Open Access Article
Open Access ArticleMitochondria-targeting cyclometallated rhodium(III) complexes appended with two rhodamine units as Type I photosensitisers for bioimaging and photocytotoxicity applications by inducing pyroptosis
Katherine Gui-Min
Jiang
 ab,
Guang-Xi
Xu
ab,
Guang-Xi
Xu
 a,
Lawrence Cho-Cheung
Lee
a,
Lawrence Cho-Cheung
Lee
 a,
Fangfang
Wei
a,
Fangfang
Wei
 b,
Siye
Wu
b,
Keith Man-Chung
Wong
b,
Siye
Wu
b,
Keith Man-Chung
Wong
 *b and
Kenneth Kam-Wing
Lo
*b and
Kenneth Kam-Wing
Lo
 *ac
*ac
aDepartment of Chemistry, City University of Hong Kong, Tat Chee Avenue, Kowloon, Hong Kong, P. R. China. E-mail: bhkenlo@cityu.edu.hk
bDepartment of Chemistry, Southern University of Science and Technology, 1088 Xueyuan Boulevard, Shenzhen 518055, P. R. China. E-mail: keithwongmc@sustech.edu.cn
cState Key Laboratory of Terahertz and Millimetre Waves, City University of Hong Kong, Tat Chee Avenue, Kowloon, Hong Kong, P. R. China
First published on 5th March 2026
Abstract
Cyclometallated rhodium(III) complexes have been underexplored as photosensitisers due to their low-lying d–d excited states, which result in weak visible-light absorption and non-emissive properties, coupled with a modest heavy atom effect that limits reactive oxygen species (ROS) generation. In this work, a series of cyclometallated rhodium(III) polypyridine complexes appended with two rhodamine units [Rh(N^C)2(bpy-diRho)](PF6)3 was rationally designed as Type I photosensitisers. These complexes exhibited intense absorption in the visible region and moderate rhodamine fluorescence in solution upon photoexcitation. Time-resolved transient absorption spectroscopy revealed a long-lived rhodamine-based triplet excited state as the lowest-lying excited state in this hybrid system, which is attributed to the presence of the rhodium(III) centre and is responsible for ROS photosensitisation. Notably, these rhodium(III) complexes efficiently generated superoxide anion (O2˙−) and hydroxyl (HO˙) radicals via the Type I pathway upon photoirradiation, likely via intramolecular electron transfer between the two adjacent excited rhodamine units within the complex to form radical cation and anion. Cellular colocalisation studies demonstrated that these complexes predominantly accumulated in mitochondria, where the photosensitised ROS triggered significant mitochondrial dysfunction, resulting in their outstanding photocytotoxicity under both normoxic and CoCl2-induced hypoxic conditions. Further mechanistic investigations revealed that the photoinduced mitochondrial ROS generation triggered cancer cell death via gasdermin D-mediated pyroptosis. This rhodium(III)–dirhodamine system further explores the utilisation of rhodium(III) complexes as phototheranostic agents and underscores their potential in this role.
Introduction
Photodynamic therapy (PDT) offers a promising approach for clinical disease treatment with minimal invasiveness, high spatiotemporal precision and no drug resistance.1–3 The photosensitiser (PS), as the key component of PDT, is activated by light of an appropriate wavelength, generating reactive oxygen species (ROS) to destroy cancer cells or tissues.4,5 An ideal PS is expected to exhibit low dark cytotoxicity, high photocytotoxicity, good biocompatibility, strong absorption at longer wavelengths and efficient ROS generation.6,7 Facilitating the intersystem crossing (ISC) process is critical to increase the population of the triplet excited state of the PS, thereby boosting ROS production. Over the past decade, significant efforts have focused on developing effective PSs based on transition metal complexes, leveraging their inherent heavy atom effect to achieve superior ROS photosensitisation.8,9 However, these complexes often require high excitation energy, which results in low tissue penetration and restricts their clinical applications. Additionally, their relatively weak absorption beyond 450 nm hinders efficient utilisation of irradiation sources.10,11 To address these challenges, integrating chromophore groups into these complexes is anticipated as a versatile strategy to lower the excitation energy and enhance absorptivity at the excitation wavelength, improving their PDT efficacy.Organelle targeting has emerged as a potent method to enhance the therapeutic efficacy of PSs. Mitochondria, referred to as the cellular “powerhouse”, play crucial roles in adenosine triphosphate (ATP) production, intracellular ROS generation and the regulation of programmed cell death, rendering them prime targets for cancer therapy.12 The highly negative membrane potential of mitochondria in cancer cells promotes the mitochondrial uptake of lipophilic cationic compounds.13 Particularly, many cationic transition metal complexes and rhodamine derivatives exhibit specific mitochondrial targeting.14–20 It is worth mentioning that mitochondria are susceptible to excessive ROS levels, leading to mitochondrial dysfunction.21,22 Thus, selective delivery of PSs to mitochondria is an effective approach to improve their efficacy in cancer therapy.
Pyroptosis, identified as a pro-inflammatory form of cell death, is characterised by cellular swelling, the formation of large bubbles from the plasma membrane and eventual plasma membrane rupture.23 Throughout the pyroptotic process, the gasdermin D (GSDMD) protein acts as the key executor, cleaved by activated caspase-1 to produce a perforating N-terminal fragment (GSDMD-N) that punctures the cell membrane.24 This leads to cell swelling and eventual rupture, releasing cellular contents and triggering a robust local inflammatory response.25 It has been reported that elevated levels of mitochondrial ROS (mtROS) can result in mitochondrial damage and induce cell death through pyroptosis.24,26,27 Consequently, the induction of pyroptosis through mitochondria-targeting PSs represents a promising strategy to enhance therapeutic outcomes in cancer treatment.
Cyclometallated rhodium(III) complexes are a well-explored class of transition metal complexes that serve as vital metalloinsertors due to their exceptional selectivity for DNA mismatches.28–31 These complexes also function as inhibitors of specific protein–protein interactions.32–34 However, their emission and photosensitisation capabilities are limited by the low-lying non-radiative d–d excited states. Through the strategic modification of their cyclometallating and ancillary ligands, these complexes can be tailored to exhibit tunable photophysical properties. Notably, the incorporation of potent σ-donating cyclometallating ligands into rhodium(III) complexes has enabled luminescence from a lower-lying ligand-centred state.35 Rhodamines, a representative class of organic dyes, exhibit strong absorption in the visible region and high fluorescence quantum yields, making them widely used as bioimaging probes and sensors.36–38 However, rhodamine and its derivatives have been less used as PSs due to inefficient ISC processes and low population of the triplet excited state.
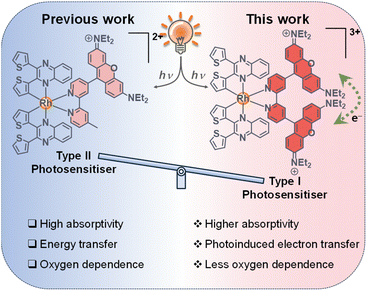
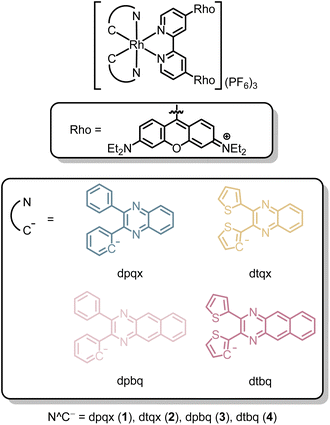
To promote triplet-state population, rhodamine has been incorporated into cyclometallated iridium(III) polypyridine scaffolds, which enables efficient population of the rhodamine triplet state (T1) via two mechanisms.39,40 One pathway involves a cascade of energy transfer through the [Ir(N^C)2]-based triplet excited states. In the other pathway, the exceptionally large spin–orbit coupling (SOC) constant of iridium (ca. 2500–3500 cm−1) also facilitates highly efficient direct ISC from the rhodamine singlet excited state (S1) to the rhodamine T1 state. This direct ISC pathway remains highly effective even when the indirect cascade mechanism is energetically inaccessible or suppressed, ensuring the population of the rhodamine T1 state and, consequently, the generation of ROS for PDT. However, the rhodamine fluorescence quantum yields of these complexes are quite low (Φem = 0.004–0.037), making them less ideal for bioimaging. In our previous work, we designed a series of cyclometallated rhodium(III) monorhodamine complexes.41 The relatively smaller SOC constant of rhodium (ca. 1200–1400 cm−1) renders direct ISC from the rhodamine S1 to T1 state inefficient. As a result, this rhodium(III)–monorhodamine system displays more intense fluorescence (Φem = 0.039–0.25). Thus, the selection of rhodium(III) as the metal centre affords a delicate balance between fluorescence and triplet excited-state population for ROS generation, which enables the rhodium(III) monorhodamine complexes to serve as versatile theranostic agents with both real-time imaging and ROS generation capabilities. However, these complexes primarily generate singlet oxygen (1O2) via the Type II photoreaction, and their strong O2 dependence limits effectiveness in hypoxic solid tumours arising from rapid cancer cell proliferation.42,43 To advance the intriguing rhodium(III)–rhodamine hybrid system and develop less O2-dependent Type I PSs, we reasoned that introducing an additional rhodamine group into the ancillary ligand would improve the PDT performance of these complexes. This modification is anticipated to not only significantly enhance the molar absorptivity of the rhodium(III) complexes, but also increase the possibility of intramolecular electron transfer between the two proximate rhodamine units via symmetry-breaking charge transfer (SBCT) upon photoexcitation, thereby improving the efficiency of Type I ROS photosensitisation (Fig. 1).44 Additionally, complexes bearing an extra positive charge are expected to facilitate intermolecular electron transfer to the surrounding substrates, further promoting ROS generation via the Type I pathway.45,46 Given these features, installing two rhodamine units is anticipated to optimise the performance of cyclometallated rhodium(III) complexes as PSs for bioimaging and therapeutic applications. Herein, we report the design of a series of cyclometallated rhodium(III) complexes appended with two rhodamine units [Rh(N^C)2(bpy-diRho)](PF6)3 (bpy-diRho = 4,4′-(9-(3,6-bis(diethylamino))xanthylium)-2,2′-bipyridine; HN^C = 2,3-diphenylquinoxaline (Hdpqx) (1), 2,3-dithienylquinoxaline (Hdtqx) (2), 2,3-diphenylbenzo[g]quinoxaline (Hdpbq) (3) and 2,3-dithienylbenzo[g]quinoxaline (Hdtbq) (4)) (Chart 1). The photophysical properties and ROS photosensitisation abilities of these complexes were evaluated in solution. Additionally, the cellular uptake, subcellular localisation, intracellular ROS photosensitisation and (photo)cytotoxicity of the free ligand and the rhodium(III) complexes were investigated. Western blot analysis and lactate dehydrogenase (LDH) release assays were also conducted to examine the induction of pyroptosis in cells.
Results and discussion
Synthesis and characterisation
The synthesis of the free ligand (bpy-diRho)(PF6)2 involved the reaction of 4,4′-dicarbaldehyde-2,2′-bipyridine with 3-(diethylamino)phenol, followed by the addition of p-toluenesulfonic acid. Then the anion of the product was exchanged with NH4PF6 and the product was purified by column chromatography. The final compound was recrystallised from CH2Cl2/Et2O, yielding purple crystals. The dirhodamine-containing cyclometallated rhodium(III) complexes were prepared from the reaction of the corresponding rhodium(III) dimers [Rh2(N^C)4Cl2] with the ligand (bpy-diRho)(PF6)2 in CH2Cl2/MeOH, followed by anion exchange with NH4PF6. These complexes were purified by column chromatography and recrystallised from CH2Cl2/Et2O as purple crystals. The ligand and the four complexes were characterised by 1H and 13C NMR and high-resolution electrospray ionisation mass spectrometry (HR-ESI-MS), and their elemental analyses were found to be satisfactory.The single crystal structure of complex 1 was established by X-ray crystallography, and the crystallographic data are listed in Table S1. The perspective view of the cation is depicted in Fig. 2. All the bond lengths (Å) and angles (°) of the rhodium(III) chromophore and rhodamine moieties are in the normal range, and selected bond lengths and angles are provided in Table S2. The rhodium(III) centre adopted a distorted octahedral geometry, and the trans angles at the metal centre ranged from 169.8(7) to 173.1(5)°. The two xanthene planes of the rhodamine moieties are oriented orthogonally to the bipyridine plane, displaying dihedral angles of 56.2(9)° and 76.4(0)°, respectively. For the cyclometallating ligand dpqx, the dihedral angles between the coordinating phenyl rings and the quinoxaline planes are 24.4(2)° and 12.1(5)°, while those of the pendant phenyl rings are much larger (50.1(3)° and 59.0(1)°, respectively).
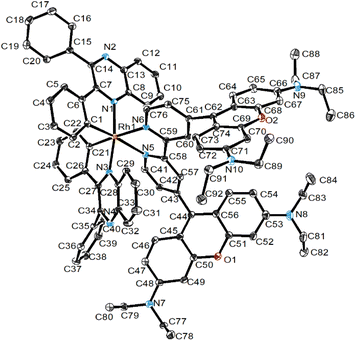 | ||
| Fig. 2 Perspective drawing of the cation of complex 1. Hydrogen atoms, counterions and solvent molecules are omitted for clarity. | ||
Photophysical properties
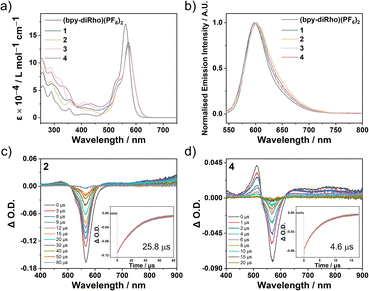
The electronic absorption spectral data of (bpy-diRho)(PF6)2 and the corresponding rhodium(III) complexes in CH3CN at 298 K are presented in Table S3 and their electronic absorption spectra are displayed in Fig. 3a. The free ligand (bpy-diRho)(PF6)2 showed moderate absorption bands in the UV region (ca. 250–350 nm), which is attributed to the spin-allowed (n → π* and π → π*) transitions of bipyridine and rhodamine, and displayed an intense absorption band at 562 nm, characteristic of rhodamine (π → π*) transition. Compared to the analogous monorhodamine ligand (bpy-Rho)(PF6) (ε = ca. 1.0 × 105 L mol−1 cm−1 at 562 nm),47 (bpy-diRho)(PF6)2 exhibited higher absorptivity (ε = 1.7 × 105 L mol−1 cm−1 at 562 nm) due to the presence of an additional rhodamine chromophore, facilitating visible light-harvesting in imaging and PDT applications. For the four rhodium(III) complexes, absorption bands were observed in the UV region (ca. 250–350 nm), assigned to the overlap of spin-allowed intraligand (1IL) (π → π*) (N^C) and 1IL (n → π* and π → π*) (bipyridine and rhodamine) transitions.41 The relatively weak bands in the visible region (ca. 350–500 nm) are attributed to the 1IL transitions from phenyl or thienyl rings to quinoxaline or benzo[g]quinoxaline groups, mixing with some spin-allowed metal-to-ligand charge transfer (1MLCT) (dπ(Rh) → π*(N^C)) transitions. Notably, these rhodium(III) complexes also displayed the characteristic rhodamine absorption at 572 nm. The red shift of the rhodamine absorption band is due to the incorporation of the electron-withdrawing [Rh(N^C)2] moiety, indicative of the interaction between the rhodium(III) metal centre and the rhodamine units.41Upon photoexcitation at 525 nm, (bpy-diRho)(PF6)2 and rhodium(III) complexes 1–4 showed red emission in CH3CN under ambient conditions. The photophysical properties of these compounds are summarised in Table 1 and their emission spectra are presented in Fig. 3b. The free ligand (bpy-diRho)(PF6)2 exhibited characteristic rhodamine fluorescence with a high emission quantum yield (0.14). The dirhodamine-containing rhodium(III) complexes also displayed similar emission spectra. However, the emission quantum yields of these complexes dramatically decreased (0.012–0.052), suggesting the presence of another decay pathway. Notably, a negligible peak at ca. 820 nm was observed in the emission spectra of complexes 2–4 in alcohol glass at 77 K (Fig. S1), which can be attributed to the triplet excited state of rhodamine.
| Compound | λ em/nma | Φ em | τ TA/µsc | Φ Δ | Φ Δ |
|---|---|---|---|---|---|
| a Emission maximum wavelength (λex = 525 nm). b The absolute emission quantum yields were measured using an integrating sphere (λex = 570 nm). c The TA lifetimes at 575 nm were measured in deaerated CH3CN (λex = 532 nm). d The 1O2 quantum yields were determined based on the emission of 1O2 at 1270 nm using Rose Bengal (ΦΔ = 0.45 in aerated CH3CN) as a reference (λex = 570 nm). e The 1O2 quantum yields were determined by using DPBF as a 1O2 scavenger. f Could not be determined. | |||||
| (bpy-diRho)(PF6)2 | 598 | 0.14 | —f | —f | —f |
| 1 | 599 | 0.052 | —f | —f | —f |
| 2 | 602 | 0.018 | 25.8 | 0.06 | 0.05 |
| 3 | 601 | 0.012 | 10.3 | 0.05 | 0.05 |
| 4 | 601 | 0.016 | 4.6 | 0.22 | 0.25 |
Our previous study has demonstrated that the coordination of rhodamine to rhodium(III) facilitates the population of the rhodamine-based triplet excited state.41 To investigate the triplet excited states of (bpy-diRho)(PF6)2 and the rhodium(III) complexes, nanosecond time-resolved transient absorption (TA) spectroscopy was conducted in deaerated CH3CN, with results presented in Table 1, and Fig. 3c, d, S2 and S3. Upon pulse laser excitation at 532 nm, the free ligand (bpy-diRho)(PF6)2 showed no detectable signal in the TA difference spectrum, illustrating negligible triplet excited state population. Conversely, the dirhodamine-decorated rhodium(III) complexes exhibited a prominent photobleaching signal at 575 nm, which is attributed to the depletion of ground-state rhodamine absorption and ascribed to the 3IL excited state (T1) localised on the rhodamine moieties.39–41,47,48 For complex 4, the positive peak at ca. 512 nm was attributed to the [Rh(N^C)2] moiety-based 3IL/3MLCT excited state, demonstrating a thermally accessible equilibrium between closely-lying triplet excited states of the rhodamine unit and [Rh(N^C)2] moiety.41 Additionally, the lifetimes of the rhodamine T1 state in complexes 2–4 were relatively long, ranging from 4.6 to 25.8 µs. Compared to their monorhodamine-containing rhodium(III) counterparts (4.1–8.7 µs),41 these dirhodamine-containing complexes exhibited significantly prolonged triplet excited-state lifetimes. Since the population of the rhodamine T1 state arises from rhodium(III) coordination, introducing a second rhodamine unit likely attenuates the heavy atom effect of the rhodium centre on any individual chromophore, which diminishes the ISC-mediated non-radiative decay from the rhodamine T1 state to the ground state and thereby extends rhodamine T1 state lifetimes.49
An energy level diagram was constructed to delineate the excited-state pathways in this system (Fig. S4). The energies of the rhodamine S1 and T1 states were estimated from their emission spectra, while the energies of the triplet excited state of the [Rh(N^C)2] moiety (T1′) were proposed according to the emission data of the rhodamine-free complexes.41 The T1′ state of these complexes served as a cascade to facilitate population of the rhodamine T1 state from the rhodamine S1 state via sequential singlet–triplet energy transfer (STET) and triplet–triplet energy transfer (TTET). Notably, for complexes 3 and 4, the relatively small energy gaps between T1′ and T1 states (0.24 and 0.22 eV, respectively) render reversible TTET thermally accessible, and thus both triplet manifolds are likely populated simultaneously. This is supported by the TA spectra of complex 4 (Fig. 3d), which display features consistent with two triplet manifolds, demonstrating a thermally accessible equilibrium between the T1′ and T1 states. In contrast, for complexes 1 and 2, the larger T1′ ‒ T1 energy gaps (0.77 and 0.54 eV, respectively) disfavour this process, and thus only the T1 state was predominantly populated.
ROS photosensitisation
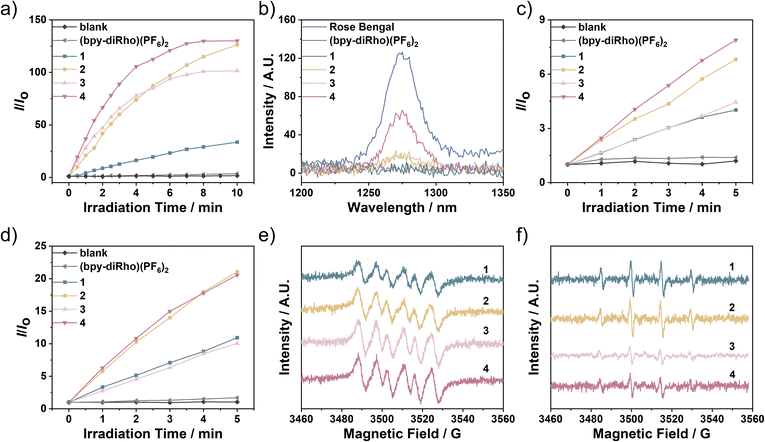
Given the effective population of the rhodamine T1 state, the dirhodamine-decorated rhodium(III) complexes were expected to exhibit efficient ROS photosensitisation. To assess their total ROS generation capabilities in aqueous solutions semi-quantitatively, the ROS probe 2′,7′-dichlorodihydrofluorescein diacetate (DCFH-DA) was employed. After hydrolysis of DCFH-DA into 2′,7′-dichlorodihydrofluorescein (DCFH), the non-emissive DCFH can be oxidised by ROS into 2′,7′-dichlorofluorescein (DCF), which gives strong green fluorescence. Upon white-light irradiation (400–700 nm, 2 mW cm−2), the emission intensity of DCF was monitored at 525 nm, and the emission enhancement is displayed in Fig. 4a. Significant fluorescence enhancement was observed upon irradiation for 3 min in the presence of complexes 2–4, indicating their robust ROS photosensitisation. However, negligible ROS was sensitised by (bpy-diRho)(PF6)2 and only a small amount of ROS was generated in the presence of complex 1. To identify specific ROS generated by these complexes, their 1O2 generation quantum yields (ΦΔ) were quantitatively determined in aerated CH3CN by measuring the 1O2 emission and the photooxidation of 1,3-diphenylisobenzofuran (DPBF), a 1O2 scavenger. The 1O2 emission spectra and the decay rates of DPBF are presented in Fig. 4b and S5. Upon 570-nm excitation, no detectable 1O2 emission was observed for (bpy-diRho)(PF6)2 or complex 1. A weak emission band was measured for complexes 2 and 3, and a quantifiable 1O2 emission band was detected for complex 4. Using a commercially available 1O2 PS, Rose Bengal, as the standard (ΦΔ = 0.45 in CH3CN),50 the ΦΔ value of complex 4 was determined to be 0.22, consistent with DPBF photooxidation results (0.25). In contrast to the efficient 1O2 photosensitisation of rhodium(III) complexes appended with only one rhodamine unit,41 these dirhodamine-decorated rhodium(III) complexes possessed relatively low ΦΔ values, demonstrating that they were poor 1O2 PSs. Considering their remarkable total ROS generation capabilities, it is reasonable to infer that other ROS species were likely photosensitised.Consequently, two commercially available ROS indicators, dihydrorhodamine 123 (DHR123) and hydroxyphenyl fluorescein (HPF), were applied to semi-quantitatively examine the generation of superoxide anion (O2˙−) and hydroxyl (HO˙) radicals, respectively, in aqueous medium by the complexes, which can be oxidised to rhodamine 123 (R123) and HPF-ox, giving strong fluorescence. Upon irradiation with white light (400–700 nm, 2 mW cm−2), negligible fluorescence enhancement of DHR123 was detected in the presence of (bpy-diRho)(PF6)2 (Fig. 4c), indicating minimal O2˙− production. Moderate emission increases were observed for complexes 1 and 3 within 5 min of light irradiation, while significant enhancement was observed for complexes 2 and 4, demonstrating their superior O2˙− photosensitisation among these samples. Their O2˙− generation efficiencies followed the order: (bpy-diRho)(PF6)2 < 1 < 3 < 2 < 4. Similarly, in studies of HO˙ radicals, no notable emission increase from HPF was observed in the presence of (bpy-diRho)(PF6)2 (Fig. 4d), suggesting that it was unable to generate HO˙. However, upon incorporation of a rhodium(III) centre, complexes 1 and 3 were capable of producing HO˙, whereas complexes 2 and 4 exhibited substantial HO˙ photosensitisation. Additionally, electron paramagnetic resonance (EPR) spectroscopy confirmed the generation of free radicals for complexes 1–4 upon white-light irradiation (400–700 nm, 10 mW cm−2) using 5,5-dimethyl-1-pyrroline N-oxide (DMPO) as a spin-trapping reagent. As shown in Fig. 4e, distinct DMPO-O2˙− signals were detected for complexes 1–4 upon light irradiation in aerated MeOH. Furthermore, a characteristic 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 quartet signal was observed in aerated aqueous solutions (Fig. 4f), which indicated the formation of the DMPO-HO˙ adduct. The efficient generation of O2˙− and HO˙ can be attributed to the long-lived rhodamine T1 state of the dirhodamine-decorated rhodium(III) complexes, which facilitate electron transfer to the surrounding molecules.51 We also evaluated and compared the ROS generation efficiencies of the dirhodamine complex 2 and the monorhodamine analogue [Rh(dtqx)2(bpy-Rho)](PF6)2 (2a) and their rhodamine-free counterpart [Rh(dtqx)2(Me2-bpy)](PF6) (2b).41 Remarkably, complex 2 showed dramatically increased ROS generation upon photoexcitation via the Type I pathway due to the additional rhodamine unit (Fig. S6 and S7). These findings demonstrate that rhodium(III) dirhodamine complexes can sensitise ROS via a Type I mechanism under light exposure, with complexes 2 and 4 showing particularly high efficiencies.
1 quartet signal was observed in aerated aqueous solutions (Fig. 4f), which indicated the formation of the DMPO-HO˙ adduct. The efficient generation of O2˙− and HO˙ can be attributed to the long-lived rhodamine T1 state of the dirhodamine-decorated rhodium(III) complexes, which facilitate electron transfer to the surrounding molecules.51 We also evaluated and compared the ROS generation efficiencies of the dirhodamine complex 2 and the monorhodamine analogue [Rh(dtqx)2(bpy-Rho)](PF6)2 (2a) and their rhodamine-free counterpart [Rh(dtqx)2(Me2-bpy)](PF6) (2b).41 Remarkably, complex 2 showed dramatically increased ROS generation upon photoexcitation via the Type I pathway due to the additional rhodamine unit (Fig. S6 and S7). These findings demonstrate that rhodium(III) dirhodamine complexes can sensitise ROS via a Type I mechanism under light exposure, with complexes 2 and 4 showing particularly high efficiencies.
Given their enhanced abilities to photosensitise radicals via electron transfer, the excited-state redox potentials of (bpy-diRho)(PF6)2 and complexes 1–4 were estimated using Latimer diagrams (Table S4, Fig. S8 and S9). They exhibited quasi-reversible oxidation couples (E° [PS(S0)4+/PS(S0)3+]) at approximately +1.25 V versus SCE, which is ascribed to the oxidation of rhodamine units. Considering the rhodamine T1 state energy of 1.53 eV, their excited-state reductional potentials (E° [PS(S0)4+/PS(T1)3+]) were calculated to be −0.30 V (Fig. S9). The free energy (ΔG) for the electron transfer reaction from complexes 1–4 to 3O2, forming O2˙− (E° (O2/O2˙−) = 0.084 V versus SCE) (1), was determined to be ca. −0.36 eV, suggesting that it is a thermodynamically favourable reaction. Alternatively, the triplet excited-state PS(T1) can undergo reduction to its radical anion form (PS˙−) through electron transfer from electron-rich substrates in the cellular environment, such as certain amino acid residues, nucleic acid components, coenzymes and antioxidant vitamins.45,46 The 3O2 is reduced by the resultant PS˙− in close proximity to produce O2˙−(2). Considering the relatively long triplet excited-state lifetimes and the presence of two adjacent rhodamine units, it is plausible to hypothesise that SBCT occurs within the triplet manifold.52–55 Although direct spectroscopic evidence, such as TA data for SBCT, is not available, most likely because radical-ion intermediates often have lifetimes on the picosecond scale and fall outside the temporal window of our instrumentation, the transient formation of rhodamine radical cation and anion cannot be excluded, given that SBCT can proceed via ultrafast charge separation without yielding detectable long-lived radical-ion bands in nanosecond TA spectra.56 The efficient generation of ROS via the Type I pathway differentiates the rhodium(III) dirhodamine complexes from related monorhodamine systems for PDT.
| PS(T1) + 3O2 → PS˙+ + O2˙− | (1) |
| PS˙− + 3O2 → PS + O2˙− | (2) |
Intracellular localisation
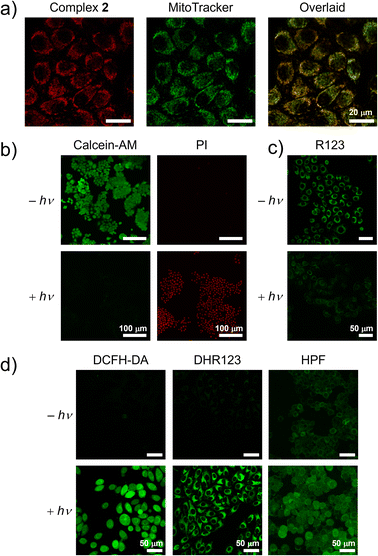
To assess the chemical stability and speciation of the dirhodamine-decorated rhodium(III) complexes under biologically relevant conditions, we selected complex 2 as a model and incubated it in cell culture medium at 37 °C for 24 h, followed by ESI-MS analysis. The complex remained intact in the medium with no discernible changes in speciation (Fig. S10), underscoring its stability and suitability for cellular applications. The biological properties of the ligand and rhodium(III) complexes were then investigated using MCF-7 cells as a model cell line. Intracellular localisation of (bpy-diRho)(PF6)2 and complexes 1–4 was examined by laser-scanning confocal microscopy (LSCM). As illustrated in Fig. 5a and S11, distinct rhodamine fluorescence was observed in the perinuclear region with a network morphology characteristic of mitochondria. Costaining experiments using MitoTracker Green confirmed mitochondrial accumulation of the ligand and complexes, supported by high Pearson's correlation coefficients (PCCs; 0.82–0.91). To further verify the mitochondrial localisation of these dirhodamine-containing rhodium(III) complexes, complex 2-treated MCF-7 cells were fractionated into mitochondria, nucleus and cytosol using a commercial kit and the rhodium content in each fraction was quantified using inductively coupled plasma-mass spectrometry (ICP-MS). Remarkably, the complex predominantly accumulated in the mitochondria (81% of total rhodium), with minor nuclear localisation and negligible cytosolic levels (Fig. S12). These quantitative data corroborate the costaining experiments (Fig. 5a) and confirm the mitochondrial specificity of the rhodium(III) complexes, which is likely related to their tricationic charge. The intense intracellular emission of MCF-7 cells treated with these complexes indicates their potential as efficient mitochondrial imaging agents.Cellular uptake and (photo)cytotoxicity studies
The cellular uptake efficiencies of the rhodium(III) complexes were quantified by ICP-MS, and the results are listed in Table S5. Upon incubation of MCF-7 cells with the complexes, the average rhodium content per cell ranged from 0.15 to 0.27 fmol. Compared to complexes 1 and 2 (0.22 and 0.27 fmol, respectively), complexes 3 and 4 exhibited lower uptake efficiencies (0.15 and 0.20 fmol, respectively), potentially due to the larger size of their cyclometallating ligands.The (photo)cytotoxicity of complexes 1–4 towards MCF-7 cells under normoxic conditions was assessed using 3-(4,5-diphenyl-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assays, and the results are presented in Table 2, Fig. S13 and S14. Upon incubation with the complexes in the dark for 6 h, no considerable decrease in cell viability was observed, with the IC50,dark values of these complexes exceeding 50 µM, indicating their low dark cytotoxicity. Upon irradiation with white light (400–700 nm, 10 mW cm−2, 30 min), cell viability decreased dramatically, with IC50,light ranging from 0.37 to 2.8 µM.
| Complex | Normoxia | CoCl2-induced hypoxia | ||||
|---|---|---|---|---|---|---|
| IC50,dark/µM | IC50,light/µM | Photocytotoxicity index | IC50,dark/µM | IC50,light/µM | Photocytotoxicity index | |
| 1 | >50 | 1.4 ± 0.1 | >36 | >50 | 1.6 ± 0.1 | >31 |
| 2 | >50 | 0.37 ± 0.07 | >135 | >50 | 0.74 ± 0.01 | >68 |
| 3 | >50 | 2.8 ± 0.1 | >18 | >50 | 3.0 ± 0.1 | >17 |
| 4 | >50 | 2.2 ± 0.1 | >23 | >50 | 2.5 ± 0.1 | >20 |
Considering the enhanced hypoxia tolerance of Type I PSs, the (photo)cytotoxicity of these complexes was further examined under CoCl2-induced hypoxia (Table 2, Fig. S15 and S16). Cells were pre-treated with 150 µM CoCl2 for 12 h before experiments to simulate a hypoxic environment. Similarly, the complexes had a negligible effect on cell viability in the dark (IC50,dark >50 µM). However, upon exposure to light, the viability of cells incubated with the rhodium(III) complexes significantly decreased. These dirhodamine-decorated rhodium(III) complexes exhibited remarkable photocytotoxicity with IC50,light values of 0.74–3.0 µM under mimicked hypoxia, resulting from the photoinduced ROS generation via the Type I pathway. Among these rhodium(III) complexes, complex 2 exhibited the highest photocytotoxicity indices (>135 and >68 under normoxia and CoCl2-induced hypoxia, respectively). Consequently, the photocytotoxicity of complex 2 was further investigated using Calcein-AM/propidium iodide (PI) assays (Fig. 5b). Upon incubation with complex 2 in the dark, the LSCM images of MCF-7 cells showed intense green fluorescence from Calcein-AM and minimal red emission from PI, revealing excellent biocompatibility of complex 2. Importantly, when cells were treated with complex 2 followed by light irradiation, the green fluorescence of Calcein-AM diminished, while the red emission from PI increased, illustrating a reduction in cell viability due to the superior photocytotoxicity of complex 2. In view of its mitochondrial localisation, the mitochondrial membrane potential (MMP) was evaluated using R123, the uptake of which depends on MMP. Upon incubation with complex 2, the cells exhibited strong green fluorescence in the mitochondria, indicating normal accumulation of R123 (Fig. 5c and S17). Upon exposure to white-light irradiation (400–700 nm, 10 mW cm−2) for 10 min, the emission from R123 significantly decreased, suggesting a reduction in MMP and mitochondrial dysfunction of the treated cells, ultimately leading to cell death.
Intracellular ROS photosensitisation
Given the effective ROS photosensitisation in aqueous solutions and the promising anticancer performance of complex 2, its intracellular ROS generation was investigated using DCFH-DA. As displayed in Fig. 5d, MCF-7 cells treated with complex 2 in the dark showed negligible green emission from DCF. However, cells exhibited strong DCF fluorescence upon white-light irradiation (400–700 nm, 10 mW cm−2) for 5 min, indicating significant ROS photosensitisation by complex 2 in cells. The intracellular photogenerated O2˙− and HO˙ were assessed using DHR123 and HPF, respectively, which showed similarly strong fluorescence in cells treated with complex 2 followed by light irradiation. Given that DHR123 is known to accumulate in mitochondria, the strong fluorescence observed in cells treated with complex 2 and DHR123 followed by light irradiation indicates mitochondrial O2˙− generation, which is consistent with the specific mitochondrial accumulation of complex 2 (Fig. 5a and S12). To further verify mtROS generation, the treated cells were costained with MitoTracker Deep Red. The strong overlap between the signals of DHR123 and the mitochondrial marker (Fig. S18) confirms that complex 2 specifically localised in the mitochondria and generated ROS in this organelle upon light irradiation, leading to mitochondrial dysfunction (vide supra). Since reduced O2-dependence is considered one of the advantages of Type I PSs, the ROS generation ability of complex 2 was also confirmed under CoCl2-induced hypoxic conditions (Fig. S19). Under these conditions, the cells treated with complex 2 and stained with DCFH-DA, DHR123 or HPF displayed distinct green fluorescence upon photoirradiation, confirming efficient Type I ROS photosensitisation by complex 2. These results demonstrate that enhanced intracellular O2˙− and HO˙ photosensitisation during the complex 2-based PDT process contributes to its high efficiency in inhibiting cancer cell proliferation.Induction of cell pyroptosis
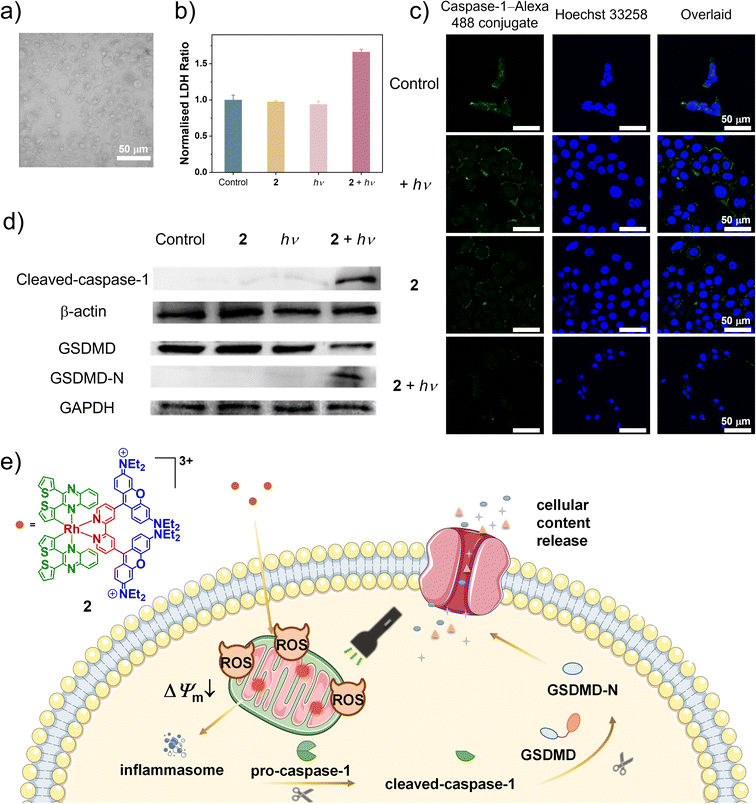
Mitochondrial dysfunction can disrupt cellular redox homeostasis and energy metabolism, resulting in cell death.57 Motivated by the generation of mtROS and the excellent anticancer performance of complex 2, the underlying mechanism of cell death was further explored. The effects of various cell death pathway inhibitors on the viability of MCF-7 cells treated with complex 2 followed by photoirradiation were evaluated to preliminarily investigate the cell death mechanism (Fig. S20). Ferrostatin-1 (Fer-1; an inhibitor of ferroptosis), z-VAD-fmk (an inhibitor of apoptosis) and necrostatin-1 (Nec-1; an inhibitor of necrosis) were ineffective in mitigating cell death. However, pretreatment of the cells with necrosulfonamide (NSA; an inhibitor of pyroptosis) led to a notable increase in cell viability from 39% to 57%, suggesting pyroptosis as the primary cell death pathway. The bright-field LSCM image shows that cells treated with complex 2 and photoirradiation exhibited swelling and membrane blebbing, characteristic features of pyroptosis (Fig. 6a). As an important indicator of intracellular content leakage, the release of lactate dehydrogenase (LDH) into the cell culture medium was further studied.58 Notably, cells treated with complex 2 and light displayed higher extracellular LDH levels compared to other treatment groups (Fig. 6b). It has been reported that ROS can activate the NOD-like receptor protein 3 (NLRP3) inflammasome, which subsequently promotes the activation and cleavage of caspase-1, ultimately leading to the cleavage of GSDMD and induction of pyroptosis.59–61 Therefore, the expression of caspase-1 was studied using an immunofluorescence staining assay. As depicted in Fig. 6c, when the cells were treated with complex 2 and light irradiation, they exhibited a significant decrease in fluorescence from caspase-1–Alexa 488, illustrating the cleavage of pro-caspase-1. Then, Western blot assays were performed to study the expression of cleaved caspase-1, full-length GSDMD and N-terminal GSDMD-N, which are the crucial executors to mediate pyroptosis. Cells treated with both complex 2 and light irradiation exhibited significantly elevated expression levels of cleaved-caspase-1 (Fig. 6d). Concurrently, the expression levels of GSDMD decreased, while GSDMD-N expression increased compared to other groups, confirming that GSDMD was cleaved by activated caspase-1, thereby inducing pyroptosis. These results collectively indicate that mtROS-induced, GSDMD-mediated pyroptosis is the predominant mechanism of photoinduced cell death for complex 2. As illustrated in Fig. 6e, complex 2 primarily targeted mitochondria and, upon photoirradiation, generated ROS. The elevated mtROS induced MMP loss and mitochondrial dysfunction, which serve as upstream triggers for inflammasome priming. This drives the cleavage of pro-caspase-1 to its active N-terminal fragment, which subsequently cleaves GSDMD to its pore-forming N-terminal fragment, GSDMD-N. The resulting membrane pore formation causes the release of LDH and other intracellular contents, indicative of pyroptosis.Conclusions
In summary, a series of cyclometallated rhodium(III) complexes incorporating two rhodamine units was designed and evaluated for their potential as bioimaging and phototherapeutic agents. These complexes showed intense absorption at 575 nm and moderate rhodamine fluorescence at 600 nm in solution upon photoexcitation. Originating from the long-lived rhodamine 3IL excited states, these complexes exhibited efficient O2˙− and HO˙ photosensitisation, likely via electron transfer from the rhodamine radical cation and anion generated via SBCT within the excited complexes. Cellular studies revealed that these rhodium(III) complexes localised in mitochondria and exhibited high photocytotoxicity under both normoxic and CoCl2-induced hypoxic conditions, with complex 2 showing the highest photocytotoxicity index. Further investigation demonstrated that complex 2 efficiently photosensitised substantial ROS generation within cellular environments, contributing to its high therapeutic efficacy. This ROS generation led to mitochondrial dysfunction, ultimately triggering cell death through GSDMD-mediated pyroptosis. We believe these findings offer valuable insights into the development of luminescent and photofunctional cyclometallated rhodium(III) complexes as promising bioimaging and phototherapeutic agents.Author contributions
K. G.-M. J.: conceptualisation, data curation, formal analysis, investigation, writing – original draft, writing – review & editing; G.-X. X.: conceptualisation, data curation; L. C.-C. L.: formal analysis, writing – original draft; F. W.: conceptualisation, data curation; S. W.: data curation; K. M.-C. W.: conceptualisation, funding acquisition, project administration, resources, supervision, writing – review & editing; K. K.-W. L.: conceptualisation, funding acquisition, project administration, resources, supervision, writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
CCDC 2469873 (1) contains the supplementary crystallographic data for this paper.62Supplementary information (SI): instrumentation, methods, synthetic details, characterisation, analytical and photophysical data and results of cellular studies, microscopy experiments and bioassays. See DOI: https://doi.org/10.1039/d5sc09339b.
Acknowledgements
We thank the Hong Kong Research Grants Council (project no. CityU 11317022, CityU 11309423, CityU 11304524 and C7075-21GF), the Hong Kong Research Grants Council and National Natural Science Foundation of China (project no. N_CityU104/21), the National Natural Science Foundation of China (grant no. 21771099) and the Science, Technology and Innovation Commission of Shenzhen Municipality (grant no. JCYJ20190809165411528) for financial support. We also acknowledge funding support from the “Laboratory for Synthetic Chemistry and Chemical Biology” under the Health@InnoHK Programme launched by Innovation and Technology Commission, the Government of Hong Kong SAR, P. R. China. We are also grateful to Dr Jie Wang for his help in biological studies. K. G.-M. J. acknowledges the receipt of a Postgraduate Studentship administered by City University of Hong Kong and Southern University of Science and Technology.References
- D. Chen, Q. Xu, W. Wang, J. Shao, W. Huang and X. Dong, Small, 2021, 17, 2006742 CrossRef CAS PubMed.
- J.-J. Hu, Y. Chen, X. Lou, F. Xia, X. Wu and J. Yoon, Coord. Chem. Rev., 2025, 532, 216526 CrossRef CAS.
- Y. Wan, L.-H. Fu, C. Li, J. Lin and P. Huang, Adv. Mater., 2021, 33, 2103978 CrossRef CAS.
- S. Kwiatkowski, B. Knap, D. Przystupski, J. Saczko, E. Kędzierska, K. Knap-Czop, J. Kotlińska, O. Michel, K. Kotowski and J. Kulbacka, Biomed. Pharmacother., 2018, 106, 1098 CrossRef.
- L. C.-C. Lee and K. K.-W. Lo, Small Methods, 2024, 8, 2400563 CrossRef CAS PubMed.
- T. C. Pham, V.-N. Nguyen, Y. Choi, S. Lee and J. Yoon, Chem. Rev., 2021, 121, 13454 CrossRef CAS PubMed.
- L. Conti, E. Macedi, C. Giorgi, B. Valtancoli and V. Fusi, Coord. Chem. Rev., 2022, 469, 214656 CrossRef CAS.
- X. Zhang, Y. Hou, X. Xiao, X. Chen, M. Hu, X. Geng, Z. Wang and J. Zhao, Coord. Chem. Rev., 2020, 417, 213371 CrossRef CAS.
- L. K. McKenzie, H. E. Bryant and J. A. Weinstein, Coord. Chem. Rev., 2019, 379, 2 CrossRef CAS.
- Y. Wu, S. Li, Y. Chen, W. He and Z. Guo, Chem. Sci., 2022, 13, 5085 RSC.
- X. Zhao, J. Liu, J. Fan, H. Chao and X. Peng, Chem. Soc. Rev., 2021, 50, 4185 RSC.
- M.-M. Wang, F.-J. Xu, Y. Su, Y. Geng, X.-T. Qian, X.-L. Xue, Y.-Q. Kong, Z.-H. Yu, H.-K. Liu and Z. Su, Angew. Chem., Int. Ed., 2022, 61, e202203843 CrossRef CAS PubMed.
- X. Guo, N. Yang, W. Ji, H. Zhang, X. Dong, Z. Zhou, L. Li, H.-M. Shen, S. Q. Yao and W. Huang, Adv. Mater., 2021, 33, 2007778 CrossRef CAS.
- H. Shigemitsu, K. Sato, S. Hagio, Y. Tani, T. Mori, K. Ohkubo, Y. Osakada, M. Fujitsuka and T. Kida, ACS Appl. Nano Mater., 2022, 5, 14954 CrossRef CAS.
- X. Geng, Y. Sun, Z. Li, R. Yang, Y. Zhao, Y. Guo, J. Xu, F. Li, Y. Wang, S. Lu and L. Qu, Small, 2019, 15, 1901517 CrossRef CAS PubMed.
- S. Li, H. Yuan, X.-Z. Yang, X. Xu, W. Yu, Y. Wu, S. Yao, J. Xie, W. He, Z. Guo and Y. Chen, ACS Cent. Sci., 2025, 11, 441 CrossRef CAS PubMed.
- G. Kumari, A. Gupta, R. K. Sah, A. Gautam, M. Saini, A. Gupta, A. K. Kushawaha, S. Singh and P. K. Sasmal, Adv. Healthcare Mater., 2023, 12, 2202411 CrossRef CAS.
- Q. Chen, C. Jin, X. Shao, R. Guan, Z. Tian, C. Wang, F. Liu, P. Ling, J.-L. Guan, L. Ji, F. Wang, H. Chao and J. Diao, Small, 2018, 14, 1802166 CrossRef.
- Y.-B. Peng, C. Tao, C.-P. Tan and P. Zhao, J. Inorg. Biochem., 2021, 218, 111400 CrossRef CAS PubMed.
- Y.-B. Peng, W. He, Q. Niu, C. Tao, X.-L. Zhong, C.-P. Tan and P. Zhao, Dalton Trans., 2021, 50, 9068 RSC.
- S. Luo, X. Tan, S. Fang, Y. Wang, T. Liu, X. Wang, Y. Yuan, H. Sun, Q. Qi and C. Shi, Adv. Funct. Mater., 2016, 26, 2826 Search PubMed.
- F. Ding, J. Liu, K. Ai, C. Xu, X. Mao, Z. Liu and H. Xiao, Adv. Mater., 2024, 36, 2306419 CrossRef CAS.
- K. Peng, Y. Zheng, W. Xia and Z.-W. Mao, Chem. Soc. Rev., 2023, 52, 2790 RSC.
- J. Zhuang, S. Song, L. Yang, Q. Pan, Y. He, L. Huo, N. Li and N. Zhao, Adv. Healthcare Mater., 2025, 14, 2501346 Search PubMed.
- Y. Ye, K. Ren, Y. Dong, L. Yang, D. Zhang, Z. Yuan, N. Ma, Y. Song, X. Huang and H. Qiao, ACS Appl. Mater. Interfaces, 2023, 15, 26285 CrossRef CAS PubMed.
- W. Zhang, Z. Liu, J. Zhu, Z. Liu, Y. Zhang, G. Qin, J. Ren and X. Qu, J. Am. Chem. Soc., 2023, 145, 16658 CrossRef CAS PubMed.
- Y. Peng, R. Mo, M. Yang, H. Xie, F. Ma, Z. Ding, S. Wu, J. W. Y. Lam, J. Du, J. Zhang, Z. Zhao and B. Z. Tang, ACS Nano, 2024, 18, 26140 CAS.
- K. M. Boyle and J. K. Barton, J. Am. Chem. Soc., 2018, 140, 5612 CrossRef CAS PubMed.
- A. Nano, J. Dai, J. M. Bailis and J. K. Barton, Biochemistry, 2021, 60, 2055 CrossRef CAS PubMed.
- R. J. Ernst, H. Song and J. K. Barton, J. Am. Chem. Soc., 2009, 131, 2359 CrossRef CAS.
- A. C. Komor and J. K. Barton, J. Am. Chem. Soc., 2014, 136, 14160 CrossRef CAS PubMed.
- G.-J. Yang, H.-J. Zhong, C.-N. Ko, S.-Y. Wong, K. Vellaisamy, M. Ye, D.-L. Ma and C.-H. Leung, Chem. Commun., 2018, 54, 2463 RSC.
- D.-L. Ma, L.-J. Liu, K.-H. Leung, Y.-T. Chen, H.-J. Zhong, D. S.-H. Chan, H.-M. D. Wang and C.-H. Leung, Angew. Chem., Int. Ed., 2014, 53, 9178 CrossRef CAS PubMed.
- G.-J. Yang, W. Wang, S. W. F. Mok, C. Wu, B. Y. K. Law, X.-M. Miao, K.-J. Wu, H.-J. Zhong, C.-Y. Wong, V. K. W. Wong, D.-L. Ma and C.-H. Leung, Angew. Chem., Int. Ed., 2018, 57, 13091 CrossRef CAS.
- F. Wei, S.-L. Lai, S. Zhao, M. Ng, M.-Y. Chan, V. W.-W. Yam and K. M.-C. Wong, J. Am. Chem. Soc., 2019, 141, 12863 CrossRef CAS.
- M. Beija, C. A. M. Afonso and J. M. G. Martinho, Chem. Soc. Rev., 2009, 38, 2410 RSC.
- H. N. Kim, M. H. Lee, H. J. Kim, J. S. Kim and J. Yoon, Chem. Soc. Rev., 2008, 37, 1465 RSC.
- Y. Wang, X. Wang, W. Ma, R. Lu, W. Zhou and H. Gao, Chemosensors, 2022, 10, 399 CrossRef CAS.
- F. Wei, F. Chen, S. Wu, M. Zha, J. Liu, K.-L. Wong, K. Li and K. M.-C. Wong, Inorg. Chem., 2024, 63, 5872 CrossRef CAS PubMed.
- D. Shi, X. Zhang, S. Wu, L. Wei, K. Li and K. M.-C. Wong, Inorg. Chem., 2025, 64, 20229 CrossRef CAS.
- K. G.-M. Jiang, F. Wei, P. K.-K. Leung, S. Wu, K. K.-W. Lo and K. M.-C. Wong, Chem. Sci., 2025, 16, 22127 RSC.
- A. L. Harris, Nat. Rev. Cancer, 2002, 2, 38 CrossRef CAS PubMed.
- L. C.-C. Lee and K. K.-W. Lo, J. Am. Chem. Soc., 2022, 144, 14420 CrossRef CAS PubMed.
- X. Li, F. Han, X. Zhou, H. Zhang, T. Shi, L. Cai, D. Zhou, W. Chi, S. Long, W. Sun, J. Du, J. Fan and X. Peng, Aggregate, 2025, 6, e70208 CrossRef CAS.
- B. Lu, L. Wang, H. Tang and D. Cao, J. Mater. Chem. B, 2023, 11, 4600 RSC.
- T. Xiong, Y. Chen, M. Li, X. Chen and X. Peng, Small, 2025, 21, 2501911 CrossRef CAS.
- C. Liu, L. Zhou, F. Wei, L. Li, S. Zhao, P. Gong, L. Cai and K. M.-C. Wong, ACS Appl. Mater. Interfaces, 2019, 11, 8797 CrossRef CAS PubMed.
- L. Zhou, F. Wei, J. Xiang, H. Li, C. Li, P. Zhang, C. Liu, P. Gong, L. Cai and K. M.-C. Wong, Chem. Sci., 2020, 11, 12212 RSC.
- K. Chen, Y. Dong, X. Zhao, M. Imran, G. Tang, J. Zhao and Q. Liu, Front. Chem., 2019, 7, 821 CrossRef CAS PubMed.
- F. Stracke, M. Heupel and E. Thiel, J. Photochem. Photobiol., A, 1999, 126, 51 CrossRef CAS.
- Y. Yue, B. Li, D. Wang, C. Wu, Z. Li and B. Liu, Adv. Funct. Mater., 2025, 35, 2414542 CrossRef CAS.
- M. T. Whited, N. M. Patel, S. T. Roberts, K. Allen, P. I. Djurovich, S. E. Bradforth and M. E. Thompson, Chem. Commun., 2012, 48, 284 RSC.
- A. Yahagh, R. R. Kaswan, S. Kazemi, P. A. Karr and F. D'Souza, Chem. Sci., 2024, 15, 906 RSC.
- J. H. Golden, L. Estergreen, T. Porter, A. C. Tadle, M. R. D. Sylvinson, J. W. Facendola, C. P. Kubiak, S. E. Bradforth and M. E. Thompson, ACS Appl. Energy Mater., 2018, 1, 1083 CrossRef CAS.
- C. Trinh, K. Kirlikovali, S. Das, M. E. Ener, H. B. Gray, P. Djurovich, S. E. Bradforth and M. E. Thompson, J. Phys. Chem. C, 2014, 118, 21834 CrossRef CAS PubMed.
- Y. Liu, J. Zhao, A. Iagatti, L. Bussotti, P. Foggi, E. Castellucci, M. Di Donato and K.-L. Han, J. Phys. Chem. C, 2018, 122, 2502 CrossRef CAS.
- M. D. Yaqoob, L. Xu, C. Li, M. M. L. Leong and D. D. Xu, Photodiagnosis Photodyn. Ther., 2022, 38, 102830 CrossRef CAS PubMed.
- M. Wu, X. Liu, H. Chen, Y. Duan, J. Liu, Y. Pan and B. Liu, Angew. Chem., Int. Ed., 2021, 60, 9093 CrossRef CAS PubMed.
- Y. Chen, Y. Zhang, N. Li, Z. Jiang and X. Li, Inflamm. Res., 2023, 72, 829 CrossRef CAS PubMed.
- S. Paik, J. K. Kim, H. J. Shin, E.-J. Park, I. S. Kim and E.-K. Jo, Cell. Mol. Immunol., 2025, 22, 563 CrossRef CAS PubMed.
- S. Paik, J. K. Kim, P. Silwal, C. Sasakawa and E.-K. Jo, Cell. Mol. Immunol., 2021, 18, 1141 Search PubMed.
- CCDC 2469873: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2nx3b1.
| This journal is © The Royal Society of Chemistry 2026 |