Open Access Article
Open Access ArticleA chemo-selective deprotection–cyclization strategy enables fluorescence imaging of hydroxylamine and reveals its pathological role in Parkinson's disease
Shunping Zang†
a,
Jia Ke†b,
Hanbing Zhenga,
Qing Liua,
Liu Yanga,
Chaoyi Yao*a,
Benhua Wang *a,
Minhuan Lan
*a,
Minhuan Lan a and
Xiangzhi Song
a and
Xiangzhi Song *a
*a
aCollege of Chemistry and Chemical Engineering, Central South University, Changsha, Hunan 410083, P.R. China. E-mail: chaoyiyao@csu.edu.cn; benhuawang@csu.edu.cn; xzsong@csu.edu.cn
bLiangzhu Laboratory, Zhejiang University Medical Center, Hangzhou, Zhejiang 311121, P.R. China
First published on 28th April 2026
Abstract
Hydroxylamine (HA), a reactive nitrogen species generated by neuronal nitric oxide synthase (nNOS), has been largely overlooked in neurodegenerative disorders. Herein, we report the first identification of HA overexpression in Parkinson's disease (PD) and elucidate its pathological role using chemo-selective fluorescence imaging combined with proteomic analysis. A general HA-responsive probe platform was developed by introducing a 1-(4,4-dimethyl-2,6-dioxacyclohexylidene)ethyl (Dde) unit as a highly specific HA-recognition motif. Among the resulting probes, DCI-HA-2 exhibited remarkable sensitivity, a fast response, and excellent selectivity toward HA. Using DCI-HA-2, elevated HA levels were visualized in PD cells, PD model mice, and brain tissues for the first time. Mechanistically, nNOS-derived HA suppresses cystathionine β-synthase (CBS) expression, leading to impaired hydrogen sulfide (H2S) biosynthesis, and simultaneously induces adenosine-5′-triphosphate (ATP) depletion, thereby disrupting adenosine 5′-monophosphate-activated protein kinase (AMPK) signaling in PD pathology. This work identifies HA as a previously unrecognized pathological regulator in PD, and establishes a powerful chemical strategy for probing HA-associated pathological processes in complex biological systems.
Introduction
Accumulating evidence suggests that reactive nitrogen species (RNS) derived from neuronal nitric oxide synthase (nNOS) have been closely implicated in Parkinson's disease (PD) pathogenesis, a progressive neurodegenerative disorder without effective therapeutic intervention.1–6 While canonical RNS such as nitric oxide (NO), nitroxyl (HNO), and peroxynitrite (ONOO−) have been extensively investigated in PD, comparatively little attention has been devoted to hydroxylamine (HA), an important downstream metabolite intimately linked to nNOS-associated nitrogen redox pathways.7–9 Emerging studies reveal that elevated levels of HA can inhibit cystathionine β-synthase (CBS) and succinate oxidase activity.10–12 HA has also been reported to possess a neurological relaxation effect.13 Despite these observations, the exact level of HA in PD and how it may contribute to PD progression remain unknown. The limited exploration of HA in PD is largely due to the lack of reliable methods for selective HA detection under complex pathological conditions.Fluorescence imaging has emerged as a powerful tool for biomarker detection and therapeutic evaluation in neurological diseases owing to its high sensitivity, selectivity, noninvasiveness, and in situ imaging capability.14–19 However, selective HA recognition remains a challenge, as HA is a highly reactive nitrogen-containing nucleophile that coexists with numerous endogenous interferents. To date, only a few fluorescent probes for HA detection have been reported (Table S1), and most rely on the nucleophilic addition or condensation with electrophilic motifs.20–25 Although such designs are synthetically accessible, they often suffer from cross-reactivity with endogenous nucleophiles, including hydrogen sulfide (H2S), thiols, sulfur dioxide derivatives, and ONOO−.26–29 For example, probe 1 responds to both HA and H2S, while the HA probe RhChr displays cross-reactivity toward ONOO− (Table S1).30,31 Therefore, the development of a fundamentally distinct recognition mechanism that enables highly selective detection of HA for elucidating its fluctuations and mechanistic role in PD is highly desirable.
Inspired by classic deprotection chemistry, we turn our attention to the 1-(4,4-dimethyl-2,6-dioxacyclohexylidene)ethyl (Dde) group, an amine-protecting moiety that undergoes selectively cleavage by HA under mild conditions.32,33 Based on this unique reactivity, we envisioned that the Dde motif could be repurposed as a HA-specific chemical trigger in living systems. Such a strategy exploits the intrinsic and preferential reactivity of HA toward Dde deprotection, thereby averting interference from biological nucleophilic species.34 Additionally, when coupled to fluorophores bearing masked electron-donating groups, HA-mediated Dde cleavage can initiate an intramolecular cyclization cascade, resulting in fluorophore release and signal amplification. This deprotection–cyclization mechanism offers a conceptually generalizable platform for HA-selective probe development.
Herein, we report a chemo-selective “deprotection–cyclization” strategy for HA recognition based on Dde chemistry and establish a general fluorescent probe platform applicable to multiple fluorophore scaffolds (Scheme 1). The optimized probe, DCI-HA-2, enabled rapid, sensitive, and exceptionally selective detection of HA in cells and in vivo. Using this probe, we achieved, for the first time, the real-time imaging of elevated HA levels in PD focal cells and PD model mice. Importantly, by integrating fluorescence imaging with proteomic analysis, we uncovered a previously unrecognized pathogenic pathway in which nNOS-derived HA suppresses the expression of CBS, leading to reduced cellular H2S production, and concurrently induces ATP depletion with subsequent downregulation of the AMPK signaling pathway during PD progression. Overall, this study not only provides a chemo-selective strategy for HA detection, but also unveils HA-associated pathogenic mechanisms in PD, thereby opening new avenues for improved pathological research and targeted therapeutic intervention.
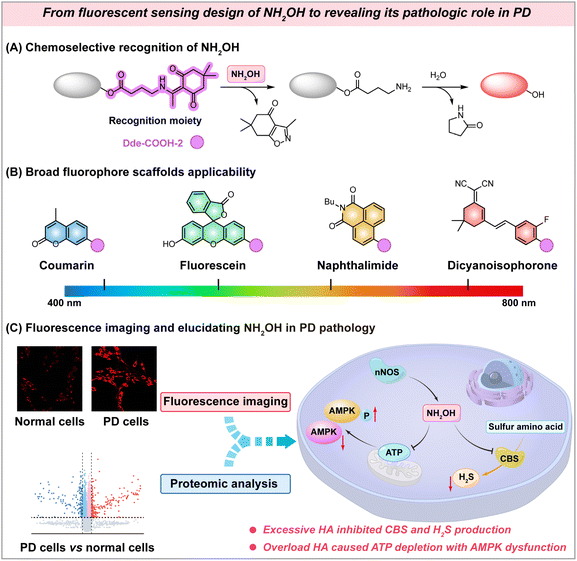 | ||
| Scheme 1 Design strategy for HA-activated fluorescent probes and schematic illustration of elucidating its pathological role in PD. | ||
Results and discussion
Molecular design and synthesis of probes
To enable deeper brain imaging in PD models, the dicyanoisophorone (DCI) fluorophore was first selected as the fluorophore due to its favorable blood–brain barrier (BBB) permeability and red-emission optical properties.35,36 To further optimize the probe's kinetics and sensitivity, a series of Dde-protected probes (DCI-HA-n, n = 1–4) with different linker lengths between the free amine and the carbonyl group were synthesized (Schemes S1 and S2) for investigation. Upon exposure to HA, the Dde-protected amine is first deprotected to generate a free primary amine (Step 1), which subsequently initiates intramolecular nucleophilic cyclization and releases the active fluorophore (Step 2), thereby restoring fluorescence (Scheme 2). All compounds were fully characterized by HRMS, 1H NMR and 13C NMR spectroscopy (Fig. S29–S68).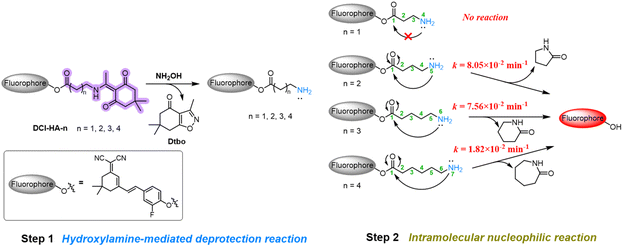 | ||
| Scheme 2 Investigation and the proposed process of HA recognition in probes with various length cleavable linkers. | ||
Photophysical properties and the response mechanism of probes towards HA
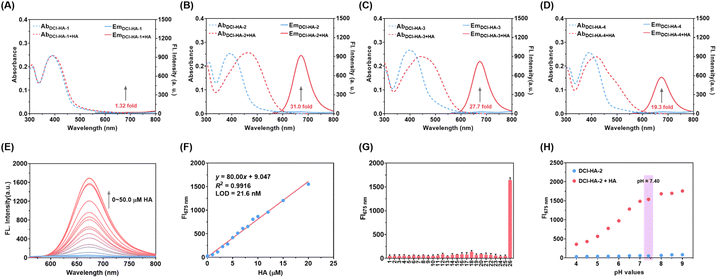
The photophysical properties and response kinetics of probes DCI-HA-n towards HA were first investigated in PBS buffer (10.0 mM, pH = 7.40, containing 1% DMSO). As seen in Fig. 1A–D, in the absence of HA, all probes displayed a characteristic absorption peak around 395 nm. Upon addition of HA (10.0 µM), DCI-HA-2, DCI-HA-3, and DCI-HA-4 displayed a pronounced red shift to 470 nm, indicative of Dde deprotection and subsequent structural transformation, whereas DCI-HA-1 exhibited almost no spectral change, hinting ineffective cyclization due to its short linker. Correspondingly, DCI-HA-2, DCI-HA-3 and DCI-HA-4 showed obvious fluorescence emission at 675 nm, with 31.0-fold, 27.7-fold and 19.3-fold fluorescence enhancement, respectively. In addition, the response rate of these probes was strongly dependent on the length of the cleavable linker, following the order: DCI-HA-2 (30.0 min) > DCI-HA-3 (45.0 min) > DCI-HA-4 (120 min) (Fig. S1).To verify the proposed sensing mechanism, HRMS analysis was performed. As illustrated in Fig. S2–S5, the mass peaks at [M + H]+ = 180.1025 and [M + Na]+ = 202.0844 correspond to the expected deprotected product, Dtbo (Scheme 2, M = 179.0946), confirming HA-mediated cleavage of the Dde group. For the mixture of DCI-HA-1 with HA, a peak at m/z = 380.1768 was observed, corresponding to the expected intermediate DCI-HA-1-NH2, consistent with its inability to undergo intramolecular cyclization (Fig. S2). In contrast, cyclic intermediates and the released fluorophore DCI-OH were observed in the mixtures of DCI-HA-2, DCI-HA-3, and DCI-HA-4 with HA, respectively, validating the proposed cyclization–release mechanism (Fig. S3–S5). The variation in response times among DCI-HA-2, DCI-HA-3, and DCI-HA-4 is attributed to linker-length-dependent intramolecular reaction rates.37–39 Among these probes, DCI-HA-2 exhibited the most favorable response characteristics and was selected for further studies.
Subsequently, we systematically investigated the spectral response of DCI-HA-2 towards HA. As shown in Fig. 1E, the fluorescence intensity at 675 nm gradually increased after adding HA (0–50.0 µM). A good linear relationship between fluorescence intensity and HA concentration was observed in the range of 0–20.0 µM, and the limit of detection (LOD) was calculated to be 21.6 nM (Fig. 1F). To validate the specificity, the fluorescence responses of DCI-HA-2 toward various biologically relevant species were examined. As shown in Fig. 1G, common metal ions (Fe3+, Fe2+, Zn2+, Mg2+, Cu2+ and Ca2+), reactive sulfur species (HSO3−, SO32−, S2O32− and S2−), reactive oxygen species (H2O2, ˙OH, O2˙− and HOCl), reactive nitrogen species (NO2−, NO3−, ONOO−, HNO and NO), biothiols (Cys, Hcy, GSH, and GSSG), and L-arginine induced negligible fluorescence changes, whereas HA triggered a pronounced fluorescence enhancement. Moreover, the fluorescence response of DCI-HA-2 toward HA was not affected in the presence of these potential interferents (Fig. S6).
Despite the fact that N2H4 is another reagent that can deprotect Dde (Fig. S7), it is unlikely to interfere with the detection of HA by using DCI-HA-2 because N2H4 is not present in most living systems. The performance of DCI-HA-2 under physiological conditions was further investigated. As seen in Fig. 1H, DCI-HA-2 remained stable over a broad pH range, and a significant fluorescence increase was observed upon HA addition in the pH range of 7.00–9.00, indicating its suitability for HA detection in living systems. Therefore, these results verify that DCI-HA-2 enables highly sensitive and selective detection of HA under biological conditions, highlighting its potential for real-time imaging of HA dynamics in living cells and the brain.
To further verify the universality of the design strategy, coumarin, fluorescein, and naphthalimide fluorophores were employed to develop HA-activated probes, namely Cou-HA, Fluo-HA, and Naph-HA (Fig. 2). Their synthetic routes are shown in Scheme S3 and the fluorescence behaviors of these probes toward HA were investigated. As seen in Fig. 2A and C, all three probes presented marked fluorescence enhancements at 460 nm, 525 nm and 555 nm, respectively, upon the addition of HA, accompanied by corresponding absorption changes (Fig. S8). Moreover, each probe exhibited a good linear fluorescence response toward HA: 0–15.0 µM with R2 = 0.9903 for Cou-HA (Fig. 2D), 0–10.0 µM with R2 = 0.9908 for Fluo-HA (Fig. 2E), and 0–20.0 µM with R2 = 0.9919 for Naph-HA (Fig. 2F). The LOD values were determined to be 68.7, 467.0, and 92.9 nM, respectively. In addition, all three probes exhibited high selectivity (Fig. S9), anti-interference performance (Fig. S10) and good photostability (Fig. S11). Overall, these results confirm that the Dde-COOH-2 moiety serves as a robust and versatile chemoselective handle for constructing HA-responsive probes.
Fluorescence imaging of exogenous and endogenous cellular HA
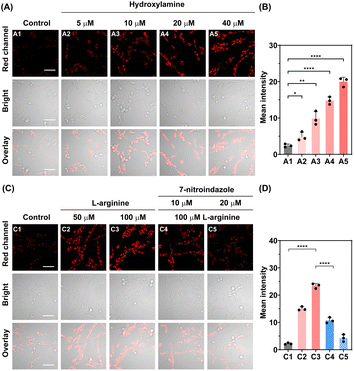
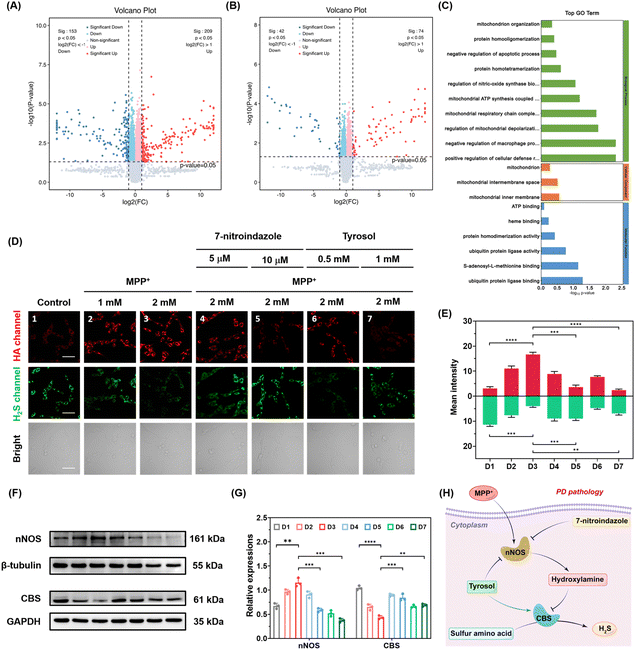
Low cytotoxicity and good biocompatibility are crucial for fluorescent probes used in cellular and brain imaging. To evaluate these, the cytotoxicity of DCI-HA-2 was examined using MTT assays. As seen in Fig. S12, PC-12 cells treated with DCI-HA-2 (10.0 µM) for 24 h exhibited a viability of 93.6%. Moreover, hematoxylin–eosin (H&E) staining indicated no discernible histological or morphological differences between the sham group and the probe-treated group (Fig. S13), confirming the biocompatibility of DCI-HA-2. We first investigated the ability of DCI-HA-2 to visualize the fluctuations in exogenous HA in living cells. As depicted in Fig. 3A, PC-12 cells incubated with DCI-HA-2 alone exhibited faint red fluorescence, indicating a low intrinsic level of intracellular HA under normal conditions. In contrast, when cells were pre-incubated with increasing concentrations of HA (0.0, 5.0, 10.0, 20.0 and 40.0 µM) prior to DCI-HA-2 staining, the red fluorescence intensity increased progressively (Fig. 3B), implying that DCI-HA-2 is capable of sensitively monitoring exogenous HA levels in living cells. Next, we evaluated the capability of DCI-HA-2 to track endogenous HA production.L-arginine was used as a precursor for endogenous HA generation via nNOS-catalyzed reaction,40 while 7-nitroindazole, a selective inhibitor of nNOS, was used to suppress HA production.41 As shown in Fig. 3C, incubation of PC-12 cells with L-arginine (50.0 and 100.0 µM) led to a marked enhancement of red fluorescence, corresponding to 6.81-, and 9.98-fold increases, respectively, confirming the endogenous nNOS-mediated generation of intracellular HA. However, pre-treatment of cells with 7-nitroindazole (10.0 and 20.0 µM), followed by stimulation with L-arginine (100 µM), led to a pronounced attenuation of fluorescence signals to 0.18- and 0.12-fold, respectively (Fig. 3D), suggesting effective inhibition of nNOS-dependent HA generation. These results indicate that DCI-HA-2 can sensitively monitor endogenous HA fluctuations in living PC-12 cells under pharmacological modulation.
Fluorescence imaging of HA changes in PD model cells
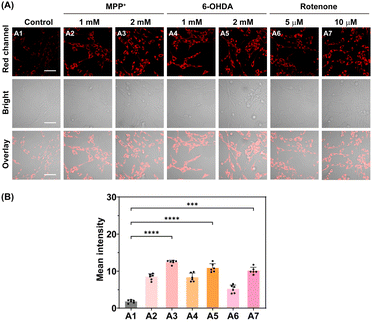
To further evaluate the HA dynamics under PD pathological conditions, PD cell models were established by exposing PC-12 cells to several neurotoxins, including 1-methyl-4-phenylpyridinium iodide (MPP+), 6-hydroxydopamine (6-OHDA), and rotenone, which are known to induce dopaminergic neuron loss.42 As shown in Fig. 4A, cells treated with each neurotoxin exhibited greatly enhanced red fluorescence compared with the control group, implying elevated intracellular HA levels in PD model cells. Quantitative analysis indicated that treatment with MPP+ (2.00 mM) resulted in a 6.92-fold increase in fluorescence intensity, while 6-OHDA (2.00 mM) and rotenone (10.0 µM) led to 5.95- and 5.63-fold increases, respectively (Fig. 4B). In addition, elevated HA levels were also observed in PD model SH-SY5Y cells and BV2 cells (Fig. S14). These findings convincingly demonstrate that HA is upregulated in PD model cells and further demonstrate that DCI-HA-2 can reliably monitor intracellular HA fluctuations under PD-related pathological conditions.Fluorescence imaging of HA variation in PD model mice
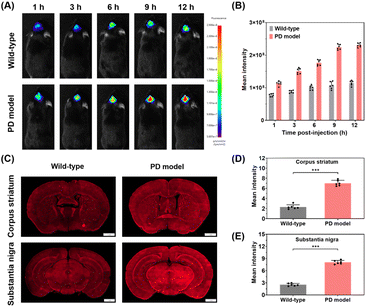
To investigate intracerebral levels of HA under PD conditions, PD model mice were established via intraperitoneal injection of 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP) for 7 consecutive days. DCI-HA-2 was then administered via tail vein injection to each group (n = 6 per group), and HA levels were evaluated by in vivo fluorescence imaging. As illustrated in Fig. 5A and S15, PD mice displayed markedly stronger fluorescence in the brain compared with the wild-type group, with intensity gradually increasing over time during blood circulation. After 12 h, the fluorescence intensity in the brains of PD mice was up to 2.05-fold higher compared with the control group (Fig. 5B). Then, brain tissues from each group were collected after euthanasia and subjected to imaging. As seen in Fig. S16, fluorescence imaging of brain tissues revealed a 2.15-fold increase in PD mice compared with wild-type mice. In contrast, imaging of major organs showed minimal probe accumulation (Fig. S17), except in metabolic organs such as the liver and kidney, suggesting good BBB permeability and brain enrichment of DCI-HA-2. Additionally, the pharmacokinetics of DCI-HA-2 were investigated by collecting plasma and brain tissues. As seen in Fig. S18 and Table S2, the half-life time of DCI-HA-2 was 5.34 h in the plasma and 9.44 h in the brain, suggesting relatively long systemic circulation time. Meanwhile, tissue distribution and clearance kinetics in the liver and kidney were further evaluated. As depicted in Fig. S19 and S20, DCI-HA-2 was mainly distributed in the liver and kidney, with higher accumulation in the liver than the kidney. The clearance rates in the liver and kidney were 1.11 mL min−1 and 3.68 mL min−1, respectively, suggesting that renal excretion is the primary clearance pathway (Table S3). Given the pathological relevance of the corpus striatum and substantia nigra in PD,43 brain slices were prepared and stained with DAPI for regional analysis. As shown in Fig. 5C–E, significantly brighter red fluorescence signals were observed in these regions in PD mice than in the wild-type group, with 3.05- and 3.18-fold increases in the corpus striatum and substantia nigra, respectively. These results revealed that DCI-HA-2 enables effective and specific visualization of elevated HA levels in the brains of PD mice.To address this question, we investigated the connection between HA and H2S in MPP+-induced PD model cells using the HA probe DCI-HA-2 and the H2S fluorescent probe N-H2S (Scheme S4). As depicted in Fig. 6D, MPP+-treated PD model cells displayed stronger HA-associated red fluorescence and weaker H2S-associated green fluorescence than the control group. Notably, the increase in red fluorescence and the decrease in green fluorescence became more pronounced with increasing MPP+ concentration. In particular, PD cells induced by MPP+ (2.00 mM) showed a 5.41-fold increase in red fluorescence and a 0.35-fold decrease in green fluorescence, suggesting HA overexpression accompanied by reduced H2S levels in PD cells. To prove the cascade relationship between HA and H2S, cells were successively pre-treated with 7-nitroindazole (20.0 µM) and MPP+ (2.00 mM), followed by staining with DCI-HA-2 and N-H2S. Under these conditions, the red fluorescence decreased to 0.22 times that in MPP+ (2.00 mM)-incubated cells, while the green fluorescence increased to 2.19-times that in MPP+ (2.00 mM)-incubated cells, suggesting that inhibition of HA generation leads to recovery of intracellular H2S levels. In addition, tyrosol, a known CBS activator, was used to upregulate the activity of CBS.45 Cells pre-treated with tyrosol (1.00 mM) prior to MPP+ (2.00 mM) exposure showed a 1.72-fold increase in H2S-related green fluorescence and a 0.15-fold decrease in HA-related red fluorescence (Fig. 6E), implying that tyrosol promotes H2S production while reducing cellular HA levels. To fully understand the above phenomenon, western blot (WB) analysis was performed to analyse the expression of nNOS and CBS under the above conditions (Fig. S18 and S19). As illustrated in Fig. 6F and G, compared with the control group, cells treated with 2.00 mM MPP+ showed upregulated nNOS expression (1.71-fold) and downregulated CBS expression (0.42-fold), which was consistent with the imaging results. Moreover, pre-treatment with 7-nitroindazole (20.0 µM) decreased the nNOS level to 0.49 times and enhanced the CBS level to 1.90 times that in the MPP+ (2.00 mM)-treated group, demonstrating a negative correlation between HA production and CBS expression. Similarly, cells pre-treated with tyrosol (1.0 mM) followed by MPP+ (2.00 mM) resulted in a 1.51-fold increase in the CBS level. Notably, a pronounced decrease in nNOS expression (0.34-fold) was also observed, likely due to the inhibitory effect of tyrosol on nNOS activity,46 which also explains the reduced HA-related red fluorescence observed in Fig. 5D. Taken together, these results reveal, for the first time, that elevated levels of HA derived from nNOS inhibit the expression of CBS, thereby diminishing the production of H2S in PD pathological cells (Fig. 6H).
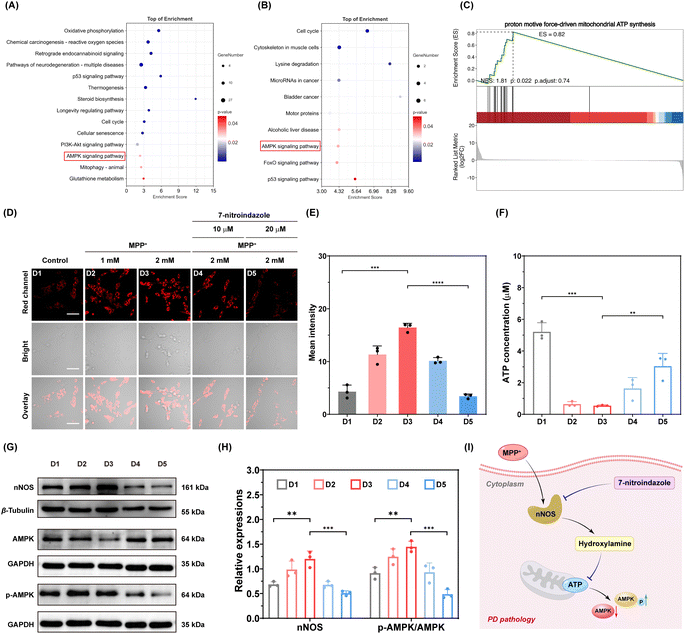
Next, we additionally delved into the regulatory role of HA in PD cells. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis revealed two biological pathways in both the PD group vs. control group (Fig. 7A) and 7-nitroindazole-treated PD group vs. PD group (Fig. 7B), namely the p53 signaling pathway and the AMPK signaling pathway. In particular, gene set enrichment analysis also showed that 7-nitroindazole treatment markedly upregulated genes associated with ATP synthesis in PD cells (Fig. 7C and S17), prompting us to investigate the involvement of HA in cellular energy metabolism. Similar to the above, an elevated level of HA was detected in PD cells (Fig. 7D), and pre-treatment with 7-nitroindazole can significantly inhibit the generation of cellular HA in PD cells (Fig. 7E). To determine whether the alteration in HA is associated with ATP synthesis-related signaling pathways in PD, a commercial ATP assay kit was employed to quantify the intracellular ATP levels under above conditions. It was seen that the level of ATP in 2.00 mM MPP+-treated cells was markedly reduced to 0.55 µM, corresponding to an 89.44% decrease compared with the control group (5.21 µM), revealing pronounced ATP depletion under PD conditions (Fig. 7F). In contrast, pre-treatment with 7-nitroindazole (20.0 µM) led to a 5.63-fold recovery of ATP levels (3.04 µM) relative to MPP+ treatment alone, indicating that suppression of HA production effectively promotes ATP synthesis. To further assess the involvement of HA in ATP-related AMPK signaling, WB analysis was performed to examine the expression of nNOS, AMPK and phosphorylated-AMPK (p-AMPK) under different treatment conditions (Fig. S20–S22). As shown in Fig. 7G and H, nNOS expression increased by 1.75-fold following 2.00 mM MPP+ treatment relative to control cells, consistent with the elevated HA level (brighter red fluorescence), whereas 7-nitroindazole (20.0 µM) treatment reduced nNOS expression by 57.6%. Moreover, the p-AMPK/AMPK ratio increased by 1.32-fold following 2.00 mM MPP+ treatment, implying the dysregulated AMPK signalling in PD. Notably, pre-treatment with 7-nitroindazole (20.0 µM) decreased the p-AMPK/AMPK ratio by 66.2% relative to 2.00 mM MPP+ treatment alone, suggesting that the inhibition of HA generation ameliorates ATP synthesis and normalizes AMPK signaling. Overall, these results demonstrate that nNOS-derived HA inhibits ATP synthesis and disrupts the AMPK signaling pathway during PD pathology, highlighting HA as a critical metabolic regulator in PD progression (Fig. 7I).
Conclusions
In conclusion, we have established a universal strategy for constructing highly specific HA-activated fluorescent probes across various fluorescent scaffolds based on chemo-selective cleavage and intramolecular nucleophilic cascade reactions, enabling sensitive and selective detection of HA in complex biological environments. The optimized probe, DCI-HA-2, enabled, for the first time, the real-time visualization of HA overexpression in PD cells and PD model mice. By integrating fluorescence imaging with comprehensive proteomic analysis, we revealed that nNOS-derived HA suppresses CBS, impairing H2S biosynthesis, and simultaneously induces ATP depletion, thereby disrupting AMPK signaling in PD pathology. These results establish HA as a previously unrecognized pathological mediator and highlight DCI-HA-2 as a versatile tool for probing HA-associated signaling in complex biological systems. This work offers new mechanistic insights into PD progression and lays the foundation for future diagnostic and therapeutic strategies targeting HA-mediated pathways.Ethical statement
All animal experiments were carried out in accordance with national guidelines and all procedures were approved by the ethical committees of Central South University (No. 430727211101478756) and Zhejiang University (ZJU20250124).Author contributions
S. Z. and J. K. contributed equally to this work. S. Z. and X. S. conceived the ideas and designed the studies. S. Z. and H. Z. carried out the synthesis and photophysical studies. S. Z. and J. K. performed the in vivo imaging and protein studies. S. Z., B. W. and X. S. wrote the manuscript. B. W., C. Y., M. L. and X. S. supervised the project.1–46 All authors reviewed and approved the final version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data of this study are available in the supplementary information (SI). Supplementary information: experimental materials, synthetic routes, characterizations, and other figures. See DOI: https://doi.org/10.1039/d6sc01655c.Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 22278447 and 22178395) and the Natural Science Foundation of Hunan Province (No. 2024JJ4050 and 2023JJ40793).Notes and references
- Y. Ben-Shlomo, S. Darweesh, J. Llibre-Guerra, C. Marras, M. San Luciano and C. Tanner, Lancet, 2024, 403, 283–292 CrossRef PubMed.
- W. Poewe, K. Seppi, C. M. Tanner, G. M. Halliday, P. Brundin, J. Volkmann, A.-E. Schrag and A. E. Lang, Nat. Rev. Dis. Primers, 2017, 3, 17013 CrossRef PubMed.
- H. R. Morris, M. G. Spillantini, C. M. Sue and C. H. Williams-Gray, Lancet, 2024, 403, 293–304 CrossRef CAS PubMed.
- M. J. Armstrong and M. S. Okun, JAMA, 2020, 323, 548–560 CrossRef PubMed.
- S. Przedborski and H. Ischiropoulos, Antioxid. Redox Signaling, 2005, 7, 685–693 CrossRef CAS PubMed.
- S. Z. Imam, G. D. Newport, Y. Itzhak, J. L. Cadet, F. Islam, W. Slikker Jr and S. F. Ali, J. Neurochem., 2001, 76, 745–749 CrossRef CAS PubMed.
- J. L. O'Connor, D. M. Fountos, B. Firouzan, F. Azizi, R. Ghasemi and K. Kashfi, Neurotherapeutics, 2025, 22, e00710 CrossRef PubMed.
- A. J. Väänänen, M. Moed, R. K. Tuominen, T. H. Helkamaa, M. Wiksten, P. Liesi, C. C. Chiueh and P. Rauhala, Free Radical Res., 2003, 37, 381–389 CrossRef PubMed.
- H. C. Çubukçu, M. Yurtdaş, Z. E. Durak, B. Aytaç, H. N. Güneş, B. G. Çokal, T. K. Yoldaş and İ. Durak, Neurol. Sci., 2016, 37, 1793–1798 CrossRef PubMed.
- Y. Kono and I. Fridovich, J. Biol. Chem., 1983, 258, 13646–13648 CrossRef CAS PubMed.
- P. Gross and R. P. Smith, CRC Crit. Rev. Toxicol., 1985, 14, 87–99 CrossRef CAS PubMed.
- N. Aleksidze and C. Blomstrand, Brain Res., 1968, 11, 717–719 CrossRef CAS PubMed.
- N. A. Correia, R. B. Oliveira and G. Ballejo, Life Sci., 2000, 68, 709–717 CrossRef CAS PubMed.
- H. Zhu, Z. Ma, X. Wang, L. Ji and B. Zhu, Coord. Chem. Rev., 2025, 544, 216963 CrossRef CAS.
- S. Wang, H. Song, C. Yin and F. Huo, Coord. Chem. Rev., 2025, 543, 216940 CrossRef CAS.
- L. Gao, W. Wang, X. Wang, F. Yang, L. Xie, J. Shen, M. A. Brimble, Q. Xiao and S. Q. Yao, Chem. Soc. Rev., 2021, 50, 1219–1250 RSC.
- J. Zhou, P. Jangili, S. Son, M. S. Ji, M. Won and J. S. Kim, Adv. Mater., 2020, 32, 2001945 CrossRef CAS PubMed.
- J. Wang, M. Cao, L. Han, P. Shangguan, Y. Liu, Y. Zhong, C. Chen, G. Wang, X. Chen, M. Lin, M. Lu, Z. Luo, M. He, H. H. Y. Sung, G. Niu, J. W. Y. Lam, B. Shi and B. Z. Tang, J. Am. Chem. Soc., 2024, 146, 28783–28794 CrossRef CAS PubMed.
- H. Li, J. Wang, H. Kim, X. Peng and J. Yoon, Angew. Chem., Int. Ed., 2024, 63, e202311764 CrossRef CAS PubMed.
- A. C. Sedgwick, R. S. L. Chapman, J. E. Gardiner, L. R. Peacock, G. Kim, J. Yoon, S. D. Bull and T. D. James, Chem. Commun., 2017, 53, 10441–10443 RSC.
- B. Dong, M. Tian, X. Kong, W. Song, Y. Lu and W. Lin, Anal. Chem., 2019, 91, 11397–11402 CrossRef CAS PubMed.
- P. Rana, L. Panda, N. Murmu, B. P. Bag and S. N. Sahu, Org. Biomol. Chem., 2020, 18, 5963–5971 RSC.
- W. Du, Y. Tian, Y.-A. Feng, X.-W. Cong, R. Tan, Y.-W. Wang and Y. Peng, Anal. Chim. Acta, 2024, 1318, 342941 CrossRef CAS PubMed.
- M. Du, H. Jiang, M. Song, Y. Zhang, H. Lv, S. Zhao, H. Du and Z. Dong, Anal. Methods, 2024, 16, 4843–4855 RSC.
- J. Ma, M. Zhao, X. Kong, Y. Liu, D. Meng and Z. Zhang, Dyes Pigm., 2026, 245, 113218 CrossRef CAS.
- A. C. Sedgwick, R. S. L. Chapman, J. E. Gardiner, L. R. Peacock, G. Kim, J. Yoon, S. D. Bull and T. D. James, Chem. Commun., 2017, 53, 10441–10443 RSC.
- Z. Liu, X. Zhou, Y. Miao, Y. Hu, N. Kwon, X. Wu and J. Yoon, Angew. Chem., Int. Ed., 2017, 56, 5812–5816 CrossRef CAS PubMed.
- Y. Yue, F. Huo, P. Ning, Y. Zhang, J. Chao, X. Meng and C. Yin, J. Am. Chem. Soc., 2017, 139, 3181–3185 CrossRef CAS PubMed.
- D. Cheng, Y. Pan, L. Wang, Z. Zeng, L. Yuan, X. Zhang and Y.-T. Chang, J. Am. Chem. Soc., 2017, 139, 285–292 CrossRef CAS PubMed.
- A. C. Sedgwick, R. S. L. Chapman, J. E. Gardiner, L. R. Peacock, G. Kim, J. Yoon, S. D. Bull and T. D. James, Chem. Commun., 2017, 53, 10441–10443 RSC.
- D. Cheng, Y. Pan, L. Wang, Z. Zeng, L. Yuan, X. Zhang and Y.-T. Chang, J. Am. Chem. Soc., 2017, 139, 285–292 CrossRef CAS PubMed.
- J. J. Díaz-Mochón, L. Bialy and M. Bradley, Org. Lett., 2004, 6, 1127–1129 CrossRef PubMed.
- J. Andersonmckay, G. P. Savage and G. W. Simpson, Aust. J. Chem., 1996, 49, 163–166 CrossRef CAS.
- E. Suárez-Picado, S. K. Ahirwar and L. R. Malins, Org. Lett., 2026, 28, 1815–1820 CrossRef PubMed.
- W. Zhang, F. Huo and C. Yin, J. Mater. Chem. B, 2018, 6, 6919–6929 RSC.
- L. Dai, Q. Zhang, Q. Ma and W. Lin, Coord. Chem. Rev., 2023, 489, 215193 CrossRef CAS.
- F. Liu, X. Ding, X. Xu, F. Wang, X. Chu and J.-H. Jiang, Angew. Chem., Int. Ed., 2022, 61, e202203243 CrossRef CAS PubMed.
- A. Alouane, R. Labruère, T. Le Saux, F. Schmidt and L. Jullien, Angew. Chem., Int. Ed., 2015, 54, 7492–7509 CrossRef CAS PubMed.
- W. Wei, C. Huang, J. Zhang, Q. Chen, Z. Liu, X. Ren, S. Gan, P. Wu, D. Wang, B. Z. Tang and H. Sun, Anal. Chem., 2024, 96, 2406–2414 CrossRef CAS PubMed.
- T. L. Poulos and H. Li, Acc. Chem. Res., 2013, 46, 390–398 CrossRef CAS PubMed.
- P. E. Erik, A. L. Elizabeth, S. Jiwon, Z. Yaoqiu, J. Haitao and B. S. Richard, Curr. Top. Med. Chem., 2005, 5, 603–624 CrossRef PubMed.
- D. Blum, S. Torch, N. Lambeng, M.-F. Nissou, A.-L. Benabid, R. Sadoul and J.-M. Verna, Prog. Neurobiol., 2001, 65, 135–172 CrossRef CAS PubMed.
- Y. Lu, X. Zhang, L. Zhao, C. Yang, L. Pan, C. Li, K. Liu, G. Bai, H. Gao and Z. Yan, Front. Neurosci., 2018, 12, 2018 Search PubMed.
- Y.-W. Wang, T. Chu, X.-L. Wang, Y.-Q. Fan, L. Cao, Y.-H. Chen, Y.-W. Zhu, H.-X. Liu, X.-Y. Ji and D.-D. Wu, Cell. Signalling, 2024, 124, 111406 CrossRef CAS PubMed.
- L. K. Sarna, V. Sid, P. Wang, Y. L. Siow, J. D. House and K. O, Lipids, 2016, 51, 583–590 CrossRef CAS PubMed.
- P. Dewapriya, S. W. A. Himaya, Y.-X. Li and S.-K. Kim, Food Chem., 2013, 141, 1147–1157 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |