Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Conformational switching modulates excited-state pathways in a cofacial perylene dimer
Giovanni
Bressan
 *a,
Denis
Hartmann
*a,
Denis
Hartmann
 b,
Jonathan
Brouwer
a,
Erico M.
Braun
b,
Jonathan
Brouwer
a,
Erico M.
Braun
 ac,
James N.
Bull
ac,
James N.
Bull
 a and
Timothy A.
Barendt
a and
Timothy A.
Barendt
 *b
*b
aSchool of Chemistry, University of East Anglia, Norwich NR4 7TJ, UK. E-mail: g.bressan@uea.ac.uk
bSchool of Chemistry, University of Birmingham, Birmingham B15 2TT, UK. E-mail: t.a.barendt@bham.ac.uk
cInstituto de Fisica, Universidade Federal do Rio Grande do Sul, Av. Bento Gonçalves, Porto Alegre, 9500, Brazil
First published on 22nd January 2026
Abstract
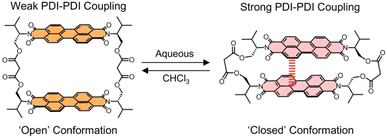
Controlling excited-state pathways in supramolecular chromophore assemblies is key to designing next-generation optoelectronic and photonic materials. Here we elucidate the conformation-dependent photophysics of a flexible perylene diimide (PDI) dimer, valPDI2, which undergoes reversible solvent-driven switching between two distinct dimer geometries, within the same structure. Exciton-coupling calculations and ultrafast spectroscopy show that in chloroform the dimer adopts an open, weakly coupled geometry that supports slow, partial excimer formation due to structural inhomogeneity within the excited state potential. In contrast, in polar DMSO/water the dimer collapses into a cofacial stacked conformer that enables barrierless, sub-200 femtosecond excimer formation, a subset of which forms a multiexciton state over tens of picoseconds. Half-broadband 2D electronic spectroscopy reveals conformation-dependent vibrational coherences, with nuclear wavepacket motion along the π-stacking coordinate promoting vibrationally coherent excimer formation in the closed conformer. These findings demonstrate that environmentally driven conformational control offers a powerful non-covalent strategy to modulate excimer and multiexciton dynamics in PDI assemblies. More broadly, they establish supramolecular switching as a general design principle for tuning excited-state behaviour in flexible organic chromophore arrays, with implications for the development of responsive optoelectronic and energy-conversion materials.
Introduction
Excited-state interactions in multichromophore assemblies lie at the heart of many optoelectronic processes, from energy transport to charge separation and luminescence. Among these interactions, excimer formation in π-stacked systems has attracted considerable interest since its first observation in concentrated pyrene solutions 70 years ago.1 Excimers (EX) represent adiabatic mixtures of singlet excitons 1(S1S0)-also known as local excited (LE)- and charge-transfer (CT) wavefunctions and can either act as traps or facilitate useful processes such as charge separation and multiexciton (ME) generation via singlet fission (SF).2–10 Supramolecular architectures based on the perylene diimide (PDI) building block provide an ideal platform for probing these phenomena, combining strong absorption in the green region of the visible spectrum, high photostability, and structural tunability via functionalisation at the imide or bay positions.11–14 Upon photoexcitation, rylene dimers access mixed electronic states with EX, CT, and correlated triplet-pair ME character, whose balance and time evolution are strong function of relative chromophore orientation, distance and environment and ultimately dictate the dimer's photophysics.4,7,15–23Therefore, understanding how geometry and environment jointly govern the mixed state behaviour is of central importance to the development of photoactive organic materials. Further, coherently excited nuclear wavepackets have been shown to drive sub-picosecond relaxation out of the Franck–Condon region, suggesting the possibility of harnessing vibrational coherence to control the fate of mixed excited states.4,16,17,19,24,25
However, mixed state character and dynamics are difficult to control within a single dimer structure. Typical tuning strategies rely on covalent modifications, which may alter the electronic structure, and require the synthesis of a series of dimer structures to systematically vary interchromophore distance and/or orientation. An alternative strategy leverages non-covalent stimuli, allowing a single dimer structure to explore multiple conformations and thus excitonic coupling regimes and excited state evolution in distinct dimer geometries. A notable example of such behaviour was recently reported by Thakur et al.,26 who demonstrated solvent polarity-controlled switching between open and folded conformations of a flexible PDI dimer, accompanied by drastic changes in charge separation dynamics. Despite their relevance to solid-state devices, the photophysics of chromophore dimers capable of reversibly switching between weak (or null) and strong excitonic couplings remain scarcely explored, motivating the present study.
Here, we investigate the conformation dependent photophysics and ultrafast (coherent) dynamics of valPDI2, a switchable and discrete PDI dimer recently reported by some of us, whose structure is shown in Scheme 1.27 This macrocycle incorporates a flexible L-valinol-malonate ester linker to connect two PDIs via their imide N positions which, in contrast to our previous perylene core-connected PDI dimer,24,28 affords valPDI2 with molecular recognition properties. This includes interactions with solvent molecules, generating distinct solvent-dependent conformations of valPDI2. In moderately polar CHCl3 (εr = 4.81), the discrete PDI dimer adopts an “open” geometry supporting weak excitonic coupling; in polar, protic 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 DMSO/water (εr = 66),29valPDI2 collapses into a cofacial, strongly coupled “closed” conformation due to PDI hydrophobicity. Such solvent-controlled switching enables reversible modulation of the excitonic coupling and associated excited-state dynamics without the need for alterations of the covalent structure.
1 DMSO/water (εr = 66),29valPDI2 collapses into a cofacial, strongly coupled “closed” conformation due to PDI hydrophobicity. Such solvent-controlled switching enables reversible modulation of the excitonic coupling and associated excited-state dynamics without the need for alterations of the covalent structure.
Our study combines fs transient absorption (TA) and half-broadband two-dimensional electronic spectroscopy (HB2DES), supported by quantum chemical modelling. Density functional theory (DFT) and extended tight-binding (XTB)30 calculations confirm the presence of solvent-dependent conformers, which populate EX and, in DMSO/water, ME states over markedly different timescales. In CHCl3, EX formation occurs over several ps following photoexcitation via LE (i.e. the Franck–Condon region of the S1 potential energy surface, PES) state branching. This arises from multiple dimer conformers in thermal equilibrium within the S0 PES. In contrast, in DMSO/water, the preorganised collapsed cofacial geometry enables near-barrierless and sub-200 fs EX formation, a non-negligible fraction of which evolves into a ME correlated triplet pair state over tens of ps, in parallel with charge separation and recombination repopulating S0. This indicates that solvent polarity, intermolecular H-bonding and conformational switching can accelerate EX formation by more than two orders of magnitude and gate ME generation and charge separation. Further, 2DES beatmap analysis suggests that the sub-200 fs excimer formation is driven by coherent nuclear motion along the PDI–PDI π–π coordinate, aiding population transfer from the LE to the EX regions of the excited state PES.
Together, our findings show that conformational switching, medium polarity, H-bond networks and nuclear wavepackets collectively modulate EX and ME state generation in flexible PDI dimers in solution, offering new design principles for organic materials with tuneable excited-state properties. Moreover, our results indicate that the supramolecular arrangement -and, consequently, the optical response- of PDI-based systems in solid-state devices could be controlled by the dielectric constant and the protic nature of the surrounding polymer matrix. This environmentally driven, reversible control mechanism is a novel tool for tuning excited-state behaviour via conformational regulation, rather than synthetic redesign.
Results and discussion
Steady-state spectroscopy, molecular structure and excitonic coupling
The normalised steady-state electronic absorption and emission spectra of valPDI2 in CHCl3 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 DMSO/water are reported in Fig. 1A and B. For comparison, the absorption and emission spectra of the reference monomer (refPDI, structure shown in SI, Fig. S1) in the same solvents are overlaid.
1 DMSO/water are reported in Fig. 1A and B. For comparison, the absorption and emission spectra of the reference monomer (refPDI, structure shown in SI, Fig. S1) in the same solvents are overlaid.
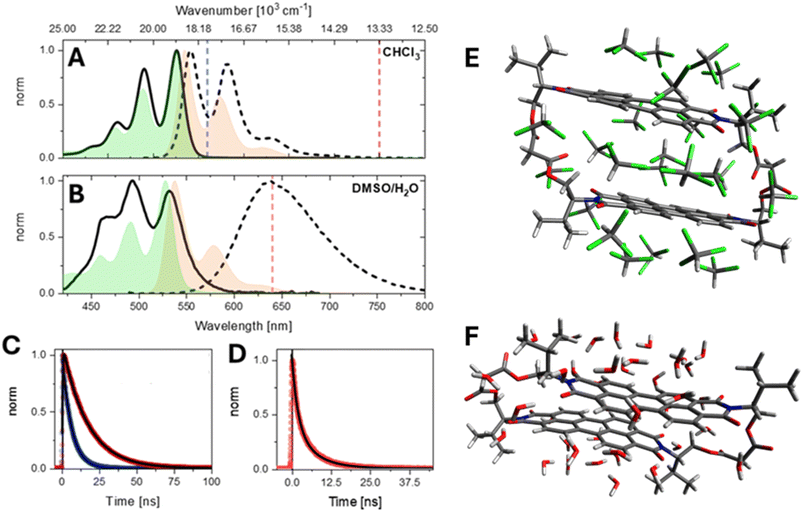 | ||
Fig. 1 Solvent-dependent spectroscopy and conformers of valPDI2. (A) Normalised steady-state absorption (solid) and emission (dashed) spectra of valPDI2 and the reference monomer, refPDI (absorption in green and emission in orange). Fluorescence spectra were excited at νExc = 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cm−1 (478 nm). (B) Same as (A) for 1 000 cm−1 (478 nm). (B) Same as (A) for 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 DMSO/water. (C) ns fluorescence emission traces of valPDI2 in CHCl3 measured at 18 1 DMSO/water. (C) ns fluorescence emission traces of valPDI2 in CHCl3 measured at 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cm−1 (blue) and 13 000 cm−1 (blue) and 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 cm−1 (dark red). Emission wavelength marked also with dash line in (A) and (B). (D) Same as (C) for valPDI2 in 1 500 cm−1 (dark red). Emission wavelength marked also with dash line in (A) and (B). (D) Same as (C) for valPDI2 in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 DMSO/water at 15 1 DMSO/water at 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 700 cm−1; single/multiexponential fits (fitted time constants given in text) are shown as black solid lines in (C) and (D). All traces were recorded after photoexcitation at 21 700 cm−1; single/multiexponential fits (fitted time constants given in text) are shown as black solid lines in (C) and (D). All traces were recorded after photoexcitation at 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cm−1. (E and F) Molecular structures of the valPDI2 dimer with the proposed “open” (0.7 nm PDI–PDI separation) and “closed” (0.35 nm PDI–PDI separation) macrocycle conformations in CHCl3 and DMSO/water. Both conformers have been obtained by optimisation of XRD structures27 with 25 explicit CHCl3 or 40 explicit H2O solvent molecules using XTB. XTB trajectories out to 200 ps at 300 K show that the cavity in the open geometry (E) supports five or six CHCl3 molecules. In the closed geometry shown in (F), the PDI units remain closely π-stacked with no solvent intercalation. 000 cm−1. (E and F) Molecular structures of the valPDI2 dimer with the proposed “open” (0.7 nm PDI–PDI separation) and “closed” (0.35 nm PDI–PDI separation) macrocycle conformations in CHCl3 and DMSO/water. Both conformers have been obtained by optimisation of XRD structures27 with 25 explicit CHCl3 or 40 explicit H2O solvent molecules using XTB. XTB trajectories out to 200 ps at 300 K show that the cavity in the open geometry (E) supports five or six CHCl3 molecules. In the closed geometry shown in (F), the PDI units remain closely π-stacked with no solvent intercalation. | ||
In CHCl3, valPDI2 (1A, solid) shows an absorption maximum at 526 nm (19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cm−1) with a well-resolved vibronic progression (≈1350 cm−1 spacing), attributed to Raman-active C
000 cm−1) with a well-resolved vibronic progression (≈1350 cm−1 spacing), attributed to Raman-active C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretches displaced along the S1 ← S0 transition, as typically encountered in polycyclic aromatic hydrocarbons (PAHs).14,31 This spectrum closely resembles that of the monomeric refPDI (green shade), though valPDI2 displays a more intense 0–1 vibronic peak at 492 nm (20
C stretches displaced along the S1 ← S0 transition, as typically encountered in polycyclic aromatic hydrocarbons (PAHs).14,31 This spectrum closely resembles that of the monomeric refPDI (green shade), though valPDI2 displays a more intense 0–1 vibronic peak at 492 nm (20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 360 cm−1) relative to the 0–0. Its fluorescence (1A, dashed; fluorescence quantum yield (FQY) = 0.91) upon 478 nm excitation has a quasi-mirror image relation to the absorption but is slightly broadened and redshifted by ≈300 cm−1 compared to refPDI. A weak emission tail extending to 13
360 cm−1) relative to the 0–0. Its fluorescence (1A, dashed; fluorescence quantum yield (FQY) = 0.91) upon 478 nm excitation has a quasi-mirror image relation to the absorption but is slightly broadened and redshifted by ≈300 cm−1 compared to refPDI. A weak emission tail extending to 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 cm−1 – absent in refPDI– is observed. Nanosecond time-resolved fluorescence (478 nm excitation, 1C) reveals wavelength-dependent lifetimes: 6 ns at 570 nm (blue, 17
300 cm−1 – absent in refPDI– is observed. Nanosecond time-resolved fluorescence (478 nm excitation, 1C) reveals wavelength-dependent lifetimes: 6 ns at 570 nm (blue, 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 540 cm−1) and 17 ns at 750 nm (dark red, 13
540 cm−1) and 17 ns at 750 nm (dark red, 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 330 cm−1), the latter has a ≈0.5 ns rise time. This suggests multiple emissive states or an excited-state equilibrium for valPDI2 in CHCl3. Conversely, in CHCl3refPDI exhibits a wavelength independent 5 ns lifetime (SI, Fig. S2).
330 cm−1), the latter has a ≈0.5 ns rise time. This suggests multiple emissive states or an excited-state equilibrium for valPDI2 in CHCl3. Conversely, in CHCl3refPDI exhibits a wavelength independent 5 ns lifetime (SI, Fig. S2).
In a polar 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 DMSO/water solvent mixture (1B, solid), the absorption of valPDI2 broadens and redshifts (onset at ca 600 nm), with an inversion of the vibronic peak intensities (0–0 at 533 nm, 0–1 at 494 nm). Similar behaviour was reported by Cadena et al. upon addition of increasing amounts of water to a DMF solution of a covalently tethered PDI oligomer.32 In DMSO/water, fluorescence (1B, dashed) is weaker (FQY = 0.065), redshifted (λmax = 640 nm), lacks vibronic structure, and exhibits a biexponential fluorescence decay (1.9 and 6.7 ns; weighted average = 3 ns, 1D) independent of emission wavelength. By comparison, refPDI in DMSO/water has a 1.5 ns fluorescence lifetime (SI, Fig. S2). This is expected, as water quenches photoexcited dyes via electronic-vibrational excitation energy transfer.33 The hypothesis of such quenching being due to PDI aggregation in polar media is ruled out by the strong similarity between the absorption spectra of refPDI in CHCl3 and DMSO/water. The ≈10-fold decrease in FQY of valPDI2 in DMSO/water (vs. in CHCl3) and their comparable lifetimes indicate a reduced emission transition dipole moment (TDM) in polar solvents. Weak, broad and redshifted emission is commonly observed for PDI excimers, whose role in mediating or hindering ME generation has been debated over the past decade.4,7,9,10,34 We also note that, despite ME states being dark, they can gain oscillator strength via Herzberg–Teller coupling to energetically close bright states (such as S1). Intensity borrowing leads to redshifted emission, strongly resembling EX features.34
1 DMSO/water solvent mixture (1B, solid), the absorption of valPDI2 broadens and redshifts (onset at ca 600 nm), with an inversion of the vibronic peak intensities (0–0 at 533 nm, 0–1 at 494 nm). Similar behaviour was reported by Cadena et al. upon addition of increasing amounts of water to a DMF solution of a covalently tethered PDI oligomer.32 In DMSO/water, fluorescence (1B, dashed) is weaker (FQY = 0.065), redshifted (λmax = 640 nm), lacks vibronic structure, and exhibits a biexponential fluorescence decay (1.9 and 6.7 ns; weighted average = 3 ns, 1D) independent of emission wavelength. By comparison, refPDI in DMSO/water has a 1.5 ns fluorescence lifetime (SI, Fig. S2). This is expected, as water quenches photoexcited dyes via electronic-vibrational excitation energy transfer.33 The hypothesis of such quenching being due to PDI aggregation in polar media is ruled out by the strong similarity between the absorption spectra of refPDI in CHCl3 and DMSO/water. The ≈10-fold decrease in FQY of valPDI2 in DMSO/water (vs. in CHCl3) and their comparable lifetimes indicate a reduced emission transition dipole moment (TDM) in polar solvents. Weak, broad and redshifted emission is commonly observed for PDI excimers, whose role in mediating or hindering ME generation has been debated over the past decade.4,7,9,10,34 We also note that, despite ME states being dark, they can gain oscillator strength via Herzberg–Teller coupling to energetically close bright states (such as S1). Intensity borrowing leads to redshifted emission, strongly resembling EX features.34
The pronounced solvent dependence of the valPDI2 photophysics can be attributed to solvent-induced conformational changes of the discrete macrocycle dimer structure. In CDCl3, 1H NMR spectroscopy indicates the presence of an open conformation where the flexible ligands expand to accommodate solvent between the PDI units.27 This assignment is supported by XTB calculations, in which the crystal structure of the host–guest valPDI2-coronene cocrystal27 (with the coronene guest removed) was optimised using the SOLVATOR method with 25 explicit CHCl3 solvent molecules, as implemented in ORCA 6.1.0,35 revealing solvation of the interchromophoric cavity by 5–6 CHCl3 molecules, resulting in an estimated PDI–PDI separation of approximately 0.7 nm (see Fig. 1E). For PDIs, chlorinated solvents preferentially stabilise solute–solvent interactions, hence inhibiting the π–π stacking solute–solute interactions that are typically responsible for the formation of dimers or extended aggregates.36
In DMSO/water, the valPDI2 macrocycle adopts a closed collapsed conformer geometry (shown in Fig. 1F), driven by intramolecular π–π stacking between the two PDIs since, in this medium, these interactions dominate over solute-(polar) solvent interactions. This conformer was characterised by 1D and 2D 1H NMR spectroscopy in solution.27 Importantly, the addition of increasing amounts of water to DMSO does not drive a continuous evolution of the valPDI2 geometry. Instead, isosbestic points in the absorption/circular dichroism solvent–solvent titrations27 indicate a two-state equilibrium, i.e. switching between “open” and “closed” conformations, rather than a gradual evolution through intermediate conformers with multiple equilibria.
Further support for this conformer structure was obtained by optimising the crystal structure27 of valPDI2 in SOLVATOR with 40 explicit water molecules using the XTB method. While the SOLVATOR framework requires neat water, this is an adequate approximation to the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 DMSO/water solvent used, since intramolecular π–π aggregation is observed for valPDI2 in both 1
1 DMSO/water solvent used, since intramolecular π–π aggregation is observed for valPDI2 in both 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 DMSO/water and water with only 0.1% DMSO. Significantly, the calculated structure does not show any water molecules intercalated between the PDI chromophores. This conformer displays a reduced PDI–PDI separation, ≈0.35 nm, accompanied by a longitudinal slip along the PDI long axis (i.e., aligned with the PDI S1 ← S0 TDM).
1 DMSO/water and water with only 0.1% DMSO. Significantly, the calculated structure does not show any water molecules intercalated between the PDI chromophores. This conformer displays a reduced PDI–PDI separation, ≈0.35 nm, accompanied by a longitudinal slip along the PDI long axis (i.e., aligned with the PDI S1 ← S0 TDM).
Excitonic coupling strength (JTot) was estimated using Spano's method,37,38 which relates the 0–1/0–0 vibronic peak area ratio R0–1/R0–0 to excitonic coupling magnitude and sign. In CHCl3, R0–1/R0–0 = 0.8 (vs. 0.68 for refPDI); in DMSO/water, R0–1/R0–0 = 1.13, indicating positive (H-type) excitonic interactions in both solvents. Peak area ratios were extracted by fitting absorption spectra to three Gaussian components (see SI, Fig. S3). For a vibronic spacing of 1350 cm−1, the determined R0–1/R0–0 values yield total excitonic coupling strengths JTot of +210 cm−1 in CHCl3 and +473 cm−1 in DMSO/water (details of the calculation are reported in the SI). These results indicate that valPDI2 behaves as a weakly and strongly coupled H-dimer in CHCl3 and in DMSO/water, respectively (Scheme 1), consistent with the proposed and calculated open and closed conformers populated in the two solvent media (Fig. 1E and F).
Following Spano's work,39 the total excitonic coupling JTot is dictated by the interference between long-range coulombic JCoul and short-range charge-transfer JCT terms:
| JTot = JCoul + JCT | (1) |
Whilst JCoul can be approximated using the point-dipole model,40 where the coupling scales as the inverse cube of the distance between TDMs, this model tends to overestimate coupling strengths at chromophore separations comparable to molecular dimensions. For a more reliable estimate, we used the transition electrostatic potential (trESP) method, as implemented in Multiwfn, with TDMs obtained from time-dependent (TD)-DFT (Gaussian 16.B01).24,41–43
The trESP method (details given in the SI) yielded JCoul values of +261 cm−1 and +462 cm−1 for the open and closed valPDI2 conformers, respectively, aligning well with Spano's fit results (+210 and +473 cm−1). The 25% overestimate for the open form likely arises from the omission of dielectric screening by CHCl3 solvent in the vacuum TD-DFT calculations. A possible additional contribution from a small negative i.e. “J-type” JCT term (≈−50 cm−1) was considered. This arises from a superexchange mechanism via a virtual CT state, and is defined as:24
 | (2) |
In contrast, the closed conformer (0.35 nm PDI–PDI distance) populated in DMSO/water may support non-negligible short-range coupling (JCT). th and te for the “closed” conformer are +182 and −112 cm−1, respectively, and were obtained using CATNIP.44 Details of the method are given in Bressan et al.24 and in the SI. Using eqn (2), these values yield a JCT value of +29 cm−1, an order of magnitude smaller than that determined for similar PDI structures with comparable interchromophore separation.45,46 The weak positive JCT contribution to JTot in the closed conformer of valPDI2 is attributed to the ≈0.38 nm longitudinal slip between PDIs disrupting frontier MO overlap, consistent with calculations showing that such longitudinal displacement makes the product teth negligible.13,38 A cofacial PDI dimer displaying large positive JCoul and small negative JCT has been reported by Hong et al.47 Therefore, even in the closed form, short-range coupling plays a marginal role in valPDI2, and changes in JTot are dominated by modulation of JCoul and solvent dielectric screening effects.
Overall, the combined evidence from NMR, XRD, DFT, XTB and excitonic coupling analysis supports solvent-controlled switching between weakly coupled (“open”, CHCl3) and strongly coupled (“closed”, DMSO/water) valPDI2 conformers. We next investigate how these structural differences affect excited-state dynamics probed first by femtosecond transient absorption (fsTA) and then by half-broadband 2D electronic spectroscopy (HB2DES).
Population dynamics
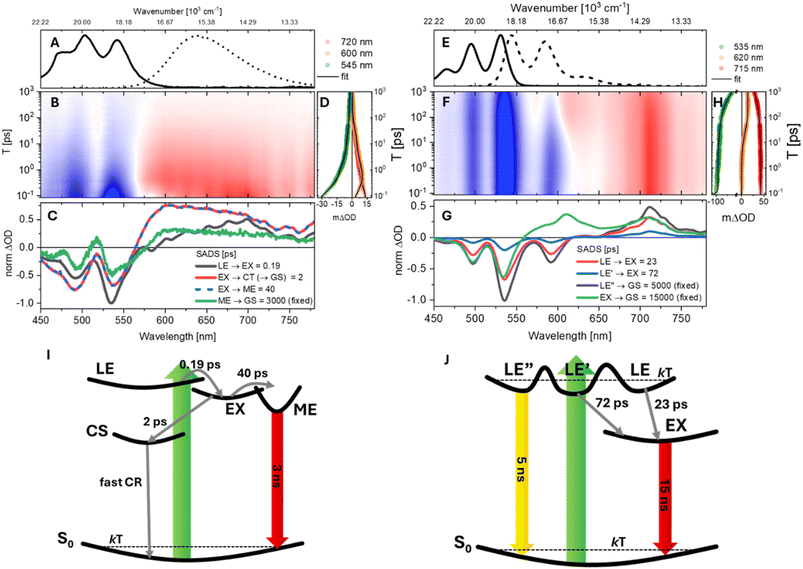
The solvent-dependent population dynamics of valPDI2 were investigated by fsTA spectroscopy. The data are presented on a log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) T scale contour from 0.08 to 1000 ps (Fig. 2B and F). Data for refPDI and transient spectra at selected T values for valPDI2 in both solvents are provided in the SI (Fig. S4 and S5). Steady-state absorption and emission spectra of valPDI2 in DMSO/water and CHCl3 are reproduced in Fig. 2A and E for reference.
T scale contour from 0.08 to 1000 ps (Fig. 2B and F). Data for refPDI and transient spectra at selected T values for valPDI2 in both solvents are provided in the SI (Fig. S4 and S5). Steady-state absorption and emission spectra of valPDI2 in DMSO/water and CHCl3 are reproduced in Fig. 2A and E for reference.
Following excitation in DMSO/water (Fig. 2B), valPDI2 shows an initial spectrum featuring a negative band between 450 and 590 nm due to ground-state bleach (GSB) and stimulated emission (SE), and a broad excited-state absorption (ESA) extending to the red edge of the probed spectral window. Within 1 ps, both the 535 nm GSB/SE and 700 nm ESA decay by ≈50% whilst a prominent ESA rises at 605 nm (Fig. 2D). Over tens of ps the ESA signal decays nonexponentially, as the GSB refills and a weak positive feature centred at 525 nm appears, which persists for longer than our time range (>1 ns, SI, Fig. S5).
In CHCl3 (Fig. 2F), valPDI2 exhibits a promptly formed broad negative feature attributed to overlapping structured GSB and SE. This band matches the inverted steady-state spectra. A positive ESA appears promptly at 700 nm (extending from 750 to 650 nm) and is nearly identical to that of refPDI (Fig. S4) and is thus assigned to an Sn ← S1 transition from the LE state. The spectra match similar data in the literature for related PDI molecules.20,48,49 The early time evolution of the open conformer of valPDI2 closely resembles that of refPDI under comparable conditions. However, over tens of ps, the SE (550–600 nm) partially recovers as a new ESA emerges between 600 and 650 nm (Fig. 2H). In contrast, refPDI shows nonexponential decay without any new ESA formation (SI, Fig. S4). After 100 ps valPDI2 exhibits ≈20% overall decay in signal amplitude, consistent with its ns fluorescence lifetime.
Such spectral evolution suggests branching within the excited state PES, not accounted for by a sequential model. Hence, we performed target analysis using GloTarAn50 (results of a global fit to a sequential model are reported in the SI). Four components were needed to yield an accurate fit to either dataset.
In DMSO/water, the components are τ1–4 = 0.19, 2, 40, and 3000 ps, whilst in CHCl3, τ1–4 = 23, 72, 5000 and 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ps were required. Traces of the fits (solid black) are superimposed to the experimental traces in Fig. 2D and H. The ns lifetimes were constrained to values obtained by time-resolved fluorescence emission (Fig. 1C and D). Species Associated Difference Spectra (SADS) are shown in Fig. 2C (DMSO/water) and G (CHCl3).
000 ps were required. Traces of the fits (solid black) are superimposed to the experimental traces in Fig. 2D and H. The ns lifetimes were constrained to values obtained by time-resolved fluorescence emission (Fig. 1C and D). Species Associated Difference Spectra (SADS) are shown in Fig. 2C (DMSO/water) and G (CHCl3).
In DMSO/water, the initial SADS (Fig. 2C, dark grey) exhibits a 0–0/0–1 intensity ratio that does not match the steady-state absorption. The negative band at 535 nm is stronger than the 0–1 GSB at 500 nm because it contains both GSB and monomer-like SE originating from the promptly populated LE state. Within 0.19 ps, this component forms a SADS characterised by ESA growth at 605 nm, decay at 700 nm, and quenching of SE at 535 nm. As PDI excimers display broad, nearly featureless ESA extending from ∼600 nm into the NIR,4,20,21,51–54 we assign this ultrafast evolution to LE → EX formation. The sub-ps timescale reflects the closed conformer geometry, which is preorganised for essentially barrierless population flow from LE to EX. Similar 100–200 fs EX formation has been reported for multiple cofacial PDI dimers by the Kim and Wasielewski groups, and by us for a bay-connected PDI dimer.4,24,55,56 EX formation in π-stacked PDI systems is governed by subtle geometric factors rather than by excitonic coupling strength alone. For example, in PDI nanowires57 and crystals58 a significant rotational displacement between chromophores can, in line with theory,59 suppress population of the EX minimum. In contrast, the closed conformer of valPDI2 combines a short π–π separation with negligible rotational (and transverse) displacements, features that enhance frontier MO overlap and promote structural relaxation along the π-stacking coordinate on the excited-state potential energy surface (vide infra). These structural characteristics promote ultrafast (vibrationally coherent) excimer formation in the “closed” conformer of valPDI2.
The EX state (red SADS) loses a substantial fraction of its initial ESA amplitude, accompanied by significant GSB recovery, with a 2 ps timescale. We attribute this rapid relaxation to symmetry-breaking charge separation (SBCS) i.e. formation of a PDI˙+/˙− pair. SBCS is typically driven by dipolar solvation and should therefore occur on solvent-relaxation timescales. While bulk water solvation is complete within ∼2 ps, solvation in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 DMSO/water mixture is slower, extending over several ps, due to extended H-bond networks, as reported by Roy et al.60 We propose that the observed GSB refilling arises from SBCS followed by faster charge recombination (CS → GS). Because CR is faster than SBCS, the characteristic 615/710 nm PDI˙+/˙− transients are not detected.49,61
1 DMSO/water mixture is slower, extending over several ps, due to extended H-bond networks, as reported by Roy et al.60 We propose that the observed GSB refilling arises from SBCS followed by faster charge recombination (CS → GS). Because CR is faster than SBCS, the characteristic 615/710 nm PDI˙+/˙− transients are not detected.49,61
The thermodynamic feasibility of photoinduced charge separation was evaluated using the Rehm–Weller formalism, which estimates the free energy change associated with electron transfer following photoexcitation ΔGCS, defined as:
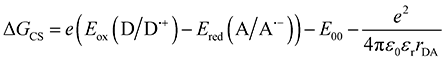 | (3) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 DMSO/water solvent (εr), the centre-to-centre separation between the donor and acceptor moieties (rDA) i.e. the PDI–PDI distance.
1 DMSO/water solvent (εr), the centre-to-centre separation between the donor and acceptor moieties (rDA) i.e. the PDI–PDI distance.
For PDI, the one-electron half-wave oxidation and reduction potentials versus SCE have been reported by Lee et al.62 as +1.56 V and −0.55 V, respectively. Using these values, together with the experimentally determined E00 (2.36 eV), εr = 66, and rDA = 0.35 nm (from XRD and XTB calculations, see above) yield a ΔGCS = −0.282 eV, indicating thermodynamically favourable SBCS. The free energy of charge recombination can then be expressed as:
| ΔGCR = −(ΔGCS + E00) | (4) |
which gives ΔGCR ≈ −2.05 eV. The strongly exergonic nature of this process provides a rationale for the fast CR following SBCS, preventing direct observation of the spectral features of the PDI˙+/˙− pair.
A parallel, slower (40 ps) EX decay channel forms a SADS (green) whose distinctive lineshape in the 450–550 nm region is nm due to overlap of negative singlet GSB and positive Tn ← T1 absorption at ≈525 nm.15,63–67 This component develops far more rapidly than typical ns PDI intersystem crossing and is therefore assigned to singlet fission (SF). In this process, a non-negligible fraction of the EX populates a multiexciton 1(T1T1) state. As PDI has E(S1) = 2.36 eV and E(ME) ≈ 2 × E(T1) = 2.4 eV, ME formation is slightly endothermic, implying that thermal activation contributes to its relatively slow onset.68 Contributions from CT states close in energy to S1 can lower the 1(T1T1) energy through superexchange, enabling SF even when the ME state would be otherwise inaccessible.4 We propose that the highly polar DMSO/water environment (εr ≈ 66) increases the CT character of the EX state, as suggested by the fast CS channel, enhancing S1–ME mixing, thereby promoting SF in the closed conformer of valPDI2. EX-mediated ME formation over comparable timescales was reported for a slip-stacked terrylene diimide dimer by Margulies et al.,69 and a CT-enabled SF mechanism in pentacene dimers in polar benzonitrile was recently proposed by Thiel et al.5
The ME of valPDI2 persists for a few ns and, through vibronic intensity borrowing, produces the weak, red-shifted, and featureless fluorescence described earlier. Such long-lived emission indicates that the ME state does not dissociate, preventing quantification of the free triplet yield. In any case, such yield is expected to be modest, as a large fraction of the EX relaxes back to S0via CS followed by CR within less than 2 ps from photoexcitation. Notably, the FQY of the ME emission shows only a minor change beyond 50% water content (SI, Fig. S11) indicating that its formation and relaxation are primarily governed by solvent-induced conformational changes, with further increases in the dielectric constant of the environment having little influence on its behaviour.
We propose that the bifurcation within the EX PES originates from structural heterogeneity associated with the malonate ester linkers. The oxygen lone pairs can participate in intermolecular H-bonding with water, affecting both electrostatics and dimer geometry. Small variations in the PDI–PDI rotational displacement strongly affecting CT coupling, and in the long-axis slip coordinate, critical for SF,37,39,70,71 lead to a distribution of EX conformations for which the former or latter relaxation pathways are preferred. The photoinduced dynamics of valPDI2 in DMSO/water are summarised in Fig. 2I.
In CHCl3, both the excited-state timescales and spectral evolution differ markedly from those observed in the polar, protic medium. Upon photoexcitation, multiple spectroscopically indistinguishable minima of the LE PES are populated. These correspond to conformers arising from the flexible architecture of valPDI2. Such conformers are in thermal equilibrium in S0 but separated by barriers larger than kT in the excited state and are labelled in Fig. 2G as LE (red), LE′ (light blue), and LE″ (dark grey). From these minima, the population decays either directly to the ground state or to a long-lived component (green), assigned to the EX state based on its relatively flat ESA spanning 560–700 nm coupled to SE quench. We note that the long-lived SADS presents a peak at 615 nm which is not present in the data (see SI) and presumably arises from the fit compensating positive ESA and negative structured SE.
Although the PDI˙+ D1 ← D0 absorption also lies near 610 nm, the formation of a new ESA in this region cannot be attributed to SBCS: this mechanism would require a corresponding risetime at ∼710 nm, where the D1 ← D0 absorption of PDI˙− contributes, but no such feature is observed.49,61 Instead, the strong positive signal at 700 nm, present from the earliest delay times and also observed for refPDI (SI), is assigned to the Sn ← S1 transition from the LE region of photoexcited valPDI2. The absence of any redshift or spectral reshaping of this band (see comparison with the absorption spectrum of chemically reduced refPDI in Fig. S5) confirms that PDI˙− is not formed. The data therefore support EX formation rather than SBCS. Further, EX formation aligns with the weak, long-lived 750 nm emission, which a dark CS state fails to account for.
The fastest component (red, Fig. 2G) corresponds to a subset of the LE population that converts to EX with a 23 ps timescale. Over ∼72 ps, a smaller LE′ subpopulation (blue) also relaxes to EX. A substantial fraction of the initially excited population (dark grey) resides in LE″ and does not reach the EX region; instead, it decays radiatively back to S0 over a few ns, consistent with the LE lifetime of refPDI in CHCl3. The EX state persists for ∼15 ns, in agreement with the long-lived weak fluorescence observed at 750 nm for valPDI2 in CHCl3 (Fig. 1C). This suggests that over several ps, the LE and LE’ macrocycle subpopulations adopt a “closed” conformation expelling the intercalated CHCl3 solvent molecules.
Because SADS1-3 are spectrally identical, their relative amplitudes directly reflect the relative populations of the corresponding LE subsets, i.e. conformers. From this, we conclude that a branching mechanism governed by conformational heterogeneity causes more than 60% of photoexcited valPDI2 (LE″, dark grey) to relax radiatively to S0, producing monomer-like structured fluorescence and SE, whereas the remaining fraction (red and blue SADS) undergoes LE/LE′ → EX conversion on tens of ps timescales. The full photoinduced dynamics for valPDI2 in CHCl3 are summarised in Fig. 2J.
Overall, the fsTA measurements show that the excited state dynamics of valPDI2 proceed through conformation-dependent branching pathways. In DMSO/water, the collapsed conformer enables rapid, sub-ps EX formation, a fraction of which forms a ME state, whist most of the population quickly refills the ground state via CS followed by (faster) CR. In CHCl3 the open conformers undergo slower structural dynamics, leading to a mixture of monomer-like and EX excited state populations. The sub-ps EX formation observed in the collapsed conformer suggests that vibrationally coherent dynamics may assist the system in escaping the LE region of the excited-state PES. We now investigate with HB2DES whether coherent nuclear-wavepacket motion influences the ultrafast photophysics of valPDI2.
Coherent vibrational dynamics
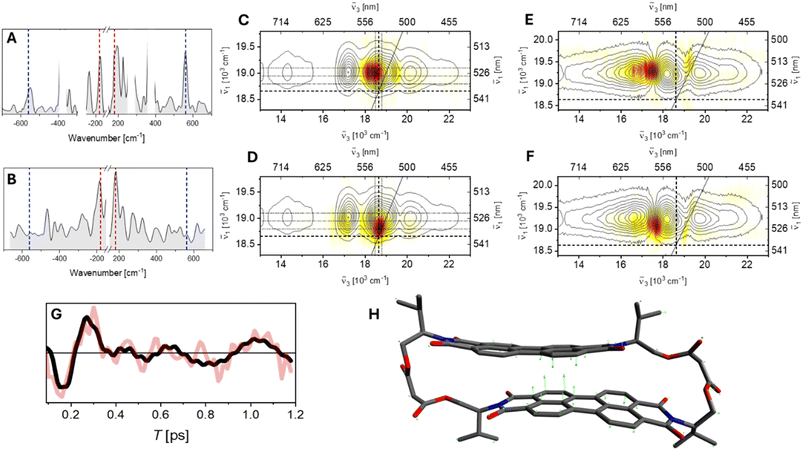
We employed HB2DES to investigate the potential role of S1 vibrational coherence in the excited state dynamics of valPDI2. The waiting time T (analogous to the pump-probe delay time in fsTA) was scanned in 10 fs steps between 0 and 1200 fs to capture ground and excited state wavepacket motion, detected as amplitude oscillations during T superimposed to the population dynamics. From a Fourier transform (FT) of these data we construct a 3D spectrum, which is then “sliced” at frequencies of specific resonance Raman active molecular vibrations to create their “beatmaps”. Beatmaps are thus a full frequency domain representation of nuclear wavepacket dynamics in which the contribution due to a specific molecular vibration is resolved as a function of the excitation (pump, ν1) and detection (probe, ν3) dimensions and with respect to its beat frequency and phase over T. Details of the HB2DES beatmap construction are presented in the SI (Fig. S7).Our experimental implementation of HB2DES enables the retrieval of both the real and imaginary components of the rephasing (photon-echo) and non-rephasing (free induction decay) signals. Acquiring the full complex-valued response is necessary for the unambiguous assignment of vibrational coherence to either the electronic ground (S0) or excited (LE, S1) state PESs, which is based on the sign (i.e. the phase) of the oscillatory response following complex FT. The discussion below is restricted to the rephasing data, as the nonrephasing response does not here contain additional information. For rephasing signals, positive coherence frequencies correspond to density matrices |sn〉〈sn+1| evolving during T, (where s is an arbitrary electronic state and n ≥ 0 is the number of vibrational quanta of a resonance Raman active vibrational mode) and encode contributions from wavepackets evolving in S0 and S1, whereas negative frequencies, corresponding to |sn+1〉〈sn| density matrices exclusively arise from vibrational wavepacket evolution on the excited state potential energy surface. It should be emphasised that this correlation is rigorously valid only for vibrational mode energies significantly exceeding kT, as discussed below. The selection of specific vibrational wavenumbers for further analysis in beatmaps is guided by the inspection of the rephasing vibrational spectra integrated over the excitation (ν1) and detection (ν3) frequency axes (“summary Raman”) shown in Fig. 3A and B for valPDI2 in CHCl3 and DMSO/water respectively.72,73
The summary Raman spectrum of valPDI2 in CHCl3 (Fig. 3A) reveals a few resonantly enhanced vibrational modes between 100–600 cm−1, along with nonresonant CHCl3 solvent contributions at 261 and 369 cm−1 (masked in white). In PDI dimers, modes in the 100–400 cm−1 range correspond to delocalised PDI and scaffold deformations. DFT calculations and the absence of signals in this spectral region for refPDI support assignment of the strong ±180 cm−1 signals (red dashes) to an out-of-plane (OOP) PDI bending mode (Fig. 3H). The 300–600 cm−1 region corresponds to PDI ring breathing modes, with the ±550 cm−1 peaks (blue dashes) attributed to a strongly Raman allowed ring breathing vibration, confirmed via DFT, literature, and comparison with refPDI.14 Both 180 and 550 cm−1 modes contribute to the negative side of the Raman spectrum, indicating S1 activity.73,74 However, their presence on the positive side suggests ground state wavepackets also contribute. To confirm the frequencies observed in the summary Raman of valPDI2 in CHCl3, we fitted the time-domain oscillations of rephasing HB2DES residuals (integrated over a 300 cm−1 square centred on the GSB + SE feature maximum) to a sum of damped cosines (Fig. S8). The 550 cm−1 and 180 cm−1 components are damped with a ≈650 fs rate, a timescale typical of vibrational dephasing for chromophores in solution.75 The strong CHCl3 solvent mode at 359 cm−1, with dephasing time fixed to 1 ps, was included to yield a good fit.
The amplitude ratio of rephasing positive to negative peaks reflects the total number of allowed Liouville-space pathways containing oscillatory terms of the type |sn〉〈sn+1| vs. |sn+1〉〈sn|.73,76,77 For the 550 cm−1 mode, this ratio is ≈2, consistent with the displaced harmonic oscillator (DHO) model. Beatmaps relative to this mode can be fully assigned following the DHO model and are reported and discussed in the SI (Fig. S8). As a reference, the rephasing ±550 cm−1 beatmaps of refPDI are also reported in Fig. S10. In contrast, the 180 cm−1 mode shows a ≈1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ± amplitude ratio, diverging from DHO expectations. We address this point below.
1 ± amplitude ratio, diverging from DHO expectations. We address this point below.
Rephasing positive and negative beatmaps of the 180 cm−1 mode of valPDI2 in CHCl3 (Fig. 3C and D) show that the positive features are blue-shifted along ν1 by (at least) one quantum of vibrational excitation relative to their negative counterpart, located at the 0–0 transition. Comparison with HB2DES absorptive data (black contours, reproduced from Fig. S6) shows coherence amplitude contributions, along ν3, between 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 and 18
800 and 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 cm−1 consistent with GSB (+180 cm−1) and SE (±180 cm−1) pathways. Absence of amplitude at ν3 = 14
900 cm−1 consistent with GSB (+180 cm−1) and SE (±180 cm−1) pathways. Absence of amplitude at ν3 = 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 cm−1 (Sn ← S1 LE ESA) implies minimal displacement along this mode between S1 and higher excited states. Hence, the coherent vibrational response of the 180 cm−1 mode of valPDI2 in CHCl3 can be fully rationalised by a two-level DHO model including the S0 and LE PESs.
300 cm−1 (Sn ← S1 LE ESA) implies minimal displacement along this mode between S1 and higher excited states. Hence, the coherent vibrational response of the 180 cm−1 mode of valPDI2 in CHCl3 can be fully rationalised by a two-level DHO model including the S0 and LE PESs.
For such a model system, GSB and SE coherence amplitudes are both governed by the displacements between the S0 and S1 (LE) PESs. Hence, it is unphysical for a vibrational mode to be S0 silent while being active in S1. Nonetheless, the nearly 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 amplitude ratio observed in the summary Raman spectrum in Fig. 3A suggests S1 only activity. This apparent inconsistency can be explained by thermal population of vibrational levels within kT (≤200 cm−1) at room temperature. Assuming that thermal population is significant only for v = 1, four additional ‘anti-Stokes-like’ vibrationally coherent rephasing GSB pathways must be considered, originating from thermally populated |g1〉〈g1| and beating at negative frequency during T, as shown by Green et al.77 The corresponding double-sided Feynman diagrams (DSFDs) are provided in the SI (Fig. S9). We refer to such pathways as hot ground state bleach (HGSB) pathways. These shift spectral weight from positive to negative peaks of the integrated vibrational spectrum according to a Maxwell–Boltzmann distribution, flattening the amplitude ratio of the low frequency region of the summary Raman spectrum. HGSB contributions also broaden beatmap features due to increased numbers of possible quasi-degenerate DSFDs involving multiple thermally populated states excited by the pump, as we have previously reported.24,78
1 amplitude ratio observed in the summary Raman spectrum in Fig. 3A suggests S1 only activity. This apparent inconsistency can be explained by thermal population of vibrational levels within kT (≤200 cm−1) at room temperature. Assuming that thermal population is significant only for v = 1, four additional ‘anti-Stokes-like’ vibrationally coherent rephasing GSB pathways must be considered, originating from thermally populated |g1〉〈g1| and beating at negative frequency during T, as shown by Green et al.77 The corresponding double-sided Feynman diagrams (DSFDs) are provided in the SI (Fig. S9). We refer to such pathways as hot ground state bleach (HGSB) pathways. These shift spectral weight from positive to negative peaks of the integrated vibrational spectrum according to a Maxwell–Boltzmann distribution, flattening the amplitude ratio of the low frequency region of the summary Raman spectrum. HGSB contributions also broaden beatmap features due to increased numbers of possible quasi-degenerate DSFDs involving multiple thermally populated states excited by the pump, as we have previously reported.24,78
Thus, the vibrational response of valPDI2 in CHCl3 is fully explained by a two-state DHO model incorporating HGSB pathways for the 180 cm−1 mode. These results do not therefore suggest involvement of coherently excited nuclear motion in S1 in the formation of the valPDI2 excimer. This is expected, as the main LE → EX timescale (23 ps, Fig. 2G) is ≈40 times slower than the 550 cm−1 vibrational dephasing time (650 fs) retrieved from the damped cosine fit.
In DMSO/water, valPDI2 exhibits different vibrational features to the dimer or monomer (refPDI, Fig. S10) in CHCl3. Its summary Raman spectrum (Fig. 3B) retains the ±180 cm−1 OOP bending mode observed in CHCl3 but lacks the ±550 cm−1 ring breathing mode. These differences likely arise from the “open” and “closed” conformers having different displacements along these normal mode coordinates and differing electronic transitions, consistent with the observed differences between their electronic spectra, affecting resonance enhancement.79,80 Coherent vibrational responses dominated by low frequency vibrations were previously reported, among others, by Kim et al. and O'Connor et al. for EX forming cofacial PDI dimers16,19 and by us in a more rigid macrocyclic PDI dimer.24
The ±180 cm−1 beatmaps in DMSO/water (Fig. 3E and F) resemble those in CHCl3 along ν1, but differ along ν3: amplitude is localised near the 0-crossing between GSB and ESA and extends toward the EX ESA at 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 cm−1. Beatmap amplitude localised in such region suggests contributions from ESA pathways i.e. vibrational coherence in the S1 PES. The lack of GSB features suggests minimal S0–S1 displacement along this coordinate for the strongly coupled valPDI2 conformer. This implies the S0–S1 displacement is insufficient to generate appreciable GSB coherence amplitude (which scales with the 4th power of the S0–S1 displacement along this vibrational coordinate) but large enough to launch S1 wavepacket motion modulating the ESA band, dependent on the product of squared displacements between S0–S1 and S1–Sn.73,78 This suggests that the 180 cm−1 mode is significantly displaced along S1–Sn for the “closed” conformer but only weakly along S0–S1, in contrast to the “open” conformer.
300 cm−1. Beatmap amplitude localised in such region suggests contributions from ESA pathways i.e. vibrational coherence in the S1 PES. The lack of GSB features suggests minimal S0–S1 displacement along this coordinate for the strongly coupled valPDI2 conformer. This implies the S0–S1 displacement is insufficient to generate appreciable GSB coherence amplitude (which scales with the 4th power of the S0–S1 displacement along this vibrational coordinate) but large enough to launch S1 wavepacket motion modulating the ESA band, dependent on the product of squared displacements between S0–S1 and S1–Sn.73,78 This suggests that the 180 cm−1 mode is significantly displaced along S1–Sn for the “closed” conformer but only weakly along S0–S1, in contrast to the “open” conformer.
The OOP PDI core bending coordinate corresponds to a contraction of the interchromophore spacing and hence promotes ultrafast, vibrationally coherent, EX formation, revealed by the dephasing time of such vibration matching the EX formation timescale.16,24,81 Therefore, we analysed time-domain oscillations of rephasing HB2DES residuals (integrated over a 300 cm−1 side square centred at 19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 250, 16
250, 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 950 cm−1) using damped cosine fits, as shown in Fig. 3G. The 180 cm−1 component is damped with a 180 fs dephasing time, a timescale closely matching the 190 fs EX formation. The ps-lived modes at 90 and 670 (DMSO solvent) cm−1 were needed to yield a good fit. The similar vibrational dephasing and EX formation timescales suggest a role for the 180 cm−1 OOP bending mode in enabling efficient LE-to-EX population transfer promoted via reduction of the PDI–PDI interchromophoric coordinate, as previously shown for structurally related PDI dimers.16,19,24,25,56
950 cm−1) using damped cosine fits, as shown in Fig. 3G. The 180 cm−1 component is damped with a 180 fs dephasing time, a timescale closely matching the 190 fs EX formation. The ps-lived modes at 90 and 670 (DMSO solvent) cm−1 were needed to yield a good fit. The similar vibrational dephasing and EX formation timescales suggest a role for the 180 cm−1 OOP bending mode in enabling efficient LE-to-EX population transfer promoted via reduction of the PDI–PDI interchromophoric coordinate, as previously shown for structurally related PDI dimers.16,19,24,25,56
Conclusions
Ultrafast coherent spectroscopies reveal that the photophysics of a macrocyclic perylene diimide dimer, valPDI2, are dictated by a pronounced solvent-induced conformational switch that modulates both excitonic coupling and excited-state relaxation. In weakly polar CHCl3, valPDI2 adopts an “open” macrocycle geometry with a large PDI–PDI separation, yielding weak H-type coupling and steady-state spectra closely resembling those of monomeric refPDI. Under these conditions, photoexcitation largely populates a monomer-like local excited state, while conformational heterogeneity enables a fraction of the local excited state population to form an excimer over several tens of ps. Half-broadband two-dimensional electronic spectroscopy further shows that vibrational coherences in the “open” conformer follow a simple displaced-harmonic-oscillator description of the ground and excited state potential energy surfaces, with no evidence for coherent nuclear motion funnelling population toward the excimer region of the excited state, in agreement with its slow formation.In sharp contrast, in polar DMSO/water the dimer collapses into a tightly π-stacked cofacial conformer characterized by strong H-type coupling and broadened, red-shifted emission. This “closed” geometry is preorganised to support sub-200 fs barrierless excimer formation, after which the excimer partitions between rapid charge separation and recombination and slower multiexciton generation via excimer-mediated singlet fission. The weakly emissive multiexciton state persists on the nanosecond timescale without dissociating into free triplets. Half-broadband two-dimensional electronic spectroscopy reveals that, unlike in CHCl3, the dominant coherence corresponds to an out-of-plane 180 cm−1 PDI bending mode with a dephasing time (∼180 fs) in close agreement with the excimer formation timescale. Localisation of the coherence amplitude within the ESA region further indicates wavepacket motion in the excited state potential along a nuclear coordinate that contracts the PDI–PDI separation, thereby supporting vibrationally coherent LE → EX population transfer.
Importantly, our results demonstrate that in the strongly coupled structure, dimer conformation promotes excimer formation and solvent polarity enhances its charge transfer character, opening a channel for multiexcition state generation. However, the stabilisation of the radical pair state also accelerates relaxation to the ground state via ultrafast charge separation and recombination, creating a competing decay pathway. Optimising multiexciton generation, and therefore triplet yield, requires striking a balance: polarity-driven conformational tuning must be sufficient to promote short-range coupling and thus charge transfer-assisted excimer-to-multiexciton conversion, but not so strong as to favour fast charge separation/recombination pathways that short-circuit multiexciton formation yield.
Experimental
Synthesis
The macrocycle dimer valPDI2 and the monomer refPDI were prepared and characterised as previously described.27fsTA and HB2DES
Ultrafast spectroscopy data were acquired using a previously reported home-built actively referenced fsTA/HB2DES spectrometer72,82 in 1 mm optical path static fused silica cells (Starna), with the concentration of valPDI2 adjusted to a peak OD of 0.5 in CHCl3 and 0.3 in DMSO/water. Sample and solvents were used as received. Briefly, 500 µJ pulses from the output of a Ti:Sa regenerative amplifier (Spitfire Ace, Spectra-Physics) operating at 800 nm and 1 kHz repetition rate pump a commercial noncollinear optical parametric amplifier (NOPA, Topas White, Light Conversion) to generate pulses centred at 520 nm (19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 cm−1). The broadband visible pulse is pre-compressed by a commercial folded grism compressor (Fastlite) to achieve close to FT limited pulses at the sample position. Downstream, a commercial acousto-optical programmable dispersive filter (AOPDF, Dazzler, Fastlite) creates a pair of pump pulses with programmable time delay (coherence time τ) and relative carrier-envelope phase. τ is scanned shot-to-shot from −95 to 0 fs in 0.792 fs steps. Real and imaginary parts of the rephasing and nonrephasing responses are obtained by a 3 × 1 phase-cycling method83 and summed to obtain absorptive 2D spectra. Each 2D spectrum is averaged over 270 laser shots per τ value. The waiting time delay T between the second pump and the probe is introduced with a retroreflector mounted on a mechanical linear translation stage (Physik Instrumente). Data are measured with 10 fs steps from 0 to 1200 fs to record vibrationally coherent dynamics (HB2DES) or at increasing T steps between −1500 fs and 1 ns to recover population dynamics (fsTA). Pump scatter and fluorescence contributions are removed by subtracting a 2DES spectrum acquired at T < 0 fs. The probe pulse (WLC) is generated by focusing ≈1 µJ of the 800 nm regenerative amplifier output into a 3 mm static sapphire plate and spans 13
200 cm−1). The broadband visible pulse is pre-compressed by a commercial folded grism compressor (Fastlite) to achieve close to FT limited pulses at the sample position. Downstream, a commercial acousto-optical programmable dispersive filter (AOPDF, Dazzler, Fastlite) creates a pair of pump pulses with programmable time delay (coherence time τ) and relative carrier-envelope phase. τ is scanned shot-to-shot from −95 to 0 fs in 0.792 fs steps. Real and imaginary parts of the rephasing and nonrephasing responses are obtained by a 3 × 1 phase-cycling method83 and summed to obtain absorptive 2D spectra. Each 2D spectrum is averaged over 270 laser shots per τ value. The waiting time delay T between the second pump and the probe is introduced with a retroreflector mounted on a mechanical linear translation stage (Physik Instrumente). Data are measured with 10 fs steps from 0 to 1200 fs to record vibrationally coherent dynamics (HB2DES) or at increasing T steps between −1500 fs and 1 ns to recover population dynamics (fsTA). Pump scatter and fluorescence contributions are removed by subtracting a 2DES spectrum acquired at T < 0 fs. The probe pulse (WLC) is generated by focusing ≈1 µJ of the 800 nm regenerative amplifier output into a 3 mm static sapphire plate and spans 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500–23 000 cm−1. Two pairs of dispersive mirrors at 5° and 19° angles of incidence (PC 1332, Ultrafast Innovations) recompress the WLC which is split before the sample by a 50
500–23 000 cm−1. Two pairs of dispersive mirrors at 5° and 19° angles of incidence (PC 1332, Ultrafast Innovations) recompress the WLC which is split before the sample by a 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 beamsplitter to provide probe and reference. The probe is crossed at 4° with the collinear pump pair at sample position. Pump and probe spot sizes are 80 and 160 µm, respectively. Pump pulse energy and duration at sample position are ≈300 nJ and 30 fs, respectively. The signal is recollimated after the sample and the signal and reference are focused into a home-built dual channel prism-based spectrometer and recorded shot-to-shot by a pair of 1024 pixels CCD detectors (Stresing) synchronised to the AOPDF. The signal is referenced using an active noise reduction method proposed by Feng et al.84 The instrument response function (ca. 45 fs) is measured by spectrally resolving the instantaneous nonresonant solvent response. All measurements were conducted at the magic angle to rule out contributions from rotational diffusive motion to the measured signal decay. HB2DES beatmaps are obtained by a procedure previously discussed in the literature and outlined in the SI (Fig. S6).72,73,76
50 beamsplitter to provide probe and reference. The probe is crossed at 4° with the collinear pump pair at sample position. Pump and probe spot sizes are 80 and 160 µm, respectively. Pump pulse energy and duration at sample position are ≈300 nJ and 30 fs, respectively. The signal is recollimated after the sample and the signal and reference are focused into a home-built dual channel prism-based spectrometer and recorded shot-to-shot by a pair of 1024 pixels CCD detectors (Stresing) synchronised to the AOPDF. The signal is referenced using an active noise reduction method proposed by Feng et al.84 The instrument response function (ca. 45 fs) is measured by spectrally resolving the instantaneous nonresonant solvent response. All measurements were conducted at the magic angle to rule out contributions from rotational diffusive motion to the measured signal decay. HB2DES beatmaps are obtained by a procedure previously discussed in the literature and outlined in the SI (Fig. S6).72,73,76
Computational methods
Crystallographic structures27 of the “open” and “closed” valPDI2 were solvated with CHCl3 (25 explicit) and H2O (40 explicit) using the SOLVATOR algorithm as part of the XTB package in ORCA 6.1.0.30,35 After solvent optimisation using molecular dynamics trajectories, the solvated structures were reoptimised using XTB. A series of 10 molecular dynamics trajectories using random seed velocities from a Wigner distribution for the open structure were performed at 300 K using canonical sampling velocity rescaling (CSVR) out to 200 ps in 0.5 fs steps to assess the average solvent occupancy of the cavity.85 It is not feasible to perform long-timescale simulations with only 40 water molecules due to the strong intermolecular hydrogen-bonding interactions inherent to water. Accurately capturing bulk-like behaviour in water would require hundreds of explicit molecules. In contrast, for chloroform, thermal energy at ambient temperatures is sufficient to overcome intermolecular forces. As a result, simulations with 25 chloroform molecules were found to yield reliable trajectories.Author contributions
Giovanni Bressan: conceptualisation; methodology; investigation; formal analysis; writing – original draft; writing – review and editing; funding acquisition; supervision. Denis Hartmann: investigation; methodology; resources; writing – review and editing. Jonathan Brouwer: investigation; methodology; writing – review and editing. Erico M. Braun: formal analysis; methodology; writing – review and editing. James N. Bull: software; formal analysis; writing – review and editing. Timothy A. Barendt: conceptualisation; funding acquisition; methodology; resources; supervision; writing – review and editing.Conflicts of interest
There are no conflicts to declare.Note added after first publication
This version replaces the manuscript published on 22nd January 2026 which contained an incorrect version of eqn (3). The RSC apologises for any confusion.Data availability
All the data that support the findings of this study are available in this article and its supplementary information (SI). The raw data are available from the corresponding authors upon request. Supplementary information: structure of reference PDI, refPDI ns fluorescence, discussion and calculations of solvent-dependent excitonic coupling, refPDI fsTA in CHCl3 and DMSO/water, fsTA and global fit analysis of valPDI2 in CHCl3 and DMSO/water and chemically reduced radical anion absorption, HB2DES of valPDI2 in CHCl3 and DMSO/water, scheme of the beatmap calculation method, valPDI2 550 cm−1 ring breathing coherence analysis, double-sided Feynman diagrams for hot ground state bleach, refPDI rephasing beatmaps of the 550 cm−1 mode, fluorescence quantum yields vs. water/DSMO volume ratio, coordinates. See DOI: https://doi.org/10.1039/d5sc09512c.Acknowledgements
G. B. is grateful to the Leverhulme Trust for funding him through an Early Career Fellowship (grant no. ECF-2023-195) and to EPSRC for supporting the Ultrafast Laser laboratory (EP/V00817X/1, EP/J009148/1). T. A. B. and D. H. thank the Engineering and Physical Sciences Research Council (EP/W037661/1) for funding. E. M. B. thanks the CNPq for his studentship (SWE200581/2025-2). The quantum chemical calculations presented in this work were carried out on the High-Performance Computing Cluster supported by the Research and Specialist Computing Support service at the University of East Anglia.References
- Th. Förster and K. Kasper, Ber. Bunsenges. Phys. Chem., 1955, 59, 976–980 CrossRef.
- J. Zirzlmeier, D. Lehnherr, P. B. Coto, E. T. Chernick, R. Casillas, B. S. Basel, M. Thoss, R. R. Tykwinski and D. M. Guldi, Proc. Natl. Acad. Sci. U. S. A., 2015, 112, 5325–5330 CrossRef CAS PubMed.
- R. Montero, V. Martínez-Martínez, A. Longarte, N. Epelde-Elezcano, E. Palao, I. Lamas, H. Manzano, A. R. Agarrabeitia, I. López Arbeloa, M. J. Ortiz and I. Garcia-Moreno, J. Phys. Chem. Lett., 2018, 641–646 CrossRef CAS PubMed.
- R. M. Young and M. R. Wasielewski, Acc. Chem. Res., 2020, 53, 1957–1968 CrossRef CAS PubMed.
- D. Thiel, H. Gotfredsen, P. M. Greißel, L. Chen, M. Krug, I. Papadopoulos, M. J. Ferguson, T. Torres, T. Clark, C. Neiss, A. Görling, M. B. Nielsen, R. R. Tykwinski and D. M. Guldi, J. Am. Chem. Soc., 2025, 147, 32124–32144 CrossRef CAS PubMed.
- A. K. Le, J. A. Bender, D. H. Arias, D. E. Cotton, J. C. Johnson and S. T. Roberts, J. Am. Chem. Soc., 2018, 140, 814–826 Search PubMed.
- C. M. Mauck, P. E. Hartnett, E. A. Margulies, L. Ma, C. E. Miller, G. C. Schatz, T. J. Marks and M. R. Wasielewski, J. Am. Chem. Soc., 2016, 138, 11749–11761 CrossRef CAS PubMed.
- A. Thampi, H. L. Stern, A. Cheminal, M. J. Y. Tayebjee, A. J. Petty, J. E. Anthony and A. Rao, J. Am. Chem. Soc., 2018, 7b06274 Search PubMed.
- C. B. Dover, J. K. Gallaher, L. Frazer, P. C. Tapping, A. J. Petty, M. J. Crossley, J. E. Anthony, T. W. Kee and T. W. Schmidt, Nat. Chem., 2018, 10, 305–310 Search PubMed.
- Y. Huang, I. A. Buyanova, C. Phansa, M. E. Sandoval-Salinas, D. Casanova, W. K. Myers, N. C. Greenham, A. Rao, W. M. Chen and Y. Puttisong, Cell Rep. Phys. Sci., 2021, 2, 100339 CrossRef CAS.
- H. Langhals and W. Jona, Angew. Chem., Int. Ed., 1998, 37, 952–955 CrossRef CAS PubMed.
- U. Selig, P. Nuernberger, V. Dehm, V. Settels, M. Gsänger, B. Engels, F. Würthner and T. Brixner, ChemPhysChem, 2013, 14, 1413–1422 CrossRef CAS PubMed.
- P. P. Das, A. Mazumder, M. Rajeevan, R. S. Swathi and M. Hariharan, Phys. Chem. Chem. Phys., 2024, 26, 2007–2015 RSC.
- G. Bressan, D. Green, Y. Chan, P. C. Bulman Page, G. A. Jones, S. R. Meech and I. A. Heisler, J. Phys. Chem. A, 2019, 123, 1594–1601 CrossRef CAS PubMed.
- P. Spenst, R. M. Young, M. R. Wasielewski and F. Würthner, Chem. Sci., 2016, 7, 5428–5434 RSC.
- T. Kim, C. Lin, J. D. Schultz, R. M. Young and M. R. Wasielewski, J. Am. Chem. Soc., 2022, 144, 11386–11396 CrossRef CAS PubMed.
- C. Lin, T. Kim, J. D. Schultz, R. M. Young and M. R. Wasielewski, Nat. Chem., 2022, 14, 786–793 CrossRef CAS PubMed.
- C. Ramanan, A. L. Smeigh, J. E. Anthony, T. J. Marks and M. R. Wasielewski, J. Am. Chem. Soc., 2012, 134, 386–397 CrossRef CAS PubMed.
- J. P. O’Connor, J. D. Schultz, N. A. Tcyrulnikov, T. Kim, R. M. Young and M. R. Wasielewski, J. Chem. Phys., 2024, 161, 074306 CrossRef PubMed.
- K. E. Brown, W. A. Salamant, L. E. Shoer, R. M. Young and M. R. Wasielewski, J. Phys. Chem. Lett., 2014, 5, 2588–2593 Search PubMed.
- R. E. Cook, B. T. Phelan, R. J. Kamire, M. B. Majewski, R. M. Young and M. R. Wasielewski, J. Phys. Chem. A, 2017, 121, 1607–1615 CrossRef CAS PubMed.
- J. D. Schultz, J. Y. Shin, M. Chen, J. P. O'Connor, R. M. Young, M. A. Ratner and M. R. Wasielewski, J. Am. Chem. Soc., 2021, 143, 2049–2058 CrossRef CAS PubMed.
- A. Mandal, M. Chen, E. D. Foszcz, J. D. Schultz, N. M. Kearns, R. M. Young, M. T. Zanni and M. R. Wasielewski, J. Am. Chem. Soc., 2018, 140, 17907–17914 Search PubMed.
- G. Bressan, S. E. Penty, D. Green, I. A. Heisler, G. A. Jones, T. A. Barendt and S. R. Meech, Angew. Chem., Int. Ed., 2024, 63, e202407242 Search PubMed.
- J. Sung, P. Kim, B. Fimmel, F. Würthner and D. Kim, Nat. Commun., 2015, 6, 8646 CrossRef CAS PubMed.
- K. Thakur, S. Datta, P. W. M. Blom, D. Chaudhuri and C. Ramanan, J. Phys. Chem. B, 2024, 128, 1760–1770 CrossRef CAS PubMed.
- D. Hartmann, S. E. Penty, M. A. Zwijnenburg, R. Pal and T. A. Barendt, Angew. Chem., Int. Ed., 2025, 64, e202501122 CrossRef CAS PubMed.
- S. E. Penty, M. A. Zwijnenburg, G. R. F. Orton, P. Stachelek, R. Pal, Y. Xie, S. L. Griffin and T. A. Barendt, J. Am. Chem. Soc., 2022, 144, 12290–12298 Search PubMed.
- S. M. Puranik, A. C. Kumbharkhane and S. C. Mehrotra, J. Chem. Soc., Faraday Trans., 1992, 88, 433–435 RSC.
- C. Bannwarth, S. Ehlert and S. Grimme, J. Chem. Theory Comput., 2019, 15, 1652–1671 Search PubMed.
- J. Rühe, K. Vinod, H. Hoh, K. Shoyama, M. Hariharan and F. Würthner, J. Am. Chem. Soc., 2024, 146(41), 28222–28232 Search PubMed.
- D. M. Cadena, V. Paulino, S. R. Allen, J. A. Maleszka, I. Tsironi, D. A. Darbah, J.-H. Olivier and S. T. Roberts, Chem. Mater., 2025, 37, 4059–4070 Search PubMed.
- J. Maillard, K. Klehs, C. Rumble, E. Vauthey, M. Heilemann and A. Fürstenberg, Chem. Sci., 2021, 12, 1352–1362 Search PubMed.
- J. Hausch, A. J. Berges, C. Zeiser, T. Rammler, A. Morlok, J. Bredehöft, S. Hammer, J. Pflaum, C. J. Bardeen and K. Broch, J. Phys. Chem. C, 2022, 126, 6686–6693 Search PubMed.
- F. Neese, F. Wennmohs, U. Becker and C. Riplinger, J. Chem. Phys., 2020, 152, 224108 Search PubMed.
- Z. Chen, B. Fimmel and F. Würthner, Org. Biomol. Chem., 2012, 10, 5845–5855 RSC.
- N. J. Hestand and F. C. Spano, Chem. Rev., 2018, 118, 7069–7163 CrossRef CAS PubMed.
- N. J. Hestand and F. C. Spano, J. Chem. Phys., 2015, 143, 244707 CrossRef PubMed.
- N. J. Hestand and F. C. Spano, Acc. Chem. Res., 2017, 50, 341–350 CrossRef CAS PubMed.
- M. Kasha, Radiat. Res., 1963, 20, 55–70 Search PubMed.
- A. Benny, R. Ramakrishnan and M. Hariharan, Chem. Sci., 2021, 12, 5064–5072 RSC.
- T. Lu and F. Chen, J. Comput. Chem., 2012, 33, 580–592 CrossRef CAS PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery Jr., J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, 2016, preprint.
- J. Brown, GitHub - JoshuaSBrown/QC_Tools: This Small Repository Provides Functionality for Calculating the Charge Transfer Integrals between Two Molecules., https://Github.Com/JoshuaSBrown/QC_Tools, accessed 29 May 2024.
- Y. Hong, M. Rudolf, M. Kim, J. Kim, T. Schembri, A. M. Krause, K. Shoyama, D. Bialas, M. I. S. Röhr, T. Joo, H. Kim, D. Kim and F. Würthner, Nat. Commun., 2022, 13, 4488 CrossRef CAS PubMed.
- C. Kaufmann, D. Bialas, M. Stolte and F. Würthner, J. Am. Chem. Soc., 2018, 140, 9986–9995 CrossRef CAS PubMed.
- Y. Hong, J. Kim, W. Kim, C. Kaufmann, H. Kim, F. Würthner and D. Kim, J. Am. Chem. Soc., 2020, 142, 7845–7857 Search PubMed.
- M. Son, K. H. Park, M.-C. Yoon, P. Kim and D. Kim, J. Phys. Chem. A, 2015, 119, 6275–6282 Search PubMed.
- J. Heitmüller, R. Fröhlich, R. Renner, F. Würthner and T. Brixner, Phys. Chem. Chem. Phys., 2023, 25, 17214–17229 Search PubMed.
- J. J. Snellenburg, S. P. Laptenok, R. Seger, K. M. Mullen and I. H. M. van Stokkum, J. Stat. Softw., 2012, 49, 1–22 Search PubMed.
- J. Sung, A. Nowak-Król, F. Schlosser, B. Fimmel, W. Kim, D. Kim and F. Würthner, J. Am. Chem. Soc., 2016, 138, 9029–9032 CrossRef CAS PubMed.
- S. Samanta and D. Chaudhuri, J. Phys. Chem. Lett., 2017, 3427–3432 CrossRef CAS PubMed.
- E. Sebastian, J. Sunny and M. Hariharan, Chem. Sci., 2022, 13, 10824–10835 Search PubMed.
- M. Son, K. H. Park, C. Shao, F. Würthner and D. Kim, J. Phys. Chem. Lett., 2014, 5, 3601–3607 Search PubMed.
- C. Kaufmann, W. Kim, A. Nowak-Król, Y. Hong, D. Kim and F. Würthner, J. Am. Chem. Soc., 2018, 140, 4253–4258 Search PubMed.
- Y. Hong, W. Kim, T. Kim, C. Kaufmann, H. Kim, F. Würthner and D. Kim, Angew. Chem., Int. Ed., 2022, 61, 2021144 Search PubMed.
- D. Chaudhuri, D. Li, Y. Che, E. Shafran, J. M. Gerton, L. Zang and J. M. Lupton, Nano Lett., 2011, 11, 488–492 Search PubMed.
- S. Samanta, S. K. Ray, S. Deolka, S. Saha, P. K. R., R. Bhowal, N. Ghosh and D. Chaudhuri, Chem. Sci., 2020, 11, 5710–5715 Search PubMed.
- R. F. Fink, J. Seibt, V. Engel, M. Renz, M. Kaupp, S. Lochbrunner, H.-M. Zhao, J. Pfister, F. Würthner and B. Engels, J. Am. Chem. Soc., 2008, 130, 12858–12859 CrossRef CAS PubMed.
- S. Roy and B. Bagchi, J. Chem. Phys., 2013, 139, 034308 Search PubMed.
- J. M. Alzola, N. A. Tcyrulnikov, P. J. Brown, T. J. Marks, M. R. Wasielewski and R. M. Young, J. Phys. Chem. A, 2021, 125, 7633–7643 CrossRef CAS PubMed.
- S. K. Lee, Y. Zu, A. Herrmann, Y. Geerts, K. Müllen and A. J. Bard, J. Am. Chem. Soc., 1999, 121, 3513–3520 Search PubMed.
- K. M. Lefler, K. E. Brown, W. A. Salamant, S. M. Dyar, K. E. Knowles and M. R. Wasielewski, J. Phys. Chem. A, 2013, 117, 10333–10345 CrossRef CAS PubMed.
- M. Mayländer, O. Nolden, M. Franz, S. Chen, L. Bancroft, Y. Qiu, M. R. Wasielewski, P. Gilch and S. Richert, Chem. Sci., 2022, 13, 6732–6743 Search PubMed.
- A. K. Le, J. A. Bender, D. H. Arias, D. E. Cotton, J. C. Johnson and S. T. Roberts, J. Am. Chem. Soc., 2018, 140, 814–826 Search PubMed.
- A. K. Le, J. A. Bender and S. T. Roberts, J. Phys. Chem. Lett., 2016, 7, 4922–4928 Search PubMed.
- Y. Yang, W. Su, H. Wang, X. Bao, X. Liu, Z. Bo and W. Zhang, J. Phys. Chem. B, 2024, 128, 7219–7226 Search PubMed.
- D. Veldman, S. M. A. Chopin, S. C. J. Meskers, M. M. Groeneveld, R. M. Williams and R. A. J. Janssen, J. Phys. Chem. A, 2008, 112(26), 5846–5857 CrossRef CAS PubMed.
- E. A. Margulies, J. L. Logsdon, C. E. Miller, L. Ma, E. Simonoff, R. M. Young, G. C. Schatz and M. R. Wasielewski, J. Am. Chem. Soc., 2017, 139, 663–671 CrossRef CAS PubMed.
- S. W. Eaton, L. E. Shoer, S. D. Karlen, S. M. Dyar, E. A. Margulies, B. S. Veldkamp, C. Ramanan, D. A. Hartzler, S. Savikhin, T. J. Marks and M. R. Wasielewski, J. Am. Chem. Soc., 2013, 135, 14701–14712 Search PubMed.
- F. C. Spano, J. Phys. Chem. C, 2024, 128, 248–260 CrossRef CAS.
- G. Bressan, I. A. Heisler, G. M. Greetham, A. Edmeades and S. R. Meech, Opt. Express, 2023, 31, 42687 Search PubMed.
- G. Bressan, D. Green, G. A. Jones, I. A. Heisler and S. R. Meech, J. Phys. Chem. Lett., 2024, 15, 2876–2884 Search PubMed.
- F. V. A. Camargo, H. L. Anderson, S. R. Meech and I. A. Heisler, J. Phys. Chem. A, 2014, 119, 95–101 Search PubMed.
- J. C. Dean and G. D. Scholes, Acc. Chem. Res., 2017, 50, 2746–2755 CrossRef CAS PubMed.
- F. V. de A. Camargo, L. Grimmelsmann, H. L. Anderson, S. R. Meech and I. A. Heisler, Phys. Rev. Lett., 2017, 118, 033001 Search PubMed.
- D. Green, G. Bressan, I. A. Heisler, S. R. Meech and G. A. Jones, J. Chem. Phys., 2024, 160, 1–12 Search PubMed.
- S. R. Meech, J. N. Bull and G. Bressan, J. Chem. Phys., 2025, 162, 184302 CrossRef CAS PubMed.
- G. Bressan, D. Green, G. A. Jones, I. A. Heisler and S. R. Meech, J. Phys. Chem. Lett., 2024, 15, 2876–2884 CrossRef CAS PubMed.
- A. C. Albrecht, J. Chem. Phys., 1961, 34, 1476–1484 Search PubMed.
- J. Hoche, H.-C. Schmitt, A. Humeniuk, I. Fischer, R. Mitrić and M. I. S. Röhr, Phys. Chem. Chem. Phys., 2017, 19, 25002–25015 Search PubMed.
- C. R. Hall, J. Conyard, I. A. Heisler, G. Jones, J. Frost, W. R. Browne, B. L. Feringa and S. R. Meech, J. Am. Chem. Soc., 2017, 139, 7408–7414 Search PubMed.
- S. Yan and H. S. Tan, Chem. Phys., 2009, 360, 110–115 Search PubMed.
- Y. Feng, I. Vinogradov and N.-H. Ge, Opt. Express, 2017, 25, 26262 CrossRef CAS PubMed.
- G. Bussi, D. Donadio and M. Parrinello, J. Chem. Phys., 2007, 126, 014101 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |