Open Access Article
Open Access ArticleBlack-box data: a new paradigm for biomedicine in the AI era
Luca
Naef
*a and
Michael
Bronstein
 *abc
*abc
aProxima Bio, USA. E-mail: luca@proximabio.com
bUniversity of Oxford, UK. E-mail: michael.bronstein@cs.ox.ac.uk
cAITHYRA, Austria
First published on 8th April 2026
Abstract
As artificial Intelligence cements its role as a cornerstone of scientific discovery, the field is undergoing a fundamental shift beyond the current transition from “white-box” first-principles models to “black-box” deep learning. We argue that a parallel, necessary transformation is emerging in data generation: the rise of “black-box data.” These data sources are intentionally optimized for machine consumption rather than human intuition—a trade-off we contend is essential to achieving the scale required for high-capacity biological foundation models. This article defines the “black-box data” paradigm, explores the necessity of this shift for the future of AI-driven science, and provides a unifying taxonomy illustrated by both historical precedents and contemporary breakthroughs.
In the past decade, we have witnessed a dramatic expansion of Artificial Intelligence (AI) and Machine Learning (ML) into a broad range of industries and scientific domains, from natural language processing (NLP) and computer vision (CV) to particle physics and chemistry. AI has beaten human players in games such as Go† or Starcraft‡, previously thought to be unconquerable. Most recently, it achieved top human-level performance in international math and programming olympiads.3
What characterizes these achievements is that they are done in software. In experimental fields like the life sciences and biology, where predictions typically need to be validated by a lab experiment, the number of AI successes has so far been more limited. DeepMind's AlphaFold2 (AF2), recognized by the 2024 Nobel Prize in Chemistry, stands out as perhaps the most prominent example, revolutionizing protein monomer structure prediction and, more recently, protein binder design.4
We have recently hypothesized5 that AF2's success stems partly from the significant degeneracy of protein structure space: the large, diverse PDB may provide good coverage of naturally occurring folds.6–8 Supporting this, recent large-scale applications of AF2 predicted only a single new fold across known protein sequences,9 suggesting either limited generalizability or, alternatively, that few novel folds remain to be discovered. Studies on other modalities corroborate the importance of data coverage: performance on protein–protein and protein-ligand complexes, where training data covers less of the interaction space, show strong dependence on similarity to training examples.10–12
The AlphaFold breakthrough has spurred ambition to replicate this success elsewhere in biology to produce “the next AlphaFold”. But it may be more critical to consider “the next PDB,” i.e., new experimental data sources1 for training future biological ML.13 The PDB originated in the 1970s, predating modern ML and its relevance to biology. It represents five decades of effort by thousands of structural biologists at an estimated cost of up to $50 billion.14 Building comparable datasets for new domains requires experimental approaches that bridge orders-of-magnitude gaps in cost and throughput.
Taking a historical perspective on algorithmic advances, from first-principles “white-box” models to present-day “black-box” deep learning, we explore the emergence of data sources optimized for ML rather than human consumption. Just as trading handcrafted features for learned representations drove breakthroughs in CV and NLP, we argue that an analogous tradeoff in data generation could unlock the scale needed for the next generation of biological foundation models. We call this paradigm “black-box data” and present a unifying taxonomy2 with historical and recent examples.
Software 2.0, the bitter lesson and black-box methods
The 2012 “ImageNet moment” is generally considered the Rubicon when end-to-end deep learning started to progressively overtake human-designed algorithms across a wide spectrum of domains. ImageNet was a large-scale computer vision competition consisting of classifying images of everyday objects from 1000 classes (such as cats, dogs, etc.), a notoriously hard challenge in the field. At the time, most state-of-the-art computer vision algorithms were based on extracting carefully hand-crafted features from images. In 2012, for the first time, the competition was won by AlexNet, a method that completely removed the human interpretable features in favor of data-driven features directly learned by a deep Convolutional Neural Network.15This transition was dubbed the “Software 2.0″ paradigm by Andrej Karpathy.16 In traditional “Software 1.0,” humans explicitly program algorithms that operate on data. Software 2.0 is programmed through data: a neural network learns from data examples without explicit instructions, resulting in a black-box system whose rules are encoded implicitly in weights rather than interpretable code. In the ImageNet example, traditional computer vision approaches (“Software 1.0”) attempted to derive an “equation of a cat” via hand-crafted features distinguishing cats from dogs. Modern deep-learning based approaches (“Software 2.0”) instead present many labeled examples to a neural network, which learns the distinction on its own (Fig. 1).
Over the past decade, the abandonment of handcrafted inductive biases and features towards ever more general learning methods has been a predominant trend, often referred to as “The Bitter Lesson”: simple systems that scale with available computational power will eventually outperform more complex systems relying on human knowledge.17 One could perhaps see black-box data as the natural continuation of this trend beyond algorithms, by removing handcrafted human biases also from the data generation process.
Protein structure prediction methods are a prime example of this trend. Early works on protein folding exploited first-principle physics-based models. AlphaFold1 (2018), DeepMind's first foray into the field, still combined three separate deep learning models to predict distances and confidence, and then folded proteins with predicted distances given “handcrafted” Multiple Sequence Alignments (MSAs).18 AlphaFold2 (2021) was a single end-to-end trained model with strong domain-specific inductive biases, including a geometric frame-representation and roto-translation-invariant and chirally-aware Frame-Aligned Point Error (FAPE), geometry-inspired triangle operations, and a roto-translation Invariant Point Attention (IPA).19 AlphaFold3 (AF3, 2024) removed most inductive biases (IPA, frame representation, FAPE, and invariant featurization) in favor of data augmentation to learn the required invariances with a general Transformer-inspired architecture.20 Yet, the further expected simplification has not yet happened: AF3 still notoriously relies on MSA and a geometry-inspired architecture. More “general” methods that remove MSA and/or triangle operations (ESMFold, SimpleFold, ESM3) perform significantly worse.21–23 Hence, while this has been a trend, it is not yet a fait accompli.
Black-box AI methods require AI-first data sources
One of the unique aspects of the PDB is that it provided not only a large, clean dataset covering many folds occurring in nature, but also a clear scientific question with a benchmark (CASP competition) allowing to measure progress. This is far from being the case in many other life science domains, such as single-cell biology, protein-ligand complexes,10 protein–protein complexes,11 disease biology, human phenotypes, and other biological subdomains.24,25 Not only is there less homogenous and complete data, but in some cases, even the question one tries to answer from the data is not well defined.ML models in biology have traditionally relied on data produced as a byproduct of scientific inquiry. Repositories like ChEMBL,26 PDB,27 PRIDE,28 GEO,29 OpenTargets30 aggregate and standardize this data, but they are not purpose-generated for ML. This leads to poor standardization, limited scale, and systematic biases. Unlike the diverse naturally occurring protein structures in the PDB, protein–drug complexes, for example, are often derived from drug discovery campaigns biased toward a narrow set of medically relevant targets. Freitas et al. found that of 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 016 high-quality protein-ligand structures spanning over 500 families, more than 45% came from just ten families of enzymes and receptors.31 This lack of diversity contributes to ML performance on protein-ligand complexes significantly trailing protein monomers.20
016 high-quality protein-ligand structures spanning over 500 families, more than 45% came from just ten families of enzymes and receptors.31 This lack of diversity contributes to ML performance on protein-ligand complexes significantly trailing protein monomers.20
Since existing data sources are optimized for human-driven discovery, they have tradeoffs often suboptimal for ML. In traditional drug discovery, for example, deep coverage of individual targets is critical, whereas ML methods likely require diverse targets.32 Humans generally prefer small, low-noise datasets for deductive reasoning, while ML is noise-tolerant but sample-inefficient, benefiting more from large noisy datasets. Rolnick et al. demonstrated that models achieve over 90% accuracy on MNIST even with 100 mislabeled examples for every true label, but a minimum absolute number of correct labels is critical.33 The adage “quality over quantity” is inverted in the black-box era: “quantity is quality,” as single-cell pioneer Aviv Regev puts it.34
Current foundation models bear this out. Large language models are pretrained on massive, diverse, noisy web-crawled corpora (C4, CommonCrawl35), then fine-tuned on small high-quality datasets via reinforcement learning from human feedback.36 Intentional noise injection is also broadly useful in ML for improving generalization.37 Denoising diffusion models38 take this further, training explicitly on data synthetically noised to varying levels. This also allows naturally noisy experimental data to be integrated at higher-noise training steps, where inherent measurement noise becomes indistinguishable from synthetic noise.39
Another problem with “data as a byproduct of scientific inquiry” is the lack of true negatives. Partly a consequence of the academic incentive and publication model, unsuccessful experiments are rarely reported, also known as the “file drawer problem”.40 Yet such data is critical for AI.41
Above all, the cost of generating datasets such as the PDB is likely in the billions,14 making a fundamental paradigm shift on biological data generation all but necessary.
Black-box data: a paradigm of AI-first, scalable data sources
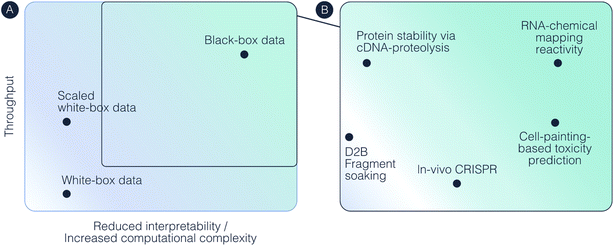
In recent years, we observe the emergence of black-box data sources that focus directly on ML consumption. The leitmotif is moving away from tradeoffs optimized for human consumption – high signal-to-noise protocols, expensive low-noise readouts, limited diversity and human interpretability – enables orders of magnitude improvements in throughput. We can see a clear analogy to the emergence of black-box algorithms – trading off human interpretability in favor of scalability optimized for ML. In the most extreme version of this paradigm, such datasets can become completely unintelligible to humans, hence our term “black-box data”.5Computationally, the defining characteristic of black-box data is the complexity of the inverse problem separating the raw measurement from the quantity of interest. In white-box data, this mapping is simple, often identity or a known analytical transformation. In black-box data, reconstruction requires algorithms with high representational complexity (whether classical optimization or learned mappings) and strong priors drawn from domain knowledge or adjacent examples. The distinction is not binary, illustrated in Fig. 2. The degree of “black-boxness” corresponds to how much algorithmic machinery, and how strong the priors, must sit between measurement and useful output. At one end is simply the DL-based denoising of assay outputs (i.e., output data structure is fully preserved), e.g. Direct-to-Biology (D2B).42 At the other are measurements used purely as pretraining signal, where the relationship to downstream outputs is learned implicitly and not fully known explicitly (e.g., chemical probing reactivity as latent supervision signal for RNA structure prediction43). Hence “black-box data” is defined by emphasizing trade-offs that optimize for ML rather than the lack of interpretability – similar to state-of-the-art “black-box models,” i.e., neural networks that often still contain interpretable elements such as attention matrices or equivariant layers.44
We note that “black-box” refers to the relationship between measurement and quantity of interest from the perspective of unaided human interpretation, not to the rigor of data generation. As with black-box algorithms, which require careful architectural and training choices despite producing opaque internal representations, black-box data demands deep domain expertise to hypothesize which modalities carry sufficient (possibly latent) signal and to design the corresponding reconstruction machinery.
Historical examples of black-box data
Our notion of “black-box data” in fact predates the recent dominance of black-box methods and was successfully applied in the past with white-box methods. Perhaps the most extreme example of black-box data is from the mathematical theory of compressed sensing,45 providing a theoretical guarantee of the ability to recover sparse signals from a few incoherent random projections. The data in compressed sensing intentionally looks like noise illegible to a human. The measurement process (represented by a random sensing matrix) is designed together with the reconstruction algorithm (based on solving an optimization problem) and requires a good understanding of the underlying problem (e.g. the assumption that data has a sparse representation in some basis, and the sensing matrix is incoherent with it). Compressed sensing techniques have been used in the medical imaging domain to design faster MRI acquisition protocols.46 Multiple-access protocols, such as CDMA (Code-Division Multiple Access) and OFDM (Orthogonal Frequency-Division Multiplexing), are further examples, critical in mobile communications such as 5 G. They allow transmitting information from multiple users over a single channel by combining the different users' signals via a convolution scheme. This reduces latency and increases throughput by avoiding time-dividing the channel (TDMA, Time-Division Multiple Access). The combined signal is completely uninterpretable without the deconvolution algorithm.47Within biology, the strategy of trading interpretability for scale is not new either. The Human Genome Project succeeded in part due to “whole-genome shotgun sequencing” (WGS) pioneered by Craig Venter and Celera Genomics.48 Rather than assembling the genome sequentially, it is fragmented into millions of short pieces that are reassembled using alignment methods. The individual reads are too short to carry useful information, but in conjunction with alignment they can reconstruct (nearly) the whole genome. Shifting away from sequential assembly is what ultimately made the completion of the human genome possible.
Modern examples and a taxonomy of black-box data generation tricks
While not new conceptually, we now increasingly see the emergence of new experimental “black-box data” sources developed in conjunction with ML. Excitingly, such black-box methods have already been employed across the whole lifecycle of basic biological research and drug discovery and development, as depicted in Fig. 3.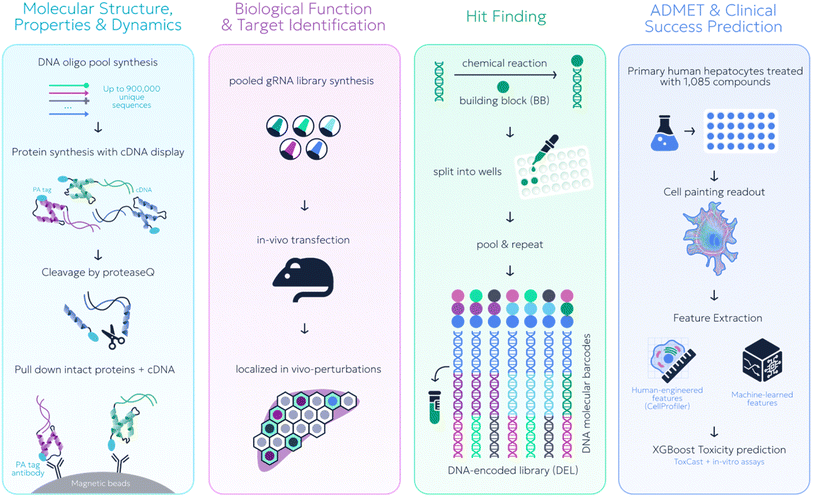 | ||
| Fig. 3 Black-box data paradigms have been applied to the whole drug discovery spectrum. From left to right: high-throughput protein stability via cDNA proteolysis (Main text),49 target identification through in vivo CRISPR (Appendix),50 DNA-encoded libraries for hit finding (Appendix),51 liver toxicity prediction using cell painting readouts (main text).52 | ||
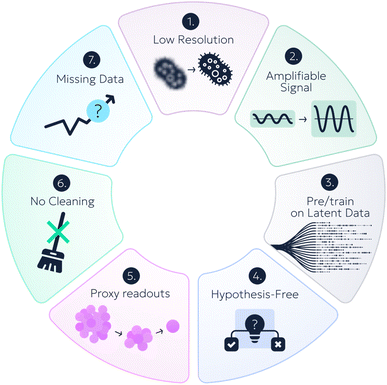
Despite divergent objectives and modalities, black-box data approaches share recurring structural themes or “tricks”. The examples listed below and in the Appendix can be organized into a seven-trick taxonomy (Fig. 4). Many methods combine multiple tricks, often synergistically increasing throughput (Table 1).
| Name | Primary trick | Primary application area | Short description | Location |
|---|---|---|---|---|
| Cross-linking Mass spectrometry | Low resolution | Molecular structure, properties & dynamics | Whole-proteome structure determination via covalent distance markers | Main text |
| CMap | Low resolution | Biological function & target identification | Perturbomics via landmark gene arrays and imputation (L1000) | Appendix |
| cDNA proteolysis | Amplifiable signal | Molecular structure, properties & dynamics | Mega-scale protein folding stability measurements via sequencing | Main text |
| Human domainome 1 | Amplifiable signal | Molecular structure, properties & dynamics | Yeast protein complementation assay to link domain stability to growth read via sequencing readout | Appendix |
| DNA-encoded library | Amplifiable signal | Molecular structure, properties & dynamics | Combinatorial library construction with DNA tags for massive-scale small molecule screens with sequencing readout | Appendix |
| Chemical mapping | Latent data | Molecular structure, properties & dynamics | Mutational profiling of RNA after secondary structure-dependent reagent exposure for structure prediction via reactivity pre-training | Main text |
| ESMFold | Latent data | Molecular structure, properties & dynamics | Sequence-only pretraining extracts co-evolutionary motifs benefiting downstream structure prediction | Appendix |
| Sup35-aggregation screen | Hypothesis-free | Molecular structure, properties & dynamics | Massive random peptide screening via yeast Sup35 aggregation measured through ade1 reporter to train aggregation predictors | Main text |
| Cell painting to predict toxicity | Proxy readout | ADMET & clinical success prediction | In vivo (and in vitro) toxicity prediction at high-throughput via ML on in vitro cell painting outputs from hepatocytes post compound exposure | Main text |
| L1000 toxicity translation | Proxy readout | ADMET & clinical success prediction | In vivo rat kidney toxicity prediction at high-throughput via L1000 human cell line profiling and ML | Appendix |
| Organoid toxicity translation | Proxy readout | ADMET & clinical success prediction | Human in vivo GIT toxicity prediction at high-throughput via human ileal organoids and ML | Appendix |
| In vivo CRISPR | Proxy readout | ADMET & clinical success prediction | Pooled, localized in vivo CRISPR for high-throughput in vivo target identification | Appendix |
| CRM fragment screening | No cleaning | Hit finding | Crystallographic fragment screening of crude reaction mixtures denoised via SAR model for high-throughput hit-to-lead/lead optimization | Main text |
| Dyna-1 | Missing data | Molecular structure, properties & dynamics | Reinterpret systematic absences in solution NMR tables as dynamics to train protein dynamics DL model Dyna-1 | Main text |
| Diffuse scattering | Missing data | Molecular structure, properties & dynamics | Reinterpret diffuse scattering signal around bragg-peaks as dynamics to train protein-dynamics DL models | Appendix |
| AlphaMissense | Missing data | Biological function & target identification | Reinterpret missing variants as purifying selection to weakly label pathogenic variants and train DL variant effect predictor AlphaMissense | Appendix |
Trick 1: leverage low resolution readouts
Resolution and throughput are often inversely related. Low resolution manifests in two forms: sparse sampling (measuring a subset of features, as in microarrays capturing landmark transcripts rather than full transcriptomes53) or increased noise (losing high-frequency information, as in low-resolution Cryo-EM maps). Deep learning is particularly well-suited to both cases because learned priors can reconstruct missing or corrupted information from partial observations. Examples include cross-linking mass spectrometry (below), mutate-and-map chemical probing, and NMR-derived distance restraints (both covered later).Example: Sparse distance restraints combined with structural foundation models unlock atomic-accuracy structures at a fraction of the cost
One of the central tenets of protein biology is that structure defines function.54 However, as mentioned previously, current methods for structure elucidation are cost prohibitive without advances in data generation.14 Structural proteomics provides one pathway to address this by switching to a readout that enables characterizing complex mixtures of many proteins in parallel (Mass Spectrometry). Cross-Linking Mass Spectrometry (XLMS) is a type of structural proteomics which allows to obtain sparse distance restraints at a whole proteome scale.55 By incubating cells or lysates with covalent probes called Cross-Linkers, residues that are at a distance no further than the maximum length of the extended crosslinker are connected covalently. Through these covalent markers, along with specialized spectral search methods,56 cross-linked peptides can be detected using MS, yielding a potentially whole-proteome, sparse residue contact map. Traditionally, this was predominantly employed to validate or guide biophysics approaches with orthogonal structural data (HDX-MS, NMR, templates).57 However, with deep learning-based structure prediction methods, models can use these structural restraints at inference time to dramatically increase success rates in structures prediction.58,59 Recently, a biotech startup, Proxima Bio, has massively scaled this data source and unveiled their foundation model, Neo-1, making use of this data. They demonstrated that they can leverage their XLMS-based platform to predict novel protein interfaces that cannot be predicted by current models alone, and dramatically increase accuracy of small-molecule induced protein complex prediction.60Trick 2: link quantity of interest to amplifiable signal
Increasing signal-to-noise ratio is a central problem in any assay. Including signal amplification steps can maintain high signal while increasing throughput. This has made two general readout categories “privileged”: sequencing readouts, allowing for amplification via PCR; and positive selection assays where the quantity is tied to growth. Examples of the former include DNA-encoded libraries (Appendix), mRNA-display,61 cDNA-linked proteolysis as in the MEGAScale experiment (covered below) and many more. Examples of the latter include yeast-two-hybrid screens62 and the Sup35 and aPCA aggregation and stability assays covered later.Example: MEGAscale proteolysis to train protein dynamics foundation models
Beyond structure, understanding how sequence and structural variation affect protein function is critical for protein engineering, therapeutic design, and disease mechanism. An example of addressing this data-need is cDNA display proteolysis, developed by the Rocklin lab.49 In the MEGAscale assay, large DNA pooled libraries are produced with a puromycin linker, yielding protein–cDNA fusions captured on resin, making the proteins easily identifiable via the cDNA. The pool is exposed to titrations of proteases such as trypsin and chymotrypsin and after washing away cleaved proteins, intact proteins can be sequenced and counted. For each protein, the decline in counts across titrations yields a K50 value which is converted to folding free energies. Unlike direct structure determination, this gives indirect structural data via folding free energies from a massively scalable assay, enabling over 776![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 measurements across hundreds of domains and dense mutational scans.
000 measurements across hundreds of domains and dense mutational scans.
This data was used by a team at Microsoft Research to train BioEmu, a generative model that emulates protein equilibrium ensembles.63 To obtain paired structural data, short molecular dynamics (MD) simulations for over 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 proteins from the dataset were generated to produce ensembles of folded and unfolded structures with experimental ΔG values used to reweight the simulation, creating a training set where the ratio of folded to unfolded structures reflects the measured stability. They further developed Property-Prediction Fine-Tuning (PPFT) to train the model directly to produce a distribution matching the experimental ΔG values without expensive paired simulation data.
000 proteins from the dataset were generated to produce ensembles of folded and unfolded structures with experimental ΔG values used to reweight the simulation, creating a training set where the ratio of folded to unfolded structures reflects the measured stability. They further developed Property-Prediction Fine-Tuning (PPFT) to train the model directly to produce a distribution matching the experimental ΔG values without expensive paired simulation data.
Trick 3: pre-train models on large abundant data with related but latent information
Black-box methods learn transferable representations rather than fixed algorithms. This is significant because it enables pretraining on abundant data from adjacent domains and fine-tuning for data-limited tasks without an explicit known relationship between pretraining and target tasks. RibonanzaNet2 and Dyna-1, covered in the following, as well as AlphaMissense and ESMFold (Appendix) all pretrain on data containing latent but not directly mappable task-relevant information.Example: chemical mapping data for RNA structure prediction and design
Structure prediction of nucleic acids currently significantly lags the performance of proteins. At CASP16, all top methods were expert predictors, with ML methods significantly behind.64 Data scarcity is a core contributor: the PDB contained over 227![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 protein structures in 2025,27 but only 14
000 protein structures in 2025,27 but only 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 750 DNA/RNA-protein structures. And the 6500 RNA structures are biased towards a few families (tRNAs, riboswitches, and ribozymes).65 A promising black-box remedy for this is chemical mapping43 where reagents preferentially mutate flexible and/or unpaired nucleotides to yield reactivity values when sequenced under mutational profiling (MaP). This provides indirect RNA structural data that is many orders of magnitude more economic than traditional, direct structure determination (Cryo-EM/crystallography). Two popular reagents are dimethyl sulfate (DMS) for N1-adenine and N3-cytosine methylation for solvent-exposed nucleotides, and 2-aminopyridine-3-carboxylic acid imidazolide (2A3) for conformationally dynamic nucleotides. Multiple extensions exist: Mutate-and-map (M2)66 systematic substitutions change reactivity of nucleotide pairs, creating correlated off-diagonal signals providing a contact map. MOHCA-seq67 and KARR-seq68 correlate with distograms. Recent efforts69 have scaled this data source significantly. As part of the 2025 Stanford 3D RNA Folding Kaggle challenge, 40 M sequences chemical mapping profiles were produced and used to pre-train a range of different models, including an encoder predicting sequence-wise chemical reactivity. Combined with a diffusion structure prediction head trained on RNA structures, structure prediction performance dramatically improved compared to a model that was not pretrained on this task.70,71 Representations learned from these tasks were also leveraged in a recent model, RNAPro, to surpass AF3 in RNA structure prediction.72 The practical therapeutic relevance of this data source for RNA design was further demonstrated by Joshi et al. which combined a RNA language model (gRNAde) filtered by the first version of RibonanzaNet to yield high success rates surpassing previous models.73
750 DNA/RNA-protein structures. And the 6500 RNA structures are biased towards a few families (tRNAs, riboswitches, and ribozymes).65 A promising black-box remedy for this is chemical mapping43 where reagents preferentially mutate flexible and/or unpaired nucleotides to yield reactivity values when sequenced under mutational profiling (MaP). This provides indirect RNA structural data that is many orders of magnitude more economic than traditional, direct structure determination (Cryo-EM/crystallography). Two popular reagents are dimethyl sulfate (DMS) for N1-adenine and N3-cytosine methylation for solvent-exposed nucleotides, and 2-aminopyridine-3-carboxylic acid imidazolide (2A3) for conformationally dynamic nucleotides. Multiple extensions exist: Mutate-and-map (M2)66 systematic substitutions change reactivity of nucleotide pairs, creating correlated off-diagonal signals providing a contact map. MOHCA-seq67 and KARR-seq68 correlate with distograms. Recent efforts69 have scaled this data source significantly. As part of the 2025 Stanford 3D RNA Folding Kaggle challenge, 40 M sequences chemical mapping profiles were produced and used to pre-train a range of different models, including an encoder predicting sequence-wise chemical reactivity. Combined with a diffusion structure prediction head trained on RNA structures, structure prediction performance dramatically improved compared to a model that was not pretrained on this task.70,71 Representations learned from these tasks were also leveraged in a recent model, RNAPro, to surpass AF3 in RNA structure prediction.72 The practical therapeutic relevance of this data source for RNA design was further demonstrated by Joshi et al. which combined a RNA language model (gRNAde) filtered by the first version of RibonanzaNet to yield high success rates surpassing previous models.73
Trick 4: hypothesis-free assays – unbiased, multiplexed and highly pooled readouts
Hypothesis-driven fields like drug discovery test limited, targeted hypotheses with low-throughput, low-noise, interpretable readouts. Unbiased “hypothesis-free” approaches sacrifice these properties but provide the diversity ML models need. Examples include the aforementioned structural proteomics analyzing whole-cell lysates or Sup35-based aggregation screens testing random 20-mer peptides rather than curated sets that we discuss below. Multiplexed and pooled readouts typically enable this scale.Example: massively scaled random peptide screening to train aggregation predictors
Thompson et al. used a hypothesis-free approach to predict protein aggregation.74 Instead of a biased library, they constructed a pooled library of 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 random 20-mer peptides, each cloned in-frame upstream of the yeast prion nucleation domain of Sup35. In this system, peptides that nucleate or promote amyloid-like assembly of the Sup35 fusion trigger nonsense suppression of an ade1 reporter, causing growth on adenine-deficient media. Over several growth cycles fitness differences are amplified and barcodes are quantified by sequencing. They then developed a DL model (CANYA) with this data, outperforming all existing aggregation predictors trained on smaller, human-curated datasets.
000 random 20-mer peptides, each cloned in-frame upstream of the yeast prion nucleation domain of Sup35. In this system, peptides that nucleate or promote amyloid-like assembly of the Sup35 fusion trigger nonsense suppression of an ade1 reporter, causing growth on adenine-deficient media. Over several growth cycles fitness differences are amplified and barcodes are quantified by sequencing. They then developed a DL model (CANYA) with this data, outperforming all existing aggregation predictors trained on smaller, human-curated datasets.
Trick 5: use proxy readouts that are less costly and higher throughput
Proxy readouts are ubiquitous in science: biomarkers, cell cultures, or model organisms. In our context, the use of proxies is a special case of Trick 3 (pretraining on latent information), where the readout derives from a proxy system sharing relevant biology but without direct translation to the target. Black-box methods excel here because they can learn the proxy-to-target mapping. Examples include Cell Painting to predict hepatotoxicity, ileal organoids to predict gastrointestinal toxicity (Appendix), in vivo localized CRISPR to enable pooled in vivo target discovery (Appendix).Example: high-throughput in vitro readouts to predict in vivo toxicity across species
Cell Painting75 has historically been extensively used as a biological proxy-readout in conjunction with black-box methods, perhaps most prominently by the startup Recursion Pharmaceuticals. In Cell Painting, cells are imaged after staining with six dyes highlighting distinct morphological features. Embeddings are typically extracted via hand-crafted pipelines (CellProfiler) or learned models. For example, in a seminal 2024 publication, Recursion leveraged black-box embeddings of Cell Painting data through their proprietary model to discover a hitherto unknown undesired genomic off-target pattern in CRISPR-Cas9 screens.76Cell Painting was recently also applied in the context of toxicity prediction. Next to potency and affinity, developing drugs involves optimizing many other properties such as Absorption, Distribution, Metabolism and Toxicity (ADMET). Since these are often measured in vivo, throughput is limited. Data sources which alleviate this bottleneck are needed, with an early example being pooling multiple compounds into a single animal via Cassette Dosing.77
To address these throughput limitations, in a recent preprint by a team from MIT, the Broad Institute and the startup Axiom Bio, authors exposed primary human hepatocytes to 1085 compounds with previously known in vivo hepatotoxicity readouts from ToxCast.78 They assessed whether Cell Painting is simultaneously predictive of hundreds of in vitro toxicity readouts, including assays for metabolic activity and membrane damage measured by the authors and 412 cytotoxicity and mode-of-action endpoints measured by the ToxCast screening effort.52 They extracted various features from the images, including hand-crafted CellProfiler features and “black-box” embeddings (CellPainting CNN and DINOv2), and used the features to predict assay activity using XGBoost.
In a similar spirit, Gardiner et al. developed a black-box inspired cross-species toxicity translation: they train a ML model on the molecular structure and their associated in vitro human cell line landmark gene array profiles to predict in vivo rat kidney toxicity measured via blood urea nitrogen (combining both the “proxy” and “low resolution theme” via the L1000 assay also leveraged in CMap, see appendix).79 Researchers at Celgene employed a similar idea – in this case leveraging Machine Learning and human ileal organoid models to recapitulate human clinical in vivo Gastrointestinal toxicity (GIT) with 90% accuracy.80
Trick 6: omit cleaning or filtering of readouts
Purification and isolation are often rate-limiting: protein production, small molecule synthesis, and sample preparation are all bottlenecked by purification. Skipping purification yields noisier readouts, connecting this trick to Trick 1 (low-resolution readouts) through shared reliance on DL noise tolerance and priors. The distinction: Trick 1 changes the readout modality (e.g., Cryo-EM to XLMS), while Trick 6 removes a protocol step within the same modality. Direct-to-Biology (D2B, below) and DNA-encoded libraries (appendix) exemplify this approach. Trick 6 also intersects with Trick 4 (hypothesis-free assays), which often measure unpurified, unbiased samples by design.Example: crude reaction mixtures unlock high-throughput lead optimization
Nowadays, many thousands of DNA sequences or proteins can be synthesized and tested on the order of weeks.81 Yet for small molecules, custom synthesis is significantly slower, often requiring months, years, or even decades in extreme cases to establish synthetic pathways for just a single new molecule,82 dramatically slowing hit-to-lead campaigns in drug discovery. The large synthesis quantities required (due to material losses during purification) and the slow speed of purification are dominant bottlenecks in synthesis.83 In D2B this is addressed by testing Crude Reaction Mixtures (CRM) directly without purification, allowing well-based parallel nanoscale synthesis with dramatic throughput increase at the cost of increased false negatives/positives due to impurities.42 Again, ML is used to compensate for this. An example is McCorkindale et al., who predicted potency of each molecule, allowing to rescue mixtures that are incorrectly labelled as weak binders in the assay but labeled positive by the model.84A promising combination has been with fragment screening. As opposed to traditional high-throughput screening, where often billions of compounds are tested for binding, in fragment screening a smaller set of diverse (e.g. few thousand) of low-molecular weight fragments (120–250 dalton) are screened.85 These can be merged, linked or elaborated into complete and high affinity binders, allowing to explore a wide range of chemical space with a small “basis set” of tested fragments. Since initial fragments are often of lower affinity, highly sensitive readouts, such as crystallographic soaking are needed.86 Grosjean et al. employed this to the bromodomain of PHIP(2).87 CRMs of 957 analogs of a fragment hit yielded 22 binders. A simple structure–activity-relationship algorithm built on the 22 binders could denoise the results by rescuing false negatives, identifying 26 additional, mislabeled binders. They further used it in virtual screening, identifying 9 binders. Recently, the OpenBind consortium was formed to systematically scale this data source for Machine Learning purposes.88
D2B is also extensively used by many larger companies and startups.89 Kimia Therapeutics employs it in conjunction with Chemotype Evolution90 where “generation of molecules” are produced starting from a target-binding bait fragment with a reactive handle that is combinatorially coupled to fragments with further reactive handles, enabling iterative “activity-based” evolution. They demonstrated this for a selective covalent KRAS-G12C inhibitor series.91 Octant is another example coupling D2B to multiplexed reporter assays.92–94
Trick 7: leverage signal from missing, or weakly-labeled data
Missing or noisy data is often systematic rather than random. While insufficient for human interpretation, the structure implicit in the missing data can provide valuable weak supervision for ML. Examples: AlphaMissense95 treats unobserved variants as pathogenic (purifying selection), diffuse X-ray scattering96 extracts dynamics from traditionally discarded scattering noise (all in appendix), and missing NMR peaks can indicate conformational exchange rather than experimental failure (below). Trick 7 differs from Trick 1 since here the perceived “noise” is the primary source of signal and not noise after all. This is related to Trick 3 as the patterns in the noise may be primarily latent.Example: missing NMR peaks is supervision to learn protein dynamics
Experimental data on protein dynamics is hard to obtain at scale. Models are often exclusively trained on a limited amount of simulated data obtained at high computational cost. Wayment-Steele, El Nesr et al. developed a method to extract microsecond-to-millisecond (µs–ms) protein dynamics from missing NMR peak assignments in the Biological Magnetic Resonance Data Bank (BMRB) and trained Dyna-1, a deep learning model predicting these dynamics, to demonstrate the data's potential.97 The main realization was that “missing” NMR peaks (systematic absences) from chemical shift datasets can indicate µs–ms dynamics. In the regime where the interconversion rate is comparable to the angular frequency separation of exchanging states, transverse relaxation acquires an exchange contribution, producing line broadening and, in the limit, loss of peaks in standard heteronuclear experiments. Solution NMR assignment tables in the BMRB often contain these absences which were traditionally interpreted as experimental artifacts. To extract signals from this noisy weak supervision, they built on a frozen pre-trained language model, training a prediction head on data extracted from ∼10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 proteins from the BMRB (two orders of magnitude larger than existing datasets) to produce Dyna-1. It accurately predicts µs–ms dynamics when evaluated against high quality curated datasets (RelaxDB, RelaxDB-CPMG). This demonstrates another critical component of black-box data approaches – the requirement for high-quality “white-box” (often smaller sized) benchmark data.
000 proteins from the BMRB (two orders of magnitude larger than existing datasets) to produce Dyna-1. It accurately predicts µs–ms dynamics when evaluated against high quality curated datasets (RelaxDB, RelaxDB-CPMG). This demonstrates another critical component of black-box data approaches – the requirement for high-quality “white-box” (often smaller sized) benchmark data.
Future outlook: the scientific process in the black-box age
We believe that black-box data provides a viable path to overcome data limitations that bottleneck the advancement of biological applications of AI. As the field is increasingly driven through model-led discoveries, much of the way data is generated will change to adapt to ML as the primary consumer. This will drive a fundamental shift in the types of data created and there will be critical considerations to make this transition work.The generation of black-box data should not be viewed solely as a static, feed-forward process. With lab-in-the-loop and active learning approaches, instead of passively mining existing datasets, models can evaluate their own uncertainty to dictate the next batch of massively multiplexed experiments. This closed-loop experimental design will ensure that high-throughput, noisy assays are deployed where the model needs them most, maximizing data efficiency and accelerating the discovery process in a way that traditional, hypothesis-driven screening cannot.
Since black-box datasets sacrifice human interpretability for scale, they might be fundamentally unintelligible without algorithmic deconvolution. Standardizing raw readouts, precise experimental metadata, and noise profiles across different laboratories is thus an important hurdle. We believe that more generally, the field requires a shift toward “AI-first” data repositories moving away from the traditional scientific publishing route—where only cleaned, processed conclusions are shared—toward a pipeline where raw, unpurified readouts are standardized and fed directly into foundation models.
Nascent consortium efforts provide an early blueprint for this infrastructure. For example, the OpenBind consortium was formed to systematically scale and standardize crude reaction mixture fragment screening data for ML purposes. Similarly, the Diffuse Project aims to systematically collect and standardize previously discarded X-ray scattering noise into a shared repository for learning protein dynamics.
The importance of human intuition remains undiminished in the black-box data era. Even though the required signal-to-noise of black-box data is reduced, and the signal can be indirect and latent, there still needs to be sufficient signal for the models to extract. Hypothesizing and testing which black-box data methods contain sufficient signal will need to be driven by domain expertise, as will be evaluating the resulting models.
White-box and noise-free data will be needed to verify and benchmark black-box methods. While the learning process of these methods is noise tolerant, their evaluation is less so.98 Being black-box makes this even more critical as we cannot always understand, but only assess these methods for their accuracy, making evaluation a critical fail-safe. In practice, we expect white-box and black-box data to exist in a symbiotic relationship: small curated white-box datasets can be used to construct initial priors, calibrate reconstruction algorithms, validate model predictions, and it will often be desirable to “post-train” models trained on more noisy “black-box” data with noise-minimized high-quality white-box data.
This lack of interpretability is a common criticism of black-box data and algorithms. The overarching concern is that black-box approaches usher in a “post-theory” science where we can simulate, predict, yet not understand future scientific phenomena.99 While black-box methods are not directly interpretable, there exists nonetheless a plethora of tools from the field of Mechanistic Interpretability (MI) to extract knowledge from these models such as sparse autoencoders.100 A biological example is the categorical Jacobian, applied to language-model based folding methods, to understand the motif-driven recall process that underlies the success of the current generation of folding methods.101 In another case, teams from Prima Mente and Goodfire used MI on a large foundation model, Pleiades, pretrained on large amounts of patient genomic and ctDNA, to extract the finding that ctDNA fragment length could serve as diagnostic marker for Alzheimer's from the weights of the black-box model.102 These point to a future where there might be an inversion of the scientific method from understand, encode, and then simulate – a Software 1.0 paradigm – to encode, simulate, understand (via MI) – a Software 2.0 approach in which logic is not an input into the scientific discovery process but its output. Hence mechanistic interpretability will become critical to extract knowledge about scientific processes from the weights of neural networks that simulate them – standing on the shoulders of giants increasingly made of silicon.
Author contributions
L. N. and M. B. conceived the idea and wrote the manuscript.Conflicts of interest
L. N. and M. B. have employement relations with and hold equity in Proxima Bio, whose technology is mentioned in the paper. M. B. is on the SAB of Recursion Pharmaceuticals whose technology is mentioned in the paper.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this Perspective paper.Appendix
Below we present an expanded example list of black-box data sources that have been combined with ML. We group them by the corresponding application area.Structural biology
Protein biophysical properties
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 534 variant–abundance measurements across 522 domains, correlating strongly with independent biophysical measurements and the MEGAscale experiment. They combined the data with the protein language model ESM1v107 correlating measured stability scores with predicted fitness scores, identifying mutations whose effect on fitness was not explained by stability. Outlier mutations were strongly enriched in functional sites like DNA/protein-binding interfaces, yielding additional structural readouts when combined with Deep Learning models.
534 variant–abundance measurements across 522 domains, correlating strongly with independent biophysical measurements and the MEGAscale experiment. They combined the data with the protein language model ESM1v107 correlating measured stability scores with predicted fitness scores, identifying mutations whose effect on fitness was not explained by stability. Outlier mutations were strongly enriched in functional sites like DNA/protein-binding interfaces, yielding additional structural readouts when combined with Deep Learning models.
Biological function
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 080 perturbations (19
080 perturbations (19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 811 small molecule compounds, 18
811 small molecule compounds, 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 493 shRNAs, 3462 cDNAs, and 314 biologics). Its high throughput stems from the sparse L1000 assay, which measures a pre-defined reduced set of landmark genes per sample via an array instead of performing bulk or single-cell RNA sequencing. This significantly reduces cost and increases throughput (1.3 million profiles). 978 “landmark” transcripts are measured (selected to span diverse pathways and be representative for co-expressed genes) – in addition to 80 invariant controls. This is done via ligation-mediated amplification (LMA) and bead-based fluorescent detection.108 In CMap, cells from a standardized nine-line panel are exposed to small molecules or genetic perturbations over multiple concentrations and short exposure times. Transcriptome-wide values for the 11
493 shRNAs, 3462 cDNAs, and 314 biologics). Its high throughput stems from the sparse L1000 assay, which measures a pre-defined reduced set of landmark genes per sample via an array instead of performing bulk or single-cell RNA sequencing. This significantly reduces cost and increases throughput (1.3 million profiles). 978 “landmark” transcripts are measured (selected to span diverse pathways and be representative for co-expressed genes) – in addition to 80 invariant controls. This is done via ligation-mediated amplification (LMA) and bead-based fluorescent detection.108 In CMap, cells from a standardized nine-line panel are exposed to small molecules or genetic perturbations over multiple concentrations and short exposure times. Transcriptome-wide values for the 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 350 non-measured genes are inferred by regression trained on external RNA-sequencing compendia, with 9196 of 11
350 non-measured genes are inferred by regression trained on external RNA-sequencing compendia, with 9196 of 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 350 inferred genes (81%) achieving correlations in the top 95th percentile relative to their RNA-sequencing counterparts, indicating adequate performance.
350 inferred genes (81%) achieving correlations in the top 95th percentile relative to their RNA-sequencing counterparts, indicating adequate performance.
Target identification
Clustered Regularly Interspaced Short Palindromic Repeats114 (CRISPR) is a naturally occurring adaptive immune system in bacteria that has been repurposed as a programmable genome-editing technology and promises to help address the target discovery problem. Its key components are a Cas effector protein, most commonly Cas9, which acts as a nuclease, and a guide RNA (gRNA) that directs Cas9 to a specific genomic locus through base-pairing with the target DNA. CRISPR has revolutionized the scope and scale at which we can conduct genetic interventions to elucidate the biological function of proteins. Nowadays, CRISPR is heavily used in functional genomics applications to understand how gene-level perturbations, often obtained in high-throughput, pooled fashion, affect phenotypes.115 This has already led to the discovery of novel clinical targets.116 Traditionally, these screens have been conducted most frequently in cell lines due to their cost and scalability. In vivo CRISPR has recently emerged allowing for local genetic perturbations in the complete biological context of a living organism in a pooled fashion to test hundreds or thousands of target hypotheses.50 Perturbations are generally done at low transfection multiples such that most cells only obtain a single perturbation and perturbed cells are isolated by non-perturbed tissue to localize perturbation effects, acting as a proxy readout for the full organism (Trick 5). The perturbagen is tracked through barcodes. Phenotypic readouts e.g. single-cell transcriptomics, spatial profiling, imaging or others, enable the reconstruction of genotype–phenotype relationships within a physiologically relevant milieu. The startup Gordian Biotechnology is further leveraging perturbation-local single-cell sequencing readouts to predict in vivo disease responses for a given perturbation using paired disease-response and sequencing readouts.117
Hit Finding
McCloskey51et al. screened three targets (soluble epoxide hydrolase, estrogen receptor-α, c-KIT), trained classifiers purely on DEL selections (graph convolutional models trained on 355![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 804, 74
804, 74![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 741, and 50
741, and 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 186 positive training examples for sEH, ERα, and c-KIT), virtually screened 88 million catalog compounds, tested 2,000, and obtained a 30% hit rate at 30 µM with sub-10 nM actives for every target.
186 positive training examples for sEH, ERα, and c-KIT), virtually screened 88 million catalog compounds, tested 2,000, and obtained a 30% hit rate at 30 µM with sub-10 nM actives for every target.
Iqbal et al.121 targeted CK1α/δ with three differently sized libraries: MS (10 million), DD (11 million), HG (1 billion). They trained five models on orthosteric-binding site enriched outputs and screened a 140![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000-member test set. This yielded 808 compounds tested via SPR with 80 hits (9.9%), including two nanomolar (187 nM, 69.6 nM).
000-member test set. This yielded 808 compounds tested via SPR with 80 hits (9.9%), including two nanomolar (187 nM, 69.6 nM).
Wellnitz et al.99 screened WDR91 via HitGen OpenDEL (3 billion) and applied the resulting models on Enamine REAL (37 billion members). They nominated 50 molecules, tested 48, and confirmed 7 hits (14.5%, 2.7–21 µM) and generated co-crystal structures with WDR91.
The startup Leash Bio has scaled DEL against many targets and recently unveiled Hermes, a model achieving competitive binding prediction accuracy on non-DEL data despite exclusively being trained on DEL readouts.122
Acknowledgements
We thank Abhishaike Mahajan, Imran Haque, Hannah Wayment-Steele, Jess Ewald, Chaitanya Joshi, Ian Quigley and Martin Borch Jensen for thoughtful feedback.References
- Mastering the game of Go with deep neural networks and tree search | Nature. https://www.nature.com/articles/nature16961 (accessed 2026-01-03).
- Grandmaster level in StarCraft II using multi-agent reinforcement learning | Nature. https://www.nature.com/articles/s41586-019-1724-z (accessed 2026-01-03).
- D. Castelvecchi, DeepMind and OpenAI Models Solve Maths Problems at Level of Top Students, Nature, 2025, 644(8075), 20, DOI:10.1038/d41586-025-02343-x.
- M. Pacesa, L. Nickel, C. Schellhaas, J. Schmidt, E. Pyatova, L. Kissling, P. Barendse, J. Choudhury, S. Kapoor, A. Alcaraz-Serna, Y. Cho, K. H. Ghamary, L. Vinué, B. J. Yachnin, A. M. Wollacott, S. Buckley, A. H. Westphal, S. Lindhoud, S. Georgeon, C. A. Goverde, G. N. Hatzopoulos, P. Gönczy, Y. D. Muller, G. Schwank, D. C. Swarts, A. J. Vecchio, B. L. Schneider, S. Ovchinnikov and B. E. Correia, One-Shot Design of Functional Protein Binders with BindCraft, Nature, 2025, 1–10, DOI:10.1038/s41586-025-09429-6.
- L. Naef and M. Bronstein, The Road to Biology 2.0 Will Pass Through Black-Box Data. TDS Archive. https://medium.com/data-science/the-road-to-biology-2-0-will-pass-through-black-box-data-bbd00fabf959 (accessed 2025-09-24) Search PubMed.
- M. Gao and J. Skolnick, Structural Space of Protein-Protein Interfaces Is Degenerate, Close to Complete, and Highly Connected, Proc. Natl. Acad. Sci. U. S. A., 2010, 107(52), 22517–22522, DOI:10.1073/pnas.1012820107.
- S.-J. Chen, M. Hassan, R. L. Jernigan, K. Jia, D. Kihara, A. Kloczkowski, S. Kotelnikov, D. Kozakov, J. Liang, A. Liwo, S. Matysiak, J. Meller, C. Micheletti, J. C. Mitchell, S. Mondal, R. Nussinov, K. Okazaki, D. Padhorny, J. Skolnick, T. R. Sosnick, G. Stan, I. Vakser, X. Zou and G. D. Rose, Protein Folds vs. Protein Folding: Differing Questions, Different Challenges, Proc. Natl. Acad. Sci. U. S. A., 2023, 120(1), e2214423119, DOI:10.1073/pnas.2214423119.
- C. O. Mackenzie, J. Zhou and G. Grigoryan, Tertiary Alphabet for the Observable Protein Structural Universe, Proc. Natl. Acad. Sci. U. S. A., 2016, 113(47), E7438–E7447, DOI:10.1073/pnas.1607178113.
- J. Yeo, Y. Han, N. Bordin, A. M. Lau, S. M. Kandathil, H. Kim, E. L. Karin, M. Mirdita, D. T. Jones, C. Orengo, M. SteineggerMetagenomic-Scale Analysis of the Predicted Protein Structure Universe, bioRxiv, 2025, preprint, DOI:10.1101/2025.04.23.650224.
- J. Durairaj, Y. Adeshina, Z. Cao, X. Zhang, V. Oleinikovas, T. Duignan, Z. McClure, X. Robin, D. Kovtun, E. Rossi, G. Zhou, S. Veccham, C. Isert, Y. Peng, P. Sundareson, M. Akdel, G. Corso, H. Stärk, Z. Carpenter, M. Bronstein, E. Kucukbenli, T. Schwede and L. NaefPLINDER: The Protein-Ligand Interactions Dataset and Evaluation Resource, bioRxiv, 2024, preprint, DOI:10.1101/2024.07.17.603955.
- D. Kovtun, M. Akdel, A. Goncearenco, G. Zhou, G. Holt, D. Baugher, D. Lin, Y. Adeshina, T. Castiglione, X. Wang, C. Marquet, M. McPartlon, T. Geffner, G. Corso, H. Stärk, Z. Carpenter, E. Kucukbenli, M. Bronstein and L. NaefPINDER: The Protein Interaction Dataset and Evaluation Resource, bioRxiv, 2024, preprint, DOI:10.1101/2024.07.17.603980.
- P. Škrinjar, J. Eberhardt, J. Durairaj and T. SchwedeHave Protein-Ligand Co-Folding Methods Moved beyond Memorisation?, bioRxiv, 2025, preprint, DOI:10.1101/2025.02.03.636309.
- E. DeBenedictis, How do we make the next PDB? Introducing Align's Data Platform, NeurIPS Machine Learning in Structural Biology Workshop, https://slideslive.com/embed/presentation/39031188?js_embed_version=3&embed_init_token=eyJhbGciOiJIUzI1NiJ9.eyJpYXQiOjE3NjE4MDAxNDksImV4cCI6MTc2MTkyOTc0OSwidSI6eyJ1dWlkIjoiYmExYTViNzYtZjM1Zi00NThlLTljYTAtOGY3OWI3ZDcyZDk2IiwiaSI6bnVsbCwiZSI6bnVsbCwibSI6ZmFsc2V9LCJkIjoibmV1cmlwcy5jYyJ9.emkI0Ur-Pg-TPByWfqZPE406jDNCy55AFvlhHlf3q_w, accessed 2025-10-29 Search PubMed.
- Protein Data Bank Marks 50 Years Helping to Unlock Mysteries of Human Disease, Rutgers University, https://www.rutgers.edu/news/protein-data-bank-marks-50-years-lifesaving-scientific-collaboration-and-coordination, accessed 2025-09-24 Search PubMed.
- A. Krizhevsky, I. Sutskever and G. E. HintonImageNet Classification with Deep Convolutional Neural Networks, in Advances in Neural Information Processing Systems, Curran Associates, Inc., 2012, vol. 25 Search PubMed.
- A. Karpathy, Software 2.0. Medium, https://karpathy.medium.com/software-2-0-a64152b37c35, accessed 2025-09-24 Search PubMed.
- R. Sutton, The Bitter Lesson. The Bitter Lesson, https://www.cs.utexas.edu/~eunsol/courses/data/bitter_lesson.pdf, accessed 2025-09-24.
- A. W. Senior, R. Evans, J. Jumper, J. Kirkpatrick, L. Sifre, T. Green, C. Qin, A. Žídek, A. W. R. Nelson, A. Bridgland, H. Penedones, S. Petersen, K. Simonyan, S. Crossan, P. Kohli, D. T. Jones, D. Silver, K. Kavukcuoglu and D. Hassabis, Protein Structure Prediction Using Multiple Deep Neural Networks in the 13th Critical Assessment of Protein Structure Prediction (CASP13), Proteins:Struct., Funct., Bioinf., 2019, 87(12), 1141–1148, DOI:10.1002/prot.25834.
- J. Jumper, R. Evans, A. Pritzel, T. Green, M. Figurnov, O. Ronneberger, K. Tunyasuvunakool, R. Bates, A. Žídek, A. Potapenko, A. Bridgland, C. Meyer, S. A. A. Kohl, A. J. Ballard, A. Cowie, B. Romera-Paredes, S. Nikolov, R. Jain, J. Adler, T. Back, S. Petersen, D. Reiman, E. Clancy, M. Zielinski, M. Steinegger, M. Pacholska, T. Berghammer, S. Bodenstein, D. Silver, O. Vinyals, A. W. Senior, K. Kavukcuoglu, P. Kohli and D. Hassabis, Highly Accurate Protein Structure Prediction with AlphaFold, Nature, 2021, 596(7873), 583–589, DOI:10.1038/s41586-021-03819-2.
- J. Abramson, J. Adler, J. Dunger, R. Evans, T. Green, A. Pritzel, O. Ronneberger, L. Willmore, A. J. Ballard, J. Bambrick, S. W. Bodenstein, D. A. Evans, C.-C. Hung, M. O'Neill, D. Reiman, K. Tunyasuvunakool, Z. Wu, A. Žemgulytė, E. Arvaniti, C. Beattie, O. Bertolli, A. Bridgland, A. Cherepanov, M. Congreve, A. I. Cowen-Rivers, A. Cowie, M. Figurnov, F. B. Fuchs, H. Gladman, R. Jain, Y. A. Khan, C. M. R. Low, K. Perlin, A. Potapenko, P. Savy, S. Singh, A. Stecula, A. Thillaisundaram, C. Tong, S. Yakneen, E. D. Zhong, M. Zielinski, A. Žídek, V. Bapst, P. Kohli, M. Jaderberg, D. Hassabis and J. M. Jumper, Accurate Structure Prediction of Biomolecular Interactions with AlphaFold 3, Nature, 2024, 1–3, DOI:10.1038/s41586-024-07487-w.
- Z. Lin, H. Akin, R. Rao, B. Hie, Z. Zhu, W. Lu, N. Smetanin, R. Verkuil, O. Kabeli, Y. Shmueli, A. Costa, S. dos, M. Fazel-Zarandi, T. Sercu, S. Candido and A. Rives, Evolutionary-Scale Prediction of Atomic Level Protein Structure with a Language Model, , bioRxiv, , 2022, preprint, DOI:10.1101/2022.07.20.500902.
- Y. Wang, J. Lu, N. Jaitly, J. Susskind and M. A. Bautista, SimpleFold: Folding Proteins Is Simpler than You Think, arXiv, prperint, arXiv:2509.18480, DOI:10.48550/arXiv.2509.18480.
- T. Hayes, R. Rao, H. Akin, N. J. Sofroniew, D. Oktay, Z. Lin, R. Verkuil, V. Q. Tran, J. Deaton, M. Wiggert, R. Badkundri, I. Shafkat, J. Gong, A. Derry, R. S. Molina, N. Thomas, Y. Khan, C. Mishra, C. Kim, L. J. Bartie, M. Nemeth, P. D. Hsu, T. Sercu, S. Candido and A. Rives, Simulating 500 Million Years of Evolution with a Language Model, bioRxiv, 2024, preprint, DOI:10.1101/2024.07.01.600583.
- S. E. Antonsson and P. Melsted, Batch Correction Methods Used in Single Cell RNA-Sequencing Analyses Are Often Poorly Calibrated, bioRxiv, 2024, preprint, DOI:10.1101/2024.03.19.585562.
- A. Spinner, E. DeBenedictis and C. M. Hudson, Scaling and Data Saturation in Protein Language Models, arXiv, 2025, preprint, DOI:10.48550/arXiv.2507.22210.
- B. Zdrazil, Fifteen Years of ChEMBL and Its Role in Cheminformatics and Drug Discovery, J. Cheminf., 2025, 17(1), 32, DOI:10.1186/s13321-025-00963-z.
- S. K. Burley, R. Bhatt, C. Bhikadiya, C. Bi, A. Biester, P. Biswas, S. Bittrich, S. Blaumann, R. Brown, H. Chao, V. R. Chithari, P. A. Craig, G. V. Crichlow, J. M. Duarte, S. Dutta, Z. Feng, J. W. Flatt, S. Ghosh, D. S. Goodsell, R. K. Green, V. Guranovic, J. Henry, B. P. Hudson, M. Joy, J. T. Kaelber, I. Khokhriakov, J.-S. Lai, C. L. Lawson, Y. Liang, D. Myers-Turnbull, E. Peisach, I. Persikova, D. W. Piehl, A. Pingale, Y. Rose, J. Sagendorf, A. Sali, J. Segura, M. Sekharan, C. Shao, J. Smith, M. Trumbull, B. Vallat, M. Voigt, B. Webb, S. Whetstone, A. Wu-Wu, T. Xing, J. Y. Young, A. Zalevsky and C. Zardecki, Updated Resources for Exploring Experimentally-Determined PDB Structures and Computed Structure Models at the RCSB Protein Data Bank, Nucleic Acids Res., 2025, 53(D1), D564–D574, DOI:10.1093/nar/gkae1091.
- L. Martens, H. Hermjakob, P. Jones, M. Adamski, C. Taylor, D. States, K. Gevaert, J. Vandekerckhove and R. Apweiler, PRIDE: The Proteomics Identifications Database, Proteomics, 2005, 5(13), 3537–3545, DOI:10.1002/pmic.200401303.
- E. Clough and T. Barrett, The Gene Expression Omnibus Database, Methods Mol Biol., 2016, 1418, 93–110, DOI:10.1007/978-1-4939-3578-9_5.
- G. Koscielny, P. An, D. Carvalho-Silva, J. A. Cham, L. Fumis, R. Gasparyan, S. Hasan, N. Karamanis, M. Maguire, E. Papa, A. Pierleoni, M. Pignatelli, T. Platt, F. Rowland, P. Wankar, A. P. Bento, T. Burdett, A. Fabregat, S. Forbes, A. Gaulton, C. Y. Gonzalez, H. Hermjakob, A. Hersey, S. Jupe, Ş. Kafkas, M. Keays, C. Leroy, F.-J. Lopez, M. P. Magarinos, J. Malone, J. McEntyre, A. Munoz-Pomer Fuentes, C. O'Donovan, I. Papatheodorou, H. Parkinson, B. Palka, J. Paschall, R. Petryszak, N. Pratanwanich, S. Sarntivijal, G. Saunders, K. Sidiropoulos, T. Smith, Z. Sondka, O. Stegle, Y. A. Tang, E. Turner, B. Vaughan, O. Vrousgou, X. Watkins, M.-J. Martin, P. Sanseau, J. Vamathevan, E. Birney, J. Barrett and I. Dunham, Open Targets: A Platform for Therapeutic Target Identification and Validation, Nucleic Acids Res., 2017, 45(Database issue), D985–D994, DOI:10.1093/nar/gkw1055.
- R. Ferreira de Freitas and M. A. Schapira, Systematic Analysis of Atomic Protein–Ligand Interactions in the PDB, MedChemComm, 2017, 8(10), 1970–1981, 10.1039/C7MD00381A.
- G. Ahdritz, N. Bouatta, S. Kadyan, Q. Xia, W. Gerecke, T. J. O'Donnell, D. Berenberg, I. Fisk, N. Zanichelli, B. Zhang, A. Nowaczynski, B. Wang, M. M. Stepniewska-Dziubinska, S. Zhang, A. Ojewole, M. E. Guney, S. Biderman, A. M. Watkins, S. Ra, P. R. Lorenzo, L. Nivon, B. Weitzner, Y.-E. A. Ban, P. K. Sorger, E. Mostaque, Z. Zhang, R. Bonneau and M. AlQuraishi, OpenFold: Retraining AlphaFold2 Yields New Insights into Its Learning Mechanisms and Capacity for Generalization, bioRxiv, 2022, preprint, DOI:10.1101/2022.11.20.517210.
- D. Rolnick, A. Veit, S. Belongie and N. Shavit, Deep Learning Is Robust to Massive Label Noise, arXiv, 2018, preprint, arXiv:1705.10694, DOI:10.48550/arXiv.1705.10694.
- When Quantity Becomes Quality with Aviv Regev, Andreessen Horowitz. https://a16z.com/podcast/when-quantity-becomes-quality-with-aviv-regev/, accessed 2025-10-27.
- J. Dodge, M. Sap, A. Marasović, W. Agnew, G. Ilharco, D. Groeneveld, M. Mitchell and M. Gardner, Documenting Large Webtext Corpora: A Case Study on the Colossal Clean Crawled Corpus, arXiv, 2021, arXiv:2104.08758, DOI:10.48550/arXiv.2104.08758.
- N. Lambert, Reinforcement Learning from Human Feedback, arXiv, 2025, arXiv:2504.12501, DOI:10.48550/arXiv.2504.12501.
- C. K. Joshi, X. Fu, Y.-L. Liao, V. Gharakhanyan, B. K. Miller, A. Sriram and Z. W. Ulissi, All-Atom Diffusion Transformers: Unified Generative Modelling of Molecules and Materials, arXiv, 2025, preprint, arXiv:2503.03965, DOI:10.48550/arXiv.2503.03965.
- J. Ho, A. Jain and P. Abbeel, Denoising Diffusion Probabilistic Models, arXiv, 2020, preprint, arXiv:2006.11239, DOI:10.48550/arXiv.2006.11239.
- G. Daras, K. Shah, Y. Dagan, A. Gollakota, A. G. Dimakis and A. Klivans. Ambient Diffusion: Learning Clean Distributions from Corrupted Data, arXiv, 2023, preprint, arXiv;2305.19256, DOI:10.48550/arXiv.2305.19256.
- R. Rosenthal, The File Drawer Problem and Tolerance for Null Results, Psychol. Bull., 1979, 86(3), 638–641, DOI:10.1037/0033-2909.86.3.638.
- M. P. Maloney, C. W. Coley, S. Genheden, N. Carson, P. Helquist, P.-O. Norrby and O. Wiest, Negative Data in Data Sets for Machine Learning Training, J. Org. Chem., 2023, 88(9), 5239–5241, DOI:10.1021/acs.joc.3c00844.
- R. P. Thomas, R. E. Heap, F. Zappacosta, E. K. Grant, P. Pogány, S. Besley, D. J. Fallon, M. M. Hann, D. House, N. C. O. Tomkinson and J. T. Bush, A Direct-to-Biology High-Throughput Chemistry Approach to Reactive Fragment Screening, Chem. Sci., 2021, 12(36), 12098–12106, 10.1039/D1SC03551G.
- C. Y. Cheng, W. Kladwang, J. D. Yesselman and R. Das, RNA Structure Inference through Chemical Mapping after Accidental or Intentional Mutations, Proc. Natl. Acad. Sci. U. S. A., 2017, 114(37), 9876–9881, DOI:10.1073/pnas.1619897114.
- A. Vaswani, N. Shazeer, N. Parmar, J. Uszkoreit, L. Jones, A. N. Gomez, Ł. Kaiser and I. Polosukhin, Attention Is All You Need, in Advances in Neural Information Processing Systems, ed. Guyon, I., Luxburg, U. V., Bengio, S., Wallach, H., Fergus, R., Vishwanathan, S., Garnett, R., Curran Associates, Inc., 2017, vol. 30 Search PubMed.
- E. Candes, J. Romberg, T. Tao, Stable Signal Recovery from Incomplete and Inaccurate Measurements, arXiv, 2005, preprint, arXiv:math/0503066, DOI:10.48550/arXiv.math/0503066.
- M. Lustig, D. Donoho and J. M. Pauly, Sparse MRI: The Application of Compressed Sensing for Rapid MR Imaging, Magn. Reson. Med., 2007, 58(6), 1182–1195, DOI:10.1002/mrm.21391.
- G. L. Stüber, Principles of Mobile Communication, Springer International Publishing, Cham, 2017, DOI:10.1007/978-3-319-55615-4.
- J. C. Venter, M. D. Adams, G. G. Sutton, A. R. Kerlavage, H. O. Smith and M. Hunkapiller, Shotgun Sequencing of the Human Genome, Science, 1998, 280(5369), 1540–1542, DOI:10.1126/science.280.5369.1540.
- K. Tsuboyama, J. Dauparas, J. Chen, E. Laine, Y. Mohseni Behbahani, J. J. Weinstein, N. M. Mangan, S. Ovchinnikov and G. J. Rocklin, Mega-Scale Experimental Analysis of Protein Folding Stability in Biology and Design, Nature, 2023, 620(7973), 434–444, DOI:10.1038/s41586-023-06328-6.
- A. J. Santinha, A. Strano and R. J. Platt, Methods and Applications of in Vivo CRISPR Screening, Nat. Rev. Genet., 2025, 26(10), 702–718, DOI:10.1038/s41576-025-00873-8.
- K. McCloskey, E. A. Sigel, S. Kearnes, L. Xue, X. Tian, D. Moccia, D. Gikunju, S. Bazzaz, B. Chan, M. A. Clark, J. W. Cuozzo, M.-A. Guié, J. P. Guilinger, C. Huguet, C. D. Hupp, A. D. Keefe, C. J. Mulhern, Y. Zhang and P. Riley, Machine Learning on DNA-Encoded Libraries: A New Paradigm for Hit Finding, J. Med. Chem., 2020, 63(16), 8857–8866, DOI:10.1021/acs.jmedchem.0c00452.
- J. D. Ewald, K. L. Titterton, A. Bäuerle, A. Beatson, D. A. Boiko, Á. A. Cabrera, J. Cheah, B. A. Cimini, B. Gorissen, T. Jones, K. J. Karczewski, D. Rouquie, S. Seal, E. Weisbart, B. White, A. E. Carpenter and S. Singh, Cell Painting for Cytotoxicity and Mode-of-Action Analysis in Primary Human Hepatocytes, bioRxiv, 2025, preprint, DOI:10.1101/2025.01.22.634152.
- A. Subramanian, R. Narayan, S. M. Corsello, D. D. Peck, T. E. Natoli, X. Lu, J. Gould, J. F. Davis, A. A. Tubelli, J. K. Asiedu, D. L. Lahr, J. E. Hirschman, Z. Liu, M. Donahue, B. Julian, M. Khan, D. Wadden, I. C. Smith, D. Lam, A. Liberzon, C. Toder, M. Bagul, M. Orzechowski, O. M. Enache, F. Piccioni, S. A. Johnson, N. J. Lyons, A. H. Berger, A. F. Shamji, A. N. Brooks, A. Vrcic, C. Flynn, J. Rosains, D. Y. Takeda, R. Hu, D. Davison, J. Lamb, K. Ardlie, L. Hogstrom, P. Greenside, N. S. Gray, P. A. Clemons, S. Silver, X. Wu, W.-N. Zhao, W. Read-Button, X. Wu, S. J. Haggarty, L. V. Ronco, J. S. Boehm, S. L. Schreiber, J. G. Doench, J. A. Bittker, D. E. Root, B. Wong and T. R. Golub, A Next Generation Connectivity Map: L1000 Platform and the First 1,000,000 Profiles, Cell, 2017, 171(6), 1437–1452.e17, DOI:10.1016/j.cell.2017.10.049.
- L. Konieczny, I. Roterman-Konieczna and P. Spólnik, The Structure and Function of Living Organisms, in Systems Biology: Functional Strategies of Living Organisms, Springer, 2nd edn, 2023, DOI:10.1007/978-3-031-31557-2_1.
- F. J. O'Reilly and J. Rappsilber, Cross-Linking Mass Spectrometry: Methods and Applications in Structural, Molecular and Systems Biology, Nat. Struct. Mol. Biol., 2018, 25(11), 1000–1008, DOI:10.1038/s41594-018-0147-0.
- M. A. Clasen, M. Ruwolt, C. Wang, J. Ruta, B. Bogdanow, L. U. Kurt, Z. Zhang, S. Wang, F. C. Gozzo, T. Chen, P. C. Carvalho, D. B. Lima and F. Liu, Proteome-Scale Recombinant Standards and a Robust High-Speed Search Engine to Advance Cross-Linking MS-Based Interactomics, Nat. Methods, 2024, 21(12), 2327–2335, DOI:10.1038/s41592-024-02478-1.
- W. S. Bush, Introduction to Bioinformatics and Computational Biology, in Proceedings of the fourteenth international conference on Genetic and evolutionary computation conference companion - GECCO Companion ’12, ACM Press, Philadelphia, Pennsylvania, USA, 2012, p 1141, DOI:10.1145/2330784.2330935.
- K. Stahl, A. Graziadei, T. Dau, O. Brock and J. Rappsilber, Protein Structure Prediction with In-Cell Photo-Crosslinking Mass Spectrometry and Deep Learning, Nat. Biotechnol., 2023, 41(12), 1810–1819, DOI:10.1038/s41587-023-01704-z.
- K. Gilep, A. Obarska-Kosinska and J. Kosinski, Improving AlphaFold 3 Structural Modeling by Incorporating Explicit Crosslinks, bioRxiv, 2025, preprint, DOI:10.1101/2024.12.03.626671.
- C. Isert, M. Pun, E. Rossi, D. Tischer, M. Akdel, D. Kovtun, M. Pegoraro, T. Duignan, A. Zhang, V. Oleinikovas, G. Holt, Y. Adeshina, P. Kunzmann, A. Ramesh, D. Wu, A. Goncearenco, L. Foguel, D. Felker, D. Sabbadin, V. Lam, M. Grass, Z. Carpenter, M. Bronstein and L. Naef, NEO-1 - Decode and Design the Structure of Life. NEO-1 - Decode and Design the Structure of Life, https://www.proximabio.com/neo-1, accessed 2025-09-24.
- D. S. Wilson, A. D. Keefe and J. W. Szostak, The Use of mRNA Display to Select High-Affinity Protein-Binding Peptides, Proc. Natl. Acad. Sci. U. S. A., 2001, 98(7), 3750–3755, DOI:10.1073/pnas.061028198.
- A. Brückner, C. Polge, N. Lentze, D. Auerbach and U. Schlattner, Yeast Two-Hybrid, a Powerful Tool for Systems Biology, Int. J. Mol. Sci., 2009, 10(6), 2763–2788, DOI:10.3390/ijms10062763.
- S. Lewis, T. Hempel, J. Jiménez-Luna, M. Gastegger, Y. Xie, A. Y. K. Foong, V. G. Satorras, O. Abdin, B. S. Veeling, I. Zaporozhets, Y. Chen, S. Yang, A. Schneuing, J. Nigam, F. Barbero, V. Stimper, A. Campbell, J. Yim, M. Lienen, Y. Shi, S. Zheng, H. Schulz, U. Munir, C. Clementi and F. Noé, Scalable Emulation of Protein Equilibrium Ensembles with Generative Deep Learning. , bioRxiv, 2024, preprint, DOI:10.1101/2024.12.05.626885.
- R. C. Kretsch, A. M. Hummer, S. He, R. Yuan, J. Zhang, T. Karagianes, Q. Cong, A. Kryshtafovych and R. Das, Assessment of Nucleic Acid Structure Prediction in CASP16, bioRxiv, 2025, preprint, DOI:10.1101/2025.05.06.652459.
- E. Laine, S. Grudinin, R. Klypa and I. C. de. Beauchêne, Navigating Protein–Nucleic Acid Sequence-Structure Landscapes with Deep Learning, Curr. Opin. Struct. Biol., 2025, 95, 103162, DOI:10.1016/j.sbi.2025.103162.
- P. Cordero, W. Kladwang, C. C. VanLang and R. Das, The Mutate-and-Map Protocol for Inferring Base Pairs in Structured RNA, Methods Mol Biol., 2014, 1086, 53–77, DOI:10.1007/978-1-62703-667-2_4.
- C. Y. Cheng, F.-C. Chou, W. Kladwang, S. Tian, P. Cordero and R. Das, Consistent Global Structures of Complex RNA States through Multidimensional Chemical Mapping, eLife, 2015, 4, e07600, DOI:10.7554/eLife.07600.
- T. Wu, A. Y. Cheng, Y. Zhang, J. Xu, J. Wu, L. Wen, X. Li, B. Liu, X. Dou, P. Wang, L. Zhang, J. Fei, J. Li, Z. Ouyang and C. He, KARR-Seq Reveals Cellular Higher-Order RNA Structures and RNA–RNA Interactions, Nat. Biotechnol., 2024, 42(12), 1909–1920, DOI:10.1038/s41587-023-02109-8.
- S. He, R. Huang, J. Townley, R. C. Kretsch, T. G. Karagianes, D. B. T. Cox, H. Blair, D. Penzar, V. Vyaltsev, E. Aristova, A. Zinkevich, A. Bakulin, H. Sohn, D. Krstevski, T. Fukui, F. Tatematsu, Y. Uchida, D. Jang, J. S. Lee, R. Shieh, T. Ma, E. Martynov, M. V. Shugaev, H. S. T. Bukhari, K. Fujikawa, K. Onodera, C. Henkel, S. Ron, J. Romano, J. J. Nicol, G. P. Nye, Y. Wu, C. Choe, W. Reade, E. Participants and R. Das, Ribonanza: Deep Learning of RNA Structure through Dual Crowdsourcing, bioRxiv, 2024, preprint, DOI:10.1101/2024.02.24.581671.
- S. He, C. organizers, C. R. experimentalists, R.-P. consortium, V. team, R. Kretsch, A. Hummer, A. Favor, W. Reade, M. Demkin, R. Das, others, Stanford RNA 3D Folding, 2025, https://kaggle.com/competitions/stanford-rna-3d-folding.
- RibonanzaNet2 alpha release, https://www.kaggle.com/competitions/stanford-rna-3d-folding/discussion/571704, accessed 2025-12-14 Search PubMed.
- Y. Lee, S. He, T. Oda, G. J. Rao, Y. Kim, R. Kim, H. Kim, C. K. Heng, D. Kowerko, H. Li, H. Nguyen, A. Sampathkumar, R. E. Gómez, M. Chen, A. Yoshizawa, S. Kuraishi, K. Ogawa, S. Zou, A. Paullier, B. Zhao, H.-L. Chen, T.-A. Hsu, T. Hirano, W. Chiu, J. G. Gezelle, D. Haack, Y. Hong, S. Jadhav, D. Koirala, R. C. Kretsch, A. Lewicka, S. Li, M. Marcia, J. Piccirilli, B. Rudolfs, Y. Srivastava, A.-L. Steckelberg, Z. Su, N. Toor, L. Wang, Z. Yang, K. Zhang, J. Zou, D. Baker, S.-J. Chen, M. Demkin, A. Favor, A. M. Hummer, C. K. Joshi, A. Kryshtafovych, E. Küçükbenli, Z. Miao, J. Moult, C. Munley, W. Reade, T. Viel, E. Westhof, S. Zhang and R. Das, Template-Based RNA Structure Prediction Advanced through a Blind Code Competition, bioRxiv, 2025, preprint, DOI:10.64898/2025.12.30.696949.
- C. K. Joshi, E. Gianni, S. L. Y. Kwok, S. V. Mathis, P. Liò and P. HolligerGenerative Inverse Design of RNA Structure and Function with gRNAde, bioRxiv, 2025, preprint, DOI:10.1101/2025.11.29.691298.
- M. Thompson, M. Martín, T. S. Olmo, C. Rajesh, P. K. Koo, B. Bolognesi and B. Lehner, Massive Experimental Quantification Allows Interpretable Deep Learning of Protein Aggregation, Sci. Adv., 2025, 11(18), eadt5111, DOI:10.1126/sciadv.adt5111.
- M.-A. Bray, S. Singh, H. Han, C. T. Davis, B. Borgeson, C. Hartland, M. Kost-Alimova, S. M. Gustafsdottir, C. C. Gibson and A. E. Carpenter, Cell Painting, a High-Content Image-Based Assay for Morphological Profiling Using Multiplexed Fluorescent Dyes, Nat. Protoc., 2016, 11(9), 1757–1774, DOI:10.1038/nprot.2016.105.
- N. H. Lazar, S. Celik, L. Chen, M. M. Fay, J. C. Irish, J. Jensen, C. A. Tillinghast, J. Urbanik, W. P. Bone, C. C. Gibson and I. S. Haque, High-Resolution Genome-Wide Mapping of Chromosome-Arm-Scale Truncations Induced by CRISPR–Cas9 Editing, Nat. Genet., 2024, 56(7), 1482–1493, DOI:10.1038/s41588-024-01758-y.
- R. E. White and P. Manitpisitkul, Pharmacokinetic Theory of Cassette Dosing in Drug Discovery Screening, Drug Metab. Dispos. Biol. Fate Chem., 2001, 29(7), 957–966 CAS.
- A. M. Richard, R. S. Judson, K. A. Houck, C. M. Grulke, P. Volarath, I. Thillainadarajah, C. Yang, J. Rathman, M. T. Martin, J. F. Wambaugh, T. B. Knudsen, J. Kancherla, K. Mansouri, G. Patlewicz, A. J. Williams, S. B. Little, K. M. Crofton and R. S. Thomas, ToxCast Chemical Landscape: Paving the Road to 21st Century Toxicology, Chem. Res. Toxicol., 2016, 29(8), 1225–1251, DOI:10.1021/acs.chemrestox.6b00135.
- L.-J. Gardiner, A. P. Carrieri, J. Wilshaw, S. Checkley, E. O. Pyzer-Knapp and R. Krishna, Using Human in Vitro Transcriptome Analysis to Build Trustworthy Machine Learning Models for Prediction of Animal Drug Toxicity, Sci. Rep., 2020, 10(1), 9522, DOI:10.1038/s41598-020-66481-0.
- D. G. Belair, R. J. Visconti, M. Hong, M. Marella, M. F. Peters, C. W. Scott and K. L. Kolaja, Human Ileal Organoid Model Recapitulates Clinical Incidence of Diarrhea Associated with Small Molecule Drugs, Toxicol. In Vitro, 2020, 68, 104928, DOI:10.1016/j.tiv.2020.104928.
- Y. Ma, Z. Zhang, B. Jia and Y. Yuan, Automated High-Throughput DNA Synthesis and Assembly, Heliyon, 2024, 10(6), e26967, DOI:10.1016/j.heliyon.2024.e26967.
- A. J. Zhang and K. Burgess, Total Syntheses of Vancomycin, Angew Chem. Int. Ed. Engl., 1999, 38(5), 634–636, DOI:10.1002/(SICI)1521-3773(19990301)38:5%253C634::AID-ANIE634%253E3.0.CO;2-G.
- M. Jones and R. L. Goodyear, High-Throughput Purification in Drug Discovery: Scaling New Heights of Productivity, ACS Med. Chem. Lett., 2023, 14(7), 916–919, DOI:10.1021/acsmedchemlett.3c00073.
- W. McCorkindale, M. Filep, N. London, A. A. Lee and E. King-Smith, Deconvoluting Low Yield from Weak Potency in Direct-to-Biology Workflows with Machine Learning, RSC Med. Chem., 2024, 15(3), 1015–1021, 10.1039/D3MD00719G.
- D. C. Rees, M. Congreve, C. W. Murray and R. Carr, Fragment-Based Lead Discovery, Nat. Rev. Drug Discovery, 2004, 3(8), 660–672, DOI:10.1038/nrd1467.
- D. Fearon, A. Powell, A. Douangamath, A. Dias, C. W. E. Tomlinson, B. H. Balcomb, J. C. Aschenbrenner, A. Aimon, I. A. Barker, F. Bertram, J. Brandão-Neto, P. A. Coe, P. Collins, L. E. Dunnett, M. Fairhead, R. J. Gildea, M. Golding, T. Gorrie-Stone, P. V. Hathaway, L. Koekemoer, T. Krojer, R. M. Lithgo, E. M. Maclean, P. G. Marples, H. Mikolajek, X. Ni, K. H. V. Nidamarthi, G. O'Donnell, R. Skyner, R. Talon, W. Thompson, G. Watt, C. F. Wild, M. A. Williams, M. Winokan, N. D. Wright, G. Winter, E. J. Shotton and F. von Delft, Accelerating Drug Discovery With High-Throughput Crystallographic Fragment Screening and Structural Enablement, Appl. Res., 2025, 4(1), e202400192, DOI:10.1002/appl.202400192.
- H. Grosjean, K. Fieseler, R. Sanchez-Garcia, W. Thompson, C. M. Deane, F. von Delft and P. C. Biggin, Structure-Activity-Relationships Can Be Directly Extracted from High-Throughput Crystallographic Evaluation of Fragment Elaborations in Crude Reaction Mixtures, ChemRxiv, 2025, preprint, DOI:10.26434/chemrxiv-2025-bg2ll.
- OpenBind – Diamond Light Source, https://www.diamond.ac.uk/Science/Collaborations/openbind.html, accessed 2025-11-08 Search PubMed.
- C. E. Hendrick, J. R. Jorgensen, C. Chaudhry, I. I. Strambeanu, J.-F. Brazeau, J. Schiffer, Z. Shi, J. D. Venable and S. E. Wolkenberg, Direct-to-Biology Accelerates PROTAC Synthesis and the Evaluation of Linker Effects on Permeability and Degradation, ACS Med. Chem. Lett., 2022, 13(7), 1182–1190, DOI:10.1021/acsmedchemlett.2c00124.
- Science and Tech, https://www.kimiatx.com/science-tech, accessed 2025-10-28.
- B. A. Lanman, J. R. Allen, J. G. Allen, A. K. Amegadzie, K. S. Ashton, S. K. Booker, J. J. Chen, N. Chen, M. J. Frohn, G. Goodman, D. J. Kopecky, L. Liu, P. Lopez, J. D. Low, V. Ma, A. E. Minatti, T. T. Nguyen, N. Nishimura, A. J. Pickrell, A. B. Reed, Y. Shin, A. C. Siegmund, N. A. Tamayo, C. M. Tegley, M. C. Walton, H.-L. Wang, R. P. Wurz, M. Xue, K. C. Yang, P. Achanta, M. D. Bartberger, J. Canon, L. S. Hollis, J. D. McCarter, C. Mohr, K. Rex, A. Y. Saiki, T. San Miguel, L. P. Volak, K. H. Wang, D. A. Whittington, S. G. Zech, J. R. Lipford and V. J. Cee, Discovery of a Covalent Inhibitor of KRASG12C (AMG 510) for the Treatment of Solid Tumors, J. Med. Chem., 2020, 63(1), 52–65, DOI:10.1021/acs.jmedchem.9b01180.
- A. M. Zahm, W. S. Owens, S. R. Himes, K. E. Rondem, B. S. Fallon, A. N. Gormick, J. S. Bloom, S. Kosuri, H. Chan and J. G. English, Discovery and Validation of Context-Dependent Synthetic Mammalian Promoters, bioRxiv, 2023, preprint, DOI:10.1101/2023.05.11.539703.
- Octant Blog: Miniaturizing Drug Discovery, https://www.octant.bio/blog-posts/miniaturizing-drug-discovery-using-high-throughput-chemistry, accessed 2025-10-02 Search PubMed.
- Octant Blog: Harnessing Multiplexing for Cellular Assay and Reporter Development, https://www.octant.bio/blog-posts/harnessing-multiplexing-for-cellular-assay-and-reporter-development, accessed 2025-10-02 Search PubMed.
- J. Cheng, G. Novati, J. Pan, C. Bycroft, A. Žemgulytė, T. Applebaum, A. Pritzel, L. H. Wong, M. Zielinski, T. Sargeant, R. G. Schneider, A. W. Senior, J. Jumper, D. Hassabis, P. Kohli and Ž. Avsec, Accurate Proteome-Wide Missense Variant Effect Prediction with AlphaMissense, Science, 2023, 381(6664), eadg7492, DOI:10.1126/science.adg7492.
- S. P. Meisburger, D. A. Case and N. Ando, Diffuse X-Ray Scattering from Correlated Motions in a Protein Crystal, Nat. Commun., 2020, 11(1), 1271, DOI:10.1038/s41467-020-14933-6.
- H. K. Wayment-Steele, G. E. Nesr, R. Hettiarachchi, H. Kariyawasam, S. Ovchinnikov and D. Kern, Learning Millisecond Protein Dynamics from What Is Missing in NMR Spectra, bioRxiv, 2025, preprint, DOI:10.1101/2025.03.19.642801.
- C. G. Northcutt, A. Athalye and J. Mueller, Pervasive Label Errors in Test Sets Destabilize Machine Learning Benchmarks, arXiv, 2021, preprint, arXiv:2103.14749, DOI:10.48550/arXiv.2103.14749.
- J. Wellnitz, S. Ahmad, N. Begale, J. Joseph, H. Zeng, A. Bolotokova, A. Dong, S. Reza, P. Ghiabi, G. Elisa, X. Cheng, G. Tu, X. Li, J. Liu, D. Dou, J. Li, R. J. Harding, A. M. Edwards, B. Haibe-Kains, L. Halabelian, A. Tropsha and R. Couñago, Enabling Open Machine Learning of DNA Encoded Library Selections to Accelerate the Discovery of Small Molecule Protein Binders, ChemRxiv, 2024, preprint, DOI:10.26434/chemrxiv-2024-xd385.
- O. Gujral, M. Bafna, E. Alm and B. Berger, Sparse Autoencoders Uncover Biologically Interpretable Features in Protein Language Model Representations, Proc. Natl. Acad. Sci. U. S. A., 2025, 122(34), e2506316122, DOI:10.1073/pnas.2506316122.
- Z. Zhang, H. K. Wayment-Steele, G. Brixi, H. Wang, M. D. Peraro, D. Kern and S. Ovchinnikov, Protein Language Models Learn Evolutionary Statistics of Interacting Sequence Motifs, bioRxiv, 2024, preprint, DOI:10.1101/2024.01.30.577970.
- N. Wang, C. Fang, M. Bissell, A. Jain and D. Balsam, 2026, Using Interpretability to Identify a Novel Class of Alzheimer's Biomarkers, https://www.goodfire.ai/research/interpretability-for-alzheimers-detection, accessed 2026-01-30.
- U. Göbel, C. Sander, R. Schneider and A. Valencia, Correlated Mutations and Residue Contacts in Proteins, Proteins, 1994, 18(4), 309–317, DOI:10.1002/prot.340180402.
- S. P. Meisburger, D. A. Case and N. Ando, Robust Total X-Ray Scattering Workflow to Study Correlated Motion of Proteins in Crystals, Nat. Commun., 2023, 14(1), 1228, DOI:10.1038/s41467-023-36734-3.
- S. Chou, From systems operators to systems architects, Seemay’s blog. https://seemay.substack.com/p/from-systems-operators-to-systems, accessed 2025-09-24 Search PubMed.
- A. Beltran, X. Jiang, Y. Shen and B. Lehner, Site-Saturation Mutagenesis of 500 Human Protein Domains, Nature, 2025, 637(8047), 885–894, DOI:10.1038/s41586-024-08370-4.
- J. Meier, R. Rao, R. Verkuil, J. Liu, T. Sercu and A. Rives, Language Models Enable Zero-Shot Prediction of the Effects of Mutations on Protein Function, bioRxiv, 2021, preprint, DOI:10.1101/2021.07.09.450648.
- D. Peck, E. D. Crawford, K. N. Ross, K. Stegmaier, T. R. Golub and J. Lamb, A Method for High-Throughput Gene Expression Signature Analysis, Genome Biol., 2006, 7(7), R61, DOI:10.1186/gb-2006-7-7-r61.
- C. H. Emmerich, L. M. Gamboa, M. C. J. Hofmann, M. Bonin-Andresen, O. Arbach, P. Schendel, B. Gerlach, K. Hempel, A. Bespalov, U. Dirnagl and M. J. Parnham, Improving Target Assessment in Biomedical Research: The GOT-IT Recommendations, Nat. Rev. Drug Discovery, 2021, 20(1), 64–81, DOI:10.1038/s41573-020-0087-3.
- L. M. McNamee, M. J. Walsh and F. D. Ledley, Timelines of Translational Science: From Technology Initiation to FDA Approval, PLoS One, 2017, 12(5), e0177371, DOI:10.1371/journal.pone.0177371.
- J. W. Scannell, J. Bosley, J. A. Hickman, G. R. Dawson, H. Truebel, G. S. Ferreira, D. Richards and J. M. Treherne, Predictive Validity in Drug Discovery: What It Is, Why It Matters and How to Improve It, Nat. Rev. Drug Discovery, 2022, 21(12), 915–931, DOI:10.1038/s41573-022-00552-x.
- P. Geeleher, N. J. Cox and R. S. Huang, Clinical Drug Response Can Be Predicted Using Baseline Gene Expression Levels and in Vitro Drug Sensitivity in Cell Lines, Genome Biol., 2014, 15(3), R47, DOI:10.1186/gb-2014-15-3-r47.
- How to build the virtual cell with artificial intelligence: Priorities and opportunities: Cell, https://www.cell.com/cell/fulltext/S0092-8674(24)01332-1, accessed 2025-10-29.
- R. Jansen, J. D. A. van Embden, W. Gaastra and L. M. Schouls, Identification of Genes That Are Associated with DNA Repeats in Prokaryotes, Mol. Microbiol., 2002, 43(6), 1565–1575, DOI:10.1046/j.1365-2958.2002.02839.x.
- B.-S. Park, M. Lee, J. Kim and T. P. Kim, CRISPR–Cas Screening-Based Functional Genomics Approach for Drug Target Discovery, Exp. Mol. Med., 2025, 57(7), 1443–1454, DOI:10.1038/s12276-025-01487-0.
- F. Vazquez and W. R. Sellers, Are CRISPR Screens Providing the Next Generation of Therapeutic Targets?, Cancer Res., 2021, 81(23), 5806–5809, DOI:10.1158/0008-5472.CAN-21-1784.
- Gordian Biotechnology Keystone 2025 - Heart - Martin - YouTube, https://www.youtube.com/watch?v=MVhaSfktffw, accessed 2025-09-25.
- C. V. Dang, E. P. Reddy, K. M. Shokat and L. Soucek, Drugging the “undruggable” Cancer Targets, Nat. Rev. Cancer, 2017, 17(8), 502–508, DOI:10.1038/nrc.2017.36.
- A. Gironda-Martínez, E. J. Donckele, F. Samain and D. Neri, DNA-Encoded Chemical Libraries: A Comprehensive Review with Successful Stories and Future Challenges, ACS Pharmacol. Transl. Sci., 2021, 4(4), 1265–1279, DOI:10.1021/acsptsci.1c00118.
- C. A. Reiher, D. P. Schuman, N. Simmons and S. E. Wolkenberg, Trends in Hit-to-Lead Optimization Following DNA-Encoded Library Screens, ACS Med. Chem. Lett., 2021, 12(3), 343–350, DOI:10.1021/acsmedchemlett.0c00615.
- S. Iqbal, W. Jiang, E. Hansen, T. Aristotelous, S. Liu, A. Reidenbach, C. Raffier, A. Leed, C. Chen, L. Chung, E. Sigel, A. Burgin, S. Gould and H. H. Soutter, Evaluation of DNA Encoded Library and Machine Learning Model Combinations for Hit Discovery, NPJ Drug Discov., 2025, 2(1), 5, DOI:10.1038/s44386-025-00007-4.
- M. Kleinsasser and I. Quigley, Good binding data is all you need, Leash, 2025 Search PubMed . https://leashbio.substack.com/p/good-binding-data-is-all-you-need.
Footnotes |
| † We would like to emphasize our distinction between a data set — a fixed collection of data produced by some experimental technology (such as X-ray crystallography or cryogenic electron microscopy, in the case of the PDB) — and a data source, a process or technological capability to produce new data for purposes that might not have been envisaged at the time of the technology's development (e.g. cross-linking mass-spectrometry was not conceived to be used for generative ML). |
| ‡ Here, we use the term “taxonomy” referring to different experimental data generation strategies, rather than human-curated classification of biological entities and relationships (e.g., the Gene Ontology). A defining feature of the “Software 2.0″ and black-box data paradigms is that models often bypass the need for rigid, predefined biological taxonimies or ontologies. Instead of relying on human-annotated labels, modern neural architectures learn their own implicit representations and latent spaces directly from raw, uncurated experimental readouts, allowing the data itself to dictate the structural relationships. |
| This journal is © The Royal Society of Chemistry 2026 |