exhibited pronounced signal drift within hours of continuous interrogation in biological media, whereas an improved monolayer cohesion and antifouling surface chemistries achieve stable operation over multiple days, and in some cases up to one week, in serum at physiological temperature.57 In that work, optimized passivation layers demonstrated markedly improved robustness under frequent electrochemical interrogation, with approximately 50% less signal loss over seven days compared to conventional mercaptohexanol-based monolayers.57 Complementary biomimetic strategies, such as phosphatidylcholine-terminated monolayers, have further reduced baseline drift in flowing whole blood from ∼70% to only a few percent over several hours, enabling in vivo deployment without physical membranes or active drift-correction algorithms.58 Importantly, analytical performance in E-AB sensors is not defined by detection limits alone. Signal gain and drift are strongly governed by interface chemistry, redox reporter selection, and electrochemical interrogation parameters. Systematic optimization of square-wave voltammetry conditions has been shown to yield greater than two-fold increases in signal gain in E-DNA and E-AB platforms,53 while comparative studies of redox reporters consistently identify methylene blue as providing superior long-term stability under repeated interrogation in complex biological matrices.49 Collectively, these quantitative benchmarks demonstrate that recent progress in interfacial engineering has transformed E-AB sensors from short-term proof-of-concept devices into robust platforms increasingly capable of high-accuracy operation in physiologically relevant environments.
They examined the sensor stability, the detection limit, the reusability, and the selectivity and were able to show that a redox reporter positioned at the distal end of the aptamer probe generated the best performances for this sensor design without interfering with monolayer formation and target binding. Furthermore, they argued that the internally conjugated methylene blue is not advantageous as it not only affects the probe structure but also imposes a potential interference in target binding. Therefore, the proximal attached methylene blue tends to negatively impact the monolayer formation, which can compromise the stability and thereby the performances of E-AB sensor.
4. Operational parameters
The signal changes upon target binding are not only dependant on the sensor fabrication but also are strongly impacted by the electrochemical interrogation parameters. Given that the motional dynamics and the change in dynamics of the probe aptamer play a pivotal role in the signalling current behaviour, it is reasonable to argue that the time scale and frequency of the voltammetric interrogation are critical to the sensor optimization and fabrication.53Most of the techniques featuring frequency element in their current data acquisition including cyclic, alternating-current (ACV), square-wave (SWV), and differential-pulse voltammetry are all suitable methods for monitoring aptamer conformation changes and the resulting changes in corresponding charge kinetics. The signal evolution from E-AB sensor involves two different striking dynamics (bound and unbound aptamer conformations) which is disturbed with target biding. E-AB sensors will have an optimum interrogation frequency range where the measured current exclusively originated from the faradic conversion of aptamer-bound reporters.59
At interrogation frequencies below 10 Hz, the current evolution is sensitive to non-faradic currents originating for example from side reactions from metal impurities or gases (e.g., the reduction of oxygen) and vibrational or electronic noise. At frequencies higher than 1000 Hz the contribution of the double layer60 and electronic noise are more pronounced.61 Additionally, the capacitive background in biological matrices can severely affect measurement reproducibility, especially at sub-optimal frequencies. This makes frequency-dependent optimization critical for achieving reliable signal output in complex samples such as serum or plasma. As shown in the Fig. 10, plotting the peak current/frequency vs. log frequency before and after target addition gives a map that helps navigation of the optimal interrogation frequency where the current is solely acquired from the bound and unbound probe electrons transfer. Here it shows a maximum electron transfer rate of 60 Hz for the unbound and of 500 Hz for target-bound states of the E-AB sensor.59 This divergence underscores the need for frequency scanning during sensor development to locate the differential signal window, which enables high signal-to-noise ratios and target specificity. Moreover, beyond frequency alone, the amplitude and pulse width in SWV or DPV can also affect the redox signal resolution and should be co-optimized with frequency. Therefore, the development of a map, in which the signal gain is plotted against large ranges of interrogating frequencies is highly recommended. It will allow for the rational identification of optimal sensing windows that maximize discrimination between baseline and target-bound states.
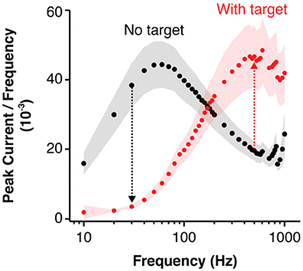 | ||
| Fig. 10 E-AB sensors have an optimal interrogation frequency range irrespective of interrogation technique. Here, one can determine the frequency ranges in which the bound and un-bound probe aptamer each produces the highest faradic signal current. Reproduced from ref. 59 with permission from American Chemical Society,59 copyright 2019. | ||
5. Conclusion
Recent advancements in biosensing have positioned DNA and RNA aptamer-based technologies, particularly E-AB sensors, as powerful platforms for real-time, label-free, and highly selective detection. Their operation through physical mechanisms such as conformational switching and electron transfer modulation, rather than enzymatic catalysis, enables exceptional stability in complex biological matrices and supports applications ranging from intraoperative diagnostics to continuous molecular monitoring. However, the clinical translation of E-AB sensors remains constrained by challenges in interfacial chemistry, probe design, biocompatibility, and electroanalytical robustness. This review has highlighted strategies addressing these barriers, including advances in surface functionalization, redox reporter selection, antifouling monolayers, and frequency-optimized interrogation methods that collectively improve sensitivity, reproducibility, and operational stability. By integrating insights from materials science and electrochemistry, this work outlines a framework for rational E-AB sensor design. The implementation of E-AB sensors in clinical settings could rely on several emerging technologies. The rise of artificial intelligence, machine-learning, and computational modelling could be explored to further optimize sensor interfacial design aspects, including probe packing density, backfilling agent, as well as operational parameters. In parallel, the integration of E-AB sensors with microfluidics offers an interesting approach towards miniaturization of such sensors. Future progress will also hinge on achieving long-term stability, antifouling resilience, and seamless integration into real-world diagnostic platforms, paving the way for next-generation biosensors capable of reliable in vivo and point-of-care operation.Author contributions
The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript Conceptualization, A. K. and G. E. M.; formal analysis, A. K.; resources, G. E. M.; writing – original draft preparation, A. K. and Y. R.; writing – review and editing, E. J. H. and G. E. M.; supervision, G. E. M.; funding acquisition, G. E. M.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included, and no new data were generated or analysed as part of this review.Acknowledgements
This research was funded by Merle, NSERC discovery and FRQS chercheur boursier J1.References
- C. Wang, M. Liu, Z. F. Wang, S. Li, Y. Deng and N. Y. He, Nano Today, 2021, 37, 101092 CrossRef CAS PubMed.
- J. H. T. Luong, K. B. Male and J. D. Glennon, Biotechnol. Adv., 2008, 26, 492–500 CrossRef CAS PubMed.
- (a) N. S. Que-Gewirth and B. A. Sullenger, Gene Ther., 2007, 14, 283–291 CrossRef CAS PubMed; (b) C. M. Dollins, S. Nair and B. A. Sullenger, Hum. Gene Ther., 2008, 19, 443–450 CrossRef CAS PubMed; (c) S. P. Song, L. H. Wang, J. Li, J. L. Zhao and C. H. Fan, TrAC, Trends Anal. Chem., 2008, 27, 108–117 CrossRef CAS.
- R. Reid, B. Chatterjee, S. J. Das, S. Ghosh and T. K. Sharma, Anal. Biochem, 2020, 593, 113574 CrossRef CAS PubMed.
- Y. Xiao, A. A. Lubin, A. J. Heeger and K. W. Plaxco, Angew. Chem., Int. Ed., 2005, 44, 5456–5459 CrossRef CAS PubMed.
- (a) A. D. Ellington and J. W. Szostak, Nature, 1990, 346, 818–822 CrossRef CAS PubMed; (b) L. C. Bock, L. C. Griffin, J. A. Latham, E. H. Vermaas and J. J. Toole, Nature, 1992, 355, 564–566 CrossRef CAS PubMed; (c) L. Gold, J. Mol. Evol., 2015, 81, 140–143 CrossRef CAS PubMed.
- E. Roth, A. Glick Azaria, O. Girshevitz, A. Bitler and Y. Garini, Nano Lett., 2018, 18, 6703–6709 CrossRef CAS PubMed.
- J. Wang, Biosens. Bioelectron., 2006, 21, 1887–1892 CrossRef CAS PubMed.
- (a) F. Ricci, A. Valée-Bélisle, A. J. Simon, A. Porchetta and K. W. Plaxco, Acc. Chem. Res., 2016, 49, 1884–1892 CrossRef CAS; (b) A. A. Lubin and K. W. Plaxco, Acc. Chem. Res., 2010, 43, 496–505 CrossRef CAS PubMed.
- P. D. P. Swetha, J. Sonia, K. Sapna and K. S. Prasad, Curr. Opin. Electrochem., 2021, 30, 100829 CrossRef PubMed.
- Z. Y. Lu, K. Xu, K. Xiao, Q. B. Xu, L. Wang, P. Li, J. H. Zhou, D. Zhao, L. B. Bai, Y. H. Cheng and W. Huang, npj Flexible Electron., 2025, 9, 9 CrossRef.
- Z. R. Liao, Y. Zhang, Y. R. Li, Y. F. Miao, S. M. Gao, F. K. Lin, Y. L. Deng and L. N. Geng, Biosens. Bioelectron., 2019, 126, 697–706 CrossRef CAS PubMed.
- C. A. Vu and W. Y. Chen, Sensors, 2019, 19, 4214 CrossRef CAS PubMed.
- L. R. Schoukroun-Barnes, F. C. Macazo, B. Gutierrez, J. Lottermoser, J. Liu and R. J. White, Annu. Rev. Anal. Chem., 2016, 9, 163–181 CrossRef CAS.
- H. Yoo, H. Jo and S. S. Oh, Mater. Adv., 2020, 1, 2663–2687 RSC.
- N. Fontaine and P. Dauphin-Ducharme, Curr. Opin. Electrochem., 2023, 41, 101361 CrossRef CAS.
- N. Arroyo-Currás, P. Dauphin-Ducharme, K. Scida and J. L. Chávez, Anal. Methods, 2020, 12, 1288–1310 RSC.
- K. Han, L. Chen, Z. S. Lin and G. X. Li, Electrochem. Commun., 2009, 11, 157–160 CrossRef CAS.
- J. W. Liu and Y. Lu, Angew. Chem., Int. Ed., 2006, 45, 90–94 CrossRef CAS PubMed.
- W. A. Zhao, W. Chiuman, J. C. F. Lam, S. A. McManus, W. Chen, Y. G. Cui, R. Pelton, M. A. Brook and Y. F. Li, J. Am. Chem. Soc., 2008, 130, 3610–3618 CrossRef CAS PubMed.
- J. Wang, W. Y. Meng, X. F. Zheng, S. L. Liu and G. X. Li, Biosens. Bioelectron., 2009, 24, 1598–1602 CrossRef CAS PubMed.
- Y. Liu, Z. Matharu, M. C. Howland, A. Revzin and A. L. Simonian, Anal. Bioanal. Chem., 2012, 404, 1181–1196 CrossRef CAS PubMed.
- A. L. Furst, M. G. Hill and J. K. Barton, Langmuir, 2015, 31, 6554–6562 CrossRef CAS PubMed.
- S. W. Abeykoon and R. J. White, ACS Meas. Sci. Au, 2022, 3, 1–9 Search PubMed.
- Y. P. Liu, B. B. Zhang, X. Y. Wu, F. Wang, Z. Y. Yang, M. Y. Li, K. X. Sheng, Y. Yan, L. Zhu, H. Jing, Y. M. Wu, L. L. Hu, Y. Y. Yu and C. L. Li, Biosens. Bioelectron., 2025, 275, 117236 CrossRef CAS PubMed.
- B. R. Baker, R. Y. Lai, M. S. Wood, E. H. Doctor, A. J. Heeger and K. W. Plaxco, J. Am. Chem. Soc., 2006, 128, 3138–3139 CrossRef CAS PubMed.
- R. Nutiu and Y. F. Li, J. Am. Chem. Soc., 2003, 125, 4771–4778 CrossRef CAS PubMed.
- R. J. White, N. Phares, A. A. Lubin, Y. Xiao and K. W. Plaxco, Langmuir, 2008, 24, 10513–10518 CrossRef CAS PubMed.
- A. W. Peterson, R. J. Heaton and R. M. Georgiadis, Nucleic Acids Res., 2001, 29, 5163–5168 CrossRef CAS PubMed.
- (a) A. B. Steel, T. M. Herne and M. J. Tarlov, Anal. Chem., 1998, 70, 4670–4677 CrossRef CAS PubMed; (b) L. M. Demers, C. A. Mirkin, R. C. Mucic, R. A. Reynolds, 3rd, R. L. Letsinger, R. Elghanian and G. Viswanadham, Anal. Chem., 2000, 72, 5535–5541 CrossRef CAS PubMed.
- K. K. Leung, A. D. Gaxiola, H. Z. Yu and D. Bizzotto, Electrochim. Acta, 2018, 261, 188–197 CrossRef CAS.
- P. V. Riccelli, F. Merante, K. T. Leung, S. Bortolin, R. L. Zastawny, R. Janeczko and A. S. Benight, Nucleic Acids Res., 2001, 29, 996–1004 CrossRef CAS PubMed.
- (a) A. Meller, L. Nivon and D. Branton, Phys. Rev. Lett., 2001, 86, 3435–3438 CrossRef CAS PubMed; (b) A. Reuter, W. U. Dittmer and F. C. Simmel, Eur. Phys. J. E: Soft Matter Biol. Phys., 2007, 22, 33–40 CrossRef CAS PubMed.
- A. L. Furst, M. G. Hill and J. K. Barton, Langmuir, 2013, 29, 16141–16149 CrossRef CAS PubMed.
- K. S. Sykes, L. F. L. Oliveira, G. Stan and R. J. White, Langmuir, 2019, 35, 12962–12970 CrossRef CAS PubMed.
- N. Khuda, S. Somasundaram, A. B. Urgunde and C. J. Easley, ACS Appl. Mater. Interfaces, 2023, 15, 5019–5027 CrossRef CAS PubMed.
- Y. Xiao, T. Uzawa, R. J. White, D. Demartini and K. W. Plaxco, Electroanalysis, 2009, 21, 1267–1271 CrossRef CAS PubMed.
- (a) H. Ron and I. Rubinstein, J. Am. Chem. Soc., 1998, 120, 13444–13452 CrossRef CAS; (b) D. E. Weisshaar, B. D. Lamp and M. D. Porter, J. Am. Chem. Soc., 1992, 114, 5860–5862 CrossRef CAS.
- F. Y. Ma and R. B. Lennox, Langmuir, 2000, 16, 6188–6190 CrossRef CAS.
- (a) D. Jambrec, F. Conzuelo, A. Estrada-Vargas and W. Schuhmann, ChemElectroChem, 2016, 3, 1484–1489 CrossRef CAS; (b) D. Jambrec, M. Gebala, F. La Mantia and W. Schuhmann, Angew. Chem., Int. Ed., 2015, 54, 15064–15068 CrossRef CAS PubMed.
- (a) R. Levicky, T. M. Herne, M. J. Tarlov and S. K. Satija, J. Am. Chem. Soc., 1998, 120, 9787–9792 CrossRef CAS; (b) K. A. Peterlinz, R. M. Georgiadis, T. M. Herne and M. J. Tarlov, J. Am. Chem. Soc., 1997, 119, 3401–3402 CrossRef CAS; (c) T. M. Herne and M. J. Tarlov, J. Am. Chem. Soc., 1997, 119, 8916–8920 CrossRef CAS.
- P. Gong, C. Y. Lee, L. J. Gamble, D. G. Castner and D. W. Grainger, Anal. Chem., 2006, 78, 3326–3334 CrossRef CAS PubMed.
- F. Ricci, N. Zari, F. Caprio, S. Recine, A. Amine, D. Moscone, G. Palleschi and K. W. Plaxco, Bioelectrochemistry, 2009, 76, 208–213 CrossRef CAS PubMed.
- M. Mrksich, G. B. Sigal and G. M. Whitesides, Langmuir, 1995, 11, 4383–4385 CrossRef CAS.
- H. Li, P. Dauphin-Ducharme, N. Arroyo-Currás, C. H. Tran, P. A. Vieira, S. G. Li, C. Shin, J. Somerson, T. E. Kippin and K. W. Plaxco, Angew. Chem., Int. Ed., 2017, 56, 7492–7495 CrossRef CAS PubMed.
- V. Clark, M. A. Pellitero and N. Arroyo-Currás, Anal. Chem., 2023, 95, 4974–4983 CrossRef CAS PubMed.
- A. B. Steel, R. L. Levicky, T. M. Herne and M. J. Tarlov, Biophys. J., 2000, 79, 975–981 CrossRef CAS PubMed.
- (a) B. Tinland, A. Pluen, J. Sturm and G. Weill, Macromolecules, 1997, 30, 5763–5765 CrossRef CAS; (b) D. Rekesh, Y. Lyubchenko, L. S. Shlyakhtenko and S. M. Lindsay, Biophys. J., 1996, 71, 1079–1086 CrossRef CAS PubMed; (c) C. A. Mirkin, R. L. Letsinger, R. C. Mucic and J. J. Storhoff, Nature, 1996, 382, 607–609 CrossRef CAS PubMed.
- D. Kang, F. Ricci, R. J. White and K. W. Plaxco, Anal. Chem., 2016, 88, 10452–10458 CrossRef CAS PubMed.
- H. Li, N. Arroyo-Currás, D. Kang, F. Ricci and K. W. Plaxco, J. Am. Chem. Soc., 2016, 138, 15809–15812 CrossRef CAS PubMed.
- J. D. Mahlum, M. A. Pellitero and N. Arroyo-Currás, J. Phys. Chem. C, 2021, 125, 9038–9049 CrossRef CAS.
- A. R. K. R. Prins and A. G. T. G. Kortbeek, J. Organomet. Chem., 1972, 39, 335–344 CrossRef.
- P. Dauphin-Ducharme and K. W. Plaxco, Anal. Chem., 2016, 88, 11654–11662 CrossRef CAS PubMed.
- M. D. Mayer and R. Y. Lai, Talanta, 2018, 189, 585–591 CrossRef CAS PubMed.
- J. C. Cunningham, N. J. Brenes and R. M. Crooks, Anal. Chem., 2014, 86, 6166–6170 CrossRef CAS PubMed.
- (a) R. Y. Lai, K. W. Plaxco and A. J. Heeger, Anal. Chem., 2007, 79, 229–233 CrossRef CAS PubMed; (b) C. E. Immoos, S. J. Lee and M. W. Grinstaff, J. Am. Chem. Soc., 2004, 126, 10814–10815 CrossRef CAS PubMed.
- Z. Watkins, A. Karajic, T. Young, R. White and J. Heikenfeld, ACS Sens., 2023, 8, 1119–1131 CrossRef CAS PubMed.
- H. Li, P. Dauphin-Ducharme, N. Arroyo-Currás, C. H. Tran, P. A. Vieira, S. Li, C. Shin, J. Somerson, T. E. Kippin and K. W. Plaxco, Angew. Chem., Int. Ed., 2017, 56, 7492–7495 CrossRef CAS PubMed.
- S. D. Curtis, K. L. Ploense, M. Kurnik, G. Ortega, C. Parolo, T. E. Kippin, K. W. Plaxco and N. Arroyo-Currás, Anal. Chem., 2019, 91, 12321–12328 CrossRef CAS PubMed.
- P. Dauphin-Ducharme, N. Arroyo-Currás, R. Adhikari, J. Somerson, G. Ortega, D. E. Makarov and K. W. Plaxco, J. Phys. Chem. C, 2018, 122, 21441–21448 CrossRef CAS.
- R. J. White and K. W. Plaxco, Anal. Chem., 2010, 82, 73–76 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |