Open Access Article
Open Access ArticleThe hidden language of gut-derived lipopolysaccharides: fine chemistry, huge immunological consequences
Marcello
Mercogliano
a,
Valentina
Mazziotti
ab,
Alba
Silipo
 ab,
Antonio
Molinaro
ab,
Antonio
Molinaro
 ab and
Flaviana
Di Lorenzo
ab and
Flaviana
Di Lorenzo
 *ab
*ab
aDepartment of Chemical Sciences and Task Force for Microbiome Studies, University of Naples Federico II, Via Cinthia 4, Naples, 80126, Italy. E-mail: flaviana.dilorenzo@unina.it
bCEINGE Biotecnologie Avanzate Franco Salvatore, Via Gaetano Salvatore, 486, Naples, 80131, Italy
First published on 4th February 2026
Abstract
Lipopolysaccharides (LPSs) from Gram-negative bacteria are traditionally viewed as potent “endotoxins” recognized by the immune system and capable of triggering robust inflammation. However, increasing evidence from gut commensals is dismantling this one-dimensional view. The gastrointestinal tract is indeed the major reservoir of LPSs, owing to the dense Gram-negative community inhabiting the small and large intestine, with total weight in healthy individuals estimated to exceed one gram. This necessarily means that the mere presence of LPSs cannot be directly linked to inflammation. Moreover, chronic exposure to low-potency or atypical LPSs can recalibrate innate immunity, fostering tolerance or, conversely, failing to provide adequate tonic stimulation and thereby predisposing the system to aberrant activation. Understanding this delicate balance and the structural and cellular mechanisms that sustain it, is essential to interpret the immunological impact of the gut LPSs in health and disease. In this Perspective, we highlight recent advances revealing the remarkable chemical diversity of commensal-derived LPSs and illustrate how subtle variations in LPS lipid A acylation and phosphorylation, core oligosaccharide architecture, O-antigen composition, and overall supramolecular organization profoundly rewire receptor usage and downstream immune outcomes. These insights underscore the enormous, still largely untapped potential of gut LPS chemistry to reveal unifying structural hallmarks that distinguish inflammatory, tolerogenic, and immunologically “tuned” features. Although fragments of this logic are beginning to emerge, a comprehensive framework remains urgently needed. Decoding the chemical language adopted by LPSs in the gut will be essential to reclassify LPSs not merely as dangerous molecules, but as a potential source of immunomodulators and as a blueprint for next-generation tools enabling precision control of host–microbe interactions.
Introduction
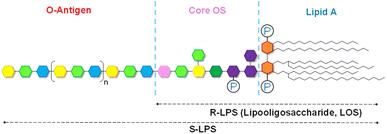
Lipopolysaccharides (LPSs) are major constituents of the outer leaflet of the Gram-negative bacterial outer membrane and key mediators of host–microbe interactions. They are widely regarded as among the microbial molecules with a profound impact on the host immune system.1 Owing to this central role, LPSs hold considerable promise for translational applications across medicine, immunology, and vaccinology.1–3 An LPS exerts its action through interaction with pattern-recognition receptors (PRRs) expressed by innate immune cells, thereby triggering activation of multiple signal transduction cascades.4,5 Among these, the best characterized is the pathway initiated by binding of an LPS molecule to the Toll-Like Receptor 4/Myeloid Differentiation Protein-2 (TLR4/MD-2) complex, which represents the “canonical” axis of LPS sensing.1,4,6 Upon binding to the TLR4/MD-2 complex, LPS drives the formation of an activated TLR4/MD-2/LPS dimer, which triggers intracellular signaling, most prominently the NF-κB cascade, by inducing conformational rearrangements in the Toll/IL-1R (TIR) domains of TLR4 and culminating in the transcription of pro-inflammatory cytokines and immune mediators.7 Consequently, based largely on studies of pathogens, LPSs have historically been regarded as potent pro-inflammatory molecules, known as “endotoxins” and considered potential threats to human health.8 Nevertheless, and fortunately, the magnitude of the inflammatory response they induce is inextricably linked to their chemical structure.9–11 The region of an LPS that primarily engages the TLR4/MD-2 complex is the lipid A: the membrane-anchoring glycolipid consisting of a β(1→6)-linked diglucosamine backbone substituted with phosphate groups (typically at position 4′ of the non-reducing glucosamine and at position 1 of the reducing one) as well as acyl chains of varying lengths and number (Fig. 1). Beyond lipid A, a smooth-type LPS (S-LPS) also comprises a heteropolysaccharide composed of the core oligosaccharide (core OS) and the O-polysaccharide domain (or O-antigen). In contrast, when the O-antigen is absent, the molecule is referred to as rough-type LPS (R-LPS) or lipooligosaccharide (LOS) (Fig. 1).1,9 Although lipid A represents the immunodominant and most bioactive portion of an LPS, its fine structure varies substantially across species, strains, and even within a single bacterial population.1 Such variations profoundly influence TLR4 engagement and ultimately dictate the strength and quality of the ensuing immune response. The core OS (for R-LPSs) and the O-antigen (for S-LPSs) are the most diverse segments of the molecule and major determinants of antigenic specificity, virulence potential, and the capacity of Gram-negative bacteria to attach to, colonize, or evade the host.12,13 These structural domains also contribute to the intrinsic antimicrobial resistance of Gram-negative bacteria by forming a permeability barrier at the outer membrane, underscoring the dual role of LPSs as both immune triggers and a physical shield.1,14Over the years, therefore, it has become increasingly evident that LPS chemistry and biology are far more complex than initially assumed. The molecular architecture of an LPS is highly variable: each of its domains can differ markedly in chemical composition depending on the bacterial species, environmental cues, and host-associated factors. These structural variations not only allow bacteria to adapt and thrive in specific niches, but also profoundly reshape immune recognition by engaging a broad spectrum of receptors and signaling pathways. Consequently, LPS molecules can elicit remarkably diverse immunological outputs, ranging from robust inflammatory responses to weak activation or even complete immunological silence.9,15 Despite decades of investigation, the mechanisms by which different LPS structures trigger, or suppress, innate immune responses remain only partially understood. What is becoming increasingly evident, however, is the existence of a complex interaction network in which a given LPS engages multiple immune receptors, challenging the classical view of TLR4 as the sole, or even primary, sensor in all contexts.6,16 This underscores an urgent need to delineate how an LPS, on the basis of its structural characteristics, engages alternative signaling pathways that, although still poorly characterized, may play crucial roles in fine-tuning immune responses.6 Identifying the key structural determinants and related signaling pathways activated is essential for developing new biomedical strategies with far-reaching implications for human health. Yet, despite this tremendous biomedical potential, the structure–function landscape of LPSs still remains surprisingly underexplored, representing one of the most pressing and promising frontiers in (glyco)immunology. One reason for this knowledge gap is that, to date, most studies have disproportionately focused on the pro-inflammatory activity of LPSs, often using Escherichia coli LPS as a representative model for the entire class. In contrast, far less attention has been devoted to LPSs from non-pathogenic bacteria, including those that inhabit our intestines, i.e. the gut microbiota. This microbial community, densely populated and highly diverse, plays a central role in host physiology, supporting nutrient digestion, metabolite production, metabolic regulation, and, critically, the development and calibration of the immune system.17 A continuous bidirectional dialogue between microbiota and host immunity dictates whether responses remain tolerogenic or shift toward inflammation. When this equilibrium is perturbed, a state referred to as dysbiosis, microbiota–immune interactions can skew toward pathology,18–20 contributing to metabolic disorders such as obesity and type 2 diabetes,21 chronic inflammatory diseases,22 autoimmune conditions,23,24 and allergy.25 Within this framework, a central question in (glyco)immunology emerges: how are commensal and mutualistic bacteria recognized by the immune system, and how do their LPSs guide the immune system toward homeostasis rather than inflammation? Indeed, circulating gut-derived LPSs can be detected in healthy individuals without triggering overt inflammation,26 thus further highlighting that not all LPSs are created equal. Their chemical structures and, consequently, the receptors they engage dictate whether they promote tolerance, immune education, or pathology. This highlights the need to reevaluate the statement that circulating LPSs directly cause inflammatory responses. But then, what is the role of non-inflammatory LPSs? Are they merely passive molecular bystanders, active modulators of immune tone, or unrecognized contributors to immune training and disease susceptibility? And, given their abundance and capacity to tune immune responses, should certain gut-derived LPSs be considered potential postbiotics rather than “endotoxins”?
This Perspective aims to spotlight the chemical and biological significance of recently characterized “harmless” LPSs expressed by gut commensals. We examine how their structural features underpin unconventional immunomodulatory activities and summarize what is currently known, and still unresolved, about the signaling pathways they engage. Deciphering these mechanisms and the structural determinants that shape specific immune outcomes will be crucial for identifying new therapeutic targets and for developing selective immunomodulators with translational potential.
Beyond the pro-inflammatory paradigm: the case of Bacteroides spp. LPSs
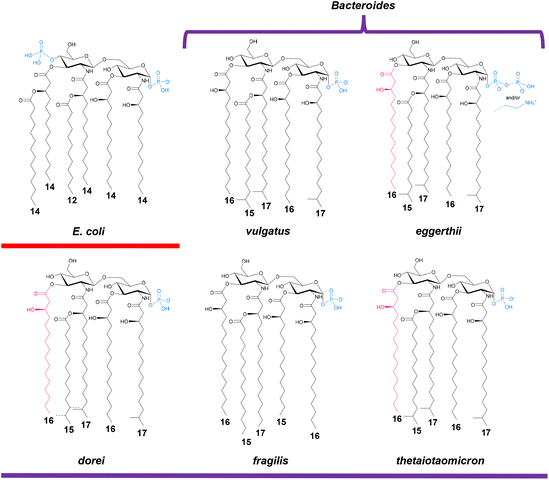
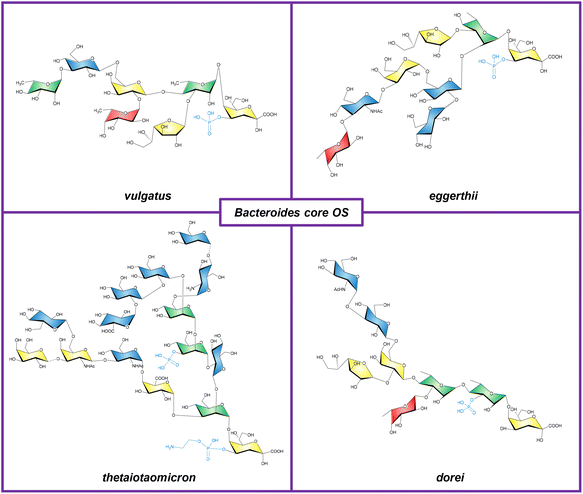
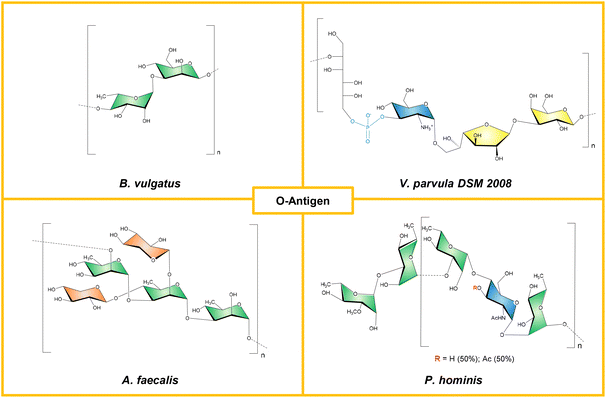
A growing body of evidence shows that many commensal and mutualistic bacteria produce LPS variants whose molecular architecture is tuned to promote immune tolerance rather than inflammation.9,27 Although a unified framework for distinguishing pro-inflammatory from “tolerogenic” LPSs is still lacking, emerging patterns are remarkably consistent: hypo-acylation and reduced phosphorylation in the lipid A are commonly found in microbiota-derived lipid A and correlated with weakened TLR4 engagement and attenuated signaling.9 This is particularly evident in Bacteroides species (Fig. 2). Despite the limited number of fully elucidated structures, Bacteroides spp. remain one of the most intriguing, and widespread, sources of immunomodulatory LPSs within the human gut.28 As dominant early colonizers, they contribute to gut ecological stability through nutrient biosynthesis, metabolic cooperation, toxin neutralization, and competitive exclusion of pathogens.29–32 Across the genus, including B. vulgatus, B. eggerthii, B. dorei, B. fragilis, and B. thetaiotaomicron and others, LPS lipid A structures consistently depart from the canonical hexa-acylated blueprint of E. coli (Fig. 2). Missing acyl chains and, in some cases, reduced phosphorylation impede the precise “lock-and-key” interactions required to stabilize the TLR4/MD-2 heterotetramer. Without this stabilization the full downstream signaling is compromised, resulting in markedly attenuated or entirely absent pro-inflammatory responses.The case of B. vulgatus (recently reclassified as Phocaeicola vulgatus) illustrates this principle with exceptional clarity. Its LPS lipid A consists of a mixture of tetra- and penta-acylated species carrying a single phosphate group at position 1 of the reducing glucosamine (Fig. 2).33,34 The absence of the 4′ phosphate, known to reduce endotoxic activity by up to two orders of magnitude,35 further contributes to the weak agonistic profile of this LPS. Notably, no additional negative charge sources are found in this LPS apart from the phosphate group on the 3-deoxy-D-manno-oct-2-ulosonic acid (Kdo) unit (the “sugar marker” of LPS molecules), further contributing to the weak receptor engagement and subsequent inflammatory signaling activation. Consistently, experimental studies demonstrated that B. vulgatus LPS elicits minimal pro-inflammatory cytokine release in human dendritic cells while promoting elevated anti-inflammatory IL-10 production, a hallmark of tolerogenic immune engagement.33 This was an unprecedented observation for an LPS molecule and became even more compelling in light of in vivo findings showing that, in mouse models of colitis, B. vulgatus LPS alleviates intestinal inflammation by inducing a distinct form of LPS tolerance.36 Mechanistically, this effect appears to operate through the TLR4/MD-2 axis in lamina propria CD11c+ cells, pointing to a specialized, tissue-adapted tolerogenic response driven by this commensal-derived LPS.36 While lipid A provides the primary determinant of attenuated TLR4 activation, unusual features in the saccharide region of B. vulgatus LPS add further layers of complexity. Notably, as shown in Fig. 3, the core OS lacks the heptose residue typically found directly linked to the Kdo unit. In heptose-deficient LPS, this position is commonly occupied by an α-mannose residue; however, in B. vulgatus it is instead replaced by a 6-deoxy-α-mannose, an α-rhamnose. The occurrence of rhamnose at this position is not an isolated peculiarity, but rather a conserved feature shared by all Bacteroides LPS structures characterized to date, with the exception of B. thetaiotaomicron (Fig. 3).37 The functional significance of this conserved substitution remains unknown, but its recurrence strongly suggests an evolutionarily selected role for its glycosyltransferase. However, it is also tempting to speculate that α-rhamnose may represent an early structural cue of a core OS configured for non-canonical receptor engagement. Adding further interest, this rhamnose residue is itself substituted with another highly unusual sugar for LPS core regions: β-galactofuranose (Fig. 3). Strikingly, galactofuranose has also been identified in the core OS of B. eggerthii and B. dorei (now Phocaeicola dorei), whose complete structural elucidation has only recently been achieved (Fig. 3).38,39 As observed for rhamnose, galactofuranose appears to be a recurring feature of Bacteroides LPS core OSs, again with the exception of B. thetaiotaomicron, suggesting a non-random and potentially functional role that warrants further investigation. One plausible hypothesis is that galactofuranose acts as a molecular signal that biases host immune recognition toward a more tolerogenic program, thereby enhancing bacterial fitness within the intestinal environment.36 In this frame, it has been shown that the C-type lectin DC-SIGN, expressed on dendritic cells and regulatory M2 macrophages,40 recognizes specific epitopes within the B. vulgatus LPS structure, thus possibly playing a key role in shaping immune responses. Unlike TLR4, this lectin does not directly activate NF-κB, while it modulates TLR-dependent signaling and can skew responses toward IL-10 production, anti-inflammatory programs, and tissue repair when engaged by particular glycan signatures,41–43 including those present in B. vulgatus LPS.33,44 Supporting this view, an elegant study using synthetic fragments of B. vulgatus LPS revealed that several core OS residues, not only galactofuranose but also fucose, galactose, and Kdo, contribute to DC-SIGN recognition.44 More recently, an integrated approach combining chemical synthesis of core OS derivatives, STD Nuclear Magnetic Resonance (NMR) spectroscopy, and molecular simulations elucidated the structural determinants underlying this interaction.45 The study demonstrated that fucosylated ligands play a dominant role through a specific fucose-calcium coordination and a subsequent hydrogen-bonding network within the DC-SIGN carbohydrate-recognition domain.45 However, galactofuranose further modulates the conformation and presentation of the binding epitope, fine-tuning receptor engagement.45 These insights highlight a finely tuned glycan-lectin code and reinforce the need to pinpoint the minimal structural motifs capable of driving such immunomodulatory effects. Adding another layer of complexity to an already intricate signaling landscape, B. vulgatus LPS has also been shown to activate TLR2 in addition to modulating TLR4. Using human embryonic kidney (HEK)293 cells engineered to co-express human TLR4 and TLR2, it was observed that B. vulgatus LPS induces stronger NF-κB activation than in cells expressing either receptor alone, suggesting a cooperative or synergistic signaling mechanism.33 As a matter of fact, the formation of TLR2/TLR4 heterodimers in response to non-canonical LPS structures has been already proposed by Francisco et al.46 whose molecular docking analyses of LPS from the human opportunistic pathogen Ochrobactrum intermedium predicted a favorable TLR2/TLR4/MD-2 heterodimeric interface, a configuration subsequently validated experimentally by fluorescence resonance energy transfer in live cells.46 Notably, the uncommon core OS structure of O. intermedium LPS emerged as a key contributor to this interaction, further highlighting how subtle chemical features in LPSs can diversify receptor engagement and downstream immune signalling.46 However, whether B. vulgatus LPS can actually induce TLR2/TLR4 heterodimerization, and through which structural features, remain unresolved. Of note, other Bacteroides LPSs, such as B. eggerthii LOS and B. dorei LPS, also show weak inflammatory profiles and only limited TLR4 activation, even though B. eggerthii contains minor lipid A species further decorated with an additional phosphate and an aminoethyl group (Fig. 2).38 Yet, despite sharing the same dominant lipid A species and carrying non-canonical core OS signatures, i.e. they express fucose, galactofuranose, and rhamnose, these LPS molecules appear unable to trigger TLR2 signaling. Though, it should be noted that functional assays in cellular systems co-expressing both TLR2 and TLR4 have not yet been performed for these LPSs, and such studies will be essential to clarify receptor usage and to unravel the principles governing this intriguing signaling divergence.
One important member of the Bacteroides genus is B. fragilis, which constitutes 1–2% of the normal intestinal microflora.47,48 It plays multiple roles within the host, largely due to the production of diverse bioactive molecules, including a peculiar LPS and several capsular polysaccharides such as polysaccharide A, both of which have drawn considerable attention for their immunological relevance.49–52 Beyond the typical hypo-acylated and mono-phosphorylated lipid A species shared across Bacteroides spp., Yang and co-workers recently provided a near-complete characterization of the B. fragilis core OS using top-down and tandem mass spectrometry (MS), showcasing the power of this analytical approach.50 Their work revealed a complex core OS architecture extending from Kdo and comprising polygalactose and deoxy-hexoses, with potential hexuronic acid residues and 2-aminoethyl phosphate (PEtN) modifications on the lipid A moiety.50 The immunological behavior of B. fragilis LPS closely mirrors that of B. vulgatus. In vitro, it suppresses cytokine production in THP-1 monocytes and human dendritic cells, displaying markedly reduced immunostimulatory activity compared to E. coli LPS and, under certain conditions, even functioning as a TLR4 antagonist.51,52 In collagen antibody-induced arthritis models, B. fragilis LPS showed a significantly diminished capacity to induce disease and could even prevent its development when administered at higher doses.52 Additional evidence in healthy mice and E. coli LPS-primed mice confirmed these effects: B. fragilis LPS lowered TNF-α levels, increased IL-10 release, restored regulatory T-cell populations, without amplifying NF-κB signaling.51,53 Nevertheless, the precise molecular epitopes underlying this behavior remain undefined, highlighting the need for structure to function studies of this characteristic LPS.
Famous, and perhaps even more intriguing than B. fragilis and B. vulgatus, is B. thetaiotaomicron, a deeply studied human gut symbiont capable of producing at least eight distinct capsular polysaccharides that directly shape host–microbe crosstalk, although the underlying mechanisms remain incompletely understood. The first structural elucidation of B. thetaiotaomicron LOS was achieved using a mutant strain in which all gene clusters encoding the extensive capsular polysaccharide layer had been deleted.37,54,55 This strategy revealed an unprecedented LOS architecture that diverges radically from previously characterized core OS structures: two complex oligosaccharide branches emanating from an α-mannose linked to Kdo and incorporating uronic acids, PEtN, and an uncommon non-acetylated glucosamine residue that locally modulates charge (Fig. 3).37 Such an arrangement generates a finely tuned electrostatic landscape that might represent a charge-masking strategy adopted by bacteria to persist in the colonized host.56 Although this structural characterization provides a compelling blueprint, it is based on a CPS-deficient mutant; confirming whether the same LOS is produced in the wild-type strain will be an important future step, despite the considerable challenges posed by its dense and heterogeneous capsule repertoire. Crucially and in contrast to B. vulgatus, B. eggerthii, and B. dorei, B. thetaiotaomicron lacks galactofuranose and fucose in its LPS, and does not feature α-rhamnose as the sugar immediately following Kdo (Fig. 3), likely underscoring a distinct mechanism underlying its immunomodulatory behavior, which was associated, also in this case, with an attenuated TLR4 activation, engaging of TLR2 pathways, while selectively shaping cytokine profiles.37
Taken together, the case of Bacteroides spp. LPSs highlights that, despite architectural diversity, certain structural themes, such as lipid A hypo-acylation, reduced phosphorylation, and specific core OS signatures, recur across species and consistently align with attenuated or tolerogenic immune profiles.57 Yet, key comparative experiments are still missing, and the field lacks a unifying structure–function framework capable of predicting immunological behavior across Bacteroides spp. LPSs. This motivates extending the analysis to other Bacteroides to explore whether other LPSs from this complex and highly heterogenous genus converge on similar molecular solutions, or reveal new strategies, to shape the inflammatory landscape.
Location matters? Niche-dependent chemistry of microbiota-derived LPS
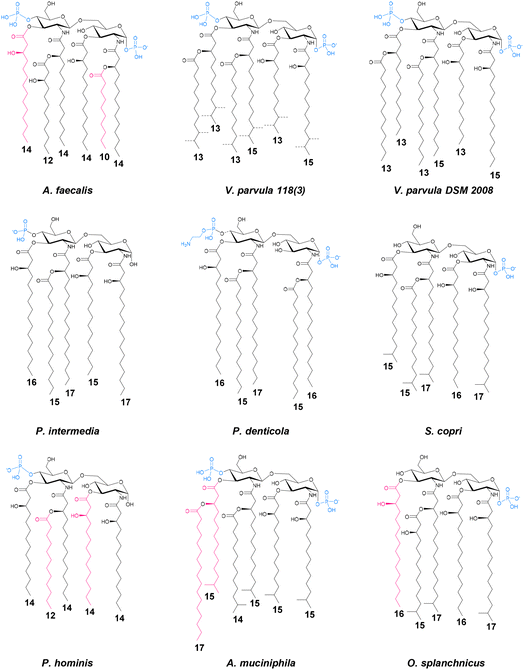
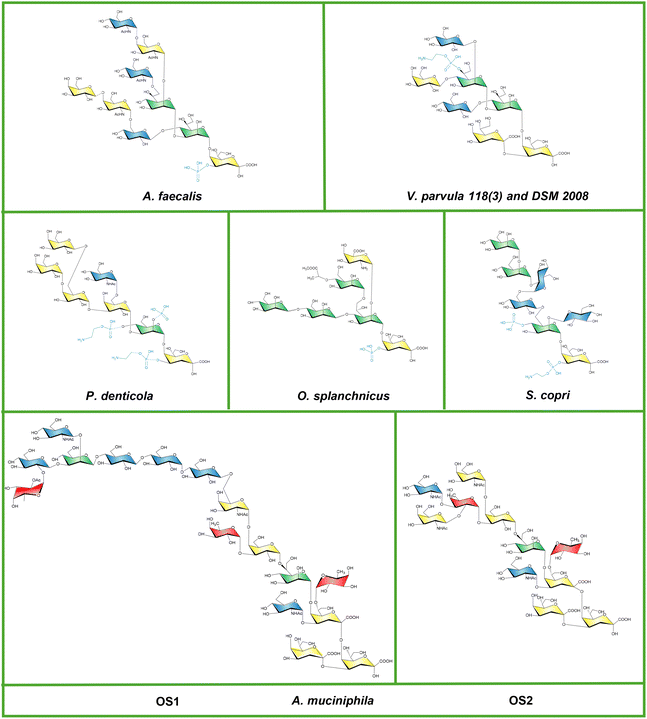
Atypical LPSs are widespread among Gram-negative members of the gut microbiota characterized thus far, reflecting the profound selective pressure exerted by the host environment, which shapes both bacterial physiology and LPS architecture. A striking example of this phenomenon is Veillonella parvula, a bacterium commonly found in the oral cavity and gastrointestinal tract. Although generally a commensal, it can become opportunistic under conditions of dysbiosis or inflammation.58–60 A recent work revealed that two strains, DSM 2008 (intestinal) and 118(3) (oral), produce chemically distinct LPS molecules, despite belonging to the same species.61 NMR and MS analyses uncovered deep structural divergence: DSM 2008 synthesizes an LPS with an unprecedented O-antigen repeating unit, whereas strain 118(3) produces a LOS characterized by striking heterogeneity in its lipid A. DSM 2008 displays a relatively homogeneous lipid A enriched in hexa-acylated, mono- and bis-phosphorylated species with linear acyl chains, features typically associated with strong TLR4 agonists (Fig. 4). In contrast, strain 118(3) exhibits extraordinary lipid A heterogeneity, ranging from tetra- to hexa-acylated forms, variable phosphorylation states, and a high abundance of branched acyl chains (Fig. 4).61 Finding two lipid A architectures so profoundly different within the same species, especially considering that lipid A is traditionally regarded as the most conserved region of an LPS molecule, is remarkable and strongly suggests a major influence of the ecological niche from which each strain was isolated (intestine vs. oral cavity). In this frame, bioinformatic and chemical biology approaches will be pivotal to understanding how such extensive structural remodeling arises in strain 118(3). As expected, this lipid A complexity has dramatic consequences for TLR4 activation: the LOS from the oral strain acts as a very weak TLR4 agonist, whereas the intestinal DSM 2008 LPS elicits a noticeably stronger response, though still weaker than E. coli LPS, in line with the atypical acyl chain lengths (i.e. C13 in place of C14 of E. coli LPS). Adding yet another layer of complexity, immunological assays revealed that despite being a potent TLR4 activator in reporter cells, the intestinal LPS induces lower TNF-α release in dendritic cells, behaving overall as a weak inflammatory molecule.61 This unexpected behavior was attributed to the presence of a highly unusual mannitol-containing tetrameric O-antigen repeating unit, found exclusively in the intestinal strain, composed of β-D-galactofuranose, β-D-galactopyranose, and α-D-glucosamine, polymerised through a phosphodiester linkage between O-6 of mannitol and O-3 of α-D-glucosamine (Fig. 5). This O-antigen is unprecedented not only for incorporating galactofuranose and mannitol phosphate, but also for its zwitterionic character, arising from the coexistence of a phosphate group and a free amino function within the repeating unit. Such zwitterionic motifs have previously been shown to modulate dendritic cell responses in other gut-derived polysaccharides, suggesting that the charge distribution of this O-antigen may directly contribute to its immunomodulatory effect and warrants further investigation. In parallel, intestinal lectins such as DC-SIGN, Langerin, and the Mannose Receptor exhibited markedly stronger binding to the intestinal LPS than to the oral LOS, supporting the notion that C-type lectins may cooperate with TLR4 to fine-tune downstream signaling and steer immune responses toward either activation or tolerance.61 In this context, future experiments using targeted antagonists or inhibitors to block C-type lectins and/or TLR4 will be essential to definitively demonstrate the regulatory contribution of the DSM 2008 O-antigen. Overall, beyond underscoring the importance of bacterial niche in shaping LPS chemistry, this study clearly highlighted that the perception of complex molecules such as V. parvula LPS relies on the activation of multiple, interdependent pathways that may converge at different nodes to shape the final immune outcome.A comparable scenario is observed for Alcaligenes faecalis, a peculiar Gram-negative commensal known to colonize Peyer's patches, a major gut-associated lymphoid tissue in the small intestine.62A. faecalis persists within these immunologically active niches, where it contributes to mucosal homeostasis by protecting the epithelial surface from pathogen invasion, regulating dendritic cell activation, and promoting IgA production through an IL-6-dependent mechanism.63 Its ability to induce IgA secretion enables A. faecalis to persist in Peyer's patches by evading excessive immune surveillance, thereby avoiding the pathological inflammation typically associated with enterobacterial LPSs while still supporting mucosal monitoring.63,64 Remarkably, its LPS was shown to enhance IgA production without eliciting a strong TLR4 response, a rare combination of features that aligns with the key hallmarks sought in next-generation vaccine adjuvants.65 Consistent with this profile, synthetic analogues of its lipid A demonstrated excellent safety and potent adjuvanticity when co-administered intranasally or subcutaneously with antigens in murine models,66 enhancing OVA-specific IgG titers and preferentially skewing the response toward Th17 rather than Th1/Th2 immunity.67 Moreover, when delivered intranasally together with pneumococcal surface protein A, a universal antigen of Streptococcus pneumoniae, the lipid A derivative significantly boosted pneumococcal surface protein A-specific nasal IgA and a protective Th17 response, conferring resistance against pneumococcal challenge.66A. faecalis produces both a LOS and an LPS whose structures are far from canonical.65 They comprise a heterogeneous mixture of tetra- to hexa-acylated lipid A species, in both mono- and bis-phosphorylated forms, and features 14:0 (3-OH) as the primary and 12:0 (3-OH) and 10:0 as secondary acyl chains (Fig. 4). Even though the hexa-acylated species contain two phosphates and share several acyl chain identities with E. coli, multiple structural modifications likely underpin their distinct immunological behavior. Among these, (i) the symmetric acyl-chain distribution (3 + 3 in place of 4 + 2 observed in E. coli lipid A), (ii) the presence of a shorter secondary chain (10:0 instead of 14:0) positioned differently relative to E. coli, (iii) the inclusion of a hydroxylated secondary acyl moiety, and (iv) the coexistence of hypo-acylated, mono-phosphorylated forms, stand out as particularly relevant (Fig. 4).65 Collectively, these features are predicted to weaken TLR4 engagement perturbing the alignment of lipid A within the TLR4/MD-2 pocket, impairing productive dimerization of the receptor complex and ultimately resulting in a poor inflammatory response. Moreover, A. faecalis LOS displays a mono-phosphorylated core OS composed of nine monosaccharide units also comprising two N-acetyl D-glucosamine, two N-acetyl D-galactosamine, one D-glucose, and one D-galactose in the outermost region (Fig. 6). This unusual composition tempting suggests that this specific glycan signature might be recognized by lectins expressed in Peyer's patches, potentially contributing to the bacterium ability to persist within this niche. Notably, the Syk-coupled C-type lectin receptor Mincle is expressed in Peyer's patches, where its stimulation by commensal-derived ligands induces IL-6 and IL-23p19 production, thereby regulating intestinal Th17 cells and IL-17-secreting Innate Lymphoid Cells.68 This cascade ultimately promotes IgA secretion and supports host-microbiota mutualism while preventing systemic inflammation. Despite some discrepancies in the literature, Mincle has been reported to recognize not only mannose and glucose residues but also galactose and N-acetyl galactosamine, both of which are present in the A. faecalis core OS (Fig. 6). This raises the intriguing possibility that direct engagement of Mincle by the A. faecalis core region contributes to the striking ability of this bacterium to colonize Peyer's patches and sustain a mutually beneficial relationship with the host.
These two examples, A. faecalis and V. parvula, illustrate a fascinating principle: anatomical niche might be not merely a background variable, but a powerful evolutionary force that sculpts LPS chemistry and, in turn, rewires innate immune recognition. Deciphering this niche–structure–function axis is essential to identify the molecular rules that distinguish “beneficial” from “harmful” LPSs, an endeavor that stands at the frontier of microbiota-driven immunomodulation.
Friends or foes?
Resolving the chemistry of microbiota-derived LPSs might be essential for explaining why some gut microbes exert beneficial effects in certain contexts yet appear detrimental in others. Few cases illustrate this ambiguity as clearly as Segatella copri (formerly Prevotella copri), a dominant member of non-Westernized gut communities whose impact on human health remains highly debated. A recent study revealed that S. copri produces a heptose-deficient LOS, in which α-mannose replaces the canonical heptose as the first sugar linked to Kdo, forming a core OS essentially composed of glucose and mannose and featuring an intriguing α-1,2-linked terminal di-mannose at the non-reducing end (Fig. 6).69 Moreover, its lipid A is extremely heterogenous, mono-phosphorylated and hypo-acylated (tetra and penta-acylated forms were identified) with minor species carrying an additional phosphate on the disaccharide backbone (Fig. 4). Even within the Prevotella/Segatella lineage, this architecture stands apart. For example, P. intermedia lipid A is mostly mono-phosphorylated and penta-acylated, with a relatively homogeneous profile,70 whereas P. denticola displays a predominantly penta-acylated lipid A decorated with PEtN and a distinct combination of C17(3-OH), C16(3-OH), and C15:0 acyl chains (Fig. 4).71 Functionally, S. copri LOS behaves as a weak TLR4 agonist, yet it can dampen E. coli-induced NF-κB activation and selectively preserve CD14+CD16+ monocytes.69 This distinctive immunological signature suggests that S. copri LOS is not perceived as a danger signal but rather acts as a finely tuned modulator of host immunity. Its unusual chemical structure may therefore not elicit a strong alarm response, even at higher doses, but instead shape an attenuated, yet still immunologically competent, innate response. In this light, pinpointing the precise epitopes responsible for this peculiar behavior becomes essential, especially given the likely contribution of a coordinated crosstalk between TLR4 and mannose-recognizing lectins such as DC-SIGN and MBL. Deciphering this phenomenon may ultimately resolve the “Segatella paradox” by revealing when S. copri acts as a protective symbiont or as a potential driver of pathology.A similar ambiguity emerges with Paenalcaligenes hominis, a bacterium increasingly detected in the gut microbiota of elderly individuals,72 and frequently linked to disorders involving the gut–brain axis.73 Lee and colleagues demonstrated that P. hominis is associated with suppression of brain-derived neurotrophic factor, mediated by the activation of the TLR4-NF-κB signaling pathway in microglia, the resident macrophages of the central nervous system.73 These findings suggest that bacterial components, particularly LPSs, can translocate from the gut to the brain, triggering neuroinflammatory responses and establishing a mechanistic link between gut dysbiosis and cognitive decline.74 How bacteria reach the brain remains an intriguing and largely unresolved question. However, it is hypothesized that outer membrane vesicles may play a key role in this interaction, as they can cross the blood–brain barrier and deliver immunologically active components like LPSs. A recent study showed that P. hominis produces an LPS whose O-antigen, built of rhamnose-glucosamine repeating units and decorated by nonstoichiometric acetylation (Fig. 5), adopts a flexible, loosely coiled conformation and forms highly hydrated aggregates, properties that likely influence receptor accessibility and immune signalling.75 Moreover, consistent with its penta-acylated lipid A, P. hominis LPS activates TLR4 only weakly, while eliciting a moderate cytokine response in dendritic cells and engaging a distinct set of innate immune lectins, including ficolins and galectin-4.75 These selective lectin-glycan interactions once again point to non-canonical recognition pathways and is particularly interesting in the case of P. hominis as it appears contradictory: while it exhibits structural features typically associated with low inflammatory potential, functional studies associate this bacterium to colitis and neuroinflammation. This data suggested that P. hominis might therefore exploit the peculiar composition of its LPS to sustain chronic low-grade inflammatory states rather than acute responses.
Among the most emblematic examples of the “beneficial LPS concept” is Akkermansia muciniphila, a mucin-degrading symbiont whose presence is consistently linked to improved metabolic health, enhanced intestinal barrier integrity, and expansion of regulatory T cells.76 Remarkably, both live and pasteurized bacterial cells exert comparable, sometimes even superior, beneficial effects in mice and humans, a property largely attributed to the heat-stable outer membrane protein Amuc_1100, known to signal through TLR2.77,78 Yet A. muciniphila, as a Gram-negative member of the Verrucomicrobia, also produces LPSs, raising the question of how molecules classically associated with inflammation can coexist within a bacterium universally regarded as beneficial and considered a next-generation probiotic. Recent structural investigations resolved this apparent contradiction by demonstrating that A. muciniphila strain MucT synthesizes two unusual LOSs, one of which bearing a strikingly large, non-repetitive core OS including a distinctive fucosylation pattern (Fig. 6, OS1 and OS2), coupled to an atypical lipid A moiety characterized by the presence of branched acyl chains and made up of a complex blend of mono- and bis-phosphorylated, tetra to hexa-acylated lipid A species.79 Strikingly, this LOS engages both TLR4 and TLR2, with the TLR2 arm providing a compensatory, anti-inflammatory counterbalance to the weak canonical TLR4 activation.79 The peculiar chemistry of its lipid A once again appears central to the LOS attenuated TLR4 agonism, while its extended carbohydrate region likely contributes to lectin-mediated modulation at the mucosal interface. A comparable paradigm emerges from recent work on Odoribacter splanchnicus, another promising next-generation probiotic. Its exceptionally short LOS, enriched in unusual sugars such as D-4-O-(1-carboxyethyl)mannose (D-Man4Lac) and carrying a Bacteroides-like lipid A, forms supramolecular aggregates encased in an extended oligosaccharide shell (Fig. 6).80 This kind of “structural masking” sterically shields lipid A from the TLR4/MD-2 complex, thereby dampening receptor activation and favoring immune tolerance, epithelial preservation, and competition against more inflammatory E. coli LPS species.
Together, these examples further underline that within the gut microbiota, the boundary between “good” and “bad” LPS is not defined by taxonomy, but by chemistry. Only by dissecting the fine structural motifs, and the signaling pathways they engage, we can begin to predict whether an LPS will trigger inflammation, modulate immune response, or even serve as a therapeutic tool. In this sense, mapping the chemical logic of microbiota-derived LPSs is not an academic exercise; it is the key, instead, to decoding microbial behavior and engineering the next generation of immunomodulators.
Conclusions
The examples discussed in this Perspective collectively challenge the traditional view of LPSs as unidimensional pro-inflammatory triggers sensed exclusively through TLR4. Instead, gut-derived LPSs emerge as chemically diverse and functionally nuanced mediators that can span a continuum from strong agonism to near-immunological silence, and even active gut mucosal tolerance. This spectrum is not random: it is encoded in precise structural features of lipid A (acylation pattern, phosphorylation, charge distribution), in the architecture of the core OS and O-antigen, and in how these motifs are presented at the supramolecular level. A recurring theme across all species characterized thus far is that small chemical changes can produce disproportionately large shifts in immune outcome. Hypo-acylation and reduced phosphorylation of lipid A, zwitterionic or charge-masked saccharide regions, rare sugars such as galactofuranose or D-Man4Lac, and atypical aggregation states all converge to weaken or rewire canonical TLR4 signaling. These LPS structures, in fact, frequently engage a broader receptor repertoire, including TLR2, C-type lectins, galectins, ficolins, and other soluble or membrane-associated pattern-recognition molecules. The resulting responses are not simply “less inflammatory”, but qualitatively distinct, often favoring IL-10 production, Treg expansion, barrier maintenance, and tissue repair, or sustaining chronic low-grade inflammation instead of acute responses.1Another key message is that context matters: anatomical niche, host age, inflammatory tone, and microbial lifestyle (symbiont vs. pathobiont vs. opportunist) all shape both LPS chemistry and its interpretation by the host. The comparison between oral and intestinal V. parvula and the Peyer's patches-adapted A. faecalis underscores that LPSs are not static molecular patterns, but an evolvable interface optimized for specific ecological and immunological landscapes. Understanding this niche–structure–function axis will be essential for predicting when a given LPS behaves as a beneficial signal, a tolerogenic buffer, or a driver of pathology. In this spirit, we hope that future work will increasingly focus on LPS heterogeneity not only across species, but within the same species isolated from different anatomical sites or even from different individuals. Such efforts may reveal personalized patterns of LPS chemistry that mirror, or even shape, individual immune tone. Ultimately, expanding LPS research into this intra-species and inter-individual dimension could lay the foundation for a new frontier in precision (glyco)immunology, where bacterial surface glycans are actionable targets for personalised interventions.
From a translational viewpoint, these insights open a double opportunity. On one hand, they highlight the potential of microbiota-derived LPSs as templates for next-generation immunomodulators, ranging from anti-inflammatory agents and adjuvants to mucosal vaccine platforms. Second, it highlights the need for diagnostic tools capable of distinguishing “good” from “bad” LPSs in complex samples, moving beyond crude LPS measurements toward molecularly resolved profiling of LPS mixtures. In this context, analytical advances are indispensable. While wet organic chemistry conjugated with NMR spectroscopy remain foundational for defining saccharide chemistry, high-resolution MS, still underused beyond lipid A, has the potential to transform LPS structural biology. Modern MS allows simultaneous interrogation of lipid and carbohydrate moieties with high accuracy, often requiring minimal material. When combined with ion mobility, MS can resolve isobaric species based on size, shape, and charge, generating drift-time maps that disentangle heterogeneous mixtures. Given the remarkable diversity of LPS structures even within single species, ion mobility-assisted MS is poised to become a central tool for deconvoluting this complexity. It also remains crucial to develop multiomic approaches capable of profiling lipid A and extended LPS structures in human-derived samples (i.e. stool, saliva, blood) and linking them to functional outputs.25 Further validation in physiologically relevant in vivo systems is required, considering host physiology, microbial–microbial interactions, and environmental cues. While germ-free and TLR-knockout mice have provided valuable insights, their limitations are well known: several LPS chemotypes that are silent or antagonistic in mice remain stimulatory in humans due to species-specific differences in MD-2.81,82
Moreover, large portions of the Gram-negative gut microbiota remain chemically unexplored. Parabacteroides species (P. distasonis, P. goldsteinii, P. johnsonii, P. merdae), reduced in obesity, metabolic syndrome, and IBD, still lack any structural definition of their LPSs.83 The same holds for other abundant Bacteroides members, despite mounting functional evidence of immunoregulatory potential. Together, these blind spots reveal a simple truth: most immunologically relevant LPS structures in the gut microbiota remain undiscovered. Mapping this hidden chemical space will be essential to resolve conflicting phenotypes, identify novel immunomodulatory archetypes, and refine the notion of what makes an LPS “beneficial” or “pathogenic”.
Looking ahead, what the field still lacks, and what now appears within reach, is a unifying structure–function framework for gut commensal LPSs: experimentally grounded rules linking discrete chemical motifs to reproducible immunological outputs across receptors, cell types, and anatomical sites. Building such a framework will require systematic comparisons across multiple gut Gram-negative species, integrating analytical chemistry, synthetic glycoengineering, biophysics, and immunology. Yet the potential payoff is enormous: (i) the ability to classify microbiota-derived LPSs as inflammatory, tolerogenic, or “tuned”; (ii) to predict their behavior in vivo; and ultimately (iii) to engineer synthetic LPS mimetics that recalibrate host immunity with unprecedented precision. In this sense, deciphering the chemical logic of gut-derived LPSs is far more than an exercise in structural curiosity. It is the necessary step to transform LPSs from just feared “endotoxins” into designable immunological tools that will help us understand, and eventually control, the thin line that separates microbial harm from microbial benefit.
Author contributions
Conceptualization, F. D. L.; writing – original draft, M. M., V. M. and F. D. L.; writing – review & editing, M. M., V. M., A. S., A. M. and F. D. L.; supervision, F. D. L.Conflicts of interest
There are no conflicts to declare.Data availability
Data for this perspective article are available from the references.Acknowledgements
This work was supported by the European Research Council (ERC) under the Horizon Europe program under grant agreement no. 101039841 (DEBUGGING LPS) to F. D. L. and under grant agreement no. 101198490 (LPS-CODE) to A. M. It was also supported by the project “CLariFY” and “GIVING” funded by the Italian Ministry of University and Research, PRIN MUR 2022 (2022SHW3KY) and PRIN MUR PNRR 2022 (P202293ZMC) to F. D. L. This work was also supported in part by the Italian Ministry of Foreign Affairs and International Cooperation (Italy Germany Science and Technology Cooperation – Call for joint research proposals for the years 2023–2025). This research was funded under the National Recovery and Resilience Plan (NRRP), Mission 4 Component 2 Investment 1.3—Call for tender no. 341 of 15 March 2022 of Italian Ministry of University and Research funded by the European Union—NextGenerationEU. Project code PE00000003, Concession Decree No. 1550 of 11 October 2022, adopted by the Italian Ministry of University and Research, CUP E63C22002030007, Project title “ON Foods—Research and Innovation Network on Food and Nutrition Sustainability, Safety and Security—Working ON Foods”.References
- F. Di Lorenzo, K. A. Duda, R. Lanzetta, A. Silipo, C. De Castro and A. Molinaro, Chem. Rev., 2022, 122, 15767–15821 Search PubMed.
- H. Zhu, C. S. Rollier and A. J. Pollard, Expert Rev. Vaccines, 2021, 20, 1515–1538 Search PubMed.
- E. Könönen, S. Asikainen and H. Jousimies-Somer, Oral Microbiol. Immunol., 1992, 7, 28–31 CrossRef.
- R. Luo, Y. Yao, Z. Chen and X. Sun, Cell Commun. Signal., 2025, 23, 142 CrossRef CAS PubMed.
- B. S. Park, D. H. Song, H. M. Kim, B. S. Choi, H. Lee and J. O. Lee, Nature, 2009, 458, 1191–1195 CrossRef CAS.
- S. De Chiara, L. De Simone Carone, R. Cirella, E. Andretta, A. Silipo, A. Molinaro, M. Mercogliano and F. Di Lorenzo, ChemMedChem, 2025, 20, e202400780 Search PubMed.
- T. Kawai and S. Akira, Trends Mol. Med., 2007, 13, 460–469 CrossRef CAS.
- E. R. Davenport, J. G. Sanders, S. J. Song, K. R. Amato, A. G. Clark and R. Knight, BMC Biol., 2017, 15, 127 Search PubMed.
- F. Di Lorenzo, C. De Castro, A. Silipo and A. Molinaro, FEMS Microbiol. Rev., 2019, 43, 257–272 Search PubMed.
- F. Di Lorenzo, C. De Castro, R. Lanzetta, M. Parrilli, A. Silipo and A. Molinaro, in Carbohydrates in Drug Design and Discovery, ed. J. Jimenez-Barbero, F. J. Canada and S. Martin-Santamaria, RSC, 2015, vol. 3, pp. 38–63 Search PubMed.
- P. Kumar, E. A. Schroder, M. V. S. Rajaram, E. N. Harris and L. P. Ganesan, Cells, 2024, 13, 1590 CrossRef CAS.
- J. K. Kim, H. A. Jang, M. S. Kim, J. H. Cho, J. Lee, F. Di Lorenzo, L. Sturiale, A. Silipo, A. molinaro and B. L. Lee, J. Biol. Chem., 2017, 292, 19226–19237 Search PubMed.
- F. Di Lorenzo, A. Silipo, R. Lanzetta, M. Parrilli and A. Molinaro, in Carbohydrate Chemistry: State of the Art and Challenges for Drug Development, ed. Field, R. J. and Hill, H. A. O., World Scientific Publishing, 2015, vol. 3, pp. 57–89 Search PubMed.
- A. De Nicola, C. Montis, G. Donati, A. Molinaro, A. Silipo, A. Balestri, D. Berti, F. Di Lorenzo, Y.-L. Zhu and G. Milano, Nanoscale, 2023, 15, 8988–8995 Search PubMed.
- F. Di Lorenzo, J. Billod, S. Martìn-Santamarìa, A. Silipo and A. Molinaro, Eur. J. Org Chem., 2017, 28, 4055–4073 Search PubMed.
- A. Molinaro, O. Holst, F. Di Lorenzo, M. Callaghan, A. Nurisso, G. D'Errico, A. Zamyatina, F. Peri, R. Berisio, R. Jerala, J. Jimènez-Barbero, A. Silipo and S. Martìn-Santamarìa, Chem.–Eur. J., 2015, 21, 500–519 CrossRef CAS PubMed.
- J. R. Brestoff and D. Artis, Nat. Immunol., 2013, 14, 676–684 CrossRef CAS.
- M. Shabani, A. Ghoshehy, A. M. Mottaghi, Z. Chegini, A. Kerami, A. Shariati and M. Taati Moghadam, Front. Cell. Infect. Microbiol., 2025, 15, 1516010 CrossRef.
- A. B. Shreiner, J. Y. Kao and V. B. Young, Curr. Opin. Gastroenterol., 2015, 31, 69–75 CrossRef CAS.
- K. Hou, Z. Wu, X. Chen, J. Wang, D. Zang, C. Xiao, D. Zhu, J. B. Koya, L. Wei, J. Li and Z. Chen, Signal Transduct. Targeted Ther., 2022, 7, 135 Search PubMed.
- S. S. Pillai, C. A. Gagnon, C. Foster and A. P. Ashraf, J. Clin. Endocrinol. Metab., 2024, 109, 2709–2719 CrossRef PubMed.
- A. Vargas, B. L. Robinson, K. Houston, A. R. Vilela Sangay, M. Saadeh, S. D'Souza and D. A. Johnson, Explor. Med., 2025, 6, 1001275 Search PubMed.
- F. Sadeghpour Heravi, Curr. Clin. Microbiol. Rep., 2024, 11, 177–191 CrossRef.
- A. Sai, G. B. Shetty, P. Shetty and H. L. Nanjeshgowda, Influence of gut microbiota on autoimmunity: A narrative review, Brain Behav. Immun. Integr., 2024, 5, 100046 Search PubMed.
- F. Di Lorenzo, L. Paparo, L. Pisapia, F. Oglio, M. D. Pither, R. Cirella, R. Nocerino, L. Carucci, A. Silipo, F. De Filippis, D. Ercolini, A. Molinaro and R. Berni Canani, Front. Mol. Biosci., 2023, 10, 1266293 Search PubMed.
- Z. Nádházi, A. Takàts, K. Offenmüller and L. Bertòk, Acta Microbiol. Immunol. Hung., 2002, 49, 151–157 Search PubMed.
- E. d'Hennezel, S. Abubucker, L. O. Murphy and T. W. Cullen, mSystems, 2017, 2, e00046–17 Search PubMed.
- K. Nishiyama, T. Yokoi, M. Sugiyama, R. Osawa, T. Mukai and N. Okada, Front. Microbiol., 2021, 12, 754819 CrossRef CAS.
- J. Tan, C. McKenzie, M. Potamitis, A. N. Thorburn, C. R. Mackay and L. Macia, Adv. Immunol., 2014, 121, 91–119 CAS.
- J. H. Shin, G. Tillotson, T. N. Mackensie, C. A. Warren, H. M. Wexler and E. J. C. Goldstein, Anaerobe, 2024, 85, 102819 CrossRef CAS.
- C. K. Fisher and P. Mehta, PLoS One, 2014, 9, e102451 Search PubMed.
- H. M. Wexler, Clin. Microbiol. Rev., 2007, 20, 593–621 CrossRef CAS.
- F. Di Lorenzo, M. D. Pither, M. Martufi, I. Scarinci, J. Guzmán-Caldentey, E. Łakomiec, W. Jachymek, S. C. M. Bruijns, S. M. Santamaria, J. Frick, Y. van Kooyk, F. Chiodo, A. Silipo, M. L. Bernardini and A. Molinaro, ACS Cent. Sci., 2020, 6, 1602–1616 Search PubMed.
- M. Hashimoto, F. Kirikae, T. Dohi, S. Adachi, S. Kusumoto, Y. Suda, T. Fujita, H. Naoki and T. Kirikae, Eur. J. Biochem., 2002, 269, 3715–3721 CrossRef PubMed.
- E. T. Rietschel, T. Kirikae, F. U. Schade, U. Mamat, G. Schmidt, H. Loppnow, A. J. Ulmer, U. Zähringer, U. Seydel, F. Di Padova, M. Schreier and H. Brade, FASEB J., 1994, 8, 217–225 CrossRef CAS.
- A. Steimle, L. Michaelis, F. Di Lorenzo, T. Kliem, T. Münzner, J. K. Maerz, A. Schäfer, A. Lange, R. Parusel, K. Gronbach, K. Fuchs, A. Silipo, H. H. Öz, B. J. Pichler, I. B. Autenrieth, A. Molinaro and J. Frick, Mol. Ther., 2019, 27, 1974–1991 CrossRef CAS PubMed.
- M. D. Pither, A. Illiano, C. Pagliuca, A. Jacobson, G. Mantova, A. Stornaiuolo, R. Colicchio, M. Vitiello, G. Pinto, A. Silipo, M. A. Fischbach, P. Salvatore, A. Amoresano, A. Molinaro and F. Di Lorenzo, Carbohydr. Polym., 2022, 297, 120040 CrossRef CAS PubMed.
- M. Tiemblo Martín, M. Coccimiglio, E. Andretta, L. De Simone Carone, A. Bell, T. Gerpe-Amor, C. Di Carluccio, A. Molinaro, Y. Van Kooyk, N. Juge, F. Chiodo, F. Di Lorenzo and A. Silipo, Carbohydr. Polym., 2025, 348, 122833 CrossRef.
- Z. Lin, H. Tang, L. Che, M. Chen, Z. Ye, X. Yang, L. Sun, K. H. Mayo, G. Tai, L. Cui and Y. Zhou, Carbohydr. Polym., 2026, 373, 124566 CrossRef CAS.
- G. Lugo-Villarino, A. Troegeler, L. Balboa, C. Lastrucci, C. Duval, I. Mercier, A. Bènard, F. Capilla, T. Al Saati, R. Poincloux, I. Kondova, F. A. W. Verreck, C. Cougoule, I. Maridonneau-Parini, M. del Carmen Sasiain and O. Neyrolles, Front. Immunol., 2018, 9, 1123 CrossRef.
- H. Baek, S.-W. Yang, S. Kim, Y. Lee, H. Park, M. Park, B.-J. Jeon, H. Park, H.-S. Hwang, J.-Y. Kim, J.-H. Kim and Y.-S. Kang, Int. J. Mol. Sci., 2025, 26, 2329 CrossRef CAS PubMed.
- S. I. Gringhuis, J. den Dunnen, M. Litjens, M. van der Vlist and T. B. H. Geijtenbeek, Nat. Immunol., 2009, 10, 1081–1088 CrossRef CAS PubMed.
- T. B. H. Geijtenbeek, S. J. van Vliet, E. A. Koppel, M. Sanchez-Hernandez, C. M. J. E. Vandenbroucke-Grauls, B. Appelmelk and Y. van Kooyk, J. Exp. Med., 2003, 197, 7–17 CrossRef CAS.
- F. Nieto-Fabregat, Q. Zhu, C. Vivès, Y. Zhang, A. Marseglia, F. Chiodo, M. Thépaut, D. Rai, S. S. Kulkarni, F. Di Lorenzo, A. Molinaro, R. Marchetti, F. Fieschi, G. Xiao, B. Yu and A. Silipo, JACS Au, 2024, 4, 697–712 CrossRef CAS PubMed.
- D. Rai, F. Nieto-Fabregat, R. Dikshit, M. Thépaut, F. Fieschi, A. Silipo and S. S. Kulkarni, ACS Omega, 2025, 10, 51985–52000 CrossRef.
- S. Francisco, J.-M. Billod, J. Merino, C. Punzòn, A. Gallego, A. Arranz, S. Martin-Santamaria and M. Fresno, Front. Immunol., 2022, 12, 748303 CrossRef.
- S. K. Mazmanian, C. H. Liu, A. O. Tzianabos and D. L. Kasper, Cell, 2005, 122, 107–118 CrossRef CAS.
- C. L. Sears, A. L. Geis and F. Housseau, J. Clin. Invest., 2014, 124, 4166–4172 CrossRef CAS.
- E. B. Troy and D. L. Kasper, Front. Biosci., 2010, 15, 25–34 CrossRef CAS PubMed.
- T. Yang, J. Daugherti and D. L. Kasper, bioRxiv, 2024, preprint, DOI:10.1101/2024.04.13.589361.
- K. Kitamura, M. Sasaki, M. Matsumoto, H. Shironoya and K. Iida, Immunol. Lett., 2021, 233, 48–56 CrossRef CAS.
- A. K. Sulit, M. Daigneault, E. Allen-Vercoe, O. K. Silander, B. Hock, J. McKenzie, J. Person, F. A. Frizelle, S. Schmeier and R. Purcell, npj Biofilms Microbiomes, 2023, 9, 59 CrossRef CAS.
- Q. He, M. Niu, J. Bi, N. Du, S. Liu, K. Yang, H. Li, J. Yao, Y. Du and Y. Duan, Sci. Rep., 2023, 13, 15842 CrossRef CAS PubMed.
- S. Hsieh, N. T. Porter, D. L. Donermeyer, S. Horvath, G. Strout, B. T. Saunders, N. Zhang, B. Zinselmeyer, E. C. Martens, T. S. Stappenbeck and P. M. Allen, J. Immunol., 2020, 204, 1035–1046 CrossRef CAS.
- A. N. Jacobson, B. P. Choudhury and M. A. Fischbach, mBio, 2018, 9, e02289–17 CrossRef CAS.
- M. D. Pither, A. Silipo, F. Di Lorenzo and A. Molinaro, in Comprehensive Glycoscience, ed. N. Taniguchi and J. Barchi Jr, Elsevier, 2nd edn, 2021, vol. 5, pp. 233–249 Search PubMed.
- F. Di Lorenzo, Ł. Kubik, A. Oblak, N. I. Lorè, C. Cigana, R. Lanzetta, M. Parrilli, M. A. Hamad, A. De Soyza, A. Silipo, R. Jerala, A. Bragonzi, M. A. Valvano, S. Martìn-Santamaria and A. Molinaro, J. Biol. Chem., 2015, 290, 21305–21319 CrossRef CAS.
- P. Zhou, D. Manoil, G. N. Belibasakis and G. A. Kotsakis, Front. Oral Health, 2021, 2, 774115 CrossRef.
- D. I. Poppleton, M. Duchateau, V. Hourdel, M. Matondo, J. Flechsler, A. Klingl, C. Beloin and S. Gribaldo, Front. Microbiol., 2017, 8, 1215 CrossRef.
- C. V. Hughes, P. E. Kolenbrander, R. N. Andersen and L. V. Moore, Appl. Environ. Microbiol., 1988, 54, 1957–1963 CrossRef CAS PubMed.
- M. D. Pither, E. Andretta, G. Rocca, F. Balzarini, A. Matamoros-Recio, R. Colicchio, P. Salvatore, Y. Van Kooyk, A. Silipo, F. Granucci, S. Martìn-Santamaria, F. Chiodo, A. Molinaro and F. Di Lorenzo, Angew. Chem., Int. Ed., 2024, 63, e202401541 CrossRef CAS.
- T. Obata, Y. Goto, J. Kunisawa, S. Sato, M. Sakamoto, H. Setoyama, T. Matsuki, K. Nonaka, N. Shibata, M. Gohda, Y. Kagiyama, T. Nochi, Y. Yuki, Y. Fukuyama, A. Mukai, S. Shinzaki, K. Fujihashi, C. Sasakawa, H. Iijima, M. Goto, Y. Umesaki, Y. Benno and H. Kiyono, Proc. Natl. Acad. Sci. U.S.A., 2010, 107, 7419–7424 CrossRef CAS PubMed.
- N. Shibata, J. Kunisawa, K. Hosomi, Y. Fujimoto, K. Mizote, N. Kitayama, A. Shimoyama, H. Mimuro, S. Sato, N. Kishishita, K. J. Ishii, K. Fukase and H. Kiyono, Mucosal Immunol., 2018, 11, 693–702 CrossRef CAS.
- L. Ding, X. Chen, H. Cheng, T. Zhang and Z. Li, Front. Chem., 2022, 10, 974854 CrossRef CAS.
- A. Shimoyama, F. Di Lorenzo, H. Yamaura, K. Mizote, A. Palmigiano, M. D. Pither, I. Speciale, T. Uto, S. Masui, L. Sturiale, D. Garozzo, K. Hosomi, N. Shibata, Z. Kabayama, Y. Fujimoto, A. Silipo, J. Kunisawa, H. Kiyono, A. Molinaro and K. Fukase, Angew. Chem., Int. Ed., 2021, 60, 10023–10031 CrossRef CAS.
- K. Yoshii, K. Hosomi, A. Shimoyama, Y. Wang, H. Yamaura, T. Nagatake, H. Suzuki, H. Lan, H. Kiyono, K. Fukase and J. Kunisawa, Microorganisms, 2020, 8, 1102 CrossRef CAS.
- Y. Wang, K. Hosomi, A. Shimoyama, K. Yoshii, H. Yamaura, T. Nagatake, T. Nishino, H. Kiyono, K. Fukase and J. Kunisawa, Vaccines, 2020, 8, 395 CrossRef CAS.
- M. Martínez-López, S. Iborra, R. Conde-Garrosa, A. Mastrangelo, C. Danne, E. R. Mann, D. M. Reid, V. Gaboriau-Routhiau, M. Chaparro, M. P. Lorenzo, L. Minnerup, P. Saz-Leal, E. Slack, B. Kemp, J. P. Gisbert, A. Dzionek, M. J. Robinson, F. J. Rupérez, N. Cerf-Bensussan, G. D. Brown, D. Bernardo, S. LeibundGut-Landmann and D. Sancho, Immunity, 2019, 50, 446–461 CrossRef.
- L. De Simone Carone, G. Barra, R. Cirella, M. Ziaco, M. Mercogliano, F. Olmeo, E. Andretta, V. Mazziotti, C. Fusco, G. D'Ippolito, K. A. Duda, F. M. Farquharson, P. Louis, A. Fontana, A. Silipo, A. Molinaro and F. Chiodo, Angew. Chem., Int. Ed., 2025, e202512947 CAS.
- M. Hashimoto, Y. Asai, R. Tamai, T. Jinno, K. Umatani and T. Ogawa, FEBS Lett., 2003, 543, 98–102 CrossRef CAS.
- F. Di Lorenzo, A. Silipo, T. Matier, A. Hanuszkiewicz, J. S. Elborn, R. Lanzetta, L. Sturiale, A. Scamporrino, D. Garozzo, M. A. Valvano, M. M. Tunney and A. Molinaro, Eur. J. Org Chem., 2016, 2016, 1732–1738 CrossRef CAS.
- S.-H. Park, K.-A. Kim, Y.-T. Ahn, J.-J. Jeong, C.-S. Huh and D.-H. Kim, BMC Microbiol., 2015, 15, 49 CrossRef PubMed.
- K.-E. Lee, J.-K. Kim, S.-K. Han, D. Y. Lee, H.-J. Lee, S.-V. Yim and D.-H. Kim, Microbiome, 2020, 8, 107 CrossRef CAS PubMed.
- H. J. Kim, H. Kim, J.-H. Lee and C. Hwangbo, Immun. Ageing, 2023, 20, 67 CrossRef CAS.
- F. Nieto-Fabregat, M. Mercogliano, A. Cangiano, G. Vitiello, E. Andretta, L. A. Clifton, A. Vanacore, L. Buono, M. Asunción Campanero-Rhodes, D. Solís, C. Di Carluccio, G. Pecoraro, A. Molinaro, G. Smaldone, J.-K. Kim, D.-H. Kim, L. Paduano, F. Di Lorenzo and A. Silipo, JACS Au, 2025, 5, 3311–3327 CrossRef PubMed.
- F. Gao, C. Cheng, R. Li, Z. Chen, K. Tang and G. Du, Front. Med., 2025, 12, 1484656 CrossRef.
- J. Wang, W. Xu, R. Wang, R. Cheng, Z. Tang and M. Zhang, Food Funct., 2021, 12, 3597–3610 RSC.
- P. Garcia-Vello, H. L. P. Tytgat, J. Gray, J. Elzinga, F. Di Lorenzo, J. Biboy, D. Vollmer, C. De Castro, W. Vollmer, W. M. de Vos and A. Molinaro, Glycobiology, 2022, 32, 712–719 CrossRef CAS.
- P. Garcia-Vello, H. L. P. Tytgat, J. Elzinga, M. Van Hul, H. Plovier, M. Tiemblo-Martin, P. D. Cani, S. Nicolardi, M. Fragai, C. De Castro, F. Di Lorenzo, A. Silipo, A. Molinaro and W. M. de Vos, Nat. Commun., 2024, 15, 8411 CrossRef.
- M. Tiemblo Martín, M. Mercogliano, K. Hiippala, L. De Simone Carone, M. A. Campanero-Rhodes, M. Masiello, A. Cangiano, A. Molinaro, L. Paduano, D. Solís, R. Satokari, F. Di Lorenzo and A. Silipo, Chem. Sci., 2025, 17(4), 2281–2294 RSC.
- A. M. Hajjar, R. K. Ernst, E. S. Fortuno, A. S. Brasfield, C. S. Yam, L. A. Newlon, T. R. Kollmann, S. I. Miller and C. B. Wilson, PLoS Pathog., 2002, 8, e1002963 CrossRef.
- J. Vašl, A. Oblak, T. T. Peternelj, J. Klett, S. Martìn-Santamaria, T. L. Gioannini, J. P. Weiss and R. Jerala, J. Immunol., 2016, 196, 2309–2318 CrossRef.
- Y. Cui, L. Zhang, X. Wang, Y. Yi, Y. Shan, B. Liu, Y. Zhou and X. Lü, FEMS Microbiol. Lett., 2022, 369, fnac072 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |