Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Stereoselective hydrogen isotope exchange on nicotinamide cofactors through flavoenzyme microscopic reversibility
Christopher W. Otun a,
Michael Yuen
a,
Michael Yuen a,
Harry J. Spaceya,
Carlo Bawn
a,
Harry J. Spaceya,
Carlo Bawn a,
Matthew J. Cliffa,
Francesco Falcionib,
Ryan A. Bragg
a,
Matthew J. Cliffa,
Francesco Falcionib,
Ryan A. Bragg b,
Charles S. Elmore
b,
Charles S. Elmore c,
Sam Hay
c,
Sam Hay a and
Jack S. Rowbotham
a and
Jack S. Rowbotham *a
*a
aManchester Institute of Biotechnology, Department of Chemistry, University of Manchester, Manchester, M1 7DN, UK. E-mail: jack.rowbotham@manchester.ac.uk
bEarly Chemical Development, Pharmaceutical Sciences, R&D, AstraZeneca, Cambridge, UK
cEarly Chemical Development, Pharmaceutical Sciences, R&D, AstraZeneca, Gothenburg, Sweden
First published on 24th April 2026
Abstract
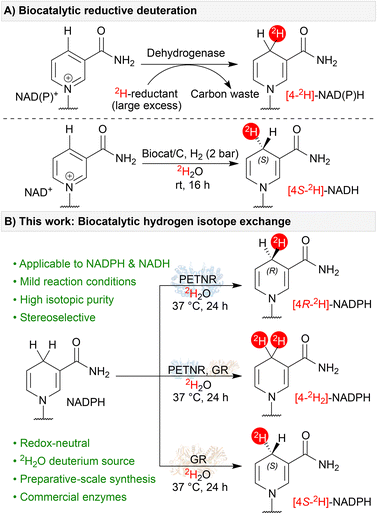
We describe a panel of flavoenzymes with the ability to catalyse stereoselective hydrogen isotope exchange (HIE) between 2H2O (D2O) and reduced nicotinamide cofactors, enabling a simple and redox-neutral route to deuterated NAD(P)H isotopomers. In screening the FMN- and FAD-dependent enzymes, which have diverse native functions, we identified catalysts (many of which are commercially available) that selectively yield the full suite of [4-2H]-NAD(P)H stereoisotopomers in a single step. In combining stereo-complementary enzymes, we also identify simple one-pot routes to dideuterated [4-2H2]-NAD(P)H. The biocatalytic methods provide near-quantitative 2H-incorporation for both NADPH and NADH under mild conditions, using readily available 2H2O as the isotope source. The observed activity indicates a reversible flavoenzyme hydride-transfer cycle, reliant on hydrogen/deuterium exchange on the transiently reduced flavin cofactors. We provide computational analysis to rationalise the stereochemical outcomes of the screened reactions. Finally, preparative-scale syntheses are described that deliver isolated deuterated cofactors in excellent yield and isotopic purity. We envisage this easily implemented procedure will simplify access to these important biochemical compounds for mechanistic studies, and may open up wider 2H2O-driven biocatalytic deuteration reactions.
Introduction
The range of catalytic approaches to hydrogen isotope labelling has been significantly expanded in recent years,1 driven in part to support the development of deuterated pharmaceuticals.2 Here, biocatalysis offers an attractive option for preparing deuterated compounds with high selectivity and low environmental impact.3 Enzymatic strategies for installing deuterium atoms (D or 2H) broadly fall into three categories, all of which have seen significant recent developments: (i) reductive deuteration,4 (ii) defunctionalisation deuteration (e.g. decarboxylation and dehalogenation),5 and (iii) hydrogen isotope exchange (HIE).6 Of the three, HIE is particularly advantageous as it enables late-stage installation of deuterium atoms by incubating the enzyme and target in 2H2O – a relatively low cost, readily available, and easy to handle isotope source.Many of the recently developed enzymatic HIE approaches exploit microscopic reversibility in biocatalytic mechanisms across a range of enzymes and cofactors. For instance, pyridoxal 5′-phosphate (PLP)-dependent enzymes have been exploited for both the regio- and stereo-selective deuteration of amino acids and derivatives,6a–c thiamine diphosphate (ThDP)-dependent enzymes can perform aldehyde HIE,6d and decarboxylases utilising a prenylated flavin cofactor (prFMN) can exchange alkene C(sp2)-1H for C(sp2)-2H.6e HIE in relation to NAD(P)H-dependent enzymes has been comparatively underexplored. Given the widespread synthetic applications of [4-2H]-NAD(P)H for reductive deuteration (see, for example, previous work by Rowbotham et al., Fig. 1A),4,7 a better understanding of redox neutral HIE could be of significant benefit.
In addition to their synthetic utility, isotopomers of [4-2H]-NAD(P)H are useful for a variety of applications in kinetic and mechanistic enzymology.8 As such, a number of routes for their synthesis have been published.4a,7,9 These examples, which often involve multiple steps, rely on sacrificial redox chemistry to drive the reductive deuteration of oxidised nicotinamide cofactors (NAD(P)+, Fig. 1A). However, in seminal work by Rabinowitz and colleagues wherein in vivo isotope distribution pathways were examined,10 the authors reported that flavoenzymes can catalyse hydride/deuteride exchange on nicotinamide cofactors in their reduced form. Furthermore, they hypothesised that this exchange should be stereoselective and identified both NADH- and NADPH-active enzymes capable of incorporating 2H at the 4-position of the nicotinamide ring. Motivated by these observations, we set out to explore and characterise this novel flavoenzyme activity in more detail and develop a panel of enzymes capable of easily preparing the full suite of [4-2H]-NAD(P)H stereoisotopomers (Fig. 1B). Developing this panel will, in turn, open up opportunities for other biocatalytic deuteration strategies that can exploit the very wide array of NAD(P)H-dependent enzymes to prepare selectively deuterated chiral amines and alcohols.
Results and discussion
Preparing a panel of flavoenzymes to test for HIE of NAD(P)H
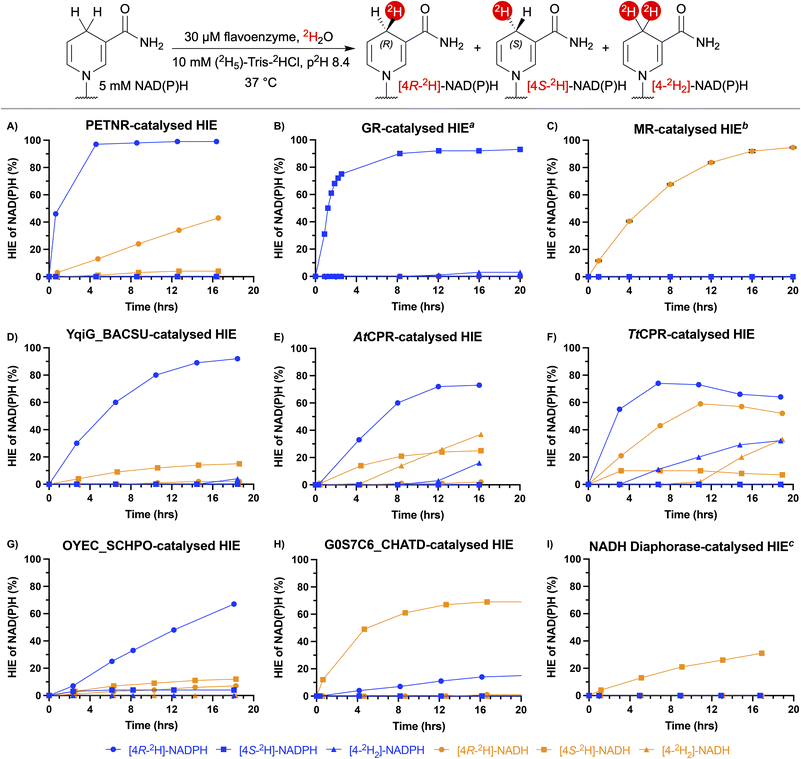
A panel of flavoenzymes was assembled to screen for HIE activity (Fig. 2). The full details of these varied enzyme candidates are summarised in the supplementary Table S1. Included in the enzyme panel were the reductase domains of various truncated thermostable cytochrome P450 monooxygenases (CYP450): CYP116B65 from Amycolatopsis thermoflava (AtCPR), CYP116B29 from Thermobispora bispora (TbCPR) and CYP116B46 from Tepidiphilus thermophilus (TtCPR). All of these enzymes are isostructural with the phthalate dioxygenase reductase from Pseudomonas cepacia and contain a flavin mononucleotide (FMN) cofactor, as well as a [2Fe-2S] iron-sulfur cluster within their structures.11 Also included in the flavoenzyme panel were the FMN-containing enzymes NADPH dehydrogenase C23G7.10c (OYEC_SCHPO) from Schizosaccharomyces pombe, commercially available NADH diaphorase from Clostridium kluyveri, pentaerythritol tetranitrate reductase (PETNR) from Enterobacter cloacae, morphinone reductase (MR) from Pseudomonas putida, thermophilic old yellow enzyme (TOYE) from Thermoanaerobacter pseudethanolicus, NADH:flavin oxidoreductase from Bacillus subtilis (YqiG_BACSU) and NADH:flavin oxidoreductase from Chaetomium thermophilum (G0S7C6_CHATD). Previously identified glutathione reductase (GR) from Saccharomyces cerevisiae was also included in the panel as a commercially available flavoenzyme which uses flavin adenine dinucleotide (FAD) instead of FMN.10A diagnostic 1H NMR signal at δ = 2.7–2.9 ppm, corresponding to the redox-active protons at the 4-position of the nicotinamide ring of NAD(P)H, was used to quantify 2H-incorporation and to determine the stereoselectivity of the HIE reactions.9g 1H NMR signals in the aromatic region were used to quantify NAD(P)H oxidation. Reaction mixtures were analysed by UPLC-MS to confirm monodeuteration by an increase in the molecular weight of NAD(P)H by +1.0. Instances of dideuteration were evidenced by a +2.0 increase in the molecular weight of NAD(P)H compared to commercial standards of NAD(P)H of natural isotopic abundance (see SI sections S5.2.1 and S5.2.2). Additionally, ultraviolet-visible (UV-vis) spectrophotometry was used to further monitor cofactor oxidation or degradation occurring during the HIE reactions by comparing A260![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) A340 ratios to known standards (see SI sections S5.2.3 and S5.2.4).12
A340 ratios to known standards (see SI sections S5.2.3 and S5.2.4).12
To minimise the oxidation or degradation of NAD(P)H in solution, all reactions and analytical samples were prepared under nitrogen. Unless otherwise stated, analytical-scale reaction mixtures containing 5 mM NAD(P)H, 30 µM flavoenzyme and 10 mM (2H5)-Tris-2HCl, p2H 8.4 in 2H2O were incubated at 37 °C (see SI section S3 for detailed methods).
Initial HIE screening using 1H NMR time-course experiments
The initial screen for flavoenzyme HIE activity was performed as a high-throughput 1H NMR time-course experiment, with 24 reactions run and monitored in parallel over a 20 hour period (Fig. 2). This system enabled the reactions to be incubated at the desired temperature, with periodic NMR sampling as necessary. In analysing these reactions, it was apparent that PETNR and GR can rapidly catalyse HIE between 2H2O and NADPH with >99% and 97% 2H-incorporation, respectively (Fig. 2A and B). In addition to the faster HIE rates exhibited by PETNR and GR compared to the other flavoenzymes tested, PETNR and GR were found to be both stereoselective and mutually stereo-complementary, with PETNR catalysing the formation of [4R-2H]-NADPH and GR catalysing the formation of [4S-2H]-NADPH. The HIE of NADH by PETNR resulted in lower levels of 2H-incorporation compared to HIE of NADPH, but stereoselectivity was maintained for the most part (43% [4R-2H]-NADH after 16 hours, Fig. 2A). GR was unable to catalyse HIE of NADH (Fig. 2B). Conversely, MR proved to be capable of catalysing highly stereoselective HIE of NADH (95% [4R-2H]-NADH, Fig. 2C), while incubation of MR with NADPH resulted in no 2H-incorporation (unsurprisingly, as MR is known to be highly specific for NADH).13 Similarly to PETNR, YqiG_BACSU was able to catalyse stereoselective HIE of NADPH (92% [4R-2H]-NADPH, Fig. 2D). However, the use of NADH as the cofactor resulted in a large decrease in YqiG_BACSU-catalysed HIE activity and was accompanied by a reversal of stereo-preference (15% [4S-2H]-NADH, Fig. 2D). This reversal of stereoselectivity was also exhibited by AtCPR, which resulted in the formation of 73% [4R-2H]-NADPH and 25% [4S-2H]-NADH in the HIE of NADPH and NADH, respectively (Fig. 2E). More generally, AtCPR-catalysed HIE of NADPH and NADH proceeded with less stereochemical control compared to most of the other enzymes screened. As such, AtCPR formed appreciable amounts of dideuterated product ([4-2H2]-NAD(P)H) by the end of the time-course experiments (16% [4-2H2]-NADPH and 37% [4-2H2]-NADH, Fig. 2E). Of all of the flavoenzymes studied, TtCPR showed the least cofactor specificity, with moderate 2H-incorporation levels for monodeuteration reached by the end of the experiment for both NADPH and NADH (64% [4R-2H]-NADPH and 52% [4R-2H]-NADH, Fig. 2F). While no reversal of stereoselectivity between TtCPR-catalysed HIE of NADPH and NADH was observed, TtCPR still exhibited a lack of strict stereochemical control, resulting in the formation of 32% [4-2H2]-NADPH and 33% [4-2H2]-NADH (Fig. 2F). With 67% [4R-2H]-NADPH detected at the end of the time-course, OYEC_SCHPO catalysed HIE of NADPH at a rate slower than any of the other flavoenzymes which possess a strong stereochemical preference for [4R-2H]-NADPH (Fig. 2G). Incubation of NADH with OYEC_SCHPO again resulted in a reversal of stereochemical preference and lower total levels of 2H-incorporation compared to the reaction with NADPH (7% [4R-2H]-NADH and 12% [4S-2H]-NADH, Fig. 2G). G0S7C6_CHATD showed a preference for NADH, as HIE of NADH resulted in the stereoselective formation of [4S-2H]-NADH (69%, Fig. 2H). G0S7C6_CHATD-catalysed HIE on NADPH also resulted in a reversal of stereochemistry (15% [4R-2H]-NADPH, Fig. 2H). Although selective for [4S-2H]-NADH, NADH diaphorase-catalysed HIE of NADH proceeded at a relatively slow rate, with levels of 2H-incorporation reaching 31% by the final measurement of the experiment (Fig. 2I). HIE catalysed by TbCPR and TOYE resulted in relatively low levels of 2H-incorporation for both NADPH and NADH. The kinetic profiles for these enzymes can be found in SI section S5.1. All of the promising reactions from the initial screening were repeated for 24 hours to substantiate the identification of flavoenzymes capable of complete deuteration of the redox active hydrogen of NAD(P)H (Table S4).Incubation of the flavoenzymes with the synthetic nicotinamide cofactor 1-methyl-1,4-dihydronicotinamide (MNAH), which can be used as a biomimetic cofactor in place of NAD(P)H,14 resulted in no deuteration in any instance (see SI section S4.3).
Interestingly, the UPLC-UV chromatograms and mass spectra obtained showed that, under some circumstances, the biologically active β-NAD(P)+ undergoes anomerisation to result in the formation of biologically inactive α-NAD(P)+ (see SI section S4.4 for a more detailed discussion and UPLC-UV peak assignment). However, this was only observed in rare cases where significant NAD(P)H oxidation had occurred, indicating that the α-NAD(P)+ might form from unstable mixtures of NAD(P)H and NAD(P)+.
Comparison of observed HIE against known enzyme catalytic parameters
An examination of published kinetic data for some of the flavoenzymes in their native reactions could explain the rates of HIE observed in Fig. 2. For example, glutathione reduction by GR has the NADPH reduction of the FAD cofactor as the rate determining step. However, this reductive half-reaction step occurs relatively rapidly, even at 5 °C (153 s−1).15 This could explain the particularly fast rate of GR-catalysed HIE of NADPH at 37 °C (Fig. 2B). GR is known to exhibit low Km values for NADPH (∼3 – 20 µM, depending on the organism),15,16 while Km values for NADH tend to be around 100 times higher.16 These values are consistent with the cofactor specificity exhibited by GR for NADPH over NADH under the HIE reaction conditions (Fig. 2B).Reports show that the rate of GR-bound FAD reduction is significantly faster than that of MR-bound FMN reduction (23.4 s−1 at 5 °C).15,17 The data obtained in this study is consistent with these reported kinetics as GR-catalysed HIE of NADPH (Fig. 2B) occurred at a rate faster than MR-catalysed HIE of NADH (Fig. 2C).
PETNR-catalysed HIE of NADPH occurred at a rate comparable to the GR-catalysed reaction (Fig. 2A), however, reported kinetic constants measured at 5 °C for the reduction of GR-bound FAD are significantly higher than that of PETNR-bound FMN.15,17b,18 This suggests that the kinetics of flavoenzyme-catalysed HIE between 2H2O and NAD(P)H cannot be accurately explained by looking at known rate constants for the reductive half-reactions alone. Nevertheless, the cofactor specificity exhibited by PETNR for NADPH over NADH in the HIE reaction is in agreement with reported Km values for PETNR (0.08 ± 0.01 mM for NADPH, 1.5 ± 0.7 mM for NADH),19 suggesting that the cofactor specificity observed in the native reactions is maintained in the flavoenzyme-catalysed HIE reactions. The difference in Km between NADPH and NADH for GR is far greater than that of PETNR,15,16,19 indicating that GR is more cofactor specific than PETNR. This trend in specificity is also observed in the HIE reactions (Fig. 2A versus 2B).
Observed non-enzymatic reactivity
Control experiments where NAD(P)H was incubated in 2H2O in the absence of any enzyme resulted in no 2H-incorporation, indicating that the protons at the 4-position of the nicotinamide ring do not freely undergo exchange with 2H2O (Table S4, entry 1). Experiments where enzyme-free FAD was incubated with NAD(P)H also resulted in no 2H-incorporation (Table S4, entry 2).The 1H NMR spectra obtained from the time-course experiments showed that a small proportion of the starting 1,4-NAD(P)H cofactor can undergo slow isomerisation to form redox-inactive 1,6-NAD(P)H. It is likely that this reactivity has been overlooked until now, because it is easily missed without the aid of a high field (800 MHz) spectrometer. In addition to unlabelled 1,6-NAD(P)H, our 1H NMR analysis revealed the formation of deuterated 1,6-NAD(P)H (assumed to be [6R-2H]-NAD(P)H from analogy of the peak shape with [4R-2H]-NAD(P)H). It is probable that, under the standard HIE reaction conditions, isomerisation of 1,4-NAD(P)H proceeds via a non-enzymatic hydride transfer from NAD(P)H to NAD(P)+.20 The observed stereo-preference of the reaction could be explained by the asymmetric conformation adopted by the cofactor in solution. For reactions which efficiently produced [4-2H]-NAD(P)H (e.g. GR-catalysed HIE of NADPH), levels of 1,6-NAD(P)H were around 2%. The highest levels of 1,6-NAD(P)H (≤10%) were observed only in reactions where significant cofactor oxidation took place (see SI section S4.5 for a more detailed discussion, NMR peak assignment and a scheme depicting the proposed mechanism for 1,4-NAD(P)H isomerisation).
Synthesis of dideuterated NAD(P)H
Having identified enzymes capable of performing efficient HIE for the synthesis of [4R-2H]-NADPH (PETNR, OYEC_SCHPO and YqiG_BACSU), [4S-2H]-NADPH (GR), [4R-2H]-NADH (MR) and [4S-2H]-NADH (G0S7C6_CHATD), we envisioned the possibility for complete dideuteration of NAD(P)H by using stereo-complementary flavoenzymes in one-pot. Through this approach, PETNR and GR successfully catalysed the formation of [4-2H2]-NADPH with 99% 2H-incorporation (Fig. 3A). NADH underwent dideuteration in a one-pot, two-step reaction where G0S7C6_CHATD was initially incubated with NADH for 24 hours, followed by the addition of MR to give [4-2H2]-NADH with 97% 2H-incorporation after a further 24 hours of incubation (Fig. 3B). It is worth noting that, for very high levels of deuteration, additional precautions are required to remove adventitious protons such as pre-exchanging the enzymes into 2H2O buffer, drying and exchanging out labile protons on other salts and reagents, and working in a dry atmosphere.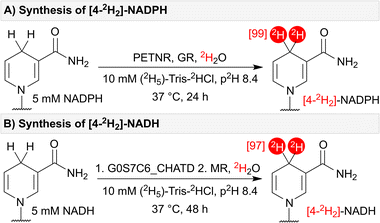 | ||
| Fig. 3 (A) Synthesis of [4-2H2]-NADPH – catalysed by stereo-complementary enzymes PETNR and GR. (B) Synthesis of [4-2H2]-NADH – catalysed by stereo-complementary enzymes G0S7C6_CHATD and MR. | ||
HIE mechanism and molecular dynamics
The proposed mechanism for flavoenzyme-catalysed HIE proceeds via microscopic reversibility in the half-reaction between NAD(P)H and the flavin cofactor (Fig. 4).10 NAD(P)H first reduces the flavin via hydride transfer, forming a labile N5–H bond that can exchange readily with 2H2O.17b The resulting deuterated reduced flavin then transfers deuterium to NAD(P)+ (via deuteride transfer) to generate [4-2H]-NAD(P)H.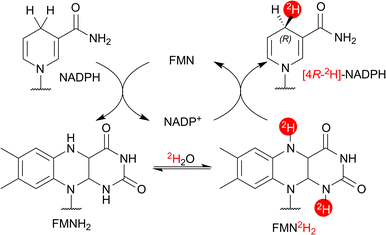 | ||
| Fig. 4 Speculative mechanism for AtCPR-catalysed HIE of NADPH.10 | ||
Truncation of AtCPR for the removal of the [2Fe-2S] cluster was performed to assess whether the [2Fe-2S] clusters in the CPRs affect HIE activity (see SI section S4.6). No significant difference in NADPH HIE activity was observed between truncated AtCPR (AtCPR-FMN) and full-length AtCPR, suggesting that the [2Fe-2S] cluster plays no role in the HIE mechanism.
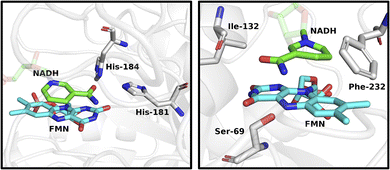
To explore differences in cofactor specificity and stereoselectivity between the flavoenzymes, molecular dynamics (MD) simulations were used to examine interactions of NADH with AtCPR and PETNR. For AtCPR, simulations were conducted from two different NADH binding poses that are consistent with either pro-(R) or pro-(S) hydride transfer to the flavin. These simulations revealed a lack of H-bonding interactions between the nicotinamide ring and the enzyme active site (Fig. 5 right for binding pose consistent with pro-(S) hydride transfer. See Fig. S6 for binding pose consistent with pro-(R) hydride transfer). Conversely, simulations of PETNR revealed interactions between the amide group of the nicotinamide ring and two histidine residues (His-181 and His-184, Fig. 5 left. See Fig. S7 for RMSD data).21 Experimental studies have shown that PETNR catalyses pro-(R) hydride transfer from NAD(P)H to alkene substrates, a stereo-preference likely governed by these nicotinamide interactions.8c,19,22 In contrast, the same interactions appear to be absent in AtCPR, enabling either pro-(R) or pro-(S) hydride transfer from the nicotinamide ring. This analysis supports the switch in stereoselectivity observed between AtCPR-catalysed HIE of NADPH and NADH (Fig. 2E). Since the 2′-phosphate of NADPH is distal from the active site, identifying specific H-interactions with this group is more difficult.23 The 2′-phosphate may constrain the cofactor in a binding orientation that favours pro-(R) hydride transfer, whereas the absence of this group in NADH may allow greater conformational flexibility and permit a binding pose that results in the formation of [4S-2H]-NADH. The conformational flexibility of NADH in the active site of AtCPR could also explain why significant levels of [4-2H2]-NADH were detected as, despite a preference for pro-(S) deuteration, deuteride transfer could occur on either face of the nicotinamide ring (Fig. 2E).
With the mechanism of flavoenzyme microscopic reversibility in mind, it is worth addressing the means by which the considerable generation of [4-2H2]-NADPH in the AtCPR- and TtCPR-catalysed HIE reactions may have arisen. In the time-course reactions, both enzymes initially produced [4R-2H]-NADPH as the sole product, however as time progressed, [4-2H2]-NADPH formation was observed, without any emergence of [4S-2H]-NADPH (Fig. 2E and F). The most plausible explanation for this occurrence may be a kinetic isotope effect (KIE) in the oxidation of [4R-2H]-NADPH after re-entry into the active sites of AtCPR and TtCPR (Fig. S8). This KIE may be large enough such that (S)-hydride transfer from [4R-2H]-NADPH to FMN becomes kinetically favourable over (R)-deuteride transfer, resulting in the generation of a planar [4-2H]-NADP+ intermediate. Subsequent deuteride transfer from FMN2H2 then leads to the formation of [4-2H2]-NADPH (see Fig. S8 for a proposed reaction scheme).
Preparative-scale synthesis of [4R-2H]-, [4S-2H]- and [4-2H2]-NADPH
Based on the levels of 2H-incorporation observed in the analytical-scale reactions, PETNR and GR were selected for preparative-scale HIE of NADPH. Firstly, a HIE reaction was performed with 25 mg NADPH (0.03 mmol) in the presence of 0.6 mol% PETNR to give [4R-2H]-NADPH in near quantitative yield and 99% 2H-incorporation (as measured by MS). The same was conducted with 34 U GR to afford [4S-2H]-NADPH in near quantitative yield and 96% 2H-incorporation. Finally, 25 mg NADPH was treated with PETNR (0.6 mol%) and GR (34 U) in one-pot to give [4-2H2]-NADPH in near quantitative yield and 94% 2H-incorporation. In all cases of preparative-scale synthesis, deuterated nicotinamide cofactor products were purified by anion exchange chromatography and easily isolated as white solids by lyophilisation.Conclusions and outlook
In summary, a panel of flavoenzymes was shown to catalyse regio- and stereoselective HIE between 2H2O and NAD(P)H. Flavoenzymes were identified for the efficient synthesis of [4R-2H]-NADPH (PETNR, OYEC_SCHPO and YqiG_BACSU), [4S-2H]-NADPH (GR) and [4R-2H]-NADH (MR). [4S-2H]-NADH could also be generated (G0S7C6_CHATD), albeit with a lower level of 2H-incorporation compared to the other NAD(P)H isotopologues mentioned under the conditions that we studied. It is likely that more successful candidates exist for this target, which may become apparent with future research.† [4-2H2]-NAD(P)H was also easily accessed upon incubation of NAD(P)H with the relevant stereo-complementary flavoenzymes in 2H2O in one-pot. Enzymatic HIE was successfully monitored over time with a parallel 1H NMR spectroscopic approach. Molecular docking of NADH in the active site of different candidates provided plausible explanations for the varying cofactor specificity and stereoselectivity of these enzymes. Moreover, the demonstration of preparative-scale enzymatic HIE shows how this method could be employed as a convenient and atom economical method for the preparation of deuterated cofactors to be used in the study of NAD(P)H-dependent enzyme kinetics and mechanisms. Our future work will focus on exploiting the observed activity in synthetic HIE strategies using NAD(P)H-dependent enzymes to prepare valuable deuterated chiral amines and alcohols.Author contributions
J. S. R. conceptualised and supervised the work. F. F., R. A. B. and C. S. E. provided industrial supervision and steered the applicative focus of the project. C. W. O., M. Y. and H. J. S. performed protein expression and purification. C. W. O. designed and carried out all deuteration experiments. C. B and M. J. C assisted C. W. O. with the execution of the NMR time-course experiments. M. Y. and S. H. carried out computational work and helped with interpretation. C. W. O. and J. S. R contributed to data analysis. C. W. O. prepared the original manuscript draft, and all authors assisted with the reviewing and editing process.Conflicts of interest
There are no conflicts to declare.Data availability
The datasets supporting this article have been uploaded as part of the SI. Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d6sc01003b.Acknowledgements
Funding from the EPSRC Future Biomanufacturing Research Hub (EP/S01778X/1) with additional support from AstraZeneca is gratefully acknowledged. J. S. R. also acknowledges funding from the BBSRC (BB/V005464/1 and UKRI2412) and Royal Society (URF\R1\231404). This work was also partly funded by BBSRC grant BB/X000974/1 and M. Y. is supported by EPSRC grant EP/S023755/1. The authors would like to acknowledge the equipment grant BB/X019217/1 as well.Notes and references
- (a) Y. Y. Loh, K. Nagao, A. J. Hoover, D. Hesk, N. R. Rivera, S. L. Colletti, I. W. Davies and D. W. C. MacMillan, Science, 2017, 358, 1182–1187 Search PubMed; (b) S. Kopf, F. Bourriquen, W. Li, H. Neumann, K. Junge and M. Beller, Chem. Rev., 2022, 122, 6634–6718 CrossRef CAS PubMed; (c) A. Tarasewicz and V. Derdau, Nat. Chem., 2023, 15, 1332–1333 CrossRef CAS PubMed; (d) F. Bu, Y. Deng, J. Xu, D. Yang, Y. Li, W. Li and A. Lei, Nature, 2024, 634, 592–599 Search PubMed; (e) B. I. P. Smith, N. M. L. Knight, G. J. Knox, D. M. Lindsay, L. C. Paterson, J. Bergare, C. S. Elmore, R. A. Bragg and W. J. Kerr, Angew. Chem., Int. Ed., 2025, 64, e202417179 Search PubMed; (f) S. Barranco, I. L. Eshel, J. Zhang, M. Di Matteo, A. Milo and M. H. Pérez-Temprano, Nat. Catal., 2025, 8, 1306–1313 Search PubMed; (g) Y. Q. Guan, T. Z. Wang, M. Bilal, X. R. Tan, L. Ackermann and Y. F. Liang, Nat. Commun., 2025, 16, 10235 Search PubMed; (h) Y. Koga, I. Fukumoto, K. Masui, T. Tanaka, Y. Naganawa, M. Hayashi, T. Ohshima and R. Yazaki, Nat. Catal., 2025, 8, 1062–1071 CrossRef CAS; (i) Y. Meng, B. Shu, J. Zhang, H. Rao, Z. Zhou, Z. Wang, Z. Liu, K. Liu, Y. Zhang and W. Wang, Nat. Commun., 2025, 16, 6744 Search PubMed; (j) S. E. Park, S. Choi, C. Lim, S. H. Lee, S. Jeong and J. M. Joo, Chem. Sci., 2025, 16, 10349–10356 Search PubMed.
- R. M. C. Di Martino, B. D. Maxwell and T. Pirali, Nat. Rev. Drug Discovery, 2023, 22, 562–584 Search PubMed.
- R. D. Jansen-van Vuuren, L. Raju, C. W. Otun, V. Fasano, J. S. Rowbotham and D. Kananovich, in Isotopically-Labeled Compounds: Recent Advances and Emerging Topics in Synthesis and Applications, eds. D. Hesk and V. Derdau, Elsevier, 1st edn, in press, ch. 10 Search PubMed.
- (a) J. S. Rowbotham, M. A. Ramirez, O. Lenz, H. A. Reeve and K. A. Vincent, Nat. Commun., 2020, 11, 1454 Search PubMed; (b) J. S. Rowbotham, J. H. Nicholson, M. A. Ramirez, K. Urata, P. M. T. Todd, G. Karunanithy, L. Lauterbach, H. A. Reeve, A. J. Baldwin and K. A. Vincent, Chem. Sci., 2023, 14, 12160–12165 RSC.
- (a) J. Xu, J. Fan, Y. Lou, W. Xu, Z. Wang, D. Li, H. Zhou, X. Lin and Q. Wu, Nat. Commun., 2021, 12, 3983 CrossRef CAS; (b) Y. Fu, X. Liu, Y. Xia, X. Guo, J. Guo, J. Zhang, W. Zhao, Y. Wu, J. Wang and F. Zhong, Chem, 2023, 9, 1897–1909 CrossRef CAS.
- (a) S. W. Chun and A. R. H. Narayan, ACS Catal., 2020, 10, 7413–7418 CrossRef CAS PubMed; (b) T. J. Doyon and A. R. Buller, J. Am. Chem. Soc., 2022, 144, 7327–7336 CrossRef CAS PubMed; (c) J. Gao, C. Zhou and Y. Hai, ChemBioChem, 2023, 24, e202300561 CrossRef CAS PubMed; (d) J. Xu, Y. Lou, L. Wang, Z. Wang, W. Xu, W. Ma, Z. Chen, X. Chen and Q. Wu, ACS Catal., 2021, 11, 13348–13354 CrossRef CAS; (e) L. Wang, Y. Lou, W. Xu, Z. Chen, J. Xu and Q. Wu, ACS Catal., 2021, 12, 783–788 CrossRef.
- J. S. Rowbotham, A. P. Hardy, H. A. Reeve and K. A. Vincent, J. Labelled Compd. Radiopharm., 2021, 64, 181–186 CrossRef CAS PubMed.
- (a) V. V. Pollock and M. J. Barber, Biochemistry, 2001, 40, 1430–1440 CrossRef CAS PubMed; (b) R. A. Dick and T. W. Kensler, J. Biol. Chem., 2004, 279, 17269–17277 CrossRef CAS PubMed; (c) C. R. Pudney, S. Hay, M. J. Sutcliffe and N. S. Scrutton, J. Am. Chem. Soc., 2006, 128, 14053–14058 CrossRef CAS PubMed; (d) S. Hay, C. R. Pudney and N. S. Scrutton, FEBS J., 2009, 276, 3930–3941 CrossRef CAS PubMed; (e) J. E. Longbotham, M. Delgado, M. Yuen, J. Berreur, S. Görlich, A. Iorgu, L. O. Johannissen, D. J. Heyes, N. S. Scrutton, V. Moliner, I. Tuñón and S. Hay, ACS Catal., 2025, 15, 14456–14468 CrossRef CAS.
- (a) C. Wong and G. Whitesides, J. Am. Chem. Soc., 1983, 105, 5012–5014 CrossRef CAS; (b) C. R. Pudney, S. Hay and N. S. Scrutton, in Flavins and Flavoproteins: Methods and Protocols, eds. S. Weber and E. Schleicher, Springer New York, New York, NY, 2014, DOI: DOI:10.1007/978-1-4939-0452-5_8, pp. 161–175; (c) S. S. Jeong and J. E. Gready, Anal. Biochem., 1994, 221, 273–277 CrossRef CAS PubMed; (d) A. Yahashiri, A. Sen and A. Kohen, J. Labelled Compd. Radiopharm., 2009, 52, 463–466 CrossRef CAS PubMed; (e) D. Roston and A. Kohen, J. Labelled Compd. Radiopharm., 2014, 57, 75–77 CrossRef CAS PubMed; (f) A. Kohen, P. Singh and Q. Guo, Synlett, 2017, 28, 1151–1159 CrossRef; (g) J. S. Rowbotham, H. A. Reeve and K. A. Vincent, ACS Catal., 2021, 11, 2596–2604 CrossRef CAS PubMed; (h) A. Al-Shameri, M. C. Petrich, K. Junge Puring, U. P. Apfel, B. M. Nestl and L. Lauterbach, Angew. Chem., Int. Ed., 2020, 59, 10929–10933 CrossRef CAS.
- Z. Zhang, L. Chen, L. Liu, X. Su and J. D. Rabinowitz, J. Am. Chem. Soc., 2017, 139, 14368–14371 CrossRef CAS PubMed.
- (a) C. C. Correll, C. J. Batie, D. P. Ballou and M. L. Ludwig, Science, 1992, 258, 1604–1610 CrossRef CAS PubMed; (b) M. Tavanti, J. L. Porter, S. Sabatini, N. J. Turner and S. L. Flitsch, ChemCatChem, 2018, 10, 1042–1051 CrossRef CAS.
- J. De Ruyck, M. Famerée, J. Wouters, E. A. Perpète, J. Preat and D. Jacquemin, Chem. Phys. Lett., 2007, 450, 119–122 CrossRef CAS.
- C. E. French and N. C. Bruce, Biochem. J., 1994, 301, 97–103 CrossRef CAS PubMed.
- (a) F. Friedlos, M. Jarman, L. C. Davies, M. P. Boland and R. J. Knox, Biochem. Pharmacol., 1992, 44, 25–31 CrossRef CAS PubMed; (b) Q. V. Vo, N. T. Hoa and A. Mechler, RSC Adv., 2024, 14, 37196–37201 RSC; (c) C. E. Paul and F. Hollmann, Appl. Microbiol. Biotechnol., 2016, 100, 4773–4778 CrossRef CAS PubMed; (d) K. C. Kenney, T. P. LaFortune and G. A. Weiss, ACS Catal., 2025, 15, 18591–18600 CrossRef CAS.
- P. W. Huber and K. G. Brandt, Biochemistry, 1980, 19, 4568–4575 CrossRef CAS PubMed.
- M. A. Vanoni, K. K. Wong, D. P. Ballou and J. S. Blanchard, Biochemistry, 1990, 29, 5790–5796 CrossRef CAS PubMed.
- (a) D. H. Craig, P. C. E. Moody, N. C. Bruce and N. S. Scrutton, Biochemistry, 1998, 37, 7598–7607 CrossRef CAS PubMed; (b) J. Basran, R. J. Harris, M. J. Sutcliffe and N. S. Scrutton, J. Biol. Chem., 2003, 278, 43973–43982 CrossRef CAS PubMed.
- H. Khan, R. J. Harris, T. Barna, D. H. Craig, N. C. Bruce, A. W. Munro, P. C. Moody and N. S. Scrutton, J. Biol. Chem., 2002, 277, 21906–21912 CrossRef CAS PubMed.
- C. R. Pudney, S. Hay and N. S. Scrutton, FEBS J., 2009, 276, 4780–4789 CrossRef CAS PubMed.
- (a) S. E. Godtfredsen, M. Ottesen and N. R. Andersen, Carlsberg Res. Commun., 1979, 44, 65–75 CrossRef CAS; (b) M. V. Makarov, F. Hayat, B. Graves, M. Sonavane, E. A. Salter, A. Wierzbicki, N. R. Gassman and M. E. Migaud, ACS Chem. Biol., 2021, 16, 604–614 CrossRef CAS PubMed.
- H. S. Toogood, A. Fryszkowska, M. Hulley, M. Sakuma, D. Mansell, G. M. Stephens, J. M. Gardiner and N. S. Scrutton, ChemBioChem, 2011, 12, 738–749 CrossRef CAS PubMed.
- (a) C. R. Pudney, R. S. Lane, A. J. Fielding, S. W. Magennis, S. Hay and N. S. Scrutton, J. Am. Chem. Soc., 2013, 135, 3855–3864 CrossRef CAS PubMed; (b) C. R. Pudney, S. Hay, C. Levy, J. Pang, M. J. Sutcliffe, D. Leys and N. S. Scrutton, J. Am. Chem. Soc., 2009, 131, 17072–17073 CrossRef CAS PubMed.
- A. I. Iorgu, N. J. Baxter, M. J. Cliff, C. Levy, J. P. Waltho, S. Hay and N. S. Scrutton, ACS Catal., 2018, 8, 11589–11599 CrossRef CAS PubMed.
Footnote |
| † For example, Rowbotham et al. have previously identified an atom efficient route to [4S-2H]-NADH using a hydrogenase-driven approach,7 and preliminary data indicates that the FMN-centred NAD+ reductase domain may also be a suitable candidate for HIE. |
| This journal is © The Royal Society of Chemistry 2026 |