Open Access Article
Open Access ArticlePeptide-directed folding of the elusive RNA i-motif
Lachlan B.
Cox
 ab,
Pall
Thordarson
ab,
Pall
Thordarson
 *ab and
Felix J.
Rizzuto
*ab and
Felix J.
Rizzuto
 *ab
*ab
aSchool of Chemistry, University of New South Wales, Sydney, NSW 2052, Australia. E-mail: f.rizzuto@unsw.edu.au; p.thordarson@unsw.edu.au
bUNSW RNA Institute, University of New South Wales, Sydney, NSW 2052, Australia
First published on 12th March 2026
Abstract
Folded RNA structures are increasingly being recognised as key regulators in biological processes, yet the RNA i-motif remains poorly characterised due to its low stability and lack of selective molecular probes. Here, we describe the first ligand – a short peptide – that binds the elusive RNA i-motif. Our minimalist peptide RGGFGGRGG is derived from the intrinsically disordered region of the protein nucleolin and binds to folded RNA over DNA with >5-fold selectivity. The binding of two peptide molecules folds the RNA i-motif at a higher pH than under native conditions. This folded, peptide-bound structure can still bind other guests, such as the intercalator thiazole orange, displaying heteroallosteric properties. Our peptide binding is driven by more than simple electrostatic attraction, exploiting the subtle differences in steric complementarity and hydration of the compact RNA structures relative to DNA congeners and unfolded strands. Our findings underline the potential of minimalistic peptide scaffolds as selective binders for non-canonical RNA structures, allowing for the probing and modulation of RNA topologies.
Non-coding RNA secondary structures are increasingly being recognised for their regulatory roles in transcription, translation, and epigenetic regulation.1 While canonical Watson–Crick–Franklin base pairing between cytosine/guanine and uracil/adenine forms the well-characterised RNA A-helix, the significance of non-canonical RNA secondary structures is less understood.2 Two of the most well-known examples are the guanine-rich G-quadruplex and its cytosine-rich counterpart, the intercalated-motif (i-motif).3
The i-motif consists of two parallel cytosine–cytosine duplexes, intercalating in an antiparallel orientation to form a pseudo-four-stranded structure (Fig. 1B).4 The i-motif structure only forms when the cytosines become hemi-protonated on the N3 position, allowing for a third hydrogen bond between cytosine pairs (Fig. 1C).5 Depending on the sequence and length of the nucleic acid chain, DNA i-motifs typically fold anywhere between pH 4.5 and 6.5; however, the RNA i-motif structure is significantly less stable, occurring between pH 4.0 and 4.5.6
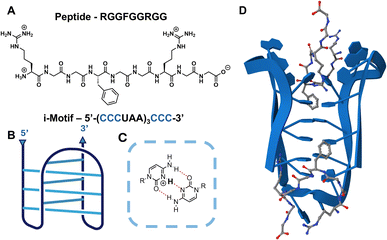 | ||
| Fig. 1 Peptide and oligonucleotide sequences used in this study. (A) Sequence of the RNA-binding peptide RGGFGGRGG. (B) Folded structure of the hTeloC i-motif structure (blue). (C) Hemi-protonated cytosines will associate via self-complementary pairing. (D) Computational interactions predicted by AlphaFold 3.17 These predictions are based on the hTeloC DNA crystal structure.12 | ||
While there has been some debate about the existence of i-motif structures in vivo,7 intact i-motif DNA structures have been found in the nuclei of human cells8 and are prevalent in cytosine-rich regions of the genome, such as the telomere and gene-promoter regions.9 High macromolecular crowding in these regions allows for the structure to form at a higher pH than typically required,10 allowing for i-motifs to be present in only slightly acidic conditions (pH ∼ 6), such as exist in tumour microenvironments, for example.11 The i-motif has also been recently linked to disease pathogenesis; point mutations in cytosine-rich sequences in the insulin-linked polymorphic region cause a loss of the DNA i-motif structure, indicating a diabetes disease state.12
An area in the human genome with i-motif structures of particular interest is the telomere. The capping region of the human telomere contains tandem repeats of 5′-(TTAGGG)n/(CCCTAA)n-3′ DNA.13 While the guanine-rich sense sequence forms G-quadruplex structures in vivo, it has also been shown that the antisense cytosine-rich 5′-(CCCTAA)n-3′ DNA strand can form offset i-motif structures in vitro.14 This cytosine-rich repeat sequence is often referred to as hTeloC.11 It is thought that the non-canonical structures formed by these tandem repeats play crucial roles in telomerase inhibition.15
Studies show that RNA i-motifs are significantly less thermodynamically stable compared to DNA congeners.6 There are currently no known ligands that are selective for the RNA i-motif over unfolded RNA or DNA i-motifs.6 We recently reported that a short RNA-binding peptide – RGGFGGRGG (Fig. 1A) – binds and stabilises RNA G-quadruplexes.16 We hypothesised that this peptide would also be capable of binding to and stabilising protonated RNA structures.
Here, we explore the interactions of our RGGFGGRGG peptide with RNA and DNA hTeloC i-motif structures with the sequence 5′-CCC UAA CCC UAA CCC UAA CCC-3′. We show that our minimal peptide discriminates for RNA i-motif structures over the DNA congener, retains selectivity for folded i-motifs over single-stranded cytosine-rich RNA, and increases the pH at which the native RNA i-motif folds.
Our RNA-binding peptide RGGFGGRGG (Fig. 1A) was derived from the intrinsically disordered region of the nucleolin protein, a multifunctional transcription regulator that has been shown in vivo to pathologically interact with cytosine/guanine-rich sequences in a host of neurodegenerative diseases.18 The peptide was synthesised via a literature procedure.16
Selective interactions between RNA and DNA by short peptides are often challenging to design, due to the similarities between RNA and DNA nucleobases.19 Furthermore, the i-motif structure is remarkably similar between RNA and DNA species.5,6 We observed, however, that the RGGFGGRGG peptide selectively interacted with RNA over DNA i-motif structures, and discriminated between folded structures over their single-stranded derivatives.
To determine association constants between the RGGFGGRGG peptide and the RNA i-motif, we employed Nuclear Magnetic Resonance (NMR) titrations,20 which were performed in duplicate and analysed using the online fitting program https://supramolecular.org (SI Section 1.9).21 Multiple binding models were compared for folded (pH 4.0) and unfolded (pH 7.0) forms of the hTeloC RNA (5′-(CCCUAA)3CCC-3′) and DNA (5′-(CCCTAA)3CCC-3′) sequences (SI Section 5). We compared fitting models for the titration of RGGFGGRGG with hTeloC RNA across the tested pH gradient. The fitting model that best described the interaction was a 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 peptide
1 peptide![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) RNA stoichiometry with a statistical model, suggesting no inherent cooperativity between the two binding events.22 Full analysis of the binding models is available in SI Section 1.12 and SI Table 4.
RNA stoichiometry with a statistical model, suggesting no inherent cooperativity between the two binding events.22 Full analysis of the binding models is available in SI Section 1.12 and SI Table 4.
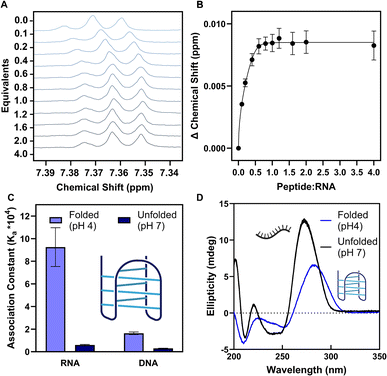
At pH 4.0, the cytosines in the hTeloC RNA sequence – 5′-(CCCUAA)3CCC-3′ – become hemi-protonated, allowing for the formation of the i-motif structure. 1H NMR titrations with the RGGFGGRGG peptide at pH 4.0 showed a moderate level of interaction (Ka = ∼93![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ± 17
000 ± 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 M−1) (SI Section 4). Conversely, titrations between our peptide and the unfolded hTeloC structure at pH 7.0 showed a >15× decrease in association constant (Ka = ∼6100 ± 400 M−1, Fig. 2A), suggesting our peptide interacts more strongly with the folded i-motif structure.
000 M−1) (SI Section 4). Conversely, titrations between our peptide and the unfolded hTeloC structure at pH 7.0 showed a >15× decrease in association constant (Ka = ∼6100 ± 400 M−1, Fig. 2A), suggesting our peptide interacts more strongly with the folded i-motif structure.
Our peptide further discriminates for RNA i-motif structures over DNA variants. When titrated with the DNA i-motif hTeloC variant 5′-(CCCTAA)3CCC-3′ at pH 4.0, the association constant for the RGGFGGRGG peptide decreased by >5× (Ka = 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ± 1000 M−1), as compared to the RNA variant (Fig. 2A). The DNA i-motif shows a similar trend to the RNA, with a >5× preference for folded i-motif DNA over unfolded ssDNA at pH 7.0 (Ka = 3000 ± 1000 M−1).
000 ± 1000 M−1), as compared to the RNA variant (Fig. 2A). The DNA i-motif shows a similar trend to the RNA, with a >5× preference for folded i-motif DNA over unfolded ssDNA at pH 7.0 (Ka = 3000 ± 1000 M−1).
We employed Circular Dichroism (CD) spectroscopy to confirm the i-motif structure at the tested pH values. The intense positive band at 288 nm and negative band at 260 nm at pH 4.0 is indicative of the i-motif structure, while the shift of the positive band towards 272 nm at pH 7.0 describes a single-stranded cytosine-rich sequence with a rigid A-form backbone (Fig. 2B).23,24 Additions of the peptide did not alter the CD spectra at either pH 4.0 or 7.0, suggesting that the peptide does not disrupt the i-motif structure (SI Fig. 3).
To further probe the selectivity of the peptide for the folded hTeloC RNA i-motif structure, we performed an additional series of NMR titrations across a range of pH points. This approach gives precise control over the extent of cytosine hemi-protonation, allowing for modulation of the proportion of folded i-motif present, as compared to ssRNA. Across the tested pH range of pH 4.0–9.0, the RGGFGGRGG peptide retains a consistent net charge of 2+, due to the calculated isoelectric point of 12.1 (SI Fig. 8).
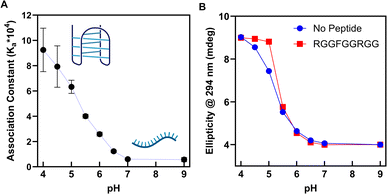
A gradual decrease in association constant was observed when increasing the pH from a fully folded, hemi-protonated state (pH 4.0) towards an eventual plateau at a single-stranded state (pH 7.0) (Fig. 3A). This relationship suggests a model where hemi-protonation induces a conformational shift in RNA structure towards an i-motif, which exposes a favourable binding pocket for the RGGFGGRGG peptide. Raw NMR titration data is available in SI Fig. 12–31.
When folded, the hemi-protonated i-motif structure has less net negative charge (overall 15−) than single stranded RNA (overall 21−) owing to cytosine protonation; the structure also becomes more compact.25 It is therefore quite unusual that a positively charged peptide would have an increased affinity towards the folded RNA structure, given that the overall negative charge of the i-motif structure lessens upon protonation and folding. This suggests that the driving force for the association of RGGFGGRGG towards i-motif RNA is governed by more than electrostatics, with steric fit likely being a driving force for association. Additionally, the hydrophobic core of the folded i-motif structure has an increased local charge density due to the aggregation of cytosines.26 Water networks are formed between cytosine base pairs to screen local charge.27 Displacement of this high-energy water may rationalise the increased association of our peptide with compact folded structures over hydrophilic unfolded strands of the same sequence.28
To correlate the association constant with the fold state of the i-motif structure, we conducted CD spectroscopy across a pH gradient of 4.0–9.0, in the absence and presence of the RGGFGGRGG peptide. By monitoring the CD ellipticity at 294 nm, we can quantify the transition from i-motif at low pH to ssRNA at neutral pH. We observed that with increasing pH, the proportion of folded i-motif strongly correlated with the association constants derived from the NMR titrations (Fig. 3B), suggesting once more that the RGGFGGRGG peptide is reliant on the i-motif structure for a strong interaction.
Upon the addition of 16 equivalents of our peptide, the stability of the RNA i-motif structure at pH 5.0 increased. Direct electrostatic interactions between peptide guanidinium and RNA phosphates may physically template the i-motif structure through kinetic pre-organisation29 (acting as a ‘molecular glue’).16 However, CD melting curves showed no change in the melting temperature of the i-motif structure at pH 4.0 or 5.0 upon additions of the RGGFGGRGG peptide (SI Fig. 4 and 7), suggesting that the mechanism of stabilisation is not purely thermodynamic in nature. Another option is that the RGGFGGRGG peptide alters the chemical environment of the compact i-motif core through multivalent weak interactions, increasing the pKa for hemi-protonation of cytosine residues and resulting in the observed stabilisation. It is known that macromolecular crowding often stabilises biomolecular interactions and nucleic acid structure through microenvironment regulation;30 this was integral to finding the DNA i-motif in the cell.8 Both methods of stabilisation – physical templation or altering of the RNA microenvironment – suggest that the RGGFGGRGG peptide is capable of increasing the transitional pH and stabilising the RNA i-motif structure at a higher pH range, as compared to untemplated native folding (Fig. 3B). The RGGFGGRGG peptide had no effect on the structure of hTeloC i-motif DNA (SI Fig. 33), making our peptide the first example of a ligand selectively stabilising the RNA i-motif structure.
To evaluate the specificity of this stabilising interaction, we synthesised a control peptide RGGRGGRGG, where the central amino acid residue phenylalanine was substituted for an additional arginine. This mutation disrupts pi-stacking with aromatic RNA bases, while strengthening electrostatic interactions. No stabilisation of the hTeloC i-motif structure was observed when titrated with RGGRGGRGG, highlighting the significance of the native nucleolin-derived sequence RGGFGGRGG in binding to the RNA i-motif (SI Fig. 10).
CD titrations of the RGGFGGRGG peptide into hTeloC RNA at pH 5 showed stepwise stabilisation of the i-motif structure. Experimental data strongly matches a simulated binding isotherm performed using https://supramolecular.org Bindsim (SI Fig. 38).
To further investigate the effect of the RGGFGGRGG peptide on the hTeloC RNA i-motif structure, we conducted a Thiazole Orange (TO) fluorescence titration in the presence and absence of our peptide. TO exhibits low fluorescence in solution, but upon intercalation with i-motif structures rotational restriction forces a planar conformation that exhibits enhanced fluorescence, as compared to single-stranded nucleic acids.31,32
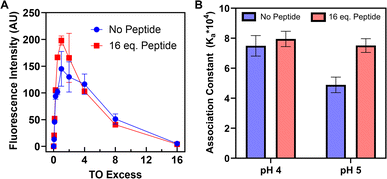
While TO binds DNA i-motif structures,21 binding to RNA i-motif structures has yet to be investigated. To assess the validity of TO as an hTeloC RNA probe, fluorescence assays were performed with 20 µM hTeloC i-motif RNA, either alone or in the presence of the RGGFGGRGG peptide (320 µM), titrated with TO (0–16 equivalents) and excited at 512 nm (Fig. 4A). At pH 4.0, where our RNA adopts a fully folded i-motif structure, TO interacted with an association constant (Ka = 75![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ± 7000 M−1) slightly lower but similar to that of literature values for DNA (270
000 ± 7000 M−1) slightly lower but similar to that of literature values for DNA (270![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ± 5000 M−1),32 as fit to a 2
000 ± 5000 M−1),32 as fit to a 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 TO
1 TO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) RNA non-cooperative titration model.21
RNA non-cooperative titration model.21
In the presence of the RGGFGGRGG peptide, TO binding remained statistically unchanged (Ka = 79![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ± 5000 M−1), suggesting that our peptide is not acting as a competitive inhibitor, nor is it altering the i-motif structure. At pH 5.0, however, where the i-motif is only partially folded in the absence of our peptide, TO interaction decreased (Ka = 49
000 ± 5000 M−1), suggesting that our peptide is not acting as a competitive inhibitor, nor is it altering the i-motif structure. At pH 5.0, however, where the i-motif is only partially folded in the absence of our peptide, TO interaction decreased (Ka = 49![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ± 5000 M−1), consistent with a lessening of the available i-motif structure to bind. Upon addition of the peptide at pH 5.0, the association of TO to RNA increased back to its baseline level (Ka = 75
000 ± 5000 M−1), consistent with a lessening of the available i-motif structure to bind. Upon addition of the peptide at pH 5.0, the association of TO to RNA increased back to its baseline level (Ka = 75![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ± 5000 M−1). These results are consistent with peptide-mediated stabilisation of the i-motif structure at pH 5.0, while it remains partially unfolded in the absence of peptide (Fig. 4B). Thiazole orange fluorescence was observed to be low at pH 7 in the absence of folded i-motif RNA (SI Fig. 33).
000 ± 5000 M−1). These results are consistent with peptide-mediated stabilisation of the i-motif structure at pH 5.0, while it remains partially unfolded in the absence of peptide (Fig. 4B). Thiazole orange fluorescence was observed to be low at pH 7 in the absence of folded i-motif RNA (SI Fig. 33).
This increase in binding affinity of TO at pH 5.0 suggests that our peptide is having a positive heteroallosteric33 effect on the i-motif structure, at pH ranges outside where it is usually stable. We hypothesise that our peptide aids the folding of the RNA i-motif at a higher pH than occurs for native folding through a templation mechanism, increasing the mole number of TO intercalation sites. Importantly, our peptide does not compete with TO binding, suggesting that RGGFGGRGG interacts with i-motif RNA primarily via external backbone interactions.
The observed heteroallosteric modulation does not disrupt the i-motif structure, as confirmed by CD spectroscopy (SI Fig. 3). Our peptide also has no such templating allosteric effect with the DNA hTeloC i-motif structure (SI Fig. 33). These findings further support our model of selective recognition of RNA i-motif structures by the RGGFGGRGG peptide, reinforcing the primary mechanisms of interaction as steric fit, hydration of folded structures, and guanidinium–phosphate interactions rather than primarily electrostatics in this RNA-peptide system.34
The RGGFGGRGG peptide is a useful model for exploring selectivity between RNA and DNA secondary structures. Our peptide is capable of selective recognition of the RNA i-motif structure over the DNA congener and corresponding single-stranded cytosine-rich oligonucleotides. We demonstrate that the binding of our peptide is pH-dependent, strongly correlating with the proportion of folded i-motif structure present.
To the best of our knowledge, our peptide is the first example of a small molecule that binds the RNA i-motif. While many DNA i-motif targeting compounds may also interact with RNA i-motif sequences, our peptide targets folded RNA selectively. Our results show that even minimalistic RNA-binding peptides can provide both structure and sequence specificity. The emerging significance of non-canonical and non-coding RNA secondary structures in transcriptional regulation and disease pathogenesis highlights the future potential of selective targeting peptides for novel therapies.16
Author contributions
L. C., P. T. and F. J. R. designed the research. L. C. performed synthesis of peptides and partially performed synthesis of oligonucleotides. L. C. performed experiments and analysed data with NMR. L. C. and P. T. analysed binding data. L. C. performed experiments and analysed data with CD. Figures were designed by L. C. and F. J. R. All the authors discussed the results and commented on the manuscript prior to submission.Conflicts of interest
There are no conflicts to declare.Data availability
This research did not generate any sensitive datasets. Unaltered raw data is available either in the supplementary information (SI), or is available upon request. Raw NMR titration data, including comparisons between fitting models, is publicly available at https://doi.org/10.6084/m9.figshare.30881207. Supplementary information: materials & methods, synthesis, characterisation, binding and titration data. See DOI: https://doi.org/10.1039/d6sc01203e.Acknowledgements
This work was supported by the Australian Research Council (ARC) through ARC Discovery Grants to F. J. R. and P. T. (DP250100081) and to P. T. (DP220101847), and an ARC DECRA Fellowship (DE220100558) to F. J. R. The authors acknowledge the UNSW Mark Wainwright Analytical Centre (MWAC) for providing access to the Nuclear Magnetic Resonance Facility, Structural Biology Facility, and the Bioanalytical Mass Spectra Facility. The authors also acknowledge Dr Hsiu Lin Li and Aidan Sajowitz at the UNSW RNA Institute for providing the RNA oligonucleotides and mass spectra analysis used in this study. https://BioRender.com was used to assist in figure creation for this manuscript.References
- J. D. Ransohoff, Y. Wei and P. A. Khavari, Nat. Rev. Mol. Cell Biol., 2018, 19, 143–157 CrossRef CAS PubMed.
- M. Zeraati, A. L. Moye, J. W. H. Wong, D. Perera, M. J. Cowley, D. U. Christ, T. M. Bryan and M. E. Dinger, Sci. Rep., 2017, 7, 708 CrossRef PubMed.
- H. A. Assi, M. Garavís, C. González and M. J. Damha, Nucleic Acids Res., 2018, 46, 8038–8056 CrossRef CAS PubMed.
- H. A. Day, P. Pavlou and Z. A. E. Waller, Bioorg. Med. Chem., 2014, 22, 4407–4418 CrossRef CAS PubMed.
- X. Han, J.-L. Leroy and M. Guéron, J. Mol. Biol., 1998, 278, 949–965 CrossRef CAS PubMed.
- K. Snoussi, S. Nonin-Lecomte and J. L. Leroy, J. Mol. Biol., 2001, 309, 139–153 CrossRef CAS PubMed.
- L. Lacroix, J.-L. Mergny, J.-L. Leroy and C. Hélène, Biochemistry, 1996, 35, 8715–8722 CrossRef CAS PubMed.
- M. Zeraati, D. B. Langley, P. Schofield, A. L. Moye, R. Rouet, W. E. Hughes, T. M. Bryan, M. E. Dinger and D. Christ, Nat. Chem., 2018, 10, 631–637 CrossRef CAS PubMed.
- J. J. King, K. L. Irving, C. W. Evans, R. V. Chikhale, R. Becker, C. J. Morris, C. D. P. Martinez, P. Schofield, D. Christ, L. H. Hurley, Z. A. E. Waller, K. S. Iyer and N. M. Smith, J. Am. Chem. Soc., 2020, 142, 20600–20604 CrossRef CAS PubMed.
- S. Takahashi, S. Ghosh, M. Trajkovski, T. Ohyama, J. Plavec and N. Sugimoto, Nucleic Acids Res., 2025, 53, gkaf500 CrossRef CAS PubMed.
- M. Martella, F. Pichiorri, R. V. Chikhale, M. A. S. Abdelhamid, Z. A. E. Waller and S. S. Smith, Nucleic Acids Res., 2022, 50, 3445–3455 CrossRef CAS PubMed.
- D. Guneri, E. Alexandrou, K. El Omari, Z. Dvořáková, R. V. Chikhale, D. T. S. Pike, C. A. Waudby, C. J. Morris, S. Haider, G. N. Parkinson and Z. A. E. Waller, Nat. Commun., 2024, 15, 7119 CrossRef CAS PubMed.
- J. J. Montero, I. López de Silanes, O. Graña and M. A. Blasco, Nat. Commun., 2016, 7, 12534 CrossRef CAS PubMed.
- Y. Cui, D. Kong, C. Ghimire, C. Xu and H. Mao, Biochemistry, 2016, 55, 2291–2299 CrossRef CAS PubMed.
- F. X. Han, R. T. Wheelhouse and L. H. Hurley, J. Am. Chem. Soc., 1999, 121, 3561–3570 CrossRef CAS.
- L. B. Cox, J. S. Mattick, I. A. Middleton, F. J. Rizzuto and P. Thordarson, Nucleic Acids Res., 2026, 54, gkaf1471 CrossRef CAS PubMed.
- J. Abramson, et al. , Nature, 2024, 630, 493–500 CrossRef CAS PubMed.
- A. Kishtagari and J. Watts, Ther. Adv. Hematol., 2017, 8, 317–326 CrossRef CAS PubMed.
- J. Ren and J. B. Chaires, Biochemistry, 1999, 38, 16067–16075 CrossRef CAS PubMed.
- P. Thordarson, Binding Constants and Their Measurement, in Supramolecular Chemistry: From Molecules to Nanomaterials, ed. P. Gale and J. Steed, John Wiley & Sons, Chichester, England, 2012, vol. 2, pp. 239–274 Search PubMed.
- https://supramolecular.org .
- D. B. Hibbert and P. Thordarson, Chem. Commun., 2016, 52, 12792–12805 RSC.
- M. A. S. Abdelhamid, A. J. Gates and Z. A. E. Waller, Biochemistry, 2019, 58, 245–249 CrossRef CAS PubMed.
- A. Pérez, A. Noy, F. Lankas, F. J. Luque and M. Orozco, Nucleic Acids Res., 2004, 32, 6144–6151 CrossRef PubMed.
- D. E. Draper, RNA, 2004, 10, 335–343 CrossRef CAS PubMed.
- W. Ren, K. Zheng, C. Liao, J. Yang and J. Zhao, Phys. Chem. Chem. Phys., 2018, 20, 916–924 RSC.
- A. Deep, A. Bhat, V. Perumal and S. Kumar, Mol. Ther.--Nucleic Acids, 2025, 36, 102474 CrossRef CAS PubMed.
- L. Liu, B. G. Kim, U. Feroze, R. B. Macgregor Jr and T. V. Chalikian, J. Am. Chem. Soc., 2018, 140, 2229–2238 CrossRef CAS PubMed.
- E. N. W. Howe, M. Bhadbhade and P. Thordarson, J. Am. Chem. Soc., 2014, 136, 7505–7516 CrossRef CAS PubMed.
- A. L. Moller, I. A. Middleton, G. E. Maynard, L. B. Cox, A. Wang, H. L. Li and P. Thordarson, Biomacromolecules, 2025, 26, 470–479 CrossRef CAS PubMed.
- J. Nygren, N. Svanvik and M. Kubista, Biopolymers, 1998, 46, 39–51 CrossRef CAS PubMed.
- Q. Sheng, J. C. Neaverson, T. Mahmoud, C. E. M. Stevenson, S. E. Matthews and Z. A. E. Waller, Org. Biomol. Chem., 2017, 15, 5669–5673 RSC.
- M. Takeuchi, M. Ikeda, A. Sugasaki and S. Shinkai, Acc. Chem. Res., 2001, 34, 865–873 CrossRef CAS PubMed.
- K. A. Schug and W. Lindner, Chem. Rev., 2005, 105, 67–114 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |