Open Access Article
Open Access ArticleMolecular basis for the biosynthetic divergence of anthracyclines kosinostatin and chartreusin
Ling Li
Fang†
a,
Chen Hao
Jia†
a,
Xuan
Zhou
a,
Jun Ying
Ma
a,
Huan Zi
Guo
a,
Xiuxiu
Ma
d,
Wei
Bai
a,
Lang
Xiang
a,
Jiapeng
Zhu
*bc,
Ren Xiang
Tan
 *ad and
Yi Shuang
Wang
*ad and
Yi Shuang
Wang
 *a
*a
aSynthetic Biology Center for Chinese Medicine, School of Pharmacy, Nanjing University of Chinese Medicine, Nanjing 210023, China. E-mail: wangys@njucm.edu.cn
bDepartment of Hepatobiliary Surgery, The First Affiliated Hospital of Anhui Medical University, Hefei, Anhui 230001, China
cMOE Innovation Center for Basic Research in Tumor Immunotherapy, Anhui Province Key Laboratory of Tumor Immune Microenvironment and Immunotherapy, Anhui Provincial Innovation Institute for Pharmaceutical Basic Research, Hefei, Anhui 230001, China
dState Key Laboratory of Pharmaceutical Biotechnology, Institute of Functional Biomolecules, School of Life Sciences, Nanjing University, 210023, China
First published on 26th February 2026
Abstract
Kosinostatin and chartreusin are both anthracycline-type antitumor agents. The structure of kosinostatin incorporates a spirally fused pyrrolopyrrole moiety attached to a five-membered ring of its anthraquinone core, whereas chartreusin possesses a distinct, rearranged pentacyclic aromatic dilactone aglycone. Biosynthetic divergence in kosinostatin and chartreusin is governed by highly homologous proteins, KstA15 and ChaU, which catalyze phenol hydroxylation and hydration, respectively. In this report, we successfully engineered a functional swap between ChaU and KstA15 via directed evolution. The crystal structure of the ChaU mutant along with molecular dynamics simulations elucidated the structural determinants of its catalytic specificity, revealing the critical residue interactions and conformational dynamics. Furthermore, a ubiquitous distribution of KstA15-like proteins across microbial genomes has been demonstrated through gene mining. These findings deliver key insights into the evolutionary trajectory of functionally divergent enzymes and create a robust framework for rationally engineering biosynthetic pathways to produce novel anthracycline derivatives with potential pharmaceutical applications.
Introduction
Anthracyclines are microbially derived natural products defined by a broad antitumor spectrum, potent efficacy, and proven clinical effectiveness, which underpin their extensive use in cancer treatment regimens.1–3 Representative anthracycline drugs include doxorubicin, daunorubicin, and aclarubicin.4–8 In terms of biosynthetic origin, anthracyclines are aromatic type II polyketides. Their carbon skeleton is assembled through nine iterative rounds of Claisen condensation between malonyl-CoA and either an acetyl-CoA or propionyl-CoA starter unit. The resulting polyketide chain is subsequently processed by conserved enzymes including ketoreductase, cyclase, and aromatase to form the characteristic tetracyclic anthracyclinone core.9–11 The structural diversity observed in anthracyclines primarily arises from a series of tailoring redox reactions that introduce differential modifications onto this common scaffold.12–18Among anthracycline derivatives, two highly oxidized molecules, kosinostatin and chartreusin, exhibit significant structural differences.19–22 Nevertheless, both originate from the common precursor auromycinone (3), and their biosynthetic pathways bifurcate at a critical step regulated by two homologous ester cyclases, KstA15 and ChaU (Fig. 1).23,24 KstA15 and its partner protein KstA16 are encoded within the kst gene cluster responsible for kosinostatin biosynthesis, whereas ChaU and its corresponding protein ChaX are found in the cha gene cluster dedicated to chartreusin construction (Fig. 1).24–26 The KstA15-KstA16 complex utilizes oxygen as a substrate to catalyze C8 phenyl ring hydroxylation, whereas the ChaU-ChaX pair promotes D-ring dehydration via a hydration step (Fig. 1). In these two catalytic systems, KstA16 and ChaX are both members of the short-chain reductase (SDR) family, sharing high sequence homology and performing identical enzymatic functions in the reduction of the para-quinone moiety in anthracyclines.24 While both KstA15 and ChaU belong to the nuclear transport factor 2 (NTF2) superfamily, they exhibit a high degree of sequence homology but markedly divergent biological functions (Fig. 1).24 The crystal structures of ChaU and KstA15 (along with its functional homolog SnoaL2) have been determined.23,24,27 Site-directed mutagenesis studies demonstrated that the catalytically essential residues in ChaU include D119, R50, E38, and S37. In contrast, the active sites in SnoaL2 are identified as H35, E106, H119, and D121, which correspond to H49, E119, H131, and D133 in KstA15 (Fig. 1).23,24 Despite their high structural similarity, the molecular basis underlying the distinct catalytic functions of ChaU and KstA15 remains elusive. Here, we identified the critical amino acid combinations that govern the catalytic functional specificity of ChaU and KstA15, thereby facilitating a switch between C8 phenol hydroxylation and their hydration activities. Through integration of the crystal structure of a hydroxylation-competent ChaU mutant with molecular dynamics (MD) simulations, we elucidated the molecular basis that drives the functional divergence toward hydration versus phenol hydroxylation in this enzyme family. Furthermore, our work reveals that homologous proteins capable of catalyzing phenol hydroxylation are widely distributed among diverse microorganisms, whereas enzymes within this class that retain hydratase activity are scarcely represented, showing the occurrence of proteins annotated as “ester cyclases” in the NTF2 superfamily.
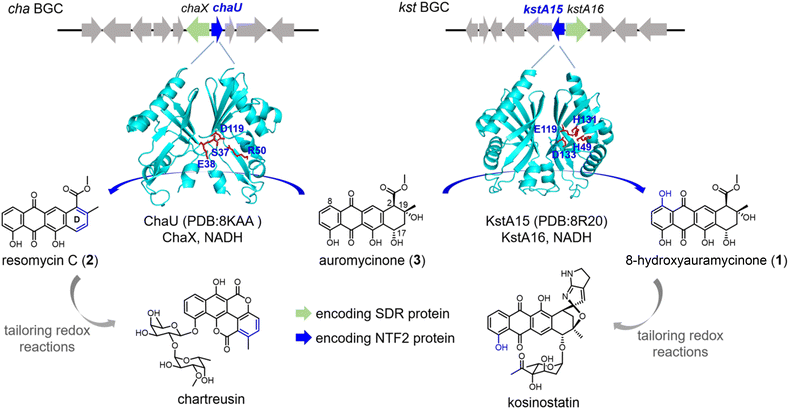 | ||
| Fig. 1 ChaU and KstA15: two annotated ester cyclases determining the biosynthetic branching points to chartreusin and kosinostatin. | ||
Results and discussion
Elucidating the core set of amino acids that bridge the catalytic properties of ChaU and KstA15
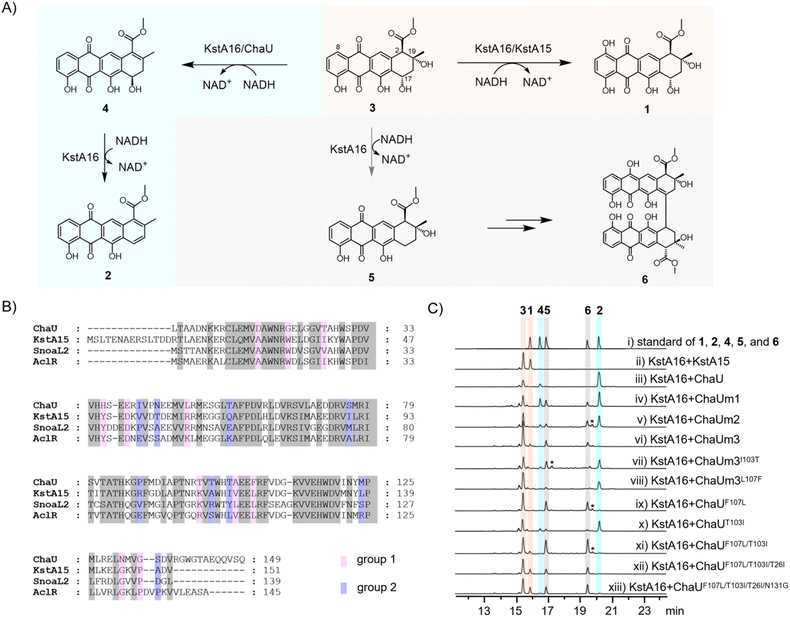
KstA15 and KstA16 jointly catalyze the hydroxylation of 3 at the C-8 position to form 1, while ChaX and ChaU together mediate the conversion of 3 to yield 2, proceeding through intermediate 4via a (de)hydration reaction (Fig. 2A). Individually, either KstA16 or ChaX catalyzes the 17-dehydroxylation of 3, producing 5 and a dimer (6), whose structure has been proposed based on molecular weight (Fig. 2A). Given the higher expression level of KstA16 relative to its isoenzyme ChaX, KstA16 was selected as the interacting partner for all subsequent catalytic reactions involving ChaU or KstA15-like proteins (Fig. S1). To pinpoint the specific amino acid residues that account for the divergent enzymatic functions of ChaU and KstA15, we chose to subject ChaU to directed evolution for phenol hydroxylation. Thus, we performed sequence alignment of ChaU with its homologous proteins (KstA15, SnoaL2, and AclR) known to catalyze such reactions (Fig. 2A and B).23 This analysis identified non-conserved residues, which we systematically categorized into two groups: “Relatively non-conserved” (a common amino acid in the homologs that differs in ChaU, group 1), and “Completely non-conserved” (divergent amino acids across all four proteins, group 2) (Fig. 2B). The initial substitution of group 1 amino acid residues in ChaU with their counterparts from KstA15 produced the mutant ChaUm1, which contains 11 mutations (Fig. 2B and S2). However, ChaUm1 failed to gain phenol hydroxylation activity and instead retained its original hydration activity to convert 3 to 4, 2, 5 and 6, the structures of which were determined by comparing their 1H NMR and HR-MS data with those previously reported (Fig. 2C(i–iv), S16–S24 and Tables S6–S9),14,24 indicating that key residues essential for phenol hydroxylation are not fully conserved among KstA15, SnoaL2, and AclR. Building upon this mutant, we performed a sequential and pairwise replacement of the “group 2” amino acids in ChaU with their KstA15 equivalents, starting from the C-terminus of the protein (Fig. 2B and S2). Surprisingly, while ChaUm2 (ChaUm1S135A/M124L) remained incapable of hydroxylating phenol (Fig. 2C(v)), the ChaUm3 (ChaUm2T103I/T100A) mutant successfully acquired the catalytic function of KstA15, allowing it to cooperate with KstA16 to convert 3 into 1 (Fig. 2C(vi), S14–S15 and Table S5).Given that the ChaUm3 mutant carries 15 amino acid substitutions, it is probable that not all residues contribute equally to its functional repertoire for C8 phenol hydroxylation. Therefore, we systematically generated a series of revertant mutants, in which each of the fifteen substituted residues was individually changed back to its wild-type counterpart in ChaU (Fig. S2). Specifically, both the ChaUm3I103T and ChaUm3L107F revertants lost phenol hydroxylation function (Fig. 2C(vii–viii)), thereby establishing that these two specific residues are essential for determining the catalytic divergence between hydration and C8 phenol hydroxylation among ChaU-like enzymes. Subsequently, we introduced single (ChaUF107L and ChaUT103I) and double (ChaUF107L/T103I) mutations into the wild-type ChaU. The data demonstrated that only the ChaUF107L/T103I double mutant could catalyze C8 phenol hydroxylation (Fig. 2C(ix–xi)), further underscoring that these two residues constitute the minimal determinant for shifting the catalytic activity of ChaU from hydration to phenol hydroxylation.
Given the limited catalytic activity of the dual mutant ChaUF107L/T103I toward phenol hydroxylation at this stage, we proceeded to generate a series of triple and quadruple mutants through its combinatorial substitution with each amino acid residue from group 1 (Fig. S2). The results demonstrated a substantial enhancement in the efficiency of phenol hydroxylation for the triple mutant ChaUF107L/T103I/T26I and quadruple mutant ChaUF107L/T103I/T26I/N131G (Fig. 2C(xii–xiii)). To confirm that these four residues are indeed critical for phenol hydroxylation, we performed site-directed mutagenesis at the equivalent positions in KstA15. Mutation of the four amino acids in KstA15 to their corresponding residues in ChaU revealed that KstA15I117T and KstA15L121F completely lost phenol hydroxylation activity, while that of KstA15I40T and KstA15 G145N was substantially diminished (Fig. S3). Notably, all four single mutants acquired the ability to catalyze the hydration reaction to yield 2 (Fig. S3). This suggested that phenol hydroxylation and hydration reactions are intricately related and the former necessitates the contribution of more key amino acid residues to retain its catalytic integrity, compared to the simpler hydration catalysis. If so, the amino acid residues critical for hydration may also impact C8 phenol hydroxylation. This hypothesis was tested by substituting the residues in KstA15 that align with ChaU's key hydration sites with alanine.24 As a result, the mutants KstA15S51A, KstA15E52A, KstA15R64A and KstA15D133A indeed lost 50–60% catalytic activity towards C8 phenol hydroxylation and retained no hydratase function (Fig. S4). These data underscored the precision of evolutionarily refined enzymatic strategies that direct specific transformations in anthracycline antibiotic biosynthesis.
Insights into the catalytic divergence between ChaU and the ChaUF107L/T103I/T26I/N131G mutant
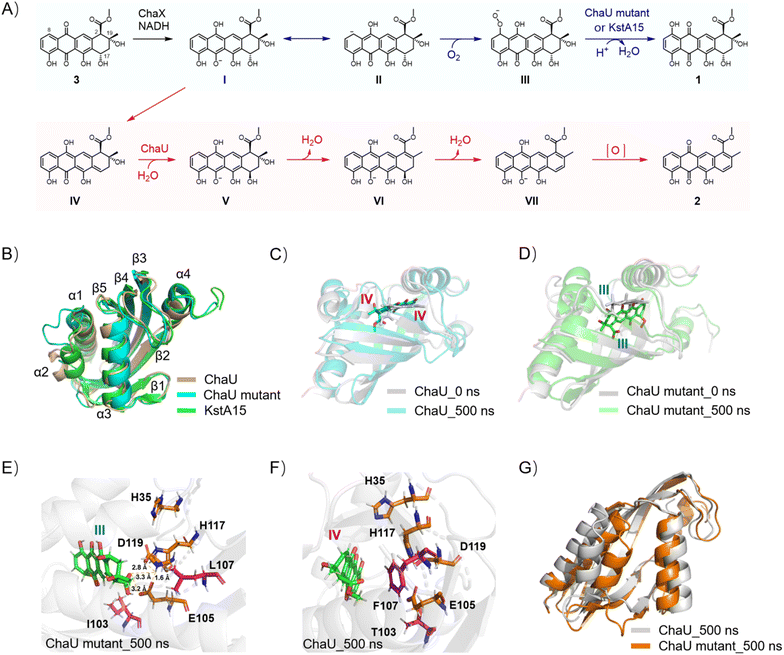
Given that the ChaUF107L/T103I/T26I/N131G mutant shares the same catalytic function for phenol hydroxylation as KstA15, we first obtained the crystal of this quadruple mutant as a tetramer, and determined its structure at a resolution of 2.70 Å (Fig. S5 and Table S4). A comparison of the crystal structures of ChaU, ChaUF107L/T103I/T26I/N131G and KstA15 indicated a conserved overall tertiary fold, while marked conformational deviation is localized to the α2-helix region (Fig. 3B). To delineate the mechanistic basis for the catalytic divergence between ChaU and ChaUF107L/T103I/T26I/N131G mutants, we performed molecular docking followed by molecular dynamics (MD) simulations. As shown in Fig. 3, a pronounced contrast in substrate-binding dynamics was observed between wild-type ChaU and ChaUF107L/T103I/T26I/N131G mutants via MD simulations. While the substrate IV (the putative direct substrate of ChaU24) retained a pose similar to its initial configuration with stability in WT ChaU (Fig. 3A and C), the mutant system exhibited a marked shift: the substrate III (the presumed direct substrate of KstA15 (ref. 14 and 23)) reoriented substantially around 80 ns, adopting a distinct binding mode that remained stable thereafter (Fig. 3A and D and S6–S7). To elucidate the structural determinants of this divergence, we analyzed hydrogen-bond interactions between the substrate and four ChaU residues (H35, E105, H117 and D119), which were selected based on their spatial positioning and functional analogy to catalytic residues in KstA15.23 Notably, substrate III in the ChaUF107L/T103I/T26I/N131G mutant sustained at least one hydrogen bond with these residues throughout most of the simulation (Fig. 3E and S8). Conversely, equivalent hydrogen-bond interactions were essentially absent in the WT ChaU system (Fig. 3F and S8). These results suggested that the mutation alters the substrate-binding microenvironment of ChaU, leading to the stabilization of a substrate conformation that is proximal to key reactive residues. This reoriented pose thereby satisfies the spatial necessary for a KstA15-type reaction, providing a plausible mechanistic basis for the observed functional shift in the ChaU mutant, which was further supported by the marked decrease in phenol hydroxylation efficiency exhibited by the variants ChaUF107L/T103I/T26I/N131G/E105A and ChaUF107L/T103I/T26I/N131G/H117A (Fig. S4). Consistent with this model, conformational analysis indicated that the mutant enzyme preferentially adopts a more closed conformation upon substrate binding compared to the WT ChaU (Fig. 3G and S9), presumably diminishing the spatial distance between the substrate and catalytic residues, which in turn consolidates the hydrogen-bonding network and thereby facilitating the establishment of a preorganized, catalytically competent architecture. We noted that in wild-type ChaU, the conformation of E105 is stabilized by a hydrogen bond with T103 and significant steric hindrance from F107 (Fig. S10). In the ChaU mutant, however, the substitution to I103 abolishes this hydrogen bond, and the replacement with L107 eliminates the steric restriction (Fig. S10). This releases E105 from its constrained state, allowing it to move closer to the incoming substrate and thereby tightening the binding cavity. The resultant contraction of the α-helical structure further enables W100 to form a hydrogen bond with the substrate (Fig. S10). Such changes synergistically promote hydroxylation at the C8 position of the substrate, driving the reaction forward.To enable a more direct comparison of the phenol hydroxylation mechanisms between the ChaUF107L/T103I/T26I/N131G mutant and KstA15, molecular docking and MD simulations of substrate III bound to KstA15 were also performed. Notably, a difference in substrate-binding conformation is observed between the ChaUF107L/T103I/T26I/N131G mutant and KstA15 (Fig. 3E and S11), despite their identical catalytic functions.23 Within KstA15, H131 (the counterpart of H117 in the ChaUF107L/T103I/T26I/N131G mutant) is positioned sufficiently close to the peroxide anion, especially during the first 80 ns, to allow direct transfer of HE2 or HD1 in H131 to the peroxide anion moiety (Fig. S11 and S12). However in the ChaUF107L/T103I/T26I/N131G mutant, H117 and E105 are located adjacent to the carboxyl group of III (Fig. 3E and S11), likely contributing to substrate stabilization. This hinted the presence of another proton transfer channel in the ChaUF107L/T103I/T26I/N131G mutant catalytic systems, potentially accounting for the approximate three-fold higher KstA15 catalytic activity relative to ChaUF107L/T103I/T26I/N131G (Fig. 2C(ii, xiii) and S4).
Collectively, these findings indicated that the mutation orchestrates a coordinated reorganization across multiple levels—including substrate binding, residue interaction networks, and global conformational dynamics—rather than simply enhancing local substrate interactions. These integrated, multiscale changes collectively promote the emergence of structural and dynamic properties in the ChaU mutant that resemble those essential for KstA15-like catalysis, thereby providing a mechanistic basis for its altered reactivity.
Broader natural occurrence of KstA15-like versus ChaU-like proteins
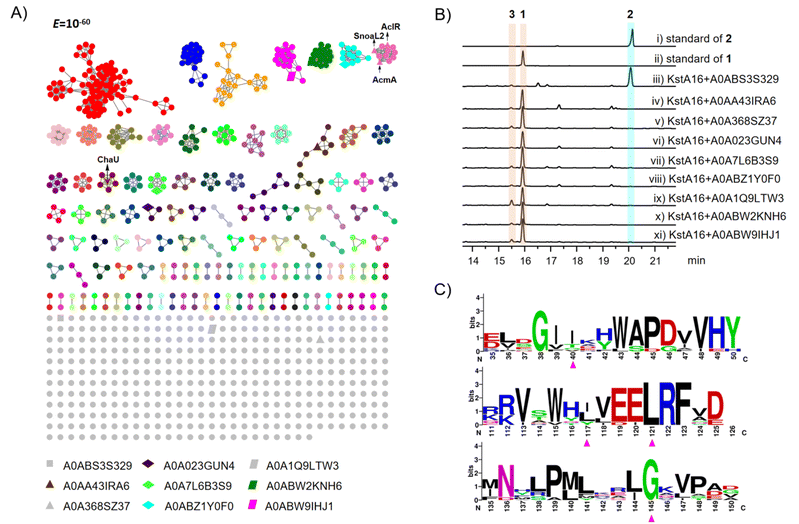
Despite the high sequence homology between KstA15 and ChaU, KstA15-like proteins, including SnoaL2, AclR, and SwaR1, have been identified in various anthracycline biosynthetic gene clusters,23,28,29 while, ChaU represents the first NFT2 superfamily member reported to possess a hydratase function.24 To explore the distribution of ChaU-like proteins across the microbial genome, we attempted to mine databases for them. Thus, using ChaU as a query, we searched against microbial proteomes in the UniProt database and retrieved 946 homologous sequences. A sequence similarity network (SSN) was constructed with these sequences (Fig. 4A).30,31 Application of an E-value cutoff of 10−60 resolved ChaU-like and KstA15-like enzymes into two distinct clusters within the network (Fig. 4A).In an attempt to identify ChaU-like enzymes within other clades, we randomly selected 9 proteins from distinct branches of diverse microorganisms for heterologous expression in Escherichia coli (Fig. 4A). Co-catalytic assays with KstA16 demonstrated that only one protein (UniProt ID: A0ABS3S329) exhibited hydratase activity, whereas all others catalyzed specifically the C8-hydroxylation reaction (Fig. 4B). Aligning the 9 selected proteins against ChaU, KstA15, AclR, and SnoaL2 revealed that the ChaU residues T103 and F107 are functionally conserved in homolog A0ABS3S329 capable of catalyzing the hydration reaction (Fig. S13). A similar conservation pattern was observed for the corresponding residues (I/L117 and L121) in KstA15 and its functionally homologous proteins (Fig. 4C and S13). Residues T26 and N131 in ChaU are not conserved across its isozyme A0ABS3S329 (Fig. S13). Intriguingly, A0ABS3S329 shares the identical amino acid at position 26 with the corresponding residue in KstA15 (Fig. S13). Although position 131 is also variable between ChaU and A0ABS3S329, the corresponding site in KstA15 (position 145) and its isozymes is consistently a conserved glycine (Fig. 4C and S13). These findings reinforce the conclusion that residues 103 and 107, along with residue131 in ChaU, play a pivotal role in dictating its functional specificity as either a hydratase or a hydroxylase.
Conclusions
In conclusion, we pinpointed two key amino acid residues responsible for the distinct catalytic activities of ChaU and KstA15 through a combined experimental approach involving sequence alignment and site-directed mutagenesis. This critical finding allowed us to rationally convert ChaU from a hydratase into a phenyl ring hydroxylase. Furthermore, protein crystallography, along with molecular dynamics simulations, ultimately elucidated the precise molecular basis for their catalytic divergence. Our study offers a paradigm for elucidating the evolutionary mechanisms of natural enzymes and establishes a foundational framework for the rational engineering of biosynthetic pathways.Author contributions
Y. S. W. designed the research and wrote the manuscript; R. X. T. provided financial support; J. Z. directed the protein crystallographic analysis; L. L. F., C. H. J., J. Y. M., H. Z. G. and X. M. performed the biochemical experiments; Y. S. W., W. B. and L. X. analyzed the data.Conflicts of interest
The authors declare no conflicts of interest.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: Tables S1–S9, Fig. S1–S24, NMR spectra and further experimental details. See DOI: https://doi.org/10.1039/d6sc00451b.Acknowledgements
This work was co-financed by the NSFC (No. 82003608), the Jiangsu Annual Basic Science (Natural Science) Research Projects of Higher Education Institutions (No. 25KJB350006), the Jiangsu Commission of Health (No. QN202405), Nanjing University of Chinese Medicine (No. ZYXJC2024-003 and No. JC202402), Jiangsu Provincial Youth Science Foundation Project grant (BK20220473) and the Interdisciplinary Funding for Synthetic Biology and Traditional Chinese Medicine by Nanjing University of Chinese Medicine.Notes and references
- M. B. Hulst, T. Grocholski, J. J. C. Neefjes, G. P. van Wezel and M. Metsä-Ketelä, Nat. Prod. Rep., 2022, 39, 814–841 Search PubMed.
- C. Hertweck, A. Luzhetskyy, Y. Rebets and A. Bechthold, Nat. Prod. Rep., 2007, 24, 162–190 Search PubMed.
- M. K. Kharel, P. Pahari, M. D. Shepherd, N. Tibrewal, S. E. Nybo, K. A. Shaaban and J. Rohr, Nat. Prod. Rep., 2012, 29, 264–325 RSC.
- D. P. A. Wander, S. Y. van der Zanden, G. A. van der Marel, H. S. Overkleeft, J. Neefjes and J. D. C. Codée, J. Med. Chem., 2020, 63, 12814–12829 CrossRef CAS PubMed.
- A. Bisht, D. Avinash, K. K. Sahu, P. Patel, G. Das Gupta and B. D. Kurmi, Drug Deliv. Transl. Res., 2025, 15, 102–133 CrossRef PubMed.
- F. Wang, L. Zhao, Y. Tan, X. Cen, H. Gao, H. Jiang, Y. Liu, Y. Li, T. Zhang, C. Zhao, T. Shi, G. Xu, C. Wang, J. Hu, X. Li, Y. Z. Qin, K. Wang, H. H. Zhu and K. Li, Nat. Commun., 2025, 16, 617 CrossRef CAS PubMed.
- E. R. Abels, E. Ter Linden, J. H. T. Rohling, L. M. Voortman, L. Janssen, M. Hornsveld, H. S. Overkleeft, S. Y. van der Zanden, M. L. D. Broekman and J. Neefjes, Proc. Natl. Acad. Sci. U. S. A., 2025, 122, e2510263122 CrossRef CAS PubMed.
- J. Yang, Q. Liu, X. Zhang, Y. Jing, N. Le, M. Li, L. Xu, W. Zhao, S. Huang, D. Liu and L. Dou, Int. Immunopharmacol., 2025, 151, 114268 CrossRef CAS PubMed.
- X. Zhu, V. Siitonen, C. E. Melançon Iii and M. Metsä-Ketelä, ACS Synth. Biol., 2021, 10, 243–251 CrossRef CAS PubMed.
- G. Wang, J. Chen, H. Zhu and J. Rohr, Org. Lett., 2017, 19, 540–543 CrossRef CAS PubMed.
- L. Y. Li, Y. L. Hu, J. L. Sun, L. B. Yu, J. Shi, Z. R. Wang, Z. K. Guo, B. Zhang, W. J. Guo, R. X. Tan and H. M. Ge, Chem. Sci., 2022, 13, 12892–12898 RSC.
- Y. S. Wang, B. Zhang, J. Zhu, C. L. Yang, Y. Guo, C. L. Liu, F. Liu, H. Huang, S. Zhao, Y. Liang, R. H. Jiao, R. X. Tan and H. M. Ge, J. Am. Chem. Soc., 2018, 140, 10909–10914 CrossRef CAS PubMed.
- F. W. Jiao, Y. S. Wang, X. T. You, W. Wei, Y. Chen, C. L. Yang, Z. K. Guo, B. Zhang, Y. Liang, R. X. Tan, R. H. Jiao and H. M. Ge, Angew. Chem., Int. Ed., 2021, 60, 26378–26384 CrossRef CAS PubMed.
- Z. Zhang, Y. K. Gong, Q. Zhou, Y. Hu, H. M. Ma, Y. S. Chen, Y. Igarashi, L. Pan and G. L. Tang, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 1554–1559 CrossRef CAS PubMed.
- B. Nji Wandi, V. Siitonen, K. Palmu and M. Metsä-Ketelä, Chembiochem, 2020, 21, 3062–3066 CrossRef CAS PubMed.
- L. Xiang, J. Shi, A. Zhu, Z. F. Xu, S. H. Liu, Y. S. Wang, Z. K. Guo, R. H. Jiao, R. X. Tan and H. M. Ge, Angew. Chem., Int. Ed., 2023, 62, e202218660 CrossRef CAS PubMed.
- S. K. Pang, Curr. Cancer Drug Targets, 2018, 18, 600–607 CrossRef CAS PubMed.
- F. F. Qi, W. Zhang, Y. Y. Xue, C. Geng, X. N. Huang, J. Sun and X. F. Lu, J. Am. Chem. Soc., 2021, 143, 16326–16331 CrossRef CAS PubMed.
- Y. Shimizu, S. Takahashi, T. Iwata, H. Shimooka, T. Okauchi and M. Kitamura, J. Org. Chem., 2025, 90, 9291–9294 CrossRef CAS PubMed.
- Y. S. Wang, W. Zheng, N. Jiang, Y. X. Jin, Z. K. Meng, M. X. Sun, Y. L. Zong, T. Xu, J. Zhu and R. X. Tan, Angew. Chem., Int. Ed., 2022, 61, e202201321 CrossRef CAS PubMed.
- H. M. Ma, Q. Zhou, Y. M. Tang, Z. Zhang, Y. S. Chen, H. Y. He, H. X. Pan, M. C. Tang, J. F. Gao, S. Y. Zhao, Y. Igarashi and G. L. Tang, Chem. Biol., 2013, 20, 796–805 CrossRef CAS PubMed.
- Z. Xu, K. Jakobi, K. Welzel and C. Hertweck, Chem. Biol., 2005, 12, 579–588 CrossRef CAS PubMed.
- B. N. Wandi, P. Dinis, V. Siitonen, G. Schneider, R. Schnell and M. Metsä-Ketelä, ACS Catal., 2024, 14, 12359–12371 CrossRef CAS.
- Y. S. Wang, H. W. Chen, Z. K. Meng, C. H. Jia, J. Q. Han, Y. X. Jin, X. X. Ma, N. X. Wang, J. P. Zhu, Y. Liang and R. X. Tan, ACS Catal., 2025, 15, 19253–19267 CrossRef CAS.
- Z. Zhang, Y. K. Gong, Q. Zhou, Y. Hu, H. M. Ma, Y. S. Chen, Y. Igarashi, L. Pan and G. L. Tang, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 1554–1559 CrossRef CAS PubMed.
- M. Niemczura, A. Nuutila, R. Wang, K. Rauhanen, S. E. Nybo and M. Metsä-Ketelä, ACS Chem. Biol., 2025, 20, 1457–1463 CrossRef CAS PubMed.
- P. Beinker, B. Lohkamp, T. Peltonen, J. Niemi, P. Mäntsälä and G. Schneider, J. Mol. Biol., 2006, 359, 728–740 CrossRef CAS PubMed.
- H. Ma, Q. Zhou, Y. Tang, Z. Zhang, Y. Chen, H. He, H. Pan, M. Tang, J. Gao, S. Zhao, Y. Igarashi and G. Tang, Chem. Biol., 2013, 20, 796–805 CrossRef CAS PubMed.
- M. Niemczura, A. Nuutila, R. Wang, K. Rauhanen, S. E. Nybo and M. Metsä-Ketelä, ACS Chem. Biol., 2025, 20, 1457–1463 CrossRef CAS PubMed.
- Y. Hu, J. Chen, S. Qi, H. Wang, Z. Zhu, Y. Peng, W. Wang, G. Huang, Z. Fang, Y. Ye, Z. Wang and K. Guo, Angew. Chem., Int. Ed., 2025, 64, e202501425 CrossRef CAS PubMed.
- K. H. O'Toole, B. Imperiali and K. N. Allen, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2018289118 CrossRef PubMed.
Footnote |
| † Contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |