Open Access Article
Open Access ArticleA high-affinity dopamine aptamer: implications of library diversity and negative selection
Yi Yuab,
Yuanli Li b and
Juewen Liu
b and
Juewen Liu *b
*b
aSchool of Laboratory Medicine, Hubei University of Chinese Medicine, 16 Huangjia Lake West Road, Wuhan, 430065, PR China
bDepartment of Chemistry, Waterloo Institute for Nanotechnology, University of Waterloo, Waterloo, Ontario N2L 3G1, Canada. E-mail: liujw@uwaterloo.ca
First published on 23rd March 2026
Abstract
In 2018, a DNA aptamer for dopamine was isolated after extensive negative selection steps. Herein, a new selection experiment was carried out under the same conditions except that no negative selections were performed. This selection yielded a single family of binding sequences, with a Kd of 424 nM from isothermal titration calorimetry, 6-fold lower compared to the previous aptamer, although they differ only by one nucleotide. Using the fluorescence strand-displacement reaction, the newly selected aptamer had a 4-fold faster release rate, 2-fold lower affinity, and 15-fold lower apparent Kd compared to a quencher-labeled DNA acting as a capture strand surrogate. All these properties would favor the selection of this new higher affinity aptamer. This aptamer was missed in the previous work likely by the overly stringent negative selection. Positive selection needs to be carried out at a concentration a few times higher than the Kd of the best aptamer, whereas negative selection needs to be done at a negative target concentration that does not induce significant removal of the best aptamers. This work not only identifies a high-affinity dopamine aptamer mutant but also prompts rethinking of strategies to increase library diversity and optimize the concentration of negative targets in aptamer selection.
Introduction
Dopamine is a critical neurotransmitter known for its roles in the regulation of reward, motivation, and pleasure pathways in the brain.1 Dopamine modulates motor control, and its deficiency is linked to Parkinson's disease, whereas its excess is tied to addiction. In addition, dopamine also influences the functions of the heart and kidneys, as well as hormone release.2 Therefore, real-time, continuous, and in situ monitoring of dopamine is essential for neuroscience.3Monitoring of dopamine is currently done by microdialysis coupled with mass spectrometry4 or by electrochemical detection based on its intrinsic redox potential.5 However, electrochemical detection is hampered by the presence of other redox-active molecules in the same redox potential window. Therefore, having a highly specific recognition molecule is critical to boost specificity. While the use of enzymes for dopamine detection was attempted with such enzymes as laccase,6 copper efflux oxidase7 and tyrosinase,8 these do not have high specificity. Antibodies have also been developed for dopamine, but they have not attracted analytical applications in vivo.9
DNA aptamers are ideal for the detection of small molecules like dopamine due to their fast binding kinetics, high stability, small size and ease of labeling.10–16 In addition, aptamers often show a large conformational change upon target binding, allowing versatile detection methods, including optical and electrochemical detection.17,18
A dopamine aptamer was reported by the Stojanovic group in 2018,19 and it has since been widely used for fundamental studies and biosensors.20–27 This aptamer has a Kd of 2.7 µM, as determined using ITC by the Johnson group,28 similar to our previous report (2.2 µM).29 Although a Kd of 150 nM was determined using the strand-displacement reaction,19 ITC is a more direct method. Our lab studied this aptamer by making mutations and found that most mutations abolished binding, suggesting that it is a highly conserved sequence.29 This aptamer was isolated using the capture-SELEX method,19 where a DNA library was immobilized via hybridization to a short capture strand attached to a bead.30–33 By flowing a dopamine solution through the immobilized DNA library, aptamer sequences that released upon binding to dopamine were collected and amplified via PCR. To achieve high selectivity, the Stojanovic group performed extensive negative selections using up to 200 µM L-DOPA, serotonin, tryptophan and tyrosine with extensive washing. Since Sanger sequencing was used, no sequence alignment information was available.19
Considering the importance of dopamine in analytical chemistry and its low concentration in physiological conditions, higher-affinity aptamers are desirable. Herein, we did another dopamine selection with the following two differences: (1) no negative selections and (2) using deep sequencing. Interestingly, we isolated a highly conserved mutant of the previously reported aptamer, and in our selection, the previous aptamer almost completely disappeared. Binding assays indicated that our mutant had a higher binding affinity to dopamine, reaching a nanomolar level by ITC. This result led us to rethink the optimal conditions for negative selection. In light of the striking similarity between the two independently selected aptamers from different laboratories, statistical analysis was conducted to rationalize and evaluate the observed convergence.
Materials and methods
Chemicals
All DNA samples were from Integrated DNA Technologies (Coralville, IA), except for the quencher-labeled DNA, which was from Eurofins. The DNA sequences are listed in Table S1. Dopamine and other chemicals were from Sigma-Aldrich. All the buffers and solutions were prepared using Milli-Q water.Capture-SELEX
Capture-SELEX was undertaken following an established strategy in our lab.34 In the first round of selection, a 500 pmol DNA library (approximately 3 × 1014 distinct sequences, each with a 36-nt random region) was hybridized with a biotinylated capture strand for immobilization to streptavidin-coated agarose resin. Unbound or weakly associated sequences were washed away using selection buffer (1× PBS, 2 mM MgCl2). A dopamine solution prepared in the same buffer was then incubated with the library for 10 min to initiate competitive elution of binding sequences. During the selection process, dopamine was at 100 µM for rounds 1–10 and 20 µM for rounds 11–15. In each round, 250 µL dopamine was added three times. DNA released from the resin was collected and subjected to PCR amplification. For the first round, all the released DNA strands were used for PCR amplification to maximally retain sequence diversity. The DNA library from the subsequent rounds was generated using PCR (typically around 15 rounds), and approximately 30% of the dopamine-eluted DNA was used as template. From round 2 onwards, the library has already become much less diverse due to the 15 rounds of PCR aimed to produce at least 100 pmol PCR products. Biotinylated PCR amplicons were then re-immobilized onto streptavidin agarose for strand separation using 0.2 M NaOH and incubated for 10 min at room temperature. The released single-stranded DNA was purified to serve as the library for the next round. The PCR products from round 15 of the selection were subjected to deep sequencing.Isothermal titration calorimetry (ITC)
Isothermal titration calorimetry (ITC) experiments were carried out on a MicroCal ITC 200 instrument. All aptamer and target solutions were prepared in the same selection buffer. Before measurement, DNA aptamers were thermally annealed, allowed to cool to room temperature, and subsequently degassed for 5 min. Fresh solutions of dopamine were prepared immediately before titration and were also degassed for 5 min to remove dissolved gases that could interfere with the baseline stability. During each experiment, the sample cell was filled with 209 µL of the aptamer solution, while the syringe was loaded with 38.5 µL of the target solution. After an initial 0.5-µL priming injection, a total of 19 injections of 2 µL each were delivered at 25 °C. Each injection lasted for 20 s, with an interval of 360 s between successive injections to allow the system to reach thermal equilibrium. The resulting thermograms were analyzed using Origin software, and the binding isotherms were fitted to a one-site binding model to obtain the binding constants and thermodynamic parameters.ThT fluorescence spectroscopy
Fluorescence measurements were performed on a Cary Eclipse fluorometer. For each experiment, a 500 µL reaction mixture was prepared by combining 5 µL of aptamer solution (100 µM), 5 µL of ThT solution (100 µM), and 485 µL of buffer (1× PBS containing 2 mM MgCl2). The mixture was transferred to a quartz cuvette, and dopamine was incrementally added to achieve a final concentration of 240 µM. The samples were excited at 440 nm, and fluorescence emission spectra were collected from 460 to 520 nm. Fluorescence intensities at 490 nm were measured for quantitative analysis. The apparent dissociation constant (Kd) was determined by fitting the data to the equation: F = F0 + aKd/(Kd + [Dopamine]), where a represents the maximal fluorescence change at saturation.Strand-displacement reaction
A sensor complex was prepared by annealing 1 µM FAM-labeled aptamer with 4 µM quencher-labeled complementary strand in the selection buffer, followed by dilution to a final concentration of 20 nM FAM-labeled aptamer prior to measurement. Kinetic assays were performed in a 96-well microplate using a Tecan F200 Pro microplate reader. For each measurement, 98 µL of the 20 nM sensor complex was dispensed into a well and allowed to equilibrate for 5 min to obtain the background signal. Subsequently, 2 µL of the target solution was added to initiate the reaction. Fluorescence was recorded with excitation and emission wavelengths set to 485 nm and 535 nm, respectively. The fluorescence signals were continuously monitored for at least 30 min with a sampling interval of 30 s.Results and discussion
Capture-SELEX for dopamine
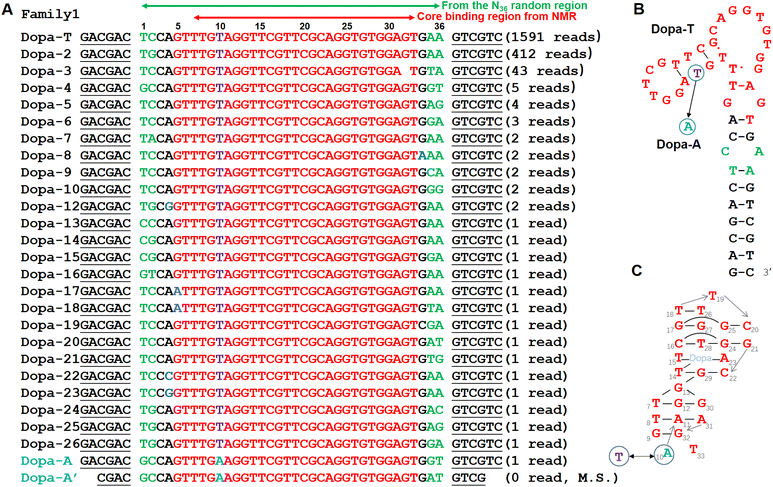
The previous dopamine selection was performed using a library containing 36 random nucleotides (N36). In this work, we adopted the same library design, the same capture-SELEX method, and also started with 100 µM dopamine.19,33,35 The main difference was that no negative selection was performed in our selection. We performed a total of 15 rounds of selection and ultimately enriched a sequence family that accounted for approximately 26% of the final library (Fig. 1A).Among these sequences, the most abundant variant was designated Dopa-T. This sequence differs from the previously reported aptamer by only a single nucleotide within the conserved region, highlighted by a circle in Fig. 1B (A or T). Accordingly, our aptamer can be regarded as a mutant variant of the earlier sequence, featuring an A-to-T substitution at this position. To explicitly distinguish this single-nucleotide difference, we refer to the corresponding adenine-containing variant as Dopa-A.
Systematic sequence analysis
As shown in Fig. 1A, Dopa-T belongs to a larger sequence family comprising 2055 copies that all share the same conserved Dopa-T core motif, whereas only a single copy of Dopa-A was identified. In other words, Dopa-T is 2055-fold more abundant than Dopa-A in the final pool. If we focus on the N36 region (highlighted by the top green arrow), sequence variations occur primarily at the terminal nucleotides marked in green and black, with the green bases showing the greatest variability. These variable terminal regions cannot form perfect base pairs, although previous studies have shown that converting this region to canonical base pairs can enhance binding affinity.28,29 Therefore, this variable region is likely introduced and enriched through PCR bias. If a perfect base pair were present at this position, the resulting long and highly stable hairpin stem could impede efficient PCR amplification. Thus, the mismatch likely represents a functional compromise by providing sufficient structural stability for target binding while avoiding amplification suppression during SELEX.Another minor difference between our newly selected aptamers and the previously reported ones is the terminal base pairs. We named the previously reported aptamer Dopa-A′ (the last row of the sequences in Fig. 1B). For a fair comparison, we mainly compared Dopa-T with Dopa-A, which have the same terminal base pairs.
Rationalizing the selection of similar aptamers: a long conserved region
At first glance, it seems statistically improbable that two independent laboratories would enrich aptamers with such a high degree of similarity. Our dopamine selection started with a 500 pmol library (3 × 1014 random sequences), while the Stojanovic group started with a larger library containing ∼1016 random sequences.19 The Dopa-T, Dopa-A and Stojanovic's Dopa-A′ aptamer share 32 identical nucleotides. Within a random library, the chance of having a motif with 32 conserved nucleotides is 1/432 ≅ 5.4 × 10−20. Therefore, the probability that this aptamer motif existed in the initial libraries of either SELEX experiment is close to zero, meaning that neither laboratory should have identified this motif if they relied solely on sequences present at the outset. Yet, both groups independently discovered the same aptamer architecture. It should be noted that we did not obtain a single sequence; rather, many closely related sequences formed a distinct family. Among the 27 sequences in this family, 9 positions are variable, leaving 27 fixed positions within the N36 region. The presence of such a large number of sequence variants within one family further rules out the possibility that all of them were present in the initial library.To further confirm our findings, we repeated the selection under the same conditions and tracked the enrichment of aptamer sequences at rounds 3, 6, 9, and 12. The motif first appeared at round 9 with a frequency of 0.3% and increased to 32.2% by round 12. At round 12, the dominant sequence still belonged to the Dopa-T group (7487 copies), whereas only two copies of Dopa-A were detected (Fig. S1). These results further indicate that the identified dopamine aptamer motif was not a coincidence but a reproducible outcome of the selection process.
This outcome can be rationalized by invoking PCR-induced mutations, which effectively expanded the sequence diversity during the selection process. As 3 × 1014 (our initial library size) is approximately 424, we can expect that our library already contained sequences that have 24 out of the 27 absolutely fixed nucleotides, and thus the library is only three mutations away from this core sequence. If we assume an error rate of 1 in 36![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 nucleotide incorporations for Taq polymerase (the typical range is 10
000 nucleotide incorporations for Taq polymerase (the typical range is 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 to 50
000 to 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000), then in an N36 library, we would expect approximately one mutation per 1000 amplified sequences. Across 15 rounds of PCR, such errors can accumulate, ultimately affecting more than 1% of the pool. This mutation rate is further compounded during each SELEX round, and any advantageous mutations are preferentially enriched under selection pressure. It is noteworthy that it is nearly impossible to distinguish whether a given sequence originated from the initial library or arose through PCR-induced mutation, since from a library of ∼1014 sequences, we typically sequence only on the order of 104. Regardless of whether a sequence is originally present in the library or generated via mutation from a previous round, it is subjected to the same selection pressure as its peer sequences in that round of selection.
000), then in an N36 library, we would expect approximately one mutation per 1000 amplified sequences. Across 15 rounds of PCR, such errors can accumulate, ultimately affecting more than 1% of the pool. This mutation rate is further compounded during each SELEX round, and any advantageous mutations are preferentially enriched under selection pressure. It is noteworthy that it is nearly impossible to distinguish whether a given sequence originated from the initial library or arose through PCR-induced mutation, since from a library of ∼1014 sequences, we typically sequence only on the order of 104. Regardless of whether a sequence is originally present in the library or generated via mutation from a previous round, it is subjected to the same selection pressure as its peer sequences in that round of selection.
Several striking examples in the literature highlight how powerful mutation can be in driving aptamer evolution. For instance, in one study, the Holliger group began with a simple polyadenine sequence flanked by primer-binding regions, yet through iterative rounds of mutation and selection, they successfully evolved aptamers capable of binding both ATP and GTP.47 To promote mutation, error-prone PCR was used. In another example, the Li laboratory began with a fixed 50-nucleotide DNA fragment as the so-called “random region” and successfully evolved a DNAzyme after 26 rounds of selection.48 Notably, this study did not intentionally employ error-prone PCR. Together, these cases demonstrate that substantial sequence diversity can emerge through directional evolution, despite minimal or nonexistent diversity in the starting pool. In this context, our libraries already contained highly diverse sequences, and even if the exact aptamer sequence was not present initially, mutations accumulated during SELEX could readily facilitate the evolution of such aptamers.
Structural biology reason for the evolution of this aptamer motif
Although this aptamer motif may not be the only sequence capable of binding dopamine, it may be the only one that does so with high affinity. Recently, the solution NMR structure of a 27-nucleotide minimal motif of the dopamine aptamer was determined,36 revealing that only two of the 18 base pairs within the binding architecture are canonical Watson–Crick pairs (Fig. 1C). The predominance of non-canonical base-pairing interactions implies that the sequence must be highly conserved to maintain the intricate three-dimensional fold required for high-affinity binding. Thus, the NMR structure confirms the practical need for a long stretch of conserved nucleotides, with the only position likely tolerating mutation being the A/T-variable site. Given that both our selection and the earlier selection performed by the Stojanovic group used a final dopamine concentration of 20 µM, the convergence on this aptamer motif (Kd in the range of 0.4 to 3.5 µM) likely reflects an evolutionary constraint, suggesting that this structure may be the only one capable of achieving high-affinity binding under these selection pressures. With a low target concentration, only high-affinity aptamers can survive.34,44Typical aptamers contain conserved sequence motifs connected by non-conserved regions, with the latter often folding into simple structural elements such as hairpins. For aptamers of this type, independent SELEX experiments performed in different laboratories against the same target frequently converge on sequences that share the same conserved motifs but vary in non-conserved hairpins. In most cases, the combined length of these conserved elements totals roughly 17 nucleotides or fewer (e.g., the classical ATP aptamer with 12 conserved nucleotides,35 a voriconazole aptamer with 9 conserved nucleotides,43 a guanine aptamer with 17 conserved nucleotides45). The chance of a 17-nt motif existing in initial libraries is high (a library containing 1014 random sequences can cover a 17-nt conserved motif approximately 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 times, since 417 ≅ 1010). In the very first paper by Ellington and Szostak, it was estimated that in 1010 random sequences, there is one aptamer.46 However, this dopamine aptamer has a much longer conserved region due to its folding to a very unique structure. Likely, any laboratory using this library design and performing a successful dopamine aptamer selection with concentrations as low as 20 µM would enrich the same motif.
000 times, since 417 ≅ 1010). In the very first paper by Ellington and Szostak, it was estimated that in 1010 random sequences, there is one aptamer.46 However, this dopamine aptamer has a much longer conserved region due to its folding to a very unique structure. Likely, any laboratory using this library design and performing a successful dopamine aptamer selection with concentrations as low as 20 µM would enrich the same motif.
Dopa-T is a higher-affinity aptamer
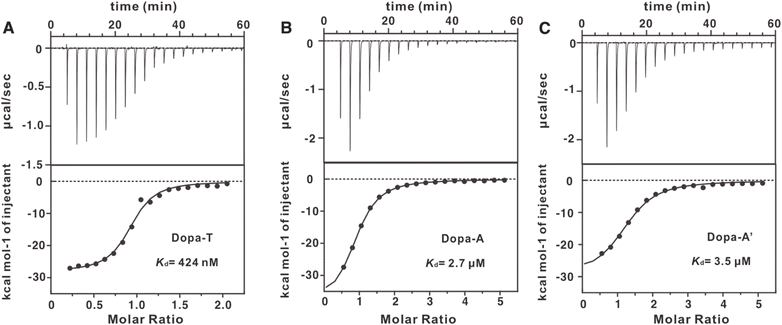
To evaluate binding of the aptamers, we performed ITC on the three sequences (Dopa-T, Dopa-A and Dopa-A′, Fig. 2). Among them, Dopa-T has the highest binding affinity, reaching a Kd of 424 nM. Dopa-A and Dopa-A′ have similar Kd values of 2.7 µM and 3.5 µM, respectively, and these values are consistent with previous ITC measurements.28,29 The thermodynamic constants are listed in Table 1. Therefore, we obtained a higher-affinity mutant.| Aptamer | Kd (µM) | ΔH (kcal mol−1) | ΔS (cal mol−1 K−1) | N |
|---|---|---|---|---|
| Dopa-T | 0.42 | −28 | −65.3 | 0.90 |
| Dopa-A | 2.7 | −30 | −74.8 | 1.26 |
| Dopa-A′ | 3.5 | −39 | −105 | 0.88 |
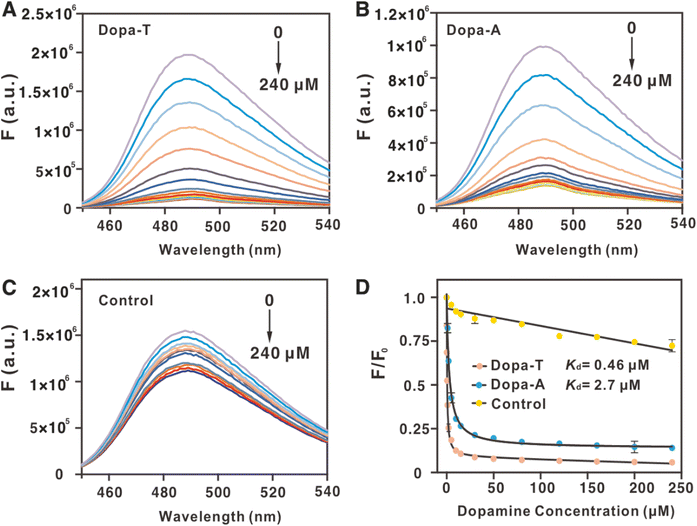
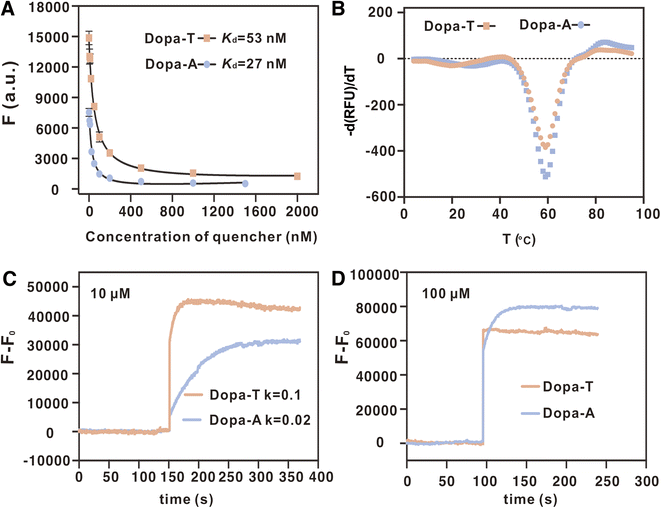
While ITC provided a gold-standard measurement of binding,37 it is a slow experiment. To further understand the binding, we also performed convenient thioflavin T (ThT) fluorescence spectroscopy.38 ThT was first mixed with an aptamer to boost its fluorescence. Then, dopamine or other molecules were titrated. If binding occurred, some ThT molecules might be released to show decreased fluorescence. We previously showed that the Kd values measured by ThT fluorescence and by ITC are quite comparable for most aptamers.39 As shown in Fig. 3A, up to 90% fluorescence quenching was achieved when titrating dopamine into the Dopa-T aptamer (Fig. 3A), and the fitted apparent Kd was 0.46 µM (Fig. 3D), very close to the 0.42 µM determined by ITC. Similarly, the Dopa-A aptamer showed an apparent Kd of 2.7 µM (Fig. 3B and D), identical to that determined by ITC. Using a control sequence, no binding was detected (Fig. 3C and D). Therefore, using an independent method, a similar conclusion on the tighter binding of the Dopa-T aptamer was also reached.
DNA strand-displacement-based fluorescent biosensors
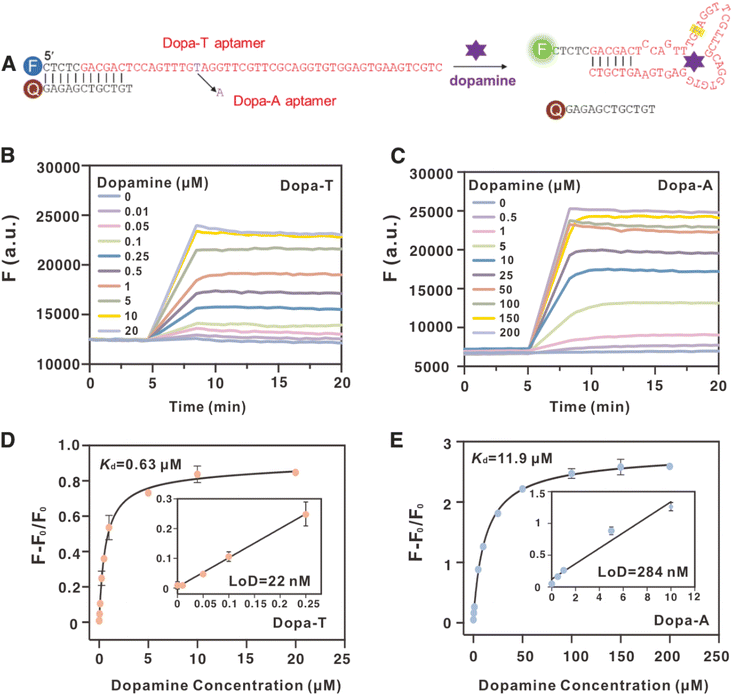
We then compared these two aptamers using the strand-displacement reaction.40 Each aptamer was extended on the 5′-end and hybridized to a short quencher-labeled strand, leading to quenched fluorescence. Upon binding to dopamine, the quencher strand is released, and fluorescence increase is expected (Fig. 4A). This system can also mimic the aptamer selection step, where the quencher-labeled strand serves the same role as the biotinylated capture strand. Therefore, studying this strand-displacement system can offer further insights into aptamer selection.We monitored the fluorescence of the two aptamer systems for 5 min to ensure that a stable background fluorescence was achieved before dopamine was added (Fig. 4B and C). The first observation is that the Dopa-A aptamer showed better fluorescence quenching, suggesting that it has a stronger affinity to the quencher-labeled strand. At 5 min, various concentrations of dopamine were added, and an immediate fluorescence enhancement was observed. A higher dopamine concentration resulted in a stronger final fluorescence. As the Dopa-A system had better initial quenching, it also had a higher fold enhancement of fluorescence. We then plotted the fluorescence change as a function of dopamine concentration (Fig. 4D). The Dopa-T aptamer had better sensitivity, with an apparent Kd of 0.63 µM (1.6-fold that of the free aptamer), whereas the Dopa-A aptamer had an apparent Kd of 11.9 µM (3.9-fold that of the free aptamer). Thus, due to hybridization to the quencher-labeled strand, the apparent affinity of the Dopa-A aptamer decreased even more. The relatively smaller change of Kd also indicated weaker hybridization of Dopa-T to the quencher-labeled strand.41 Using these two calibration curves, we calculated the limit of detection (LoD) to be 22 nM using the Dopa-T aptamer (inset of Fig. 4D) and 284 nM using the Dopa-A aptamer (inset of Fig. 4E).
Comparing the two aptamers by dissecting the capture-SELEX steps
In this work, we used the same library design, PCR primers, buffer, and initial dopamine concentrations as those used by the Stojanovic group.19 The only difference was that no negative selection was performed. The outcome of the selection, however, was different in an interesting way. While the higher affinity of Dopa-T may explain its dominance in our selection, it cannot explain why the Stojanovic group did not observe this higher affinity sequence. Instead, they observed the lower affinity one, which hardly presented in our final library. Given that these two aptamers differed by only one nucleotide, we took this opportunity to understand their enrichment by following all the steps of aptamer selection.The first step is hybridization of aptamers to the capture strand (here, we used the quencher-labeled strand as a surrogate). We measured the hybridization of the quencher-labeled strand to the FAM-labeled aptamers and calculated the Kd values (Fig. 5A). The affinity between the quencher-labeled strand and Dopa-A is slightly higher (about 1-fold difference in Kd). Based on mFold-predicted secondary structure in the absence of dopamine,42 it might be that the Dopa-A aptamer can form a more stable internal structure in the absence of dopamine, which makes the whole structure less interfering for hybridization.
The fluorophore/quencher system allowed us to compare their stability by measuring their melting curves. The first derivatives of the melting curves showed a similar melting transition at approximately 60 °C (Fig. 5B). As we used approximately 1 µM capture strand, they should both be captured efficiently and stably for the selection, and the Dopa-A aptamer might even have a slight advantage due to its slightly lower Kd with the capture strand.
After comparing the hybridization step, we then compared the target-induced aptamer release step. Both selections started with 100 µM dopamine, and this concentration should sufficiently release both aptamers (Fig. 4D and E). At lower dopamine concentrations, the Dopa-T aptamer would be released more. We recently discovered that some aptamers may be released more slowly than other aptamers.43 Thus, we compared the kinetics of release using both 10 and 100 µM dopamine (Fig. 5C and D). The Dopa-T aptamer released about 4-fold faster based on the rate fitting. Therefore, the Dopa-T aptamer has the advantage of being selected due to faster release kinetics and a smaller apparent Kd. This is exactly what we observed in our final library; Dopa-T was over 2000-fold more abundant than Dopa-A.
High-affinity aptamers might be removed by stringent negative selection
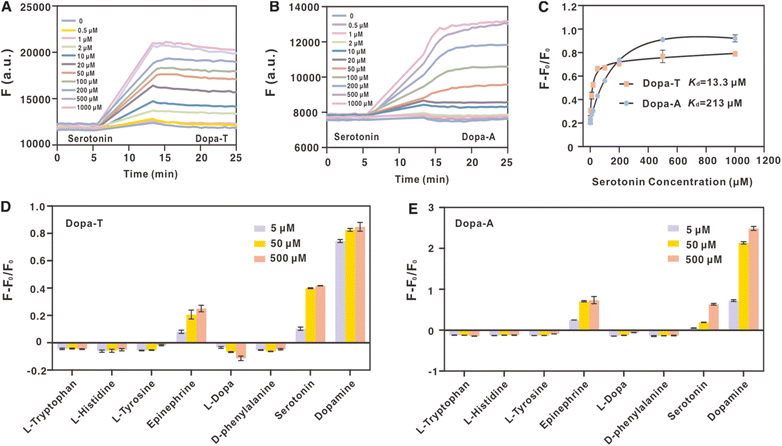
After this analysis, we could not explain why Dopa-A was enriched in Stojanovic's selection. After a careful reading of the literature, we found that the authors used a stringent negative condition. Four molecules were used for negative selection, including up to 200 µM serotonin, tyrosine, L-DOPA, and tryptophan. These negative selection targets were used in different rounds (Table S2). We hypothesized that due to the higher binding affinity of the Dopa-T aptamer, it might also have a higher affinity to the negative selection targets, and these were thus washed away during the negative selections.We first did a quick screening of the binding of the two aptamers to the four negative selection targets using ThT fluorescence spectroscopy and found that the most interfering molecule is serotonin, followed by L-DOPA, whereas tyrosine and tryptophan did not show much binding (Fig. S2). Thus, we used the strand displacement reaction to compare the responses of the two aptamers to serotonin. Both aptamers indeed showed serotonin-dependent fluorescence enhancement (Fig. 6A and B). We then plotted the binding curves (Fig. 6C). The Dopa-T aptamer has an apparent Kd of 13.3 µM, whereas the Dopa-A aptamer has an apparent Kd of 213 µM. Therefore, it is easy to understand that with 200 µM serotonin, Dopa-T was nearly fully removed, whereas about 50% Dopa-A still remained. Thus, we attributed the lack of Dopa-T in the previous work to the stringent negative selection.
The two aptamers have similar relative selectivity
Next, an important question is whether or not Dopa-A has better selectivity. Based on our strand-displacement sensors, the apparent Kd of Dopa-A with serotonin is 18-fold higher than that with dopamine, and this ratio is 21-fold for the Dopa-T aptamer. Given the similarity of the relative selectivity of the two aptamers, Dopa-T is more sensitive to both dopamine and serotonin, to a similar extent, compared to Dopa-A.We also assessed the selectivity of Dopa-T (Fig. 6D) and Dopa-A (Fig. 6E) to a series of structurally related analogs. Both aptamers are quite selective, and they only responded to serotonin and epinephrine. Therefore, Dopa-T might be a more sensitive substitution of the currently used Dopa-A.
The best negative selection condition
Aside from supplying a higher-affinity aptamer for dopamine, this work has also sparked some thoughts regarding negative selection during SELEX. Negative selection is carried out by incubating a library with potential interfering molecules and removing sequences that may bind to them. This process has so far been empirical, as it can start at any round with any negative target, concentration, incubation time or washing stringency. In this work, only one family of sequences was enriched (Dopa-A and Dopa-T are in the same family). This is a special case, since it seems that there is only one way for aptamers to tightly bind to dopamine, and negative selection may not make much of an improvement in boosting selectivity. Adding too high a concentration of the negative target eliminated only the tighter binders within that family.Negative selection has, of course, worked in many SELEX studies, and some beautiful examples include the selection of the aptamer specific to ATP. For the ATP–RNA aptamer selection, the negative target AMP and positive target ATP were both at 5 mM. The resulting aptamer had 5 µM Kd for ATP and 5.5 mM Kd for AMP.49 The Xiao group recently did an elegant selection to enrich DNA aptamers for ATP, and the final aptamers had 33-fold higher affinity for ATP than ADP.50 The highest ADP concentration was 250 µM (10.3-fold higher than the Kd of the best aptamer), whereas the ATP concentration was 50 µM for most rounds but decreased to 20 µM and even 5 µM in the final rounds. Aside from ADP, a few other counter targets, such as caffeine, were also used. A consequence of such stringent selection conditions is that the final pool remained highly diverse. The aptamer candidates were picked based on round-to-round abundance change as well as the final abundance. This is close to pushing the limits of negative selection.
In our theophylline selection, we used up to 10 mM caffeine as a counter target, with the theophylline concentration down to 50 µM.51 In this case, we could get away with extremely high caffeine concentrations with the expectation that the aptamer would not bind caffeine due to the known 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000-fold affinity difference.52 From the current work and the above examples, we reasoned that we need to have an estimation of the Kd of the best aptamer and the needed affinity difference between the target and counter targets. Overly stringent negative selections may also release good aptamers.
000-fold affinity difference.52 From the current work and the above examples, we reasoned that we need to have an estimation of the Kd of the best aptamer and the needed affinity difference between the target and counter targets. Overly stringent negative selections may also release good aptamers.
Conclusions
In this work, an aptamer selection was performed using dopamine as a target, repeating a previous selection experiment. Interestingly, our sequence and the previously published one differed only by a single nucleotide, yet the previous aptamer existed in more than a 2000-fold lower abundance. ITC and ThT fluorescence both indicated that our aptamer has approximately 6-fold higher binding affinity. In a strand-displacement system, it showed 19-fold higher affinity than the previous aptamer. We attributed the missed higher affinity aptamer in the previous selection to stringent negative selection with a too-high concentration of serotonin. This work has provided a high-affinity mutant that would be a more sensitive probe for dopamine. Given the very similar DNA sequence, most previously developed sensing methods can likely be directly adapted for this mutant.Author contributions
Yi Yu performed the aptamer selections, collected data, proposed key ideas, and drafted the initial manuscript. Yuanli Li carried out aptamer characterization and re-selection experiments. Juewen Liu contributed to conceptualization, data analysis, manuscript revision, and funding acquisition. All authors discussed the results and commented on the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data that support the findings of this study are available from the the Federated Research Data Repository (FRDR), at https://doi.org/10.20383/103.01626Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5sc09660j.
Acknowledgements
We thank the Stojanovic group for sharing their 2018 dopamine SELEX sequencing results. Dr Y. Ding for help with the second dopamine selection experiment and Y. Xie for drawing the scheme for the 3D dopamine aptamer NMR structure. Funding from this work is from the Natural Sciences and Engineering Research Council of Canada (NSERC), Canada Research Chair program, and Hubei Provincial Natural Science Foundation of China, (2025AFB765). Y. Yu received a China Scholarship Council scholarship to visit the University of Waterloo.References
- R. A. Wise, Nat. Rev. Neurosci., 2004, 5, 483–494 CrossRef CAS PubMed.
- C. Bucolo, G. M. Leggio, F. Drago and S. Salomone, Pharmacol. Ther., 2019, 203, 107392 CrossRef CAS PubMed.
- X. Liu and J. Liu, View, 2020, 2, 20200102 CrossRef.
- A. Gottås, Å. Ripel, F. Boix, V. Vindenes, J. Mørland and E. L. Øiestad, J. Pharmacol. Toxicol. Methods, 2015, 74, 75–79 Search PubMed.
- J. Yu, H. C. Lee, H. J. Yang, S. Hong and J. H. Bae, Analyst, 2025, 150, 3927–3934 RSC.
- S. Jabbari, B. Dabirmanesh, S. Daneshjou and K. Khajeh, Sci. Rep., 2024, 14, 14303 Search PubMed.
- I. Algov, A. Feiertag, R. Shikler and L. Alfonta, Biosens. Bioelectron., 2022, 210, 114264 CrossRef CAS PubMed.
- J. Njagi, M. M. Chernov, J. C. Leiter and S. Andreescu, Anal. Chem., 2010, 82, 989–996 CrossRef CAS PubMed.
- M. Geffard, O. Kah, B. Onteniente, P. Seguela, M. Le Moal and M. Delaage, J. Neurochem., 1984, 42, 1593–1599 Search PubMed.
- H. Yu, O. Alkhamis, J. Canoura, Y. Liu and Y. Xiao, Angew. Chem., Int. Ed., 2021, 60, 16800–16823 Search PubMed.
- A. Ruscito and M. C. DeRosa, Front. Chem., 2016, 4, 14 Search PubMed.
- D. Wu, C. K. L. Gordon, J. H. Shin, M. Eisenstein and H. T. Soh, Acc. Chem. Res., 2022, 55, 685–695 Search PubMed.
- S. Qian, D. Chang, S. He and Y. Li, Anal. Chim. Acta, 2022, 1196, 339511 Search PubMed.
- S. Stangherlin, N. Lui, J. H. Lee and J. Liu, TrAC, Trends Anal. Chem., 2025, 191, 118349 Search PubMed.
- A. Stuber and N. Nakatsuka, ACS Nano, 2024, 18, 2552–2563 CrossRef CAS PubMed.
- R. Walsh and M. C. DeRosa, Biochem. Biophys. Res. Commun., 2009, 388, 732–735 Search PubMed.
- A. A. Lubin and K. W. Plaxco, Acc. Chem. Res., 2010, 43, 496–505 CrossRef CAS PubMed.
- L. Wu, Y. Wang, X. Xu, Y. Liu, B. Lin, M. Zhang, J. Zhang, S. Wan, C. Yang and W. Tan, Chem. Rev., 2021, 121, 12035–12105 Search PubMed.
- N. Nakatsuka, K.-A. Yang, J. M. Abendroth, K. M. Cheung, X. Xu, H. Yang, C. Zhao, B. Zhu, Y. S. Rim, Y. Yang, P. S. Weiss, M. N. Stojanović and A. M. Andrews, Science, 2018, 362, 319–324 CrossRef CAS PubMed.
- J. T. Chen, F. Y. Xia, X. T. Ding and D. D. Zhang, Anal. Chem., 2024, 96, 10322–10331 Search PubMed.
- B. M. Wei, H. Y. Zhong, L. J. Wang, Y. Liu, Y. L. Xu, J. T. Zhang, C. Z. Xu, L. He and H. B. Wang, Int. J. Biol. Macromol., 2019, 135, 400–406 CrossRef CAS PubMed.
- Y. Y. Huang, P. W. Chen, L. Y. Zhou, J. Y. Zheng, H. T. Wu, J. X. Liang, A. X. Xiao, J. Li and B. O. Guan, Adv. Mater., 2023, 35, 2304116 CrossRef CAS PubMed.
- H. Abu-Ali, C. Ozkaya, F. Davis, N. Walch and A. Nabok, Chemosensors, 2020, 8, 28 CrossRef CAS.
- N. Nakatsuka, J. M. Abendroth, K. A. Yang and A. M. Andrews, ACS Appl. Mater. Interfaces, 2021, 13, 9425–9435 Search PubMed.
- M. A. H. Omer, M. Li, X. L. Yang, L. X. Liu and H. L. Qi, Anal. Chem., 2025, 97, 23580–23588 Search PubMed.
- A. Stuber, A. Douaki, J. Hengsteler, D. Buckingham, D. Momotenko, D. Garoli and N. Nakatsuka, ACS Nano, 2023, 17, 19168–19179 Search PubMed.
- Y. Hou, J. Hou and X. Liu, ChemBioChem, 2021, 22, 1948–1954 CrossRef CAS PubMed.
- Y. A. Kaiyum, E. H. P. Chao, L. Dhar, A. A. Shoara, M. D. Nguyen, C. D. Mackereth, P. Dauphin-Ducharme and P. E. Johnson, ChemBioChem, 2024, 25, e202400493 CrossRef CAS PubMed.
- X. Liu, Y. Hou, S. Chen and J. Liu, Biosens. Bioelectron., 2020, 173, 112798 CrossRef PubMed.
- K.-A. Yang, R. Pei and M. N. Stojanovic, Methods, 2016, 106, 58–65 Search PubMed.
- C. Lyu, I. M. Khan and Z. Wang, Talanta, 2021, 229, 122274 CrossRef PubMed.
- O. Alkhamis, J. Canoura, Y. Wu, N. A. Emmons, Y. Wang, K. M. Honeywell, K. W. Plaxco, T. E. Kippin and Y. Xiao, J. Am. Chem. Soc., 2024, 146, 3230–3240 CrossRef CAS PubMed.
- L. Zou, Y. Liu and J. Liu, Biosens. Bioelectron., 2025, 279, 117392 Search PubMed.
- Y. Ding and J. Liu, J. Am. Chem. Soc., 2023, 145, 7540–7547 CrossRef CAS PubMed.
- R. Nutiu and Y. Li, Angew. Chem., Int. Ed., 2005, 44, 1061–1065 CrossRef CAS PubMed.
- E. H. P. Chao, E. Largy, Y. A. Kaiyum, M.-D. Nguyen, B. Vialet, P. Dauphin-Ducharme, P. E. Johnson and C. D. Mackereth, bioRxiv, 2025, preprint, DOI:10.1101/2025.11.27.690961.
- S. Slavkovic and P. E. Johnson, Aptamers, 2018, 2, 45–51 Search PubMed.
- Y. Shu, S. Liu and J. Liu, Anal. Chem., 2025, 97, 19767–19774 CrossRef CAS PubMed.
- S. Stangherlin, Y. Ding and J. Liu, Small Methods, 2025, 9, 2401572 CrossRef CAS PubMed.
- R. Nutiu and Y. Li, J. Am. Chem. Soc., 2003, 125, 4771–4778 CrossRef CAS PubMed.
- J. Hu and C. J. Easley, Analyst, 2011, 136, 3461–3468 RSC.
- M. Zuker, Nucleic Acids Res., 2003, 31, 3406–3415 CrossRef CAS PubMed.
- Y. Ding, Y. Heng, K.-Y. Wong, Q. Chen and J. Liu, Angew. Chem., Int. Ed., 2026, 65, e14445 CrossRef CAS PubMed.
- O. Alkhamis and Y. Xiao, J. Am. Chem. Soc., 2023, 145, 194–206 CrossRef CAS PubMed.
- Y. Ding, Z. Zhang and J. Liu, ChemBioChem, 2024, 25, e202400570 CrossRef CAS PubMed.
- A. D. Ellington and J. W. Szostak, Nature, 1990, 346, 818–822 CrossRef CAS PubMed.
- F. Wachowius, B. T. Porebski, C. M. Johnson and P. Holliger, ChemSystemsChem, 2023, 5, e202300006 CrossRef CAS PubMed.
- R. Gysbers, K. Tram, J. Gu and Y. Li, Sci. Rep., 2015, 5, 11405 CrossRef PubMed.
- P. L. Sazani, R. Larralde and J. W. Szostak, J. Am. Chem. Soc., 2004, 126, 8370–8371 CrossRef CAS PubMed.
- A. Bacon, O. Alkhamis, C. Byrd, J. Perry, J. Canoura and Y. Xiao, Small, 2025, 21, e08898 CrossRef CAS PubMed.
- P.-J. J. Huang and J. Liu, ACS Chem. Biol., 2022, 17, 2121–2129 CrossRef CAS PubMed.
- R. D. Jenison, S. C. Gill, A. Pardi and B. Polisky, Science, 1994, 263, 1425–1429 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |