Open Access Article
Open Access ArticleTowards accurate and scalable high-throughput MOF adsorption screening: merging classical force fields and universal machine learned interatomic potentials
Satyanarayana Bonakalaa,
Mohammad Wahiduzzaman a,
Taku Watanabeb,
Karim Hamzaouib and
Guillaume Maurin
a,
Taku Watanabeb,
Karim Hamzaouib and
Guillaume Maurin *ac
*ac
aICGM, Univ. Montpellier, CNRS, ENSCM, 34293 Montpellier, France. E-mail: guillaume.maurin1@umontpellier.fr
bMatlantis Corporation, Otemachi, Chiyoda-ku, 100-0004 Tokyo, Japan
cInstitut Universitaire de France, France
First published on 5th May 2026
Abstract
High-throughput computational screening (HTCS) of gas adsorption in metal–organic frameworks (MOFs) typically relies on classical generic force fields which are computationally efficient but often fail to capture complex host–guest interactions. Universal machine learned interatomic potentials (u-MLIPs) offer near-quantum accuracy at far lower cost than density-functional theory (DFT), yet their large-scale application in adsorption screening remains limited. Here, we introduce an efficient hybrid screening workflow that merges classical generic force fields and u-MLIPs within a Monte Carlo scheme to accurately assess the adsorption performance of a large MOF structure database. As a proof-of-concept, this HTCS is applied to identify top-performing MOFs for selective ethylene capture under humid conditions, highly relevant to food-preservation packaging technologies. Our workflow demonstrates that accurate treatment of both host–guest energetics and framework flexibility, enabled by u-MLIPs, is essential to achieve reliable adsorption-performance rankings.
Introduction
Metal–Organic Frameworks (MOFs), a class of hybrid porous materials with remarkable chemical and structural diversity, hold immense promise across a wide range of applications. Yet, their full potential, particularly for sorption-based processes, remains largely untapped. One key reason is that only a small fraction of the 128![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000+ MOF structures reported in the Cambridge Structural Database (CSD)1 have been explored for specific applications. This limited exploration is primarily due to the impracticality of synthesizing and experimentally testing thousands of candidates, given the substantial investment of time, resources and specialized infrastructure required. To overcome these limitations, computational approaches have emerged as indispensable tools. In particular, high-throughput computational screening (HTCS) has revolutionized the evaluation of the properties of MOFs and related porous materials across large structural databases. This strategy enables the rapid identification of promising sorbent candidates for a wide array of adsorption/separation applications, including carbon capture, H2 storage, and natural gas purification, among others.2–4 Traditionally, these computational screenings have relied on classical interatomic potentials, commonly referred to as classical universal interatomic potentials (u-IPs) to describe the MOF−guest interactions. The most widely used classical u-IPs, e.g., UFF5 employ transferable Lennard-Jones parameters to account for van der Waals interactions, complemented by electrostatic terms to describe interatomic interactions within the system.
000+ MOF structures reported in the Cambridge Structural Database (CSD)1 have been explored for specific applications. This limited exploration is primarily due to the impracticality of synthesizing and experimentally testing thousands of candidates, given the substantial investment of time, resources and specialized infrastructure required. To overcome these limitations, computational approaches have emerged as indispensable tools. In particular, high-throughput computational screening (HTCS) has revolutionized the evaluation of the properties of MOFs and related porous materials across large structural databases. This strategy enables the rapid identification of promising sorbent candidates for a wide array of adsorption/separation applications, including carbon capture, H2 storage, and natural gas purification, among others.2–4 Traditionally, these computational screenings have relied on classical interatomic potentials, commonly referred to as classical universal interatomic potentials (u-IPs) to describe the MOF−guest interactions. The most widely used classical u-IPs, e.g., UFF5 employ transferable Lennard-Jones parameters to account for van der Waals interactions, complemented by electrostatic terms to describe interatomic interactions within the system.
While this modelling approach enables an effective qualitative assessment of thousands of MOF sorbents, it inherently lacks the precision required to capture complex host–guest interactions, especially between guest molecules and functionalized linkers as well as open metal sites (OMS) and polar groups within the inorganic nodes, where polarization effect may play a dominant role.6 For example, UFF significantly underestimates CO2 interaction energies with the OMSs of Mg-MOF-74 by 60–65% compared to density-functional theory (DFT) calculations,7 which can potentially lead to the mis-ranking of this sub-class of MOFs. To overcome these shortcomings, case-specific FF parametrization has been developed using exhaustive DFT-derived interaction energies for individual MOF–guest pairs, e.g., CO2@Mg-MOF-74,8 C2H2@Cu-BTC,9 N2@MIL-100(Cr)10 and SO2@Mg2(dobpdc).11 More recently, Machine-learned potentials (MLPs), trained on high-quality quantum mechanical datasets, have proven effective in capturing intricate host–guest interactions across a range of MOFs, achieving near quantum mechanical accuracy at significantly lower computational cost.12–14 For instance, this strategy has been successfully applied to accurately predict the H2 adsorption isotherm in the low-pressure domain of Al-soc-MOF-1d containing OMSs.15 Despite their improved accuracy in targeted studies for a few MOFs, both case-specific FFs and trained MLPs remain limited in transferability and are computationally impractical for large-scale screenings. In this context, the on-going development of universal machine learned interatomic potentials (u-MLIPs) offers a potentially transformative advance. Foundation models such as the Multi-Atomic Cluster Expansion (MACE),16 Materials Graph Network with three-body interactions (M3GNet)17 and Crystal Hamiltonian Graph Neural Network (CHGNet)18 architectures have demonstrated near-DFT accuracy for predicting MOF framework properties offering a promising pathway towards MLP-based generalizable MOF screening platforms. Notably, Meta's Equiformer V2, eSEN, or UMA models trained on the Open DAC datasets, ODAC23 and ODAC25 have shown superior performance over the UFF in predicting CO2 and H2O adsorption energies for a few MOFs, especially those featuring OMSs.19 A few recent benchmark studies extended this comparison across a small sub-set of MOFs evaluating a range of u-MLIPs including EqV2-ODAC,20 MACE, M3GNet, CHGNet, and the Equivariant Smooth Energy Network (eSEN)21 against DFT and UFF calculations for computing their CO2 and H2O adsorption energies.22
However, despite these promising developments, u-MLIPs have not yet been systematically applied to predict adsorption properties across a large and diverse set of MOFs. Their integration into HTCS pipelines is still not feasible due to computational costs, which, while significantly lower compared to DFT, are still substantially higher than those associated with classical FFs. Herein, we advance a hybrid HTCS workflow for MOFs that bridges classical u-IPs and u-MLIPs, using ethylene (C2H4) capture under humid conditions as a test case. This adsorption scenario poses a fundamental challenge: weakly physisorbed ethylene must compete with strongly interacting polar water molecules, requiring accurate modelling of the MOF–guest interactions. Ethylene was selected due to its high-impact relevance in food preservation technologies, particularly in regulating the ripening of fruits and vegetables.23 Its uncontrolled accumulation in postharvest storage and food packaging, environments typically characterized by high humidity,24 can accelerate spoilage, contributing significantly to food loss.25 A few technologies have been proposed to mitigate ethylene accumulation, including chemical inhibitors such as 1-methylcyclopropene (1-MCP),26 oxidative scavenging agents like potassium permanganate (KMnO4)27 and TiO2-based photocatalysts.28 A truly viable sorbent-based solution has yet to be realized, as conventional porous materials, such as zeolites and activated carbons, exhibit inherently low C2H4 affinity and/or inadequate C2H4/H2O selectivity.29,30 Recently, a few biodegradable and non-toxic MOFs have been explored as alternative sorbents for food-related applications.31,32 For example, cyclodextrin-based MOFs (CD-MOF-Na/K),32 synthesized from bio-derived α-cyclodextrins and alkali metal ions, have been proposed as effective C2H4 sorbents for delaying the ripening of fruits under ambient conditions. These preliminary findings highlight the potential of MOFs in selective C2H4 capture and pave the way for the identification of other existing MOFs with optimized C2H4 sorption properties.
Our proposed computational workflow begins with a HTCS of a curated database of sustainable MOFs selected from the 128![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 799 experimentally synthesized MOFs in the latest release of the Cambridge Structure Database.1 Following a rigorous MOF structure curation and geometry optimization pipeline, key adsorption descriptors, Henry's law constants (KH) and enthalpy of adsorption at infinite dilution (ΔH0,ads) are computed via the Widom insertion method at room temperature for both C2H4 and H2O as well as the resulting ideal selectivity (SC2H4/H2O = KH,C2H4/KH,H2O) using the UFF. This first conventional stage enables the identification of hydrophobic MOF candidates capable of selectively capturing ethylene over water. The Preferred Potential (PFP) u-MLIP, an equivariant graph neural network AI-model trained on a comprehensive dataset of 42 million DFT-calculated structures (including molecules, crystals, slabs, clusters and disordered systems),33 is subsequently employed to refine the performance predictions of the 88 experimentally viable ethylene selective MOFs identified in the initial screening. Following its rigorous validation against DFT-calculated adsorption energies for water and ethylene across these 88 MOFs, the PFP u-MLIP is systematically applied to compute ΔH0,ads and KH for both guest molecules and the corresponding S(C2H4/H2O). Notably, under the commonly adopted rigid-MOF framework approximation in HTCS workflows, the UFF approach demonstrates reliable performance in capturing the hydrophobic behaviour of most MOFs examined in this study, and in yielding ethylene adsorption enthalpies that are comparable to those predicted by the PFP u-MLIP model with a mean absolute deviation (MAD) of 5 kJ mol−1. The use of PFP u-MLIP becomes more critical for a small number of outlier MOFs featuring small pore sizes or polar functional groups, where UFF proves less effective at accurately capturing C2H4 and H2O adsorption energetics, particularly under high confinement or when hydrogen-bonding interactions play a significant role. While UFF performs adequately for most cases, these outliers highlight the limitations of classical generic FFs and the advantages of higher-fidelity models like PFP u-MLIP. Notably, PFP u-MLIP results show that 99% of the MOFs exhibit ideal selectivity, S(C2H4/H2O) > 1, confirming their strong ethylene preference and suitability for separation under humid conditions.
799 experimentally synthesized MOFs in the latest release of the Cambridge Structure Database.1 Following a rigorous MOF structure curation and geometry optimization pipeline, key adsorption descriptors, Henry's law constants (KH) and enthalpy of adsorption at infinite dilution (ΔH0,ads) are computed via the Widom insertion method at room temperature for both C2H4 and H2O as well as the resulting ideal selectivity (SC2H4/H2O = KH,C2H4/KH,H2O) using the UFF. This first conventional stage enables the identification of hydrophobic MOF candidates capable of selectively capturing ethylene over water. The Preferred Potential (PFP) u-MLIP, an equivariant graph neural network AI-model trained on a comprehensive dataset of 42 million DFT-calculated structures (including molecules, crystals, slabs, clusters and disordered systems),33 is subsequently employed to refine the performance predictions of the 88 experimentally viable ethylene selective MOFs identified in the initial screening. Following its rigorous validation against DFT-calculated adsorption energies for water and ethylene across these 88 MOFs, the PFP u-MLIP is systematically applied to compute ΔH0,ads and KH for both guest molecules and the corresponding S(C2H4/H2O). Notably, under the commonly adopted rigid-MOF framework approximation in HTCS workflows, the UFF approach demonstrates reliable performance in capturing the hydrophobic behaviour of most MOFs examined in this study, and in yielding ethylene adsorption enthalpies that are comparable to those predicted by the PFP u-MLIP model with a mean absolute deviation (MAD) of 5 kJ mol−1. The use of PFP u-MLIP becomes more critical for a small number of outlier MOFs featuring small pore sizes or polar functional groups, where UFF proves less effective at accurately capturing C2H4 and H2O adsorption energetics, particularly under high confinement or when hydrogen-bonding interactions play a significant role. While UFF performs adequately for most cases, these outliers highlight the limitations of classical generic FFs and the advantages of higher-fidelity models like PFP u-MLIP. Notably, PFP u-MLIP results show that 99% of the MOFs exhibit ideal selectivity, S(C2H4/H2O) > 1, confirming their strong ethylene preference and suitability for separation under humid conditions.
Although guest-triggered flexibility in MOFs is well documented and has been shown to significantly affect host–guest interactions in specific cases,34 incorporating such structural responsiveness into HTCS workflows remains computationally prohibitive. Assessing whether the rigid-framework approximation introduces systematic deviations or remains within acceptable error margins is critical to ensuring the reliability of HTCS-derived adsorption properties. To this end, the PFP u-MLIP model was also employed to refine the predictions of ΔH0,ads(C2H4) for the 88 top MOF candidates, explicitly accounting for adsorption-induced framework flexibility, including atomic displacements alone or in combination with unit cell relaxations. It was found that for most MOFs exhibiting guest-induced unit cell volume changes below 10% led to only moderate deviations in ΔH0,ads(C2H4) relative to the rigid-framework approximation (mean absolute deviation (MAD) of 1.5 kJ mol−1). This refined step enables the exclusion of a few otherwise attractive MOFs that undergo significantly larger guest-induced unit cell expansion, accompanied by a notable decrease in C2H4 affinity.
Beyond introducing a robust data-driven strategy for identifying MOFs suitable for ethylene removal under humid food storage conditions as a test case, this work proposes a generalized hybrid HTCS workflow that balances computational efficiency with predictive accuracy. We show that the classical generic force field UFF, when combined with rigid framework approximations, provides a reliable and scalable first-pass filter to identify top-performing MOF candidates for sorption-based applications. For the top-ranked MOFs, we then leverage PFP u-MLIP to refine UFF-derived performance predictions, capturing host–guest interactions and structural flexibility more accurately. Our findings provide valuable insights into the transformative potential of u-MLIPs in HTCS, offering a powerful strategy to refine classical u-IPs predictions and to enable more accurate and resource-efficient screening pipelines for MOF-based sorption properties.
Computational methodology
Collation and curation of a sustainable MOF structure database
A high-quality and structurally consistent MOF database is a prerequisite for reliable HTCS of MOF properties.35 As such, a systematic curation pipeline was implemented starting from the CSD MOF database (version 2024.3.0)1 as outlined in Fig. 1. We cross-referenced the initial CSD entries with two well-established molecular simulation-ready MOFs: QMOF36 and MOSAEC-DB.37 To avoid redundant, time- and computation-intensive curation steps, we excluded CSD MOF entries that overlapped with QMOF (15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 966) and/or MOSAEC-DB (71
966) and/or MOSAEC-DB (71![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 138), retaining a total of 25
138), retaining a total of 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 388 MOFs from the non-disordered subset of the CSD MOF database. These MOFs were subsequently curated using SAMOSA (Structural Activation via Metal Oxidation State Analysis) tool,38 which removes free solvent molecules while preserving chemically relevant monodentate species. This yielded a combined baseline dataset of 112
388 MOFs from the non-disordered subset of the CSD MOF database. These MOFs were subsequently curated using SAMOSA (Structural Activation via Metal Oxidation State Analysis) tool,38 which removes free solvent molecules while preserving chemically relevant monodentate species. This yielded a combined baseline dataset of 112![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 492 unique MOF structures for our HTCS pipeline. To ensure the accessibility of ethylene (kinetic diameter of 4.163 Å), Zeo++ software was used to filter MOFs with a pore-limiting diameter (PLD) > 4.1 Å. Additionally, MOFs were pre-selected based on their environmentally benign metal ions (Al, Zn, Hf, Fe or Ti) to align with sustainability and health safety standards relevant to food-related applications. Combining these geometric and sustainability criteria yielded a refined subset of 4166 MOFs.
492 unique MOF structures for our HTCS pipeline. To ensure the accessibility of ethylene (kinetic diameter of 4.163 Å), Zeo++ software was used to filter MOFs with a pore-limiting diameter (PLD) > 4.1 Å. Additionally, MOFs were pre-selected based on their environmentally benign metal ions (Al, Zn, Hf, Fe or Ti) to align with sustainability and health safety standards relevant to food-related applications. Combining these geometric and sustainability criteria yielded a refined subset of 4166 MOFs.
We then implemented a rigorous refinement process to ensure the highest possible chemical correctness and to eliminate structural redundancy. First, we applied the Metal Oxidation State Automated Error Checker (MOSAEC)39 to identify chemically inconsistent structures by analysing metal oxidation states. MOFs flagged as “BAD” by MOSAEC, typically due to unbalanced charges or chemically implausible valence assignments, were excluded. Next, we addressed structural redundancy by evaluating both topological and geometric similarity using the CoRE-MOF-Tools package.35 Specifically, we employed the StructureMatcher module to identify duplicate or near-duplicate geometries. MOFs with charged frameworks were finally excluded, as these require additional considerations, such as counterion placement, that fall beyond the scope of this study. This multi-stage refinement process yielded a final dataset of 1881 neutral and computation-ready MOFs that were geometry optimized using the EqV2-ODAC u-MLIP via BFGS optimization in the Atomic Simulation Environment (ASE), with a force convergence threshold of 0.05 eV Å−1.40
Monte Carlo simulations
PFP u-MLIP Widom's insertion simulations
The PFP u-MLIP, version 6.0.0,33 was implemented within Widom's test particle insertion method to assess KH and ΔH0,ads for both water and ethylene in the 88 top-performing MOFs identified by UFF-based HTCS. To accurately account for dispersion interactions, the PBE-D3 correction47,48 was applied via the CRYSTAL_U0_PLUS_D3 module, improving the description of the van der Waals interactions between MOFs and the guests. Each simulation was carried out for 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 MC cycles. To further investigate the impact of MOF framework flexibility on ΔH0,ads of ethylene, simulations were carried out using three distinct structural configurations for each of the 88 MOFs: (i) MOFreference – structures optimized (atomic positions relaxed) in the absence of guest molecules using the EqV2-ODAC u-MLIP model; (ii) MOFgeometry-opt – structures with atomic positions relaxed in the presence of a single ethylene molecule using the PFP u-MLIP; (iii) MOFcell-opt – structures with both atomic positions and unit cell parameters relaxed in the presence of a single ethylene molecule using the PFP u-MLIP. The initial structures for (ii) and (iii) were derived from the minimum energy ethylene adsorption configurations identified during the PFP u-MLIP based Widom insertion simulations. These geometry optimizations were performed using the BFGS optimizer49 with a force convergence threshold of 0.005 eV Å−1. All Widom insertion and geometry optimization calculations using the PFP u-MLIP were performed via the MATLANTIS platform,50 under an academic license. In this framework, the PFP u-MLIP is implemented through the Estimator module interfaced with the ASECalculator enabling seamless integration with the Atomic Simulation Environment (ASE) for energy and force evaluations.
000 MC cycles. To further investigate the impact of MOF framework flexibility on ΔH0,ads of ethylene, simulations were carried out using three distinct structural configurations for each of the 88 MOFs: (i) MOFreference – structures optimized (atomic positions relaxed) in the absence of guest molecules using the EqV2-ODAC u-MLIP model; (ii) MOFgeometry-opt – structures with atomic positions relaxed in the presence of a single ethylene molecule using the PFP u-MLIP; (iii) MOFcell-opt – structures with both atomic positions and unit cell parameters relaxed in the presence of a single ethylene molecule using the PFP u-MLIP. The initial structures for (ii) and (iii) were derived from the minimum energy ethylene adsorption configurations identified during the PFP u-MLIP based Widom insertion simulations. These geometry optimizations were performed using the BFGS optimizer49 with a force convergence threshold of 0.005 eV Å−1. All Widom insertion and geometry optimization calculations using the PFP u-MLIP were performed via the MATLANTIS platform,50 under an academic license. In this framework, the PFP u-MLIP is implemented through the Estimator module interfaced with the ASECalculator enabling seamless integration with the Atomic Simulation Environment (ASE) for energy and force evaluations.
PFP u-MLIP bulk modulus calculations
To evaluate the mechanical properties of the 88 MOFs, we computed their elastic response across an isotropic volumetric strain of ±8% over 10 equidistant steps, utilizing the PFP u-MLIP. At each step, the lattice vectors were scaled to the target volume, and internal atomic positions were relaxed via the BFGS algorithm until the interatomic forces converged below 0.01 eV Å−1. Finally, the bulk modulus was computed by fitting the resulting energy–volume (E–V) data to the Vinet equation of state (EOS). To assess the reliability of the u-MLIP in describing framework mechanical stiffness, A520 was considered as a test-case, with a calculated bulk modulus (4.9 GPa) in excellent agreement with available experimental data (5 GPa).51PFP u-MLIP benchmarking against DFT calculations
To validate the accuracy of the PFP u-MLIP used in this study for modelling MOF–guest interactions, specifically in the case of ethylene and water, the current release of the PFP u-MLIP model (version 6.0.0)33 was rigorously benchmarked against 0 K DFT calculations. The analysis focused on ethylene and water adsorption in 88 top-performing MOFs (average system size of 366 atoms; Fig. S2), with guest positions taken from the lowest-energy adsorption configurations identified via PFP-based Widom insertion MC simulations on MOFreference. Each resulting MOF + guest configuration, containing a single guest molecule per simulation box, was then subjected to PFP u-MLIP and DFT geometry optimization, with only the atomic positions relaxed. The MOF–guest interaction energies (ΔE) for ethylene and water were then evaluated using the following expression:| ΔE = E(MOF + guest) − E(MOF) − E(guest) |
DFT calculations were carried out using the Quickstep module in the CP2K package.52 We used the Perdew–Burke–Ernzerhof (PBE)53 exchange–correlation functional within the Gaussian and plane-wave (GPW) formalism. Double-zeta valence polarized MOLOPT basis sets (DZVP-MOLOPT)54 and Goedecker–Teter–Hutter (GTH) pseudopotentials55 were used. Long-range dispersion interactions were included via Grimme's D3 correction,47 consistent with the treatment employed in the PFP u-MLIP framework. A density cutoff of 450 Ry was applied and the geometry optimizations were performed until maximum residual forces were less than 2 × 10−4 a.u. Å−1.
Results and discussion
UFF-guided identification of MOFs for ethylene capture under humidity
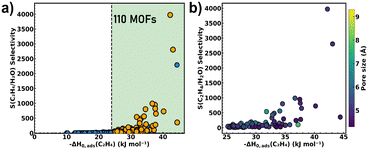
Our multi-stage screening pipeline started with 1881 charge-neutral, computation-ready MOFs, validated via MOSAEC and CoRE MOF protocols and geometry-optimized at the u-MLIP (EqV2-ODAC) level. The hydrophobic character of these MOFs was first assessed by computing their KH,H2O using Widom's test particle insertion MC simulations with UFF. Using ZIF-8 (a well-established benchmark highly hydrophobic MOF with KH,H2O ≈ 2.5 × 10−6 mol kg−1 Pa−1,56) as a reference, we applied a conservative threshold for KH,H2O = 1 × 10−5 mol kg−1 Pa−1 to ensure the selection of a broader range of hydrophobic MOFs. This resulted in a subset of 264 hydrophobic MOFs suitable for ethylene capture in moisture-rich environments.The ethylene/water selective adsorption of this subset of hydrophobic MOFs was then assessed by calculating their ideal selectivity values S(C2H4/H2O) (=KH,C2H4/KH,H2O). MOFs with S(C2H4/H2O) > 1 were considered as promising sorbents for ethylene adsorption over water. Additionally, to ensure practical relevance, we imposed a thermodynamic constraint, requiring the adsorption enthalpy of ethylene (ΔH0,ads(C2H4)) to exceed 25 kJ mol−1 (in absolute value) significantly higher than ethylene's vaporization enthalpy (14 kJ mol−1).57 This criterion ensured strong host–guest interactions, critical for trace ethylene capture. These combined filtering, hydrophobicity, S(C2H4/H2O) > 1 and high C2H4 affinity narrowed the list of potential candidates to 110 high-performance MOFs. These materials, which collectively satisfy all three criteria, are highlighted in the shaded green region of Fig. 2a. To ensure practical applicability, we rigorously evaluated the synthetic feasibility of linkers in the 110 candidate MOFs using the MOFid toolkit.58 Through systematic filtering and manual inspection (Fig. S1), we excluded MOFs containing excessively long linkers (backbone lengths > 3 nm), highly complex architectures including polycyclic or sterically hindered structures, and non-commercially available linkers. This comprehensive screening process eliminated 22 impractical candidates, yielding a final set of 88 synthetically viable MOFs highlighted in orange colour in Fig. 2a. Fig. 2b shows that these selected MOFs predominantly feature moderate pore sizes ranging from 4.5 to 6.0 Å underscoring the importance of pore confinement in promoting selective ethylene adsorption over water. To validate the reliability of this first-pass filter, UFF-predicted ethylene adsorption enthalpies were benchmarked against available experimental isosteric heats of adsorption from the literature, showing overall good agreement (Table S1). In addition, comparison of the UFF-based workflow with the DREIDING force field (Fig. S3) confirms that the relative screening trends are robust, with strong correlations observed for both ethylene adsorption enthalpy (R2 = 0.89) and ideal selectivity (R2 = 0.96), indicating very limited sensitivity to the specific choice of generic non-bonded LJ parameters.
Beyond the LJ parameters, we also assessed the sensitivity of the screening results to the electrostatic description of the MOF/guest interactions by comparing three machine-learning-based charge assignment schemes for MOF atoms: MEPO-ML, PACMAN,59 and PACMOF.60 As shown in Fig. S4, when combined with UFF, the three charge models lead to predicted S(C2H4/H2O) consistently yielding high pairwise correlation coefficients (R2 > 0.94), thereby demonstrating the robustness of our electrostatic description for performance ranking.
Benchmarking u-MLIP against DFT calculations
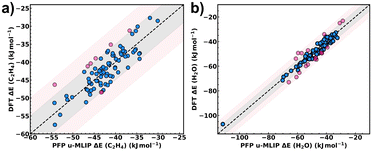
We first validated the applicability of PFP u-MLIP to our test case by benchmarking its predicted interaction energies (ΔE) for both C2H4 and H2O against the values obtained by DFT calculations across the representative set of 88 MOFs. Fig. 3a and b emphasize an excellent agreement for both ethylene and water with very low MAD values of 2.4 kJ mol−1 and 3.0 kJ mol−1 respectively. Notably, 91% of the investigated MOFs exhibit deviations within ±5 kJ mol−1 for ethylene, while 78% MOFs fall within the same range for water. The remaining MOFs show moderate deviations of 5–10 kJ mol−1. This benchmarking also confirms PFP u-MLIP's quantitative reliability in modelling MOF–guest interactions for both polar (H2O) and non-polar (C2H4) guests across diverse MOF frameworks.Comparison of classical generic FF and PFP u-MLIP adsorption predictions
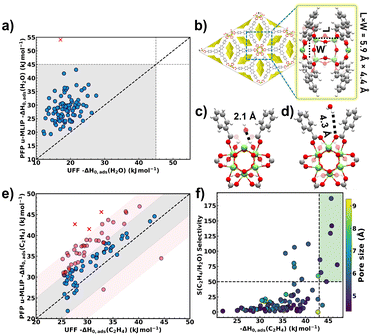
To evaluate the reliability of the conclusions drawn from our initial UFF-based HTCS, we conducted a systematic comparison of adsorption energetics predicted by UFF and those calculated by PFP u-MLIP across the final set of 88 MOF structures. As a critical first validation step, we examined whether PFP u-MLIP confirms the hydrophobic character of these MOFs by computing their ΔH0,ads(H2O) under the same rigid MOF framework approximation applied in the initial UFF calculations.As shown in Fig. 4a, u-MLIP predicts ΔH0,ads(H2O) values below the water vaporization enthalpy threshold (ΔHvap(H2O) ≈ 44 kJ mol−1) for 87 out of 88 MOFs, indicating weak water–MOF affinity and thereby supporting the hydrophobic classification originally established using UFF, which predicted all values below 30 kJ mol−1. Although UFF does not quantitatively agree with the adsorption enthalpy values computed with PFP u-MLIP, it effectively captures the qualitative trend, demonstrating its utility in discriminating hydrophobic MOFs in large-scale screening efforts.
The single MOF outlier ZSTU-3 (ref. 61) (CSD ref. code TOYHOH; marked with a red cross in Fig. 4a) leads to a PFP u-MLIP calculated −ΔH0,ads(H2O) of ∼54 kJ mol−1, which exceeds ΔHvap(H2O), indicating significantly stronger water–MOF framework interactions compared to UFF predictions. The pore architecture of this MOF is shown in Fig. 4b, where polar –OH moieties are pointing towards a narrow pore space of 5.9 Å × 4.4 Å. Typically, this type of pore confinement plays a dominant role in enhancing water/polar functions interactions, an effect UFF fails to accurately capture, and would require tailored LJ parameters,62 which is impractical in a HTCS workflow. Analysis of the lowest-energy adsorption configurations deduced from Widom insertion simulations confirms that PFP u-MLIP captures this confinement effect, placing water molecules as close as 2.1 Å to framework atoms near the –OH moieties (Fig. 4c). In contrast, UFF predicts the nearest interaction at 4.3 Å, occurring at carboxylate oxygen atoms (Fig. 4d).
Following the confirmation of the overall hydrophobic ranking of the MOF candidates, we evaluated their ethylene affinity using both UFF- and PFP u-MLIP-derived ΔH0,ads(C2H4). Fig. 4e reveals a very good quantitative agreement between both values across all 88 MOFs, with a MAD of 5.1 kJ mol−1. This comparative analysis highlights that 52% of the MOFs (blue points) show an excellent correlation between u-MLIP and UFF calculations, with deviations below 5 kJ mol−1. Another 44% (pink points) show moderate deviations in the range of 5–10 kJ mol−1, while only three MOFs (3%, marked with red crosses) exhibit significant deviations greater than 10 kJ mol−1. A closer inspection of these three outlier MOFs: ZTUS-3,61,63 and Fe-CFA-6 (ref. 64) reveals they all contain bridging µ-OH sites confined within angular (‘V-shaped’) pockets defined by two adjacent linkers (Fig. S5). In the PFP u-MLIP simulations, ethylene molecules are positioned in such a way to favour short π(C2H4)⋯OH contacts (∼2.5 Å). In contrast, UFF does not capture this interaction mode, instead predicting different ethylene orientations and associated longer host–guest separating distances (3.0–4.1 Å). These differences in predicted geometries illustrate how the choice of potential model can substantially influence the representation of adsorption environments and associated interaction energies in confined pores. Similar to the adsorbed water geometries discussed above, analysis of the ethylene adsorption modes also revealed noticeable differences in the orientation of ethylene molecules between PFP u-MLIP and UFF, underscoring that classical generic force fields may overlook key directional interactions in spatially constrained environments.
We further evaluated the ethylene/water ideal selectivity, S(C2H4/H2O), using KH,C2H4 and KH,H2O values obtained from the PFP u-MLIP model (Fig. S6). The ranking of the top 99% ethylene selective MOFs remains largely consistent with those derived from UFF predictions, with 87 out of the 88 investigated MOFs exhibiting S(C2H4/H2O) > 1. ZSTU-3 was the sole exception to exhibit a selectivity value of 0.2 arising from a rather hydrophilic behaviour predicted by PFP u-MLIP simulations as discussed above. In general, the absolute values of S(C2H4/H2O) predicted by the PFP u-MLIP (Fig. 4f) are consistently lower than those obtained using UFF (Fig. 2b) mostly due to a more accurate assessment of KH,H2O with PFP u-MLIP paving the way towards a more realistic estimation of the selectivity.
Further analysis of Fig. 4f enables the identification, among these 87 MOFs, of seven top-performing candidates combining high S(C2H4/H2O) > 50 and strong ethylene affinity (−ΔH0,ads(C2H4) > 43 kJ mol−1). These MOFs are JNU-90 (CSD ref. code: NOPGUY65), ZnPF-1 (CSD ref. code: XEGXUF66), EVONOS,67 A520 (CSD ref. code: DOYBEA68), CYCU-5 (CSD ref. code: HONCEU69), ATULIM,70 and PERROW71 (Fig. S7). Structurally, these MOFs typically possess microporous cavities in the range of 4.5–5.5 Å, as indicated by the pore size colour gradient represented in Fig. 4f. The combination of high ethylene/water selectivity and strong ethylene affinity positions these materials as promising candidates for targeted ethylene capture, particularly in low-trace applications such as food preservation.
MOF framework flexibility and its ethylene affinity
Most HTCS workflows inherently assume rigid MOF frameworks, prompting a critical question: to what extent explicit consideration of framework flexibility is essential to refine the prediction of the adsorption properties? To address this question, we equally simulated ΔH0,ads(C2H4) for our shortlisted 88 MOFs via a PFP u-MLIP based Widom insertion MC method, applied to geometry optimized MOFs, relaxed either by atomic positions alone or by both atomic positions and unit cell parameters.The comprehensive analyses presented in Fig. 5a reveal that, for the majority of the MOFs, the relaxation of their atomic positions exerts only a limited effect on ethylene adsorption energetics, largely preserving the overall ranking as predicted using the rigid MOF framework approximation. Specifically, when only atomic positions are relaxed, 98% of the MOFs (86 out of 88) exhibit ΔH0,ads(C2H4) variations within 5 kJ mol−1, with a very low MAD of 0.72 kJ mol−1. Only two structures show a more substantial deviation that remains however moderate in the range of 5–10 kJ mol−1. This supports those tiny localized atomic rearrangements have negligible influence on ethylene affinity for this MOF set.
In the case of full structure optimization, which includes both atomic positions and unit cell parameter relaxation, our u-MLIP calculations revealed that a significant fraction of the 88 MOFs show a substantial unit cell volume change upon C2H4 adsorption. To better understand the origin of these adsorption-induced structural responses, we evaluated the intrinsic mechanical stiffness of the 88 MOFs through bulk modulus calculations at the u-MLIP level (Fig. S8). The results show that larger adsorption-induced volume changes tend to occur preferentially in MOF frameworks with low to moderate bulk moduli as shown in Fig. 6, consistent with a higher ability for geometrical deformation in mechanically compliant MOFs.
 | ||
| Fig. 6 Correlation between C2H4-induced absolute unit cell volume change (|ΔV|%) and bulk modulus (B0) predicted for the 88 MOFs by PFP u-MLIP. | ||
Fig. 5b further shows that 80% of MOFs (70 out of 88) still show moderate ΔH0,ads(C2H4) changes within 5 kJ mol−1. 17 MOFs exhibit deviations of 5–10 kJ mol−1, and just one structure shows a deviation exceeding 10 kJ mol−1. Correspondingly, the MAD increases to 2.4 kJ mol−1, which however remains modest overall but clearly reflects the larger variability introduced by lattice flexibility for certain MOF frameworks. These results highlight the critical role of lattice flexibility in modulating host–guest interactions, particularly for MOFs whose structural adaptability significantly influences adsorption behaviour. Microscopic insight into these effects is provided in Fig. 5c, which compares rigid and fully cell-optimized configurations for the outlier MOF QAQTEJ72 (red cross in Fig. 5b), a layered structure which displays a distinct flexibility mode: while its internal square nets remain largely intact, unit cell optimization triggers a lateral shift of the layers, reducing the interlayer spacing from 9 to 6 Å. This structural rearrangement results in 35% reduction in unit cell volume and increases −ΔH0,ads(C2H4) from 31 to 51 kJ mol−1. The enhancement in adsorption enthalpy can be attributed to increased confinement and stronger π⋯π interactions between ethylene and the pyridine rings of the linkers, with contact distances shortening from 4.5 Å to 3.6 Å.
Notably, among the seven identified top-performing MOFs, six of them display small volume changes of 1–4% leading to a tiny variation of their adsorption enthalpy of ethylene (0.4–4 kJ mol−1) (Table S2). In addition, while A520 (CSD ref. code: DOYBEA68) undergoes a substantial relative volume variation of 19%, yet the change in adsorption enthalpy is only 0.39 kJ mol−1. In the relaxed state, the framework retains nearly identical host–guest geometries, with –OH⋯π interactions (2.6 → 2.1 Å) and C–H⋯π contacts (2.9 → 3.1 Å) showing only marginal adjustments, as illustrated in Fig. 5c.
Conclusions
This work presents a scalable, multi-scale HTCS framework that couples the speed of classical force fields with the quantum-level accuracy of universal machine learning potentials, validated against DFT, for the identification of high-performance MOF adsorbents. This strategy uses ethylene capture under humid conditions as a challenging test case, which is highly relevant to food-preservation packaging technologies. We demonstrate how classical u-IPs and u-MLIPs can be synergistically integrated to achieve both computational efficiency and accurate predictions for a large library of MOFs.Specifically, our results show that classical u-IP methods like UFF-based HTCS remain a robust first-pass filter, correctly capturing adsorption trends in ∼97–99% of cases under the rigid-MOF framework approximation. u-MLIPs are essential for further refining the adsorption predictions for top-performing candidates, especially for MOFs involving strong hydrogen bonding, directional π⋯H/π⋯π interactions, or guest-induced framework flexibility, where deviations in ΔH0,ads(C2H4) can reach ∼20 kJ mol−1. This workflow enables the identification of top MOF performers combining microporous confinement (4.5–6.0 Å), strong ethylene affinity (ΔH0,ads(C2H4) > 43 kJ mol−1), and high selectivity (S(C2H4/H2O) > 50). These computational findings are expected to prioritize MOFs for experimental validation and accelerate their integration into scalable postharvest ethylene management technologies. Beyond ethylene capture, the proposed hybrid classical generic FF/u-MLIP strategy offers a transferable computational workflow for other adsorption-driven separations, enabling the screening of large structural databases while retaining the resolution needed to treat complex host–guest interactions and framework flexibility.
Author contributions
G. M. conceived the idea and supervised the project. S. B. performed and interpreted the computational calculations. M. W. developed the u-MLIP-based Widom simulation code and contributed to the analysis of the simulated data. S. B. wrote the original draft with input from M. W. and G. M. T. W. and K. H. provided support with the MATLANTIS framework and contributed to manuscript improvement.Conflicts of interest
There are no conflicts to declare.Data availability
The code supporting the PFP u-MLIP-based Widom simulations and the data (Excel file) generated in this study are available on GitHub (https://github.com/gmaurin-group/MLP-WIDOM-SIM).Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d6sc00831c.
Acknowledgements
The computational work was performed using HPC resources from GENCI-CINES (grant A0180907613) and MATLANTIS. G. M. thanks Institut Universitaire de France for the Senior Chair.Notes and references
- P. Z. Moghadam, A. Li, S. B. Wiggin, A. Tao, A. G. P. Maloney, P. A. Wood, S. C. Ward and D. Fairen-Jimenez, Chem. Mater., 2017, 29, 2618–2625 Search PubMed.
- H. Daglar and S. Keskin, Coord. Chem. Rev., 2020, 422, 213470 CrossRef CAS.
- Y. Qiu, L. Wang, C. Liu, X. Zhang, Y. Tian and Z. Zhou, Mater. Today, 2025, 91, 103–123 CrossRef CAS.
- S. Ga, N. An, G. Y. Lee, C. Joo and J. Kim, Renewable Sustainable Energy Rev., 2024, 192, 114275 CrossRef CAS.
- A. K. Rappe, C. J. Casewit, K. S. Colwell, W. A. Goddard and W. M. Skiff, J. Am. Chem. Soc., 1992, 114, 10024–10035 CrossRef CAS.
- M. Fischer, J. R. B. Gomes, M. Fröba and M. Jorge, Langmuir, 2012, 28, 8537–8549 CrossRef CAS PubMed.
- L.-C. Lin, K. Lee, L. Gagliardi, J. B. Neaton and B. Smit, J. Chem. Theory Comput., 2014, 10, 1477–1488 CrossRef CAS PubMed.
- A. L. Dzubak, L.-C. Lin, J. Kim, J. A. Swisher, R. Poloni, S. N. Maximoff, B. Smit and L. Gagliardi, Nat. Chem., 2012, 4, 810–816 CrossRef CAS PubMed.
- A. R. Kulkarni and D. S. Sholl, J. Phys. Chem. C, 2016, 120, 23044–23054 CrossRef CAS.
- J. W. Yoon, H. Chang, S.-J. Lee, Y. K. Hwang, D.-Y. Hong, S.-K. Lee, J. S. Lee, S. Jang, T.-U. Yoon, K. Kwac, Y. Jung, R. S. Pillai, F. Faucher, A. Vimont, M. Daturi, G. Férey, C. Serre, G. Maurin, Y.-S. Bae and J.-S. Chang, Nat. Mater., 2017, 16, 526–531 Search PubMed.
- E. Martínez-Ahumada, D. won Kim, M. Wahiduzzaman, P. Carmona-Monroy, A. López-Olvera, D. R. Williams, V. Martis, H. A. Lara-García, S. López-Morales, D. Solis-Ibarra, G. Maurin, I. A. Ibarra and C. S. Hong, J. Mater. Chem. A, 2022, 10, 18636–18643 RSC.
- H. Kraß, J. Huang and S. M. Moosavi, MOFSimBench: Evaluating Universal Machine Learning Interatomic Potentials in Metal–Organic Framework Molecular Modeling, npj Comput. Mater., 2026, 12, 4 CrossRef PubMed.
- R. Goeminne and V. Van Speybroeck, J. Am. Chem. Soc., 2025, 147, 3615–3630 CrossRef CAS PubMed.
- D. Fan, F. L. Oliveira, M. Wahiduzzaman and G. Maurin, Decoding local framework dynamics in the ultra-small pore MOF MIL-120(Al) CO2 sorbent with Machine Learned Potentials, Nat. Commun., 2026, 17, 3235 CrossRef PubMed.
- S. Liu, R. Dupuis, D. Fan, S. Benzaria, M. Bonneau, P. Bhatt, M. Eddaoudi and G. Maurin, Chem. Sci., 2024, 15, 5294–5302 RSC.
- I. Batatia, P. Benner, Y. Chiang, A. M. Elena, D. P. Kovács, J. Riebesell, X. R. Advincula, M. Asta, M. Avaylon, W. J. Baldwin, F. Berger, N. Bernstein, A. Bhowmik, F. Bigi, S. M. Blau, V. Cărare, M. Ceriotti, S. Chong, J. P. Darby, S. De, F. Della Pia, V. L. Deringer, R. Elijošius, Z. El-Machachi, E. Fako, F. Falcioni, A. C. Ferrari, J. L. A. Gardner, M. J. Gawkowski, A. Genreith-Schriever, J. George, R. E. A. Goodall, J. Grandel, C. P. Grey, P. Grigorev, S. Han, W. Handley, H. H. Heenen, K. Hermansson, C. H. Ho, S. Hofmann, C. Holm, J. Jaafar, K. S. Jakob, H. Jung, V. Kapil, A. D. Kaplan, N. Karimitari, J. R. Kermode, P. Kourtis, N. Kroupa, J. Kullgren, M. C. Kuner, D. Kuryla, G. Liepuoniute, C. Lin, J. T. Margraf, I.-B. Magdău, A. Michaelides, J. H. Moore, A. A. Naik, S. P. Niblett, S. W. Norwood, N. O'Neill, C. Ortner, K. A. Persson, K. Reuter, A. S. Rosen, L. A. M. Rosset, L. L. Schaaf, C. Schran, B. X. Shi, E. Sivonxay, T. K. Stenczel, C. Sutton, V. Svahn, T. D. Swinburne, J. Tilly, C. van der Oord, S. Vargas, E. Varga-Umbrich, T. Vegge, M. Vondrák, Y. Wang, W. C. Witt, T. Wolf, F. Zills and G. Csányi, J. Chem. Phys., 2025, 163, 184110 CrossRef CAS PubMed.
- C. Chen and S. P. Ong, Nat. Comput. Sci., 2022, 2, 718–728 CrossRef PubMed.
- B. Deng, P. Zhong, K. Jun, J. Riebesell, K. Han, C. J. Bartel and G. Ceder, Nat. Mach. Intell., 2023, 5, 1031–1041 CrossRef.
- A. Sriram, L. M. Brabson, X. Yu, S. Choi, K. Abdelmaqsoud, E. Moubarak, P. de Haan, S. Löwe, J. Brehmer, J. R. Kitchin, M. Welling, C. L. Zitnick, Z. Ulissi, A. J. Medford and D. S. Sholl, arXiv, 2025, preprint, arXiv:2508.03162, DOI:10.48550/arxiv.2508.03162.
- A. Sriram, S. Choi, X. Yu, L. M. Brabson, A. Das, Z. Ulissi, M. Uyttendaele, A. J. Medford and D. S. Sholl, ACS Cent. Sci., 2024, 10, 923–941 Search PubMed.
- X. Fu, B. M. Wood, L. Barroso-Luque, D. S. Levine, M. Gao, M. Dzamba and C. L. Zitnick, arXiv, 2025, preprint, arXiv:2502.12147, DOI:10.48550/arxiv.2502.12147.
- Y. Lim, H. Park, A. Walsh and J. Kim, Matter, 2025, 8, 102203 CrossRef CAS.
- C. S. Barry and J. J. Giovannoni, J. Plant Growth Regul., 2007, 26, 143–159 CrossRef CAS.
- N. Y. State, C. B. Watkins, J. F. Nock and J. Reid, Production Guide for Storage of Organic Fruits and Vegetables, 2012 Search PubMed.
- J. Gustavsson, C. Cederberg, U. Sonesson, R. van Otterdijk and A. Meybeck, Food and Agriculture Organization of the United Nations, 2011 Search PubMed.
- L.-Y. Chang and J. K. Brecht, Sci. Hortic., 2023, 309, 111636 Search PubMed.
- M. H. Álvarez-Hernández, G. B. Martínez-Hernández, F. Avalos-Belmontes, F. D. Miranda-Molina and F. Artés-Hernández, Postharvest Biol. Technol., 2020, 160, 111061 CrossRef.
- H. Li, F. Li, L. Wang, J. Sheng, Z. Xin, L. Zhao, H. Xiao, Y. Zheng and Q. Hu, Food Chem., 2009, 114, 547–552 CrossRef CAS.
- W. You, Y. Liu, J. D. Howe and D. S. Sholl, J. Phys. Chem. C, 2018, 122, 8960–8966 CrossRef CAS.
- Y. Qi, H. Yang, C. Li and H. Li, Sep. Purif. Technol., 2024, 347, 127634 CrossRef CAS.
- Z. Zhang, R. Zhou, L. Ke, J. Li, H. Jayan, H. R. El-Seedi, X. Zou and Z. Guo, Trends Food Sci. Technol., 2024, 154, 104771 CrossRef CAS.
- H. Li, L. Shi, C. Li, X. Fu, Q. Huang and B. Zhang, ACS Appl. Mater. Interfaces, 2020, 12, 34095–34104 Search PubMed.
- S. Takamoto, C. Shinagawa, D. Motoki, K. Nakago, W. Li, I. Kurata, T. Watanabe, Y. Yayama, H. Iriguchi, Y. Asano, T. Onodera, T. Ishii, T. Kudo, H. Ono, R. Sawada, R. Ishitani, M. Ong, T. Yamaguchi, T. Kataoka, A. Hayashi, N. Charoenphakdee and T. Ibuka, Nat. Commun., 2022, 13, 2991 CrossRef CAS PubMed.
- L. M. Brabson, A. J. Medford and D. S. Sholl, J. Phys. Chem. C, 2025, 129, 16811–16825 CrossRef CAS PubMed.
- X. Jin, K. M. Jablonka, E. Moubarak, Y. Li and B. Smit, Digital Discovery, 2025, 4, 1560–1569 RSC.
- A. S. Rosen, S. M. Iyer, D. Ray, Z. Yao, A. Aspuru-Guzik, L. Gagliardi, J. M. Notestein and R. Q. Snurr, Matter, 2021, 4, 1578–1597 CrossRef CAS.
- M. Gibaldi, A. Kapeliukha, A. White, J. Luo, R. A. Mayo, J. Burner and T. K. Woo, Chem. Sci., 2025, 16, 4085–4100 RSC.
- M. Gibaldi, A. Kapeliukha, A. White and T. K. Woo, J. Chem. Inf. Model., 2025, 65, 275–287 Search PubMed.
- M. Gibaldi, A. Kapeliukha, A. White, J. Luo, R. A. Mayo, J. Burner and T. K. Woo, Chem. Sci., 2025, 16, 4085–4100 Search PubMed.
- A. Hjorth Larsen, J. Jørgen Mortensen, J. Blomqvist, I. E. Castelli, R. Christensen, M. Dułak, J. Friis, M. N. Groves, B. Hammer, C. Hargus, E. D. Hermes, P. C. Jennings, P. Bjerre Jensen, J. Kermode, J. R. Kitchin, E. Leonhard Kolsbjerg, J. Kubal, K. Kaasbjerg, S. Lysgaard, J. Bergmann Maronsson, T. Maxson, T. Olsen, L. Pastewka, A. Peterson, C. Rostgaard, J. Schiøtz, O. Schütt, M. Strange, K. S. Thygesen, T. Vegge, L. Vilhelmsen, M. Walter, Z. Zeng and K. W. Jacobsen, J. Phys.: Condens. Matter, 2017, 29, 273002 Search PubMed.
- D. Frenkel and B. Smit, Understanding Molecular Simulation: From Algorithms to Applications, Academic Press, San Diego, 2nd edn, 2002, vol. 1 Search PubMed.
- P. P. Ewald, Ann. Phys., 1921, 369, 253–287 Search PubMed.
- J. Luo, O. Ben Said, P. Xie, M. Gibaldi, J. Burner, C. Pereira and T. K. Woo, npj Comput. Mater., 2024, 10, 224 CrossRef CAS PubMed.
- B. Liu, B. Smit, F. Rey, S. Valencia and S. Calero, J. Phys. Chem. C, 2008, 112, 2492–2498 Search PubMed.
- C. Vega, J. L. F. Abascal and I. Nezbeda, J. Chem. Phys., 2006, 125, 034503 CrossRef CAS PubMed.
- Q. Yang and C. Zhong, J. Phys. Chem. B, 2006, 110, 17776–17783 CrossRef CAS PubMed.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, J. Chem. Phys., 2010, 132, 154104 CrossRef PubMed.
- S. Grimme, S. Ehrlich and L. Goerigk, J. Comput. Chem., 2011, 32, 1456–1465 Search PubMed.
- R. Battiti and F. Masulli, in International Neural Network Conference: July 9–13, 1990, Palais Des Congres—Paris—France, Springer, Netherlands, Dordrecht, 1990, pp. 757–760 Search PubMed.
- MATLANTIS, Preferred Computational Chemistry Inc., https://matlantis.com/ Search PubMed.
- P. Ramaswamy, J. Wieme, E. Alvarez, L. Vanduyfhuys, J.-P. Itié, P. Fabry, V. Van Speybroeck, C. Serre, P. G. Yot and G. Maurin, J. Mater. Chem. A, 2017, 5, 11047–11054 RSC.
- J. Hutter, M. Iannuzzi, F. Schiffmann and J. VandeVondele, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2014, 4, 15–25 CAS.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1996, 77, 3865–3868 CrossRef CAS PubMed.
- J. VandeVondele and J. Hutter, J. Chem. Phys., 2007, 127, 114105 CrossRef.
- S. Goedecker, M. Teter and J. Hutter, Phys. Rev. B: Condens. Matter Mater. Phys., 1996, 54, 1703–1710 CrossRef CAS PubMed.
- G. Ercakir, G. O. Aksu, C. Altintas and S. Keskin, ACS Eng. Au, 2023, 3, 488–497 CrossRef CAS PubMed.
- R. M. Stephenson and S. Malanowski, Handbook of the Thermodynamics of Organic Compounds, Springer, Dordrecht, 1st edn, 1987 Search PubMed.
- B. J. Bucior, A. S. Rosen, M. Haranczyk, Z. Yao, M. E. Ziebel, O. K. Farha, J. T. Hupp, J. I. Siepmann, A. Aspuru-Guzik and R. Q. Snurr, Cryst. Growth Des., 2019, 19, 6682–6697 CrossRef CAS.
- G. Zhao and Y. G. Chung, J. Chem. Theory Comput., 2024, 20, 5368–5380 CrossRef CAS.
- S. Kancharlapalli, A. Gopalan, M. Haranczyk and R. Q. Snurr, J. Chem. Theory Comput., 2021, 17, 3052–3064 CrossRef CAS PubMed.
- C. Li, H. Xu, J. Gao, W. Du, L. Shangguan, X. Zhang, R.-B. Lin, H. Wu, W. Zhou, X. Liu, J. Yao and B. Chen, J. Mater. Chem. A, 2019, 7, 11928–11933 RSC.
- J. Shen, A. Kumar, M. Wahiduzzaman, D. Barpaga, G. Maurin and R. K. Motkuri, Chem. Rev., 2024, 124, 7619–7673 CrossRef CAS.
- E. Alvarez, N. Guillou, C. Martineau, B. Bueken, B. Van de Voorde, C. Le Guillouzer, P. Fabry, F. Nouar, F. Taulelle, D. de Vos, J.-S. Chang, K. H. Cho, N. Ramsahye, T. Devic, M. Daturi, G. Maurin and C. Serre, Angew. Chem., Int. Ed., 2015, 54, 3664–3668 CrossRef CAS PubMed.
- S. Spirkl, M. Grzywa, C. S. Zehe, J. Senker, S. Demeshko, F. Meyer, S. Riegg and D. Volkmer, CrystEngComm, 2015, 17, 313–322 Search PubMed.
- Y. Wang, Z.-J. Jiang, D.-R. Wang, W. Lu and D. Li, J. Am. Chem. Soc., 2024, 146, 6955–6961 CrossRef CAS PubMed.
- C. Castillo-Blas, V. A. de la Peña-O’Shea, I. Puente-Orench, J. R. de Paz, R. Sáez-Puche, E. Gutiérrez-Puebla, F. Gándara and Á. Monge, Sci. Adv., 2025, 3, e1700773 CrossRef PubMed.
- Y. Xu, R.-H. Wang, B.-Y. Lou, L. Han and M.-C. Hong, Acta Crystallogr., Sect. C: Cryst. Struct. Commun., 2004, 60, m296–m298 Search PubMed.
- E. Alvarez, N. Guillou, C. Martineau, B. Bueken, B. Van de Voorde, C. Le Guillouzer, P. Fabry, F. Nouar, F. Taulelle, D. de Vos, J.-S. Chang, K. H. Cho, N. Ramsahye, T. Devic, M. Daturi, G. Maurin and C. Serre, Angew. Chem., Int. Ed., 2015, 54, 3664–3668 CrossRef CAS PubMed.
- C.-Y. Wu, D. S. Raja, C.-C. Yang, C.-T. Yeh, Y.-R. Chen, C.-Y. Li, B.-T. Ko and C.-H. Lin, CrystEngComm, 2014, 16, 9308–9319 Search PubMed.
- M. Armaghan, X.-J. Shang, Y.-Q. Yuan, D. J. Young, W.-H. Zhang, T. S. A. Hor and J.-P. Lang, ChemPlusChem, 2015, 80, 1231–1234 CrossRef CAS PubMed.
- X.-G. Liu, C.-L. Tao, H.-Q. Yu, B. Chen, Z. Liu, G.-P. Zhu, Z. Zhao, L. Shen and B. Z. Tang, J. Mater. Chem. C, 2018, 6, 2983–2988 RSC.
- C. J. Adams, M. C. Muñoz, R. E. Waddington and J. A. Real, Inorg. Chem., 2011, 50, 10633–10642 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |