Open Access Article
Open Access ArticleMAPLE: a machine-learning force-field-native platform for automated reaction modeling and enzyme design
Xujian Wang abc,
Zeyu Suna,
Yilu Zhanga,
Carlo Asamb,
Ruzhan Zhud,
Wan-Lu Li
abc,
Zeyu Suna,
Yilu Zhanga,
Carlo Asamb,
Ruzhan Zhud,
Wan-Lu Li *be and
Junmei Wang*a
*be and
Junmei Wang*a
aDepartment of Pharmaceutical Sciences and Computational Chemical Genomics Screening Center, School of Pharmacy, University of Pittsburgh, Pittsburgh, Pennsylvania 15261, USA. E-mail: junmei.wang@pitt.edu
bAiiso Yufeng Li Family Department of Chemical and Nano Engineering, University of California San Diego, CA 92093, USA. E-mail: wal019@ucsd.edu
cDepartment of Computational and Systems Biology, School of Medicine, University of Pittsburgh, Pittsburgh, Pennsylvania 15261, USA
dSchool of Engineering, Computer and Mathematical Sciences, Auckland University of Technology, Auckland 1010, New Zealand
eProgram of Materials Science and Engineering, University of California San Diego, CA 92093, USA
First published on 30th April 2026
Abstract
Machine-learning force fields (MLFFs) are reshaping computational chemistry and biology by delivering near-quantum mechanical accuracy at a computational cost comparable to conventional force fields, enabling applications in biomolecular simulation, catalysis, and materials science. However, despite these advances, a unified and automated computational platform enabling the broader application of MLFFs is still lacking. Here, we present MAPLE (MAchine learning Potential for Landscape Exploration), a computational toolkit specially developed for MLFF-based molecular modeling, featuring a tailored software framework and parallelized algorithms for large-scale and versatile molecular modeling tasks. We demonstrated the robustness and usability of MAPLE through systematic benchmarking of state-of-the-art reactive MLFFs and applications to multiple biocatalytic scenarios, highlighting its capability for fast yet accurate simulation of catalytic reactions. By integrating accurate and efficient MLFFs with parallelized algorithms in a highly optimized and flexible software framework, MAPLE serves as a next-generation, physically informed, machine-learning-driven molecular modeling platform with broad applicability to rational catalyst design and drug discovery.
1 Introduction
The development of molecular mechanics force fields (MMFFs) has enabled atomistic simulations of molecular systems,1,2 providing a foundation for studying molecular interactions,3 biomolecular conformations,4 and catalytic mechanisms.5 Despite these successes, the applicability of MMFFs remains intrinsically limited by their simplified functional forms, which are not suitable for studying nonequilibrium systems and struggle to capture chemically complex phenomena, particularly bond breaking and formation. In contrast, quantum mechanical (QM) methods provide a more rigorous description of these effects, but at a substantially higher computational cost, thereby limiting QM simulations to relatively small systems and short timescales.During the past decade, machine-learning force fields (MLFFs) have substantially narrowed the gap between conventional MMFFs and QM methods by learning potential energy surfaces (PESs) directly from QM data, achieving near-QM accuracy at computational cost of the MMFF-level.6–8 Among them, the AIMNet2 family has demonstrated robust performance in a wide range of chemical scenarios, including catalysis9,10 and biomolecular simulations.11 More recently, the release of the Open Molecules 2025 dataset (OMol25)12 enabled the training of the Universal Model Atom (UMA),7 a large-scale MLFF that covers 83 elements and is capable of describing chemical reactions and open-shell systems. Together, these advances represent an important step toward a universal MLFF.
Despite rapid progress in MLFFs, their practical deployment remains constrained by the lack of dedicated, MLFF-native computational infrastructures. Most existing toolkits, such as Sella,13 MLatom 3,14 and the Atomic Simulation Environment (ASE),15 are designed as general-purpose frameworks and are not specifically optimized for MLFF-driven workflows, and also do not fully leverage a key advantage of MLFFs: their suitability for large-scale parallel evaluation across large ensembles of molecular systems. This capability, which enables high-throughput molecular simulations, can be realized through parallel optimization algorithms. This limitation becomes especially pronounced for reaction discovery and pathway sampling, where the intrinsic parallelism of MLFFs could enable high-throughput exploration across multiple configurations, intermediates, or reaction channels. A representative example is the batched nudged elastic band (B-NEB) method,9 which demonstrates how an MLFF can be coupled with parallel algorithms to efficiently explore reaction pathways. However, effective use of current toolkits depends heavily on users' programming skills, creating a high technical barrier that restricts accessibility beyond specialist developers and hinders broader adoption in applied and industrial settings. Together, these challenges highlight the urgent need for MLFF-native software infrastructures that are both GPU-oriented and user-friendly, supporting advanced parallel algorithms while remaining practical for a broad research community.
In this work, we developed MAPLE (MAchine learning Potential for Landscape Exploration), a computational platform specifically designed to enable the effective application of MLFF across diverse research fields. The current release of MAPLE offers a comprehensive set of core capabilities, including single-point energy calculations, geometry optimizations, PES scans, transition state (TS) searches, and frequency analyses. We demonstrated the robustness and reliability of MAPLE's core algorithms in transition state optimizations using three general-purpose reactive MLFFs. We further applied MAPLE to investigate enzymatic catalysis across three distinct enzymes and their corresponding mutants, demonstrating that the platform consistently predicts accurate potential energy profiles and reliable geometric structures in catalytic processes. These results underscore MAPLE's potential to enable efficient, interpretable, and high-resolution computational molecular design. We envision MAPLE as a next-generation MLFF-based computational chemistry platform that not only broadens their applications but also provides a foundation for developing advanced MLFF algorithms, ultimately reshaping the computational paradigm of the field.
2 Results
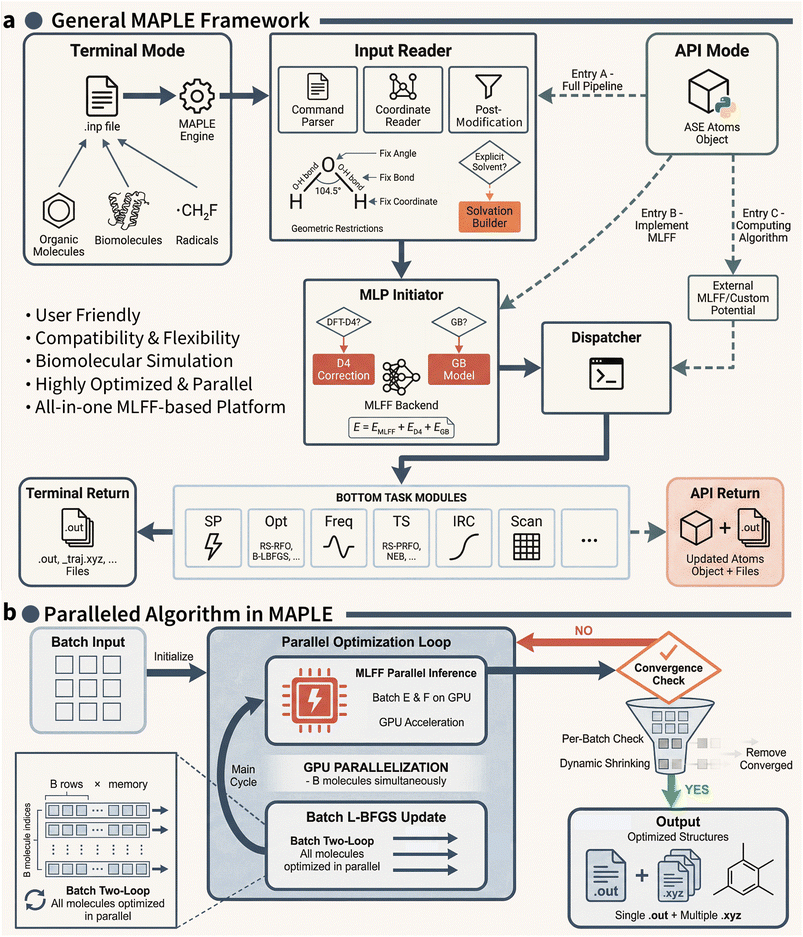
2.1 The MAPLE framework
MAPLE is a molecular modeling platform developed from scratch, designed to provide a unified and extensible framework for modern MLFF-based molecular modeling. MAPLE consists of three core components (Fig. 1a): an Input Reader, a Machine-Learning Potential (MLP) Initiator, and a suite of task modules.The Input Reader serves as the central control unit of MAPLE, translating user-defined inputs into customized computational workflows, including task selection, parameter setting, and device allocation. Unlike conventional computational chemistry packages, MAPLE supports concurrent loading of multiple molecular systems in various coordinate formats, enabling efficient parallel execution. The Input Reader further integrates post-modification utilities for enforcing structural constraints, such as fixed coordinates or bond lengths, as well as a built-in explicit solvation builder. Leveraging the efficiency of MLFFs, MAPLE enables routine simulations of large systems and supports explicit treatment of various solvent effects by providing a library of nearly one hundred equilibrium explicit solvent environments.
The MLP Initiator interfaces with multiple state-of-the-art MLFFs, including the ANI,8,16–18 AIMNet2,6,19 MACE,20–22 and UMA.7 Users may employ built-in calculators or define custom potentials. Moreover, to improve robustness and transferability, MAPLE incorporates DFT-D4 dispersion corrections23,24 and generalized Born solvation (GB) models25,26 as optional corrections to the underlying potentials.
Based on these components, MAPLE supports a broad range of molecular simulation tasks through MLFF-compatible parallel algorithms. Notably, a batched L-BFGS optimizer enables the simultaneous optimization of large ensembles of molecular systems within a single workflow (Fig. 1b). Simulation results are exported in ORCA-27 and Gaussian-style28 formats, ensuring compatibility with standard analysis and visualization tools. All workflows can be defined through a unified input file, with extended control available via an application programming interface (API). A complete summary of all functionalities currently available in MAPLE is provided in Table S1 of the SI. Comprehensive documentation and tutorials are provided at https://www.maplechem.org/.
2.2 Assessing reactive MLFFs for transition-state optimization
Non-equilibrium properties, such as TS, are central to chemical reactivity and catalysis. Therefore, a comprehensive benchmark of reactive MLFF performance in TS optimization is essential. Here, 100 test cases were randomly selected from the Transition1x dataset,29 each comprising a reactant–product pair, for more details about the test set, please refer to Table S2. All MLFF-based calculations were performed using the MAPLE program, while DFT calculations at the ωB97M-V/def2-TZVPD level30,31 were carried out using ORCA 6.0.First, each reactant–product pair was subjected to climbing-image nudged elastic band (CI-NEB)32 calculations using the same number of images to generate reaction pathways. MAPLE demonstrated high robustness, with all four MLFFs achieving success rates of at least 78% (Table 1). Although exact HEI positions often differed from DFT, relaxed criteria (within one frame of the DFT reference) yielded significantly improved agreement, with UMA achieving a success rate over 95%. These results indicate that MAPLE, in conjunction with MLFFs, can generate physically meaningful reaction pathways. Consistently, geometric analysis showed that MAPLE-optimized structures agree very well with their DFT counterparts, as indicated by small RMSDs of ≤0.4 Å for both the full pathways and HEI structures (Table 1). In the energetic assessment, ANI-1xnr shows pronounced deviations, with errors exceeding 40 kcal mol−1, reflecting the limited quantitative reliability of this early-generation reactive MLFF. Inspection of the largest ANI-1xnr outliers indicates that these errors are concentrated in highly strained ring-opening or fragmentation reactions, as well as heteroatom rearrangements, suggesting that such motifs are not adequately covered by the training set of this earlier-generation reactive potential. In contrast, the other models exhibit substantially improved agreement with typical energy deviations below 10 kcal mol−1, indicating their suitability for accurate evaluation of reaction-pathway energetics.
| Potentials | Success ratea | HEI prediction | Geometry RMSD | Activation energy | |||
|---|---|---|---|---|---|---|---|
| Exactb | ±1 framec | Pathwayd | HEIe | Opt.f | Ref. geom.g | ||
| a Successful NEB convergence out of 100 samples (DFT reference: 92%).b Percentage where predicted highest-energy image (HEI) matches DFT-identified frame.c Percentage where predicted HEI is within one frame of DFT result.d Mean RMSD across all pathway images (exclude end-points) relative to DFT geometries.e RMSD of highest-energy image relative to DFT geometry.f Unsigned energy difference between MLFF-optimized CI-NEB barriers and DFT reference.g Unsigned energy difference from single-point calculations on DFT-optimized geometries. Units: RMSD in Å, energy in kcal mol−1. The most accurate results are highlighted in bold. | |||||||
| ANI-1xnr | 78% | 27.6% | 65.8% | 0.38 ± 0.19 | 0.40 ± 0.06 | 44.2 ± 24.6 | 27.4 ± 20.0 |
| ANI-1xnr-D4 | 76% | 28.2% | 71.8% | 0.32 ± 0.03 | 0.33 ± 0.03 | 44.5 ± 25.5 | 27.4 ± 20.0 |
| AIMNet2-NSE | 85% | 34.6% | 88.9% | 0.26 ± 0.14 | 0.24 ± 0.02 | 6.3 ± 4.5 | 9.8 ± 7.2 |
| UMA | 93% | 39.8% | 95.5% | 0.24 ± 0.14 | 0.19 ± 0.03 | 2.0 ± 4.5 | 0.6 ± 1.2 |
| MACE-POL-S | 79% | 32.9% | 86.8% | 0.26 ± 0.16 | 0.24 ± 0.05 | 5.9 ± 7.6 | 4.0 ± 3.0 |
| MACE-POL-L | 89% | 32.9% | 85.9% | 0.26 ± 0.20 | 0.22 ± 0.06 | 4.8 ± 12.9 | 1.7 ± 1.3 |
We further assessed the Hessian quality of the MLFFs from both analytic evaluation via PyTorch automatic differentiation33 and numerical differentiation. The quality of the resulting Hessians was evaluated using two criteria: the number of imaginary vibrational modes and the transition vector, defined as the eigenvector corresponding to the single negative eigenvalue of the Hessian at the first-order saddle point. The transition vector directly encodes the reaction coordinate direction and is thus the most chemically significant vibrational mode for TS characterization. Across all performance metrics, MACE-POL-L22 performs particularly well, with the lowest mean absolute error (MAE) of the imaginary frequency and the highest transition vector overlap relative to the DFT references (Table 2), indicating its strong capability for accurate TS characterization. MACE-POL-S is less accurate but still yields chemically meaningful transition vectors and TS geometries. A more detailed decomposition of the imaginary-frequency count mismatches is provided in Table S4. Notably, none of the evaluated MLFFs were explicitly trained using Hessian information, which suggests that MLFFs can implicitly encode physically meaningful second-derivative information by learning solely from energies and forces.
| Potentialsa | Hessian quality | TS optimization | |||||
|---|---|---|---|---|---|---|---|
| Acc.b | MAE | Overlapc | Angled | TS geom. | ΔIter | Runtimee | |
| a Frequency calculation methods: anal. = analytic; num. = numerical.b Percentage with correct number of imaginary frequencies.c Transition vector overlap with DFT (mass-weighted).d Calculated as arccos (overlap).e DFT takes 191.6 minutes on average. MLFF runtimes measured on an Intel Xeon w7-3455 CPU with a single NVIDIA RTX 5070 Ti GPU units: MAE in cm−1, angle in degrees, TS geom. in Å, runtime in minutes. The most accurate results are highlighted in bold. | |||||||
| ANI-1xnr (anal.) | 6.2% | 291 ± 265 | 0.83 ± 0.24 | 28.6 ± 21.2 | 0.59 ± 0.33 | 53.4 | 0.4 |
| ANI-1xnr (num.) | 6.2% | 292 ± 265 | 0.83 ± 0.24 | 28.6 ± 21.2 | 0.59 ± 0.32 | 51.1 | 0.8 |
| AIMNet2-NSE (anal.) | 31.2% | 529 ± 600 | 0.85 ± 0.24 | 25.9 ± 20.6 | 0.23 ± 0.19 | 10.8 | 0.1 |
| AIMNet2-NSE (num.) | 32.3% | 521 ± 567 | 0.85 ± 0.24 | 25.9 ± 20.6 | 0.22 ± 0.16 | 15.7 | 0.5 |
| UMA (num.) | 86.5% | 75 ± 109 | 0.99 ± 0.05 | 6.1 ± 6.6 | 0.09 ± 0.20 | 3.2 | 0.9 |
| MACE-POL-S (anal.) | 47.9% | 182 ± 187 | 0.94 ± 0.10 | 16.2 ± 12.2 | 0.24 ± 0.22 | 17.2 | 1.0 |
| MACE-POL-S (num.) | 47.9% | 182 ± 187 | 0.94 ± 0.10 | 16.2 ± 12.2 | 0.24 ± 0.22 | 19.5 | 1.5 |
| MACE-POL-L (anal.) | 85.4% | 54 ± 56 | 0.99 ± 0.01 | 5.4 ± 2.6 | 0.09 ± 0.13 | −1.6 | 3.5 |
| MACE-POL-L (num.) | 86.5% | 55 ± 55 | 0.99 ± 0.01 | 5.9 ± 2.8 | 0.11 ± 0.19 | −1.8 | 3.7 |
Finally, we performed TS optimizations using the proposed Dual-Shift Partitioned Rational Function Optimization (DS-P-RFO) algorithm, which is better suited for MLFF-based applications. MACE-1 POL-L performs best in TS localization, achieving a geometric deviation of 0.09 Å relative to the DFT references (Table 2). In comparison, UMA and AIMNet2-NSE also performed robustly, with deviations below 0.25 Å. Additional TS optimizations initialized from MLFF-derived HEI structures resulted in similarly modest geometric deviations (Table S1), underscoring the capability of MLFFs in providing high-quality initial guesses. In terms of computational efficiency, although AIMNet2-NSE and ANI-1xnr required more iterations to converge than UMA and MACE (Table 2), their overall wall-clock time remained significantly shorter and were orders of magnitude faster than the corresponding DFT calculation. These results highlight the strong potential of MLFF-based TS optimization workflows for high-throughput reaction screening applications.
It is worth noting that ANI-1xnr and AIMNet2-NSE were trained on independent DFT datasets at different levels of theory, which likely contributes to their comparatively reduced accuracy in reproducing DFT reference results in the above tests. Overall, UMA and MACE-POL deliver consistently high accuracy across TS-related tasks, while AIMNet2-NSE offers a favorable balance between accuracy and computational efficiency. Importantly, these benchmarks validate the reliability and robustness of MAPLE's core functionalities.
2.3 Whole-enzyme modeling of Kemp eliminase with MLFFs
We further validate MAPLE's functionalities in a real-world enzymatic setting. As an initial test case, we selected Kemp eliminase (KE),34 a well-studied computationally designed enzyme that catalyzes a single-step Kemp elimination reaction via proton transfer35–39 (Fig. 2a). The full enzyme system, comprising nearly 4000 atoms, was explicitly modeled using UMA MLFF. We note that all energies reported from MAPLE in the enzymatic case studies correspond to PES values, which serve as approximations to activation free energies.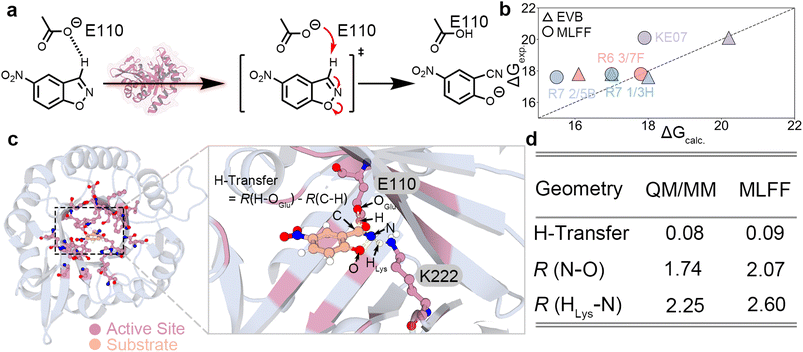 | ||
| Fig. 2 MLFF modeling of Kemp eliminase catalysis. (a) Reaction scheme of the Kemp elimination reaction catalyzed by Kemp eliminase. (b) Comparison of MLFF-derived activation energies and EVB activation free energies36 with experimentally measured values (ΔGexp) for KE07 and selected directed-evolution mutants. All values in kcal mol−1. (c) Representative TS structure obtained from MLFF simulations for KE07. The hydrogen-transfer coordinate is defined as the difference between the breaking C–H bond and the forming H–OGlu bond. All values in Å. (d) Comparison of selected TS geometric parameters between QM/MM35 and MLFF calculations. | ||
Specifically, KE07 and three of its directed-evolution mutants were investigated using MAPLE. In these simulations, residues within 8 Å of the reaction center were treated as flexible, while the remaining residues were kept fixed. The reported MLFF activation energies are derived from the PES. For these systems, empirical valence bond (EVB) activation free energies and experimentally characterized ΔG‡ values34,36 have been reported previously. Comparison of the MLFF-predicted activation energies (ΔG‡, approximated from PES barriers ΔE‡) with the experimental values (Fig. 2b) reveals that both MLFF and EVB exhibit only one outlier (R7 2/5B for MLFF and R6 3/7F for EVB). In most cases, however, the predictions agree well with experiment, with deviations within a few kcal mol−1 despite the fundamentally different levels of theory. To further assess the quality of the MLFF description, we analyzed the TS structure of KE07 obtained from our simulations (Fig. 2c). A key descriptor of the reaction coordinate is the hydrogen-transfer metric,35 defined as the difference between the breaking C–H bond length and the forming H–OGlu bond length (Fig. 2c). This metric shows excellent agreement with QM/MM references at the B3LYP/6-31G*//OPLS-AA level,40–42 with deviations of only 0.01 Å (Fig. 2d). Other geometric parameters, such as N–O and HLys–N distances, exhibit slightly larger deviations but remain well within physically reasonable ranges.
Overall, this example demonstrates the successful application of MLFFs to enzymatic reactions, yielding reasonable activation energies and physically meaningful TS geometries. Notably, unlike theozyme models that approximate second-shell residues using implicit solvent, or QM/MM approaches that treat indirectly involved residues at the MM level, MLFFs enable explicit modeling of the entire enzyme system at near-QM accuracy. This unified treatment provides a physically consistent and holistic description of enzymatic catalysis.
2.4 Quantifying stereoselectivity in Diels–Alderase catalysis
The MaDA family represents a recently discovered class of stand-alone intramolecular Diels–Alderases capable of producing highly stereoselective products43–47 (Fig. 3a), highlighting their potential for industrial applications. Benefiting from the computational efficiency of MLFFs, MAPLE enables explicit treatment of solvent effects, facilitating a realistic description of the reaction system. We first scanned the PESs of the reactants (1 and 2) in vacuum, aqueous solution, and ether for both enantiomeric pathways using UMA. The resulting PESs reveal asynchronous formation of the two C–C bonds (Fig. S1), consistent with prior experimental and computational studies,43,44 and capture solvent cage effects that modulate the chemical reaction (Fig. 3b). Quantitative activation energies obtained from CI-NEB calculations agree well with previous DFT43,44 and ML/MM results48 (Fig. 3c). Distinct treatment of solvent effects, for instance, the use of explicit solvent models in DFT and ensemble averaging in ML/MM calculations, likely contribute substantially to the observed variations in activation energy predictions.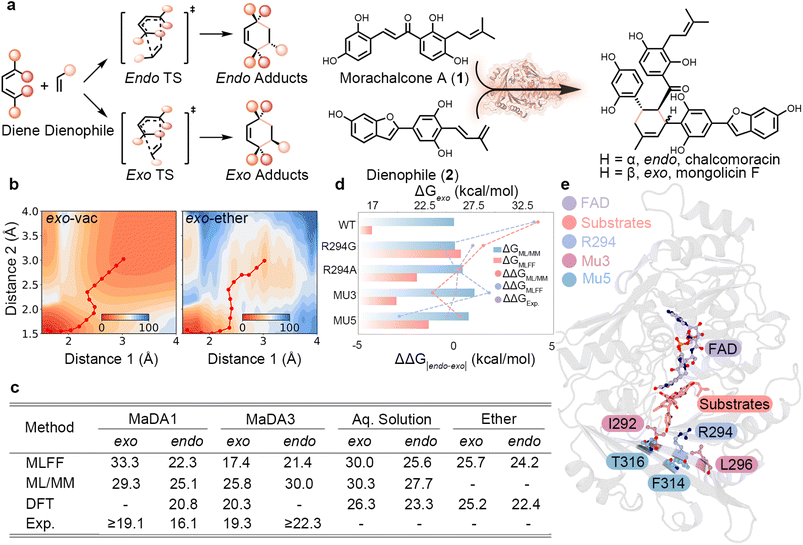 | ||
| Fig. 3 Stereoselectivity in MaDA-catalyzed intramolecular Diels–Alder reactions. (a) Reaction scheme illustrating competing endo and exo transition states and products in Diels–Alder reactions. (b) PESs for the exo pathway in vacuum and ether environments, computed with UMA, highlighting minimum-energy pathways (red lines). (c) Summary of free energies (ΔG‡, kcal mol−1) for endo and exo pathways in MaDA1, MaDA3, and solution-phase reactions, comparing MLFF, ML/MM,48 DFT,43,44 and experimental data.43,44 (d) Comparison of computed and experimental stereoselectivity across wild-type MaDA3 and selected mutants, expressed as ΔΔGendo–exo. (e) Representative active-site model of MaDA showing the substrate bound in proximity to the FAD cofactor and key residues involved in stereocontrol. | ||
Next, we investigated enzymatic catalysis in the MaDA family by studying MaDA1 and MaDA3, which preferentially yield endo and exo products, respectively. Full enzyme models containing over 8000 atoms were constructed. As in the Kemp eliminase case, only the residues within 8 Å of the reaction center were allowed to relax. However, all atoms, including constrained ones, were included in the MLFF energy and force evaluation. Estimated activation energies are comparable to conventional theozyme-based DFT models,43,44 with residual errors of approximately 3 kcal mol−1. While DFT and ML/MM48 calculations typically require days to weeks, MLFF-based calculations achieve a favorable balance between accuracy and efficiency, with most simulations completed within a day, underscoring their potential for rapid characterization of enzymatic catalysis.
We last examined MaDA3 and a series of mutants exhibiting systematic changes in stereoselectivity, including R294G, R294A, MU3 (I292F, R294G, L296R), and MU5 (MU3 plus F314I and T316S) (Fig. 3e). The wild-type MaDA3 yields exclusively exo products, with mutations progressively shifting selectivity toward the endo pathway (exo/endo down to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2). Although ML/MM and MLFF approaches differ in absolute ΔG‡ values, both capture consistent trends in ΔΔG‡, reflecting changes in stereoselectivity (Fig. 3d). It is noted that stereoselectivity depends exponentially on ΔΔG‡, such that even modest errors in ΔΔG‡ on the order of 1–2 kcal mol−1 can lead to substantial changes in the predicted endo/exo ratio. Therefore, while the consistent reproduction of experimental selectivity trends across MaDA3 variants validates MLFF-based ΔΔG‡ as a practical tool for ranking candidate mutations and identifying selectivity switches, the absolute selectivity ratios should be interpreted with caution. In the absence of experimental reference values for absolute activation energies, accurate prediction of ΔΔG‡ provides a practical and efficient means to screen candidate mutations, substantially reducing experimental trial-and-error and guiding rational enzyme design.
2). Although ML/MM and MLFF approaches differ in absolute ΔG‡ values, both capture consistent trends in ΔΔG‡, reflecting changes in stereoselectivity (Fig. 3d). It is noted that stereoselectivity depends exponentially on ΔΔG‡, such that even modest errors in ΔΔG‡ on the order of 1–2 kcal mol−1 can lead to substantial changes in the predicted endo/exo ratio. Therefore, while the consistent reproduction of experimental selectivity trends across MaDA3 variants validates MLFF-based ΔΔG‡ as a practical tool for ranking candidate mutations and identifying selectivity switches, the absolute selectivity ratios should be interpreted with caution. In the absence of experimental reference values for absolute activation energies, accurate prediction of ΔΔG‡ provides a practical and efficient means to screen candidate mutations, substantially reducing experimental trial-and-error and guiding rational enzyme design.
2.5 Photoactivated DNA repair by photolyase
DNA is vulnerable to ultraviolet irradiation, which can induce dimerization between adjacent pyrimidines to form pyrimidine(6-4)pyrimidone photoproducts49,50 ((6-4)-PP; Fig. 4a), leading to mutagenesis and genetic disease. DNA photolyase is a key repair enzyme that reverses such lesions through a photoactivated radical mechanism.51 Here, this complex photoinduced repair process was investigated using MAPLE, which enables MLFF-driven simulation of large-scale, non-equilibrium biochemical reactions. UMA was employed here to simulate the repair reaction, and previously reported quantum mechanics/molecular mechanics (QM/MM) studies52 were adopted as references.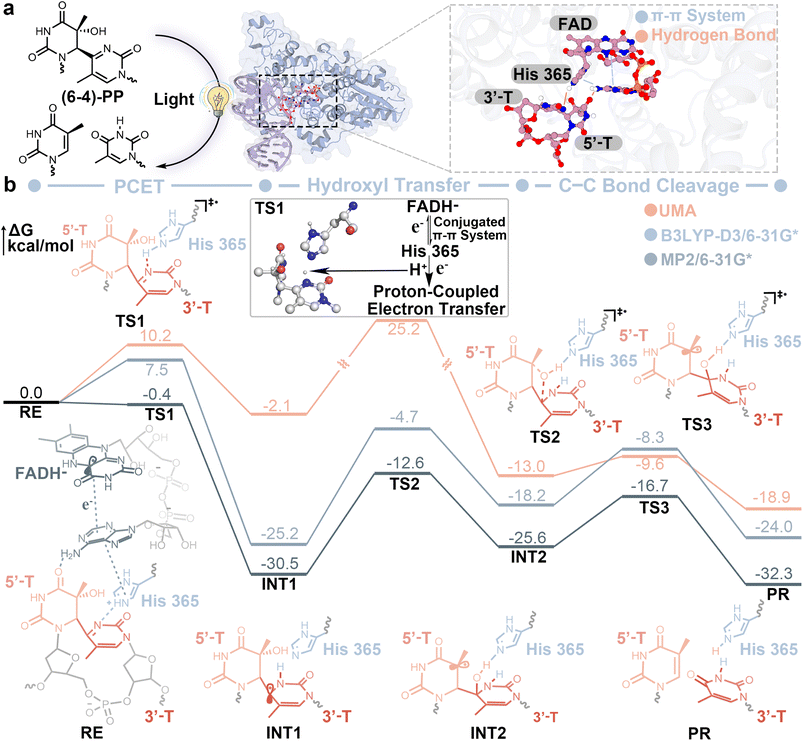 | ||
| Fig. 4 Photoactivated DNA repair mechanism catalyzed by DNA photolyase. (a) Schematic illustration of light-induced repair of (6-4)-PP by DNA photolyase and a representative active-site configuration highlights key interactions between FAD, His365, and the damaged DNA bases. (b) Energy profiles of the repair reaction obtained from UMA (orange), B3LYP-D3/6-31G* QM/MM (blue), and MP2/6-31G* QM/MM (gray) calculations.52 Representative structures along the reaction pathway are shown with corresponding reactant (RE), transition states (TS1–TS3), intermediates (INT1–INT2), and repaired product (PR). | ||
Using MAPLE, we modeled the whole DNA photolyase–DNA complex explicitly, resulting in a system of nearly 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 atoms. The reaction is initiated from reduced flavin adenine dinucleotide (FADH−) bound to (6-4)-PP in the enzyme active site, with the system set to a doublet state to mimic post-photoactivation.52 MAPLE enables efficient propagation of such non-equilibrium reactive trajectories by coupling MLFF-based force evaluations with automated pathway construction and reaction-pathway analysis, allowing direct interrogation of multi-step biochemical mechanisms at atomistic resolution. Upon excitation, MAPLE captures the redistribution of electron density through the π–π conjugated active-site network toward H365 (Fig. 4a). The overall repair process can be divided into three elementary steps: proton-coupled electron transfer (PCET), hydroxyl transfer, and C–C bond cleavage (Fig. 4b).
000 atoms. The reaction is initiated from reduced flavin adenine dinucleotide (FADH−) bound to (6-4)-PP in the enzyme active site, with the system set to a doublet state to mimic post-photoactivation.52 MAPLE enables efficient propagation of such non-equilibrium reactive trajectories by coupling MLFF-based force evaluations with automated pathway construction and reaction-pathway analysis, allowing direct interrogation of multi-step biochemical mechanisms at atomistic resolution. Upon excitation, MAPLE captures the redistribution of electron density through the π–π conjugated active-site network toward H365 (Fig. 4a). The overall repair process can be divided into three elementary steps: proton-coupled electron transfer (PCET), hydroxyl transfer, and C–C bond cleavage (Fig. 4b).
In the first step, UMA captures the PCET barrier with a deviation of 2.7 kcal mol−1 (10.2 vs. 7.5 kcal mol−1) relative to the QM/MM reference at the B3LYP-D3/6-31G* level.40,41,53 However, the stability of the resulting intermediate INT1 is severely underestimated by UMA, which predicts a relative energy of approximately −2 kcal mol−1 compared to approximately −30 kcal mol−1 at the DFT/MM and MP2/MM levels. This large deviation suggests that the PCET mechanism, involving tightly coupled electron and proton transfer within a radical active site, is likely underrepresented in current MLFF training datasets. In the second step (hydroxyl transfer), UMA overestimates the barrier height, yielding a maximum ΔG of 27.3 kcal mol−1 compared to 20.5 and 17.9 kcal mol−1 from two QM/MM calculations. In the final step, which involves pyrimidone dissociation and electron back-transfer to regenerate FADH−, UMA underestimates the barrier, predicting a ΔG of 3.4 kcal mol−1 relative to 9.9 and 8.9 kcal mol−1 reported from QM/MM. Notably, the later steps (TS2, INT2 to PR) show considerably better agreement with QM/MM, suggesting that the model generalizes reasonably well for conventional bond-breaking processes but remains limited for the complex electronic structure rearrangements characteristic of radical biochemistry. The remaining discrepancies between UMA and QM/MM can be attributed to two main factors: limitations in the description of long-range interactions and the absence of extensive biochemical reaction data during training. It is noted that the accurate treatment of long-range electrostatics remains an open challenge for both current MLFFs and early QM/MM implementations. Addressing this limitation in MLFF architectures is an active area of research.54 Overall, despite these quantitative discrepancies, it is worth noting that different QM-based approaches themselves can exhibit large variations, and UMA is trained on datasets generated at levels of theory that differ from those used in the two QM/MM studies. Within this context, MAPLE together with UMA reproduces the established DNA photolyase repair mechanism and correctly identifies hydroxyl transfer as the rate-limiting step, demonstrating its promise for modeling complex biochemical systems while highlighting clear areas for improvement in MLFF training and architecture.
3 Discussion
In this work, we developed MAPLE, a general framework that integrates MLFFs into automated molecular modeling and simulation workflows. By unifying MLFF evaluation, parallel optimization algorithms, and reaction-pathway analysis within a single platform, MAPLE addresses a critical gap between recent advances in MLFF development and their practical deployment in chemically complex systems.Through systematic benchmarks, we assessed the performance of contemporary general-purpose reactive MLFFs across a range of non-equilibrium tasks, including reaction-pathway construction, transition-state localization, and Hessian-based vibrational analysis. These results not only highlight their current applicability, but also provide insights into future methodological development. From a dataset preparation perspective, our findings underscore the need for larger and more diverse biochemical reaction datasets. From a model-design perspective, while existing architectures already support simulations of systems containing thousands of atoms, improved treatment of long-range interactions and higher-order effects will be critical for further advances.
To the best of our knowledge, this work represents the first systematic application of MLFFs to full enzymatic systems, producing chemically meaningful energy profiles. To validate MAPLE as a general-purpose framework, we present case studies spanning the well-established Kemp eliminase, the newly discovered Diels–Alderases, and the highly complex DNA photolyase. In particular, the MaDA mutation screening workflow demonstrates MAPLE's utility for automated enzyme design through high-throughput variant evaluation. Together, these results establish MLFFs as a powerful and versatile tool for rapid and accurate modeling of catalytic processes, with broad potential for future applications.
A key advantage of MAPLE lies in its unified treatment of the entire system without predefined QM/MM partitioning. By modeling all atoms on the same MLFF potential energy surface, MAPLE avoids artificial boundary effects and enables a physically consistent description of catalytic environments, including second-shell residues and solvent interactions. This capability is particularly valuable for studying mutational effects, stereoselectivity, and long-range electrostatic contributions, where local approximations may obscure mechanistic insights.
We envision MAPLE as a long-term, actively maintained platform. Beyond the enzymatic catalysis case studies presented here, MAPLE's functionalities extend to general molecular modeling tasks, including geometry optimization, transition state search, IRC, frequency analysis, PES scanning, and molecular dynamics (Table S1). Future development directions include extended molecular dynamics capabilities, molecular docking workflows, and enhanced sampling methods for free energy calculations. On the algorithmic side, MAPLE currently implements batched parallel L-BFGS optimization and an automatic-differentiation-based Dimer method; further parallelized algorithms, including batched NEB and batched IRC, are under active development. We also plan to incorporate established generative models to enable rapid and physically meaningful predictions. Overall, MAPLE aims to serve a broad research community, from MLFF and algorithm developers to applied researchers in biocatalysis and materials science, bridging methodological innovation with practical scientific applications.
4 Methods
4.1 Batched L-BFGS optimization with dynamic batch shrinking
Geometry optimizations are performed using a GPU-based batched L-BFGS optimizer, where multiple molecular structures are optimized simultaneously. Each structure maintains an independent L-BFGS history, while all operations are executed in parallel using batched tensor algebra.All molecular coordinates are stored in a fixed-shape tensor of size (B, nmax), where nmax = 3Nmax corresponds to the largest system in the batch. Smaller systems are zero-padded and excluded from all dot products and updates using a binary mask, ensuring that padding does not affect optimization.
Independent quasi-Newton histories are maintained for each structure and stored as batched tensors. The L-BFGS update and two-loop recursion follow the standard formulation55 and are evaluated in parallel across the batch.
Structures that satisfy convergence criteria are removed from the batch at each iteration, and all associated history tensors are sliced accordingly. This dynamic batch shrinking avoids unnecessary computation on converged systems.
Atomic displacements are clipped to a maximum step of 0.2 Å per iteration. The history length is set to m = 5, the initial inverse Hessian scaling parameter is γ0 = 70Eh Å−2, and all calculations are performed in double precision.
4.2 Dual-shift partitioned rational function optimization
Transition-state searches are performed using partitioned rational function optimization (P-RFO),56,57 where the optimization space is decomposed into an uphill subspace associated with the negative curvature mode and a downhill subspace spanning all remaining modes.We introduce a dual-shift P-RFO (DS-P-RFO) scheme in which independent shift parameters are applied to the uphill and downhill subspaces. The total step is decomposed as
| s = s− + s+, | (1) |
| |s−|2 ≤ R−2, |s+|2 ≤ R+2, | (2) |
The trust-region budget is adaptively partitioned according to the unconstrained Newton steps in each subspace,
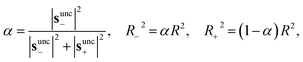 | (3) |
Independent shift parameters µ− and µ+ are then determined for the uphill and downhill subspaces to satisfy their respective constraints,
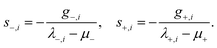 | (4) |
By decoupling the trust-region control of the uphill and downhill modes, DS-P-RFO avoids the iterative micro-cycles required in conventional RS-P-RFO while preserving numerical stability and convergence behavior.
4.3 Dimer method with automatic-differentiation Hessian–vector products
Saddle-point searches without reliable initial guesses are performed using the dimer method.58 Instead of estimating Hessian–vector products (HVPs) by finite differences, we compute them analytically using automatic differentiation.Specifically, for a given dimer orientation vector n, the Hessian–vector product is evaluated as
| Hn = ∇R(∇RE(R) × n), | (5) |
The analytically evaluated HVP is used in both the rotation step, to align the dimer with the lowest-curvature mode, and the translation step, to follow the minimum mode toward a first-order saddle point. All remaining aspects of the dimer algorithm follow the standard formulation.58
4.4 Enzymatic Catalysis Simulations
Enzyme structures were obtained from the Protein Data Bank (PDB) and preprocessed using AMBERTools59 to repair missing residues and heavy atoms, followed by adding hydrogen atoms. The corrected structures were then converted to XYZ format using Open Babel. Global geometry optimizations were performed in MAPLE using the L-BFGS algorithm with superloose convergence criteria to relax the full enzyme systems.
to repair missing residues and heavy atoms, followed by adding hydrogen atoms. The corrected structures were then converted to XYZ format using Open Babel. Global geometry optimizations were performed in MAPLE using the L-BFGS algorithm with superloose convergence criteria to relax the full enzyme systems.
Based on the optimized structures, residues within 8 Å of the reaction center were defined as flexible, while all remaining residues were kept fixed to reduce the computational cost. Constrained optimizations were subsequently carried out using MAPLE's default L-BFGS settings. The optimized reactant and product structures were then used to conduct climbing-image nudged elastic band (CI-NEB) calculations. After iterative refinement, converged minimum-energy pathways were obtained, and the highest-energy image (HEI) along each pathway was taken as the transition-state structure.
Author contributions
Xujian Wang: conceptualization, methodology, software, investigation, writing original draft. Zeyu Sun: investigation. Carlo Asam, Yilu Zhang and Ruzhan Zhu: software. Junmei Wang and Wan-Lu Li: review, editing, supervision.Conflicts of interest
There are no conflicts to declare.Data availability
The MAPLE program and source code developed in this work are openly available at GitHub (https://github.com/ClickFF/MAPLE). Documentation and tutorials are available at https://www.maplechem.org/. All data supporting this study, including reactive potential benchmarks, enzymatic simulation data, and related computational results, are openly available at Zenodo: https://doi.org/10.5281/zenodo.18135653. MAPLE is also accessible through the Qbics-MolStar platform at https://molstar.szbl.ac.cn/viewer/.Supplementary information (SI): computational methods, algorithmic implementations, benchmark datasets, additional figures and tables, protein and DNA sequence information, reaction pathway analyses, and supporting results for all simulations described in this study. See DOI: https://doi.org/10.1039/d6sc01279e.
Acknowledgements
This work was supported by the following funds from the National Institutes of Health (NIH) and the National Science Foundation (NSF): NIH R01GM147673, R01GM149705 and NSF 1955260. Authors also acknowledge Rongfa Bai and Prof. Jun Zhang for their assistance in integrating MAPLE into the Qbics-MolStar platform. W.-L. Li acknowledges support by the ACS Petroleum Research Fund under Doctoral New Investigator Grant 69037-DNI10, the National Science Foundation through the DMREF program under award No. 2522541, and the Hellman Fellowship, University of California San Diego. X. W. acknowledges Dean's fellowship from School of Medicine, University of Pittsburgh. Computational resources were partly provided by the San Diego Supercomputer Center through ACCESS allocations of CHM250110 and CHM250109.References
- C. Tian, K. Kasavajhala, K. A. A. Belfon, L. Raguette, H. Huang, A. N. Migues, J. Bickel, Y. Wang, J. Pincay, Q. Wu and C. Simmerling, ff19SB: Amino-Acid-Specific Protein Backbone Parameters Trained against Quantum Mechanics Energy Surfaces in Solution, J. Chem. Theory Comput., 2020, 16, 528–552 CrossRef PubMed.
- J. Wang, R. M. Wolf, J. W. Caldwell, P. A. Kollman and D. A. Case, Development and Testing of a General Amber Force Field, J. Comput. Chem., 2004, 25, 1157–1174 CrossRef CAS PubMed.
- X. Wang, H. Liu, Y. Li, J. Li and W.-L. Li, TinkerModeller: An Efficient Tool for Building Biological Systems in Tinker Simulations, J. Chem. Theory Comput., 2025, 21, 2712–2722 CrossRef CAS PubMed.
- X. Wang, X. Wu, B. R. Brooks and J. Wang, Accurate Free Energy Calculation via Multiscale Simulations Driven by Hybrid Machine Learning and Molecular Mechanics Potentials, J. Chem. Theory Comput., 2025, 21, 6979–6987 CrossRef CAS PubMed.
- X. Wang, H. Liu, J. Wang, L. Chang, J. Cai, Z. Wei, J. Pan, X. Gu, W.-L. Li and J. Li, Enzyme Tunnel Dynamics and Catalytic Mechanism of Norcoclaurine Synthase: Insights from a Combined LiGaMD and DFT Study, J. Phys. Chem. B, 2024, 128, 9385–9395 CrossRef CAS PubMed.
- D. M. Anstine, R. Zubatyuk and O. Isayev, AIMNet2: A Neural Network Potential to Meet Your Neutral, Charged, Organic, and Elemental-Organic Needs, Chem. Sci., 2025, 16, 10228–10244 RSC.
- B. M. Wood, et al., UMA: A Family of Universal Models for Atoms, arXiv, 2025, preprint, arXiv:2506.23971, DOI:10.48550/arXiv.2506.23971.
- S. Zhang, M. Z. Makoś, R. B. Jadrich, E. Kraka, K. Barros, B. T. Nebgen, S. Tretiak, O. Isayev, N. Lubbers, R. A. Messerly and J. S. Smith, Exploring the Frontiers of Condensed-Phase Chemistry with a General Reactive Machine Learning Potential, Nat. Chem., 2024, 16, 727–734 CrossRef CAS PubMed.
- D. M. Anstine, Q. Zhao, R. Zubatiuk, S. Zhang, V. Singla, F. Nikitin, B. M. Savoie and O. Isayev, AIMNet2-rxn: A Machine Learned Potential for Generalized Reaction Modeling on a Millions-of-Pathways Scale, arXiv, 2025, preprint, arXiv:2505.13963, DOI:10.48550/arXiv.2505.13963.
- D. Anstine, R. Zubatyuk, L. Gallegos, R. Paton, O. Wiest, B. Nebgen, T. Jones, G. Gomes, S. Tretiak, O. Isayev, Transferable Machine Learning Interatomic Potential for Pd-Catalyzed Cross-Coupling Reactions, ChemRxiv, 2025, DOI:10.26434/chemrxiv-2025-0k4kf.
- R. Zubatyuk, M. Biczysko, K. Ranasinghe, N. W. Moriarty, H. Gokcan, H. Kruse, B. K. Poon, P. D. Adams, M. P. Waller, A. E. Roitberg, O. Isayev and P. V. Afonine, AQuaRef: Machine Learning Accelerated Quantum Refinement of Protein Structures, Nat. Commun., 2025, 16, 9224 CrossRef CAS PubMed.
- D. S. Levine, et al., The Open Molecules 2025 (OMol25) Dataset, Evaluations, and Models, arXiv, 2025, preprint, arXiv:2505.08762, DOI:10.48550/arXiv.2505.08762.
- E. Hermes, K. Sargsyan, H. N. Najm and J. Zádor, Accelerated Saddle Point Refinement through Full Exploitation of Partial Hessian Diagonalization, J. Chem. Theory Comput., 2019, 15, 6536–6549 CrossRef CAS PubMed.
- P. O. Dral, et al., MLatom 3: A Platform for Machine Learning-Enhanced Computational Chemistry Simulations and Workflows, J. Chem. Theory Comput., 2024, 20, 1193–1213 CrossRef CAS PubMed.
- A. Hjorth Larsen, et al., The Atomic Simulation Environment—A Python Library for Working with Atoms, J. Phys.: Condens. Matter, 2017, 29, 273002 CrossRef PubMed.
- J. S. Smith, B. Nebgen, N. Lubbers, O. Isayev and A. E. Roitberg, Less is More: Sampling Chemical Space with Active Learning, J. Chem. Phys., 2018, 148, 241733 CrossRef PubMed.
- J. S. Smith, B. T. Nebgen, R. Zubatyuk, N. Lubbers, C. Devereux, K. Barros, S. Tretiak, O. Isayev and A. E. Roitberg, Approaching Coupled Cluster Accuracy with a General-Purpose Neural Network Potential through Transfer Learning, Nat. Commun., 2019, 10, 2903 CrossRef PubMed.
- C. Devereux, J. S. Smith, K. K. Huddleston, K. Barros, R. Zubatyuk, O. Isayev and A. E. Roitberg, Extending the Applicability of the ANI Deep Learning Molecular Potential to Sulfur and Halogens, J. Chem. Theory Comput., 2020, 16, 4192–4202 CrossRef CAS PubMed.
- B. Kalita, R. Zubatyuk, D. M. Anstine, M. Bergeler, V. Settels, C. Stork, S. Spicher and O. Isayev, AIMNet2-NSE: A Transferable Reactive Neural Network Potential for Open-Shell Chemistry, Angew. Chem., Int. Ed., 2026, 65, e202516763 CrossRef PubMed.
- D. P. Kovács, J. H. Moore, N. J. Browning, I. Batatia, J. T. Horton, Y. Pu, V. Kapil, W. C. Witt, I.-B. Magdău, D. J. Cole and G. Csányi, MACE-OFF: Short-Range Transferable Machine Learning Force Fields for Organic Molecules, J. Am. Chem. Soc., 2025, 147, 17598–17611 CrossRef PubMed.
- E. L. Mann, C. C. Wagen, J. E. Vandezande, A. M. Wagen and S. C. Schneider, Egret-1: Pretrained Neural Network Potentials for Efficient and Accurate Bioorganic Simulation, arXiv, 2025, preprint, arXiv:2502.02765, DOI:10.48550/arXiv.2502.02765.
- I. Batatia, W. J. Baldwin, D. Kuryla, J. Hart, E. Kasoar, A. M. Elena, H. Moore, M. J. Gawkowski, B. X. Shi, V. Kapil, P. Kourtis, I.-B. Magdău, G. Csányi, MACE-POLAR-1: A Polarisable Electrostatic Foundation Model for Molecular Chemistry, arXiv, 2026, preprint, arXiv:2602.19411, DOI:10.48550/arXiv.2602.19411.
- E. Caldeweyher, J.-M. Mewes, S. Ehlert and S. Grimme, Extension and evaluation of the D4 London-dispersion model for periodic systems, Phys. Chem. Chem. Phys., 2020, 22, 8499–8512 RSC.
- M. Friede, C. Hölzer, S. Ehlert and S. Grimme, dxtb—An efficient and fully differentiable framework for extended tight-binding, J. Chem. Phys., 2024, 161, 062501 CrossRef CAS PubMed.
- H. Nguyen, D. R. Roe and C. Simmerling, Improved Generalized Born Solvent Model Parameters for Protein Simulations, J. Chem. Theory Comput., 2013, 9, 2020–2034 CrossRef CAS PubMed.
- M. S. Lee, J. Salsbury, R. Freddie, I. Brooks and L. Charles, Novel Generalized Born Methods, J. Chem. Phys., 2002, 116, 10606–10614 CrossRef CAS.
- F. Neese, Software Update: The ORCA Program System—Version 6.0, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2025, 15, e70019 Search PubMed.
- M. J. Frisch et al., Gaussian 16, Revision C.01, Gaussian, Inc., Wallingford CT, 2016 Search PubMed.
- M. Schreiner, A. Bhowmik, T. Vegge, J. Busk and O. Winther, Transition1x—a dataset for building generalizable reactive machine learning potentials, Sci. Data, 2022, 9, 779 CrossRef PubMed.
- N. Mardirossian and M. Head-Gordon, ωB97M-V: A combinatorially optimized, range-separated hybrid, meta-GGA density functional with VV10 nonlocal correlation, J. Chem. Phys., 2016, 144, 214110 CrossRef PubMed.
- F. Weigend and R. Ahlrichs, Balanced basis sets of split valence, triple zeta valence and quadruple zeta valence quality for H to Rn: Design and assessment of accuracy, Phys. Chem. Chem. Phys., 2005, 7, 3297–3305 RSC.
- G. Henkelman, B. P. Uberuaga and H. Jónsson, A climbing image nudged elastic band method for finding saddle points and minimum energy paths, J. Chem. Phys., 2000, 113, 9901–9904 CrossRef CAS.
- A. Paszke, et al., PyTorch: An Imperative Style, High-Performance Deep Learning Library, Adv. Neural Inf. Process. Syst., 2019, 8024–8035 Search PubMed.
- D. Röthlisberger, O. Khersonsky, A. M. Wollacott, L. Jiang, J. DeChancie, J. Betker, J. L. Gallaher, E. A. Althoff, A. Zanghellini, O. Dym, S. Albeck, K. N. Houk, D. S. Tawfik and D. Baker, Kemp Elimination Catalysts by Computational Enzyme Design, Nature, 2008, 453, 190–195 CrossRef PubMed.
- A. N. Alexandrova, D. Röthlisberger, D. Baker and W. L. Jorgensen, Catalytic Mechanism and Performance of Computationally Designed Enzymes for Kemp Elimination, J. Am. Chem. Soc., 2008, 130, 15907–15915 CrossRef CAS PubMed.
- M. P. Frushicheva, J. Cao, Z. T. Chu and A. Warshel, Exploring Challenges in Rational Enzyme Design by Simulating the Catalysis in Artificial Kemp Eliminase, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 16869–16874 CrossRef CAS PubMed.
- A. Labas, E. Szabó, L. Mones and M. Fuxreiter, Optimization of Reorganization Energy Drives Evolution of the Designed Kemp Eliminase KE07, Biochim. Biophys. Acta, 2013, 1834, 908–917 CrossRef CAS PubMed.
- N.-S. Hong, et al., The Evolution of Multiple Active Site Configurations in a Designed Enzyme, Nat. Commun., 2018, 9, 3900 CrossRef PubMed.
- O. Khersonsky, D. Röthlisberger, O. Dym, S. Albeck, C. J. Jackson, D. Baker and D. S. Tawfik, Evolutionary Optimization of Computationally Designed Enzymes: Kemp Eliminases of the KE07 Series, J. Mol. Biol., 2010, 396, 1025–1042 CrossRef CAS PubMed.
- A. D. Becke, Density-Functional Thermochemistry. III. The Role of Exact Exchange, J. Chem. Phys., 1993, 98, 5648–5652 CrossRef CAS.
- P. C. Hariharan and J. A. Pople, The Influence of Polarization Functions on Molecular Orbital Hydrogenation Energies, Theor. Chim. Acta, 1973, 28, 213–222 CrossRef CAS.
- W. L. Jorgensen, D. S. Maxwell and J. Tirado-Rives, Development and Testing of the OPLS All-Atom Force Field on Conformational Energetics and Properties of Organic Liquids, J. Am. Chem. Soc., 1996, 118, 11225–11236 CrossRef CAS.
- L. Gao, et al., FAD-dependent enzyme-catalysed intermolecular [4+2] cycloaddition in natural product biosynthesis, Nat. Chem., 2020, 12, 620–628 CrossRef CAS PubMed.
- L. Gao, Y. Zou, X. Liu, J. Yang, X. Du, J. Wang, X. Yu, J. Fan, M. Jiang, Y. Li, K. N. Houk and X. Lei, Enzymatic control of endo- and exo-stereoselective Diels–Alder reactions with broad substrate scope, Nat. Catal., 2021, 4, 1059–1069 CrossRef CAS.
- Q. Ding, N. Guo, L. Gao, M. McKee, D. Wu, J. Yang, J. Fan, J.-K. Weng and X. Lei, The evolutionary origin of naturally occurring intermolecular Diels-Alderases from Morus alba, Nat. Commun., 2024, 15, 2492 CrossRef CAS PubMed.
- L. Gao, Q. Ding and X. Lei, Hunting for the Intermolecular Diels–Alderase, Acc. Chem. Res., 2024, 57, 2166–2183 CrossRef CAS PubMed.
- N. Guo, J. Gu, Q. Zhou, F. Liu, H. Dong, Q. Ding, Q. Wang, D. Wu, J. Yang, J. Fan, L. Gao, K. N. Houk and X. Lei, Aspartic acid residues in BBE-like enzymes from Morus alba promote a function shift from oxidative cyclization to dehydrogenation, Proc. Natl. Acad. Sci. U. S. A., 2025, 122, e2504346122 CrossRef CAS PubMed.
- X. Wang, H. Tang, X. Wu, B. Brooks, J. Wang, W.-L. Li, Redefining Computational Enzymology with Multiscale Machine Learning/Molecular Mechanics Metadynamics: Deciphering Catalytic Mechanism and Stereoselectivity in Diels–Alderases, ChemRxiv, 2025, DOI:10.26434/chemrxiv-2025-x2k6j.
- J.-S. Taylor, DNA, Sunlight and Skin Cancer, Pure Appl. Chem., 1995, 67, 183–190 CrossRef CAS.
- H. Kamiya, S. Iwai and H. Kasai, The (6-4) Photoproduct of Thymine–Thymine Induces Targeted Substitution Mutations in Mammalian Cells, Nucleic Acids Res., 1998, 26, 2611–2617 CrossRef CAS PubMed.
- R. M. Herriott, Formation of Heterozygotes by Annealing a Mixture of Transforming DNAs, Proc. Natl. Acad. Sci. U. S. A., 1961, 47, 146–153 CrossRef CAS PubMed.
- S. Faraji, G. Groenhof and A. Dreuw, Combined QM/MM Investigation on the Light-Driven Electron-Induced Repair of the (6–4) Thymine Dimer Catalyzed by DNA Photolyase, J. Phys. Chem. B, 2013, 117, 10071–10079 CrossRef CAS PubMed.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, A Consistent and Accurate Ab Initio Parametrization of Density Functional Dispersion Correction (DFT-D) for the 94 Elements H–Pu, J. Chem. Phys., 2010, 132, 154104 CrossRef PubMed.
- X. Wang, J. Wang and W.-L. Li, Machine learning/molecular mechanics enzymology for the next generation of computational enzymatic catalysis, Chem, 2026, 6, 101658 Search PubMed.
- D. C. Liu and J. Nocedal, On the limited memory BFGS method for large scale optimization, Math. Program., 1989, 45, 503–528 CrossRef.
- J. Baker, An algorithm for the location of transition states, J. Comput. Chem., 1986, 7, 385–395 CrossRef CAS.
- E. Besalú and J. M. Bofill, On the automatic restricted-step rational-function-optimization method, Theor. Chem. Acc., 1998, 100, 265–274 Search PubMed.
- G. Henkelman and H. Jónsson, A dimer method for finding saddle points on high dimensional potential surfaces using only first derivatives, J. Chem. Phys., 1999, 111, 7010–7022 CrossRef CAS.
- D. A. Case, et al., AmberTools, J. Chem. Inf. Model., 2023, 63, 6183–6191 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |