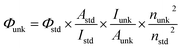Open Access Article
Open Access ArticleMolecular jackhammers induce intracellular calcium release and skeletal muscle contraction by vibronic-driven action
Yuchen
Rui†
a,
Bowen
Li†
*a,
Vardan
Vardanyan
a,
Soonyoung
Kim
d,
Dallin
Arnold
 a,
Jacob L.
Beckham
a,
Ciceron
Ayala-Orozco
a,
Ana L.
Santos
a,
Gautam
Chaudhry
a,
Jacob L.
Beckham
a,
Ciceron
Ayala-Orozco
a,
Ana L.
Santos
a,
Gautam
Chaudhry
 a,
Lixin
Zhou
a,
Shichen
Xu
a,
Lixin
Zhou
a,
Shichen
Xu
 a,
Tengda
Si
a,
Zicheng
Wang
a,
Angel A.
Martí
a,
Tengda
Si
a,
Zicheng
Wang
a,
Angel A.
Martí
 abc,
Anatoly
Kolomeisky
abc,
Anatoly
Kolomeisky
 *a,
Jacob T.
Robinson
*bd and
James M.
Tour
*a,
Jacob T.
Robinson
*bd and
James M.
Tour
 *acef
*acef
aDepartment of Chemistry, Rice University, Houston, Texas 77005, USA. E-mail: libowen924@gmail.com; tolya@rice.edu; tour@rice.edu
bDepartment of Bioengineering, Rice University, Houston, Texas 77005, USA. E-mail: jtrobinson@rice.edu
cDepartment of Materials Science and Nanoengineering, Rice University, Houston, Texas 77005, USA
dDepartment of Electrical Engineering, Rice University, Houston, Texas 77005, USA
eSmalley-Curl Institute, Rice University, Houston, Texas 77005, USA
fNanoCarbon Center and the Rice Advanced Materials Institute, Rice University, Houston, Texas 77005, USA
First published on 13th April 2026
Abstract
Intracellular calcium release (ICR) is of fundamental importance for numerous physiological and pathological processes, from cell differentiation to neurotransmitter release and muscle contraction. The ability to precisely control ICR is crucial for understanding cellular signaling mechanisms and developing therapeutic interventions for calcium-related disorders. Light-induced ICR offers numerous advantages over conventional approaches, including superior specificity, precise spatiotemporal control, excellent reproducibility, and minimal invasiveness. Here, we demonstrate that cyanine dyes function as molecular jackhammers (MJHs) to induce ICR in human embryonic kidney (HEK) 293 and immortalized mouse myoblast C2C12 cells through vibronic-driven action (VDA) following red light (640 nm) activation. VDA generates longitudinally and axially coordinated whole-molecule vibrations, enabling mechanical interactions with cells. Structure–activity relationship studies of twenty synthesized MJHs demonstrated that sulfonate-containing derivatives most effectively triggered intracellular calcium release while exhibiting minimal cytotoxicity. Mechanistic studies showed that MJH-induced calcium release operates through the inositol-triphosphate (IP3) pathway rather than reactive oxygen species generation. Finally, we demonstrated light-activated MJH-induced calcium waves in vivo using transgenic Hydra vulgaris expressing a fluorescent calcium indicator. This work establishes MJHs as molecular-scale devices for remote control of cellular signaling, expanding the utility of cyanine dyes for modulating physiological processes with potential therapeutic applications.
Introduction
Intracellular calcium release (ICR) is the process by which calcium ions stored in cellular organelles, such as the endoplasmic reticulum (ER) and sarcoplasmic reticulum (SR), are released into the cytoplasm.1 ICR plays a fundamental role in regulating various physiological and pathological processes,2,3 including signal transduction,4 gene expression,5,6 cell proliferation,7 neurotransmitter release,8,9 and muscle contraction.10,11 Understanding ICR mechanisms is crucial for elucidating cell signaling pathways and disease pathogenesis, with implications for drug development and clinical diagnostics.Multiple channels, pumps, and signaling pathways regulate ICR. Traditional methods for inducing ICR include: (1) chemical agonists such as ionomycin12 and thapsigargin,13,14 or hormones such as parathyroid15 and acetylcholine;16 (2) electrical stimulation via membrane depolarization, which opens voltage-gated calcium channels in specific tissues to increase intracellular calcium levels;17–19 and (3) mechanical stimuli such as cell compression and stretching.20,21 However, these approaches have limitations, including potential cytotoxicity, poor specificity, limited temporal and spatial resolution, and low reproducibility. In contrast, light-activated molecular strategies offer several advantages: they are minimally invasive, highly specific and reproducible, precisely controllable, and fully reversible, enabling dynamic regulation of calcium release.22–26
Our laboratory previously developed the biological applications of Feringa-type molecular motors (MMs), overcrowded alkenes composed of a rotor and stator that perform unidirectional mechanical work upon activation with 400 nm light.27–30 These motors can stimulate intercellular calcium waves31 through the inositol-triphosphate (IP3) signaling pathway.32–34 However, their requirement for 400 nm light activation limits broader applications due to potential tissue damage and poor penetration depth.35
Cyanine dyes are fluorescent compounds characterized by two nitrogen-containing heterocycles connected by a conjugated odd-numbered carbon chain, with one heterocycle carrying a positive charge.36,37 These dyes exhibit high molar extinction coefficients, narrow absorption and emission bands, tuneable fluorescence, and excellent biocompatibility.36–38 We recently demonstrated that cyanine dyes activated by low-intensity near-infrared (NIR) light (730 nm, 80 mW cm−2) for 10 min can function as molecular jackhammers (MJHs) to eradicate cancer cells through vibronic-driven action (VDA).39–42 VDA emerges from the coupling between the electronic and vibrational modes of a molecule, creating a molecular plasmon that induces concerted whole-molecule vibrations on picosecond timescales when activated by light. These vibrations cause the entire molecule to stretch both longitudinally and axially, generating mechanical forces capable of disrupting cell membranes.39,42 In giant unilamellar vesicles (GUVs), light activation of MJHs has been shown to induce vesicle shrinkage and collapse, which provides direct evidence of mechanically driven effects on lipid membranes. This mechanically driven process represents a new therapeutic approach distinct from traditional photodynamic and photothermal therapies, as evidenced by the fact that it is neither inhibited by reactive oxygen species (ROS) scavengers nor accompanied by media temperature increases.39,41,42
Different from inducing cancer cell death, here we demonstrate that MJHs activated by red light (640 nm) for only 250 ms effectively stimulate ICR via VDA by mechanically activating the IP3 pathway. We show the efficacy of MJH-induced ICR both in vitro using HEK293 cells,43 C2C12 cells44 and in vivo using Hydra vulgaris.45,46 Unlike our previous work on cancer cell eradication, brief MJH activation efficiently induces ICR while maintaining cell viability. The use of red light provides reduced autofluorescence, enhanced tissue penetration, and minimal tissue damage.35 During the past decades, cyanine dyes have been found to have numerous applications in biomedical imaging, drug delivery and in vivo sensing.47–49 However, this study introduces their capability to control intracellular calcium release through mechanical action, expanding its application as precise molecular tools for manipulating cellular processes in living systems.
Results and discussion
Cyanine 5.5 is a versatile far-red fluorescent dye comprising two benzoindole chromophores connected by five-methine units.39,42 It exhibits both longitudinal molecular plasmons (LMP) and transversal molecular plasmons (TMP). Time-dependent density functional theory (TD-DFT) calculations confirmed the presence of a vibronic mode (coupled vibrational oscillations and electronic states) at 640 nm,39,42 as shown by the induced charge density of the molecular plasmon in Fig. 1a.50–52 | ||
| Fig. 1 Design, synthesis, and cellular activity of molecular jackhammers. (a) Chemical structure of Cy5.5 showing longitudinal molecular plasmon (LMP, red horizontal arrow) and transversal molecular plasmon (TMP, blue vertical arrow), with calculated induced charge density from time-dependent density functional theory (TD-DFT).42 (b) General synthetic pathway to molecule 3. Reaction conditions (a) (1) Ac2O, reflux; (2) NaOAc, Ac2O, 80 °C; reaction conditions (b) NaOAc, Ac2O, 80 °C. (c) Representative molecular structures 3r, 3m, 3s and 3p. (d) UV-vis absorption spectra of 3r, 3m, and 3s, highlighting the vibronic mode at the shoulder position (640 nm). (e) Confocal microscopy images showing rapid intracellular calcium release in HEK293 cells treated with molecular jackhammer 3p (2 µM) and calcium indicator Fluo-4 (2 µM) after 640 nm laser stimulation. Red arrows indicate cells before and after stimulation. Scale bar = 50 µm (applies to all images). (f) Normalized Fluo-4 fluorescence intensity traces HEK293 cells treated with each MJHs. The solid line represents the average responses of n = 6 independent cells and the shaded region represents the standard error of the mean. 3r, 3m, 3s and 3p were treated to cells at 2 µM. All the stimuli were delivered with a 640 nm laser of 250 ms with a power of 0.73 mW to a circular area of diameter 5 µm. For all the plots, the cyan line indicates the time of stimulus presentation. | ||
The synthetic approach for both symmetrical and non-symmetrical MJHs is shown in Fig. 1b. Symmetrical MJHs 3 were synthesized from heterocyclic salts 1 and commercially available malondianil or glutacondianil 2 in the presence of NaOAc and Ac2O, affording good yields. Furthermore, non-symmetrical MJHs 3 required a two-step process: first, heterocyclic salts 1 and 2 were reacted in Ac2O to form acetylated precursors, that were then condensed with equimolar amounts of differently substituted heterocyclic salts 1′, providing satisfactory yields (Schemes S1 and S2).41 The photophysical properties including absorption and emission spectra of all synthesized MJHs were determined (Table S1 and Fig. S1–S21).
To investigate structure–activity relationships and the importance of VDA, we selected four representative MJHs (Fig. 1c). The non-symmetrical Cy5.5-type molecule 3m incorporates one heterocyclic moiety with a methyl group and another with a six-carbon alkyl chain dimethylamino group. The Cy5-type molecule 3r shares identical functional groups with 3m but lacks one phenyl ring on each heterocyclic moiety. The Cy7-type molecule 3s features two indole-based nitrogen atoms linked by seven methyl units. Molecule 3p differs from 3m by incorporating a three-carbon alkyl chain sulfonate group on one heterocyclic moiety. As shown in Fig. 1d, all molecules display a sharp, intense absorption band with a high-energy shoulder. At 640 nm, Cy5 (3r) shows maximum absorption at its main peak, while for Cy5.5 (3m) this wavelength matches its vibronic shoulder. In contrast, Cy7 (3s) shows minimal absorption at 640 nm as this wavelength is far from both its main peak and vibronic shoulder.
To investigate ICR responses to MJH stimulation, we incubated HEK293 cells with MJHs (2 µM) and Fluo-4 (2 µM), which fluoresces green upon calcium binding (Fig. S22 and S23) for 45 min. Fig. 1e and SI Video 1 shows calcium response localization in cells treated with the Cy5.5-type compound 3p that exhibits the strongest calcium release efficacy. Both MJH and Fluo-4 penetrated the cell membrane during incubation. Stimulation of a single 3p-treated cell with a 640 nm laser at a power of 0.73 mW, scanning a 5 µm diameter circular area for 250 ms resulted in a significant increase in Fluo-4 fluorescence intensity in the targeted cell. The observed calcium release was laser-power dependent (Fig. S24), repeatable (Fig. S25), and did not cause cell death (Fig. S26 and S27). The time of 250 ms was chosen as the shortest effective irradiation duration that reliably induces robust calcium release without additional benefit at longer exposure times (Fig. S28).
Calcium responses in cells treated with 2 µM of each representative MJH and irradiated with 640 nm light revealed distinct trends (Fig. 1f). In all cases, cellular calcium intensity peaked approximately 20 s post-stimulation before gradually declining. Despite 640 nm corresponding to its absorption maximum, Cy5-type 3r induced only weak calcium release. Cy7-type 3s showed modestly higher amplitude than 3r, while Cy5.5-type 3m demonstrated enhanced amplitude, underscoring the importance of VDA for the observed calcium responses. The sulfonate-containing molecule 3p exhibited the highest amplitude in Fluo-4 fluorescence, revealing the importance of different ionic substituents to MJH-induced calcium release. Activation of 3p at both 640 and 680 nm supports that the shoulder peak excitation stimulates greater calcium release than the main absorption peak, further suggesting VDA as the primary mode of response (Fig. S29). Moreover, we employed a control molecule 4 which lacks a positive charge in the conjugated structure and consequently does not support vibronic coupling.53 Under activation of light with wavelengths from 400 to 640 nm, molecule 4 did not induce any detectable calcium release (Fig. S30). Control experiments using only DMSO solvent showed no detectable calcium responses under identical conditions (Fig. 1f).
To thoroughly investigate the relationship between their molecular structure and calcium release activity, we synthesized 20 symmetrical and non-symmetrical MJHs with diverse functional groups, including benzyl (3b), morpholino (3e), ester (3f), amide (3g), and carboxylic acid (3n) (Scheme S2).41 The MJH structures and their normalized calcium release intensities are shown in Fig. 2a and b, respectively. Non-symmetrical Cy5.5 MJHs with sulfonate groups (3p, 3q) demonstrated the highest calcium release activity. MJHs with dimethylamine groups (3c, 3d, 3l, 3m) showed intermediate activity, performing better than those with ester (3f), carboxylic acid (3n), or other groups (3b, 3g, 3o, 3i). As expected, Cy5 and Cy7 derivatives (3j, 3k, 3s) showed relatively poor performance compared to Cy5.5 derivatives with the same groups, highlighting the importance of wavelength-specific activation of the molecular vibronic shoulder and, therefore, confirming the significance of VDA for the observed calcium responses.
To gain deeper insight into these structure–activity relationships, we calculated the experimental plasmonicity index (EPI) for selected MJHs using our previously established methodology.53 The EPI was evaluated by monitoring solvent-dependent variations in the absorption maxima of MJHs across environments with different dielectric properties. The magnitude of the solvatochromic response serves as an indicator of collective charge oscillation within the molecular framework. Increased dielectric sensitivity is associated with stronger plasmonic character, allowing distinction between delocalized plasmon-like excitations and localized electronic transitions. This method provides a practical experimental descriptor of molecular plasmonicity. EPI values for representative MJHs were measured (Fig. S31 and S32), and 3p exhibits the highest EPI among the tested compounds. We also noticed that in certain cases, the extent of calcium release induced by MJHs does not directly correlate with the measured EPI. For instance, although 3n exhibits high EPI among the molecules, it induces only a limited calcium release. These results suggest that, although EPI captures the molecular intrinsic properties, extrinsic factors such as membrane insertion and functional group-dependent interactions with specific cellular components also play critical roles in modulating intracellular calcium release in cellular contexts. Recognizing the limitations of EPI measurements, we anticipate that more rigorous physicochemical characterization, such as ultrafast spectroscopic approaches in future studies, will refine the mechanism.
While the relative effectiveness of symmetrical versus non-symmetrical MJHs remains unclear for most functional groups, a notable observation emerged with compound 3h. This symmetrical MJH, containing the same functional group as 3p, showed remarkably poor activity, lower than Cy5 and Cy7 derivatives, likely due to its two negatively charged sulfonate groups impairing cell membrane binding (Fig. S33). Importantly, we demonstrated that MJH 3p-induced calcium release could be achieved not only in HEK293 cells but also in skeletal muscle C2C12 cells (Fig. 2c and S34),44 suggesting its broader potential applications across diverse cell types with distinct basal calcium levels. Also, the calcium release in C2C12 cells is repeatable (Fig. S35).
Beyond their ability to elicit calcium release, we also evaluated cytotoxicity as a critical factor for molecule optimization. We selected ten representative MJHs to investigate their impact on cell viability using the CCK-8 assay54 in HEK293 cells. As shown in Fig. 2d and e, most MJHs exhibited high toxicity at 20 µM. However, sulfonate-containing compounds 3p and 3q displayed lower toxicity even at higher concentrations. IC50 values were determined for each MJH (Fig. 2f). The methyl-substituted MJH 3a showed the highest toxicity. Amine-containing compounds 3l and 3t showed decreased yet significant toxicity. Compound 3n with carboxylic acid showed improved but still suboptimal toxicity profiles. Notably, sulfonate-containing 3p and 3q not only exhibited the highest calcium release activity but also the lowest toxicity among all MJHs tested. For instance, MJH 3p showed a 4.7-fold higher IC50 value than 3n. These results establish 3p as the optimal MJH for inducing intracellular calcium release.
Next, we investigated the biological mechanisms of MJH-induced calcium release. Our previous work showed that MMs induce intracellular calcium release through the IP3 pathway.32–34 Given that both MJH and MMs generate mechanical work, we examined whether MJHs operate through the same pathway. The IP3 pathway is initiated by the activation of G protein-coupled receptors (GPCRs),55 which triggers the G protein subunit Gαq/11 to separate from the Gβγ subunit. Activated Gαq/11 stimulates phospholipase C (PLC),56 an enzyme responsible for breaking down phosphatidylinositol 4,5-bisphosphate (PIP2) into diacylglycerol (DAG) and inositol 1,4,5-trisphosphate (IP3). IP3 then binds to IP3 receptors (IP3R) on the ER membrane, triggering calcium release into the cytosol (Fig. 3a).
To systematically explore these mechanisms, we first examined whether MJH molecules colocalize with cellular membranes system, including plasma membrane, ER membrane and mitochondrial membrane.57 Colocalization images show the distribution of MitoTracker Green (in green, mitochondria) or ER-Tracker Green (in green, ER) relative to 3p (in red, Fig. 3b, and S36). The white color denotes the colocalization of green and red signals. Quantitative analysis from nine independent experiments (Fig. 3c) was conducted using Pearson's correlation coefficient (PCC, Rtotal) to assess colocalization between red and green channels (see SI for calculation details). PCC results showed moderate colocalization between 3p and ER (Rtotal = 0.68 ± 0.02) and slightly lower colocalization with mitochondria (Rtotal = 0.62 ± 0.03). Both tM1 and tM2 (calculated by Mander's overlap coefficient, MOC, see SI for calculation details) values were approximately 90% for 3p with either ER or mitochondria, confirming decent colocalization. While these results suggest that MJHs are broadly distributed within cellular membranes rather than specifically enriched in a single organelle, the presence of MJHs in the ER membrane, where IP3R is located, is consistent with a potential role of the IP3 pathway in the observed calcium response.
To identify the calcium source, we compared the responses of HEK293 cells treated with 3p and stimulated by light in the presence and absence of extracellular calcium. Identical calcium release magnitudes under both conditions (Fig. 3d) indicated an intracellular source. We then used specific inhibitors to probe the role of the IP3 pathway in the intracellular calcium release elicited by light-activated 3p (Fig. 3e–h). Treatment with either FR900359 (ref. 58) (Gα inhibitor) or gallein59 (Gβγ inhibitor), alone or in combination, reduced calcium release by ∼50%. The PLC antagonist U-73122 (ref. 60) almost completely blocked calcium release, strongly implicating IP3 signaling as the primary mechanism. However, the partial inhibition of MJH-induced calcium release by G protein inhibitors compared to complete inhibition by PLC antagonists, suggests that this effect may reflect incomplete pharmacological suppression of GPCR signaling or the contribution of additional calcium-dependent mechanisms, such as calcium influx (e.g., store-operated calcium entry, SOCE).58,61–63 Consistent with previous reports that certain GPCRs can respond to mechanical stimulation,64,65 MJH-induced mechanical perturbations may engage GPCR-associated signaling and thereby contribute to the observed calcium responses. These results highlight the potential of MJHs as a unique tool for modulating GPCR-associated calcium signaling.
To investigate potential photodynamic effects, we examined the role of ROS in MJH-induced calcium release. HEK293 cells were treated with ROS scavengers (10 mM thiourea and 0.25 mM sodium azide or MnTBAP up to 50 µM) for 45 min,39,41,42,66 and ROS levels were measured using CellROX Green Reagent.67 The fluoresces of CellROX increases when oxidized by ROS and is widely used to measure ROS generation in living cells. Although scavengers reduced cellular ROS levels, calcium release remained unchanged compared to untreated controls (Fig. 3i, j, S37 and S38), indicating that photodynamic effects do not drive MJH-induced calcium release. Moreover, while 3p induced the strongest calcium release among tested molecules (3a, 3d, 3p, and 3n), it did not generate higher ROS levels (Fig. 3k), further confirming that ROS generation is not the primary mechanism underlying MJH-induced calcium release.
The photothermal effect was also studied. HEK293 cells were transfected to be enabled to express a thermosensitive fluorescent protein ELP-TEMP. ELP-TEMP fluorescents both blue and green, and the intensities decrease when the temperature increases.68 Since the MJH molecules exhibit enhanced blue fluorescence upon light irradiation, we used the green channel to characterize the temperature changes during the experiment process. The result showed that despite of different abilities to induce calcium release, all the MJHs tested showed identical and negligible thermal effects (Fig. S39), indicating that the photothermal effect is also unlikely to be the reason for MJHs-induced calcium release.
To understand how the presence of different functional groups in MJHs affects their calcium release activity, we performed molecular dynamics simulations examining MJH interactions with cell membranes. MJHs with different functional groups showed varying depths of center of mass (COM) in the cell membrane (Fig. 4a–e and S40), with a clear negative correlation between COM depth and the efficiency of ICR induction. This relationship can be explained by the interactions of MJHs and the components of cell membranes.
Cell membranes consist primarily of phospholipids, with hydrophobic hydrocarbon tails oriented inward and hydrophilic phosphoryl head groups exposed outward. The hydrophobic core of MJHs preferentially inserts into the membrane's interior, while the polar or charged functional groups on the arm tend to interact with the phosphoryl heads of phospholipids. For MJH 3a, which lacks functional groups on its arm, exhibits the deepest COM depth as the entire molecule embeds within the membrane. In contrast, 3m, featuring a positively charged amine group on its arm, forms electrostatic interactions with the negatively charged phosphoryl head groups, limiting its insertion depth. The shallowest insertion is observed for MJHs 3p and 3q, which contain negatively charged sulfonate groups on their arms. Electrostatic repulsion between these charged sulfonate groups and the phosphoryl group prevents the arms of 3p and 3q from embedding into the membrane. As a result, only the hydrophobic core of these molecules inserts into the membrane, leading to the shallowest COM depths for 3p and 3q.
The observed negative correlation between COM depth and ICR induction efficiency could be rationalized by considering the spatial relationship between MJHs and GPCRs in the membrane. The key mechanosensitive region of GPCRs, helix 8,69 is positioned near the membrane boundary. We hypothesize that MJHs with shallower COM depths are situated closer to helix 8, which enhances their ability to interact with this mechanosensitive region (Fig. 4f). While our simulations do not include GPCR proteins and therefore do not directly capture protein–molecule interactions, this spatial proximity may increase the likelihood of mechanical perturbation of membrane-associated domains such as helix 8. This model provides a plausible explanation for the enhanced activity observed for molecules with shallower insertion depths.
Having elucidated the mechanism, we next investigated whether MJH-induced ICR can promote muscle contraction in C2C12 myotubes, differentiated muscle cells capable of contraction in response to increased intracellular calcium.70 In resting skeletal muscle (Fig. 5a), tropomyosin blocks myosin-binding sites on actin, preventing the interaction between myosin heads and actin filaments. When an action potential triggers calcium release from the sarcoplasmic reticulum, calcium binds to troponin, causing tropomyosin to shift and expose these binding sites. Myosin heads then form cross-bridges with actin and perform a power stroke, pulling actin filaments toward the sarcomere center, resulting in contraction. ATP binds to myosin, causing it to detach from actin, and its hydrolysis resets the myosin head for the next cycle. Muscle relaxation occurs as calcium is actively pumped back into the sarcoplasmic reticulum, restoring the resting state.71–73
As predicted, MJH 3p induced localized calcium release in C2C12 myotubes (Fig. 5b and SI Video 2), leading to myotube contractions (Fig. 5c and SI Videos 3 and 4). No calcium release or contraction occurred in myotubes treated with DMSO, 3p combined with U-73122, or 3n alone (Fig. 5c–g and S41). We quantified cellular contraction using image subtraction method, where pixel brightness corresponds to movement magnitude (see SI for calculation details). This analysis confirmed that 3p induced significant myotube contraction, while cells treated with DMSO, 3n, or 3p + U-73122 showed no contraction (Fig. 5h). These results demonstrate that MJHs can induce skeletal muscle contraction through intracellular calcium release, suggesting potential applications for muscle-related study or therapy.
To investigate whether calcium release could be induced in vivo, we used the model organism Hydra vulgaris, which is well-suited for this study due to its optical transparency, simple multicellular organization, and calcium-dependent physiological behavior that can be directly visualized using genetically encoded calcium indicators. Hydra is a millimeter-sized freshwater organism with two layers of tissue separated by an extracellular matrix, exhibiting radial symmetry with a cylindrical body structure. One end of its body connects to an oral region and tentacles, while the opposite end is connected to the foot (Fig. 6a).45,46 The transgenic Hydra line used in this study were genetically engineered to express GCaMP7b, enabling visualization of calcium dynamics in the endothelial epitheliomuscular cells (see Methods). Hydra were treated with each MJH (3p as the most potent MJH and 3m and 3n serve as control groups) and incubated for 24 h prior to experiments. After incubation, the MJH molecules effectively penetrated Hydra and distributed uniformly throughout its tissues (Fig. 6b).
Upon 640 nm laser stimulation, localized calcium release, indicated by green color fluorescence, was observed exclusively in Hydra treated with 3p. In contrast, Hydra treated with 3m, 3n, or DMSO exhibited minimal or undetectable responses (Fig. 6c, S42, SI Videos 5 and 6). Quantitative analysis of calcium release confirmed that only the 3p-treated group exhibited significant calcium elevation following stimulation (Fig. 6d and S43). The gradual increases in GCaMP7b fluorescence of 3n-, 3m- and DMSO-treated groups during the latter half of the experiment can be attributed to the spontaneous movements of Hydra, which exhibits calcium waves.74 These spontaneous increases were distinguished from MJH-induced responses by their full-body fluorescence accompanying Hydra contraction uncorrelated to the stimulation, whereas MJH-induced responses remained localized to the laser-irradiated area.
Notably, 3p demonstrated remarkable biocompatibility in Hydra compared to other MJHs. While Hydra treated with 3n and 3m at concentrations of 3 µM or higher all disintegrated and died, limiting their usable concentration to 1 µM, Hydra treated with up to 12 µM 3p remained healthy and continued normal growth post-experiments (Fig. S44). These results demonstrate that MJH 3p effectively induces calcium release in vivo with minimal toxicity, suggesting potential applications in biological systems.
Conclusion
Here, we demonstrate that cyanine dyes can function as MJHs to modulate intracellular calcium signaling with high spatiotemporal resolution. These light-activated molecules generate mechanical forces through vibronic-driven action (VDA), activating mechanosensitive IP3 pathways to trigger intracellular calcium release in skeletal muscle cells and in vivo in Hydra. Structure–activity relationship studies across 20 synthesized MJHs revealed that sulfonate-containing derivatives exhibited optimal calcium release efficiency with minimal toxicity. Molecular dynamics simulations provided mechanistic insights into the advantages conferred by the sulfonate group. The combination of intrinsic fluorescence for precise detection and red-light (640 nm) activation for deep tissue penetration and reduced phototoxicity, and the MJHs have the potential to become biologically safe and practically useful tools in research and, eventually, broader biomedical contexts. These properties, together with their demonstrated efficacy in vitro and in vivo, establish MJHs as a new class of molecular tools for controlling cellular signaling with potential therapeutic applications.Methods
General information of synthesis
All glassware was oven-dried overnight prior to use. All chemicals were purchased from commercial suppliers and used without further purification. All reactions were carried out under Argon atmosphere unless otherwise noted. Flash column chromatography was performed using 230–400 mesh silica gel. Thin layer chromatography (TLC) was performed using glass plates pre-coated with silica gel F254 0.25 mm layer thickness purchased from EM Science. 1H-NMR spectra were recorded at 500 MHz and 600 MHz, 13C-NMR spectra were recorded at 126 MHz and 151 MHz. Chemical shifts (δ) are reported in ppm relative to tetramethylsilane (TMS). HRMS (ESI) were conducted on Agilent Quadrupole-TOF LC/MS spectrometer. Synthesis and molecular characterization data are in the SI.Synthesis of symmetric MJHs (3a–3k)
Synthesis of non-symmetric MJHs (3l–3s)
UV-vis and emission spectra measurements
UV-vis spectra were acquired in a 1-cm quartz cuvette using a Shimadzu UV-2450 spectrophotometer. Molecules were dissolved in solvent (water, DMSO, methanol, ethanol, and isopropanol at 1 mM). Steady-state photoluminescence emission spectra were obtained on a Horiba Jobin Yvon FluoroLog exciting with a 450 W Xe lamp in front face (FF) mode. Relative quantum yield measurements were determined using ICG in DMSO as a reference (Φ = 0.13) as a standard. Photoluminescence quantum yield of unknown sample can be calculated by this equation, Φstd is the quantum yield of standard, Astd is absorbance intensity, Istd is emission intensity, nstd is the refractive index of solvent.Molecule preparation
MJH stock solutions were prepared in DMSO and diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 into the culture medium, resulting in a final DMSO concentration of 0.1% (v/v) in all MJH-treated conditions. The DMSO control was defined as 0.1% (v/v) DMSO in the absence of MJHs.
1000 into the culture medium, resulting in a final DMSO concentration of 0.1% (v/v) in all MJH-treated conditions. The DMSO control was defined as 0.1% (v/v) DMSO in the absence of MJHs.
Cell lines and culture conditions
HEK293 cells and C2C12 cells were obtained from the American Type Culture Collection (ATCC) and cultured with DMEM Medium (Corning, MT10013CV), the medium was supplemented with 10% FBS (Fetal Bovine Serum, Corning, MT35010CV) and 1% penicillin-streptomycin (Gibco, 15-140-122) in a humidified incubation system containing 5% CO2 at 37 °C. For the passage step, cells are detached with 0.05% trypsin–EDTA (Gibco, 25-300-054) and passaged every 2–3 days. For C2C12 differentiation, after the cells grew to near full confluency, media replacement was done with differentiation media (DMEM with 2% horse serum and 1% antimycin), and fresh differentiation media replacement was done every 1–2 d.Confocal microscopy
Confocal experiments were done with a Nikon A1-Rsi confocal microscope. Cells were plated in the µ-Plate 24-well plate (ibidi) with a concentration of 120![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 per well (HEK293 cells) or 7000 per well (C2C12 cells). Before imaging, the media was replaced with ECB-JR imaging buffer (119 mM NaCl, 5 mM KCl, 10 mM HEPES, 2 mM CaCl2, 1 mM MgCl2 (pH 7.2); 320 mOsm). Experiments were conducted 1–2 days after cells were plated except for the experiments with differentiated C2C12 cells, which took place after 4–9 days when differentiation media was replaced.
000 per well (HEK293 cells) or 7000 per well (C2C12 cells). Before imaging, the media was replaced with ECB-JR imaging buffer (119 mM NaCl, 5 mM KCl, 10 mM HEPES, 2 mM CaCl2, 1 mM MgCl2 (pH 7.2); 320 mOsm). Experiments were conducted 1–2 days after cells were plated except for the experiments with differentiated C2C12 cells, which took place after 4–9 days when differentiation media was replaced.
In experiments with IP3 pathway inhibitors, cells were initially incubated with MJHs and Fluo-4 and irradiated with 640 nm light as a positive control. After that, the inhibitor was immediately added to the well, and the cells were again irradiated with 640 nm light.
In experiments of ROS generation measurements, MJHs and CellROX were incubated with cells for 45 min. In experiments that ROS scavengers were used, the scavengers (thiourea 10 mM, sodium azide 0.25 mM) were incubated with cells for 45 min, and after replacing ECB-JR, the same amount of ROS scavengers were also added into ECB-JR solution.
In experiments testing photothermal effects, HEK293 cells were transfected with TEMP/pcDNA3.1(−) using lipofectamine 3000. The plasmid was a gift from Takeharu Nagai (Addgene plasmid # 178434; https://n2t.net/addgene:178434; RRID:Addgene_178434). The temperature was calibrated with a VAHeat Smart Substrate (part number SmS-SR) purchased from Interherence. VAHeat provides precise control of real-time temperature on its glass coverslip. We plated HEK293 on its accompanying cell culture reservoir and transfected them with ELP-TEMP. We increased the temperature from room temperature (22 °C) to 40 °C incrementally by 1 °C, and imaged the green channel at each degree on the Nikon A1 Rsi confocal microscope. 6 Individual cells were measured at each experiment, and 3 replications were conducted. We normalized their fluorescence to the initial 22 °C fluorescence to get the curve in Fig. S39a.
Super-resolution microscopy
Super-resolution microscope experiments were done with a Zeiss LSM980 Airyscan-2 microscope using Single Laser Point Scanning mode. Fluorescent dyes were added into the media with a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 dilution. After incubating with MJH molecule and the fluorescent dyes for 45 min, the media was replaced with ECB-JR, and the cells were imaged on the airyscan microscope. The dyes for organelle-labeling were obtained from Thermo Fisher. MitoTracker Green FM was used to label mitochondria and ER-Tracker Green was used to label ER.
1000 dilution. After incubating with MJH molecule and the fluorescent dyes for 45 min, the media was replaced with ECB-JR, and the cells were imaged on the airyscan microscope. The dyes for organelle-labeling were obtained from Thermo Fisher. MitoTracker Green FM was used to label mitochondria and ER-Tracker Green was used to label ER.
CCK-8 assay protocol on HEK293 cells
Dispense 100 µL cell suspension (20000 cells per well) in a 96-well plate. Pre-incubate the plate overnight in incubator (37 °C, 5% CO2). Add 100 µL media with various concentrations of MJHs to be tested to the plate (e.g., 1 µM concentration preparation![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 µL of 1 mM of molecules dissolved in 0.5 mL media and add 100 µL to each well of the plate without replacement of media) and incubate the plate for 3 h in the incubator. Add 10 vol% (20 µL) of CCK-8 solution to each well of the plate and keep incubating the plate for 1.5 hours in the incubator. Finaly, measure the absorbance at 450 nm using a microplate reader.
1 µL of 1 mM of molecules dissolved in 0.5 mL media and add 100 µL to each well of the plate without replacement of media) and incubate the plate for 3 h in the incubator. Add 10 vol% (20 µL) of CCK-8 solution to each well of the plate and keep incubating the plate for 1.5 hours in the incubator. Finaly, measure the absorbance at 450 nm using a microplate reader.
Molecular dynamics simulation
We performed molecular dynamics (MD) simulations to determine the equilibrium distance of dyes below the membrane surface. The details are provided in the SI (Molecular dynamics simulation section).Cell contraction magnitude quantification
Contraction magnitude was quantified using the image subtraction method. The first frame of the brightfield image was subtracted from all consecutive brightfield images, resulting in a new series of images where the first frame is totally black, and the pixel intensity represents the difference in each frame compared to the very first frame. The movements are positively correlated to the brightness of the pixel.In this new series of images, three unstimulated regions were chosen and averaged as background to rule out the effect of drift and normal cell activities during the course of the experiment. The background trace was subtracted from traces of each stimulated cells to get corrected traces. The corrected traces were processed in the same way as the fluorescence measurements (ΔF/F0) and zeroed at the point of stimulation.
Hydra transgenic line and maintenance
Hydra were maintained in a Hydra medium containing CaCl2·2H2O (1.00 mM), MgCl2·6H2O (0.10 mM), KNO3 (0.03 mM), NaHCO3 (0.50 mM) and MgSO4 (0.08 mM) in deionized water in a light-cycled (12 h![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 12 h light
12 h light![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) dark) incubator at 18 °C. Hydra were fed with freshly hatched Artemia nauplii (Brine Shrimp Direct) three times a week. All the experiments were performed at room temperature after starving the animals for two days. The transgenic line expressing the calcium indicator GCaMP7b under the Ef1α promoter in the endodermal epitheliomuscular cells was used for all experiments.
dark) incubator at 18 °C. Hydra were fed with freshly hatched Artemia nauplii (Brine Shrimp Direct) three times a week. All the experiments were performed at room temperature after starving the animals for two days. The transgenic line expressing the calcium indicator GCaMP7b under the Ef1α promoter in the endodermal epitheliomuscular cells was used for all experiments.
Hydra stimulation experiments
All Hydra experiments were performed two days after feeding. Before experiments, Hydra were incubated with MJH molecules for 24 h. Stimulation and imaging was done with a Nikon A1 confocal microscope and at room temperature. Hydra were irradiated by 640 nm laser with a power of 9.0 × 102 W cm−2 at a circular region 10 µm diameter for 3 s. Stimulations were performed when Hydra were relaxed to avoid their spontaneous contractions.Traces plot data analysis
The acquired images of confocal and super-resolution experiments were analyzed with Fiji. The fluorescence intensity was measured at each time point and processed using Python code. In the code, F0 was defined as the average fluorescence intensity in the first ten time points. The traces were normalized by F0 and then presented by ΔF/F0. Then the traces of each experiment were averaged and plotted with the gray regions as standard error of the mean.Statistical analysis
A minimum of three individual trials were conducted for each experiment and at least 6 cells were stimulated in each trial. The average maximum of each trial was calculated by a Python code to get the mean and standard error of the mean. Error bars and p values were calculated by GraphPad Prism.Author contributions
B. L., Y. R. and J. M. T. conceived the idea and designed the experiments. Y. R. conducted most of the biological experiments and characterizations with the help of B. L., D. A., J. L. B., C. A. O., A. S., G. C., S. X., T. S. and Z. W. B. L. conducted organic synthesis, characterizations and cytotoxicity tests. B. L., L. Z. and A. A. M. conducted UV-vis and spectroscopy measurement. V. V. and A. K. conducted the molecular dynamics simulations. Y. R., S. K. and J. T. R. conducted the Hydra experiments. B. L., Y. R. and J. M. T. wrote the original draft. J. M. T. was responsible for the funding acquisition, project administration, and overseeing of all phases of the project. All authors discussed the results and commented on the manuscript.Conflicts of interest
Rice University owns intellectual property on the use of MJH for permeabilization of cell membranes and use in calcium release. The intellectual property is presently unlicensed.Data availability
All data needed to support the conclusions in the paper are present in the manuscript and/or supplementary information (SI). Supplementary information: experimental procedures and characterization data for new compounds (NMR spectra, photophysical properties). See DOI: https://doi.org/10.1039/d5sc09717g.Acknowledgements
Funding was provided by The Discovery Institute and The Welch Foundation (C-2017-20220330). The authors thank Prof. Jacob T. Robinson at Rice University for kindly sharing his laboratory to culture the cells and thank Dr D. K. James for editing the manuscript. The characterization equipment used in this project is from the Shared Equipment Authority (SEA) at Rice University.References
- M. J. Berridge, M. D. Bootman and H. L. Roderick, Nat. Rev. Mol. Cell Biol., 2003, 4, 517–529 CrossRef CAS PubMed.
- M. J. Berridge, P. Lipp and M. D. Bootman, Nat. Rev. Mol. Cell Biol., 2000, 1, 11–21 CrossRef CAS PubMed.
- D. E. Clapham, Cell, 2007, 131, 1047–1058 CrossRef CAS PubMed.
- L. Fedrizzi, D. Lim and E. Carafoli, Biochem. Mol. Biol. Educ., 2008, 36, 175–180 CrossRef CAS PubMed.
- A. E. West, W. G. Chen, M. B. Dalva, R. E. Dolmetsch, J. M. Kornhauser, A. J. Shaywitz, M. A. Takasu, X. Tao and M. E. Greenberg, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 11024–11031 CrossRef CAS PubMed.
- B. K. Puri, Calcium Signaling and Gene Expression, in Calcium Signaling, ed. M. Islam, Springer, Cham, 2020, vol. 1131, pp. 537–545 Search PubMed.
- M. J. Berridge, BioEssays, 1995, 17, 491–500 CrossRef CAS PubMed.
- P. L. Greer and M. E. Greenberg, Neuron, 2008, 59, 846–860 CrossRef CAS PubMed.
- A. C. Dolphin and A. Lee, Nat. Rev. Neurosci., 2020, 21, 213–229 CrossRef CAS PubMed.
- A. G. Szent-Györgyi, Biophys. J., 1975, 15, 707–723 CrossRef PubMed.
- I. Y. Kuo and B. E. Ehrlich, Cold Spring Harbor Perspect. Biol., 2015, 7, a006023 CrossRef PubMed.
- A. J. Morgan and R. Jacob, Biochem. J., 1994, 300, 665–672 CrossRef CAS PubMed.
- O. Thastrup, P. J. Cullen, B. K. Drøbak, M. R. Hanley and A. P. Dawson, Proc. Natl. Acad. Sci. U. S. A., 1990, 87, 2466–2470 CrossRef CAS PubMed.
- T. G. Favero and J. J. Abramson, Cell Calcium, 1994, 15, 183–189 CrossRef CAS PubMed.
- K. D. Ryder and R. L. Duncan, J. Bone Miner. Res., 2001, 16, 240–248 CrossRef CAS PubMed.
- R. R. Resende, K. N. Gomes, A. Adhikari, L. R. G. Britto and H. Ulrich, Cell Calcium, 2008, 43, 107–121 CrossRef CAS PubMed.
- L. Khatib, D. E. Golan and M. Cho, FASEB J., 2004, 18, 1903–1905 CrossRef CAS PubMed.
- S. Buvinic, G. Almarza, M. Bustamante, M. Casas, J. López, M. Riquelme, J. C. Sáez, J. P. Huidobro-Toro and E. Jaimovich, J. Biol. Chem., 2009, 284, 34490–34505 CrossRef CAS PubMed.
- S. Staehlke, M. Bielfeldt, J. Zimmermann, M. Gruening, I. Barke, T. Freitag, S. Speller, U. Van Rienen and B. Nebe, Cells, 2022, 11, 2650 CrossRef CAS PubMed.
- M. J. Sanderson, A. C. Charles and E. R. Dirksen, Cell Regul., 1990, 1, 585–596 CrossRef CAS PubMed.
- M. Tsutsumi, K. Inoue, S. Denda, K. Ikeyama, M. Goto and M. Denda, Cell Tissue Res., 2009, 338, 99–106 CrossRef CAS PubMed.
- R. Zucker and J. Nolte, Nature, 1978, 274, 78–80 CrossRef CAS PubMed.
- G. H. Gold and J. I. Korenbrot, Proc. Natl. Acad. Sci. U. S. A., 1980, 77, 5557–5561 CrossRef CAS PubMed.
- P. Gorostiza and E. Y. Isacoff, Science, 2008, 322, 395–399 CrossRef CAS PubMed.
- T. Ishii, K. Sato, T. Kakumoto, S. Miura, K. Touhara, S. Takeuchi and T. Nakata, Nat. Commun., 2015, 6, 8021 CrossRef CAS PubMed.
- G. Ma, S. Wen, L. He, Y. Huang, Y. Wang and Y. Zhou, Cell Calcium, 2017, 64, 36–46 CrossRef CAS PubMed.
- V. García-López, F. Chen, L. G. Nilewski, G. Duret, A. Aliyan, A. B. Kolomeisky, J. T. Robinson, G. Wang, R. Pal and J. M. Tour, Nature, 2017, 548, 567–572 CrossRef.
- V. García-López, D. Liu and J. M. Tour, Chem. Rev., 2020, 120, 79–124 CrossRef PubMed.
- D. Arnold, B. Li, J. L. Beckham, K. Lopez-Jaime, C. Ayala-Orozco, Y. Rui, G. Chaudhry, J. M. Seminario and J. M. Tour, J. Am. Chem. Soc., 2025, 147, 22387–22401 CrossRef CAS PubMed.
- A. L. Santos, D. Liu, A. K. Reed, A. M. Wyderka, A. van Venrooy, J. T. Li, V. D. Li, M. Misiura, O. Samoylova, J. L. Beckham, C. Ayala-Orozco, A. B. Kolomeisky, L. B. Alemany, A. Oliver, G. P. Tegos and J. M. Tour, Sci. Adv., 2022, 8, eabm2055 CrossRef CAS PubMed.
- J. L. Beckham, A. R. van Venrooy, S. Kim, G. Li, B. Li, G. Duret, D. Arnold, X. Zhao, A. L. Santos, G. Chaudhry, J. T. Robinson and J. M. Tour, Nat. Nanotechnol., 2023, 18, 1051–1059 CrossRef CAS PubMed.
- M. J. Berridge, Nature, 1993, 361, 315–325 CrossRef CAS PubMed.
- K. Mikoshiba, J. Neurochem., 2007, 102, 1426–1446 CrossRef CAS PubMed.
- E. Decrock, M. De Bock, N. Wang, A. K. Gadicherla, M. Bol, T. Delvaeye, P. Vandenabeele, M. Vinken, G. Bultynck, D. V. Krysko and L. Leybaert, Biochim. Biophys. Acta, Mol. Cell Res., 2013, 1833, 1772–1786 CrossRef CAS PubMed.
- C. Ash, M. Dubec, K. Donne and T. Bashford, Laser Med. Sci., 2017, 32, 1909–1918 CrossRef PubMed.
- A. Mishra, R. K. Behera, P. K. Behera, B. K. Mishra and G. B. Behera, Chem. Rev., 2000, 100, 1973–2012 CrossRef CAS PubMed.
- W. Sun, S. Guo, C. Hu, J. Fan and X. Peng, Chem. Rev., 2016, 116, 7768–7817 CrossRef CAS PubMed.
- L. Štacková, E. Muchová, M. Russo, P. Slavíček, P. Štacko and P. Klán, J. Org. Chem., 2020, 85, 9776–9790 CrossRef PubMed.
- C. Ayala-Orozco, D. Galvez-Aranda, A. Corona, J. M. Seminario, R. Rangel, J. N. Myers and J. M. Tour, Nat. Chem., 2024, 16, 456–465 CrossRef CAS PubMed.
- C. Ayala-Orozco, G. Li, B. Li, V. Vardanyan, A. B. Kolomeisky and J. M. Tour, Adv. Mater., 2024, 36, 2309910 CrossRef CAS PubMed.
- B. Li, C. Ayala-Orozco, T. Si, L. Zhou, Z. Wang, A. A. Martí and J. M. Tour, Adv. Sci., 2024, 11, 2405965 CrossRef CAS PubMed.
- C. Ayala-Orozco, V. Vardanyan, K. Lopez-Jaime, Z. Wang, J. M. Seminario, A. B. Kolomeisky and J. M. Tour, RSC Mechanochem., 2025, 2, 706–722 RSC.
- G. Shaw, S. Morse, M. Ararat and F. L. Graham, FASEB J., 2002, 16, 869–871 CrossRef CAS PubMed.
- D. K. McMahon, P. A. Anderson, R. Nassar, J. B. Bunting, Z. Saba, A. E. Oakeley and N. N. Malouf, Am. J. Physiol.: Cell Physiol., 1994, 266, C1795–C1802 CrossRef CAS PubMed.
- T. Fujisawa, Dev. Dyn., 2003, 226, 182–189 CrossRef CAS PubMed.
- S. Ghaskadbi, Resonance, 2020, 25, 1197–1213 CrossRef.
- C. Shi, J. B. Wu and D. Pan, J. Biomed. Opt., 2016, 21, 050901 CrossRef PubMed.
- A. P. Gorka, R. R. Nani and M. J. Schnermann, Acc. Chem. Res., 2018, 51, 3226–3235 CrossRef CAS PubMed.
- T. C. Pham, V.-N. Nguyen, Y. Choi, S. Lee and J. Yoon, Chem. Rev., 2021, 121, 13454–13619 CrossRef CAS PubMed.
- Y. Cui, A. Lauchner, A. Manjavacas, F. J. García de Abajo, N. J. Halas and P. Nordlander, Nano Lett., 2016, 16, 6390–6395 CrossRef CAS PubMed.
- G. Orlandi and W. Siebrand, J. Chem. Phys., 1973, 58, 4513–4523 CrossRef CAS.
- F.-F. Kong, X.-J. Tian, Y. Zhang, Y.-J. Yu, S.-H. Jing, Y. Zhang, G.-J. Tian, Y. Luo, J.-L. Yang, Z.-C. Dong and J. G. Hou, Nat. Commun., 2021, 12, 1280 CrossRef CAS PubMed.
- C. Ayala-Orozco, B. Li, G. Li and J. M. Tour, Chem. Sci., 2025, 16, 2718–2729 RSC.
- K. Chamchoy, D. Pakotiprapha, P. Pumirat, U. Leartsakulpanich and U. Boonyuen, BMC Biochem., 2019, 20, 4 CrossRef PubMed.
- V. Dhyani, S. Gare, R. K. Gupta, S. Swain, K. V. Venkatesh and L. Giri, Cell. Signalling, 2020, 74, 109717 CrossRef CAS PubMed.
- C. A. Bill and C. M. Vines, Adv. Exp. Med. Biol., 2020, 1131, 215–242 CrossRef CAS PubMed.
- K. W. Dunn, M. M. Kamocka and J. H. McDonald, Am. J. Physiol.: Cell Physiol., 2011, 300, C723–C742 CrossRef CAS PubMed.
- R. Schrage, A.-L. Schmitz, E. Gaffal, S. Annala, S. Kehraus, D. Wenzel, K. M. Büllesbach, T. Bald, A. Inoue, Y. Shinjo, S. Galandrin, N. Shridhar, M. Hesse, M. Grundmann, N. Merten, T. H. Charpentier, M. Martz, A. J. Butcher, T. Slodczyk, S. Armando, M. Effern, Y. Namkung, L. Jenkins, V. Horn, A. Stößel, H. Dargatz, D. Tietze, D. Imhof, C. Galés, C. Drewke, C. E. Müller, M. Hölzel, G. Milligan, A. B. Tobin, J. Gomeza, H. G. Dohlman, J. Sondek, T. K. Harden, M. Bouvier, S. A. Laporte, J. Aoki, B. K. Fleischmann, K. Mohr, G. M. König, T. Tüting and E. Kostenis, Nat. Commun., 2015, 6, 10156 CrossRef CAS PubMed.
- G. Sanz, I. Leray, A. Muscat, A. Acquistapace, T. Cui, J. Rivière, S. Vincent-Naulleau, V. Giandomenico and L. M. Mir, BMC Res. Notes, 2017, 10, 541 CrossRef PubMed.
- D. MacMillan and J. McCarron, Br. J. Pharmacol., 2010, 160, 1295–1301 CrossRef CAS PubMed.
- J. G. Schlegel, M. Tahoun, A. Seidinger, J. H. Voss, M. Kuschak, S. Kehraus, M. Schneider, M. Matthey, B. K. Fleischmann, G. M. König, D. Wenzel and C. E. Müller, ACS Pharmacol. Transl. Sci., 2021, 4, 888–897 CrossRef CAS PubMed.
- G. A. Sanchez, A. V. Smrcka and E. M. Jutkiewicz, Mol. Pharmacol., 2024, 106, 47–55 CrossRef CAS PubMed.
- A. B. Parekh and J. W. Putney, Physiol. Rev., 2005, 85, 757–810 CrossRef CAS PubMed.
- K. Hardman, A. Goldman and C. Pliotas, Curr. Opin. Physiol., 2023, 35, 100689 CrossRef CAS.
- A. J. Shetty, A. Sirbu and P. Annibale, J. Mol. Endocrinol., 2025, 75, e240147 CAS.
- Q. Zhou, C. Gensch, C. Keller, H. Schmitt, J. Esser, M. Moser and J. K. Liao, Vasc. Pharmacol., 2015, 72, 163–171 CrossRef CAS PubMed.
- E. Lozoya-Gloria, I. Cornejo-Corona, H. R. Thapa, D. R. Browne and T. P. Devarenne, Bio-Protoc., 2017, 7, e2508 Search PubMed.
- C. Q. Vu, S. Fukushima, T. Wazawa and T. Nagai, Sci. Rep., 2021, 11, 16519 CrossRef CAS PubMed.
- S. Erdogmus, U. Storch, L. Danner, J. Becker, M. Winter, N. Ziegler, A. Wirth, S. Offermanns, C. Hoffmann, T. Gudermann and M. Mederos y Schnitzler, Nat. Commun., 2019, 10, 5784 CrossRef CAS PubMed.
- D. Yaffe and O. Saxel, Nature, 1977, 270, 725–727 CrossRef CAS PubMed.
- A. F. Huxley and R. Niedergerke, Nature, 1954, 173, 971–973 CrossRef CAS PubMed.
- W. Lehman, R. Craig and P. Vibert, Nature, 1994, 368, 65–67 CrossRef CAS PubMed.
- M. A. Geeves and K. C. Holmes, Adv. Protein Chem., 2005, 71, 161–193 CrossRef CAS PubMed.
- J. R. Szymanski and R. Yuste, Curr. Biol., 2019, 29, 1807–1817e3 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally: Yuchen Rui, Bowen Li. |
| This journal is © The Royal Society of Chemistry 2026 |