Open Access Article
Open Access ArticleDock & design: engineering specificity for an alternative pimaradiene outcome with the ent-kaurene synthase from Bradyrhizobium japonicum
Mark Schmidt-Dannert a,
Yue Zhangb,
Jin Liang
a,
Yue Zhangb,
Jin Liang acd,
Qiang Wang
acd,
Qiang Wang c,
Samuel Tuftsa,
Meirong Jiaae,
Dean J. Tantillo
c,
Samuel Tuftsa,
Meirong Jiaae,
Dean J. Tantillo *b,
Justin B. Siegel*bfg and
Reuben J. Peters
*b,
Justin B. Siegel*bfg and
Reuben J. Peters *a
*a
aRoy J. Carver Department of Biochemistry, Biophysics & Molecular Biology, Iowa State University, Ames, Iowa 50011, USA. E-mail: rjpeters@iastate.edu
bDepartment of Chemistry, University of California-Davis, Davis, California 65616, USA. E-mail: djtantillo@ucdavis.edu; jbsiegel@health.ucdavis.edu
cCollege of Agriculture, Guizhou University, Guiyang, Guizhou 550025, China
dState Key Laboratory of Crop Gene Exploration and Utilization in Southwest China, College of Agronomy, Sichuan Agricultural University, Chengdu, Sichuan 611130, China
eState Key Laboratory of Bioactive Substance and Function of Natural Medicines, NHC Key Laboratory of Biosynthesis of Natural Products, Institute of Materia Medica, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing, 100050, China
fDepartment of Biochemistry & Molecular Medicine, University of California-Davis, Davis, California 65616, USA
gGenome Center, University of California-Davis, Davis, California 65616, USA
First published on 26th March 2026
Abstract
The complexity of the reactions catalyzed by terpene synthases has hindered enzymatic engineering. In most cases such efforts result in non-specific product outcome, with the targeted compound being produced alongside others, hindering further use. Previous work with the structurally characterized ent-kaurene synthase from Bradyrhizobium japonicum (BjKS) identified a serine for alanine substitution (A167S) that led to premature deprotonation, yielding a pair of ent-pimaradiene double-bond isomers, with retrospective analysis by the TerDockin computational approach indicating that the introduced hydroxyl acts as a catalytic base for both. Here this route to ‘short-circuiting’ the BjKS catalyzed reaction for ent-pimaradiene production was further explored, with prospective application of TerDockin, via design–build–test cycles, enabling specific production of a novel pimaradiene isomer via introduction of a water molecule as the catalytic base. The resulting mutants, BjKS:F72S and particularly BjKS:F72Y/Y280S specifically yield the targeted ent-pimara-8,15-diene with reasonable catalytic efficiency, demonstrating the applicability of this computationally inexpensive approach to engineering terpene synthase product outcomes.
Introduction
Terpene synthases are notorious for both the frequently intricate reactions they catalyze as well as the complex mixture of products that often result from them.1 While certain terpene synthases are quite specific, forming a single predominant product, many exhibit catalytic promiscuity, forming a variety of products from the same substrate.2 The complexity of the catalyzed reactions, combined with the reactivity of the underlying carbocation cascades, complicates enzymatic engineering of terpene synthases, with relatively few successful efforts reported.3The reactions catalyzed by class I terpene synthases are initiated by divalent magnesium (Mg2+)-assisted lysis of the allylic diphosphate ester in their isoprenyl substrates, with the ensuing reactions proceeding via carbocation intermediates and being terminated by deprotonation.1 Given the highly reactive nature of carbocations, these observations imply that terpene synthases exert catalytic specificity, at least in part, via steric constraints on the conformation of the isoprenyl portion of their substrate within a largely chemically inert portion of their active sites.4 Electrostatic effects from the pyrophosphate anion co-product and associated trio of Mg2+, as well as the charged residues that ligate these (from the characteristic DDxxD and NSE/DTE motifs), which make up the remainder of the active sites, may be offset by enzymatic stabilization of key carbocation intermediates.5–7 Regardless, the ensuing carbocation cascade proceeds towards intermediate(s) that can be deprotonated by appropriately situated general base(s). For specificity, this active site template must then exhibit conformational stability, at least during substrate binding/activation, as well as control access to any potential general bases during the course of the reaction.8 The importance of this latter point is emphasized by the fact that even protein backbone carbonyl groups have been proposed to serve this role.9,10
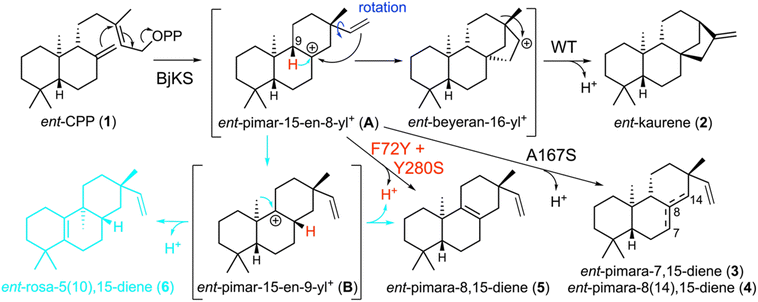
In the absence of these factors, multiple products may be formed, as highlighted by not only the known examples of native terpene synthases producing more than fifty different products,2 but also the general observation that mutational alterations typically lead to increased numbers of distinct products, hampering the design and use of these enzymes.3 The relevant example here derives from the structurally characterized ent-kaurene synthase from Bradyrhizobium japonicum (BjKS)11 which reacts with ent-copalyl diphosphate (ent-CPP, 1) to yield almost entirely ent-kaurene (2),12 in a reaction involving cyclization followed by vinyl rotation to enable concerted (secondary) cyclization and ring rearrangement (Scheme 1).13 It has been reported that substitution of serine for a particular alanine (A167S) can short-circuit this reaction after initial cyclization, largely yielding a mixture of two pimaradiene double-bond isomers (ent-pimara-7,15-diene, 3, and ent-pimara-8(14),15-diene, 4) along with small amounts of 2.14 To further explore the mechanism by which this mutant elicits this large change in product outcome, a novel computationally-inexpensive terpene-carbocation docking approach, TerDockin,15,16 was applied in a retrospective manner. Notably, TerDockin exhibited remarkable predictive efficiency for the observed product outcome when the alcohol of the serine was constrained to act as the relevant catalytic base.14 This highlights both the efficacy of this approach in predicting the outcome for direct deprotonation of intermediates as well as the ability of hydroxyl groups to act as a general base.
Further exploration of BjKS has highlighted a tyrosine for phenylalanine substitution (F72Y) leading to a mixture of products that includes ent-pimara-8,15-diene (5), for which no specific terpene synthase has yet been found.17 This presented a unique opportunity to explore the prospective capability of the TerDockin approach to design a specific product outcome. Thus, TerDockin was applied here in a novel cyclical design–build–test fashion to identify key substitutions that enabled engineering specificity for production of 5.
Methods
General
All reagents were purchased from Fisher Scientific unless otherwise specified.General TerDockin preparation
The TerDockin approach docks both the relevant carbocation moiety and the pyrophosphate–magnesium cluster (PP–3Mg2+) simultaneously, in both cases having their structures fixed during docking,16 much as previously described.14 In the case of the magnesium–pyrophosphate cluster, the structure was extracted from the closest homologue of BjKS which had the substrate bound. The carbocation used in these docking explorations was the ent-pimara-15-en-8-yl+ (A) generated by initial cyclization, a structure for which had previously been optimized and a conformer library generated. These were docked into the active site of a crystal structure of BjKS previously determined to be the best resolved (4XLX). Also docked was the ent-pimara-15-en-9-yl+ (B) generated from A by a 1,2-hydride shift from C9 to C8, which was optimized and a conformer library generated as previously described. Briefly, ent-pimara-15-en-9-yl+ was optimized using Gaussian 16 with the B3LYP functional and a 6-31+G(d,p) basis set.18 A conformer library was generated from the optimized structure using CREST19 and optimized as above. After filtering out identical structures and those with energies greater than 5 kcal mol−1 above the lowest energy structure, the resulting structures were converted into a conformer library for use in Rosetta. Also included in the docking wasa water molecule that was shown to be of high occupancy amongst all BjKS crystal structures. The constraints used in docking were those previously described for pyrophosphate–magnesium cluster coordination with constraints from both the DDxxD and NSE motifs to the magnesium atoms, as well as from either of the terminal pyrophosphate oxygens to the originally linked carbon (C16), representing the two potential orientations of this moiety. These constraints were accompanied by others representing the requisite angles and distances between atoms for the modelled chemistry as described below. After docking, the resultant structures were filtered for constraint fulfilment (constraint score <1), interface energy (lowest 20%), and total energy (lowest 20%) to select the energetically most favorable structural poses.Enzymatic assays
Recombinant constructs were generated for the selected mutants using a previously created pET100 based vector containing the gene coding for BjKS, codon optimized for expression in Escherichia coli.14 Targeted mutants were generated using overlapping primer mutagenesis with primers ordered from IDT and the Q5 polymerase system (NEB). Enzymatic activity was assayed by expression of these constructs alongside a previously described modular metabolic engineering system.20 Briefly, the expression vector was co-transformed into Bl21*(DE3) (Thermo Scientific) with a plasmid encoding a GGPP synthase and an ent-CPP synthase. Transformed E. coli were grown in 250 mL Erlenmeyer flasks with 45 mL TB and 5 mL 1 M phosphate buffer at pH 7.0. Cultures were initially grown to OD600 ∼0.6 at 37 °C and 180 rpm, following which the temperature was dropped to 16 °C for 1 hour prior to induction with 1 mM IPTG. 72 hours after induction, cultures were extracted with 50 mL hexanes, which were dried under a stream of N2 gas and resuspended in 100 µL hexanes for analysis by gas chromatography with mass spectral detection (GC-MS).GC-MS analysis
GC-MS analysis was carried out using the following Agilent equipment: an 8890 GC system with a 5977B mass spectrometer operating in 70 eV electron ionization mode using an HP-5MS column at a helium flow rate of 1.1 mL min−1. Injection was performed in splitless mode with an injector temperature of 250 °C and a 7650A autosampler. The oven temperature was kept at 50 °C for 3 minutes before being increased at 15 °C min−1 to 300 °C. This temperature was maintained for a further 3 minutes. Mass-to-charge (m/z) ratios between 90 and 300 were recorded after 13 minutes through the end of the run.Results
TerDockin analysis of BjKS:F72Y
Given the production of 5 by BjKS:F72Y,17 TerDockin was applied retrospectively to assess its capacity for accurately predicting this activity. Given that this variant also produces ent-rosadiene, requiring a hydride shift from C9 to C8 and transition through ent-pimara-15-en-9-yl+ (B), it was necessary to first identify the relevant intermediate – i.e., since 5 can be produced by deprotonation of either A or B (Scheme 1). To determine the best intermediate to use in the subsequent docking studies, the F72Y substitution was introduced in silico and docked with either A or B. The hydroxyl introduced by the tyrosine was constrained for deprotonation of A at C9 or for B at C8 instead. Deprotonation constraints consisted, as with the A167S mutant,14 of 120 ± 12° for the CTyr–OTyr–H angle and 180 ± 18° for the OTyr–H–C angle (Fig. S1). Given that either end of the pyrophosphate co-product could have been involved in the ester linkage of the substrate (1),21 for each intermediate both orientations were tested, yielding 4 constraint sets for each of which 2500 docked structures were generated. The resulting 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 structures were pooled and subsequently filtered as described in the Methods section for satisfaction of constraints, interface energy, and total energy to select the most favorable structural poses. The majority of the passing poses (>65%) were from the docking runs modeling deprotonation of A at C9 (Table S1 and Fig. S2). Hence, docking was continued with this intermediate. This further enabled analysis of the selectivity for 5 relative to the other accessible pimaradiene olefin isomers – i.e., deprotonation of A at C7 or C14 to produce 3 or 4, respectively. Accordingly, in addition to the previously used constraints for F72Y deprotonation of A at C9, identical constraint sets were generated for each of the four protons on C7 or C14, in combination with both possible pyrophosphate orientations. For each of the resulting 10 constraint sets, 5000 docked structures were generated and pooled (50
000 structures were pooled and subsequently filtered as described in the Methods section for satisfaction of constraints, interface energy, and total energy to select the most favorable structural poses. The majority of the passing poses (>65%) were from the docking runs modeling deprotonation of A at C9 (Table S1 and Fig. S2). Hence, docking was continued with this intermediate. This further enabled analysis of the selectivity for 5 relative to the other accessible pimaradiene olefin isomers – i.e., deprotonation of A at C7 or C14 to produce 3 or 4, respectively. Accordingly, in addition to the previously used constraints for F72Y deprotonation of A at C9, identical constraint sets were generated for each of the four protons on C7 or C14, in combination with both possible pyrophosphate orientations. For each of the resulting 10 constraint sets, 5000 docked structures were generated and pooled (50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 total) before filtering as above (Table S2). When grouped by the expected product of the constrained deprotonation, F72Y is predicted to almost exclusively produce 5 (>95%). This closely matches the observed production of 5 (and not 3 or 4), consistent with deprotonation of A to yield 5.
000 total) before filtering as above (Table S2). When grouped by the expected product of the constrained deprotonation, F72Y is predicted to almost exclusively produce 5 (>95%). This closely matches the observed production of 5 (and not 3 or 4), consistent with deprotonation of A to yield 5.
Prospective application of TerDockin for pimaradiene production
Given the ability of introduced hydroxyl groups to serve as a catalytic base to short-circuit the catalyzed reaction and produce pimaradienes and the remarkable efficacy of TerDockin at predicting the product outcomes of such variants, it was hypothesized that this approach could be applied prospectively. Specifically, to explore the effect introduction of hydroxyl groups at other sites around the carbocation in A might have in directing activity towards production of pimaradienes (i.e., 3–5), an in silico serine/threonine scan of the active site residues within 8 Å was conducted (Table S3). Here, the relevant angle constraints (as above) were introduced between the hydroxyl introduced by the serine or threonine substitution and each of the 5 protons around the carbocation in A (C8). Each of these 5 constraint sets was tested for both possible pyrophosphate orientations. For each constraint set (10 total), 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 structures were generated, with the resulting 250
000 structures were generated, with the resulting 250![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 docked structures pooled and then filtered to select the energetically most favorable structural poses (Fig. S3).
000 docked structures pooled and then filtered to select the energetically most favorable structural poses (Fig. S3).
The passing structures were then separated by the product of the modeled deprotonation (e.g., constraint to a proton on C7 yields 3). From this group, a set of three mutants was selected to examine the predictive capacity of TerDockin design. The BjKS L71S and I166T mutants were picked as they showed both a high number of passing poses as well as preference for a specific double bond isomer (4 and 3, respectively), while V277S was included as a control given its relatively low number of passing poses and lack of isomer preference from the docking results (Table S4). These substitutions were made and the resulting variants were tested in a metabolic engineering system – i.e., co-expressed in E. coli with enzymes producing the substrate 1.20 Although the V277S mutant did not alter product outcome and the others did lead to some production of pimaradienes, this did not generally match the predicted site of deprotonation. Instead, BjKS:L71S produced a roughly equal mixture of 3 and 4, rather than the predicted selective production of 4, and BjKS:I166T selectively produced 4 rather than the predicted 3 (Fig. S4). Visual inspection of the passing docked structures revealed improbably long distances between the hydroxyl group and the proton to be abstracted. This indicates that, in order for the passing poses to correctly represent the intended deprotonation, a distance constraint must be included alongside the angle constraints (Fig. S5).
A second docking run was performed as above with an additional constraint introduced for a distance range of 2.5 ± 0.5 Å between the relevant hydroxyl group and the proton to be abstracted. This new constraint set was applied to the previous serine/threonine substitution set, along with F72Y, and the resulting docked structures were pooled and then filtered to select the energetically most favorable structural poses as before. Given the lack of any diterpene synthase that selectively yields 5, production of this isomer was targeted, with mutants selected based on their relative proportion of passing structures for abstraction of the C9 proton from A. This docking run predicted two sites beyond F72Y to be of potential interest: Y280 and G32. When assayed in the metabolic engineering system, G32S/T and Y280S/T showed a strong switching effect, yielding 5 as their primary product (Fig. S6). This is, however, accompanied by significant losses in activity, as indicated by the accumulation of ent-copalol (1′), resulting from dephosphorylation of the substrate (1) by the endogenous (E. coli) phosphatases competing with BjKS in this system.
Designing for production of 5 with F72Y, I166F, G32X and Y280X
Given the effectiveness of F72Y, as well as serine/threonine substitutions for G32 and Y280, in switching activity to production of 5, these residues were selected for further design attempts. As the F72Y variant produces 5 without significant loss of catalytic efficiency (i.e., accumulation of 1′), the hydroxyl group on the tyrosine was constrained as the catalytic base (as described above). However, although this mutant produces around 25% 5 it also yields 50% 2. Thus, to inhibit the requisite vinyl rotation in A, an I166F mutation was included under the assumption that this would sterically block such movement. Indeed, BjKS:I166F produces a mixture of pimaradiene olefin isomers along with ∼50% 2 (Fig. S7). To this pair of substitutions, additional substitutions for positions 32 and 280 were explored. Y280 was substituted with every possible residue; however, given the proximity of G32 to the catalytic base (i.e., Y72), it was hypothesized to assist deprotonation. Therefore, its substitutions were limited to aspartate, glutamate, histidine, asparagine, glutamine, serine, threonine or tyrosine. For each possible combination of these amino acids at positions 32 and 280 (160 total, each with 2 pyrophosphate orientations), 150 docked structures were generated with constraints expected to lead to the production of 5. The resulting 48![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 docked structures were pooled and then filtered to select the energetically most favorable structural poses as before, following which the passing structures were grouped by residue pairing (i.e., identity). From the best represented mutants in this set (i.e., those with the highest proportion of passing poses), eight were selected for testing based on maximization of amino acid diversity while retaining the desired chemical reactivity – i.e., ability to act as a base to deprotonate A at C9 (Table S5). Unfortunately, while more specific for production of 5, the resulting set of variants, including the F72Y/I166F double mutant and various G32x/F72Y/I166F triple mutants made en route, showed a near complete loss of activity in the metabolic engineering system (Fig. S8). This is perhaps not unexpected given increased volume of the I166F and G32X substitutions, which reduce the size of the active site that is expected to already be a tight fit given the specificity of BjKS for production of 2. Moreover, the continued production of 2 by BjKS:I166F indicates that this substitution does not fully block rotation of the vinyl substituent in A in any case.
000 docked structures were pooled and then filtered to select the energetically most favorable structural poses as before, following which the passing structures were grouped by residue pairing (i.e., identity). From the best represented mutants in this set (i.e., those with the highest proportion of passing poses), eight were selected for testing based on maximization of amino acid diversity while retaining the desired chemical reactivity – i.e., ability to act as a base to deprotonate A at C9 (Table S5). Unfortunately, while more specific for production of 5, the resulting set of variants, including the F72Y/I166F double mutant and various G32x/F72Y/I166F triple mutants made en route, showed a near complete loss of activity in the metabolic engineering system (Fig. S8). This is perhaps not unexpected given increased volume of the I166F and G32X substitutions, which reduce the size of the active site that is expected to already be a tight fit given the specificity of BjKS for production of 2. Moreover, the continued production of 2 by BjKS:I166F indicates that this substitution does not fully block rotation of the vinyl substituent in A in any case.
Designing activity with F72X and Y280X
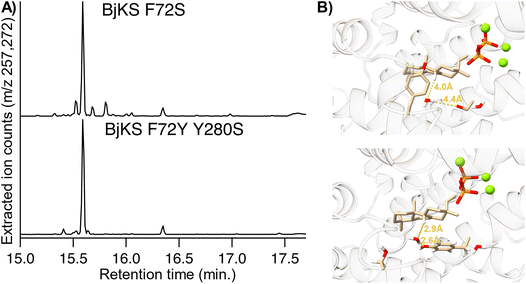
Given the loss of activity observed with the introduction of the I166F and G32X mutations, these were removed from the mutation set. Instead, substitution of the F72 and Y280 positions with any pair of residues where at least one maintained a hydrophilic nature was explored. Based on the ability of serine/threonine substitution for Y280 to lead to production of 5, both of which would expand the active site volume, it was hypothesized that this might stem from incorporation of a water molecule that acts as the catalytic base. Thus, in this cycle such a water molecule was introduced for this purpose. With this additional constraint, docking runs were performed as before, but with generation of 250 structures for each of these combinations of residues at positions 72 and 280 (256 total, each with 2 possible pyrophosphate orientations). The resulting 128![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 docked structures were pooled and filtered to select the energetically most favorable structural poses as before, although an additional filter for the individual energies of each of the two residues was included as well (Fig. S9). The passing structures were again grouped by mutations and, from the best performing, ten were selected for testing based on maximization of diversity of the introduced amino acids (Table S6). Strikingly, every variant produced primarily 5 in the metabolic engineering system, although some also produced small amounts of 2 and/or exhibited reduced activity, as indicated by the accumulation of 1′ (Fig. S10). Particularly interesting were the F72S and, especially, F72/Y280S variants, which exhibited strong selectivity for 5 and reasonable catalytic efficiency (Fig. 1A). Notably, while in BjKS:F72Y/Y280S the hydroxyl introduced by the F72Y substitution is within the hydrogen bonding distance of the water, which is optimally positioned to serve as the catalytic base, this is not the case in BjKS:F72S (Fig. 1B). This may reflect the fact that substitution of serine for phenylalanine or tyrosine presumably leaves sufficient room for multiple water molecules to bind, which might further enable the observed catalysis. Regardless, this outcome demonstrates the specificity achievable through the TerDockin based design–build–test cycle pioneered here.
000 docked structures were pooled and filtered to select the energetically most favorable structural poses as before, although an additional filter for the individual energies of each of the two residues was included as well (Fig. S9). The passing structures were again grouped by mutations and, from the best performing, ten were selected for testing based on maximization of diversity of the introduced amino acids (Table S6). Strikingly, every variant produced primarily 5 in the metabolic engineering system, although some also produced small amounts of 2 and/or exhibited reduced activity, as indicated by the accumulation of 1′ (Fig. S10). Particularly interesting were the F72S and, especially, F72/Y280S variants, which exhibited strong selectivity for 5 and reasonable catalytic efficiency (Fig. 1A). Notably, while in BjKS:F72Y/Y280S the hydroxyl introduced by the F72Y substitution is within the hydrogen bonding distance of the water, which is optimally positioned to serve as the catalytic base, this is not the case in BjKS:F72S (Fig. 1B). This may reflect the fact that substitution of serine for phenylalanine or tyrosine presumably leaves sufficient room for multiple water molecules to bind, which might further enable the observed catalysis. Regardless, this outcome demonstrates the specificity achievable through the TerDockin based design–build–test cycle pioneered here.
These results further enhance our understanding of the structure–function relationships in BjKS. Previous work with the A167S mutant suggested that it might be capable of producing all three pimaradiene products derived from deprotonation of A at any of the adjacent carbons (i.e., C7, C9 or C14). However, while 5 was predicted as a minor product it was not observed, leaving a discrepancy in the experimental versus TerDockin results. This might reflect the lack of a pre-catalytic substrate complex structure for BjKS, as only an inactive mutant has been co-crystallized with 1,11 reducing the accuracy of the docking results. Nonetheless, combining these earlier observations for A167 with the effects of altering F72 and Y280 reported here, it becomes clear that this outcome is most likely due to restricted substrate orientation, consistent with the observed catalytic specificity of BjKS. In particular, although protons can be abstracted from either face of A at C7 and C14, for C9 only one proton is available, hence restricting deprotonation to occur at this face. Given that F72 and Y280 are on the opposite side of the active site relative to A167 and I166, along with the correlation between substitutions of these residues and their effects on product outcome, this indicates restricted substrate/reactant orientation within the active site that necessitates more selective positioning of the catalytic base for deprotonation of A at C14. This further establishes constraints that will enable more accurate docking predictions for future re-design of BjKS activity. Indeed, as analogs of the A167S switch have been identified in many other diterpene synthases,22–25 this constraint may prove to be even more broadly applicable.
Conclusions
The work presented here significantly expands application of the computationally inexpensive TerDockin system beyond the previously reported retrospective analyses and establishes its effectiveness as a predictive tool for design towards a specific terpene synthase product outcome. While more intensive multiscale computational approaches for engineering such catalysis have proven to be fruitful in certain cases,26 their cost and complexity have limited their accurate application. Here, it has been shown that the inexpensive nature of TerDockin enables a cyclical design–build–test approach, which can be leveraged to generate targeted specific products resulting from premature deprotonation of the initially cyclized carbocation intermediate A (Fig. 2). While the computational methods that make up TerDockin have limitations, as shown in this study, these are expected; a cornerstone of the TerDockin ‘philosophy’ is the integration of chemical intuition alongside simulation, the combination of which, as shown here, can be used to great effect. Notably, a specific ent-pimara-8,15-diene (5) synthase does not appear to have been previously reported, and the results presented here further represent engineering of a novel enzyme. Accordingly, this approach opens new possibilities for redesigning terpene synthases to achieve a specific product outcome. Indeed, clarification of the restricted substrate/reactant orientation in the active site will allow more accurate application of TerDockin in future rational design projects.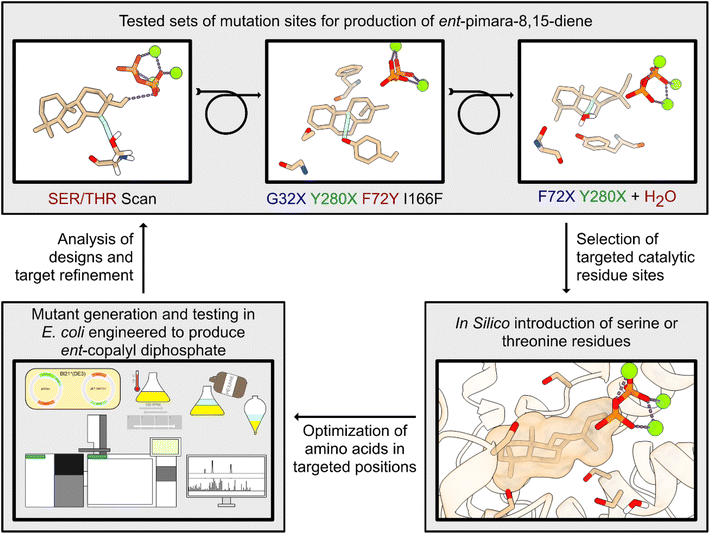 | ||
| Fig. 2 Use of TerDockin in the cyclical design–build–test approach to engineer specific BjKS variants. | ||
Author contributions
M. S. D. (investigation, formal analysis, writing – original draft), Y. Z. (investigation, formal analysis), J. L. (investigation, formal analysis), Q. W. (supervision), S. T. (investigation), M. J. (investigation, formal analysis), D. J. T. (supervision, methodology, writing – review & editing, funding acquisition), J. B. S. (supervision, methodology, writing – review & editing), R. J. P. (conceptualization, supervision, formal analysis, funding acquisition, writing – review & editing).Conflicts of interest
There are no conflicts of interest to declare.Data availability
The Rosetta score data from the TerDockin analyses as well as the PDB files of the docked structures that cleared filtering in each of the described runs have been deposited and are publicly accessible at https://doi.org/10.25380/iastate.31366222.Supplementary information (SI): figures and tables. See DOI: https://doi.org/10.1039/d5sc09199c.
Acknowledgements
This work was supported by grants from the NIH to R. J. P. and D. J. T. (currently GM156300 and GM153469), which were only used to support studies carried out in the United States.References
- D. W. Christianson, Chem. Rev., 2017, 117, 11570–11648 CrossRef CAS PubMed.
- A. Vattekkatte, S. Garms, W. Brandt and W. Boland, Org. Biomol. Chem., 2017, 16, 348–362 RSC.
- N. G. H. Leferink and N. S. Scrutton, ChemBioChem, 2022, 23, e202100484 CrossRef CAS PubMed.
- D. J. Tantillo, Angew. Chem., Int. Ed., 2017, 56, 10040–10045 CrossRef CAS PubMed.
- K. Zhou and R. J. Peters, Chem. Commun., 2011, 47, 4074–4080 RSC.
- D. T. Major and M. Weitman, J. Am. Chem. Soc., 2012, 134, 19454–19462 CrossRef CAS PubMed.
- D. T. Major, ACS Catal., 2017, 7, 5461–5465 CrossRef CAS.
- T. A. Pemberton and D. W. Christianson, J. Antibiot., 2016, 69, 486–493 CrossRef CAS PubMed.
- C. Lemke, K. Roach, T. Ortega, D. J. Tantillo, J. B. Siegel and R. J. Peters, ACS Bio Med Chem Au, 2022, 2, 490–498 CrossRef CAS PubMed.
- Y.-H. Wang, H. Xu, J. Zou, X.-B. Chen, Y.-Q. Zhuang, W.-L. Liu, E. Celik, G.-D. Chen, D. Hu, H. Gao, R. Wu, P.-H. Sun and J. S. Dickschat, Nat. Catal., 2022, 5, 128–135 CrossRef CAS.
- W. Liu, X. Feng, Y. Zheng, C.-H. Huang, C. Nakano, T. Hoshino, S. Bogue, T.-P. Ko, C.-C. Chen, Y. Cui, J. Li, I. Wang, S.-T. D. Hsu, E. Oldfield and R.-T. Guo, Sci. Rep., 2014, 4, 6214 CrossRef CAS PubMed.
- D. Morrone, J. Chambers, L. Lowry, G. Kim, A. Anterola, K. Bender and R. J. Peters, FEBS Lett., 2009, 583, 475–480 CrossRef CAS PubMed.
- Y. J. Hong and D. J. Tantillo, J. Am. Chem. Soc., 2010, 132, 5375–5386 CrossRef CAS PubMed.
- M. Jia, Y. Zhang, J. B. Siegel, D. J. Tantillo and R. J. Peters, ACS Catal., 2019, 9, 8867–8871 CrossRef CAS PubMed.
- T. E. O'Brien, S. J. Bertolani, Y. Zhang, J. B. Siegel and D. J. Tantillo, ACS Catal., 2018, 8, 3322–3330 CrossRef PubMed.
- I. S. Torrence, T. E. O'Brien, J. B. Siegel and D. J. Tantillo, Methods Enzymol., 2024, 699, 231–263 CAS.
- F. Zhang, S.-J. Wang, W. Xiao, M.-Z. Yu, F. Sha, R. Wu and Z. Xiang, Catal. Sci. Technol., 2023, 14, 306–314 RSC.
- M. D. Bojin and D. J. Tantillo, J. Phys. Chem. A, 2006, 110, 4810–4816 CrossRef CAS PubMed.
- P. Pracht, S. Grimme, C. Bannwarth, F. Bohle, S. Ehlert, G. Feldmann, J. Gorges, M. Müller, T. Neudecker, C. Plett, S. Spicher, P. Steinbach, P. A. Wesołowski and F. Zeller, J. Chem. Phys., 2024, 160, 114110 CrossRef CAS PubMed.
- A. Cyr, P. R. Wilderman, M. Determan and R. J. Peters, J. Am. Chem. Soc., 2007, 129, 6684–6685 CrossRef CAS PubMed.
- R. Schwartz, S. Zev and D. T. Major, Angew. Chem., Int. Ed., 2024, 63, e202400743 CrossRef CAS PubMed.
- P. R. Wilderman and R. J. Peters, J. Am. Chem. Soc., 2007, 129, 15736–15737 CrossRef CAS PubMed.
- D. Morrone, M. Xu, D. B. Fulton, M. K. Determan and R. J. Peters, J. Am. Chem. Soc., 2008, 130, 5400–5401 CrossRef CAS PubMed.
- C. I. Keeling, S. Weisshaar, R. P. C. Lin and J. Bohlmann, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 1085–1090 CrossRef CAS PubMed.
- M. Xu, P. R. Wilderman and R. J. Peters, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 7397–7401 CrossRef CAS PubMed.
- K. Raz, S. Levi, P. K. Gupta and D. T. Major, Curr. Opin. Biotechnol., 2020, 65, 248–258 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |