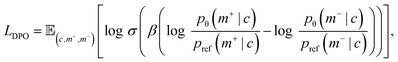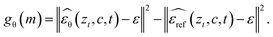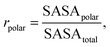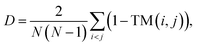Open Access Article
Open Access ArticleDesign of permeability-optimized target-binding macrocycles via direct preference optimization
Heqi Sun a,
Hong Tana,
Yanyi Chub,
Jiayi Li
a,
Hong Tana,
Yanyi Chub,
Jiayi Li a,
Ruixuan Wanga and
Dong-Qing Wei
a,
Ruixuan Wanga and
Dong-Qing Wei *acd
*acd
aState Key Laboratory of Microbial Metabolism, Joint International Research Laboratory of Metabolic & Developmental Sciences and School of Life Sciences and Biotechnology, Shanghai Jiao Tong University, Shanghai, 200040, P. R. China. E-mail: dqwei@sjtu.edu.cn
bKey Laboratory of RNA Innovation, Science and Engineering, CAS Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology, University of Chinese Academy of Sciences, Chinese Academy of Sciences, Shanghai, China
cHebi Branch, Henan Academy of Sciences, Qishui Guang East, Qibin District, Hebi Henan, 458030, P. R. China
dQihe Laboratory, Qishui Guang East, Qibin District, Hebi, Henan 458030, P. R. China
First published on 9th April 2026
Abstract
Macrocyclic peptides represent a promising therapeutic modality for challenging targets, such as protein–protein interactions. However, their clinical utility is often limited by inadequate membrane permeability, which restricts both intracellular target access and oral bioavailability. Existing structure-based generative methods for cyclic peptide design prioritize structural validity and binding affinity, yet lack mechanisms to co-optimize membrane permeability. Here we present CycDiff-DPO, a preference-aligned diffusion framework for designing target-specific macrocyclic peptide binders with optimized membrane permeability. By ranking sampled candidates with a Caco-2 permeability predictor and constructing preference pairs, CycDiff-DPO aligns the generative distribution toward permeability-favorable chemical space while preserving target binding competence. We benchmarked CycDiff-DPO across 56 protein targets, finding higher predicted Caco-2 and PAMPA permeability across multiple independent predictors, alongside superior binding energetics and comparable stereochemical quality relative to baseline methods. Case studies on Keap1–Nrf2 and SPSB2–iNOS confirm that top designs recapitulate hot-spot interactions and maintain stable bound poses in molecular dynamics simulations. CycDiff-DPO provides a framework for permeability-enhanced macrocyclic peptide design with broad therapeutic applications.
Introduction
Macrocyclic peptides have emerged as a promising therapeutic modality that occupies an intermediate space between small-molecule drugs and large biologics.1–3 While biologics can achieve high affinity and selectivity, their size and polarity generally limit them to extracellular targets.4 Small molecules, in contrast, can access intracellular targets but are often ill-suited for modulating protein–protein interactions (PPIs), which typically involve large, shallow interfaces that lack the deep hydrophobic pockets required for high-affinity binding.5,6 By bridging these two regimes, macrocyclic peptides offer sufficient molecular size to engage such challenging interfaces while benefiting from conformational constraint.7 Their rigidity can enhance binding specificity and improve proteolytic stability relative to linear counterparts.1,8,9 The translational impact of cyclic peptides is evident from their presence among approved drugs across oncology and autoimmune disease.10–12Traditionally, protein-binding macrocyclic peptides have been discovered either from natural products or through display-based screening of large randomized libraries.2,3 Although natural products can yield potent starting points, their synthetic complexity and limited amenability to systematic diversification often hinder analogue generation and lead optimization.13 Display technologies can explore broad sequence space, yet iterative selection and downstream characterization are labor-intensive and still undersample the chemical and topological diversity accessible to macrocyclic scaffolds. As a result, it is difficult to optimize binding and developability in parallel, especially membrane permeability, since this multiparameter profile depends on precise control of ring architecture and physicochemical properties that is hard to maintain during library design and selection.14
More broadly, computational methods have been applied to quantitatively model PPI interface energetics; for example, Wang et al. combined persistent-homology descriptors of 3D complex structures with a CNN and gradient-boosting ensemble to predict mutation-induced binding affinity changes.15 Building on such structure-based modeling foundations, recent advances in structure-based deep learning have enabled the de novo design of cyclic peptide binders with defined target specificity. Representative strategies include diffusion-based target-conditioned macrocycle generation, such as RFpeptides16 and CP-composer;17 AlphaFold-guided sequence optimization like EvoBind2;18 and reinforcement-learning-based methods, including CYC_BUILDER19 and HighPlay.20 Despite this progress, most models focus on optimizing binding and structural validity while neglecting membrane permeability, which is a key determinant of oral bioavailability and intracellular target access.21–23 Incorporating permeability into structure-guided design is therefore an important open challenge, yet the choice of strategy is constrained by data availability. Conditioning the generator directly on permeability requires paired structure–permeability data, which remain scarce for cyclic peptides. Preference-based alignment offers an alternative by steering a pretrained model with ranked feedback. Among such approaches, direct preference optimization (DPO) is particularly suited to this setting,24 as it learns directly from winner–loser pairs without requiring a separate reward model, and such pairs can be constructed from predictor-based rankings when experimental data are limited. DPO has recently been extended to diffusion-model alignment in molecular and antibody design.25–27
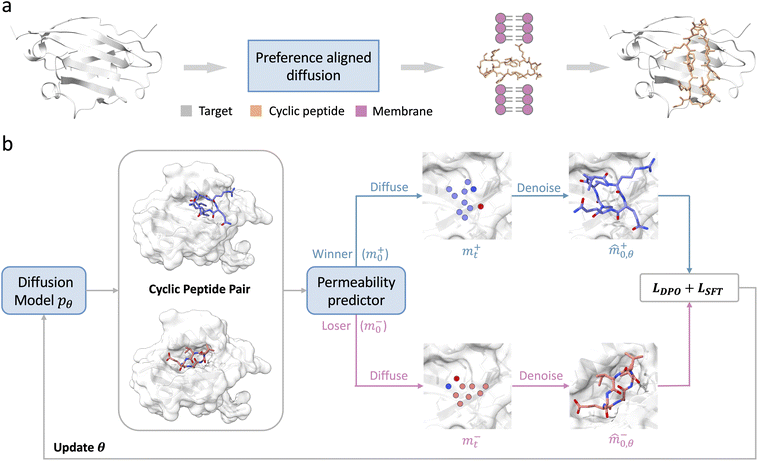
Here we present CycDiff-DPO (Fig. 1), a preference-aligned diffusion framework that generates target-binding macrocyclic peptides while co-optimizing membrane permeability. Starting from a pretrained diffusion model, CycDiff-DPO first constructs target-specific winner–loser pairs by sampling macrocycles and ranking them with a Caco-2 permeability predictor. It then applies diffusion-based DPO to shift the generative distribution toward more permeable candidates while preserving binding competence. We benchmarked CycDiff-DPO on 56 unseen protein targets and observed consistent enrichment of permeability-favorable chemical space, including higher predicted Caco-2 permeability, increased lipophilicity, and reduced exposed polarity, while maintaining stereochemical plausibility and structural diversity. These improvements generalized across multiple external predictors and extended to PAMPA, an orthogonal passive-diffusion assay. Importantly, binding-related metrics, including Rosetta interface energies and interaction patterns, remained stable. Case studies on intracellular targets Keap1–Nrf2 and SPSB2–iNOS further showed that CycDiff-DPO produced permeability-enriched candidates that retained key hot-spot contacts and stable bound poses. CycDiff-DPO thus provides a general framework for designing permeability-enhanced macrocyclic binders, and may further facilitate the development of orally bioavailable cyclic peptide therapeutics.
Results
Overview of CycDiff-DPO
CycDiff-DPO presents a direct permeability-guided preference optimization method for designing cell-permeable cyclic peptides with target specificity (Fig. 1a). It drives a controlled distributional shift by preferentially increasing the generation likelihood of highly permeable candidates while suppressing poorly permeable ones within the same target pocket. The method consists of three modules: candidate sampling, permeability evaluation, and preference-based alignment (Fig. 1b). Starting from a pretrained generator,17 we sample a pool of cyclic peptides for each target context. Each design is then scored using an in-house Caco-2 permeability predictor, and candidates are ranked within target to construct winner–loser preference pairs by matching higher-scoring peptides (winners) to lower-scoring peptides (losers). Using these target-specific preference pairs, we align the diffusion generator via diffusion-based DPO, optimizing the current model (θ) to increase the relative likelihood of winners over losers with respect to a frozen reference model (θref) initialized from the same pretrained checkpoint. To stabilize optimization and preserve sequence–structure fidelity, we additionally apply an SFT-style reconstruction regularizer on winner samples. Together, this preference-driven alignment enables an efficient shift toward permeability-favorable binders without compromising geometric validity or binding-competent conformations.Performance on target-conditioned cyclic peptide design
To systematically evaluate the stereochemical validity, physicochemical properties, and diversity of cyclic peptides generated by CycDiff-DPO, we benchmarked the model on a non-redundant dataset of 56 curated protein–peptide complexes collected from the literature.28 The reference peptide lengths ranged from 5 to 16 residues. Because CycDiff-DPO was tailored to target-conditioned macrocyclic peptide design, we restricted our comparisons to representative structure-based methods for macrocycle generation that can be run uniformly across the full 56-target benchmark. Specifically, we compared against RFpeptides16 for cyclic backbone design coupled with ProteinMPNN29 for sequence design, as well as sequence–structure co-design models CP-composer,17 PepFlow,30 and PepGLAD.31 Official pipelines and released weights were used for RFpeptides and CP-composer, while fine-tuned PepFlow and PepGLAD models trained on 71![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 867 cyclic peptide complexes were adopted from a benchmarking study.28 All generated peptides were evaluated using Ramachandran acceptance and favored rates, log
867 cyclic peptide complexes were adopted from a benchmarking study.28 All generated peptides were evaluated using Ramachandran acceptance and favored rates, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, polar SASA ratio, and structural diversity (see Methods for details).
P, polar SASA ratio, and structural diversity (see Methods for details).
As summarized in Table 1, CycDiff-DPO ranks third in stereochemical quality, achieving Ramachandran acceptance and favored rates of 72.5% and 42.2%, respectively—comparable to CP-composer (72.2%/40.0%) and PepGLAD (71.9%/40.5%). This indicates that our preference-guided optimization does not compromise structure plausibility relative to other co-design models. In terms of physicochemical profiles, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P is a widely used proxy for the balance between hydrophobicity and desolvation cost relevant to passive membrane partitioning.32 CycDiff-DPO produces the highest log
P is a widely used proxy for the balance between hydrophobicity and desolvation cost relevant to passive membrane partitioning.32 CycDiff-DPO produces the highest log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (−1.84) and the lowest polar SASA ratio (0.20) among all models, indicating reduced solvent-accessible polar surface exposure—an attribute often associated with permeable macrocycles.33 Importantly, these improvements are not achieved at the expense of molecular diversity. CycDiff-DPO attained a structural diversity score of 0.72, comparable to those of the baseline methods, demonstrating that the incorporation of DPO enhanced permeability-related properties while preserving broad coverage of the accessible conformational space.
P (−1.84) and the lowest polar SASA ratio (0.20) among all models, indicating reduced solvent-accessible polar surface exposure—an attribute often associated with permeable macrocycles.33 Importantly, these improvements are not achieved at the expense of molecular diversity. CycDiff-DPO attained a structural diversity score of 0.72, comparable to those of the baseline methods, demonstrating that the incorporation of DPO enhanced permeability-related properties while preserving broad coverage of the accessible conformational space.
| Model | Ramachandran accepted | Ramachandran favored | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P |
Polar SASA ratio | Diversity |
|---|---|---|---|---|---|
| CycDiff-DPO | 72.5% | 42.2% | −1.84 ± 3.09 | 0.20 ± 0.04 | 0.72 |
| CP-composer | 72.2% | 40.0% | −5.55 ± 3.52 | 0.23 ± 0.03 | 0.71 |
| RFpeptides | 95.8% | 83.4% | −6.72 ± 3.37 | 0.26 ± 0.06 | 0.73 |
| PepFlow (cyc) | 82.5% | 58.9% | −5.88 ± 3.01 | 0.24 ± 0.04 | 0.74 |
| PepGLAD (cyc) | 71.9% | 40.5% | −5.08 ± 2.47 | 0.23 ± 0.04 | 0.72 |
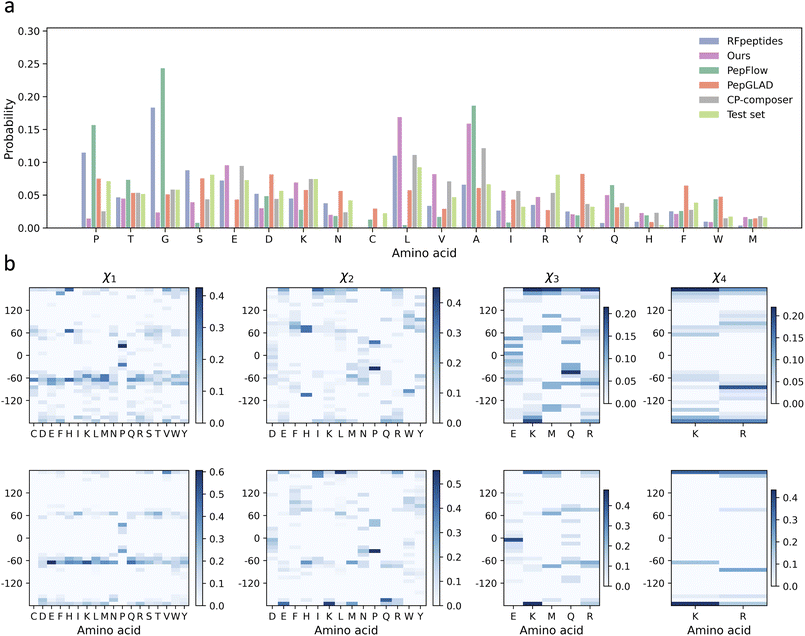
Next, we examined the amino acid composition of cyclic peptides generated by different models and compared it with the reference distribution from the test set (Fig. 2a). CycDiff-DPO showed a relatively low Kullback–Leibler (KL) divergence of 0.30. This value was substantially lower than those of RFpeptides (0.66) and PepFlow (1.25), and moderately higher than those of PepGLAD (0.11) and CP-composer (0.14). We observed modest enrichments of leucine and alanine, consistent with the highest log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P and lowest polar SASA ratio among all models (Table 1). Meanwhile, charged (E, K, R) and polar (T, Q) residues remain present at appreciable frequencies, reflecting their essential roles in forming hydrogen bonds and salt bridges at the peptide–receptor interface. These residue types are generally unfavorable for passive membrane permeability owing to desolvation penalties and increased polar surface exposure. However, the impact of the retained charged residues on membrane transit is likely mitigated by pKa shifts in the low-dielectric membrane interior, which can drive ionizable side chains toward neutralization at physiological pH, with reported shifts of +2 to +5 units for Glu34 and −4 to −5 units for Lys.35 The remaining polar residues (T, Q) are uncharged and therefore impose a much smaller barrier to membrane crossing than ionizable residues, as their membrane transfer free energies are only ∼3–5 kcal mol−1 compared to >14 kcal mol−1 for formally charged side chains.36 Importantly, we found no evidence of amino-acid-level mode collapse, a common failure mode in property-optimized generative models where the sequence space collapses to a narrow set of residue choices.
P and lowest polar SASA ratio among all models (Table 1). Meanwhile, charged (E, K, R) and polar (T, Q) residues remain present at appreciable frequencies, reflecting their essential roles in forming hydrogen bonds and salt bridges at the peptide–receptor interface. These residue types are generally unfavorable for passive membrane permeability owing to desolvation penalties and increased polar surface exposure. However, the impact of the retained charged residues on membrane transit is likely mitigated by pKa shifts in the low-dielectric membrane interior, which can drive ionizable side chains toward neutralization at physiological pH, with reported shifts of +2 to +5 units for Glu34 and −4 to −5 units for Lys.35 The remaining polar residues (T, Q) are uncharged and therefore impose a much smaller barrier to membrane crossing than ionizable residues, as their membrane transfer free energies are only ∼3–5 kcal mol−1 compared to >14 kcal mol−1 for formally charged side chains.36 Importantly, we found no evidence of amino-acid-level mode collapse, a common failure mode in property-optimized generative models where the sequence space collapses to a narrow set of residue choices.
To further assess the realism of local side-chain geometries, we analyzed the distributions of side-chain dihedral angles (χ1–χ4) for cyclic peptides generated by CycDiff-DPO and compared them with those in experimentally determined structures from the test set (Fig. 2b). Across all torsional degrees of freedom, the dihedral angle distributions closely matched the reference distributions. The agreement was strongest for χ1 and χ2. Higher-order torsions (χ3 and χ4) were more variable, but they still followed the same overall trends as the reference. Together, these results indicated that the designed cyclic peptides retained natural-like sequence statistics and locally realistic structural features.
Binding patterns of designed cyclic peptides
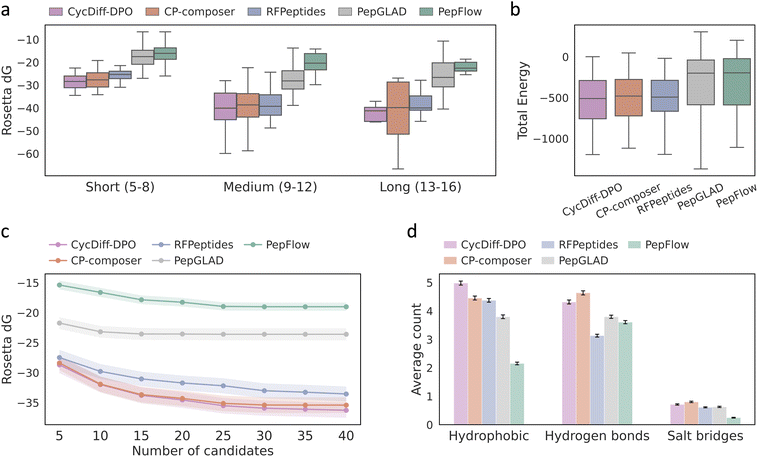
To assess whether permeability-guided preference alignment preserves target engagement, we compared binding energetics and interfacial quality for cyclic peptides generated by CycDiff-DPO and the baseline methods. For each target, peptide–receptor complexes were refined using a single round of Rosetta FastRelax and the best-scoring design per method was selected according to the Rosetta interface energy (ΔG).37 Across all peptide-length categories, CycDiff-DPO produced consistently more favorable interface energies than competing approaches (Fig. 3a). For most models, longer peptides generally exhibit lower interface energies, consistent with their ability to form larger binding interfaces and more extensive contacts with the target protein. We further examined binding quality as a function of the number of generated candidates per target. As shown in Fig. 3c, CycDiff-DPO maintained a clear advantage over baseline models across all sampling depths, with its average ΔG decreasing and gradually converging as the candidate pool expands. This convergence behavior indicated that the performance gains of CycDiff-DPO were not driven by isolated outliers, but instead reflect a systematic enrichment of high-affinity binders within the generated set. In line with the interface trends, the Rosetta total energies of the generated complexes were also shifted towards more favorable values for CycDiff-DPO (Fig. 3b), supporting that improved binding free energy was compatible with globally stable bound poses.Beyond energetic metrics, we characterized intermolecular contacts to obtain a chemically interpretable view of the binding modes. Using PLIP,38 we quantified hydrophobic contacts, hydrogen bonds, and salt bridges for the generated complexes. CycDiff-DPO designs exhibited the highest average number of hydrophobic contacts (Fig. 3d), consistent with tighter nonpolar packing and improved shape complementarity at the interface. Hydrogen-bond counts were modestly reduced relative to CP-composer, whereas salt-bridge counts remained comparable. This is expected under permeability-driven optimization, which enriches hydrophobic residues and reduces exposed polarity. Overall, CycDiff-DPO appears to preserve affinity primarily through tighter nonpolar packing at the interface.
Results on permeability optimization
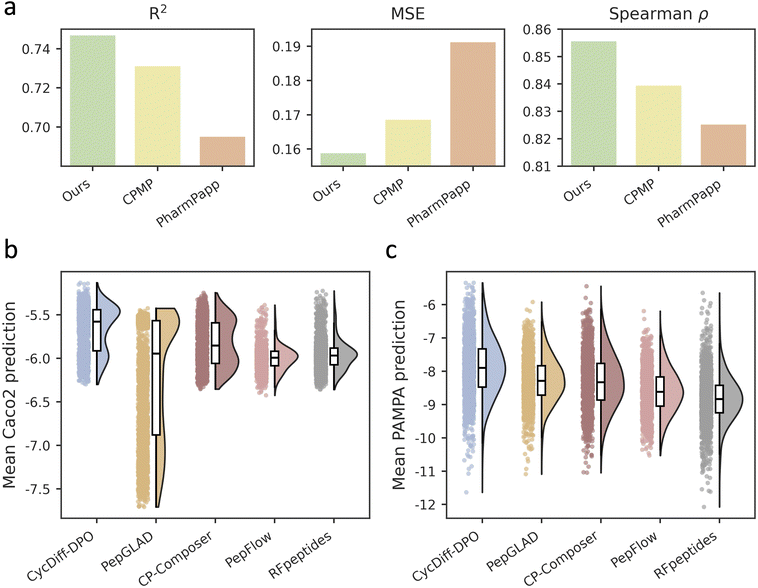
Preference optimization in this work was guided by Caco-2 permeability, a widely adopted and practically actionable metric in lead optimization that serves as a cell-based proxy for intestinal absorption. Compared with physicochemical descriptors (e.g., lipophilicity) and with PAMPA which primarily reflects passive diffusion, Caco-2 permeability provides a more integrated measure of membrane transport and can additionally reflect transporter-related liabilities such as efflux.39,40 This makes Caco-2 permeability a pragmatic optimization target for improving the developability of macrocyclic peptides. To ensure that our in-house Caco-2 predictor provides a reliable signal for constructing preference pairs, we benchmarked it on the CycPeptMPDB dataset21 using five-fold cross-validation, and compared it against two recent state-of-the-art deep learning models for cyclic peptide Caco-2 prediction (CPMP41 and PharmPapp42). As shown in Fig. 4a, our model achieved R2 = 0.747, MSE = 0.159, and Spearman's ρ = 0.856, outperforming CPMP and PharmPapp. Per-fold metrics across the five splits are shown in Table S1.Due to limited experimental throughput, we assessed membrane permeability of the designed cyclic peptides using machine-learning predictors. We evaluated each peptide set with multiple independent predictors and report results averaged across models to reduce dependence on any single evaluator, with per-predictor results provided in the Table S2 and S3. Since preference construction was guided by a Caco-2 signal, we additionally assessed PAMPA to test whether the observed gains extend to an assay that mainly reports passive transmembrane diffusion. For Caco-2 permeability, we used CPMP and PharmPapp as external predictors. For PAMPA permeability, we used DMPNN and AttentiveFP as the top-performing predictors reported in the most recent benchmarking study,43 together with CPMP. As shown in Fig. 4b, CycDiff-DPO yields the highest predicted Caco-2 permeability and the distribution is shifted towards higher values relative to all baselines. This advantage remains when each predictor is examined individually, indicating that the improvement is not driven by a single evaluation model. The same trend is observed for PAMPA (Fig. 4c), where CycDiff-DPO again attains the most favorable predictions across predictors despite PAMPA not being used to construct the preference signal. Therefore, permeability improvements generalized across predictors and assay modalities. To investigate whether the observed permeability gains are attributable to intramolecular hydrogen-bond (IMHB) shielding, we quantified IMHB counts for the generated peptides and compared them with experimentally characterized macrocycles. Across all five generative models, CycDiff-DPO did not increase the mean IMHB count relative to the baselines (Fig. S3). Consistently, analysis of CycPeptMPDB indicates that high-permeability cyclic peptides are not enriched in IMHBs; instead, the high-permeability subset shows a modestly reduced IMHB count for both Caco-2 and PAMPA measurements (Fig. S4). These observations suggest that IMHB shielding is unlikely to be the primary driver in our setting and are consistent with a permeability enhancement mechanism dominated by reduced global polarity. To account for solvent-dependent conformational reorganization, we performed CREST conformational searches under ALPB chloroform as a membrane-mimicking environment44,45 for representative subsets of experimentally characterized cyclic peptides from CycPeptMPDB dataset covering peptide lengths from 3 to 15 residues. Boltzmann-weighted IMHB counts showed no positive association with experimental permeability: in the Caco-2 dataset, a weak but statistically significant negative correlation was observed (Spearman r = −0.24, p = 0.042), although the difference between high- and low-permeability groups did not reach significance in a group comparison (Mann–Whitney p = 0.177). In the PAMPA dataset, neither correlation nor group comparison reached significance (Spearman r = −0.16, p = 0.19; Mann–Whitney p = 0.391; Fig. S5 and Table S4). In both assays, high-permeability peptides showed numerically lower mean IMHB counts, and no positive association was detected. These findings indicate that IMHB-mediated shielding does not emerge as a driver of permeability in this chemical space, consistent with a global depolarization mechanism.
Case studies
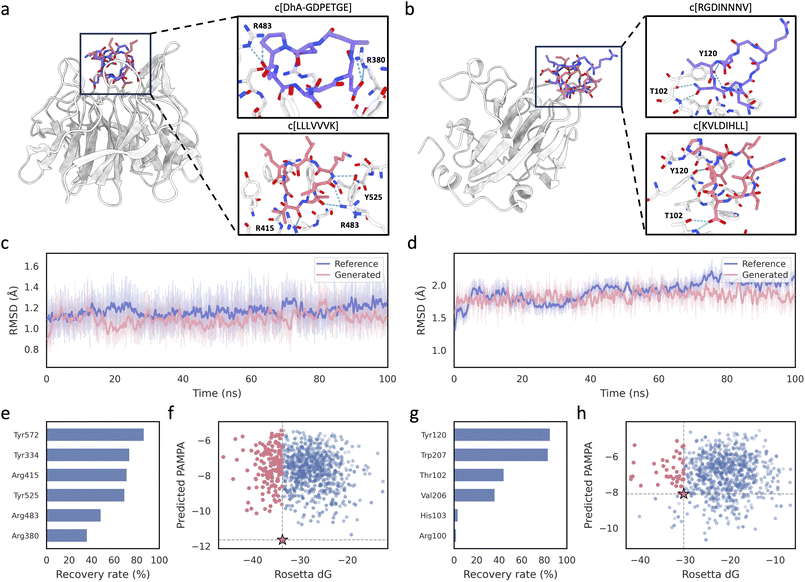
We further applied CycDiff-DPO to two clinically relevant intracellular PPI targets to test whether permeability-guided preference optimization can enrich cell-permeable cyclic peptide inhibitors without compromising target engagement. We selected Keap1–Nrf2, a central regulatory node in the cellular antioxidant response, because pharmacological modulation of the Keap1–Nrf2 axis is widely pursued for oxidative-stress–driven pathologies (e.g., chronic inflammation and degenerative disorders).46 We also selected SPSB2–iNOS, an intracellular interaction in which SPSB2 recruits ubiquitin machinery to control iNOS lifetime and thereby tunes nitric oxide output in inflammation and host defence; disrupting this interaction has been proposed as a route to prolong iNOS activity and enhance antimicrobial responses.47 For Keap1–Nrf2, we used a reported Keap1-bound cyclic peptide complex as the structural reference (PDB ID: 7K2S); for SPSB2–iNOS, we used a cyclic peptide bound to SPSB2 as the reference (PDB ID: 5XN3). In both systems, the receptor design region was defined as residues whose Cβ atoms lie within 10 Å of the ligand in the reference complex. CycDiff-DPO then generated 1000 cyclic peptides matching the reference ligand length, followed by a single round of Rosetta FastRelax refinement. We discarded candidates with positive Rosetta interface ΔG and prioritized the remaining designs by interface ΔG together with the total number of intermolecular interactions identified by PLIP.Keap1–Nrf2
For Keap1–Nrf2, the top-ranked cyclic peptide is shown in Fig. 5a. The designed peptide overlays closely with the reference macrocycle and recapitulates hot-spot recognition within the Keap1 pocket. In the refined complex, the generated peptide forms hydrogen bonds with the hot-spot residues Arg415, Arg483, and Tyr525, whereas the reference peptide engages Arg380 and Arg483 (Fig. 5a). The refined complex achieves a Rosetta interface ΔG of −43.7 REU and a DMPNN-predicted PAMPA score of −5.93, compared to −33.7 REU and −11.00 for the reference macrocycle. Evaluation across five independent predictors and two assay modalities showed that four predictors consistently indicate improved permeability for the top design over the reference, though PharmPapp showed minimal differentiation (Table S5). To assess bound-state stability beyond static scoring, we carried out explicit-solvent MD simulations for both the designed complex and the reference (Fig. 5c). The designed peptide remained stably bound over the trajectory and exhibited a lower peptide heavy-atom RMSD than the reference (1.09 Å versus 1.17 Å).Across the design set, contact recovery analysis further showed frequent engagement of experimentally validated Keap1 hot-spot residues (Fig. 5e),48 with high recovery for Tyr572, Tyr334, Arg415, and Tyr525, and additional hot spots such as Arg380 and Arg483 also recovered across designs. Consistent with these recovery profiles, individual designs typically formed contacts with 3–5 hot-spot residues (Fig. S1a), indicating that the improved permeability predictions are achieved while retaining multi-residue hot-spot engagement. The binding–permeability landscape (Fig. 5f) provides a complementary view of the design set. Relative to the reference macrocycle (star), all generated samples achieve higher predicted permeability, and 18.8% of candidates improve both predicted permeability and interface ΔG.
SPSB2–iNOS
A similar pattern is observed for SPSB2–iNOS. The representative design shown in Fig. 5b closely overlaps with the reference binding geometry and remains well accommodated in the SPSB2 pocket. The refined design shows both a slightly more favorable interface ΔG (−31.23 vs. −30.34 REU) and improved predicted PAMPA (−6.82 vs. −8.10) relative to the reference. This improvement was consistent across all five predictors, despite PharmPapp again showing minimal differentiation (Table S5). In the refined complex, the generated peptide forms hydrogen bonds with the hot-spot residues Thr102 and Tyr120, consistent with the reference interaction pattern (Fig. 5b). MD simulations further support the stability of the bound pose (Fig. 5d). The designed peptide remained associated with SPSB2 and exhibited a slightly lower mean peptide heavy-atom RMSD than the reference (1.82 Å versus 1.91 Å). Hot-spot recovery analysis indicates frequent engagement of hot-spot residues on SPSB2,47 particularly Tyr120 and Trp207, with additional contributions from Thr102 and Val206 (Fig. 5g). In line with these recovery rates, individual designs typically contacted 2–3 hot-spot residues (Fig. S1b), consistent with a compact hot-spot-centered interaction footprint across the design set. In the binding–permeability landscape (Fig. 5h), 84.7% of generated samples achieve higher predicted permeability than the reference; among these, 78.7% also show more favorable interface ΔG than the reference.Both case study top designs contain charged residues, which may raise concerns about membrane permeability. However, ionizable side chains undergo substantial pKa shifts in low-dielectric membrane environments. The Keap1–Nrf2 design c[LLLVVVK] contains a single charged residue (Lys). In aqueous solution Lys is cationic (pKa ≈ 10.4), but in the low-dielectric membrane interior its pKa shifts downward by −4 to −5 units to approximately 5.6–6.5, as demonstrated by constant-pH MD34 and solid-state NMR in DOPC bilayers.35 At physiological pH, membrane-embedded Lys would be predominantly deprotonated and neutral. The remaining six residues are all strongly hydrophobic, yielding a highly lipophilic overall composition that favors membrane partitioning. The SPSB2–iNOS design c[KVLDIHLL] contains two charged residues: one Lys and one Asp. The Lys is expected to undergo the same downward pKa shift toward neutralization. The Asp (aqueous pKa ≈ 4.0) shifts upward by +2 to +5 units in membrane environments, reaching values of 5.8–9.4,34,49 at which it would be predominantly protonated and neutral at pH 7.4. Consequently, both designs are expected to carry substantially less charge during membrane crossing than their aqueous ionization states would suggest. To provide a property-level assessment independent of ML predictors, we computed the physicochemical profiles of the top designs and reference peptides (Table S6). Across three key permeability-related descriptors—HBD, TPSA, and log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P—the top designs show substantial improvements over the reference peptides, with particularly large gains in log
P—the top designs show substantial improvements over the reference peptides, with particularly large gains in log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P and TPSA. MW increased modestly due to the incorporation of additional hydrophobic residues, but remained within the range reported for orally bioavailable macrocycles.50
P and TPSA. MW increased modestly due to the incorporation of additional hydrophobic residues, but remained within the range reported for orally bioavailable macrocycles.50
Collectively, these two intracellular case studies indicate that CycDiff-DPO can generate cyclic peptide inhibitors that preserve target-like binding modes, stable bound-state behavior, and hot-spot engagement, while enriching candidates with improved predicted membrane permeability.
Ablation analyses
To quantify how preference optimization and the inverse-temperature parameter β modulate the permeability–affinity trade-off, we compared the base generator (no DPO), SFT, and DPO across a range of β values (Tables 2, S7, and S8). Relative to the base model, SFT provides only marginal improvements in permeability (Caco-2: −5.72 vs. −5.73; PAMPA: −8.17 vs. −8.31). In contrast, DPO improves performance in both assays, with the highest mean permeability observed at an intermediate β (Caco-2: −5.66; PAMPA: −7.89). Notably, the effect of β is non-monotonic: although β acts as an inverse temperature that rescales the preference signal, performance deteriorates at both low and high extremes (e.g., Caco-2: −5.69 and −5.71; PAMPA: −8.03 and −8.10). This pattern is consistent with prior observations that the optimal β depends on the signal-to-noise ratio and information content of the preference pairs. Importantly, Rosetta interface energies remain essentially unchanged across conditions (−36.13 to −37.78), indicating that the permeability gains are achieved without detectable loss of binding energetics.| DPO setting | Caco-2 permeability | PAMPA permeability | Rosetta ΔG |
|---|---|---|---|
| SFT | −5.72 | −8.17 | −36.90 |
| Without DPO | −5.73 | −8.31 | −36.13 |
| β = 0.1 | −5.69 | −8.03 | −37.01 |
| β = 0.5 | −5.66 | −7.89 | −37.78 |
| β = 5 | −5.69 | −7.97 | −37.75 |
| β = 50 | −5.71 | −8.10 | −36.60 |
Conclusions
CycDiff-DPO introduces a preference-aligned diffusion framework that shifts a pretrained generator toward cyclic peptides with improved membrane permeability while preserving target-binding competence. Existing generative models for cyclic peptides have focused primarily on binding affinity and structural validity, without incorporating permeability during design. Conversely, methods that optimize for permeability have not addressed target-specific binding. By adapting direct preference optimization to the continuous, E(3)-equivariant latent space of a geometric diffusion model through a denoising-error surrogate, CycDiff-DPO integrates both objectives directly into the generative process, enabling exploration of chemical space that is simultaneously binding-competent and membrane-permeable, which is a critical requirement for intracellular targets.Across 56 non-redundant test targets, CycDiff-DPO achieves a systematic distribution-level shift toward permeability-favorable physicochemical space, including elevated lipophilicity, reduced polar surface exposure, and improved predicted Caco-2 and PAMPA permeability that generalizes across independent external predictors. These gains are accompanied by the most favorable Rosetta interface energies among all methods tested across peptide-length categories and sampling depths, together with stereochemical quality and structural diversity comparable to baseline approaches. Case studies on two clinically relevant intracellular PPI targets, Keap1–Nrf2 and SPSB2–iNOS, further confirm that the designed peptides retain hot-spot engagement and exhibit stable bound poses in explicit-solvent MD simulations.
The intermolecular interaction profiles offer a physicochemically coherent account of how the model balances binding and permeability. CycDiff-DPO designs exhibit an increased number of hydrophobic contacts at the peptide–receptor interface, coupled with a modest reduction in hydrogen bonds relative to some baselines (Fig. 3d). This shift toward hydrophobically driven binding is consistent with reduced polarity and improved membrane partitioning. The enrichment of leucine and alanine in the generated sequences supports this global depolarization strategy (Fig. 2a), in line with experimental evidence that reducing overall polarity and polar surface area is critical for improving membrane permeability in macrocycles.23 IMHB analysis provides further evidence for this mechanism: generated peptides show no enrichment of IMHBs relative to baselines, and experimentally characterized compounds in CycPeptMPDB reveal a significantly negative association between IMHB count and permeability (Fig. S3 and S4). This conclusion is further supported by membrane-mimicking conformational ensemble analysis, which shows no positive association between IMHB counts and permeability even when conformational changes in the membrane environment were taken into account (Fig. S5 and Table S4). These observations suggest that IMHB shielding is not the dominant permeation mechanism in the chemical space explored here, and that the permeability gains of CycDiff-DPO are more consistent with a global depolarization strategy. Importantly, binding metrics remain stable despite the absence of binding-related supervision in the preference signal. This robustness likely arises from the SFT-style reconstruction regularizer and from the fact that hydrophobic residues enriched by permeability optimization also contribute to tighter nonpolar packing at the binding interface.
More broadly, CycDiff-DPO bridges target-specific binding design and membrane permeability optimization within a single generative framework, unifying two objectives that were previously pursued independently. This establishes distributional preference alignment as a practical paradigm for designing cell-permeable cyclic peptide binders against intracellular targets. Nonetheless, several limitations should be noted. First, the design space is currently restricted to the 20 canonical amino acids, whereas non-natural modifications such as N-methylation, D-amino acids, and backbone alterations are established tools for improving membrane permeability and metabolic stability. Incorporating such building blocks into the generative model is a key direction for future work. Second, permeability assessment in this work relies entirely on computational predictors. We have mitigated this limitation through multiple orthogonal lines of evidence—multi-predictor consensus across independent models and assay modalities, residue-specific pKa analysis under membrane-mimicking conditions, and physicochemical profiling against reported property spaces for orally bioavailable macrocycles—all of which consistently support the predicted permeability gains. Nevertheless, experimental validation of membrane permeability for top-ranked designs remains a necessary next step to confirm the computational predictions and to establish the translational relevance of the framework.
Methods
Dataset
The preference-alignment data for DPO were derived from PepBench, a curated benchmark of protein–peptide complexes introduced by Kong et al.31 The PepBench training split contains 4157 complexes, with receptor proteins longer than 30 residues and peptide ligands of 4–25 residues. From this split, we randomly selected 500 receptor targets for preference-pair generation. From this split, we randomly selected 500 receptor targets for preference-pair generation. After running CP-composer to generate initial cyclic peptide structures, 9 targets were excluded because generation failed, leaving 491 unique targets. We then partitioned these targets at the receptor level into training and validation sets in a 9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio (441 targets for training and 50 for validation).
1 ratio (441 targets for training and 50 for validation).
To evaluate generalization across unseen targets, we started from the large non-redundant (LNR) dataset curated in prior work51 and further selected a cyclic-peptide–compatible subset following the ligand-length and pocket-topology criteria in CPSea.28 Complexes were first filtered by ligand length, retaining only those with peptide ligands of 5–16 residues—a length regime that typically supports head-to-tail macrocyclization without excessive ring strain or major backbone rearrangements upon closure. Complexes were further excluded if introducing a cyclic topology would be expected to cause substantial steric clashes with the receptor or destabilize the bound conformation (Fig. S2). The remaining receptors were clustered using MMseqs2 to reduce redundancy.52 To prevent overlap with the cyclic-peptide fine-tuning data used for PepFlow and PepGLAD, we excluded any receptor showing >40% sequence identity to targets in the fine-tuning data.28 After these procedures, 56 non-redundant targets were retained as the final test set.
Base model architecture
We adopt CP-composer as our base model for target-conditioned cyclic peptide generation.17 CP-composer represents the binding site and peptide as a fully-connected geometric graph G = (V, E), where each node corresponds to a residue with features (hi, Xi), including the amino-acid type encoding hi and the coordinates Xi of associated atoms. The model is a latent geometric diffusion framework with a variational autoencoder that maps peptide graphs to residue-level latent variables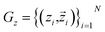 via an encoder Eϕ, and reconstructs sequence and structure via a decoder Dξ. Here zi is an E(3)-invariant scalar latent variable and
via an encoder Eϕ, and reconstructs sequence and structure via a decoder Dξ. Here zi is an E(3)-invariant scalar latent variable and 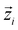 is an E(3)-invariant vector latent variable. A diffusion denoiser εθ(G(t)z, t) parameterized by an equivariant GNN is trained in this compact latent space. During sampling, latents are initialized from the prior and progressively denoised for T steps using a DDPM sampler, then decoded back to the data space using Dξ.
is an E(3)-invariant vector latent variable. A diffusion denoiser εθ(G(t)z, t) parameterized by an equivariant GNN is trained in this compact latent space. During sampling, latents are initialized from the prior and progressively denoised for T steps using a DDPM sampler, then decoded back to the data space using Dξ.
Permeability scoring model
We trained a machine-learning permeability predictor to assign each macrocyclic peptide candidate a scalar permeability score, sperm, which is subsequently used to construct preference pairs for preference optimization. We used Caco-2 measurements because they provide a higher-level proxy for cellular membrane permeation that reflects integrated transport phenomena (e.g., uptake and efflux) beyond purely passive diffusion, in contrast to artificial-membrane assays such as PAMPA.The predictor was trained on the Caco-2 permeability subset of the CycPeptMPDB database.21 After deduplication, the dataset comprises 1273 unique cyclic peptides with experimentally measured log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pexp values. Each peptide was represented by an extended feature vector that concatenates Morgan fingerprints (ECFP; radius = 6; 2048 bits) together with 35 two-dimensional physicochemical descriptors (Table S9). All features were standardized using a StandardScaler prior to model training. The predictor is an ensemble of 10 XGBoost regressors, each instantiated with a distinct hyperparameter configuration to promote diversity in model capacity and regularization strength. The 10 configurations span tree counts of 150–300, maximum depths of 5–9, learning rates of 0.05–0.10, row subsampling ratios of 0.72–0.78, and L1/L2 regularization weights of 0.01–0.20 and 1.0–2.0, respectively (Table S10). This deliberate variation in inductive biases—ranging from shallow, heavily regularized models to deeper, faster-learning ones—reduces ensemble variance and improves robustness when scoring generated peptides that may lie outside the training distribution. The final permeability score is obtained by averaging the predictions of the 10 base models. The predictor was evaluated using five-fold cross-validation. The final production model used for scoring during preference-pair construction was retrained on the full deduplicated dataset to maximize data utilization.
Pexp values. Each peptide was represented by an extended feature vector that concatenates Morgan fingerprints (ECFP; radius = 6; 2048 bits) together with 35 two-dimensional physicochemical descriptors (Table S9). All features were standardized using a StandardScaler prior to model training. The predictor is an ensemble of 10 XGBoost regressors, each instantiated with a distinct hyperparameter configuration to promote diversity in model capacity and regularization strength. The 10 configurations span tree counts of 150–300, maximum depths of 5–9, learning rates of 0.05–0.10, row subsampling ratios of 0.72–0.78, and L1/L2 regularization weights of 0.01–0.20 and 1.0–2.0, respectively (Table S10). This deliberate variation in inductive biases—ranging from shallow, heavily regularized models to deeper, faster-learning ones—reduces ensemble variance and improves robustness when scoring generated peptides that may lie outside the training distribution. The final permeability score is obtained by averaging the predictions of the 10 base models. The predictor was evaluated using five-fold cross-validation. The final production model used for scoring during preference-pair construction was retrained on the full deduplicated dataset to maximize data utilization.
Preference pair construction
To improve permeability without confounding target-dependent effects, we construct preference pairs independently within each receptor target. For each target, we generate K = 200 macrocyclic peptide candidates using the base model CP-composer, resulting in a target-conditioned candidate pool {mc,i}i=1K for each target c. Each valid cyclic peptide candidate is converted to SMILES representation and assigned a permeability score sperm(mc,i) using our permeability predictor.Preference pairs are constructed within each target to avoid confounding from cross-target differences. For a given target c, we sort candidates by predicted score in descending order and pair high-scoring samples with low-scoring samples using a symmetric rank-matching strategy: the i-th highest-scoring candidate is paired against the i-th lowest-scoring candidate. This yields up to [Nc/2] raw pairs per target, where Nc is the number of scored candidates. To ensure informative supervision, we retain a pair (m+, m−) only if the score margin satisfies
| sperm(m+) − sperm(m−) ≥ δ, |
| Dpref = {(c, m+, m−)}, |
Preference alignment with DPO
Given target-conditioned preference tuples (ci, m+i, m−i) constructed above, where m = (H, X) denotes a cyclic peptide design with sequence H and structure X under target context c, we align the pretrained generator toward higher permeability using DPO strategy. Let pθ be the current diffusion model and pref be a frozen reference initialized from the same pretrained checkpoint. DPO increases the relative likelihood of the preferred design over the dispreferred one while regularizing against the reference model. Formally, we optimizewhere σ(·) is the sigmoid function, β is an inverse temperature parameter that controls alignment strength.
Direct evaluation of log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) pθ(m|c) is intractable for diffusion generators because it requires marginalizing over the full denoising trajectory. We therefore replace the log-likelihood terms in Δθ with a tractable proxy derived from the diffusion score-matching objective, so that preference comparisons can be computed using denoising errors (MSE) rather than exact log
pθ(m|c) is intractable for diffusion generators because it requires marginalizing over the full denoising trajectory. We therefore replace the log-likelihood terms in Δθ with a tractable proxy derived from the diffusion score-matching objective, so that preference comparisons can be computed using denoising errors (MSE) rather than exact log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) p values.53 Concretely, in the latent diffusion space z produced by the autoencoder, we sample a diffusion timestep t and noise ε, form the noised latent zt via the forward process, and compute a model-vs-reference denoising-error gap for a design m:
p values.53 Concretely, in the latent diffusion space z produced by the autoencoder, we sample a diffusion timestep t and noise ε, form the noised latent zt via the forward process, and compute a model-vs-reference denoising-error gap for a design m:
Intuitively, if the current model explains m better than the reference, its denoising error is smaller and gθ(m) decreases, corresponding to a higher proxy likelihood under pθ relative to pref. We instantiate the preference logit using denoising-gap contrasts:
| Δ(c, m+, m−) ≈ −gθ(m+) + gθ(m−), |
| L = λ1LDPO + LSFT(m+), |
| LSFT(m+) = λ2LH,reconstruct(H+) + λ3LX,reconstruct(X+). |
Training and implementation details
CycDiff-DPO is trained on a single RTX 3090 GPU (24 GB) using AdamW. The model is initialized from the pretrained CP-composer checkpoint,17 and the autoencoder parameters are kept frozen throughout training. Preference alignment is performed using the DPO objective described above, with β = 0.5 and the loss weights set to λ1 = 1, λ2 = 1, and λ3 = 1. We train for up to 50 epochs and select checkpoints based on validation performance. The initial learning rate is 1 × 10−5 with weight decay 0.05, and is reduced by a factor of 0.6 if the validation loss does not improve for three consecutive epochs. Early stopping is applied with a patience of 10 epochs. We retain the top 10 checkpoints according to validation performance for downstream analysis, and report results using the best-performing checkpoint.Evaluation metrics
We evaluated the backbone stereochemical quality using Ramachandran statistics of ϕ/ψ dihedrals. reporting the mean proportion of residues in accepted and favored regions across all generated samples. To characterize surface polarity, we computed the polar solvent-accessible surface area (SASA) ratio,where SASApolar denotes the SASA contributed by polar atoms and SASAtotal is the total SASA. Lipophilicity was summarized by log
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (octanol–water partition coefficient), which serves as a compact descriptor relevant to permeability-related trade-offs. Structural diversity within a generated set was quantified as the mean pairwise dissimilarity 1 − TM, where TM-score is computed after backbone structural alignment; for N peptides, we report
P (octanol–water partition coefficient), which serves as a compact descriptor relevant to permeability-related trade-offs. Structural diversity within a generated set was quantified as the mean pairwise dissimilarity 1 − TM, where TM-score is computed after backbone structural alignment; for N peptides, we reportso that larger D indicates greater diversity. For complex-level evaluation, peptide–receptor structures were refined with one cycle of Rosetta FastRelax prior to scoring, after which we computed the Rosetta interface energy (ΔG) using the REF2015 score function and additionally reported the Rosetta total energy of the relaxed complex as a proxy for overall bound-state stability. To provide interpretable interaction readouts beyond energies, we analyzed interfaces using PLIP38 and reported the numbers of hydrophobic contacts, hydrogen bonds, and salt bridges detected for each complex.
Molecular dynamics simulations
Protein–peptide complexes were protonated at pH 7 using PROPKA in PDB2PQR.54 Systems were solvated in a 10 Å truncated octahedral TIP3P water box55 and neutralized with Na+/Cl− to 150 mM. Proteins and peptides were modeled with ff14SB.56 Simulations were performed in Amber22 using GPU-accelerated PME on RTX 3090 hardware.57 Each system was minimized for 2500 steepest-descent and 2500 conjugate-gradient steps with restraints on solvent/ions, followed by an unrestrained minimization of identical length. Velocities were assigned from a Boltzmann distribution, then heated from 0 to 310 K over 500 ps in the NVT ensemble with a Langevin thermostat and 10.0 kcal mol−1·Å−2 solute restraints. NPT equilibration (310 K, 1 bar) was run for 2.5 ns while reducing solute restraints from 5.0 to 0.1 kcal mol−1·Å−2 in four 0.5 ns stages. Production runs were conducted at 310 K and 1 bar without restraints using a Langevin thermostat and Berendsen barostat. Each production simulation was run for 100 ns, and three independent replicates with different random seeds were performed per system. A 4.0 fs timestep was enabled by hydrogen mass repartitioning,58 bonds to hydrogens were constrained with SHAKE,59 and nonbonded interactions used a 10 Å cutoff.Author contributions
Conceptualization: H. S., H. T., J. L; methodology: H. S., Y. C.; formal analysis: H. S.; writing – original draft: H. S.; writing – review & editing: H. S., H. T., R. W., D. W.; supervision: D. W.; funding acquisition: D. W.Conflicts of interest
The authors declare no conflicts of interest.Data availability
The pretrained CycDiff-DPO model weights, the preprocessed input data, and the code for running the CycDiff-DPO algorithm are available on GitHub at https://github.com/sun-heqi/CycDiff-DPO, and on Zenodo at http://zenodo.org/records/19429073.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d6sc01722c.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (32570803, 32030063) and the National Key R&D Program of China (2024YFA1306400, 2023YFE0199200).References
- L. K. Buckton, M. N. Rahimi and S. R. McAlpine, Cyclic peptides as drugs for intracellular targets: the next frontier in peptide therapeutic development, Chem, Eur. J., 2021, 27, 1487–1513 CrossRef CAS PubMed.
- A. A. Vinogradov, Y. Yin and H. Suga, Macrocyclic peptides as drug candidates: recent progress and remaining challenges, J. Am. Chem. Soc., 2019, 141, 4167–4181 CrossRef CAS PubMed.
- M. Muttenthaler, G. F. King, D. J. Adams and P. F. Alewood, Trends in peptide drug discovery, Nat. Rev. Drug Discovery, 2021, 20, 309–325 CrossRef CAS PubMed.
- N. Tsomaia, Peptide therapeutics: targeting the undruggable space, Eur. J. Med. Chem., 2015, 94, 459–470 CrossRef CAS PubMed.
- A. I. Casas, A. A. Hassan, S. J. Larsen, V. Gomez-Rangel, M. Elbatreek, P. W. M. Kleikers, E. Guney, J. Egea, M. G. López, J. Baumbach and H. H. H. W. Schmidt, From single drug targets to synergistic network pharmacology in ischemic stroke, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 7129–7136 CrossRef CAS.
- A. L. Hopkins, Network pharmacology: the next paradigm in drug discovery, Nat. Chem. Biol., 2008, 4, 682–690 CrossRef CAS PubMed.
- R. González-Muñiz, M. Á. Bonache and M. J. P. de Vega, Modulating protein–protein interactions by cyclic and macrocyclic peptides. Prominent strategies and examples, Molecules, 2021, 26, 445 CrossRef.
- J. Gavenonis, B. A. Sheneman, T. R. Siegert, M. R. Eshelman and J. A. Kritzer, Comprehensive analysis of loops at protein-protein interfaces for macrocycle design, Nat. Chem. Biol., 2014, 10, 716–722 CrossRef CAS PubMed.
- J. G. Beck, J. Chatterjee, B. Laufer, M. U. Kiran, A. O. Frank, S. Neubauer, O. Ovadia, S. Greenberg, C. Gilon, A. Hoffman and H. Kessler, Intestinal permeability of cyclic peptides: common key backbone motifs identified, J. Am. Chem. Soc., 2012, 134, 12125–12133 CrossRef CAS PubMed.
- C. Grant, F. Rahman, R. Piekarz, C. Peer, R. Frye, R. W. Robey, E. R. Gardner, W. D. Figg and S. E. Bates, Romidepsin: a new therapy for cutaneous T-cell lymphoma and a potential therapy for solid tumors, Expert Rev. Anticancer Ther., 2010, 10, 997–1008 CrossRef CAS PubMed.
- T. van Gelder, E. Lerma, K. Engelke and R. B. Huizinga, Voclosporin: a novel calcineurin inhibitor for the treatment of lupus nephritis, Expet Rev. Clin. Pharmacol., 2022, 15, 515–529 CrossRef CAS.
- J. A. Russell, K. R. Walley, J. Singer, A. C. Gordon, P. C. Hébert, D. J. Cooper, C. L. Holmes, S. Mehta, J. T. Granton, M. M. Storms, D. J. Cook, J. J. Presneill and D. Ayers, Vasopressin versus norepinephrine infusion in patients with septic shock, N. Engl. J. Med., 2008, 358, 877–887 CrossRef CAS PubMed.
- A. G. Atanasov, S. B. Zotchev, V. M. Dirsch, I. E. Orhan, M. Banach, J. M. Rollinger, D. Barreca, W. Weckwerth, R. Bauer, E. A. Bayer, M. Majeed, A. Bishayee, V. Bochkov, G. K. Bonn, N. Braidy, F. Bucar, A. Cifuentes, G. D'Onofrio, M. Bodkin, M. Diederich, A. T. Dinkova-Kostova, T. Efferth, K. El Bairi, N. Arkells, T. P. Fan, B. L. Fiebich, M. Freissmuth, M. I. Georgiev, S. Gibbons, K. M. Godfrey, C. W. Gruber, J. Heer, L. A. Huber, E. Ibanez, A. Kijjoa, A. K. Kiss, A. Lu, F. A. Macias, M. J. S. Miller, A. Mocan, R. Müller, F. Nicoletti, G. Perry, V. Pittalà, L. Rastrelli, M. Ristow, G. L. Russo, A. S. Silva, D. Schuster, H. Sheridan, K. Skalicka-Woźniak, L. Skaltsounis, E. Sobarzo-Sánchez, D. S. Bredt, H. Stuppner, A. Sureda, N. T. Tzvetkov, R. A. Vacca, B. B. Aggarwal, M. Battino, F. Giampieri, M. Wink, J. L. Wolfender, J. Xiao, A. W. K. Yeung, G. Lizard, M. A. Popp, M. Heinrich, I. Berindan-Neagoe, M. Stadler, M. Daglia, R. Verpoorte and C. T. Supuran, Natural products in drug discovery: advances and opportunities, Nat. Rev. Drug Discovery, 2021, 20, 200–216 CrossRef CAS.
- G. Bhardwaj, J. O'Connor, S. Rettie, Y. H. Huang, T. A. Ramelot, V. K. Mulligan, G. G. Alpkilic, J. Palmer, A. K. Bera, M. J. Bick, M. Di Piazza, X. Li, P. Hosseinzadeh, T. W. Craven, R. Tejero, A. Lauko, R. Choi, C. Glynn, L. Dong, R. Griffin, W. C. van Voorhis, J. Rodriguez, L. Stewart, G. T. Montelione, D. Craik and D. Baker, Accurate de novo design of membrane-traversing macrocycles, Cell, 2022, 185, 3520–3532 CrossRef CAS PubMed.
- M. Wang, Z. Cang and G.-W. Wei, A topology-based network tree for the prediction of protein–protein binding affinity changes following mutation, Nat. Mach. Intell., 2020, 2, 116–123 CrossRef.
- S. A. Rettie, D. Juergens, V. Adebomi, Y. F. Bueso, Q. Zhao, A. N. Leveille, A. Liu, A. K. Bera, J. A. Wilms, A. Üffing, A. Kang, E. Brackenbrough, M. Lamb, S. R. Gerben, A. Murray, P. M. Levine, M. Schneider, V. Vasireddy, S. Ovchinnikov, O. H. Weiergräber, D. Willbold, J. A. Kritzer, J. D. Mougous, D. Baker, F. DiMaio and G. Bhardwaj, Accurate de novo design of high-affinity protein-binding macrocycles using deep learning, Nat. Chem. Biol., 2025, 21, 1948–1956 CrossRef CAS PubMed.
- D. Jiang, X. Kong, J. Han, M. Li, R. Jiao, W. Huang, S. Ermon, J. Ma and Y. Liu, Zero-shot cyclic peptide design via composable geometric constraints, in Proc. 42nd Int. Conf. Mach. Learn., PMLR, 2025, vol. 267, pp. 27553–27568 Search PubMed.
- Q. Li, E. N. Vlachos and P. Bryant, Design of linear and cyclic peptide binders from protein sequence information, Commun. Chem., 2025, 8, 211 CrossRef PubMed.
- F. Wang, T. Zhang, J. Zhu, X. Zhang, C. Zhang and L. Lai, Reinforcement learning-based target-specific de novo design of cyclic peptide binders, J. Med. Chem., 2025, 68, 17287–17302 CrossRef CAS PubMed.
- H. Lin, C. Zhu, T. Shang, N. Zhu, K. Lin, C. Zhang, X. Shao, X. Wang and H. Duan, HighPlay: cyclic peptide sequence design based on reinforcement learning and protein structure prediction, J. Med. Chem., 2025, 68, 12047–12057 CrossRef CAS PubMed.
- J. Li, K. Yanagisawa, M. Sugita, T. Fujie, M. Ohue and Y. Akiyama, CycPeptMPDB: a comprehensive database of membrane permeability of cyclic peptides, J. Chem. Inf. Model., 2023, 63, 2240–2250 CrossRef CAS PubMed.
- C. Zhang, Z. Xu, K. Lin, N. Zhu, C. Zhang, W. Xu, J. Guo, A. Su, C. Li and H. Duan, CycleDesigner: leveraging CycRFdiffusion and HighFold to design cyclic peptide binders for specific targets, J. Chem. Inf. Model., 2025, 65, 6155–6165 CrossRef CAS PubMed.
- M. L. Merz, S. Habeshian, B. Li, J. A. G. L. David, A. L. Nielsen, X. Ji, K. I. Khwildy, M. M. Duany Benitez, P. Phothirath and C. Heinis, De novo development of small cyclic peptides that are orally bioavailable, Nat. Chem. Biol., 2024, 20, 624–633 CrossRef CAS PubMed.
- R. Rafailov, A. Sharma, E. Mitchell, S. Ermon, C. D. Manning and C. Finn, Direct preference optimization: your language model is secretly a reward model, in Adv. Neural Inf. Process. Syst., 2023, p. 36 Search PubMed.
- X. Zhou, D. Xue, R. Chen, Z. Zheng, L. Wang and Q. Gu, Antigen-specific antibody design via direct energy-based preference optimization, in Adv. Neural Inf. Process. Syst., 2024, p. 37 Search PubMed.
- S. Gu, M. Xu, A. Powers, W. Nie, T. Geffner, K. Kreis, J. Leskovec, A. Vahdat and S. Ermon, Aligning target-aware molecule diffusion models with exact energy optimization, in Adv. Neural Inf. Process. Syst., 2024, p. 37 Search PubMed.
- X. Cheng, X. Zhou, Y. Bao, Y. Yang and Q. Gu, Decomposed direct preference optimization for structure-based drug design, arXiv, 2024, preprint, arXiv:2407.13981, DOI:10.48550/arXiv.2407.13981.
- Z. Yang, H. Xie, Y. Jia, X. Kong, J. Zheng, Z. Zhang, Y. Liu, L. Liu and Y. Lan, CPSea: large-scale cyclic peptide-protein complex dataset for machine learning in cyclic peptide design, OpenReview, 2025 Search PubMed.
- J. Dauparas, I. Anishchenko, N. Bennett, H. Bai, R. J. Ragotte, L. F. Milles, B. I. M. Wicky, A. Courbet, R. J. de Haas, N. Bethel, P. J. Y. Leung, T. F. Huddy, S. Pellock, D. Tischer, F. Chan, B. Koepnick, H. Nguyen, A. Kang, B. Sankaran, A. K. Bera, N. P. King and D. Baker, Robust deep learning-based protein sequence design using ProteinMPNN, Science, 2022, 378, 49–56 CrossRef CAS PubMed.
- J. Li, C. Cheng, Z. Wu, R. Guo, S. Luo, Z. Ren, J. Peng and J. Ma, Full-atom peptide design based on multi-modal flow matching, in Proc. 41st Int. Conf. Mach. Learn., PMLR, 2024, vol. 235, pp. 27615–27640 Search PubMed.
- X. Kong, Y. Jia, W. Huang and Y. Liu, Full-atom peptide design with geometric latent diffusion, in Adv. Neural Inf. Process. Syst., 2024, vol. 37, pp. 74808–74839 Search PubMed.
- X. Liu, B. Testa and A. Fahr, Lipophilicity and its relationship with passive drug permeation, Pharm. Res., 2011, 28, 962–977 CrossRef CAS PubMed.
- M. Rossi Sebastiano, B. C. Doak, M. Backlund, V. Poongavanam, B. Over, G. Ermondi, G. Caron, P. Matsson and J. Kihlberg, Impact of dynamically exposed polarity on permeability and solubility of chameleonic drugs beyond the rule of 5, J. Med. Chem., 2018, 61, 4189–4202 CrossRef CAS PubMed.
- A. Panahi and C. L. Brooks, Membrane environment modulates the pKa values of transmembrane helices, J. Phys. Chem. B, 2015, 119, 4601–4607 CrossRef CAS.
- N. J. Gleason, V. V. Vostrikov, D. V. Greathouse and R. E. Koeppe, Buried lysine, but not arginine, titrates and alters transmembrane helix tilt, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 1692–1695 CrossRef CAS PubMed.
- J. L. MacCallum, W. F. Drew Bennett and D. P. Tieleman, Distribution of amino acids in a lipid bilayer from computer simulations, Biophys. J., 2008, 94, 3393–3404 CrossRef CAS.
- R. F. Alford, A. Leaver-Fay, J. R. Jeliazkov, M. J. O'Meara, F. P. DiMaio, H. Park, M. V. Shapovalov, P. D. Renfrew, V. K. Mulligan, K. Kappel, J. W. Labonte, M. S. Pacella, R. Bonneau, P. Bradley, R. L. Dunbrack, R. Das, D. Baker, B. Kuhlman, T. Kortemme and J. J. Gray, The Rosetta All-Atom Energy Function for Macromolecular Modeling and Design, J. Chem. Theory Comput., 2017, 13, 3031–3048 CrossRef CAS PubMed.
- M. F. Adasme, K. L. Linnemann, S. N. Bolz, F. Kaiser, S. Salentin, V. J. Haupt and M. Schroeder, PLIP 2021: expanding the scope of the protein–ligand interaction profiler to DNA and RNA, Nucleic Acids Res., 2021, 49, W530–W534 CrossRef CAS PubMed.
- A. Avdeef, S. Bendels, L. Di, B. Faller, M. Kansy, K. Sugano and Y. Yamauchi, PAMPA—critical factors for better predictions of absorption, J. Pharm. Sci., 2007, 96, 2893–2909 CrossRef CAS PubMed.
- P. Artursson, K. Palm and K. Luthman, Caco-2 monolayers in experimental and theoretical predictions of drug transport, Adv. Drug Deliv. Rev., 2001, 46, 27–43 CrossRef CAS PubMed.
- D. Jiang, Z. Chen and H. Du, Cyclic peptide membrane permeability prediction using deep learning model based on molecular attention transformer, Front. Bioinform., 2025, 5, 1566174 CrossRef PubMed.
- X. Tan, Q. Liu, Y. Fang, Y. Zhu, F. Chen, W. Zeng, D. Ouyang and J. Dong, Predicting peptide permeability across diverse barriers: a systematic investigation, Mol. Pharm., 2024, 21, 4116–4127 CrossRef CAS.
- W. Liu, J. Li, C. S. Verma and H. K. Lee, Systematic benchmarking of 13 AI methods for predicting cyclic peptide membrane permeability, J. Cheminf., 2025, 17, 129 Search PubMed.
- T. Rezai, B. Yu, G. L. Millhauser, M. P. Jacobson and R. S. Lokey, Testing the conformational hypothesis of passive membrane permeability using synthetic cyclic peptide diastereomers, J. Am. Chem. Soc., 2006, 128, 2510–2511 CrossRef CAS PubMed.
- C. A. Grambow, H. Weir, C. N. Cunningham, T. Biancalani and K. V. Chuang, CREMP: conformer-rotamer ensembles of macrocyclic peptides for machine learning, Sci. Data, 2024, 11, 859 CrossRef CAS.
- A. Cuadrado, A. I. Rojo, G. Wells, J. D. Hayes, S. P. Cousin, W. L. Rumsey, O. C. Attucks, S. Franklin, A. L. Levonen, T. W. Kensler and A. T. Dinkova-Kostova, Therapeutic targeting of the NRF2 and KEAP1 partnership in chronic diseases, Nat. Rev. Drug Discovery, 2019, 18, 295–317 CrossRef CAS PubMed.
- Z. Kuang, R. S. Lewis, J. M. Curtis, Y. Zhan, B. M. Saunders, J. J. Babon, T. B. Kolesnik, A. Low, S. L. Masters, T. A. Willson, L. Kedzierski, S. Yao, E. Handman, R. S. Norton and S. E. Nicholson, The SPRY domain-containing SOCS box protein SPSB2 targets iNOS for proteasomal degradation, J. Cell Biol., 2010, 190, 129–141 CrossRef CAS PubMed.
- S. C. Lo, X. Li, M. T. Henzl, L. J. Beamer and M. Hannink, Structure of the Keap1:Nrf2 interface provides mechanistic insight into Nrf2 signaling, EMBO J., 2006, 25, 3605–3617 CrossRef CAS PubMed.
- S. Z. Hanz, N. S. Shu, J. Qian, N. Christman, P. Kranz, M. An, C. Grewer and W. Qiang, Protonation-driven membrane insertion of a pH-low insertion peptide, Angew. Chem., Int. Ed., 2016, 55, 12376–12381 CrossRef CAS PubMed.
- E. A. Villar, D. Beglov, S. Chennamadhavuni, J. A. Porco, D. Kozakov, S. Vajda and A. Whitty, How proteins bind macrocycles, Nat. Chem. Biol., 2014, 10, 723–731 CrossRef CAS PubMed.
- T. Tsaban, J. K. Varga, O. Avraham, Z. Ben-Aharon, A. Khramushin and O. Schueler-Furman, Harnessing protein folding neural networks for peptide–protein docking, Nat. Commun., 2022, 13, 176 CrossRef CAS PubMed.
- M. Mirdita, M. Steinegger and J. Söding, MMseqs2 desktop and local web server app for fast, interactive sequence searches, Bioinformatics, 2019, 35, 2856–2858 CrossRef CAS PubMed.
- B. Wallace, M. Dang, R. Rafailov, L. Zhou, A. Lou, S. Purushwalkam, S. Ermon, C. Xiong, S. Joty and N. Naik, Diffusion model alignment using direct preference optimization, in Proc. IEEE/CVF Conf. Comput. Vis. Pattern Recognit., 2024, pp. 8228–8238 Search PubMed.
- T. J. Dolinsky, P. Czodrowski, H. Li, J. E. Nielsen, J. H. Jensen, G. Klebe and N. A. Baker, PDB2PQR: expanding and upgrading automated preparation of biomolecular structures for molecular simulations, Nucleic Acids Res., 2007, 35, W522–W525 CrossRef PubMed.
- W. L. Jorgensen, J. Chandrasekhar, J. D. Madura, R. W. Impey and M. L. Klein, Comparison of simple potential functions for simulating liquid water, J. Chem. Phys., 1983, 79, 926–935 CrossRef CAS.
- J. A. Maier, C. Martinez, K. Kasavajhala, L. Wickstrom, K. E. Hauser and C. Simmerling, ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB, J. Chem. Theory Comput., 2015, 11, 3696–3713 CrossRef CAS PubMed.
- R. Salomon-Ferrer, A. W. Götz, D. Poole, S. Le Grand and R. C. Walker, Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. Explicit solvent particle mesh Ewald, J. Chem. Theory Comput., 2013, 9, 3878–3888 CrossRef CAS PubMed.
- C. W. Hopkins, S. Le Grand, R. C. Walker and A. E. Roitberg, Long-time-step molecular dynamics through hydrogen mass repartitioning, J. Chem. Theory Comput., 2015, 11, 1864–1874 CrossRef CAS PubMed.
- J.-P. Ryckaert, G. Ciccotti and H. J. C. Berendsen, Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes, J. Comput. Phys., 1977, 23, 327–341 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |