Open Access Article
Open Access ArticleRecent advances in site-specific modifications of peptides and proteins
Xin-Rong
Duan
a,
Meng-Qian
Zhang
a,
Geng-Hui
Feng
a and
Yan-Mei
Li
 *abc
*abc
aKey Lab of Bioorganic Phosphorus Chemistry & Chemical Biology, Department of Chemistry, Tsinghua University, Beijing 100084, China. E-mail: liym@mail.tsinghua.edu.cn
bBeijing Institute for Brain Disorders, Beijing 100069, China
cCenter for Synthetic and Systems Biology, Tsinghua University, Beijing 100084, China
First published on 8th April 2026
Abstract
Modification at specific sites has long been a central goal in peptide and protein chemistry. Introducing functional groups at specific sites is an ideal state for protein function research and the modification of protein properties, as it can eliminate the interference caused by heterogeneous modifications. Recent downstream applications have placed even greater demands on site selectivity. This review examines the latest advances in the site-specific modification of peptides and proteins, organizing the reactions into four categories: exploiting disparities in chemical environments, tag-and-modify approaches, proximity-driven chemical modification, and enzymatic strategies. For each reaction, we detail the underlying design rationale and highlight downstream chemical and biological applications. Collectively, these methods have achieved remarkable gains in terms of specificity, efficiency, and scope, furnishing a versatile chemical toolbox for the site-selective functionalization of peptides and proteins. We conclude by summarizing the current state of the field and outlining prospective directions for future development.
1 Introduction
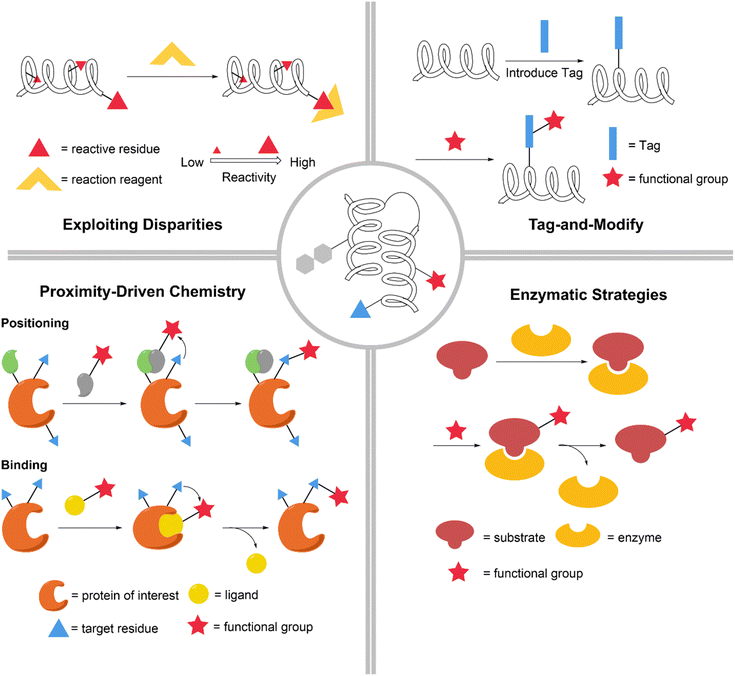
Chemical modification is a powerful tool for engineering peptides and proteins, greatly expanding their functional diversity.1–4 Site-specific modifications deliver precisely engineered products whose structures can be tailored on demand, enabling exact control over peptide and protein architecture and function and thereby deepening our understanding of structure–function relationships.5,6 Introduced functional handles, such as bioorthogonal reaction groups, enable further modification of peptides and proteins, allowing their multifunctional engineering for diverse downstream applications, including cellular imaging, omics profiling, immunotherapy7 and the design of antibody–drug conjugates.8,9 From a different perspective, the rapid evolution of these applications continually raises the bar for site selectivity.10 More precise modification translates into the more effective realization of the intended functions. Together, these factors underscore the vast potential of site-specific peptide and protein modifications.In recent years, a wealth of new site-specific modification reactions have emerged, resulting in the assembly of a rich chemical toolbox.11 Leveraging innate disparities in nucleophilicity and redox properties, researchers have developed reactions that target the N-terminus, C-terminus,12 and even specific internal residues.13 Exogenous tags can be appended to introduce new addressability, enabling direct and specific modification.14 Residue-mediated or ligand-directed positioning proximity can trigger nearby functionalization. In addition to chemistry, enzymes that recognize specific sites offer another layer of selectivity (Fig. 1). In summary, the landscape of site-specific modifications is replete with advances that merit a comprehensive review.
This review provides an up-to-date, integrated survey of recent site-specific modification reactions for peptides and proteins, with a particular emphasis on their chemical and biological applications. We also speculate on future directions, aiming to guide and accelerate the next wave of discovery.
2 Exploiting disparities in chemical environments
The same functional groups do not always exhibit the same reaction characteristics because they are affected by the chemical environment. When a specific site of a protein is chemically modified, a natural approach is to exploit differences in the chemical environment within the protein. For instance, the pKa of amines in different chemical environments varies, which in turn affects their nucleophilicity and reactivity.15,16 The C-terminal carboxyl group also exhibits distinct oxidative reactivity compared to the carboxyl groups on amino acid side chains.17 Researchers have used disparities in chemical environments to develop a variety of chemical modifications. We divide these methods into differences in nucleophilicity and oxidation–reduction properties.2.1 Differences in nucleophilicity
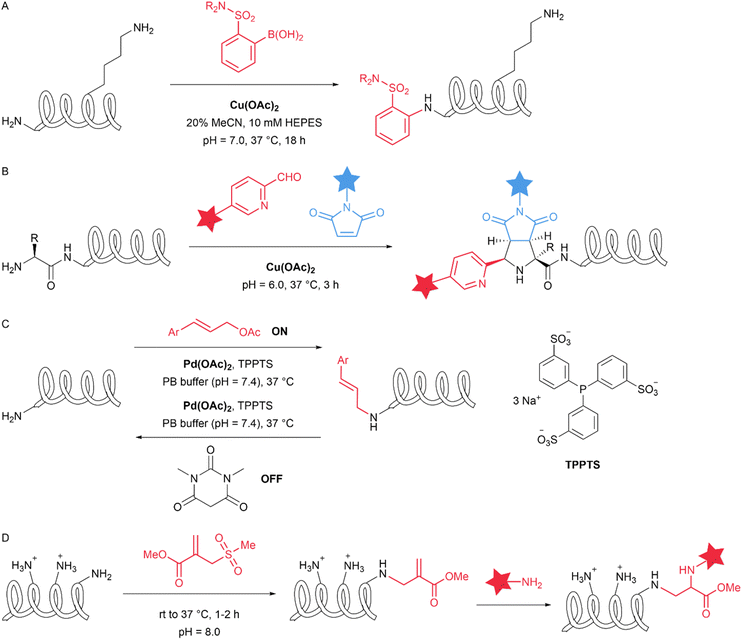
Beyond experimental strategies, computational and bioinformatic tools have emerged as valuable resources for predicting site-specific reactivity from protein sequences. Machine learning algorithms trained on structural and reactivity databases can now identify reactive hotspots and subtle electronic environment differences among identical amino acid residues. Cysteine exhibits strong nucleophilic reactivity and serves diverse functions within proteins. Predicting the reactivity of cysteine residues facilitates the annotation of protein functions. In 2012, Anandakrishnan et al. reported a tool that can predict the pKa values of cysteine thiol groups, which in turn can be used to estimate their relative nucleophilic reactivity.21 In 2015, Soylu and Marino developed an algorithm termed HAL-Cy and implemented this algorithm in a web service (Cy-preds). Cy-preds is capable of performing calculations for comprehensive analysis of Cys reactivity in proteins, encompassing reactivity predictions and functional characterization.22 In 2017, Wang et al. reported the development of a sequence-based computational algorithm named sbPCR (sequence-based prediction of cysteine reactivity). This method enables to predict cysteines with hyper-reactivity based on only local sequence features and provides a large inventory of potential hyper-reactive cysteines in proteomes.23 The ε-amino groups of different lysine residues on a protein exhibit subtle yet intrinsic differences in reactivity, thereby indicating the possibility of site-selective modification. The pKa value of the lysine amino group can also be predicted for site-selective reactions. In 2023, Wilson et al. developed a theoretical computational method for predicting the pKa values of lysine residues in proteins.24 The computational approaches complement chemical intuition and experimental screening, potentially accelerating the rational design of site-specific modification strategies. In 2018, Matos et al. computationally predicted and designed sulfonyl acrylate reagents for the targeted modification of a single lysine residue. Under mildly alkaline conditions, the kinetically favoured reaction occurs at the lysine with the lowest pKa, while exhibiting chemoselectivity in the presence of other nucleophilic residues (Fig. 2D). A transient hydrogen bond forms between the sulfonyl group of the reagent and the ε-amino group of the lysine side chain, enabling the key N–C bond to be forged via a low-energy, H-bond-assisted chair-like addition transition state. Subsequent elimination of the sulfonyl group then furnishes an additional thermodynamic driving force for the reaction. The authors achieved site-specific lysine modification in five different proteins. The resulting acrylate-bearing proteins were then conjugated in a chemoselective and regioselective manner with synthetic amine-containing cargoes, including PEG, fluorophores, and anticancer drugs, via a subsequent Michael addition.15
| Modified residue | Reagent | Highest yield (%) | Reference |
|---|---|---|---|
| N-terminal amino group | Arylboronic acid | 97 | 18 |
| Maleimides/2-pyridinecarbaldehyde | 99 | 19 | |
| Allyl ester | 98 | 20 | |
| Lysine | Sulfonyl acrylate | 98 | 15 |
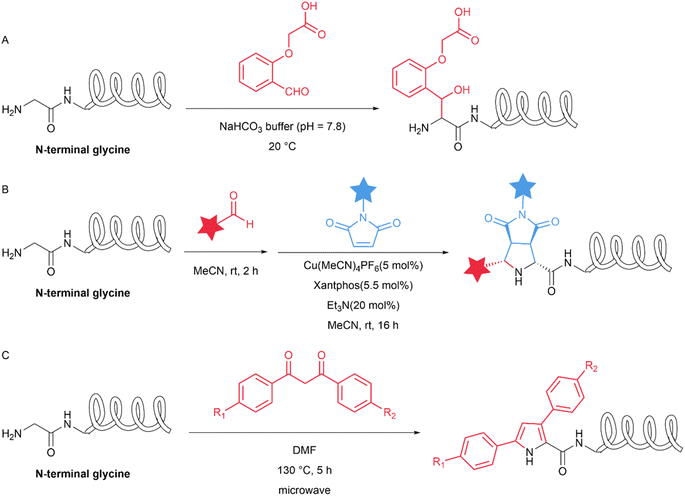
| N-terminal glycine | Ortho-substituted benzaldehyde | 71 | 25 |
| Aldehyde/maleimide | 99 | 26 | |
| Dibenzoylmethane | 76 | 27 | |
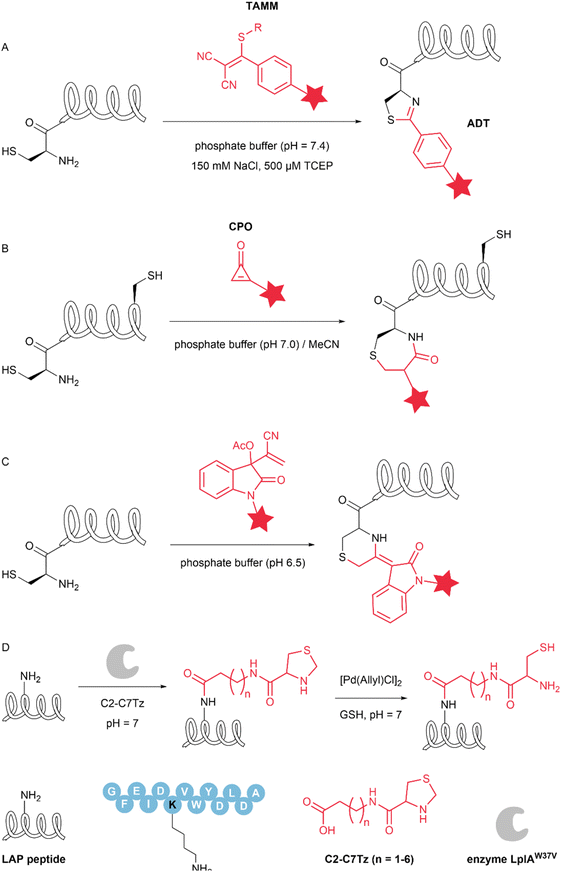
| 1,2-Aminothiol | 2-((Alkylthio)(aryl)methylene) malononitrile | >90 | 28 |
| Cyclopropenone | 88 | 29 | |
| Isatin-derived Baylis–Hillman adducts | 95 | 30 |
2.2 Differences in oxidation–reduction properties
| Modified residue | Reagent | Highest yield (%) | Reference |
|---|---|---|---|
| C-terminal carboxyl group | α,β-Unsaturated carbonyl | 95 | 17 |
| Ethynylbenziodoxolone | >95 | 32 | |
| N-terminal amino group | Ortho-quinone | 91 | 35 |
| Methionine | α,β-Unsaturated carbonyl | 96 | 36 |
| Tyrosine | Phenoxazine dialdehyde | 95 | 37 |
| Tryptophan | Biaryl N-carbamoyl pyridinium salt | 95 | 39 |
3. Tag-and-modify
In general, the inherent differences in the properties of peptide and protein substrates are limited. Relatively few examples of site-specific modification reactions relying on this difference are available. The artificial introduction of a positioning factor, which is referred to as a “Tag”, is necessary to achieve more precise and controllable modifications.1 This “Tag” can involve either altering a single amino acid with an unnatural amino acid or adding a new amino acid sequence. The introduction of unnatural amino acids allows for the precise targeting and modification of proteins at the desired site.14 Special amino acid sequences can provide unique targeting properties. We divided this part into natural amino acid derivatives, the insertion of unnatural amino acids and the insertion of amino acid sequences.3.1 Incorporation of natural amino acid derivatives
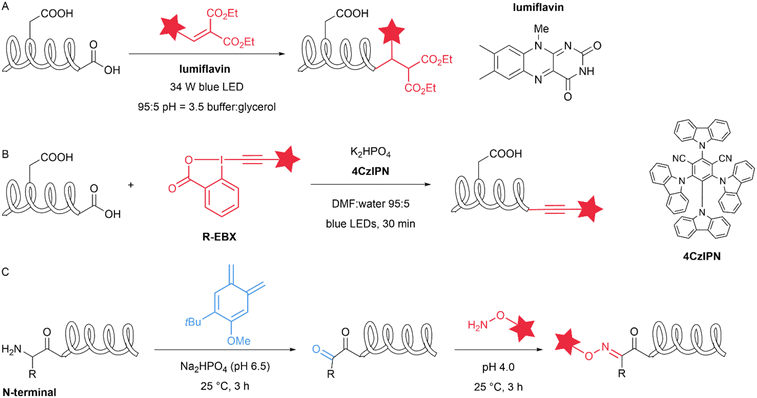
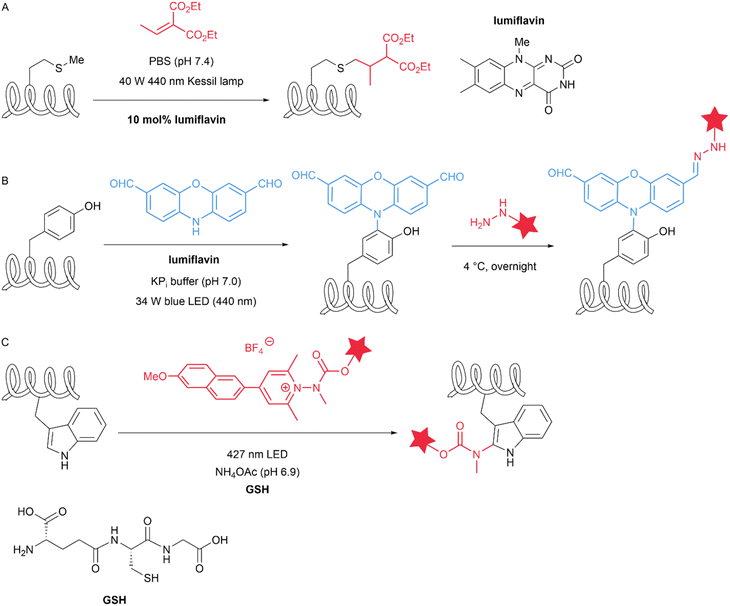
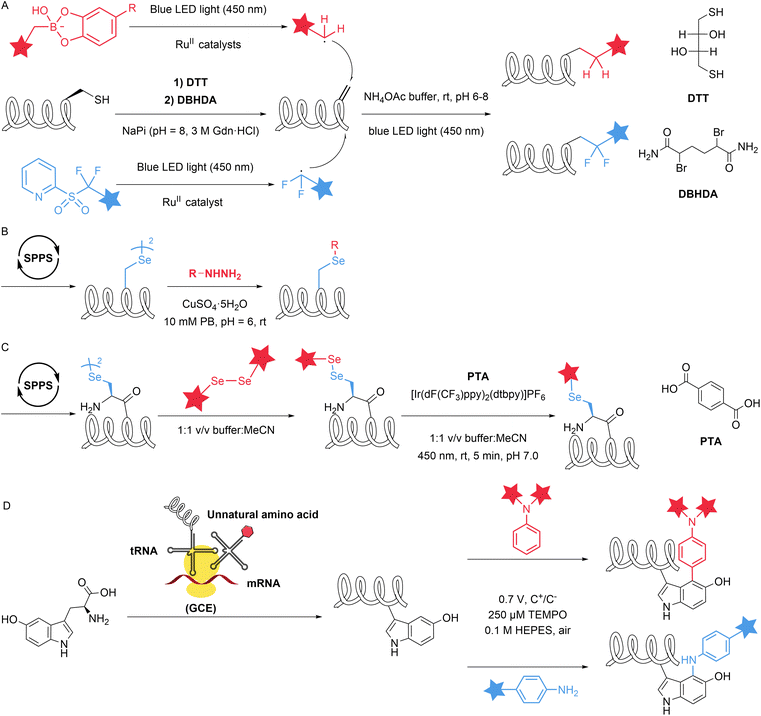
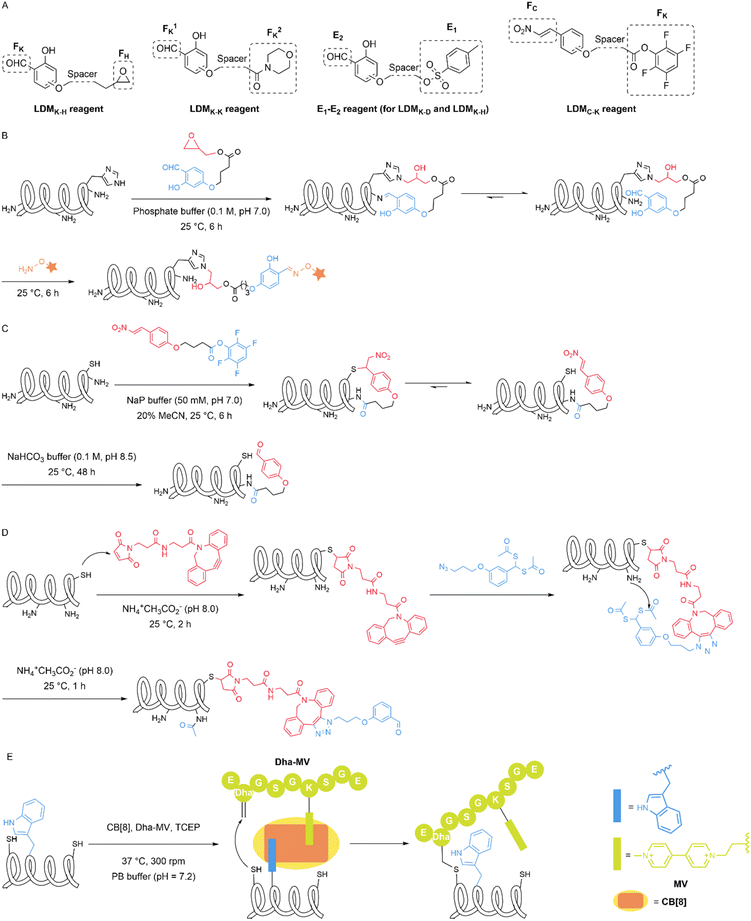
Natural amino acid derivatives exhibit only subtle differences in structure and reactivity compared with their native counterparts, yet these distinctions are sufficient for the derivatives to serve as tags that mediate site-specific modification reactions. Dehydroalanine (Dha) can be easily introduced into proteins through chemical reactions. Cysteine can react with 2,5-dibromohexanediamide (DBHDA) via desulfurization to form Dha. Its structural feature of containing an α,β-unsaturated double bond enables a variety of transformations, including nucleophilic addition, radical addition and cycloaddition.40 In 2020, Josephson et al. described the visible-light-driven installation of side chains at dehydroalanine residues in proteins through the formation of carbon-centred radicals that allow C–C bond formation in water. The authors used boronic acid catechol ester derivatives to generate RH2C˙ radicals and pyridylsulfonyl derivatives containing Fe(II) were used to generate RF2C˙ radicals. These radicals can react with dehydroalanine at specific sites to form β-CH2–γ-CH2 or β-CH2–γ-CF2 linkages, thereby incorporating a wide range of functional groups (more than 50 unique residues/side chains) into diverse protein scaffolds and sites (Fig. 7A).41 Selenocysteine is an isostere of cysteine (Cys) with notable differences in properties, including a lower pKa, high oxidation potential and nucleophilicity, relative to those of Cys. The differences in reactive properties make selenocysteine an ideal site for site-selective modification. In 2021, Zhao et al. introduced selenocysteine into peptides and proteins through solid-phase peptide synthesis (SPPS) methods, utilizing its reactions with alkyl/aryl radicals for the modification of peptides and proteins. The authors rapidly generated radicals in situ from hydrazine substrates and copper ions in an aqueous buffer at near-neutral pH values of 5–8 (Fig. 7B). This reaction is minimally affected by cysteine and methionine and involves the intramolecular cyclization of peptides and biotinylation of proteins.42 Similarly, in 2022, Dowman et al. developed a photocatalytic method using selenocysteine called photocatalytic diselenide contraction (PDC). This reaction occurs upon the irradiation at 450![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) nm of an iridium photocatalyst and a phosphine and results in rapid and clean conversion of diselenides to reductively stable selenoethers (Fig. 7C). The authors achieved dimerization of selenopeptides, late-stage modification of calmodulin (K148U) diselenide and site-specific C-terminal functionalization of ubiquitin diselenide via a PDC reaction.43
nm of an iridium photocatalyst and a phosphine and results in rapid and clean conversion of diselenides to reductively stable selenoethers (Fig. 7C). The authors achieved dimerization of selenopeptides, late-stage modification of calmodulin (K148U) diselenide and site-specific C-terminal functionalization of ubiquitin diselenide via a PDC reaction.43
Compared with canonical aromatic amino acids, the noncanonical amino acid 5-hydroxytryptophan (5HTP) possesses unique reactive properties and has substantially lower oxidation potential. Electrochemical methods, with their precise control over fundamental reaction parameters, mild reaction conditions, and ability to generate reactive intermediates in situ, have become attractive strategies for the chemoselective modification of proteins. In 2023, Loynd et al. reported an electrochemically promoted coupling reaction between 5-hydroxytryptophan and simple aromatic amines called electrochemical labelling of hydroxyindoles with chemoselectivity (eCLIC). This method selectively converts 5HTP into a reactive electrophile, enabling it to couple with nucleophilic aromatic amines that are inherently unreactive towards proteins (Fig. 7D). The authors used this method to achieve the site-specific modification of several different proteins, including full-length antibodies. The reaction is mutually compatible with the azide-alkyne cycloaddition, as well as the reaction between strained alkenes and tetrazines. The authors utilized this feature to express a protein containing 5HTP and azide-ncAA residues, successfully generating a homogeneous dual-labelled conjugate.44
3.2 Insertion of unnatural amino acids
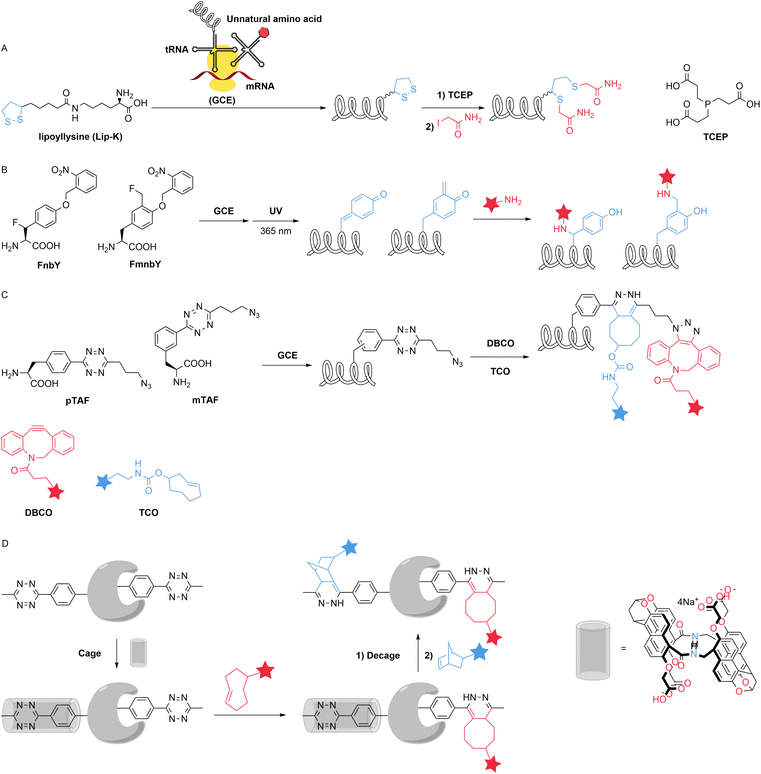
The development of the genetic code extension (GCE) technique has made the insertion of unnatural amino acids more convenient. Their incorporation creates subtle differences in reactivity. This property is discussed in Section 2.2.2. Unnatural amino acids can possess latent bioreactivity and react with target natural amino acids. This aspect will be discussed in Section 4.3. Unnatural amino acids can carry bioorthogonal handles such as alkynyl or azide groups. These groups have enabled the development of a variety of site-specific modification reactions and will be the focus of the following discussion. Introducing an unpaired cysteine thiol into proteins that already contain internal disulfide bonds without perturbing the existing disulfide network is extremely challenging. In 2020, Ling et al. used the genetic code expansion technique to introduce a surface-exposed self-paired di-thiol functional group into proteins, which can be selectively reduced to generate active thiols without disturbing the internal disulfide bond. The introduction of the dithiol-containing amino acid has no influence on protein activity or the disulfide network. The two thiol groups generated can be specifically modified by iodoacetamide (IAM) (Fig. 8A).45Quinone methides (QM) are excellent Michael acceptors for nucleophiles and have a high reaction rate. In 2020, Liu et al. introduced the amino acids FnbY and FmnbY, which contain photocaged quinone methides, into proteins using the genetic code expansion technique. Upon light-induced activation, these amino acids generate QM that enables rapid and selective conjugation with amine reagents (Fig. 8B). The authors used this method to install various functional groups onto proteins, offering advantages such as compatibility with low temperature and small linkages.46 Introducing two mutually orthogonal groups into a protein enables specific modifications of two different types. In 2023, Wang et al. designed the encodable noncanonical amino acids pTAF and mTAF that simultaneously contain azide and tetrazine reaction handles. TAFs are double bioorthogonal handles that can react with commercially available fluorophores, radioisotopes, PEGs, and drugs in one-pot (Fig. 8C). The authors can use protein dual conjugates to assess combinations of tumour diagnosis, image-guided surgery, and targeted therapy in mouse models. Furthermore, simultaneously incorporating mTAF and a ketone-containing noncanonical amino acid into one protein allows the preparation of a site-specific protein triconjugate.47 For the same type of unnatural amino acids, diverse functionalizations can be achieved by modulating their reactivity. In 2023, Cao et al. developed a supramolecular control system to reversibly regulate tetrazine reactivity through host caging. The authors synthesized naphthotubes and phenyltetrazine derivatives with high affinity and specificity as host–guest pairs, enabling the precise control of the unnatural amino acid-mediated reactions of tetrazine on demand (Fig. 8D). With respect to the tetrazine residues introduced at different sites in the proteins, the exposed residues could be reversibly caged, whereas the semiburied residues were resistant to caging and retained their reactivity. This approach allows for the preparation of dual site-specific protein conjugates and sequential labelling of dual proteins on living cells using the inverse electron demand Diels–Alder (IEDDA) reaction. This system can also be used to control IEDDA reactions in living animals.48
3.3 Insertion of amino acid sequences
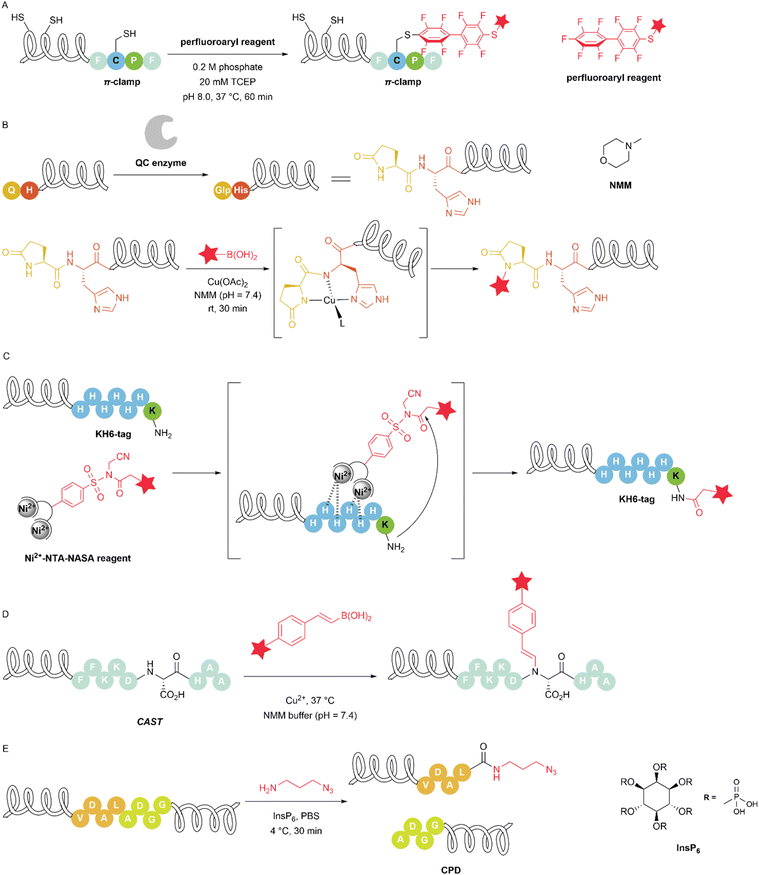
Natural proteins precisely control selective reactions and interactions through the construction of large three-dimensional structures. For example, enzymes have catalytic activity at specific sites, which is usually generated by their unique structures, resulting in the formation of a lock-and-key pairing pattern. In enzymes, particular amino acids are placed in a specialized active site environment. Similarly, researchers have designed a series of peptide sequences to adjust the reactivity of specific amino acids within the peptide sequences, thereby enabling modification reactions at specific sites. In 2016, Zhang et al. designed a four-amino-acid sequence (FCPF), which the authors call the ‘π-clamp’. The authors hypothesize that the aromatic side chains of Phe activate the cysteine thiol and enhance the interaction with the perfluoroaryl reagents used in the reaction (Fig. 9A). Moreover, Pro serves to position the Cys and Phe residues into a conformation that promotes the reaction. The authors use the π-clamp to selectively modify the cysteine in the π-clamp sequence within proteins containing multiple endogenous cysteine residues. The authors applied this method to modify antibodies and enzymes where traditional cysteine modification methods cannot readily be used. Antibodies modified using the π-clamp retained binding affinity to their targets, enabling the synthesis of site-specific antibody–drug conjugates for the selective killing of HER2-positive breast cancer cells.49 The π-clamp has been repeatedly exploited for protein modification and functionalization and has demonstrated good application prospects. In 2021, Gama-Brambila et al. inserted the π-clamp into Cas proteins (Cas9, dCas9, Cas12, and Cas13). FCPF-tagged proteins can be labelled with perfluoroarene carrying fluorescein in living cells and degraded by a perfluoroaromatic-functionalized PROTAC (PROTAC-FCPF). This strategy sufficiently suppresses the activity of Cas proteins, suggesting a wide range of applications of perfluoroaromatic-induced proximity in the regulation of stability, activity, and functionality of any FCPF-tagged protein.50 The generation of protein–protein conjugates plays an important role in advancing the fields of biotechnology and biopharmaceutical research. Posttranslational chemical modification is a powerful approach for obtaining diverse protein–protein conjugates. In 2022, Taylor et al. developed a method for preparing protein–protein conjugates through π-clamp-mediated cysteine arylation with pentafluorophenyl sulfonamide functional groups. The authors used this method to achieve the homodimerization and heterodimerization of two antibodies that have π-clamps near their C-termini, resulting in a 10–60-fold increase in binding. Even in the presence of solvent-exposed cysteine residues in the protein, the cysteine in the π-clamp is still selectively modified. The other cysteines can participate in subsequent small-molecule conjugation, thereby constructing molecule–protein–protein conjugates with complete control over the modification sites.51The limitations of traditional chemical modification methods in terms of site selectivity make fusion protein domains with unique reactivity widely valuable in protein conjugation. In addition to the π-clamp described above, researchers have developed a variety of amino acid sequences to achieve site-selective modification. In 2018, Ohata et al. discovered a dipeptide sequence, pyroglutamate–histidine (Glp–His), that can serve as the minimal tag for protein modification, featuring extremely rapid kinetics and excellent selectivity. The Glp–His sequence can be generated by the QH sequence using the glutaminyl cyclase (QC) enzyme. The use of the glutaminyl cyclase (QC) enzyme to generate specific Glp–His sequences in vivo is feasible and does not affect protein expression levels. The Glp–His dipeptide sequence exhibits remarkably fast and efficient copper-catalysed N–H coupling with boronic acids in 4-methylmorpholine (NMM) buffer, allowing specific and bioorthogonal functionalization at a natural posttranslational modification site (Fig. 9B).52 A peptide motif could specifically bind a metal ion that catalyses a conjugation reaction with the bound peptide, thus simultaneously achieving chemoselectivity and site specificity. The His tag is a very common and effective example, as it can coordinate with Ni2+ and Co2+. In 2021, Thimaradka et al. utilized a lysine in the short peptide sequence KH6 containing a His tag to react with Ni2+-NTA-tethered N-acyl-N-alkyl sulfonamide (NASA) reagent for the modification of the amino group of the lysine side chain. Upon the complexation of Ni2+-NTA with the H6 sequence, the lysine residue in the tag reacts with the NASA group to introduce a required group via the proximity effect (Fig. 9C). The authors characterized the properties of this reaction using model peptides and His-tagged enhanced green fluorescent protein (EGFP). The authors used this method to prepare a chemically functionalized nanobody and applied it to the imaging of membrane surface proteins in living cells.53 Similar “metal-binding targeting” strategies have been developed to achieve site-specific protein conjugation. In 2023, Guo et al. developed a copper-assisted sequence-specific conjugation tag (CAST) method to achieve rapid, site-specific chemical modification of the protein amide backbone with pinpoint accuracy. The authors screened peptide sequences to optimize the reaction rates of His-containing peptides capable of binding metal ions. Using scanning point mutation and preferred residue combinations for sequence optimization, the authors ultimately generated the peptide FFKKDDHAA, which has the highest reactivity with styrylboronic acid. A screen of boronic acid substrates revealed that styrene boronic acid exhibited the best reactivity (Fig. 9D). The authors inserted the CAST peptide at different locations in proteins, including the N-terminus, internal loop, and C-terminus, to efficiently and quantitatively modify a diverse range of proteins. The authors employed this method to prepare stable antibody–drug conjugates (ADCs) with potent efficacy both in vitro and in vivo. CAST conjugation is orthogonal to all currently reported chemical or enzymatic conjugation methods, enabling the preparation of dual-payload antibody–drug conjugates.54 Protease domains are frequently employed to increase the expression and purification of proteins of interest. Fusing the C-terminus of a protein with these unique domains facilitates site-specific modification by the corresponding enzymes. Unlike other tags, the cysteine protease domain (CPD) can be conveniently removed by a catalytic equivalent of inositol hexaphosphate (InsP6). In 2023, Zeng et al. fused a cysteine protease domain on the C-terminus of the protein of interest and triggered cysteine protease domain self-cleavage with InsP6 to achieve nonenzymatic C-terminal functionalization. CPD-induced C-terminal modification is compatible with diverse amine substrates and proteins (Fig. 9E). The authors leveraged this method to achieve the site-specific assembly of nanobody–drug conjugates and the construction of bifunctional antibodies (Table 3).55
| Tag type | Modified residue | Reagent | Reference |
|---|---|---|---|
| Natural amino acid derivative | Dehydroalanine | Boronic acid catechol ester/pyridylsulfonyl | 41 |
| Selenocysteine | Hydrazine substrate | 42 | |
| Selenocysteine | Diselenides | 43 | |
| 5-Hydroxytryptophan | Aromatic amines | 44 | |
| Unnatural amino acid | Lipoyllysine | Iodoacetamide | 45 |
| FnbY/FmnbY | Amine | 46 | |
| pTAF/mTAF | DBCO/TCO | 47 | |
| Phenyltetrazine | TCO | 48 | |
| Peptide sequence | FCPF | Perfluoroaryl | 49 |
| Glp–His | Boronic acid | 52 | |
| HHHHHHK | Ni2+-NTA-NASA | 53 | |
| FFKKDDHAA | Styrylboronic acid | 54 | |
| VDALADGG | InsP6 | 55 |
4 Proximity-driven chemical modification
Tags enable site-specific modification but typically restrict modifications to the introduced tag rather than allowing modification at native sites within the protein. By harnessing interactions near the protein's intrinsic modification site, the modifying agent can be confined to the vicinity of the target site, thereby increasing its local concentration. This approach exploits a local molarity effect to promote modification at a specific site, similar to the actions of enzymes. It can utilize either the amino acids adjacent to the modification site or the interactions with ligands to achieve site-specific modification. We refer to this strategy as the proximity-driven chemical modification approach. We divide these modifications into proximity-group-mediated modifications, ligand-directed modifications and affinity-peptide-directed modifications.4.1 Proximity-group-mediated modification
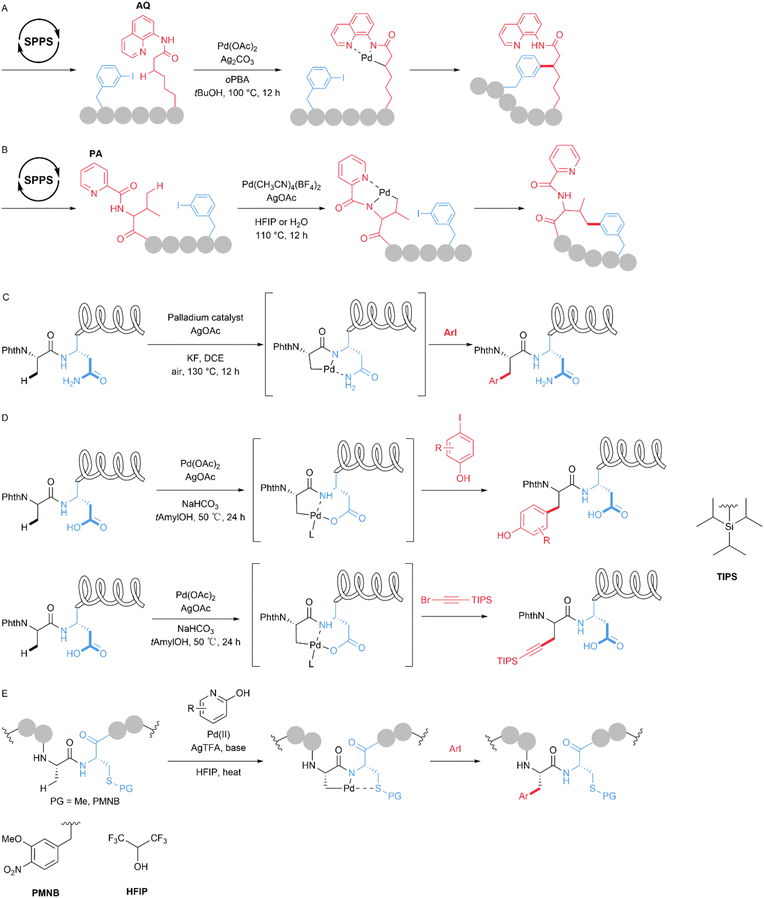
Unmodified side chains of natural amino acids can be harnessed to assist the functionalization of C(sp3)–H bonds in peptides. Asparagine, a natural amino acid bearing a primary amide side chain, can serve as a directing group. Its side chain and backbone can coordinate with palladium to form a bidentate Pd complex. In 2020, Weng et al. utilized the unmodified side chain of Asn combined with the backbone to form 5,6-fused bicyclic palladacycles that act as N,N-bidentate coordination sites, enabling the late-stage C(sp3)–H arylation of peptides (Fig. 11C). This method avoids the need to install and later remove exogenous directing groups. The authors demonstrated the coordinating role of the Asn primary amide and calculated the reaction mechanism. Using this approach, the authors achieved the arylation of Asn-containing di-, tri-, and tetrapeptides.64 In 2021, Chen et al. developed a late-stage, native Asp-directed C(sp3)–H arylation and alkynylation of peptide sequences containing L- and D-alanine (Ala), yielding a series of modified products, including tyrosine (Tyr) analogues, phenylalanine (Phe) analogues, and alkynylated Ala analogues. When conducted at a mild temperature (50 °C), the reaction delivers a variety of modified peptides without the need for expensive ligands or exogenous additives (Fig. 11D). Using this protocol, the authors successfully constructed strongly fluorescent (BODIPY)-labelled peptides.65 Transition metals tend to preferentially bind to peptide backbone amides rather than aliphatic side chains, leading to an inhibitory effect of the peptide backbone on side chain functionalization. The functionalization of aliphatic side chains at internal positions via amino acid coordination-assisted C(sp3)–H activation remains underdeveloped. In 2024, Hou et al. developed a highly site-selective β-C(sp3)–H arylation method for modifying the internal positions of both linear and cyclic peptides using S-alkyl cysteine as the directing group and 2-pyridone as the ligand for the Pd catalyst. The key to the success of the reaction lies in the formation of an N,S-bidentate directing group (Fig. 11E). The authors discovered that the protecting group on the sulfur atom is vital to preserve its coordinating ability and prevent side reactions. Based on this result, the authors developed a new protecting group, 4-methoxy-3-nitrobenzyl (PMNB), which promotes efficient C–H arylation and can be readily removed after peptide modification. The facile conversion of S-methyl cysteine to dehydroalanine (Dha) and the desulfurization of S-(p-nitrobenzyl)cysteine to alanine further expand the synthetic utility of the established C–H arylation strategy.66
4.2 Ligand-directed modification
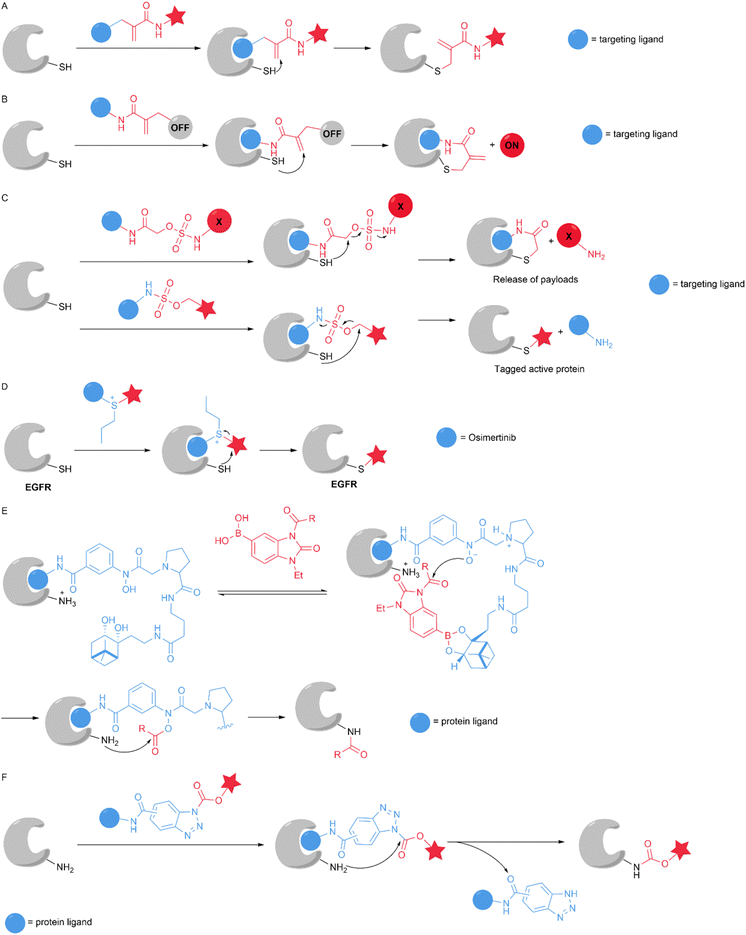
The binding pockets of proteins can engage a cognate ligand. Applying a reactive warhead to that ligand converts it into a chemical probe. After protein–ligand interactions occur, the electrophilic group of the probe is positioned near the POI, thereby facilitating nucleophilic attack by residues near the ligand binding site and inducing covalent binding. Acrylamide has been widely employed as an electrophilic warhead for the covalent labelling of many cysteine-containing proteins. London's group developed a method based on covalent ligand-directed release (CoLDR) chemistry using α-substituted methacrylamide and performed a series of studies using this method. In 2021, Reddi et al. utilized this technology to achieve the site-specific labelling of endogenous proteins. Protein labelling via traditional affinity methods often inhibits protein activity because the targeting ligand permanently occupies the target binding pocket. The authors designed α-methacrylamide reagents with the targeting ligand as a leaving group, enabling the release of the targeting ligand while various functional tags were installed on the target protein (Fig. 12A). The authors documented a variety of applications of this method, including determining the half-life of BTK in its native environment and quantifying BTK degradation via proteolysis-targeting chimaeras (PROTACs). This method is a labelling strategy that maintains target protein activity and represents an important addition to the chemical biology toolbox.67 In the same year, Reddi et al. found that hetero α-substituted methacrylamides have higher thiol reactivity and undergo a conjugate addition–elimination reaction, ultimately releasing the substituent (Fig. 12B). The authors altered the reactivity of α-methacrylamides by changing different α-substituents and used the BTK inhibitor ibrutinib as a model for modification, which exhibited low reactivity towards glutathione and high site selectivity. This method can be used to develop fluorescent turn-on probes. The authors generated a BTK CoLDR chemiluminescent probe that can be used for high-throughput screening of BTK inhibitors.68 The tunability of electrophile reactivity can help determine the optimal balance between selectivity and potency. In 2023, Reddi et al. reported that sulfamate acetamide serves as a tuneable and low-reactivity electrophile. When the reagents react with cysteine, they release sulfamic acid. This “self-immolative” feature enables their use in covalent ligand-directed release chemistry. The authors achieved the generation of “turn-on” probes and traceless ligand-directed site-specific protein labelling (Fig. 12C).69 In 2023, Wang et al. developed a strategy for the site-selective modification of the cysteine residues of target proteins using a sulfonium warhead based on a proximity-triggered approach. The authors used an inhibitor of epidermal growth factor receptor (EGFR), osimertinib, as the binding ligand and a model compound to develop a sulfonium-based reagent for EGFR modification. It can transfer different groups, such as bioorthogonal groups, dyes, and biotin, to EGFR (Fig. 12D). The authors used this method to achieve the transfer of covalent E3 ligase recruiter and hydrophobic tags to EGFR, achieving EGFR degradation based on the selectivity of the transferable PROTAC strategy and hydrophobic tagging strategy.70Selective methods for directly introducing post-translational modifications (PTMs) into proteins in living cells are highly important. In 2021, Adamson et al. targeted E. coli dihydrofolate reductase expressed within human cells as the substrate and utilized a boronate-assisted hydroxamic acid (BAHA) catalyst system to install a variety of acyl groups on the protein of interest. The system comprises a protein ligand, a hydroxamate Lewis base, and a diol moiety. When used in conjunction with a boronic acid-containing acyl donor, the catalyst can facilitate acyl transfer to the target lysine residue by binding to the protein of interest (Fig. 12E). The system affords minimal off-target protein reactivity and provides resistance to glutathione. For proteins that have a suitable ligand, this approach enables both naturally occurring and abiotic lysine PTMs to be installed within live cells.71 Most reactive groups currently used for protein affinity labelling have low reactivity towards nucleophilic residues, and the proximity effect accelerates the reaction to achieve selectivity. Great potential and an urgent need for the development of new reactive groups for affinity labelling that have fast reaction rates and good selectivity exist. In 2022, Xin et al. developed benzotriazole (BTA) chemistry that can be incorporated into affinity-labelled probes for the rapid and selective labelling of different cytosolic and membrane proteins both in vitro and in cells (Fig. 12F). Affinity-based BTA probes selectively and covalently label a lysine residue in the vicinity of the ligand binding site of a target protein with a reaction half-time of 28 s.72
4.3 Affinity biomacromolecule-directed modification
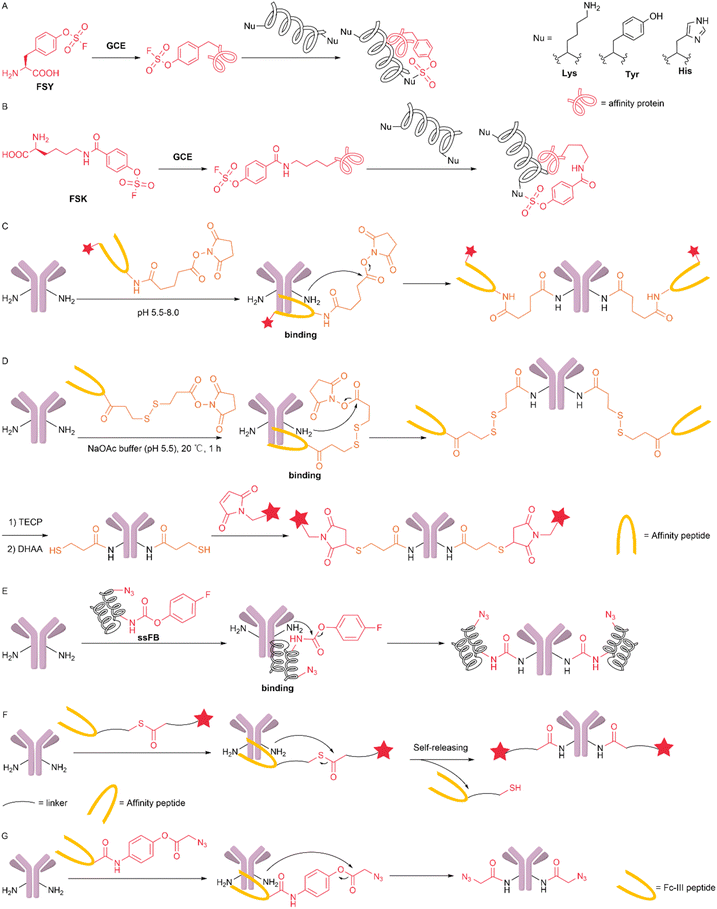
In enzymatic reactions, due to the affinity between the enzyme and the reaction substrate, the reaction substrate can move closer to the active centre of the enzyme. The affinity between substrate proteins, especially antibodies, and enzymes can also occur between the protein and affinity biomacromolecules. Similar to the proximity effect observed in natural enzymes, appending a reactive warhead to affinity biomacromolecules decreases the distance between the reaction reagent and the target residue, increasing the effective local concentration of the reagent. This strategy has been widely adopted by researchers for site-specific protein modification. Proteins can act as affinity biomacromolecules. Wang's group incorporated latent bioreactive unnatural amino acids (Uaas) into proteins to react with target natural amino acid residues through their proximity and reactivity. In 2018, Wang et al. reported the genetic encoding of fluorosulfate-L-tyrosine (FSY) as the first latent bioreactive Uaa that undergoes sulfur–fluorine exchange (SuFEx) on proteins in vivo. After FSY is incorporated into affinity proteins, it can selectively react with nearby lysine, histidine, and tyrosine residues on the protein of interest, directly generating covalent intraprotein bridges and interprotein cross-links between interacting proteins in living cells (Fig. 13A). FSY possesses proximity-activated reactivity, multitargeting capability, and excellent biocompatibility, making it highly important for in vivo protein covalent cross-links.73 FSY has a relatively rigid and short side chain, which restricts the diversity of targetable proteins and the scope of applications. In 2021, Liu et al. designed and genetically encoded a new latent bioreactive Uaa, fluorosulfonyloxybenzoyl-L-lysine (FSK), in E. coli and mammalian cells. Compared with FSY, it features an aliphatic side chain, offering greater flexibility and a longer reaction distance. FSK is particularly useful for covalently connecting protein sites that are inaccessible to FSY (Fig. 13B). The authors utilized FSY and FSK to target different sites of the epidermal growth factor receptor to combat potential mutation-induced resistance. Applying these reagents to genetically encoded chemical crosslinking can capture enzyme–substrate–protein interactions in E. coli. FSK and FSY together provide a SuFEx system for generating covalent bonds in a variety of proteins both in vitro and in vivo.74 Monoclonal antibodies (mAbs) contain a large number of reactive lysine residues, making site-specific modification with conventional chemistry challenging. Multisite reactivity typically leads to heterogeneous conjugates. Current approaches therefore rely largely on the genetic incorporation of noncanonical amino acids to introduce bioorthogonal handles. Peptides can also act as affinity biomacromolecules. Proximity-driven chemistry leverages the binding properties between affinity peptides and antibodies, eliminating the need for an additional antibody pretreatment. In 2019, Kishimoto et al. developed the method of chemical conjugation by affinity peptide (CCAP) for site-specific chemical modification using a peptide that binds with high affinity to human IgG-Fc. This method enables the rapid modification of specific residues (Lys248) in a single step under mild conditions, resulting in the formation of a stable amide bond between the peptide and the Fc (Fig. 13C). Antibody–drug conjugates and bispecific antibodies can be further developed by conjugating cytotoxic molecules or orthogonal reactive groups to affinity peptides, making it an effective conjugation method.75 However, due to the presence of a noncleavable linker, the affinity peptide remains on the modified antibody. In the same year, Yamada et al. developed a site-specific lysine modification method for native IgG antibodies using similar peptides with special affinity for the Fc region of antibodies, enabling the creation of antibody–drug conjugates. This method is called affinity peptide-mediated regiodivergent functionalization (AJICAP). When a cleavable linker is used, the affinity peptide departs and allows the introduction of a free thiol group into IgG. The authors subsequently conjugated cytotoxic molecules via thiol-maleimide coupling (Fig. 13D). This modification method did not impair antigen binding. The results from the mouse model demonstrated that the resulting ADC selectively targeted and killed HER2-positive cells.76 In 2021, Cao et al. reported a new proximity-induced site-specific antibody conjugation method (pClick) based on the proximity-induced reactivity between affinity peptide crosslinkers and a nearby antibody lysine residue. The authors designed and optimized a proximity-activated crosslinking peptide based on antibody-binding proteins (FB proteins) derived from Staphylococcus aureus protein A. The authors truncated the FB protein and modified it with 4-fluorophenyl carbamate lysine (FPheK) at position 25 (Fig. 13E). This modification resulted in excellent conjugation efficiency with the native trastuzumab antibody. The authors site-specifically conjugated the Tras antibody with monomethyl auristatin E to synthesize ADCs for the treatment of breast cancer and prepared multifunctional ADCs by coupling 5/6-carboxyfluorescein (FAM)-lysine to the N-terminus of the peptide. Using this technology, the authors prepared bispecific small molecule–antibody conjugates, which displayed excellent antitumour activity in mouse xenograft models.77In the methods described above, after modification, the affinity peptides are either conjugated to the antibody or require subsequent cleavage. The self-release of the targeting peptide in a one-step reaction can be used to develop simple and efficient traceless modifications. In 2022, Zeng et al. developed a new traceless modification strategy using thioester-based acyl transfer reagents, enabling the facile and efficient one-step synthesis of site-specific antibody–drug conjugates (ADCs). The authors designed the sequence of the Fc-binding peptide and optimized the linker. The optimized reagent can directly assemble the toxin precisely onto the K251 position and simultaneously self-release the affinity ligand in one step (Fig. 13F). Using this method, the authors synthesized a series of K251-linked ADCs, which exhibited excellent physicochemical and biological properties, including uniformity, stability, and in vitro and in vivo antitumour activity. This method enables the traceless, one-step synthesis of site-specific ADCs and provides a general model for the application of ligand-directed protein modification.78 In the same year, Yuan et al. achieved the site-selective acetylation of single lysine residues in human immunoglobulin G. The authors designed and modified the Fc-III peptide identified from the bacteriophage library. Leveraging peptide–antibody affinity, the peptide positions a phenolic ester close to Lys248. Through a proximity-driven group transfer reaction, an acetyl group is transferred to Lys248. Using this method, functional groups such as azide, alkyne, fluorescent molecules, or biotin can also be selectively installed on Lys 248, allowing further derivatization of IgG (Fig. 13G). The authors constructed antibody-conjugated liposomes (immunoliposomes) targeting HER2-positive (HER2+) cancer cells and constructed a bispecific antibody complex (bsAbC) covalently linked to anti-HER2 antibodies and anti-CD3 antibodies.79
5 Enzymatic strategies
Enzymes possess an inherent ability to recognize specific sites within peptides and proteins, such as specific amino acid sequences or defined structural motifs. This natural selectivity makes them well suited for achieving site-specific modifications in a predictable and efficient manner. Current site-specific enzymatic modification strategies rely on the recognition of chemical or spatial structures. We divided this section into enzymatic modification strategies based on the recognition of native protein motifs, introduced tags, and certain spatial structures.5.1 Engineered enzymes for the recognition of native protein motifs
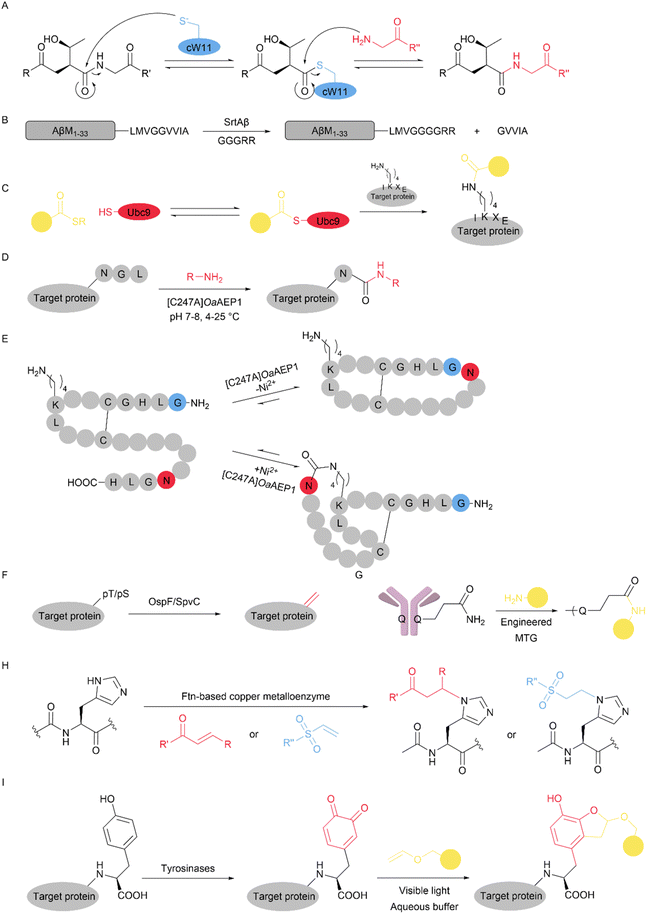
Many natural enzymes achieve site-selective catalysis by recognizing specific peptide motifs within their substrates. Similarly, proteins possess unique amino acid sequences that can serve as intrinsic recognition elements for selective enzymatic modification. For example, Staphylococcus aureus sortase A (SaSrtA) recognizes the LPXTG motif and cleaves the peptide bond between threonine and glycine. This reaction generates a thioester-linked intermediate with the enzyme, which is then resolved by nucleophilic attack from an N-terminal glycine-containing acceptor, forming a new amide bond with high site specificity.80 Thus, some strategies concentrate on engineering enzymes to recognize specific peptide sequences in natural proteins. For example, Whedon et al. engineered a sortase variant, cW11 (derived from sortase F40 reported by Schwarzer and colleagues81), which efficiently recognizes the APXTG motif, cleaves between threonine and glycine, and generates a threonine-linked product through nucleophilic attack at the threonine residue (Fig. 14A). This tool enables late-stage H3 functionalization with high precision and efficiency.82 Similarly, based on sortase A, Podracky et al. employed a yeast display selection strategy to generate a sortase variant capable of the site-specific modification of Aβ peptides (Fig. 14B). The engineered enzyme selectively recognizes the natural LMVGG motif (residues 34–38 of Aβ) and cleaves between the two glycines, allowing the installation of diverse conjugates without the need for genetically introduced tags.835.2 Tag-guided enzymatic modification
In parallel with engineering enzymes for the recognition of native sequence motifs, an alternative strategy involves the introduction of short peptide tags that are specifically recognized by modifying enzymes. These tag-guided enzymatic approaches enable site-selective functionalization in a modular and predictable manner, expanding the toolkit for precise protein modification across diverse biological contexts. Compared with engineering enzymes for each individual target, this approach is more broadly applicable because, as long as a small tag can be introduced without disrupting the protein function, the same enzyme can be used to modify many different proteins without the need for further customization. However, the requirement to genetically introduce such tags presents limitations, particularly when working with native proteins or in vivo applications where tag insertion may be impractical or interfere with biological processes.Using this tag-guided strategy, different types of modifications can be added. For example, in a 2020 study, Hofmann et al. described the use of the ubiquitin-conjugating enzyme Ubc9 to achieve site-specific lysine acylation on the tag in target proteins (Fig. 14C). Ubc9 selectively recognizes a minimal four-residue tag (IKXE, where K is lysine, X is any amino acid, and E is glutamic acid) and acylates the lysine within that tag, thereby directing the modification to a defined, tag-encoded site. This tag-guided strategy enables efficient and predictable lysine acylation under mild aqueous conditions and is compatible with a broad range of acyl donors, including those bearing biotin or biotin conjugated with PEG spacers.84
In earlier work, Ploegh, Durek and colleagues developed a C247A mutant of OaAEP1 ([C247A]OaAEP1). In this system, the NGL recognition motif is artificially introduced at the C-terminus of the protein. The enzyme cleaves the peptide bond between the asparagine and glycine residues and forms a thioester acyl-enzyme intermediate at its catalytic cysteine. This intermediate can subsequently be resolved by nucleophiles such as peptide N-termini or other primary amines, thereby producing different C-terminal ligation products (Fig. 14D).85,86 In 2024, the authors subsequently expanded the application of OaAEP1 to peptide cyclization, employing it to catalyse head-to-tail and side chain-to-C-terminal cyclization (Fig. 14E). By extending the recognition motif with a His residue, the authors achieved the modulation of cyclization. For instance, the peptide shown can be head-to-tail cyclized in the absence of Ni2+. When Ni2+ is added, the N-terminal Gly-Leu-His sequence is quenched, and C-terminus-to-side chain cyclization occurs instead.87 In addition, Veer et al. further expanded the substrate scope to include secondary amines such as proline and hydroxyproline, enabling the formation of stable tertiary amide bonds and facilitating applications such as dual labelling.88
Dha is widely used for protein modification. The bis-alkylation-elimination strategy is one of the most powerful approaches for converting Cys to Dha in peptides or proteins.89,90 However, this method may not differentiate different Cys residues. In 2022, Yang et al. reported an enzymatic approach to site-selectively introduce Dha/dehydrobutyrine (Dhb) into peptides and proteins using the bacterial phosphothreonine lyases OspF and SpvC (Fig. 14F). Using self-assembled monolayers for a matrix-assisted laser desorption/ionization mass spectrometry (SAMDI-MS)-based high-throughput screening platform, the authors identified optimized recognition motifs [F/Y/W]-pS or [F/Y/W]-pT that enable the conversion of single phosphorylated serine or threonine residues into Dha or Dhb with high efficiency. Using this method, the authors demonstrated the utility of this system in one-pot protein labelling and in modifying surface-displayed peptides on live E. coli, highlighting its potential for bioorthogonal functionalization under mild conditions.91
5.3 Structure-guided enzymatic protein modification
Another class of protein modification approaches achieves site selectivity by constructing local reaction environments that favour covalent attachment at a specific residue. Unlike sequence-guided enzymatic modification, which relies on the recognition of a defined peptide motif, structure-guided approaches instead exploit three-dimensional proximity and local surface features. Specifically, these methods typically involve engineering a catalytic pocket or reactive centre that confines reactivity to residues positioned within its spatial reach. Selectivity emerges from geometric constraints, such as coordination geometry, residue orientation, and steric accessibility. Such structure-guided strategies provide a powerful and orthogonal route to precision bioconjugation, particularly for residues traditionally considered difficult to modify selectively. For example, a microbial transglutaminase (MTG) was rationally engineered to selectively modify a native glutamine residue, Gln295, located in the hinge region of human IgG1 antibodies (Fig. 14G). The resulting MTG variant enables site-specific bioconjugation even on fully glycosylated antibodies, catalysing amide bond formation between the Gln295 side chain and amine-containing payloads, highlighting the utility of enzyme engineering in tag-free protein functionalization.92 Although the authors did not explicitly delineate the molecular basis of this selectivity, they suggested that prior reports showed that efficient Q295 conjugation typically required Fc deglycosylation, which indicates that selectivity may be governed by conformational/structural accessibility rather than recognition of a certain sequence.In 2024, Tsou et al. developed a ferritin (Ftn)-based artificial metalloenzyme platform capable of catalysing histidine-selective aza-Michael additions (Fig. 14H). By introducing a Cu(II)-binding noncanonical amino acid (such as 4-boronophenylalanine (BtA)) and fusing a substrate recognition peptide (αCT), the authors recruited insulin and stabilized a binding conformation that positioned His5 near the catalytic centre. This conformational restriction enables single-site histidine modification, even in the presence of more reactive residues, highlighting a general strategy for sequence-independent yet site-selective protein editing.93
Tyrosinases (EC 1.14.18.1) is another enzyme that catalyse the oxidation of tyrosine side chains to o-quinones. Certain tyrosine residues in peptides and proteins can also be oxidized by tyrosinases, enabling subsequent modification.94 Using this enzyme, Chen et al. developed a tyrosinase-mediated strategy to achieve the site-selective modification of antibodies (Fig. 14I). By leveraging the structural accessibility of specific tyrosine residues (Fc-Y296), the authors selectively oxidized exposed Tyr sites on heavy chains, generating reactive o-quinones that could be further modified. This approach enables precise bioconjugation without disrupting antibody function, highlighting the potential of site-selective antibody modification strategies to enhance targeted immunotherapy for cancer.95
6 Conclusions
Site-specific modification is pivotal for constructing peptides and proteins with precisely defined structures and functionalities. Homogeneously modified products are indispensable for probing structure–function relationships. This review provides a comprehensive overview of recent advances in the site-specific modification of peptides and proteins, detailing both the design principles that underpin these transformations and their innovative subsequent applications.Drawing on the intrinsic differences among protein sites, researchers have engineered site-selective reactions. By appending specifically modifiable tags or exploiting targeting moieties, they achieve precise positional control. Enzymatic strategies further extend this toolkit, offering exceptional site selectivity and broad application potential. In conclusion, site-specific modification offers innovative avenues for advancing traditional bioconjugation and bioorthogonal chemistry. Nevertheless, the field is still confronted with challenges. Some methods suffer from low conversion yields. Many efficient strategies additionally rely on the installation of exogenous tags or recognition motifs, restricting their use with native proteins in complex environments. Moreover, most current modifications are irreversible. The development of dynamically controllable labelling strategies remains a key direction for future research.
Moving forwards, the development of flexible yet precise modification strategies is crucial. Emerging computational tools, particularly machine learning algorithms trained on protein structure and reactivity databases, are increasingly enabling the prediction of subtle differences in the intrinsic properties of amino acids, potentially allowing tag-free discrimination among the same amino acid type in native proteins. Expanding the reaction toolkit through previously underexplored, orthogonal chemistries, organometallic catalysis, and photochemical and electrochemical transformations will broaden the scope of protein modifications. Notably, the rational design of novel electrophiles with tuned reactivity profiles promises to unlock the site-selective modification of traditionally challenging, low-abundance residues such as histidine, aspartic acid, and tryptophan. Directed evolution and AI-assisted enzyme design will further allow the reprogramming of natural substrate-recognition motifs, delivering artificial enzymes with superior catalytic profiles. Finally, continued progress in supramolecular and stimulus-responsive chemistry should yield reversible, on-demand protein modifications that confer spatiotemporal control over protein function in real time. As ongoing research tackles these challenges, the prospects for the ongoing evolution and broader application of site-specific modifications in chemical biology and related fields are promising.
Author contributions
Xin-Rong Duan, Meng-Qian Zhang and Yan-Mei Li organize the overall narrative of the manuscript. Xin-Rong Duan, Geng-Hui Feng and Yan-Mei Li co-wrote the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
This work was supported by the National Natural Science Foundation of China (22237003), the Tsinghua-Toyota Joint Research Fund (20253930081).Notes and references
- C. D. Spicer and B. G. Davis, Selective chemical protein modification, Nat. Commun., 2014, 5, 4740 CrossRef CAS PubMed.
- O. Boutureira and G. J. L. Bernardes, Advances in Chemical Protein Modification, Chem. Rev., 2015, 115, 2174–2195 CrossRef CAS.
- E. A. Hoyt, P. M. S. D. Cal, B. L. Oliveira and G. J. L. Bernardes, Contemporary approaches to site-selective protein modification, Nat. Rev. Chem., 2019, 3, 147–171 CrossRef CAS.
- N. C. Reddy, M. Kumar, R. Molla and V. Rai, Chemical methods for modification of proteins, Org. Biomol. Chem., 2020, 18, 4669–4691 RSC.
- D. G. Rawale, K. Thakur, S. R. Adusumalli and V. Rai, Chemical Methods for Selective Labeling of Proteins, Eur. J. Org Chem., 2019, 2019, 6749–6763 CrossRef CAS.
- Z. Sun, H. Liu and X. Li, Precision in protein chemical modification and total synthesis, Chem, 2024, 10, 767–799 CAS.
- W.-H. Li, J.-Y. Su and Y.-M. Li, Rational Design of T-Cell- and B-Cell-Based Therapeutic Cancer Vaccines, Acc. Chem. Res., 2022, 55, 2660–2671 CrossRef CAS PubMed.
- A. Rudra, J. Li, R. Shakur, S. Bhagchandani and R. Langer, Trends in Therapeutic Conjugates: Bench to Clinic, Bioconjugate Chem., 2020, 31, 462–473 CrossRef CAS PubMed.
- S.-H. Zhuo, T.-Y. Wang, L. Zhao, J.-Y. Su, J.-J. Hu, Y.-F. Zhao and Y.-M. Li, piSTING: A Pocket-Independent Agonist Based on Multivalency-Driven STING Oligomerization, Angew. Chem., Int. Ed., 2024, 63, e202407037 CrossRef CAS PubMed.
- S. J. Walsh, J. D. Bargh, F. M. Dannheim, A. R. Hanby, H. Seki, A. J. Counsell, X. Ou, E. Fowler, N. Ashman, Y. Takada, A. Isidro-Llobet, J. S. Parker, J. S. Carroll and D. R. Spring, Site-selective modification strategies in antibody–drug conjugates, Chem. Soc. Rev., 2021, 50, 1305–1353 RSC.
- G.-H. Feng, T.-Y. Wang and Y.-M. Li, Recent developments in late-stage protein modification, Chem. Commun., 2025, 61, 11095–11117 RSC.
- R. J. Spears and V. Chudasama, Recent advances in N- and C-terminus cysteine protein bioconjugation, Curr. Opin. Chem. Biol., 2023, 75, 102306 CrossRef CAS.
- A. Boto, C. C. González, D. Hernández, I. Romero-Estudillo and C. J. Saavedra, Site-selective modification of peptide backbones, Org. Chem. Front., 2021, 8, 6720–6759 RSC.
- Q. Gan and C. Fan, Orthogonal Translation for Site-Specific Installation of Post-translational Modifications, Chem. Rev., 2024, 124, 2805–2838 CrossRef CAS PubMed.
- M. J. Matos, B. L. Oliveira, N. Martínez-Sáez, A. Guerreiro, P. M. S. D. Cal, J. Bertoldo, M. Maneiro, E. Perkins, J. Howard, M. J. Deery, J. M. Chalker, F. Corzana, G. Jiménez-Osés and G. J. L. Bernardes, Chemo- and Regioselective Lysine Modification on Native Proteins, J. Am. Chem. Soc., 2018, 140, 4004–4017 CrossRef CAS PubMed.
- P.-Y. He, Y. Zhou, P.-G. Chen, M.-Q. Zhang, J.-J. Hu, Y.-J. Lim, H. Zhang, K. Liu and Y.-M. Li, A Hydroxylamine-Mediated Amidination of Lysine Residues That Retains the Protein's Positive Charge, Angew. Chem., Int. Ed., 2024, 63, e202402880 CrossRef CAS.
- S. Bloom, C. Liu, D. K. Kölmel, J. X. Qiao, Y. Zhang, M. A. Poss, W. R. Ewing and D. W. C. MacMillan, Decarboxylative alkylation for site-selective bioconjugation of native proteins via oxidation potentials, Nat. Chem., 2018, 10, 205–211 CrossRef CAS PubMed.
- M. K. Miller, H. Wang, K. Hanaya, O. Zhang, A. Berlaga and Z. T. Ball, Copper-mediated peptide arylation selective for the N-terminus, Chem. Sci., 2020, 11, 10501–10505 RSC.
- K. Hanaya, K. Taguchi, Y. Wada and M. Kawano, One-Step Maleimide-Based Dual Functionalization of Protein N-Termini, Angew. Chem., Int. Ed., 2025, 64, e202417134 CrossRef CAS PubMed.
- Z. Lin, B. Liu, M. Lu, Y. Wang, X. Ren, Z. Liu, C. Luo, W. Shi, X. Zou, X. Song, F. Tang, H. Huang and W. Huang, Controlled Reversible N-Terminal Modification of Peptides and Proteins, J. Am. Chem. Soc., 2024, 146, 23752–23763 CrossRef CAS.
- R. Anandakrishnan, B. Aguilar and A. V. Onufriev, H++ 3.0: automating pK prediction and the preparation of biomolecular structures for atomistic molecular modeling and simulations, Nucleic Acids Res., 2012, 40, W537–W541 CrossRef CAS PubMed.
- İ. Soylu and S. M. Marino, Cy-preds: An algorithm and a web service for the analysis and prediction of cysteine reactivity, Proteins: Struct., Funct., Bioinf., 2016, 84, 278–291 CrossRef PubMed.
- H. Wang, X. Chen, C. Li, Y. Liu, F. Yang and C. Wang, Sequence-Based Prediction of Cysteine Reactivity Using Machine Learning, Biochemistry, 2018, 57, 451–460 CrossRef CAS PubMed.
- C. J. Wilson, M. Karttunen, B. L. de Groot and V. Gapsys, Accurately Predicting Protein pKa Values Using Nonequilibrium Alchemy, J. Chem. Theory Comput., 2023, 19, 7833–7845 CrossRef CAS PubMed.
- L. Purushottam, S. R. Adusumalli, U. Singh, V. B. Unnikrishnan, D. G. Rawale, M. Gujrati, R. K. Mishra and V. Rai, Single-site glycine-specific labeling of proteins, Nat. Commun., 2019, 10, 2539 CrossRef PubMed.
- H. Machida and K. Kanemoto, N-Terminal-Specific Dual Modification of Peptides through Copper-Catalyzed [3 + 2] Cycloaddition, Angew. Chem., Int. Ed., 2024, 63, e202320012 CrossRef CAS.
- H. Liu, H. Deol, A. Raeisbahrami, H. Askari, C. D. Wight, V. M. Lynch and E. V. Anslyn, A Method for Rigorously Selective Capture and Simultaneous Fluorescent Labeling of N-Terminal Glycine Peptides, J. Am. Chem. Soc., 2024, 146, 13727–13732 CrossRef CAS PubMed.
- X. Zheng, Z. Li, W. Gao, X. Meng, X. Li, L. Y. P. Luk, Y. Zhao, Y.-H. Tsai and C. Wu, Condensation of 2-((Alkylthio)(aryl)methylene)malononitrile with 1,2-Aminothiol as a Novel Bioorthogonal Reaction for Site-Specific Protein Modification and Peptide Cyclization, J. Am. Chem. Soc., 2020, 142, 5097–5103 CrossRef CAS PubMed.
- A. Istrate, M. B. Geeson, C. D. Navo, B. B. Sousa, M. C. Marques, R. J. Taylor, T. Journeaux, S. R. Oehler, M. R. Mortensen, M. J. Deery, A. D. Bond, F. Corzana, G. Jiménez-Osés and G. J. L. Bernardes, Platform for Orthogonal N-Cysteine-Specific Protein Modification Enabled by Cyclopropenone Reagents, J. Am. Chem. Soc., 2022, 144, 10396–10406 CrossRef CAS.
- M. H. Mir, S. Parmar, C. Singh and D. Kalia, Location-agnostic site-specific protein bioconjugation via Baylis Hillman adducts, Nat. Commun., 2024, 15, 859 CrossRef CAS PubMed.
- Z.-Z. Chen, B. Tan, Y.-M. Li, Y.-F. Zhao, Y.-F. Tong and J.-F. Wang, Activity Difference between α-COOH and β-COOH in N-Phosphorylaspartic Acids, J. Org. Chem., 2003, 68, 4052–4058 CrossRef CAS PubMed.
- M. Garreau, F. Le Vaillant and J. Waser, C-Terminal Bioconjugation of Peptides through Photoredox Catalyzed Decarboxylative Alkynylation, Angew. Chem., Int. Ed., 2019, 58, 8182–8186 CrossRef CAS PubMed.
- K.-Q. Ling, J. Kim and L. M. Sayre, Catalytic Turnover of Benzylamine by a Model for the Lysine Tyrosylquinone (LTQ) Cofactor of Lysyl Oxidase, J. Am. Chem. Soc., 2001, 123, 9606–9611 CrossRef CAS PubMed.
- J. P. Klinman and F. Bonnot, Intrigues and Intricacies of the Biosynthetic Pathways for the Enzymatic Quinocofactors: PQQ, TTQ, CTQ, TPQ, and LTQ, Chem. Rev., 2014, 114, 4343–4365 CrossRef CAS.
- S. Wang, Q. Zhou, X. Chen, R.-H. Luo, Y. Li, X. Liu, L.-M. Yang, Y.-T. Zheng and P. Wang, Modification of N-terminal α-amine of proteins via biomimetic ortho-quinone-mediated oxidation, Nat. Commun., 2021, 12, 2257 CrossRef CAS PubMed.
- J. Kim, B. X. Li, R. Y. C. Huang, J. X. Qiao, W. R. Ewing and D. W. C. MacMillan, Site-Selective Functionalization of Methionine Residues via Photoredox Catalysis, J. Am. Chem. Soc., 2020, 142, 21260–21266 CrossRef CAS PubMed.
- B. X. Li, D. K. Kim, S. Bloom, R. Y. C. Huang, J. X. Qiao, W. R. Ewing, D. G. Oblinsky, G. D. Scholes and D. W. C. MacMillan, Site-selective tyrosine bioconjugation via photoredox catalysis for native-to-bioorthogonal protein transformation, Nat. Chem., 2021, 13, 902–908 CrossRef CAS PubMed.
- J.-J. Hu, P.-Y. He and Y.-M. Li, Chemical modifications of tryptophan residues in peptides and proteins, J. Pept. Sci., 2021, 27, e3286 CrossRef CAS.
- C. R. Hoopes, F. J. Garcia, A. M. Sarkar, N. J. Kuehl, D. T. Barkan, N. L. Collins, G. E. Meister, T. R. Bramhall, C.-H. Hsu, M. D. Jones, M. Schirle and M. T. Taylor, Donor–Acceptor Pyridinium Salts for Photo-Induced Electron-Transfer-Driven Modification of Tryptophan in Peptides, Proteins, and Proteomes Using Visible Light, J. Am. Chem. Soc., 2022, 144, 6227–6236 CrossRef CAS PubMed.
- P.-Y. He, H. Chen, H.-G. Hu, J.-J. Hu, Y.-J. Lim and Y.-M. Li, Late-stage peptide and protein modifications through phospha-Michael addition reaction, Chem. Commun., 2020, 56, 12632–12635 RSC.
- B. Josephson, C. Fehl, P. G. Isenegger, S. Nadal, T. H. Wright, A. W. J. Poh, B. J. Bower, A. M. Giltrap, L. Chen, C. Batchelor-McAuley, G. Roper, O. Arisa, J. B. I. Sap, A. Kawamura, A. J. Baldwin, S. Mohammed, R. G. Compton, V. Gouverneur and B. G. Davis, Light-driven post-translational installation of reactive protein side chains, Nature, 2020, 585, 530–537 CrossRef CAS.
- Z. Zhao, D. Shimon and N. Metanis, Chemoselective Copper-Mediated Modification of Selenocysteines in Peptides and Proteins, J. Am. Chem. Soc., 2021, 143, 12817–12824 CrossRef CAS PubMed.
- L. J. Dowman, S. S. Kulkarni, J. V. Alegre-Requena, A. M. Giltrap, A. R. Norman, A. Sharma, L. C. Gallegos, A. S. Mackay, A. P. Welegedara, E. E. Watson, D. van Raad, G. Niederacher, S. Huhmann, N. Proschogo, K. Patel, M. Larance, C. F. W. Becker, J. P. Mackay, G. Lakhwani, T. Huber, R. S. Paton and R. J. Payne, Site-selective photocatalytic functionalization of peptides and proteins at selenocysteine, Nat. Commun., 2022, 13, 6885 CrossRef CAS PubMed.
- C. Loynd, S. J. Singha Roy, V. J. Ovalle, S. E. Canarelli, A. Mondal, D. Jewel, E. D. Ficaretta, E. Weerapana and A. Chatterjee, Electrochemical labelling of hydroxyindoles with chemoselectivity for site-specific protein bioconjugation, Nat. Chem., 2024, 16, 389–397 CrossRef CAS PubMed.
- X. Ling, H. Chen, W. Zheng, L. Chang, Y. Wang and T. Liu, Site-specific protein modification by genetic encoded disulfide compatible thiols, Chin. Chem. Lett., 2020, 31, 163–166 CrossRef CAS.
- J. Liu, R. Cheng, N. Van Eps, N. Wang, T. Morizumi, W.-L. Ou, P. C. Klauser, S. Rozovsky, O. P. Ernst and L. Wang, Genetically Encoded Quinone Methides Enabling Rapid, Site-Specific, and Photocontrolled Protein Modification with Amine Reagents, J. Am. Chem. Soc., 2020, 142, 17057–17068 CrossRef CAS PubMed.
- Y. Wang, J. Zhang, B. Han, L. Tan, W. Cai, Y. Li, Y. Su, Y. Yu, X. Wang, X. Duan, H. Wang, X. Shi, J. Wang, X. Yang and T. Liu, Noncanonical amino acids as doubly bio-orthogonal handles for one-pot preparation of protein multiconjugates, Nat. Commun., 2023, 14, 974 CrossRef CAS PubMed.
- W. Cao, H. Wang, M. Quan, Y. Li, Y. Su, Y. Li, W. Jiang and T. Liu, Reversible control of tetrazine bioorthogonal reactivity by naphthotube-mediated host-guest recognition, Chem, 2023, 9, 2881–2901 CAS.
- C. Zhang, M. Welborn, T. Zhu, N. J. Yang, M. S. Santos, T. Van Voorhis and B. L. Pentelute, π-Clamp-mediated cysteine conjugation, Nat. Chem., 2016, 8, 120–128 CrossRef CAS PubMed.
- R. A. Gama-Brambila, J. Chen, Y. Dabiri, G. Tascher, V. Němec, C. Münch, G. Song, S. Knapp and X. Cheng, A Chemical Toolbox for Labeling and Degrading Engineered Cas Proteins, JACS Au, 2021, 1, 777–785 CrossRef CAS PubMed.
- R. J. Taylor, M. Aguilar Rangel, M. B. Geeson, P. Sormanni, M. Vendruscolo and G. J. L. Bernardes, π-Clamp-Mediated Homo- and Heterodimerization of Single-Domain Antibodies via Site-Specific Homobifunctional Conjugation, J. Am. Chem. Soc., 2022, 144, 13026–13031 CrossRef CAS PubMed.
- J. Ohata, Y. Zeng, L. Segatori and Z. T. Ball, A Naturally Encoded Dipeptide Handle for Bioorthogonal Chan–Lam Coupling, Angew. Chem., Int. Ed., 2018, 57, 4015–4019 CrossRef CAS PubMed.
- V. Thimaradka, J. Hoon Oh, C. Heroven, A. Radu Aricescu, M. Yuzaki, T. Tamura and I. Hamachi, Site-specific covalent labeling of His-tag fused proteins with N-acyl-N-alkyl sulfonamide reagent, Bioorg. Med. Chem., 2021, 30, 115947 CrossRef CAS PubMed.
- M. Guo, K. Zhao, L. Guo, R. Zhou, Q. He, K. Lu, T. Li, D. Liu, J. Chen, J. Tang, X. Fu, J. Zhou, B. Zheng, S. I. Mann, Y. Zhang, J. Huang, B. Yang, T. Zhou, Y. Lei and B. Dang, Copper assisted sequence-specific chemical protein conjugation at a single backbone amide, Nat. Commun., 2023, 14, 8063 CrossRef CAS.
- Y. Zeng, W. Shi, Z. Liu, H. Xu, L. Liu, J. Hang, Y. Wang, M. Lu, W. Zhou, W. Huang and F. Tang, C-terminal modification and functionalization of proteins via a self-cleavage tag triggered by a small molecule, Nat. Commun., 2023, 14, 7169 CrossRef CAS PubMed.
- S. R. Adusumalli, D. G. Rawale, U. Singh, P. Tripathi, R. Paul, N. Kalra, R. K. Mishra, S. Shukla and V. Rai, Single-Site Labeling of Native Proteins Enabled by a Chemoselective and Site-Selective Chemical Technology, J. Am. Chem. Soc., 2018, 140, 15114–15123 CrossRef CAS PubMed.
- S. R. Adusumalli, D. G. Rawale, K. Thakur, L. Purushottam, N. C. Reddy, N. Kalra, S. Shukla and V. Rai, Chemoselective and Site-Selective Lysine-Directed Lysine Modification Enables Single-Site Labeling of Native Proteins, Angew. Chem., Int. Ed., 2020, 59, 10332–10336 CrossRef CAS.
- D. G. Rawale, K. Thakur, P. Sreekumar, S. T. K, R. A, S. R. Adusumalli, R. K. Mishra and V. Rai, Linchpins empower promiscuous electrophiles to enable site-selective modification of histidine and aspartic acid in proteins, Chem. Sci., 2021, 12, 6732–6736 RSC.
- N. C. Reddy, R. Molla, P. N. Joshi, S. T. K, I. Basu, J. Kawadkar, N. Kalra, R. K. Mishra, S. Chakrabarty, S. Shukla and V. Rai, Traceless cysteine-linchpin enables precision engineering of lysine in native proteins, Nat. Commun., 2022, 13, 6038 CrossRef CAS.
- C. F. Afonso, M. C. Marques, J. P. M. António, C. Cordeiro, P. M. P. Gois, P. M. S. D. Cal and G. J. L. Bernardes, Cysteine-Assisted Click-Chemistry for Proximity-Driven, Site-Specific Acetylation of Histones, Angew. Chem., Int. Ed., 2022, 61, e202208543 CrossRef CAS PubMed.
- G. Li, J. Hu, H. Chen, Y.-X. Chen and Y.-M. Li, Cucurbit[8]uril facilitated Michael addition for regioselective cysteine modification, Chem. Commun., 2021, 57, 6086–6089 RSC.
- X. Zhang, G. Lu, M. Sun, M. Mahankali, Y. Ma, M. Zhang, W. Hua, Y. Hu, Q. Wang, J. Chen, G. He, X. Qi, W. Shen, P. Liu and G. Chen, A general strategy for synthesis of cyclophane-braced peptide macrocycles via palladium-catalysed intramolecular sp3 C–H arylation, Nat. Chem., 2018, 10, 540–548 CrossRef CAS PubMed.
- B. Li, X. Li, B. Han, Z. Chen, X. Zhang, G. He and G. Chen, Construction of Natural-Product-Like Cyclophane-Braced Peptide Macrocycles via sp3 C–H Arylation, J. Am. Chem. Soc., 2019, 141, 9401–9407 CrossRef CAS PubMed.
- Y. Weng, X. Ding, J. C. A. Oliveira, X. Xu, N. Kaplaneris, M. Zhu, H. Chen, Z. Chen and L. Ackermann, Peptide late-stage C(sp3)–H arylation by native asparagine assistance without exogenous directing groups, Chem. Sci., 2020, 11, 9290–9295 RSC.
- Z. Chen, M. Zhu, M. Cai, L. Xu and Y. Weng, Palladium-Catalyzed C(sp3)–H Arylation and Alkynylation of Peptides Directed by Aspartic Acid (Asp), ACS Catal., 2021, 11, 7401–7410 CrossRef CAS.
- Z.-L. Hou, Y. Tang, Y. Lu and B. Yao, Ligand-Enabled, Cysteine-Directed β-C(sp3)–H Arylation of Alanine in Linear and Cyclic Peptides: Overcoming the Inhibitory Effect of Peptide Bonds, ACS Catal., 2024, 14, 11026–11033 CrossRef CAS.
- R. N. Reddi, A. Rogel, E. Resnick, R. Gabizon, P. K. Prasad, N. Gurwicz, H. Barr, Z. Shulman and N. London, Site-Specific Labeling of Endogenous Proteins Using CoLDR Chemistry, J. Am. Chem. Soc., 2021, 143, 20095–20108 CrossRef CAS PubMed.
- R. N. Reddi, E. Resnick, A. Rogel, B. V. Rao, R. Gabizon, K. Goldenberg, N. Gurwicz, D. Zaidman, A. Plotnikov, H. Barr, Z. Shulman and N. London, Tunable Methacrylamides for Covalent Ligand Directed Release Chemistry, J. Am. Chem. Soc., 2021, 143, 4979–4992 CrossRef CAS PubMed.
- R. N. Reddi, A. Rogel, R. Gabizon, D. G. Rawale, B. Harish, S. Marom, B. Tivon, Y. S. Arbel, N. Gurwicz, R. Oren, K. David, J. Liu, S. Duberstein, M. Itkin, S. Malitsky, H. Barr, B.-Z. Katz, Y. Herishanu, I. Shachar, Z. Shulman and N. London, Sulfamate Acetamides as Self-Immolative Electrophiles for Covalent Ligand-Directed Release Chemistry, J. Am. Chem. Soc., 2023, 145, 3346–3360 CrossRef CAS PubMed.
- Y. Wang, R. Zhao, C. Wan, W. Kang, R. Wang, C. Chiang, X. Guo, Q. Chang, Z. Hou, Y. Ye, Q. Luo, Z. Zhou, J. Liu, S. Li, D. Wang, F. Yin and Z. Li, A Proximity-Triggered Strategy toward Transferable Proteolysis Targeting Chimeras, CCS Chem., 2022, 5, 1433–1442 CrossRef.
- C. Adamson, H. Kajino, S. A. Kawashima, K. Yamatsugu and M. Kanai, Live-Cell Protein Modification by Boronate-Assisted Hydroxamic Acid Catalysis, J. Am. Chem. Soc., 2021, 143, 14976–14980 CrossRef CAS PubMed.
- X. Xin, Y. Zhang, M. Gaetani, S. L. Lundström, R. A. Zubarev, Y. Zhou, D. P. Corkery and Y.-W. Wu, Ultrafast and selective labeling of endogenous proteins using affinity-based benzotriazole chemistry, Chem. Sci., 2022, 13, 7240–7246 RSC.
- N. Wang, B. Yang, C. Fu, H. Zhu, F. Zheng, T. Kobayashi, J. Liu, S. Li, C. Ma, P. G. Wang, Q. Wang and L. Wang, Genetically Encoding Fluorosulfate-l-tyrosine To React with Lysine, Histidine, and Tyrosine via SuFEx in Proteins in vivo, J. Am. Chem. Soc., 2018, 140, 4995–4999 CrossRef CAS PubMed.
- J. Liu, L. Cao, P. C. Klauser, R. Cheng, V. Y. Berdan, W. Sun, N. Wang, F. Ghelichkhani, B. Yu, S. Rozovsky and L. Wang, A Genetically Encoded Fluorosulfonyloxybenzoyl-L-lysine for Expansive Covalent Bonding of Proteins via SuFEx Chemistry, J. Am. Chem. Soc., 2021, 143, 10341–10351 CrossRef CAS PubMed.
- S. Kishimoto, Y. Nakashimada, R. Yokota, T. Hatanaka, M. Adachi and Y. Ito, Site-Specific Chemical Conjugation of Antibodies by Using Affinity Peptide for the Development of Therapeutic Antibody Format, Bioconjugate Chem., 2019, 30, 698–702 CrossRef CAS PubMed.
- K. Yamada, N. Shikida, K. Shimbo, Y. Ito, Z. Khedri, Y. Matsuda and B. A. Mendelsohn, AJICAP: Affinity Peptide Mediated Regiodivergent Functionalization of Native Antibodies, Angew. Chem., Int. Ed., 2019, 58, 5592–5597 CrossRef CAS PubMed.
- Y. J. Cao, C. Yu, K.-L. Wu, X. Wang, D. Liu, Z. Tian, L. Zhao, X. Qi, A. Loredo, A. Chung and H. Xiao, Synthesis of precision antibody conjugates using proximity-induced chemistry, Theranostics, 2021, 11, 9107–9117 CrossRef CAS PubMed.
- Y. Zeng, W. Shi, Q. Dong, W. Li, J. Zhang, X. Ren, C. Tang, B. Liu, Y. Song, Y. Wu, X. Diao, H. Zhou, H. Huang, F. Tang and W. Huang, A Traceless Site-Specific Conjugation on Native Antibodies Enables Efficient One-Step Payload Assembly, Angew. Chem., Int. Ed., 2022, 61, e202204132 CrossRef CAS PubMed.
- D. Yuan, Y. Zhang, K. H. Lim, S. K. P. Leung, X. Yang, Y. Liang, W. C. Y. Lau, K. T. Chow and J. Xia, Site-Selective Lysine Acetylation of Human Immunoglobulin G for Immunoliposomes and Bispecific Antibody Complexes, J. Am. Chem. Soc., 2022, 144, 18494–18503 CrossRef CAS PubMed.
- Z. Zou, Y. Ji and U. Schwaneberg, Empowering Site-Specific Bioconjugations In Vitro and In Vivo: Advances in Sortase Engineering and Sortase-Mediated Ligation, Angew. Chem., Int. Ed., 2024, 63, e202310910 CrossRef CAS PubMed.
- K. Piotukh, B. Geltinger, N. Heinrich, F. Gerth, M. Beyermann, C. Freund and D. Schwarzer, Directed Evolution of Sortase A Mutants with Altered Substrate Selectivity Profiles, J. Am. Chem. Soc., 2011, 133, 17536–17539 CrossRef CAS PubMed.
- S. D. Whedon, K. Lee, Z. A. Wang, E. Zahn, C. Lu, M. Yapa Abeywardana, L. Fairall, E. Nam, S. DuBois-Coyne, P. De Ioannes, X. Sheng, A. Andrei, E. Lundberg, J. Jiang, K.-J. Armache, Y. Zhao, J. W. R. Schwabe, M. Wu, B. A. Garcia and P. A. Cole, Circular Engineered Sortase for Interrogating Histone H3 in Chromatin, J. Am. Chem. Soc., 2024, 146, 33914–33927 CrossRef CAS PubMed.
- C. J. Podracky, C. An, A. DeSousa, B. M. Dorr, D. M. Walsh and D. R. Liu, Laboratory evolution of a sortase enzyme that modifies amyloid-β protein, Nat. Chem. Biol., 2021, 17, 317–325 CrossRef CAS PubMed.
- R. Hofmann, G. Akimoto, T. G. Wucherpfennig, C. Zeymer and J. W. Bode, Lysine acylation using conjugating enzymes for site-specific modification and ubiquitination of recombinant proteins, Nat. Chem., 2020, 12, 1008–1015 CrossRef CAS PubMed.
- F. B. H. Rehm, T. J. Harmand, K. Yap, T. Durek, D. J. Craik and H. L. Ploegh, Site-Specific Sequential Protein Labeling Catalyzed by a Single Recombinant Ligase, J. Am. Chem. Soc., 2019, 141, 17388–17393 CrossRef CAS PubMed.
- F. B. H. Rehm, T. J. Tyler, K. Yap, S. J. de Veer, D. J. Craik and T. Durek, Enzymatic C-Terminal Protein Engineering with Amines, J. Am. Chem. Soc., 2021, 143, 19498–19504 CrossRef CAS.
- F. B. H. Rehm, T. J. Tyler, Y. Zhou, Y.-H. Huang, C. K. Wang, N. Lawrence, D. J. Craik and T. Durek, Repurposing a plant peptide cyclase for targeted lysine acylation, Nat. Chem., 2024, 16, 1481–1489 CrossRef CAS PubMed.
- S. J. de Veer, Y. Zhou, T. Durek, D. J. Craik and F. B. H. Rehm, Tertiary amide bond formation by an engineered asparaginyl ligase, Chem. Sci., 2024, 15, 5248–5255 RSC.
- J. M. Chalker, S. B. Gunnoo, O. Boutureira, S. C. Gerstberger, M. Fernández-González, G. J. L. Bernardes, L. Griffin, H. Hailu, C. J. Schofield and B. G. Davis, Methods for converting cysteine to dehydroalanine on peptides and proteins, Chem. Sci., 2011, 2, 1666–1676 RSC.
- M. Zhang, P. He and Y. Li, Contemporary Approaches to α,β-Dehydroamino Acid Chemical Modifications, Chem. Res. Chin. Univ., 2021, 37, 1044–1054 CrossRef CAS.
- A. Yang, H. Tao, L. C. Szymczak, L. Lin, J. Song, Y. Wang, S. Bai, J. Modica, S.-Y. Huang, M. Mrksich and X. Feng, Efficient Enzymatic Incorporation of Dehydroalanine Based on SAMDI-Assisted Identification of Optimized Tags for OspF/SpvC, ACS Chem. Biol., 2022, 17, 414–425 CrossRef CAS PubMed.
- S. Dickgiesser, M. Rieker, D. Mueller-Pompalla, C. Schröter, J. Tonillo, S. Warszawski, S. Raab-Westphal, S. Kühn, T. Knehans, D. Könning, J. Dotterweich, U. A. K. Betz, J. Anderl, S. Hecht and N. Rasche, Site-Specific Conjugation of Native Antibodies Using Engineered Microbial Transglutaminases, Bioconjugate Chem., 2020, 31, 1070–1076 CrossRef CAS PubMed.
- J.-C. Tsou, C.-J. Tsou, C.-H. Wang, A.-L. A. Ko, Y.-H. Wang, H.-H. Liang, J.-C. Sun, K.-F. Huang, T.-P. Ko, S.-Y. Lin and Y.-S. Wang, Site-Specific Histidine Aza-Michael Addition in Proteins Enabled by a Ferritin-Based Metalloenzyme, J. Am. Chem. Soc., 2024, 146, 33309–33315 CrossRef CAS PubMed.
- S. Ito, T. Kato, K. Shinpo and K. Fujita, Oxidation of tyrosine residues in proteins by tyrosinase. Formation of protein-bonded 3,4-dihydroxyphenylalanine and 5-S-cysteinyl-3,4-dihydroxyphenylalanine, Biochem. J., 1984, 222, 407–411 CrossRef CAS PubMed.
- H. Chen, H.-C. F. Wong, J. Qiu, B. Li, D. Yuan, H. Kong, Y. Bao, Y. Zhang, Z. Xu, Y.-L. S. Tse and J. Xia, Site-Selective Tyrosine Reaction for Antibody-Cell Conjugation and Targeted Immunotherapy, Adv. Sci., 2024, 11, 2305012 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |