Open Access Article
Open Access ArticleLabel-free optical detection of protein acetylation using UV-vis charge transfer spectra
Himanshi Maniram Devi†
 a,
Apoorva Badaya†b,
Arijit Maity
a,
Apoorva Badaya†b,
Arijit Maity b,
Simangka Bor Saikia
b,
Simangka Bor Saikia a,
Ravindra Venkatramani
a,
Ravindra Venkatramani *b and
Rajaram Swaminathan
*b and
Rajaram Swaminathan *a
*a
aDepartment of Biosciences and Bioengineering, Indian Institute of Technology Guwahati, Guwahati 781039, Assam, India. E-mail: rsw@iitg.ac.in
bDepartment of Chemical Sciences, Tata Institute of Fundamental Research, Homi Bhabha Road, Colaba, Mumbai 400005, India. E-mail: ravi.venkatramani@tifr.res.in
First published on 2nd April 2026
Abstract
We report the first spectroscopic detection of protein acetylation in solution using UV-vis Protein Charge Transfer Spectra (ProCharTS). Acetylation is an important post-translational modification (PTM) that modulates diverse cellular processes, yet its detection relies on antibody-based cost-intensive and/or destructive techniques such as mass spectrometry. ProCharTS exploits the reduction of lysine charge post-acetylation to offer an alternate label-free, easily accessible, and cost-effective option to detect the PTM. Using two charge-rich proteins α3C and α3W, we demonstrate that the ProCharTS extinction coefficient between 370–800 nm monotonically decreases with increasing degree of chemical acetylation in the proteins. Complementary spectroscopic analysis and molecular dynamics (MD) simulations indicate that the ProCharTS signatures arise independent of secondary structure changes although tertiary interactions weaken post-acetylation. Using a new computational MD and Time-Dependent Density Functional Theory (TDDFT) approach to simulate ProCharTS of whole proteins from their known 3D structure, we assign the observed PTM-induced decrease in intensity to changes in the size, composition and spatial distribution of charged residue clusters. Our joint experimental–computational approach enables us to detect five or more acetylation events per protein with significant scope for further improvements in sensitivity. More broadly, this study presents a new optical mode (ProCharTSPTM) exploiting charge transfer transitions to probe/track charged residue modifications in protein solutions.
1. Introduction
Post-translational modifications (PTMs) are an important class of cellular mechanisms that expand the functional diversity, complexity and heterogeneity of the proteome by covalently modifying constituent residues in proteins.1,2 There are over 400 different types of PTMs that can influence protein function.3–5 PTMs can significantly transform the physiochemical properties of proteins including membrane binding propensities,6 site reactivity,7,8 accessibility,9 and protein stiffness.10 Acetylation is an important PTM across prokaryotes and eukaryotes, which can be achieved by both enzymatic and non-enzymatic routes.11 Acetylation adds an acetyl group (CH3CO) to the protein that removes the positive charge from the amino group of Lys or N-terminus, making it neutral. Histone acetyltransferases (HATs)12,13 are responsible for enzymatic acetylation of proteins; alternatively, acetic anhydride,14 aspirin,15 and acetyl CoA16 are the major reagents used in chemical acetylation. Extensive research over the past 50 years have revealed the role of acetylation in regulating many cellular processes including transcription, metabolic states, cell cycle, nuclear transport, and actin nucleation.17–20 Acetylation plays a role in regulating protein–protein interactions,21,22 in cancer biology,23,24 in modulating enzyme activity25 and the transport of essential molecules.26–28 Detection of acetylation in a protein is thus important for understanding many physiological and pathological conditions.At present, Lys acetylation can be confirmed by several techniques such as western blotting,29 immunoprecipitation,30 mass spectrometry,31 radiolabelling,32 NMR,33 IR spectroscopy,34 and flow cytometry.35 While western blot and ELISA36 have both high sensitivity and accuracy, these assays involve multiple washing steps and expensive antibodies, making them laborious, time-consuming and costly. Also, antibody cross-reactivity and false positive cases are limitations.37 Mass spectrometry is a definitive tool for detecting PTMs,38 as it provides a quantitative and accurate change in the mass of the protein upon addition of an external group to the parent protein. However, expensive instrumentation, limitations in maintaining labile PTMs during sample preparation, and the need for careful separation techniques39 pose challenges. The detection of sub-stoichiometric PTMs is especially difficult when the majority of the protein molecules may be unmodified.40 Radiolabelled detection based upon [14C] or [3H]-acetyl-CoA enzymatic labeling and autoradiography analysis after separation with electrophoresis is another option.37 The techniques for detection of protein acetylation described above are costly and require complex sample processing or labelling to be carried out, potentially perturbing the natural state of the proteins. Therefore, there is a need for techniques capable of detecting protein PTMs unambiguously in their native solution state without using external labels and requiring minimal sample preparation. In this context, optical spectroscopy may offer a solution. Interestingly, distinct spectral signatures for acetylation of both the N-terminus and the side chain amino groups of Lys have been reported experimentally using IR and Raman spectroscopy.41 However, these signatures have not been tested for detecting/tracking acetylation in proteins. The post-translational modification of the amino group in Lys by acetylation alters the charge on the Lys sidechain from +1 to zero. This radical change in Lys charge upon acetylation offers new possibilities to detect the PTM. Here, we propose a UV-vis spectroscopic approach to detect PTMs such as acetylation, which alter the charged state of protein residues.
Our approach is based on a novel UV-vis absorption observed in aqueous solutions of Lys-HCl42 and charged amino acid residue rich proteins such as α3C, human serum albumin and calf thymus histones43,44 This new spectrum has been assigned to photo-induced charge transfer (PICT) transitions involving anionic and cationic head groups of all charged residues (including those that become charged with PTMs such as phosphorylation) and the protein backbone.44–46 The link between chargedresidues and ProCharTS has been further confirmed in recent studies of Vázquez and co-workers, which showed that single alpha helical (SAH) peptides comprising solely of (Lys)4(Glu)4 or (Arg)4(Glu)4 repeats exhibit a broad absorption profile extending beyond 400 nm.47 Interestingly, the stability of Lys–Glu SAH peptides increases with the number of repeats, not due to formation of salt bridges as might be expected intuitively, but rather due to the increasing number of possible charged states for the individual residues.48 Our own studies have shown that within protein folds, the sidechains and neutral backbones of Lys and Glu can act as electronic donor–acceptor pairs in the absence of salt-bridges to produce broad UV-vis ProCharTS profiles. Based on the charge complementarity of the amino acid residues and their separations, five types of inter-residue and intra-residue PICT transitions were shown to arise, with diverse donor–acceptor separations ranging from 3–10 Å.46 The demonstrated sensitivity of the underlying PICT transition intensities and spectral range to the charge and clustering of residues makes ProCharTS potentially useful in identifying conformational states of proteins, their interactions, and PTMs, which alter these properties. Indeed, ProCharTS absorption has been demonstrated as an easy in vitro technique to track protein aggregation,49,50 protein unfolding51 and viral capsid assembly.52 Beyond absorption, inter- or intra-residue PICT creates electron and hole pairs, which can either undergo charge separation or recombination. The separated charges derived from ProCharTS excitations can be potentially harnessed for biochemical reactions as in the case of the light harvesting chlorophyll special pairs during photosynthesis.53 On the other hand charge recombination54 leads to excitation dependent ProCharTS luminescence, which has already been demonstrated to be useful in tracking protein aggregation49 and in photosensitizing applications.47
In this paper, we hypothesize that the ProCharTS absorption profile of proteins should be sensitive to the progressive acetylation of Lys rich proteins. To validate this hypothesis, we consider two small (67 residues), folded (3-helix bundle), synthetic proteins, α3C and α3W. These well characterized proteins, each of which possesses a high content of charged residues (∼54% of the sequence), serve as ideal models for studying the effects of acetylation on ProCharTS. We carried out systematic and progressive chemical acetylation of the proteins using acetic anhydride and employed a variety of experimental and computational methods to characterize acetylation and analyze its impact on protein structure and dynamics. Mass spectrometry (MALDI-ToF) confirmed the number of acetyl groups added to residues as a function of increasing concentration of acetic anhydride, besides confirming the heterogeneity in the acetylation levels of the sample protein. CD spectroscopy and the fluorescence of the tryptophan in α3W probed changes in protein structure post-acetylation. Additionally, we developed a computational framework based on classical molecular dynamics (MD) simulations and time-dependent density functional theory (TDDFT) calculations to predict the ProCharTS profile of whole proteins from their 3D structure. Here, absorption profiles are simulated from the spatiotemporal convolution of contributions from all charged residue chromophores within the proteins, providing residue level understanding of spectral changes between acetylated and unacetylated proteins and facilitating their deconvolution.
Our results demonstrate that ProCharTS and its associated luminescence are both sensitive to the presence of charge among the side chains of residues and their spatial proximity. Acetylation, which eliminates the positive charge of Lys, disrupts the size, composition and spatial proximities of charged residue clusters, leading to notable changes in absorbance and luminescence. Specifically, we find a monotonic quenching of the absorption spectra as a function of acetic anhydride concentration, thereby validating our hypothesis. The simulation and deconvolution of ProCharTS through computational methods further allows us to identify oppositely charged dimers as critical chromophores, which significantly contribute to the spectral changes brought about by progressive acetylation. Our study demonstrates an accessible, label-free, and non-invasive optical mode to detect acetylation in protein solutions. More generally, the results presented here highlight the potential of ProCharTS to detect any PTM that alters the charged state of residues in proteins and conceptually advance our understanding of how specific modifications influence protein electronic spectra.
2. Methods
2.1 Experimental
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 102); N-acetyl tryptophan amide (A6501) were procured from Sigma Aldrich, Bengaluru, India. Luria agar (M557); Luria broth (M1245); ampicillin sodium salt (TC021); isopropyl β-D-1-thiogalactopyranoside (I5502); HEPES (4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid) (RM380); sodium chloride (GRM853); sodium hydroxide (GRM467); tris (hydroxy methyl amino methane) (93
102); N-acetyl tryptophan amide (A6501) were procured from Sigma Aldrich, Bengaluru, India. Luria agar (M557); Luria broth (M1245); ampicillin sodium salt (TC021); isopropyl β-D-1-thiogalactopyranoside (I5502); HEPES (4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid) (RM380); sodium chloride (GRM853); sodium hydroxide (GRM467); tris (hydroxy methyl amino methane) (93![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 315); imidazole (GRM1864); MgCl2 (TC186) were obtained from HiMedia Laboratories, India. The PD-10 column (17-0851-01) was purchased from GE Healthcare. Nuvia IMAC Ni-charged resin (780–0800) was purchased from Bio-Rad. All items purchased are of analytical grade with >98% purity.
315); imidazole (GRM1864); MgCl2 (TC186) were obtained from HiMedia Laboratories, India. The PD-10 column (17-0851-01) was purchased from GE Healthcare. Nuvia IMAC Ni-charged resin (780–0800) was purchased from Bio-Rad. All items purchased are of analytical grade with >98% purity.2.1.3.1 Sample preparation. 20 µM α3C and α3W were prepared in 100 mM HEPES buffer at pH 8. We also considered NaH2PO4 buffer, where the protein was less stable and found to precipitate. The concentrations of α3C and α3W were estimated using the Lowry method.56 Here, protein absorbance with Lowry reagents (Reagent I (NaOH, Na2CO3, KNaC4H4O6·H2O, CuSO4·5H2O) and Reagent II (Folin's)) was recorded at 550 nm, and the ProCharTS contamination was eliminated by subtracting the protein absorbance in the absence of the reagents (SI Table T1).
2.1.3.2 Reaction conditions for chemical acetylation of α3C and α3W. In vitro chemical acetylation was carried out using acetic anhydride as described earlier,57 with slight modifications. Briefly, 20 µM α3C and α3W were titrated with varying concentrations (0.05, 0.1, 0.2, 2 and 20 mM) of acetic anhydride. The reaction vessel was kept at 4 °C with continuous stirring during the addition of acetic anhydride in small aliquots, and the pH of the reaction mixture was maintained between 7.5 and 8.5 by adding 0.5 N NaOH as required. The reaction was performed in HEPES buffer for 30 minutes. Subsequently the dialysis and centrifugation of the prepared samples were performed to remove the residual acetic anhydride, byproducts and any large particles, which can contribute to the scattering (details are mentioned in SI Section M1.1.1 and spectral analysis is shown in SI Fig. F3).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 ratio). Protein samples dissolved in the matrix were analyzed in linear mode by flex analysis and flex control software from Bruker Daltonics, Germany. The native (unacetylated) proteins have m/z values of 7464.2 Da (α3C) and 7545.2 Da (α3W), as shown in SI Fig. F2. The addition of an acetyl group to the protein results in a 42 Da shift in the mass spectrum. For samples containing a heterogeneous mixture of populations with varying degrees of acetylation, the mass spectra exhibit a broad distribution with multiple peaks. The number of residues acetylated in each population can be obtained from the m/z value of each acetylated species peak as (Acetylatedm/z – Nativem/z)/42.
3 ratio). Protein samples dissolved in the matrix were analyzed in linear mode by flex analysis and flex control software from Bruker Daltonics, Germany. The native (unacetylated) proteins have m/z values of 7464.2 Da (α3C) and 7545.2 Da (α3W), as shown in SI Fig. F2. The addition of an acetyl group to the protein results in a 42 Da shift in the mass spectrum. For samples containing a heterogeneous mixture of populations with varying degrees of acetylation, the mass spectra exhibit a broad distribution with multiple peaks. The number of residues acetylated in each population can be obtained from the m/z value of each acetylated species peak as (Acetylatedm/z – Nativem/z)/42.2.1.6.1 ProCharTS luminescence. Steady-state luminescence emission spectra were collected for all native and acetylated proteins in deionised water at several excitation wavelengths: 280, 310, 340, 355, 370 and 410 nm (slit width: 2 nm) and emission spectra were recorded in the range 300–500, 330–550, 360–600, 375–650, 390–700 and 430–750 nm, respectively (slit width: 15 nm). Excitation at 280 nm was not carried out for α3W due to interference from emission of indole in Trp, which has a higher quantum yield (∼0.14) compared to ProCharTS luminescence (∼0.01). For acquiring the luminescence spectra of α3C and α3W, the protein concentration was kept at ∼9–10 µM, and recorded spectra were corrected manually for Raman peaks (SI Table T2) by subtracting the emission spectra in the appropriate range from blank samples devoid of protein.
2.1.6.2 Quantum yield calculation. Quantum yields (QYs) for native and acetylated proteins (α3C and α3W) were estimated from experiments in deionised water by exciting samples at 280 and 355 nm at room temperature (25 °C). Details on the standard protocols followed58 are mentioned in SI Section M1.1.2.
2.1.6.3 ANS fluorescence. Protein-ANS binding was monitored by fluorescence emission. The ANS stock sample was prepared in deionised water with concentrations determined from extinction coefficient values at 350 nm (4900 M−1 cm−1). The stock sample was added to the protein samples (∼9–10 µM native/acetylated forms) in deionised water to achieve a final ANS concentration of 10 µM. All samples were excited at 380 nm and emission spectra were collected from 400 to 700 nm using 2 nm excitation and 15 nm emission slit widths. The ProCharTS luminescence contribution was also subtracted manually from all the samples to avoid ProCharTS contamination. All the spectra were recorded in triplicate using Fluoromax-4 spectrofluorometer (Horiba Scientific, USA) at 25 °C.
2.1.6.4 Trp fluorescence. Intrinsic fluorescence of Trp was measured using a spectrofluorometer (Fluoromax-4, Horiba, USA). Native and acetylated α3W samples (∼9–10 µM) were prepared in deionised water and excited at 280 nm. All readings were measured in an optical glass cuvette of path length 10 mm (Hellma: Z802875). Fluorescence data were acquired as S1/R1 signals with a wavelength increment of 1 nm and an integration time of 0.1 s. Emission spectra of α3W (300–500 nm) were collected with 2 nm excitation and 15 nm emission slit width. All spectra were averaged over five scans and manually corrected for Raman peaks (SI Table T2). The emission spectrum of NATA was recorded as a reference for calibrating the monochromator. Steady-state anisotropy of Trp in native and acetylated α3W (∼9–10 µM) was carried out in deionised water using the same instrument (see SI Section M1.1.2 for more details).
2.2 Computational
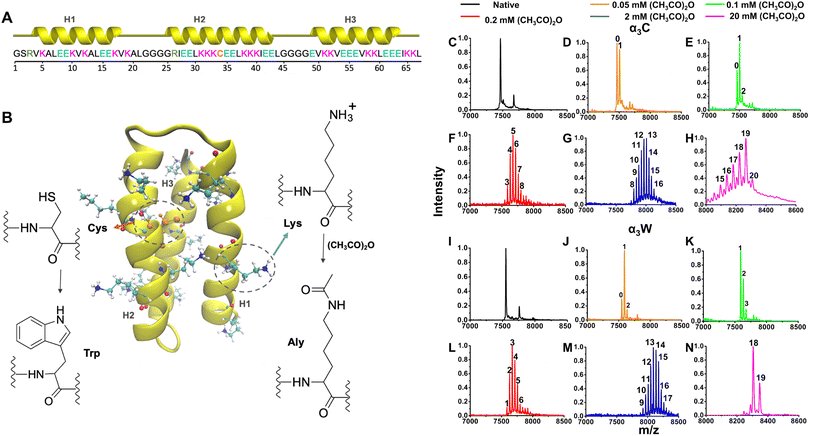
We designed a general computational workflow (SI Fig. F4 and Section M1.2) based on MD simulations and TDDFT calculations to generate the ProCharTS profile of proteins from their 3D structure. We apply this protocol to native and acetylated forms of α3C and α3W, as detailed below.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 61 were selected as starting structures for α3C (Fig. 1A and B) and α3W, respectively. These proteins are rich in charged residues (17 Lys, 17 Glu, and 2 arginine (Arg)) and contain either a single Cys (α3C) or Trp (α3W) at position 34. Acetylated variants of the proteins were generated by mutating Lys residues to acetyl-Lys (Aly) in the PDB structures (Fig. 1B). A total of six α3C models were generated: native, Ac1, Ac3, Ac5, Ac12 and Ac17 (n = number of acetylated Lys residues in Acn). These systems were chosen to match the dominant protein states obtained at different concentrations of acetic anhydride, as verified by mass spectrometry (Fig. 1C–N). The Lys residues in α3C were progressively acetylated in decreasing order of their relative solvent accessible surface area (rSASA) values (SI Section M1.2.1 and Fig. F5A–F). Furthermore, to test the sensitivity of acetylation sites on protein spectra, three variants for Ac12 were created with three different permutations of acetylated states for three specific residues: Ac12-α3C-v1 (Lys10, Aly32, Aly39), Ac12-α3C-v2 (Aly10, Lys32, Aly39), and Ac12-α3C-v3 (Aly10, Aly32, Lys39). For α3W we generated only the native and fully acetylated models (α3W and Ac17-α3W). In summary, a total of ten native and acetylated protein systems were generated for MD simulations. More details on the generation of the modelled structures are provided in SI Sections M1.2.1 and M1.2.5.
61 were selected as starting structures for α3C (Fig. 1A and B) and α3W, respectively. These proteins are rich in charged residues (17 Lys, 17 Glu, and 2 arginine (Arg)) and contain either a single Cys (α3C) or Trp (α3W) at position 34. Acetylated variants of the proteins were generated by mutating Lys residues to acetyl-Lys (Aly) in the PDB structures (Fig. 1B). A total of six α3C models were generated: native, Ac1, Ac3, Ac5, Ac12 and Ac17 (n = number of acetylated Lys residues in Acn). These systems were chosen to match the dominant protein states obtained at different concentrations of acetic anhydride, as verified by mass spectrometry (Fig. 1C–N). The Lys residues in α3C were progressively acetylated in decreasing order of their relative solvent accessible surface area (rSASA) values (SI Section M1.2.1 and Fig. F5A–F). Furthermore, to test the sensitivity of acetylation sites on protein spectra, three variants for Ac12 were created with three different permutations of acetylated states for three specific residues: Ac12-α3C-v1 (Lys10, Aly32, Aly39), Ac12-α3C-v2 (Aly10, Lys32, Aly39), and Ac12-α3C-v3 (Aly10, Aly32, Lys39). For α3W we generated only the native and fully acetylated models (α3W and Ac17-α3W). In summary, a total of ten native and acetylated protein systems were generated for MD simulations. More details on the generation of the modelled structures are provided in SI Sections M1.2.1 and M1.2.5.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 snapshots for each system, which were taken forward for stability analysis70 and subsequent sampling of chromophores. The detailed equilibration protocol and stability analysis are provided in SI Sections M1.2.2 and M1.2.3.
000 snapshots for each system, which were taken forward for stability analysis70 and subsequent sampling of chromophores. The detailed equilibration protocol and stability analysis are provided in SI Sections M1.2.2 and M1.2.3.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 snapshots in native (α3C and α3W) and fully acetylated (Ac17-α3C and Ac17-α3W) protein trajectories (Fig. F7 and F8). For each residue fragment in a cluster, the backbone of the adjacent residues was also extracted to preserve local geometry. For residue pairs that are non-adjacent in sequence, both the N − 1 and N + 1 backbone atomic positions were capped with hydrogens to create symmetric terminal methyl (CH3) groups, thereby neutralizing dangling bonds and mimicking the extended backbone environment of the native protein.46 For nearest-neighbor residue clusters, the peptide linkage between the residues was retained and only the N − 1 backbone atomic position of the first residue and the N + 1 backbone atomic position of the last residue were capped.
000 snapshots in native (α3C and α3W) and fully acetylated (Ac17-α3C and Ac17-α3W) protein trajectories (Fig. F7 and F8). For each residue fragment in a cluster, the backbone of the adjacent residues was also extracted to preserve local geometry. For residue pairs that are non-adjacent in sequence, both the N − 1 and N + 1 backbone atomic positions were capped with hydrogens to create symmetric terminal methyl (CH3) groups, thereby neutralizing dangling bonds and mimicking the extended backbone environment of the native protein.46 For nearest-neighbor residue clusters, the peptide linkage between the residues was retained and only the N − 1 backbone atomic position of the first residue and the N + 1 backbone atomic position of the last residue were capped.
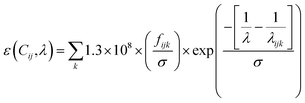 | (1) |
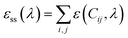 | (2) |
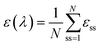 | (3) |
As discussed previously, we use N = 50 for all protein systems except for Ac3-α3C, where N = 25.
3. Results
3.1 Chemical acetylation of α3C and α3W and confirmation using mass spectrometry
We incubated α3C and α3W in acetic anhydride solutions and used mass spectrometry to quantify the number of acetylations in the two proteins as a function of acetic anhydride concentration. The mass of native α3C and α3W (Fig. 1C and I), as determined by MALDI-ToF, is consistent with the expected protein mass based on sequence. Both proteins contain 18 amino groups (17 ε-groups of Lys and 1 N-terminal), which can be potentially acetylated. Since a single acetyl group adds 42 Da to the parent protein, the increment in the protein mass relative to the native protein reveals the number of acetylated residues. The MALDI-ToF data for the proteins in the presence of acetic anhydride in Fig. 1 show multiple m/z peaks, which correspond to a distribution of protein species with different numbers of modified residues (molecular mass values for the peaks are given in SI Fig. F11 and F12). We observe two peaks in the mass spectrum for α3C in 0.05 mM acetic anhydride (Fig. 1D), corresponding to the native unacetylated protein (∼7464 Da) and to a population with one residue modified (∼7506 Da). As the concentration of acetic anhydride is increased to 0.1 mM, three peaks are observed that reveal a decrease in the native population and acetylated protein populations with one and two modified residues (Fig. 1E). At higher acetic anhydride concentrations, a broad distribution of peaks is observed (Fig. 1F and G) indicating the presence of multiple protein species. While species with 3–7 modified residues (5 acetylations for the dominant species) are observed at 0.2 mM acetic anhydride, species with 8–16 modified residues (12/13 acetylations for the largest populations) are seen for 2 mM acetic anhydride. Finally, to achieve hyperacetylated protein, 20 mM acetic anhydride was used, which modifies 17–20 residues (Fig. 1H). Since there are only 18 amino groups in α3C, one or two residues other than Lys appear to be modified. Acetic anhydride has also been reported to target the hydroxyl groups of Ser, Thr, and tyrosine (Tyr), as well as the sulfur group of Cys.73,74 In α3C, a Ser and a Cys residue are present that are presumably targeted by acetic anhydride, resulting in their acetylation. Between these two residues, the Cys side chain (RSH) is intrinsically more nucleophilic than the Ser side chain (ROH). However, because acetylation occurs within the folded protein rather than in free solution, steric effects and relative accessibility to acetic anhydride would also be important factors. A comparison of the relative SASA (rSASA) of the Ser and Cys side chains using MD trajectories of α3C before and after acetylation reveals that Ser remains moderately exposed to solvent but exhibits a significantly broader thermal distribution of accessibilities upon acetylation (rSASA of 35.7 ± 8.0% in native protein versus 36.6 ± 25.3% in Ac17-α3C). On the other hand, the relative SASA for Cys changes from (0.1 ± 0.1) % in native protein to (0.8 ± 0.7) % upon full Lys acetylation. Clearly Cys is deeply buried and is not very accessible to the solvent either before or after acetylation in MD simulations. Nevertheless, there is a relative increase in accessibility for the thiol (–SH) group post-acetylation of the 17 Lys residues. Here, the reactivity and relative accessibility of the –ROH and –SH groups have opposing influences on the acetylation potential of Ser and Cys. Based on the present analysis, it is therefore not possible to determine which of these residues is more likely to be acetylated within the folded protein environment.Similarly, α3W was progressively chemically acetylated by adding 0.05, 0.1, 0.2, 2, and 20 mM acetic anhydride. With 0.05 mM acetic anhydride, species with one modified residue is predominantly generated along with a small population of native species and an even smaller population with two modified residues (Fig. 1J). Increasing the acetylating agent concentration to 0.1 mM results in three m/z peaks, which show 1–3 modified residues, with the predominant species showing a single acetyl modification (Fig. 1K). When the acetic anhydride concentration is increased further, the distribution of modified mass peaks progressively changes, first exhibiting four prominent peaks with a range of 3–6 modified residues (0.2 mM) and then showing seven peaks with 10–16 modified residues (2 mM) (Fig. 1L and M). Finally, with a 20 mM concentration of acetic anhydride, α3W species with 18 and 19 modified residues were generated (Fig. 1N). Here, 19 modifications in the protein can only be explained if a residue other than Lys is modified. In α3W, Cys is absent; however, a Ser residue is present which along with the 18 amino groups accounts for all modifications observed.
3.2 Acetylation does not significantly alter the secondary structure of α3C and α3W but leads to weakening of tertiary interactions
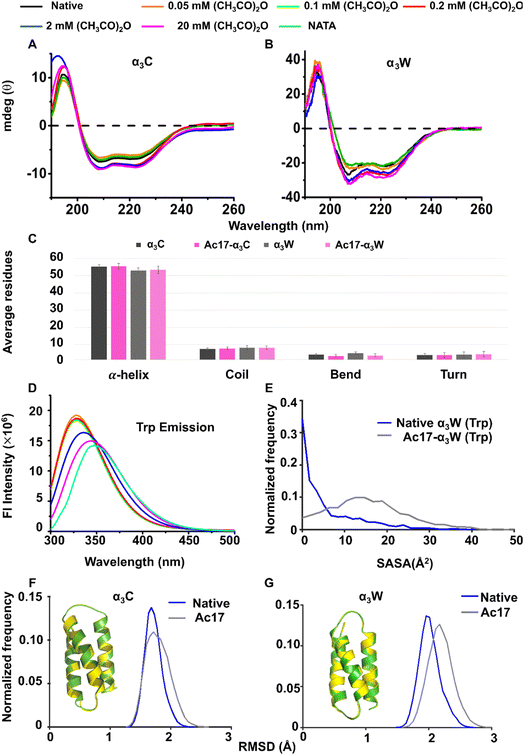
Circular dichroism (CD) spectroscopy (190–260 nm) was used to investigate changes in the secondary structure of α3C and α3W arising due to the reaction with acetic anhydride. Native α3C and α3W spectra (Fig. 2A and B) show two negative bands at 208 and 222 nm and a positive band at 195 nm, which indicates a predominantly alpha-helical protein secondary structure. The spectra do not change significantly for either protein after treatment with acetic anhydride (0.05–20 mM). Specifically, the CD spectra for α3C and α3W at low concentrations of acetic anhydride (0.05 and 0.1 mM) nearly overlap with those of the native protein. Even at higher concentrations (0.2, 2 and 20 mM) of acetic anhydride, the spectra show only minor changes indicating a slight increase in the helical content for both proteins (Fig. 2A and B). An analysis of the CD spectra using the K2D3 web server60 shows a consistent 95% helicity for α3C and α3W independent of acetic anhydride treatment (SI Table T4). Dictionary of Secondary Structure Prediction (DSSP) analyses on production MD trajectories also show that the coil, bend, turn, and alpha-helix elements of the protein are preserved following acetylation for both α3C and α3W (Fig. 2C).We next used the fluorescence intensity of Trp34 in α3W to probe potential local changes in the protein structure after acetylation. NATA (a derivative of Trp), which showed emission maxima at 347 nm, was used as a standard control. Trp34 was excited at 280 nm and its peak emission intensity was observed at 327 nm in the native protein. The emission maximum remained fixed for lower concentrations (0.05 and 0.1 mM) of acetic anhydride but exhibited a systematic red shift at higher concentrations to 329 nm (0.2 mM), 336 nm (2 mM) and 345 nm (20 mM), accompanied by a decrease in intensity (Fig. 2D). These data indicate that the indole ring of Trp34 in native α3W is buried in the hydrophobic core, and as more residues get acetylated, it gradually becomes more exposed to the polar aqueous environment. We note that, relative to the control NATA, the emission peaks for all systems are blue-shifted, indicating that even Ac17-α3W does not have a fully exposed Trp. Additionally, the integrated area of fluorescence emission for native and acetylated α3W indicates that the fluorescence emission does not change significantly upon acetylation relative to the native protein (SI Fig. F13). Fluorescence anisotropy and time-resolved data provide further insights into changes in Trp34 indole rotational motion75 and the local environment induced by acetylation. The steady state fluorescence anisotropy of the indole chromophore in native α3W was measured to be 0.083 and is sensitive to acetic anhydride concentrations (SI Fig. F14 and Table T5). The Trp34 anisotropy is comparable to the native protein at lower concentrations (0.05,0.1, and 0.2 mM) but decreases significantly at higher acetic anhydride concentrations (2 and 20 mM), where between 10–16 acetyl groups are added to the protein (Fig. 1M). The large number of modifications leads to a significant exposure of Trp34 to the solvent, which should increase its rotational mobility and decrease fluorescence anisotropy. Note that the Trp34 anisotropy in α3W is always higher than the anisotropy of NATA, as the former is linked to a polypeptide chain, unlike the latter, which can freely tumble. The anisotropy data are summarized in Table S5. Trp fluorescence intensity decay for both native and acetylated α3W (20 mM acetic anhydride) show two lifetimes, which are assigned to different Trp rotamers (SI Fig. F15 and Table T6). The native protein exhibits a mean lifetime that is slightly lower than NATA (SI Table T6). The amplitudes of native and fully acetylated (24-hour dialysis) samples are different, which reflects the differences in the exposure of Trp for the two cases, in agreement with the steady-state fluorescence data. The mean fluorescence lifetime of the native and fully acetylated (24 hours dialysis) sample are nearly the same (∼2.5 ns), consistent with nearly equal integrated areas of their emission spectra (SI Fig. F13).
Computational SASA analysis (Fig. 2E) for the Trp residue in α3W and Ac17-α3W MD trajectories reveal that acetylation indeed increases Trp solvent exposure, as shown by the broader distribution and a shift toward higher SASA values for Ac17-α3W relative to the native protein. An analysis of spatially proximal residues further reveals that two Lys residues (19 and 38) are located within 5.5 Å of Trp34 in ∼50% frames in the α3W production trajectory. Notably, Lys38 forms a stabilizing salt bridge with Glu35 in ∼66% frames in the native protein, which is disrupted in Ac17-α3W and weakens the tertiary fold near Trp. Our analysis reveals more global changes created by acetylation as well. The protein backbone for both α3C and α3W proteins tends to get slightly more flexible upon acetylation as seen by the distribution of RMSD (backbone atoms) of structures in the production MD trajectories (Fig. 2F and G). The distributions tend to get a bit broader and with a slight peak shift for both α3C and α3W upon acetylation. Finally, the radius of the gyration (Rg) value increases systematically as we increase the number of Aly residues (SI Table T8 and Fig. F16) in both α3C and α3W, indicative of a more expanded structure post-acetylation. Taken together, the computational results support the hypothesis of a loosely packed tertiary structure in α3W created by acetylation.
3.3 Acetylation makes the surface of α3C and α3W more polar and negatively charged
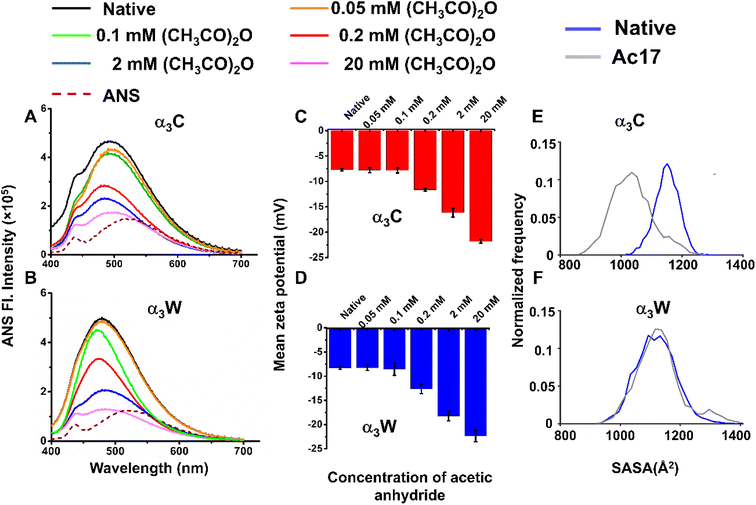
The sequence of native α3C and α3W is rich in polar, charged residues. However, the number of positive and negatively charged groups are balanced (17 Lys, 17 Glu, and 2 Arg) leading to a relatively small net charge of +2. Acetylation of Lys by acetic anhydride treatment is expected to significantly change the surface charge and polarity of the proteins, which we probe experimentally and computationally in this subsection. First, we use ANS, a hydrophobic probe that can bind to solvent-accessible clusters of non-polar atoms.76 Specifically, acetylated proteins (in 0.05–20 mM acetic anhydride concentrations) were incubated with ANS, and the fluorescence intensity of the probe was measured. In the native α3C, the ANS emission maximum appears at 490 nm, more intense than and blue shifted from the free ANS maximum at 525 nm, which confirms the binding of ANS to the hydrophobic core of the protein. With increasing acetylation, the emission maxima remain almost fixed with a slight blue shift (484 nm) at acetic anhydride concentrations of 0.2 and 2 mM (Fig. 3A). However, the emission peak intensity decreases with increasing acetic anhydride concentrations (≥0.2 mM), approaching that of free ANS. In α3W, the emission maxima in the native protein (479 nm) and its acetylated forms (for acetic anhydride concentrations ranging 0.05–20 mM) are blue-shifted relative to free ANS. The blue shift and reduction in the peak intensity above a threshold concentration (0.1 mM) of acetic anhydride were evident, while the emission spectra approach that of free ANS at 20 mM acetic anhydride concentration (Fig. 3B). These observations suggest that subjecting the protein to acetic anhydride at concentrations above a threshold reduces ANS binding in acetylated α3C and α3W.We employed further experimental and computational measurements to understand the mechanisms by which acetylation reduces ANS binding to the proteins. In addition to anilino-naphthalene, a hydrophobic group, ANS also possesses a charged sulfonate group77,78 which can bind to the positively charged Lys amino moiety. As the concentration of acetic anhydride increases, more Lys residues are derivatized with the acetyl group, making the surface more negative and creating unfavorable electrostatics for the ANS sulfonate ions. An examination of the overall charge of native and acetylated α3C/α3W by measuring their zeta potentials79 reveals that the acetylated proteins have more negative values as compared to the native form. In α3C, addition of 0.2, 2, and 20 mM acetic anhydride changes the mean zeta potential from −8.1 ± 0.7 (native protein) to −11.5 ± 1.3, −16.1 ± 0.9 and −21.4 ± 1.5, respectively (Fig. 3C). Similarly, in α3W (mean zeta potential of −8.3 ± 0.2), after acetic anhydride treatment at the same three concentrations, the zeta potentials shift to −12.6 ± 0.98, −18.2 ± 1.0 and −22.3 ± 1.1, respectively. However, at lower concentrations (0.05 and 0.1 mM), the values of mean zeta potential closely match those of native proteins with α3C showing −7.8 ± 0.5 and −7.9 ± 0.5 mV, and α3W showing −8.2 ± 0.5 and −8.5 ± 1.2, respectively (Fig. 3D). These data confirm that the protein surface becomes more negative post-acetylation. Furthermore, SASA analysis on native and acetylated proteins shows that the hydrophobic residues (alanine (Ala), glycine (Gly), valine (Val), isoleucine (Ile), leucine (Leu), phenylalanine (Phe), and methionine (Met)) are also less exposed to the solvent post-acetylation (Fig. 3E and F). The SASA distribution for hydrophobic residues in native α3C (blue line) is shifted to higher values compared to that of acetylated α3C (gray line). While the corresponding distribution for the native α3W (blue line) is also shifted to higher SASA values relative to the acetylated α3W (gray line), the shift is smaller as compared to α3C. These observations correlate with the higher threshold concentrations of acetic anhydride needed for α3W to reduce ANS binding in experiments. Furthermore, the red-shifted ANS emission maximum position in α3W (463 nm) compared to α3C (446 nm) when acetylated with 20 mM acetic anhydride also indicates that ANS resides in less non-polar regions of α3W. In summary, for both proteins (α3C and α3W), acetylation appears to reduce the solvent accessibility of hydrophobic residues and increase the surface negative charge, leading to the observed weaker binding of ANS.
3.4 The ProCharTS UV-vis absorption of α3C and α3W proteins can track their progressive chemical acetylation
The experimental absorption spectra (250–800 nm) of native and acetylated α3C and α3W in different concentrations of acetic anhydride (0.05, 0.1, 0.2, 2 and 20 mM) are compared in Fig. 4. The broad absorption profile (black curve in Fig. 4A) is characteristic of native α3C and has been previously assigned by us to PICT in closely clustered charged Lys and Glu residues within the protein fold.44,46 As the two proteins are exposed to increasing concentrations of acetic anhydride, they undergo progressive acetylation of the Lys residues (SI Fig. F17), which in turn alters the ProCharTS profiles (Fig. 4A–D). While α3C is devoid of any aromatic residues, α3W has a single Trp that shows strong absorption from 280–295 nm, which overrides changes in this wavelength range observed after acetylation (Fig. 4A and B). On the other hand, the ProCharTS from 370–800 nm for α3W is free of any contributions from Trp and matches that from α3C. In both proteins, the ProCharTS responds to acetylation only above a threshold acetic anhydride concentration of 0.2 mM, essentially overlapping with the spectra of the native protein at lower (0.05 and 0.1 mM) concentrations. Beyond this threshold, the ProCharTS intensity across all wavelengths, monotonically drops with increasing concentrations of acetic anhydride for both α3C (Fig. 4C) and α3W (Fig. 4D). The threshold acetic anhydride concentration (0.2 mM) required for ProCharTS to register a change suggests a sensitivity limit of 3–7 Lys residue modifications. Note that at shorter wavelengths (300–400 nm), the experimental spectra for α3C show well resolved shifts (Fig. 4A and C) even at lower concentrations of acetic anhydride (0.05–0.1 mM) where only a single Lys site is acetylated in the protein. However, in α3W, this sensitivity at shorter wavelengths is reduced due to the strong absorption features of Trp (Fig. 4B).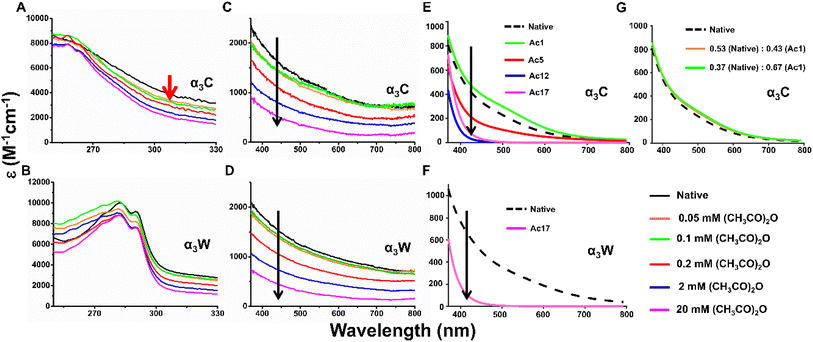 | ||
| Fig. 4 UV-visible absorption spectra track progressive acetylation in α3C and α3W. Experimental (A–D) absorption spectra between 250–330 nm (A and B) and 370–800 nm (C and D) of native/acetylated α3C and α3W. All acetylation reactions are performed with 20 µM protein in 100 mM HEPES buffer at pH 8 in 4 °C, with varying concentrations of acetic anhydride: 0 (native), 0.05, 0.1, 0.2, 2 and 20 mM. The absorption spectra of native/acetylated samples (∼9–10 µM protein) were recorded in deionised water. In α3C, the absorption is sensitive to low levels of acetylation between 300–400 nm (red arrow in panel A and short wavelength range in panel C). The presence of Trp masks this sensitivity at low acetic anhydride concentrations (B–D). Computed absorption spectra (370–800 nm) of native/acetylated proteins (E and F). Computed spectra for either the most abundant or one of the highly populated species (see Fig. 1C–N) present in the experiments at different concentrations of acetic anhydride (different colors in panels E and F). Panel G compares the computed α3C native protein spectra with that obtained from a weighted average spectrum from α3C and Ac1-α3C, as per experimental conditions (Fig. 1D–E) with 0.05 mM (orange) and 0.1 mM (green) acetic anhydride. | ||
Our computational data (Fig. 4E–G) confirm that the decrease in absorption intensity for both α3C and Ac17-α3W originates from the addition of the acetyl group to the Lys sidechains in the proteins that contribute strongly to the ProCharTS intensity. As shown in Subsection 3.3, the overall positive charge of the protein is progressively reduced with increasing acetic anhydride concentrations. Our mass spectrometry results (Fig. 1D and E) show that at low acetic anhydride concentrations (0.05 and 0.1 mM) only a single Lys residue is modified, and this species co-exists with a significant population of native species. Our calculations show a pronounced decrease in ProCharTS intensity for Ac5-α3C, Ac12-α3C, Ac17-α3C (Fig. 4E) and Ac17-α3W relative to the native protein (Fig. 4F) consistent with experimental data (Fig. 4C and D) at higher acetic anhydride concentrations (≥2 mM). The insensitivity of the spectra above 400 nm at lower concentrations is also captured by the computed spectra. In Fig. 4G, we compare the weighted average spectra (orange and green data) from native α3C and Ac1-α3C, with weights corresponding to the m/z peak ratios for the protein with 0.05- and 1-mM acetic anhydride (Fig. 1D and E). The weighted average spectra nearly overlap with that from the native α3C, reproducing the observed insensitivity of ProCharTS at lower concentrations (Fig. 4C). Our calculations predict that the spectra from Ac1-α3C (Fig. 4G) and Ac3-alone (SI Fig. F18) appear to be slightly more intense than native α3C. We are not able to comment on whether this increase is significant enough to resolve the species with the present set of calculations and experiments. The spectral changes induced by acetylation can be better understood by the UV-vis response of smaller charged residue clusters (SI Fig. F19) extracted from the Ac17-α3C MD trajectory. We find that ProCharTS in native protein clusters above 370 nm is primarily composed of SS-CT transitions between oppositely charged head groups of Lys and Glu (SI Fig. F20A and B). Upon acetylation of Lys, these low energy transitions are replaced with higher energy BS-CT transitions between the charged carboxylate and backbone of Glu (SI Fig. F20C). Note that the computed spectra in Fig. 4F and G are not expected to quantitatively reproduce the experimental data in Fig. 4C and D, since the former is calculated for a single species of protein with a distinct acetylated state, whereas the latter arises from the contribution of multiple species. A detailed analysis of these considerations is presented in the Discussion section.
We observe that the computed ProCharTS profile for Ac-α3C and Ac-α3W is not significantly altered if Aly is included along with charged residues in the clusters (SI Fig. F21A and B). This indicates that the neutral residue has a minimal impact on ProCharTS above 250 nm. This was further confirmed by computing the absorption spectra of small clusters (dimer/hexamer) of charged residues with and without Aly (SI Fig. F19). The computed spectra of the Aly–Glu dimer (blue line) are nearly identical to that of isolated Glu taken from the same dimer. The minute differences seen are probably due to electrostatic effects coming from Aly, which are absent for the Glu monomer. In the case of the hexamer, the cluster upon acetylation reduces to a Glu dimer and an isolated Glu monomer (with Rc = 6 Å). In SI Fig. F19D, cluster spectra with and without Aly show differences below 450 nm, as the latter misses the electrostatic influence of three Aly residues. While the red-most transition, BS-CT from the negatively charged carboxylate head group to the backbone of Glu, remains the same independent of inclusion of Aly (SI Fig. F20C and D), the transition is redshifted in the presence of Aly, which further confirms that electrostatic influence of the neutral residue. Nevertheless, the effect of acetylation in reducing the spectra is fully captured independent of whether Aly is included or not. These calculations further strengthen our model of including only charged residue clusters to study the effect of acetylation on ProCharTS.
3.5 ProCharTS luminescence is not an effective probe of chemical acetylation in α3C and α3W
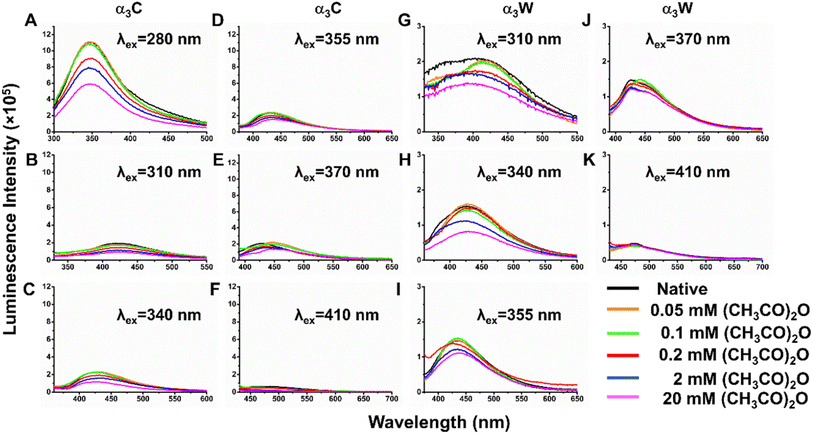
Luminescence can arise when charges separated in the excited state, subsequent to charge transfer in charge-rich proteins, undergo charge recombination. ProCharTS luminescence intensities have previously been shown to correlate with population of proximal charges within the protein.54 We therefore examined the possibility of using ProCharTS luminescence features to track acetylation in α3C and α3W. Since ProCharTS arises from the absorption of multiple chromophores (charged residue clusters) over a broad wavelength range, the resultant emission is excitation dependent. We therefore excited the native and acetylated forms of the proteins at multiple wavelengths and recorded the emission as a function of wavelength (Fig. 5). The ProCharTS luminescence in native form of α3C and α3W is retained at low acetic anhydride concentrations (0.05 and 0.1 mM) and shows reduction in intensity as acetylation increases (for acetic anhydride concentrations ≥0.2 mM) at all the excitation wavelengths. However, the decrease is not systematic with increasing acetic anhydride concentrations with trends highly sensitive to the excitation wavelength.To determine the primary cause behind the decrease in the luminescence intensity after acetylation, we calculated the integrated quantum yield (QY) of both native and acetylated forms of α3C/α3W relative to reference 9,10-diphenylanthracene (DPA) and NATA excited at 355 and 280 nm, respectively. In α3C, the luminescence from 280 nm excitation exhibits a significant reduction in quantum yield as the concentration of acetic anhydride increases beyond 0.2 mM but is nearly similar to that of the native protein at lower concentrations (SI Fig. F22). This suggests that the charge recombination is lowered post-acetylation with higher concentrations of acetic anhydride. However, the luminescence QY with 355 nm excitation in both the proteins is insensitive to the degree of acetylation (SI Table T7), which indicates that the reduction in luminescence cannot be attributed to the changes in QY. Rather, differences in protein absorption convoluted with different radiative and non-radiative relaxation mechanisms of the excitations lead to the complex changes in the luminescence of α3C and α3W post-acetylation. Furthermore, the sensitivity of the emission spectra is the same as that for the absorption profiles, both not responsive at low concentrations (≤0.1 mM) of acetic anhydride. Based on these considerations, we conclude that ProCharTS luminescence does not directly shadow charge neutralizing modifications of residues and is therefore not as effective as absorption for tracking PTMs such as acetylation in proteins.
4. Discussion
A direct label-free optical spectroscopic detection of PTMs offers many advantages in comparison to conventional techniques such as mass spectrometry or antibody-based detection. Absorption and emission spectroscopy are some of the most accessible characterization techniques in protein biophysics and biochemistry laboratories around the world. Optical techniques are least perturbative and can report on functional assays in the solution phase.49,51,80 Additionally, extensions to in vivo measurements and more advanced non-linear (ultrafast) spectroscopy is also possible.81,82 Recently, there has been growing interest in using spectroscopy to probe the collective optical properties of amino acids in supramolecular architectures and utilize them in sensing and therapeutic applications.83 However, studies in this direction have been restricted to aromatic amino acids (Phe, Tyr, and Trp). From the vantage point of protein biophysics and biochemistry, it would therefore be very useful to extend optical spectroscopy to detect PTMs in protein solutions by establishing distinct chemical signatures of modified and unmodified residues. In this direction, recently, vibrational markers were proposed for acetylated and non-acetylated forms of Lys using Raman and FTIR spectroscopy of powdered samples of the amino acids in combination with DFT calculations.41 However, these vibrational fingerprints and their sensitivity to the number of PTMs remain to be tested in the solution phase. Here, we show for the first time that emergent charge transfer transitions from collective excitations of charged residues provide distinctive signatures in electronic absorption spectroscopy to detect PTMs in the solution phase. We have demonstrated that ProCharTS absorption profiles of α3C and α3W show a monotonic decrease in intensities between 370–800 nm with progressive chemical acetylation of the proteins in acetic anhydride solutions. Mass spectrometry (Fig. 1C–N) confirms that between 1–19 acetyl groups are added to both α3C and α3W, which are assigned to modifications of 17 Lys residues, the protein N-terminus, and the single Ser residue in the proteins. Additionally, a weak signal for an extra acetyl group in α3C is assigned to the buried Cys residue in the protein. Since the ProCharTS profiles are broad and featureless, we developed a computational framework to simulate and deconvolute the spectral profile in terms of the underlying charged residue chromophores. This provides us with new insights to develop optical signatures of acetylation in terms of PICTs arising within the protein fold, as discussed below.An analysis of the computed spectra of α3C/α3W shows that the addition of acetyl groups to Lys residues significantly alters the clustering among charged residues to influence the spectra. For instance, as illustrated in SI Fig. F7 and 8, acetylation leads to an increase in the number of charged residue monomers in Ac17-α3C compared to α3C, accompanied by a corresponding decrease in the size of larger clusters, particularly tetramers to heptamers. For other acetylated systems, we find a wide range of clusters, all of which are uniformly populated with charged residues (SI Fig. F23A–E(i)) when five or less residues are acetylated. However, as the degree of acetylation progresses and charged Lys residues are neutralized, we see a gradual increase in the population of monomers and dimers at the expense of the larger clusters (SI Fig. F23A–E(ii) and Table T10, 11). Eventually, when all Lys residues are acetylated, we find most of the residues (∼85%) in monomeric form for Ac17-α3C (SI Fig. F23F(i and ii)). We find that the decrease in spectral intensity is correlated with the total number of monomers, dimers and trimers (SI Fig. F28) linking the decrease in the ProCharTS intensity of spectra post-acetylation to the reduction in the clustering propensities of oppositely charged Lys and Glu residues.
In previous computational studies, we had shown that the excess charge on the side chains of residues such as Lys and Glu creates a polarization of frontier filled orbitals to create an electronic donor–bridge–acceptor (D–B–A) molecular architecture.45 As a result, while monomeric Lys or Glu residues absorb in the deep UV, they show facile backbone-side chain (BS) PICT transitions. More interestingly, when these D–B–A residues (Lys and Glu) electrostatically interact through their charged head groups, both the nature of the PICT and their absorption wavelengths change. An oppositely charged Lys–Glu dimer, which is separated by 5–6 Å shows six different inter/intra-residue PICTs including visible side chain to side chain (SS) PICT from the Glu carboxylate group to the Lys amino group.46 SI Fig. F29 shows that the decrease in spectral intensity is strongly correlated with the number of Lys acetylated and the decrease in Lys–Glu dimers, which are separated by more than 4 Å. Thus, the intensity decrease is a direct effect of Lys charge neutralization, which is in accordance with previous findings.44 Thus, based on these results, we hypothesize that the classic ProCharTS profile of α3C in the near UV and visible regions arises from the contribution of oppositely charged residue dimers and higher order clusters. Below we carry out a deconvolution of the simulated spectra of native α3C to validate this hypothesis, which allows us to connect the spectral differences post-acetylation to the underlying changes in PICT processes.
To fully understand the contributions of charged residue clusters to ProCharTS, we first decomposed the computed spectral profiles of native and acetylated α3C into contributions from individual snapshots from the MD production trajectories. Fig. 6A and B showcase the final averaged spectra (black line) of native α3C and Ac17-α3C against a backdrop of contributions (different colours) arising from 50 snapshots from the production runs. Similar analysis for other acetylated systems is shown in SI Fig. F24. Interestingly, while the averaged ProCharTS profile for native α3C is featureless, 46% of the MD snapshots show a rich array of features including clearly defined peaks ranging from the UV to the visible (Fig. 6A and SI F25). Strikingly, in fully acetylated Ac17-α3C, all features disappear and the resultant weak ProCharTS absorption is comprised of featureless contributions from individual snapshots (Fig. 6B). Further analysis reveals that oppositely charged dimers contribute significantly to the spectra exhibiting peak features (featured spectra). For instance, the spectral profile from a representative snapshot (black arrow in Fig. 6A) arises predominantly from dimer and trimer contributions between 350–450 nm and solely from dimers above 450 nm (Fig. 6C). In this specific snapshot, there are a total of five oppositely charged dimers present but only two of them with separation distances between 5–6 Å exhibit SS-PICT transitions, which contribute to the spectra beyond 450 nm (Fig. 6D), along with a mixture of SS-PICT and BS-PICT transitions, which contribute below 450 nm. Extending this analysis to the full set of MD snapshots, we find that isolated oppositely charged dimers predominantly contribute to the native α3C absorption profile above 450 nm in more than 50% of the snapshots exhibiting featured spectra (Fig. 6E). In remaining snapshots, which exhibit featureless spectra, dimers and trimers both predominantly contribute (Fig. 6F). Similar analysis for acetylated α3C systems is also shown in SI Fig. F26 and 27. The ratio of snapshots showing featured versus those showing featureless spectra decreases as α3C is progressively acetylated and falls rapidly when 5 or more Lys residues are acetylated.
Interestingly, while the averaged absorption spectrum decreases with increasing acetylation, it does not exhibit any peak features in any of these systems despite possessing multiple snapshots with featured spectra. When the degree of acetylation is low (Ac1- and Ac3-α3C), oppositely charged dimers still dominate the spectral intensity above 450 nm and produce a slight enhancement in the spectra relative to native α3C (Fig. 4G and SI F18A). Only when five or more Lys residues are acetylated, do we observe a significant decrease in the number of snapshots showing featured spectra (SI Fig F24 and F26). Furthermore, even for the featureless spectra, the tails beyond 500 nm reduce with increasing acetylation. We find that featureless spectra in all systems arise predominantly from isolated Glu monomers and either neutral (Lys–Glu pairs forming a salt-bridge) or negatively charged Glu dimers (SI Fig. F26 and 27). These DBA systems lack SS-PICT transitions and show only BS-PICT or backbone-backbone (BB) PICT44,46 This is also true for the featured spectra, where we find dimers and trimers lacking SS-PICT transitions contributing to the spectra below 450 nm. Taken together, the analysis here shows that long range (5–6Å) SS-PICT transitions in oppositely charged dimers are responsible for the extended absorption feature above 450 nm exhibited by the native protein. With increasing acetylation, these long-range PICT transitions are turned off due to the reduction in population of oppositely charged residue pairs which manifests in the observed drop in ProCharTS intensity in the visible. Given the broad features of the spatiotemporally averaged ProCharTS profile (Fig. 4) and experimental limitations in selectively acetylating specific residues during chemical acetylation assays, which result in additional averaging over multiple species (in terms of acetylation sites), it is not possible to directly verify the transient clusters predicted here by computations. However, our sensitivity analysis of Ac12-α3C (SI Section M1.2.5 and Fig. F18) indicates that the ProCharTS profile may be much more sensitive to PTMs at certain sites in relation to others, something that can be inferred from computational screening and then verified by experiments that can carry out selective single point perturbations.
Charge neutralization of Lys (acetylation or any such PTMs) can create a number of changes, both local and global, in the protein structure, which impact the spectra to differing extents. Some changes are universal, such as changes in the composition and size distribution of charged clusters, tertiary interactions, and changes in the overall charge of the protein. Others such as changes in the protein secondary structure, conformational changes, or changes in function (e.g. enzymatic activity) are very protein specific. The sensitivity of the protein spectral changes to PTMs will therefore also be protein specific depending on whether these effects act in a concerted or opposing mode to alter the ProCharTS activity. Both computational and experimental techniques are required to understand the interplay of these effects and deconvolute them. In the present manuscript, we have attempted to do precisely this for our model systems as noted below. For our model protein systems, the CD, Trp spectra, and MD simulations (Fig. 2) indicate no major changes in the secondary structure of the protein or the overall protein fold, which leads us to the conclusion that the associated spectral changes do not arise from factors such as backbone conformational changes. On the other hand, these techniques do suggest a weakening of tertiary interactions, which leads to a systematic increase in Rg and can be attributed to changes in the surface charges and polarity of the protein, as indicated by ANS binding and zeta potential measurements (Fig. 3). Furthermore, Trp emission and SASA calculations (Fig. 2D and E) on α3W also indicate local unfolding of the structure, exposing the Trp to solvent post-acetylation. This is attributed from simulations to the disruption of a Lys38–Glu35 salt bridge in the vicinity due to charge neutralization. To more clearly show the impact of these changes on the spectra, we examined the distribution of cluster sizes as a function of acetylation, which clearly indicates a reordering of cluster sizes upon acetylation. Scatter plots (SI Fig. F28) between spectral intensity and number of clusters of smaller size (monomers, dimers and trimers) show that the systematic decrease in the intensity of spectra is correlated to the reduction in the clustering propensities of oppositely charged Lys and Glu residues. The drop in spectral intensity correlates even more strongly (SI Fig. F29 and Tables 8, 9) with protein Rg, the number of Lys acetylated or the number of Lys–Glu dimers, which are separated by more than 4 Å. Essentially, changes in cluster size reordering or reductions in the number of oppositely charged Lys–Glu dimers or even the changes in Rg can all be traced back to charge neutralization (acetylation). As demonstrated here, the combination of computational and experimental analysis can be used to dissect the nature of changes induced by PTMs, which manifest as spectral changes.
Our experimental studies and computational analysis identify a new optical mode employing charge transfer transitions to follow acetylation or indeed any PTM that alters the charged state of residues. We term the new detection mode ProCharTSPTM, which has the potential to be developed into a cheap and viable alternative to techniques such as mass spectrometry and antibody-based assays. The strategy rests on the fact that enhanced sensitivity of ProCharTS to charge altering PTMs is due to the charge transfer (CT) character of the underlying transitions. These show significant shifts and changes in intensity in response to perturbations in residue charge and their clustering.44 In the present study, we note that experimental ProCharTS spectra show noticeable sensitivity (Fig. 4A and B) in detecting lower levels of acetylation (at 0.05 and 0.1 mM acetic anhydride) in the 300–400 nm window. Mass spectrometry data indicate that only 0 and 1 Lys acetylated species are present at these low concentrations of acetic anhydride. Based on the data from α3W, the strong absorbance of tryptophan can mask this sensitivity and could possibly be deconvoluted with the aid of computations as noted above. In order to assess the applicability of ProCharTS to biologically relevant proteins, we carried out chemical acetylation studies on two proteins, the GTPase K-RAS, which regulates cellular responses and the Histone H2A, which regulates gene expression, where acetylation has been shown to play important roles.84,85 Both proteins exhibit a clear systematic monotonic decrease in ProCharTS absorbance with increasing concentrations of acetic anhydride (SI Fig. F31). Additionally, both proteins have a lower predominance of charged residues (∼30–40% of the sequence) and a higher molecular weight compared to α3C and α3W (SI Table T12). These results show that ProCharTSPTM can also be used to detect modifications in biologically relevant proteins, even when the charge content is low.
Despite these promising results, there are limitations on the ProCharTSPTM measurements and analysis, which need to be addressed in order to develop the technique further. We note that there are quantitative differences between the measured and computed spectra (Fig. 4), particularly in terms of intensity, which is lower for the latter, and the sharper drop in the ProCharTS tail for the computational data. Furthermore, the computed data cannot resolve the drop in ProCharTS intensity when more than 12 Lys are acetylated. These differences and limitations can be attributed to the diversity of protein species contributing to the experimental spectra and approximations used in our computational analysis. The mass spectrometry data reveal (Fig. 1C–N) that the experimental ProCharTS signal arises from multiple protein species with different numbers of acetylated residues. Additionally, even for a fixed number of modifications, there are multiple protein species present which differ in terms of acetylation sites. For instance, in Ac1-α3C there are 18 possible amino groups, which can be acetylated. While all modifications sites are not equally accessible, a solvent accessibility analysis (SI Fig. F5A) indicates that the number of degenerate possibilities is still large. Our computational examination of single residue permutations in the Ac12-α3W system shows that the choice of acetylated residue site can impact the ProCharTS profile above 400 nm significantly. Thus, specific acetylation sites could potentially be detected with suitable combinations of experiments and computations. For instance, experiments could be improved to selectively acetylate Lys sites or generate samples with one or a few species at most. Alternatively, the computations could be made more efficient to sample all possible permutations and combinations of PTM sites present in the sample. The prohibitive computational costs of a full detailed MD plus TDDFT framework presented can be mitigated by identifying geometric parameters/feature spaces to generate spectra using AI/ML approaches. Reducing the cost of the computations will also help improve the quality of simulated spectra. For instance, to generate the spectra in Fig. 4 at a reasonable cost, the clustering cut-off parameter Rc was set to 6 Å to restrict cluster sizes in ProCharTS simulations to decamers (Subsection 2.2.4). Increasing the value of Rc will allow larger clusters, which should produce more accurate ProCharTS profiles. Furthermore, our calculations do not include non-charged residues in the spectra simulations, which contribute to the spectra below 300 nm. Finally, even for charged residue clusters computational cost limits the maximum number of calculated transitions to 120 per cluster, covering a spectral range of 250–800 nm. These factors lead to a poor description of the spectra in the UV range (below 380 nm) and can be improved in the future. Moving forward, we suggest the following roadmap to develop ProCharTSPTM into a potent tool for biochemistry and biomedicine:
Stage 1 (Accelerating Computations and Standardizing Experimental Protocols): reducing the computational time to simulate the spectra of ProCharTS (currently two weeks on a high-performance computing cluster) to less than a day. This can be accomplished by developing either AI/ML strategies or a library of precomputed cluster spectra. On the experimental side, creating standard and systematic protocols for sample preparation (including enzyme catalyzed PTMs) that isolate specific ProCharTS active species along with their spectral response to modifications.
Stage 2 (Spectral Deconvolution and Experimental Validation): development of computational strategies to deconvolute the experimental signals arising from multiple proteins present in a sample based on individual spectral fingerprints along with experimental validation.
Stage 3 (Database of ProCharTSPTM Fingerprints): creation of a database of ProCharTS active proteins along with their spectral fingerprints. A comprehensive in silico scan on protein structures can be carried out first followed by experimental validation and spectral characterization. Such a database can be regularly updated to include newly discovered ProCharTSPTM fingerprints.
These steps lay the foundation to develop ProCharTSPTM into a quantitative spectroscopic method for detecting charge-altering PTMs such as acetylation in biological samples.
5. Conclusion
Our results show for the first time that charge transfer transitions can serve as markers to track and study PTMs in charge rich proteins. Here, we use chemical acetylation of two model charge rich proteins α3C and α3W to develop an approach (ProCharTSPTM), which combines simple UV-vis absorption spectroscopy with computational analysis to follow the progressive modifications of their charged residues. Extensive spectroscopic and biochemical characterization enables us to map ProCharTS intensity changes to the number of modified residues in the proteins. The analysis reveals that the ProCharTS profile obtained with different concentrations of acetic anhydride arises from heterogeneous protein populations with varying degrees of acetylation. Joint experiments and MD computational analysis show that while acetylation does not change the secondary structure of the proteins, their tertiary fold is weakened by acetylation. ANS binding studies, zeta potential measurements, and computational analysis indicate that acetylation makes the surface of the protein more polar and negatively charged. Further hydrophobic pockets become less accessible for ANS binding with increasing extent of acetylation. The ProCharTS emission profiles for the proteins, as a function of acetylation, are complex, showing considerable sensitivity to excitation and emission wavelengths. As such, they do not appear to be conducive to track PTM modifications in proteins. We believe that our study sets up an exciting new paradigm for tracking biologically relevant PTMs among charge rich proteins, a biomedically relevant class that includes genome regulating proteins such as histones, intrinsically disordered proteins implicated in neuropathological disorders such as tau, tumour suppressor protein p53 and gene-regulating transcription factor modifications such as TFIIE and TFIIF—proteins that are activated or inactivated by PTMs in complex networks of signaling pathways.Author contributions
Conceptualization: RV and RS; data curation: HMD, AB, AM, and SBS; formal analysis: HMD, AB, and AM; funding acquisition: RV and RS; investigation: HMD, AB, AM, and SBS methodology: HMD, AB, AM, RV and RS; project administration: RV and RS; resources: HMD and AM; software: AB and AM; supervision: RV and RS; validation: HMD, AB, and AM; visualization: HMD, AB, and AM; writing – original draft: HMD and AB writing – review and editing: all authors.Conflicts of interest
There are no conflicts to declare.Data availability
All methodology and supporting data are provided in the main manuscript and supplementary information (SI) file. Additional data for this article, including MD trajectories and analysis scripts to generate the figures, are available at the Zenodo repository at https://doi.org/10.5281/zenodo.18893836.Supplementary information: additional figures, tables, analysis, and details of computational and experimental methods. See DOI: https://doi.org/10.1039/d5sc09293k.
Acknowledgements
HMD, SBS and RS thank the Department of Biotechnology, Government of India, for providing financial support to the North East Centre for Biological Science and Healthcare Engineering (NECBH) through project number BT/NER/143/SP44675/2023. The authors acknowledge the NECBH and Central Instrument Facility (CIF) at IIT Guwahati for providing access to their instruments. AB, AM, and RV acknowledge the funding support from the Department of Atomic Energy (DAE), Government of India, under Project Identification No. 1303/10/2025-R&D-II-DAE/TIFR-17248 RTI 4015. HMD, SBS, and RS thank Prof. Bhubaneswar Mandal, Ms. Subhashree Sahu (Department of Chemistry, IIT-Guwahati), Prof. Sachin Kumar (Department of Biosciences and Bioengineering, IIT-Guwahati) and Dr. Devyani Haldar (CDFD Hyderabad) for providing acetic anhydride, for assisting with lifetime measurements, for gifting K-RAS protein, and for gifting the Histone H2A DNA construct, respectively. The authors thank Prof. Jyotishman Dasgupta (Department of Chemical Sciences, TIFR-Mumbai) for fruitful discussions on the electronic spectral profiles.References
- C. T. Walsh, S. Garneau-Tsodikova and G. J. Gatto, Angew. Chem., Int. Ed., 2005, 44, 7342–7372 CrossRef CAS PubMed.
- R. Aebersold, J. N. Agar, I. J. Amster, M. S. Baker and B. Zhang, et al., Nat. Chem. Biol., 2018, 14, 206–214 CrossRef CAS PubMed.
- S. Ramazi and J. Zahiri, Database, 2021, baab012 CrossRef CAS PubMed.
- B. Macek, K. Forchhammer, J. Hardouin, E. Weber-Ban, C. Grangeasse and I. Mijakovic, Nat. Rev. Microbiol., 2019, 17, 651–664 CrossRef CAS PubMed.
- D. G. Christensen, X. Xie, N. Basisty, J. Byrnes, S. McSweeney, B. Schilling and A. J. Wolfe, Front. Microbiol., 2019, 10, 1604 CrossRef PubMed.
- R. Bell, R. J. Thrush, M. Castellana-Cruz, M. Oeller, R. Staats, A. Nene, P. Flagmeier, C. K. Xu, S. Satapathy, C. Galvagnion, M. R. Wilson, C. M. Dobson, J. R. Kumita and M. Vendruscolo, Biochemistry, 2022, 61, 1743–1756 CrossRef CAS PubMed.
- K. W. Barber and J. Rinehart, Nat. Chem. Biol., 2018, 14, 188–192 CrossRef CAS PubMed.
- Á. Marín-Hernández, J. S. Rodríguez-Zavala, R. Jasso-Chávez, E. Saavedra and R. Moreno-Sánchez, J. Cell. Biochem., 2022, 123, 701–718 CrossRef PubMed.
- C. N. I. Pang, A. Hayen and M. R. Wilkins, J. Proteome Res., 2007, 6, 1833–1845 CrossRef CAS PubMed.
- P. Sorlier, C. Viton and A. Domard, Biomacromolecules, 2002, 3, 1336–1342 CrossRef CAS PubMed.
- M. Liu, L. Guo, Y. Fu, M. Huo, Q. Qi and G. Zhao, Biotechnol. Adv., 2021, 53, 107842 CrossRef CAS PubMed.
- E. L. Gershey, G. Vidali and V. G. Allfrey, J. Biol. Chem., 1968, 243, 5018–5022 CrossRef CAS PubMed.
- S. N. Khan and A. U. Khan, Clin. Chim. Acta, 2010, 411, 1401–1411 CrossRef CAS PubMed.
- J. F. Riordan and B. L. Vallee, in Methods in Enzymology, Elsevier, 1967, vol. 11, pp. 565–570 Search PubMed.
- K. R. Bridges, G. J. Schmidt, M. Jensen, A. Cerami and H. F. Bunn, J. Clin. Invest., 1975, 56, 201–207 CrossRef CAS PubMed.
- M.-M. Wang, D. You and B.-C. Ye, Sci. Rep., 2017, 7, 14790 CrossRef PubMed.
- E. Verdin and M. Ott, Nat. Rev. Mol. Cell Biol., 2015, 16, 258–264 CrossRef CAS PubMed.
- M. D. Shahbazian and M. Grunstein, Annu. Rev. Biochem., 2007, 76, 75–100 CrossRef CAS PubMed.
- S. Zhao, W. Xu, W. Jiang, W. Yu, Y. Lin, T. Zhang, J. Yao, L. Zhou, Y. Zeng, H. Li, Y. Li, J. Shi, W. An, S. M. Hancock, F. He, L. Qin, J. Chin, P. Yang, X. Chen, Q. Lei, Y. Xiong and K.-L. Guan, Science, 2010, 327, 1000–1004 CrossRef CAS PubMed.
- C. M. VanDrisse and J. C. Escalante-Semerena, Annu. Rev. Microbiol., 2019, 73, 111–132 CrossRef CAS PubMed.
- N. Šoštarić and V. Van Noort, PLoS Comput. Biol., 2021, 17, e1008988 CrossRef PubMed.
- S. Deota, S. Rathnachalam, K. Namrata, M. Boob, A. Fulzele, S. Radhika, S. Ganguli, C. Balaji, S. Kaypee, K. K. Vishwakarma, T. K. Kundu, R. Bhandari, A. Gonzalez De Peredo, M. Mishra, R. Venkatramani and U. Kolthur-Seetharam, J. Mol. Biol., 2019, 431, 2127–2142 CrossRef CAS PubMed.
- U. Mahlknecht and D. Hoelzer, Mod. Med., 2000, 6, 623–644 CAS.
- X. Lu, L. Wang, C. Yu, D. Yu and G. Yu, Front. Cell. Neurosci., 2015, 9, 226 Search PubMed.
- M. S. Liyasova, L. M. Schopfer and O. Lockridge, Biochem. Pharmacol., 2010, 79, 784–791 CrossRef CAS PubMed.
- I. Petitpas, C. E. Petersen, C.-E. Ha, A. A. Bhattacharya, P. A. Zunszain, J. Ghuman, N. V Bhagavan and S. Curry, Proc. Natl. Acad. Sci. U. S. A., 2003, 100, 6440–6445 CrossRef CAS PubMed.
- I. Petitpas, A. A. Bhattacharya, S. Twine, M. East and S. Curry, J. Biol. Chem., 2001, 276, 22804–22809 CrossRef CAS PubMed.
- P. A. Zunszain, J. Ghuman, A. F. McDonagh and S. Curry, J. Mol. Biol., 2008, 381, 394–406 CrossRef CAS PubMed.
- C. Krawic, M. W. Luczak and A. Zhitkovich, Chem. Res. Toxicol., 2024, 37, 1588–1597 Search PubMed.
- F. Middleton-Davis, A. Davis and K. Middleton, PLoS One, 2022, 17, e0268887 CrossRef CAS PubMed.
- K. Zhang, S. Tian and E. Fan, Analyst, 2013, 138, 1628 RSC.
- P. Li, Y. Han, Y. Li, R. Zhu, H. Wang, Z. Nie and S. Yao, Anal. Bioanal. Chem., 2016, 408, 2659–2668 CrossRef CAS PubMed.
- S. Liokatis, A. Dose, D. Schwarzer and P. Selenko, J. Am. Chem. Soc., 2010, 132, 14704–14705 CrossRef CAS PubMed.
- M. Kasaai, Carbohydr. Polym., 2008, 71, 497–508 CrossRef CAS.
- S. Ronzoni, M. Faretta, M. Ballarini, P. G. Pelicci and S. Minucci, Cytometry A, 2005, 66, 52–61 CrossRef PubMed.
- E. Hesping, T. S. Skinner-Adams, G. M. Fisher, T. Kurz and K. T. Andrews, Int. J. Parasitol. Drugs Drug Resist., 2020, 14, 249–256 CrossRef PubMed.
- I. Diallo, M. Seve, V. Cunin, F. Minassian, J.-F. Poisson, S. Michelland and S. Bourgoin-Voillard, Expet Rev. Proteonomics, 2019, 16, 139–159 CrossRef CAS PubMed.
- L. A. Murray, A. N. Combs, P. Rekapalli and I. M. Cristea, in Methods in Enzymology, Elsevier, 2019, vol. 626, pp. 587–620 Search PubMed.
- L. E. Smith and A. Rogowska-Wrzesinska, Essays Biochem., 2020, 64, 135–153 CrossRef CAS PubMed.
- A. R. Farley and A. J. Link, in Methods in Enzymology, Academic Press Inc., 2009, vol. 463, pp. 725–763 Search PubMed.
- G. Yao and Q. Huang, Spectrochim. Acta, Part A, 2022, 278, 121371 CrossRef CAS PubMed.
- L. Homchaudhuri and R. Swaminathan, Chem. Lett., 2001, 30, 844–845 CrossRef.
- L. Homchaudhuri and R. Swaminathan, Bull. Chem. Soc. Jpn., 2004, 77, 765–769 CrossRef CAS.
- S. Prasad, I. Mandal, S. Singh, A. Paul, B. Mandal, R. Venkatramani and R. Swaminathan, Chem. Sci., 2017, 8, 5416–5433 RSC.
- I. Mandal, S. Paul and R. Venkatramani, Faraday Discuss., 2018, 207, 115–135 RSC.
- I. Mandal, S. Manna and R. Venkatramani, J. Phys. Chem. B, 2019, 123, 10967–10979 CrossRef CAS PubMed.
- C. González-González, R. Lopez-Blanco, J. A. González-Vera, S. D'Ingiullo, D. Bouzada, M. Melle-Franco, A. Orte and M. E. Vázquez, Cell Rep. Phys. Sci., 2025, 6, 102631 CrossRef.
- M. J. Fossat, A. E. Posey and R. V. Pappu, ChemPhysChem, 2023, 24, e202200746 CrossRef CAS PubMed.
- S. E. Alom, S. Kalita, A. H. Kawa, B. Mandal and R. Swaminathan, Anal. Chim. Acta, 2024, 1297, 342374 CrossRef CAS PubMed.
- M. Z. Ansari, A. Kumar, D. Ahari, A. Priyadarshi, P. Lolla, R. Bhandari and R. Swaminathan, Faraday Discuss., 2018, 207, 91–113 RSC.
- A. Priyadarshi, H. M. Devi and R. Swaminathan, Biochemistry, 2023, 62, 1643–1658 CrossRef CAS PubMed.
- S. E. Alom, K. Swaminathan, V. Nuzelu, A. Singh, H. De Rocquigny and R. Swaminathan, Biomacromolecules, 2024, 25, 6425–6438 CrossRef CAS PubMed.
- N. M. Ennist, S. Wang, M. A. Kennedy, M. Curti, G. A. Sutherland, C. Vasilev, R. L. Redler, V. Maffeis, S. Shareef, A. V. Sica, A. S. Hua, A. P. Deshmukh, A. P. Moyer, D. R. Hicks, A. Z. Swartz, R. A. Cacho, N. Novy, A. K. Bera, A. Kang, B. Sankaran, M. P. Johnson, A. Phadkule, M. Reppert, D. Ekiert, G. Bhabha, L. Stewart, J. R. Caram, B. L. Stoddard, E. Romero, C. N. Hunter and D. Baker, Nat. Chem. Biol., 2024, 20, 906–915 CrossRef CAS PubMed.
- A. Kumar, D. Ahari, A. Priyadarshi, M. Ziauddin Ansari and R. Swaminathan, J. Phys. Chem. B, 2020, 124, 2731–2746 CrossRef CAS PubMed.
- C. Tommos, K. G. Valentine, M. C. Martínez-Rivera, L. Liang and V. R. Moorman, Biochemistry, 2013, 52, 1409–1418 CrossRef CAS PubMed.
- J. H. Waterborg, in The Protein Protocols Handbook, ed. J. M. Walker, Humana Press, Totowa, NJ, 2009, pp. 7–10 Search PubMed.
- A. A. Ansari, S. A. Kidwai and A. Salahuddin, J. Biol. Chem., 1975, 250, 1625–1632 CrossRef CAS PubMed.
- Principles of Fluorescence Spectroscopy, ed. J. R. Lakowicz, Springer US, Boston, MA, 2006 Search PubMed.
- S. Sahu, T. Debnath and K. Sahu, J. Phys. Chem. Lett., 2024, 15, 3677–3682 CrossRef CAS PubMed.
- C. Louis-Jeune, M. A. Andrade-Navarro and C. Perez-Iratxeta, Proteins, 2012, 80, 374–381 CrossRef CAS PubMed.
- H. M. Berman, Nucleic Acids Res., 2000, 28, 235–242 CrossRef CAS PubMed.
- S. Pronk, S. Páll, R. Schulz, P. Larsson, P. Bjelkmar, R. Apostolov, M. R. Shirts, J. C. Smith, P. M. Kasson, D. Van Der Spoel, B. Hess and E. Lindahl, Bioinformatics, 2013, 29, 845–854 CrossRef CAS PubMed.
- B. Hess, C. Kutzner, D. Van Der Spoel and E. Lindahl, J. Chem. Theory Comput., 2008, 4, 435–447 CrossRef CAS PubMed.
- D. Van Der Spoel, E. Lindahl, B. Hess, G. Groenhof, A. E. Mark and H. J. C. Berendsen, J. Comput. Chem., 2005, 26, 1701–1718 CrossRef CAS PubMed.
- K. Vanommeslaeghe, E. Hatcher, C. Acharya, S. Kundu, S. Zhong, J. Shim, E. Darian, O. Guvench, P. Lopes, I. Vorobyov and A. D. Mackerell, J. Comput. Chem., 2010, 31, 671–690 CrossRef CAS PubMed.
- P. Mark and L. Nilsson, J. Phys. Chem. A, 2001, 105, 9954–9960 CrossRef CAS.
- T. Darden, D. York and L. Pedersen, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS.
- G. Bussi, D. Donadio and M. Parrinello, J. Chem. Phys., 2007, 126, 014101 CrossRef PubMed.
- M. Parrinello and A. Rahman, J. Appl. Phys., 1981, 52, 7182–7190 CrossRef CAS.
- S. Paul, S. R. K. Ainavarapu and R. Venkatramani, J. Phys. Chem. B, 2020, 124, 4247–4262 CrossRef CAS PubMed.
- W. Humphrey, A. Dalke and K. Schulten, J. Mol. Graph., 1996, 14(27–28), 33–38 CrossRef CAS PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, D. J. Fox, et al., Gaussian 09, Revision D.01, 2013 Search PubMed.
- M. Majzoobi, E. Abedi, A. Farahnaky and M. Aminlari, Food Chem., 2012, 133(4), 1402–1407 CrossRef CAS.
- H. R. Ball and S. E. Winn, Poult. Sci., 1982, 61, 1041–1046 CrossRef CAS.
- J. R. Lakowicz, B. P. Maliwal, H. Cherek and A. Balter, Biochemistry, 1983, 22, 1741–1752 CrossRef CAS PubMed.
- L. Stryer, J. Mol. Biol., 1965, 13, 482–495 CrossRef CAS PubMed.
- D. Matulis and R. Lovrien, Biophys. J., 1998, 74, 422–429 CrossRef CAS PubMed.
- M. Cardamone and N. K. Puri, Biochem. J., 1992, 282, 589–593 CrossRef CAS PubMed.
- S. Bhattacharjee, J. Controlled Release, 2016, 235, 337–351 CrossRef CAS PubMed.
- A. Priyadarshi, S. B. Saikia and R. Swaminathan, J. Phys. Chem. B, 2024, 128, 9656–9668 CrossRef CAS PubMed.
- G. R. Fleming and G. D. Scholes, Q. Rev. Biophys., 2024, 57, e11 CrossRef CAS PubMed.
- J. Pansieri, V. Josserand, S.-J. Lee, A. Rongier, D. Imbert, M. M. Sallanon, E. Kövari, T. G. Dane, C. Vendrely, O. Chaix-Pluchery, M. Guidetti, J. Vollaire, A. Fertin, Y. Usson, P. Rannou, J.-L. Coll, C. Marquette and V. Forge, Nat. Photonics, 2019, 13, 473–479 CrossRef CAS.
- A. P. Kalra, S. Biswas, I. Mulrain, M. Wang, J. A. Tuszynski and G. D. Scholes, J. Phys. Chem. Lett., 2023, 14, 5891–5900 CrossRef CAS PubMed.
- M. H. Yang, S. Nickerson, E. T. Kim, C. Liot, G. Laurent, R. Spang, M. R. Philips, Y. Shan, D. E. Shaw, D. Bar-Sagi, M. C. Haigis and K. M. Haigis, Proc. Natl. Acad. Sci. U. S. A, 2012, 109, 10843–10848 CrossRef CAS PubMed.
- T. Nguyen, D. Sridaran, S. Chouhan, C. Weimholt, A. Wilson, J. Luo, T. Li, J. Koomen, B. Fang, N. Putluri, A. Sreekumar, F. Y. Feng, K. Mahajan and N. P. Mahajan, Nat. Commun., 2023, 14, 3357 CrossRef CAS PubMed.
Footnote |
| † Equal contributions. |
| This journal is © The Royal Society of Chemistry 2026 |