Open Access Article
Open Access ArticleFunctionalized two-dimensional nanochannel membranes to distinguish methylated/unmethylated peptides for sensing cellular G9a protein
Jing-Jing
Hu†
a,
Niya
Lin†
a,
Wenlian
Jiang
a,
Rui
Liu
a,
Xiaoding
Lou
 *ab and
Fan
Xia
*ab and
Fan
Xia
 *ab
*ab
aState Key Laboratory of Biogeology and Environmental Geology, Faculty of Materials Science and Chemistry, China University of Geosciences, Wuhan 430074, China. E-mail: louxiaoding@cug.edu.cn; xiafan@cug.edu.cn
bShenzhen Research Institute of China University of Geosciences, Shenzhen 518057, China
First published on 21st January 2026
Abstract
Histone methyltransferase G9a-catalyzed histone modifications serve as critical epigenetic marks that regulate diverse cellular functions. Aberrant enzymatic activities of G9a are closely associated with various human pathologies, making them promising biomarkers for disease diagnosis and therapeutic targeting. However, the lack of efficient assays for accurate and sensitive detection of G9a, especially in complex crude cellular extracts, limits elucidation of cancer-related mechanisms and advancements of therapeutic innovations. Here, a G9a detection system based on functionalized 2D solid-state nanochannel membranes through the specific recognition between small molecule Crown probes and dissociative peptide probes was developed. The Crown probe exhibited a strong binding force with the unmethylated peptide through hydrogen bonding, thereby altering the ion transport characteristics within the nanochannels and enabling the differentiation of methylated and unmethylated peptides followed by the detection of histone methyltransferase G9a. The excellent sensing performance enables diverse detection scenarios, including differentiation of different subtypes of breast cancer cells, screening of enzyme inhibitors and sensing of hypoxic cellular environments. This research proposed that the GO-Crown membrane could be an effective detection system for G9a, which provides a powerful tool for cancer research and clinical application.
Introduction
As one of the most important post-translational modifications, protein methylation has attracted much attention because of its important role in many biological processes such as DNA repair, RNA processing, transcriptional regulation, and protein phase separation.1–3 Methylation mostly occurs on transcription factors and histones.4,5 With the assistance of histone methyltransferases such as G9a (also known as EHMT2), histone methylation primarily takes place on the N-terminal tail regions of histone H3 and H4.6,7 Utilizing S-adenosyl-L-methionine (AdoMet) as a methyl donor, G9a catalyzes the addition of one or multiple methyl groups to lysine residues (e.g., H3K9), forming mono-, di-, or trimethylation modifications, which are closely related to the formation and maintenance of chromatin structure, silencing or activation of gene expression, etc.8–11 The aberrant expression of G9a is closely related to the occurrence and development of a variety of diseases. In breast cancer cells, aberrantly expressed G9a leads to gene silencing through hypermethylation of the tumor suppressor gene promoter region H3K9, thereby promoting tumorigenesis and progression.12,13 In addition, G9a is also highly expressed in many other cancers compared to expression in normal tissues, including human bladder cancer, lung cancer, colon cancer, etc.14–16 From a diagnostic point of view, changes in the content of G9a can serve as a potential biomarker for the diagnosis of cancer described above. In terms of treatment, G9a can also be an important therapeutic target. For cancer cells with high expression of G9a, the development of specific inhibitors can effectively block their abnormal methylation regulation, thereby inhibiting the growth and spread of cancer cells, which can provide new ideas and methods for precision treatment of cancer.17,18 Therefore, the development of G9a detection tools is crucial for elucidating cancer-related mechanisms and advancing therapeutic innovations.Current methods for detecting histone methyltransferases predominantly rely on traditional spectrophotometry,19 mass spectrometry methods,20etc., most of which involve radiolabeling, indirect detection of by-product (S)-adenosyl-L-homocysteine (AdoHcy),21 the use of specific antibodies,19 and, more recently, supramolecular replacement methods.22 While some existing approaches exhibit high sensitivity, they suffer from limitations including multistep immobilization or separation procedures, necessary post-reaction treatments, safety concerns about the use of radiotracers, interference from background fluorescence, and high costs.23,24 What's more, given G9a's potential as a tumor biomarker, few methods are capable of directly analyzing G9a in complex crude cellular extracts without prior purification or separation. There remains an urgent need for a label-free, highly selective, sensitive, and simple tool for G9a detection in native biological matrices.
Inspired by biological protein channels, the construction of artificial solid-state nanopores capable of smart responses to various targets has attracted widespread interest. Compared to biological nanopores, artificial solid-state nanopores offer distinct advantages, such as mechanical stability, controllable pore size and geometry, and ease of functional modification.25–27 By immobilizing recognition probes on nanopore surfaces, target-induced changes in transmembrane ionic currents enable the detection of analytes. Leveraging the inherent size-exclusion effect of nanopores, such biomimetic systems can filter out large-sized biological interferences such as cellular debris and organelles, simplifying sample pretreatment and realizing real-time and convenient analysis of complex samples.28–31 In previous work, many groups have achieved sensitive detection of various proteins using solid-state nanochannels.32–34 Compared with the horizontal transport path, two-dimensional (2D) nanochannel membranes can carry mass transport in the vertical direction due to the different structures of 2D layered materials, which holds promise for high-throughput membrane-based applications.35–40 In addition, the unique layered structure of 2D nanomaterials provides abundant binding sites for probes, enhancing analyte capture efficiency, and its unique mass transfer path can prolong analyte residence time within the nanochannels, thereby improving sensing performance.41,42 Among them, the graphene oxide (GO)-based 2D nanochannel stands out for biosensing due to its high sensitivity, excellent biocompatibility, facile functionalization, etc. Its surface is rich in oxygen-containing groups (e.g., hydroxyl and carboxyl), enabling covalent or non-covalent conjugation with biomolecules or functional moieties for target-specific recognition.43,44 Based on this, we propose that 2D nanochannel-based platforms may offer a groundbreaking solution to the challenges of G9a detection in complex cellular environments.
In this work, we developed a G9a detection system based on functionalized 2D solid-state nanochannels through the specific recognition between small molecule probes that anchored on the surface and dissociative peptide probes (Fig. 1). The small molecule probe 4′-aminobenzo-18-Crown-6 (briefly referred to as the Crown probe) was modified on the GO sheet, and the GO-Crown membrane was then obtained by vacuum filtration technology. The Crown probe exhibited a strong binding force with the unmethylated peptide through hydrogen bonding, thereby altering the ion transport characteristics within the nanochannels and enabling the differentiation of methylated and unmethylated peptides. Furthermore, the GO-Crown system was applied to the detection of histone methyltransferase G9a. The sensing performance in application scenarios involving different subtypes of breast cancer cells, the use of inhibitors, and simulated hypoxic environments was discussed as well. This research proposed that the GO-Crown membrane could be an effective detection system for G9a, which provides a powerful tool for cancer research and clinical application.
Results and discussion
Construction of the GO-Crown membrane
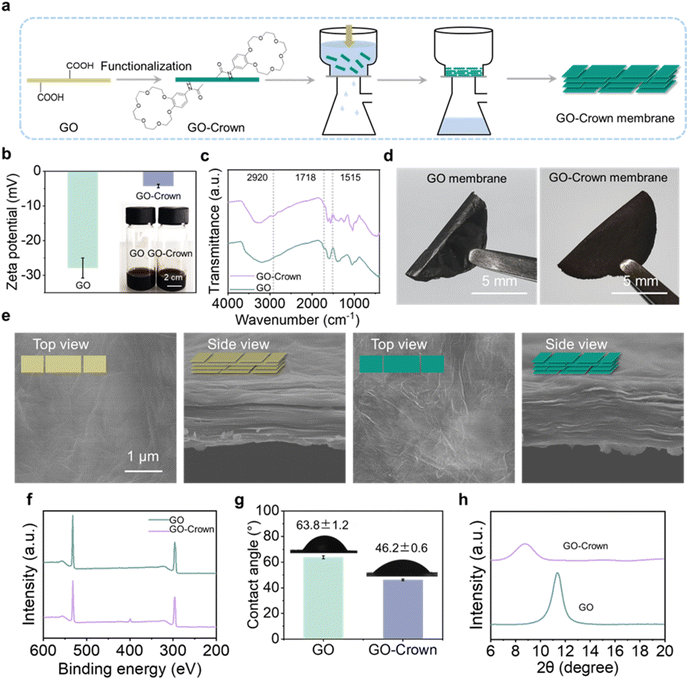
To construct the functionalized 2D nanochannel membrane, namely the GO-Crown membrane, GO was first dispersed in ultra-pure water and modified with 4′-aminobenzo-18-crown-6 (abbreviated to Crown), serving as the capture probe through an amidation reaction between the carboxyl groups on GO and the amine groups on Crown (Fig. 2a). As shown in Fig. 2b, because of the abundant negatively charged groups on the surface of GO, the zeta potential of GO dispersion liquid was −27.87 mV. After the Crown probe was successfully modified, the amounts of carboxyl groups on the GO surface decreased and the zeta potential of GO-Crown increased to −4.25 mV. The particle size increased from 1400 nm to 2200 nm (Fig. S1). Fourier transform infrared spectroscopy (FTIR) was also used to monitor the functionalization process (Fig. 2c). Compared to the absorption of GO, the absorption of functionalized GO-Crown at 1718 cm−1 from carboxyl groups declined, while the absorption at 2920 cm−1, ascribed to the asymmetric stretching of CH2 from Crown, increased. In addition, the absorption at 1515 cm−1 from stretching vibration of the CO–NH bond increased, demonstrating the formation of the amide bonds and the successful covalent conjugation of the Crown probe.45,46Subsequently, the GO-Crown membrane was obtained by pumping the functional dispersion liquid into a film by vacuum filtration technology and peeling off after drying (Fig. 2d). A scanning electron microscope (SEM) was utilized to observe the microstructure of GO and GO-Crown membranes. As shown in Fig. 2e, both membranes exhibited distinct lamellar structures; however, after modification with the probes, the GO-Crown membrane was more incompact and its surface was rougher. Consistent with the SEM images, atomic force microscopy (AFM) also showed that the roughness of the GO-Crown membrane was greater than the GO membrane (Fig. S2). X-ray photoelectron spectroscopy (XPS) illustrated that, except for the two peaks of C 1s and O 1s at 287 eV and 533 eV, respectively, in the GO membrane, a new peak corresponding to N 1s at 399.5 eV emerged for the GO-Crown membrane (Fig. 2f, Tables S1 and S2).47 Energy dispersive spectroscopy (EDS) also showed that the nitrogen content of the functionalized GO-Crown membrane was higher than that of the GO membrane, verifying the successful functionalization of the Crown probe again (Fig. S3). Moreover, the GO membrane and functionalized GO-Crown membrane were characterized by surface contact angle (CA) tests. As shown in Fig. 2g, after modification with the hydrophilic Crown probes, the CA decreased from 63.8° to 46.2°. From Raman spectra, the D peak was correlated with the vibration of carbon atoms of the sp3 bond in GO, and the G peak was correlated with the vibration of carbon atoms of the sp2 bond in GO. IG/ID (strength ratio) of GO was 1.01 and that of GO-Crown was 1.2, suggesting a reduction in structural defects48 (Fig. S4). Then the layer distance of the 2D nanochannel was analyzed by X-ray diffraction (XRD) (Fig. 2h). The GO membrane and functionalized GO-Crown membrane exhibited diffraction peaks at 11.3° and 8.8°, respectively. According to the Bragg formula d = λ/2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) sin
sin![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) θ, it could be calculated that the interlayer distance of the GO membrane was 0.78 nm and that of the GO-Crown membrane was 1.01 nm, indicating that functionalization increased the layer spacing of the 2D nanochannel. Together, the above results proved that the GO-Crown membrane with 2D nanochannels was successfully constructed.
θ, it could be calculated that the interlayer distance of the GO membrane was 0.78 nm and that of the GO-Crown membrane was 1.01 nm, indicating that functionalization increased the layer spacing of the 2D nanochannel. Together, the above results proved that the GO-Crown membrane with 2D nanochannels was successfully constructed.
Evaluating the peptide responsiveness of 2D nanochannels
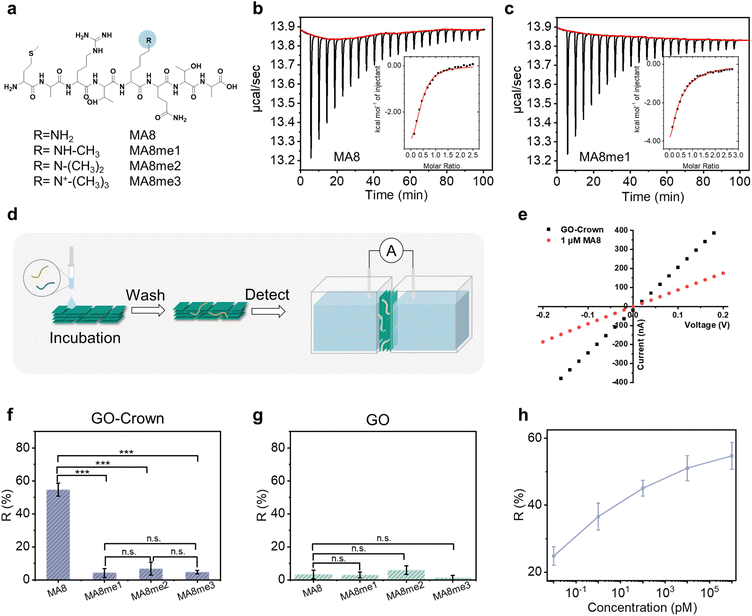
To verify the feasibility of the Crown probe for recognizing and screening unmethylated peptides and methylated peptides, a set of model peptides (NH2-MARTKQTA, termed MA8) was used, in which K could undergo methylation modification at different levels (mono-, di-, and tri-methylated, termed MA8me1, MA8me2 and MA8me3, respectively) (Fig. 3a and S5–S9).49 Then, the binding force between the Crown probe and the peptides was measured by isothermal titration calorimetry (ITC) tests. As shown in Fig. 3b, c and S10, the dissociation constant Kd of unmethylated peptides was 6.9 µM, while the Kd of methylated peptides was higher, demonstrating the higher binding force of the Crown probe toward unmethylated peptides and its strong potential to differentiate unmethylated and methylated peptides.After verifying the specific recognition ability of the Crown probe toward unmethylated peptides, the response performance of the GO-Crown system was investigated (Fig. 3d). Under incubation conditions with a peptide concentration of 1 µM, the R value of the GO-Crown membrane for MA8 peptides was 54.7% ± 4.0%, and the R values for MA8me1, MA8me2 and MA8me3 peptides were 4.2% ± 2.7%, 6.8% ± 3.9%, and 4.7% ± 1.1%, respectively. By means of incubation, the response of the GO-Crown membrane to MA8 peptides and methylated peptides showed significant differences. In contrast, for GO membranes, the responses of the R values were all less than 6% for MA8 peptides and methylated peptides (Fig. 3e–g and S11). To explore the response performance of the GO-Crown membrane toward MA8 peptides, the concentrations of MA8 peptides were set at 10 fM, 1 pM, 100 pM, 10 nM, and 1 µM. It could be seen from Fig. 3h and S12 that with the increase of peptide concentration, the R value gradually increased from 24.8% ± 2.7% to 54.7% ± 4.0%, which proved that, in the incubation form, the GO-Crown membrane could achieve sensitive response performance to changes in peptide concentration.
Next, the response performance of the GO-Crown system under electric drive was investigated (Fig. 4a). To obtain the best electric driving time, the ion current variation of the GO-Crown membrane was measured after different driving times ranging from 15 min to 10 h. As shown in Fig. 4a and S13, when unmethylated peptides passed through the membrane under electric drive from the driving chamber to the receiving chamber, they could be captured by the Crown probes on the membrane, thus inducing a decrease in current. When the electric drive time was 15 min, the R value (defined as (I0 − I)/I0, where I0 and I are the current intensities at +0.2 V before and after driving, respectively) was about 18.0% ± 3.9%. Increasing the time to 1 h increased the R value to 44.6% ± 2.8%. With further increases in time, the R value gradually increased and then reached a plateau at about 50%. Therefore, 1 h was selected as the optimal driving time. The I–T result also showed that the current gradually decreased with the increase of time and the current value stabilized around −900 nA at about 1 h (Fig. S14).
Next, the screening ability of 2D nanochannels between methylated and unmethylated peptides under electric drive for 1 h was investigated. As shown in Fig. 4b and S15, the GO membrane without the capture probe was used to detect the peptides. As expected, there was no significant difference between methylated and unmethylated peptides (p > 0.05), and the current signals of both methylated and unmethylated peptides were less than 10%, demonstrating that GO itself could not recognize and distinguish the two kinds of peptides. Meanwhile, the GO-Crown membrane efficiently captured MA8, thus leading to a remarkable decrease in ion current and the R value was 44.6% ± 2.8%. In contrast, the R values for MA8me1, MA8me2, MA8me3 were 6.2% ± 1.3%, 6.0% ± 1.1% and 3.1% ± 2.4%, respectively, which were much lower than that of MA8 (p < 0.001), indicating that the GO-Crown membrane could only capture the unmethylated peptides, instead of methylated peptides including mono-, di- and tri-methylated, and cause the ion current change.
To further confirm that the variation in current was induced by the binding between the Crown probe and unmethylated peptides, composite membrane with different contents of Crown, namely “GO/GO-Crown,” were constructed by layer-by-layer pumping. The CA variation proved their successful construction (Fig. S16). Then, the current signal response performance of the composite membrane was tested. As shown in Fig. 4c and S17, GO/GO-Crown could not bind MA8me2 peptides and did not induce current changes. Compared to the pure GO-Crown membrane, the R value of GO/GO-Crown for MA8 peptides was about 24%, demonstrating that as the proportion of the Crown probe decreased, the capture capability of the membrane for unmethylated peptides was distinctly weakened.
Subsequently, the MA8 peptide concentration–response curve of the GO-Crown membrane system was examined, as shown in Fig. 4d and S18. When the MA8 peptide was only 10 fM, the R value was 16.3% ± 1.6%. As the peptide concentration increased to 1 µM, R value gradually increased and reached ∼45%, indicating that, in the form of electric drive, the GO-Crown membrane was sensitive to unmethylated peptides, and the current decrease ratio exhibited a positive relationship to the concentration of unmethylated peptides. The plot showed good linearity at concentrations of 10 fM–1 µM, and the linear regression equation is R = 3.67![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) lg
lg![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) X [with the concentration in pM of MA8] +24.18, with a correlation coefficient of 0.997.
X [with the concentration in pM of MA8] +24.18, with a correlation coefficient of 0.997.
To explore the saturation concentration of the GO-Crown membrane, the concentration of MA8 peptides was further enhanced. As shown in Fig. S19, when the concentration was increased from 1 µM to 25 µM, the R value did not increase obviously; the R values of 5 µM and 25 µM MA8 were about 48%, respectively. After statistical analysis, there was no significant difference in response between 1 µM, 5 µM and 25 µM MA8 peptide (p > 0.05), confirming that the saturation concentration of the peptide was 1 µM. In addition, to realistically simulate the working conditions of peptide fragments from histone degradation, a mixture of methylated and unmethylated peptides, exemplified by dimethylated peptides, was detected by the GO-Crown membrane. The R value of the mixed peptide (1 µM MA8 + 0.1 µM MA8me2) was 49.5% ± 2.0%, which was nearly the same as that of 1 µM MA8 without the addition of MA8me2, and it was found that no significant difference existed. The situation of the mixed peptide with a high concentration (25 µM MA8 + 10 µM MA8me2) was similar, illustrating that the existence of methylated peptides in the mixed solution would not obviously affect the recognition of unmethylated peptides.
In histone lysates, methylated peptides generally occupy a small proportion, leading to difficulty in their detection; in this case, the enrichment of methylated peptides plays a crucial role.50,51 Thanks to the screening ability of methylated and unmethylated peptides, the GO-Crown membrane could also be utilized for enrichment of methylated peptides. When mixed peptides pass through the membrane under electric drive, unmethylated peptides bind to the Crown probe on the membrane, while the methylated peptides do not bind to the probe and are driven to the receiving chamber, achieving the enrichment of the methylated peptides in the mixed sample. UV-vis spectrophotometry and mass spectrometry were used to verify this process. As shown in Fig. 4f and S20, four peptides passed through the GO-Crown membrane under electric drive and their concentration variation was monitored by UV-vis absorbance, respectively. Compared to the approximately same absorbance (p > 0.05) before driving, the absorption value of methylated peptides was in the range of 0.1–0.2, which is about 5–10 times of unmethylated peptides (about 0.01) and showed a significant difference (p < 0.01). After enrichment, the absorption value of the unmethylated peptide was too low to obtain the converted concentration, while the concentrations of MA8me1, MA8me2 and MA8me3 were 0.0915 µM, 0.103 µM and 0.145 µM, respectively, suggesting the enrichment ability of the GO-Crown membrane for methylated peptides. Then, the solution in the receiving chamber was collected after driving a mixed solution (1 µM MA8 + 0.1 µM MA8me2) and characterized by mass spectrometry. Only the peaks from MA8me2 (m/z value of [M + H]+ 934.51 and [M + 2H]2+ 467.76) could be found, rather than MA8 (m/z value of [M + H]+ 906.48 and [M + 2H]2+ 453.75) (Fig. S21). These results further proved that the GO-Crown membrane system could be used for the separation of unmethylated and methylated peptides and had the enrichment ability for methylated peptides.
In the GO-Crown membrane system, the electric driving form played a crucial role in the process of peptides passing through the nanochannel membrane. In this complex process, both the electric field force and the concentration difference had significant effects on the transmembrane behavior of peptides. The electric field force exerted a specific driving force when the peptide molecules passed through the nanochannel membrane by its directional effect on the charged particles. The difference in concentration was generated due to the difference in the concentration of peptides between the two sides of the electrolytic cell. The imbalance of chemical potential promoted the diffusion of peptides from the region of high concentration to the region of low concentration, and then pushed the peptides through the nanochannel membrane. Three models were designed to explore the influence of FC (concentration difference) and FE (electric field force) in the electric drive model (Fig. S22 and S23). First, to investigate the influence of FE, the concentration was the same on both sides of the cell (FC = 0). The results showed that there was no significant difference between the two solutions, with the UV absorption values was around 0.7, so it was speculated that the influence of FE was tiny. Second, to investigate the influence of Fc, the electric field force was set to 0 (FE = 0). The result showed that the UV absorption value of the solution on the other side was about 0.06, which proved that there were peptides driven to the other side. Third, there were two cases in which FE and FC were in the same direction or in opposite directions.
The results showed that when FE and FC were in the same direction, the absorption value of the solution collected on the other side was about 0.13, while when FE and FC were in opposite directions, the absorption value was about 0.08. In summary, it could be concluded that both FE and FC had a certain effect on the driving force of peptides under electric drive, and it was mainly driven by FC, with the driving effect being optimal when FE and FC were combined in the same direction.
Evaluating G9a responsiveness of the system
Encouraged by the satisfactory separating ability of the GO-Crown membrane toward methylated and unmethylated peptides, its application in detecting G9a was considered, since G9a could catalyze the methylation of specific positions in histone. According to literature reports,52–54 G9a interacts specifically with 6–11 amino acid residues of H3, and the smallest in vitro substrate recognition unit is heptapeptide with the TARKSTG (TG7) sequence. In this case, the TG7 peptide was used here as a detecting probe, combined with the GO-Crown membrane to construct a G9a detection system. In the presence of G9a, the substrate TG7 peptide could be methylated to mono-, di, and tri-methylated peptides (termed TG7me1, TG7me2 and TG7me3) (Fig. S24). Then, the reaction liquid was added to the electrolytic cell for electric drive, and the current response of the GO-Crown membrane was recorded, which was related to the residual TG7 peptide content and also G9a content.First, to confirm the methylation ability and process of G9a toward the TG7 peptide, the products of the enzymatic reaction after different reaction times were characterized by mass spectrometry (Fig. 5a and S25–S27). The theoretical m/z value [M + H]+ of TG7 and TG7me1 was calculated to be 720.40 and 735.04, respectively and the peaks at 720.40 and 735.04 could be found when the reaction time was 1 h, proving the co-existence of TG7 and TG7me1 and the occurrence of methylation at this time. When the reaction time reached 2 h and 3 h, except for the peak of TG7 and TG7me1, the peak at 748.43, which was from TG7me2 (theoretical m/z value [M + H]+ of 748.43) could also be found, and the ratio of peak intensity for TG7me2/TG7me1 was increased from 2 h to 3 h. At the 4-h time point, the TG7me3 peptide emerged. The above results demonstrated that the methylation process of G9a toward the TG7 peptide occurred as expected, and with extended reaction time, the methylation degree of TG7 could be gradually strengthened.
Then, the feasibility of this combined system to detect G9a was investigated. The reaction solution of G9a (20 nM) and TG7 peptide (10 µM) after different reaction times was added to the electrolytic cell for testing. As shown in Fig. 5b, S28 and S29, when the reaction time was 0 h, the current signal response R of the GO-Crown membrane was 42.6% ± 2.0%, confirming that unmethylated TG7 could be captured by GO-Crown and lead to the ion current decrease, which was the same as that of the MA8 peptide. Meanwhile, with the increase of reaction time to 7 h, the TG7 peptide could be methylated to different degrees and thus could not bind to GO-Crown to induce current variation. The R value decreased to 34.13% at 1 h, continued to decline during 6 h and reached a plateau (3.45%) at 6 h, proving the above process and the potential of the system for detecting G9a. For comparison, the TG7 peptide solution without G9a was also added to the electrolytic cell after different times under the same conditions. With the increase of reaction time, the R value of the system for the enzyme-free mixture remained at about 42%, and there was no obvious change in current response, which proved that the current signal of the GO-Crown membrane system was caused by enzymatic action of G9a. Since after 6 h, the process of enzyme catalyzed reaction was maximal and approximately 100%, the following detection experiments selected 6 h as the reaction time (Fig. 5c).
The response performance of the GO-Crown membrane system to G9a concentration in the range of 20 fM–20 nM was investigated (Fig. 5d and S30). When the G9a concentration was in the range of 20 fM–2 pM, the GO-Crown membrane system produced an obvious current signal response with the change of G9a concentration, and the R value gradually changed from 42.1% ± 0.9% to 5.6% ± 1.0%. When the G9a concentration was in the low concentration range of 20 fM–200 fM, the fitted curve had good linearity and the linear regression equation is Y = 0.10 × [concentration of G9a] +54.70, with a correlation coefficient of 0.981 (Fig. 5e). The limit of detection (3σ/slope, σ = 0.688) is 21 fM, which was much lower than that of reported methods (Table S3). These results proved the G9a detection ability of the system and also laid a foundation for subsequent detection in complex environments. Moreover, when increasing the G9a concentration from 2 pM to 20 nM, the current signal was basically unchanged and the R value was about 4–5%, suggesting that the response of the GO-Crown membrane system had reached a saturation state.
By virtue of well G9a responsiveness ability of the system, the enzymatic reaction kinetics of G9a was further discussed (Fig. 5f and S31). To explore the effect of substrate concentration on enzymatic reaction kinetics, the temperature (25 °C), enzyme concentration (20 nM) and reaction time (6 h) were controlled. The concentrations of substrate TG7 peptides were set at 0.05 µM, 0.25 µM, 0.5 µM, 1.5 µM, 2.5 µM, and 5 µM successively. The difference of R at different concentrations was taken as the vertical coordinate. The value of ΔR gradually increased with the increase of concentration. By linear fitting of the obtained current signal data, the Km value of the enzyme activity fitted under the experimental conditions was 1.71 µM.
Specificity is an important factor in evaluating the response performance of detection systems. As shown in Fig. 5g and S32, the 1 − R of the GO-Crown membrane system towards G9a was 0.97. However, the current response of the system to other enzymes, including α-chymotrypsin, phosphatase and bovine serum albumin (BSA), was lower than 0.6. The significant difference demonstrated reliable specificity for G9a. In addition, G9a was also deactivated by heating, and the 1 − R value after the reaction of the GO-Crown membrane system with deactivated G9a was 0.57, proving that the activity of G9a was the key factor inducing the change in ionic current.
Detection of G9a from cells in different application scenarios
With good selectivity and sensitivity, the performance of the GO-Crown membrane system for detecting G9a from MCF-7 cells (breast cancer cell line with high G9a expression) was investigated (Fig. 6a, b, S33 and S34). Along with the MCF-7 cell concentration rising from 50 cells per mL to 5 × 104 cells per mL, the current change ratio R value decreased gradually, proving that there was a positive relationship between concentrations of G9a and the current change ratio. The variation tendency was similar to the G9a concentration–response curves in Fig. 5f. However, when incubated with G9a from the MCF-10A cells (normal mammary epithelial cells with low G9a expression), TG7 peptides cannot be methylated, thus resulting in an unapparent change in ion current, even when the cell concentration was up to 5 × 104 cells per mL.After successfully applying the system to precisely detect G9a from cells, we wonder if this system could be used in diverse application scenarios, including differentiation of different subtypes of breast cancer cells, evaluation of inhibitor action, and comparison of different oxygen contents, since the secretion of G9a was highly related to these factors. First, three subtypes of breast cancer cells: MCF-7 cells, MDA-231 cells and MDA-468 cells were selected. At the same time, MCF-10A was treated as the control group. The four cell lines were obtained from Wuhan Pricella Biotechnology Co., Ltd. Cell lysates from the four cell lines at a concentration of 5 × 103 cells per mL were incubated with TG7 peptides, respectively, and then added into the GO-Crown membrane system under electrical drive. As shown in Fig. 6c and S35, the GO-Crown membrane system had obvious differences in electrical signal responses to cancer cells and normal cells. the 1 − R value was 58.7% ± 0.9% for MCF-10A cells, 79.4% ± 1.7% for MCF-7 cells, 74.1% ± 0.5% for MDA-231 cells, and 68.2% ± 1.0% for MDA-468 cells, and there was a significant difference (p < 0.001) between each two groups, indicating that the system could realize the distinction between cancer cells and normal cells relying on the different expression of G9a. For different subtypes of breast cancer cells, the G9a detected by the system from high to low was in the order of MCF-7 cells, MDA-231 cells and MDA-468 cells, and the electrical signal response was significantly different, proving that the system could realize the differentiation of different subtypes of breast cancer cells. To confirm the accuracy of tendency, the enzyme linked immunosorbent assay (ELISA) analysis, as a gold standard, was further utilized to measure the G9a content in cell lysate of the four kinds of cells. The G9a concentration in the four cell lines was significantly different and was calculated to be 0.234 ng mL−1, 0.906 ng mL−1, 0.759 ng mL−1 and 0.497 ng mL−1 in MCF-10A cells, MCF-7 cells, MDA-231 cells and MDA-468 cells, respectively, which was consistent with the detection results of the functionalized GO-Crown membrane, proving the reliable detection ability of the system for differentiating different subtypes of breast cancer cells.
In another application scenario, an inhibitor of G9a UNC068 with low toxicity was used (Fig. S36), which could interfere with G9a's normal interaction with substrates and cofactors (such as SAM) by binding to the active site of G9a or key regions near it, thereby impeding its ability to catalyze methyl-transfer reactions. The concentration of the inhibitor was set at 5 nM, 150 nM, and 1.5 µM, and the expression level of G9a in the four cell lysates was detected after co-culture for 72 h (Fig. 6d and S37). With the increase of inhibitor concentration, for MCF-10A cells, the R value did not change significantly and fluctuated within the range of ∼43%. For MCF-7 cells, the R value changed from ∼21% to ∼32%. For MDA-231 cells, the R value changed from ∼26% to ∼36%. For MDA-468 cells, the R value changed from ∼32% to ∼40%. For convenience of comparison, “1 − R” was selected as the vertical coordinate. It was demonstrated that the GO-Crown membrane system had a reliable ability to detect G9a in an environment where the expression of G9a was down-regulated after inhibitor treatment. ELISA kit was used to detect cell lysate treated with 5 nM, 150 nM, and 1.5 µM inhibitor, which was consistent with the results determined by the GO-Crown membrane, proving the reliable detection capability of the system.
It has been reported that G9a could be up-regulated in hypoxic environments. In brief, under hypoxia conditions, oxygen receptors in cells would sense the reduction of oxygen content, thereby stabilizing and activating hypoxia-inducible factor (HIF-1α). Activated HIF-1α could bind to the hypoxia response elements of the G9a gene promoter, recruiting transcription-related complexes, promoting the binding of RNA polymerase to the G9a gene promoter, thus enhancing the transcription of G9a gene and making G9a expression up-regulated.55–57 In this case, different levels of oxygen content were constructed for cells, including 21% normoxia, 10% hypoxia and 1% hypoxia. The GO-Crown membrane system had a significant response to the expression of G9a in MCF-7 cells at different oxygen contents, with ∼13% at 1% oxygen concentration and ∼17% at 10% oxygen concentration (Fig. 6e and S38). For MCF-10A cells, the values were ∼42% at both 1% oxygen concentration and 10% oxygen concentration, indicating that a hypoxic environment had no significant effect on the expression of G9a in MCF-10A cells. The cell lysates cultured in hypoxic environments were detected by ELISA kit, which was consistent with the results determined by the GO-Crown system. Therefore, the system could be applied in hypoxic environments where G9a expression was upregulated with high sensitivity. The GO-Crown membrane system had the certain ability to distinguish between 1% hypoxia and 10% hypoxia conditions, so it was considered that the system had a potential application as an oxygen indicator.
Conclusions
To construct a label-free, highly selective, sensitive, and simple tool for G9a detection in native biological matrices, a G9a detection system based on functionalized 2D solid-state nanochannel membranes was developed in this study through the specific recognition between small molecule probes that anchored on the surface and dissociative peptide probes. The confined environment in nanochannels enhances interaction times and efficiency between the peptide and the crown ether receptors on the channel walls. This multivalent effect amplifies the subtle chemical difference between methylated and unmethylated peptides into a significant and measurable electrical signal. The excellent sensing performance enables diverse detection scenarios, including differentiation of different subtypes of breast cancer cells, screening of enzyme inhibitors and sensing of hypoxic cellular environments. This research proposes that the GO-Crown membrane could be an effective detection system for G9a, which provides a powerful tool for cancer research and clinical application.Author contributions
F. X., X. L. and J. J. H. conceived and supervised the project. J. J. H. and N. L. wrote the manuscript. W. J. and N. L. conducted the experiments. R. L., J. J. H., W. J. and N. L. contributed to data analysis. All authors contributed to manuscript revision.Conflicts of interest
There are no conflicts to declare.Data availability
Supplementary information: experimental procedures, characterization data, and I–V curves. See DOI: https://doi.org/10.1039/d5sc09089j.Acknowledgements
The authors acknowledge financial support by the National Natural Science Foundation of China (22090050, U24A20502, and 22474131), the National Key R&D Program of China (2020YFA0211200 and 2021YFA1200403), the Shenzhen Science and Technology Program (No. JCYJ20230807113706013 and JCYJ20220530162406014), and the Natural Science Foundation of Hubei Province (2024AFA001 and 2023AFB636).Notes and references
- D. Y. Lee, C. Teyssier, B. D. Strahl and M. R. Stallcup, Role of protein methylation in regulation of transcription, Endocr. Rev., 2005, 26, 147–170 Search PubMed.
- J. Li, M. Zhang, W. Ma, B. Yang, H. Lu, F. Zhou and L. Zhang, Post-translational modifications in liquid-liquid phase separation: a comprehensive review, Mol. Biomed., 2022, 3, 13 CrossRef PubMed.
- Y. Liu, W. Feng, Y. Wang and B. Wu, Crosstalk between protein post-translational modifications and phase separation, Cell Commun. Signaling, 2024, 22, 110 CrossRef.
- A. Bird, Methylation talk between histones and DNA, Science, 2001, 294, 2113–2115 CrossRef CAS PubMed.
- J. L. Miller and P. A. Grant, The role of DNA methylation and histone modifications in transcriptional regulation in humans, Epigenetics: Development and Disease, 2012, 289–317 CAS.
- F. Casciello, K. Windloch, F. Gannon and J. S. Lee, Functional role of G9a histone methyltransferase in cancer, Front. Immunol., 2015, 6, 487 Search PubMed.
- A. Jana, R. Naga, S. Saha, C. Griñán-Ferré and D. R. Banerjee, Integration of ligand and structure-based pharmacophore screening for the identification of novel natural leads against Euchromatic histone lysine methyltransferase 2 (EHMT2/G9a), J. Biomol. Struct. Dyn., 2024, 42, 3535–3562 CrossRef CAS PubMed.
- C.-P. Chaturvedi, B. Somasundaram, K. Singh, R. L. Carpenedo, W. L. Stanford, F. J. Dilworth and M. Brand, Maintenance of gene silencing by the coordinate action of the H3K9 methyltransferase G9a/KMT1C and the H3K4 demethylase Jarid1a/KDM5A, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 18845–18850 CrossRef CAS PubMed.
- D. Patnaik, H. G. Chin, P.-O. Esteve, J. Benner, S. E. Jacobsen and S. Pradhan, Substrate specificity and kinetic mechanism of mammalian G9a histone H3 methyltransferase, J. Biol. Chem., 2004, 279, 53248–53258 Search PubMed.
- P. Rathert, A. Dhayalan, M. Murakami, X. Zhang, R. Tamas, R. Jurkowska, Y. Komatsu, Y. Shinkai, X. Cheng and A. Jeltsch, Protein lysine methyltransferase G9a acts on non-histone targets, Nat. Chem. Biol., 2008, 4, 344–346 Search PubMed.
- S. R. Shankar, A. G. Bahirvani, V. K. Rao, N. Bharathy, J. R. Ow and R. Taneja, G9, a multipotent regulator of gene expression, Epigenetics, 2013, 8, 16–22 CrossRef CAS PubMed.
- Y. F. Wang, J. Zhang, Y. Su, Y. Y. Shen, D. X. Jiang, Y. Y. Hou, M. Y. Geng, J. Ding and Y. Chen, G9a regulates breast cancer growth by modulating iron homeostasis through the repression of ferroxidase hephaestin, Nat. Commun., 2017, 8, 274 Search PubMed.
- L. Wei, D. K. C. Chiu, F. H. C. Tsang, C. T. Law, C. L. H. Cheng, S. L. K. Au, J. M. F. Lee, C. C. L. Wong, I. O. L. Ng and C. M. Wong, Histone methyltransferase G9a promotes liver cancer development by epigenetic silencing of tumor suppressor gene RARRES3, J. Hepatol., 2017, 67, 758–769 Search PubMed.
- C. Segovia, E. San Jose-Eneriz, E. Munera-Maravilla, M. Martínez-Fernández, L. Garate, E. Miranda, A. Vilas-Zornoza, I. Lodewijk, C. Rubio and C. Segrelles, Inhibition of a G9a/DNMT network triggers immune-mediated bladder cancer regression, Nat. Med., 2019, 25, 1073–1081 Search PubMed.
- R. P. Pangeni, L. Yang, K. Zhang, J. Wang, W. Li, C. Guo, X. Yun, T. Sun, J. Wang and D. J. Raz, G9a regulates tumorigenicity and stemness through genome-wide DNA methylation reprogramming in non-small cell lung cancer, Clin. Epigenet., 2020, 12, 1–17 Search PubMed.
- S. T. Cha, C. T. Tan, C. C. Chang, C. Y. Chu, W. J. Lee, B. Z. Lin, M. T. Lin and M. L. Kuo, G9a/RelB regulates self-renewal and function of colon-cancer-initiating cells by silencing Let-7b and activating the K-RAS/β-catenin pathway, Nat. Cell Biol., 2016, 18, 993–1005 Search PubMed.
- J. R. Haebe, C. J. Bergin, T. Sandouka and Y. D. Benoit, Emerging role of G9a in cancer stemness and promises as a therapeutic target, Oncogenesis, 2021, 10, 76 Search PubMed.
- H. He, X. Li, F. Su, H. Jin, J. Zhang and Y. Wang, Current and Emerging Approaches Targeting G9a for the Treatment of Various Diseases, J. Med. Chem., 2024, 68, 1068–1089 CrossRef PubMed.
- M. Amjadi, T. Hallaj and N. Hildebrandt, A sensitive homogeneous enzyme assay for euchromatic histone-lysine-N-methyltransferase 2 (G9a) based on terbium-to-quantum dot time-resolved FRET, BioImpacts, 2020, 11, 173 Search PubMed.
- T. Bonaldi, J. T. Regula and A. Imhof, The use of mass spectrometry for the analysis of histone modifications, Methods Enzymol., 2004, 377, 111–130 Search PubMed.
- A. Dhayalan, E. Dimitrova, P. Rathert and A. Jeltsch, A continuous protein methyltransferase (G9a) assay for enzyme activity measurement and inhibitor screening, J. Biomol. Screening, 2009, 14, 1129–1133 Search PubMed.
- Y. Xiong, M. Li, Y. Cao, Z. Li, Y. Chang, X. Zhao and G. Qing, Nanofluidic device for detection of lysine methylpeptides and sensing of lysine methylation, Anal. Chem., 2023, 95, 7761–7769 CrossRef CAS PubMed.
- M. Tachibana, Y. Matsumura, M. Fukuda, H. Kimura and Y. Shinkai, G9a/GLP complexes independently mediate H3K9 and DNA methylation to silence transcription, EMBO J., 2008, 27, 2681–2690 Search PubMed.
- T. Yokochi, K. Poduch, T. Ryba, J. Lu, I. Hiratani, M. Tachibana, Y. Shinkai and D. M. Gilbert, G9a selectively represses a class of late-replicating genes at the nuclear periphery, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 19363–19368 CrossRef CAS PubMed.
- Y. Qiao, J. Hu, Y. Hu, C. Duan, W. Jiang, Q. Ma, Y. Hong, W. Huang, F. Xia and X. Lou, Detection of Unfolded Cellular Proteins Using Nanochannel Arrays with Probe-Functionalized Outer Surfaces, Angew. Chem., Int. Ed., 2023, 62, e202309671 CrossRef CAS PubMed.
- Y. Huang, L. Liu, C. Luo, W. Liu, X. Lou, L. Jiang and F. Xia, Solid-state nanochannels for bio-marker analysis, Chem. Soc. Rev., 2023, 52, 6270–6293 Search PubMed.
- J. J. Hu, W. Jiang, Y. Qiao, Q. Ma, Q. Du, J. Jiang, X. Lou and F. Xia, Enzyme regulating the wettability of the outer surface of nanochannels, ACS Nano, 2023, 17, 11935–11945 Search PubMed.
- L. Wang, H. Li, L. Shi, L. Li, F. Jia, T. Gao and G. Li, In situ peptide self-assembly on ionic nanochannel for dynamic monitoring of MMPs in extracellular matrix, Biosens. Bioelectron., 2022, 195, 113671 Search PubMed.
- N. Soni, N. Freundlich, S. Ohayon, D. Huttner and A. Meller, Single-file translocation dynamics of SDS-denatured, whole proteins through sub-5 nm solid-state nanopores, ACS Nano, 2022, 16, 11405–11414 CrossRef CAS PubMed.
- Y. Huang, W. Zhang, F. Xia and L. Jiang, Solid-state nanochannel-based sensing systems: Development, challenges, and opportunities, Langmuir, 2022, 38, 2415–2422 Search PubMed.
- Q. Ma, Y. Li, R. Wang, H. Xu, Q. Du, P. Gao and F. Xia, Towards explicit regulating-ion-transport: nanochannels with only function-elements at outer-surface, Nat. Commun., 2021, 12, 1573 CrossRef CAS PubMed.
- W. Zhang, Y. Tu, H. Liu, R. Liu, X. Zhang, L. Jiang, Y. Huang and F. Xia, A Single Set of Well-Designed Aptamer Probes for Reliable On-site Qualitative and Ultra-Sensitive Quantitative Detection, Angew. Chem., Int. Ed., 2024, 63, e202316434 CrossRef CAS PubMed.
- X. Li, T. Zhai, P. Gao, H. Cheng, R. Hou, X. Lou and F. Xia, Role of outer surface probes for regulating ion gating of nanochannels, Nat. Commun., 2018, 9, 40 Search PubMed.
- P. Gao, Q. Ma, D. Ding, D. Wang, X. Lou, T. Zhai and F. Xia, Distinct functional elements for outer-surface anti-interference and inner-wall ion gating of nanochannels, Nat. Commun., 2018, 9, 4557 CrossRef PubMed.
- Z. Shang, J. Zhao, M. Yang, Y. Xiao, W. Chu, S. Xu, X. Zhang, X. Yi, M. Lin and F. Xia, Precise control of transmembrane current via regulating bionic lipid membrane composition, Sci. Adv., 2024, 10, eadq0118 CrossRef CAS PubMed.
- Z. Zhang, L. Wen and L. Jiang, Bioinspired smart asymmetric nanochannel membranes, Chem. Soc. Rev., 2018, 47, 322 RSC.
- S. Wang, L. Yang, G. He, B. Shi, Y. Li, H. Wu, R. Zhang, S. Nunes and Z. Jiang, Two-dimensional nanochannel membranes for molecular and ionic separations, Chem. Soc. Rev., 2020, 49, 1071 RSC.
- J. Wang, H. Zhou, S. Li and L. Wang, Selective Ion Transport in Two-Dimensional Lamellar Nanochannel Membranes, Angew. Chem., Int. Ed., 2023, 62, e202218321 Search PubMed.
- A. Bhardwaj, M. V. Surmani Martins, Y. You, R. Sajja, M. Rimmer, S. Goutham, R. Qi, S. A. Dar, B. Radha and A. Keerthi, Fabrication of angstrom-scale two-dimensional channels for mass transport, Nat. Protoc., 2024, 19, 240 CrossRef CAS PubMed.
- P. M. Sajeer, A. Bhardwaj, B. Radha, M. Varma and A. Keerthi, Microtomy-fabricated two-dimensional nano-slits enable single molecule biosensing, Nanoscale, 2025, 17, 18605 RSC.
- S. K. Tiwary, M. Singh, S. V. Chavan and A. Karim, Graphene oxide-based membranes for water desalination and purification, npj 2D Mater. Appl., 2024, 8, 27 Search PubMed.
- J. Wu, Y. Zhang, J. Hu, Y. Yang, D. Jin, W. Liu, D. Huang, B. Jia and D. Moss, 2D graphene oxide films expand functionality of photonic chips, Adv. Mater., 2024, 36, 2403659 CrossRef CAS PubMed.
- J. Fan, Y. Pan, H. Wang and F. Song, Efficient reverse osmosis-based desalination using functionalized graphene oxide nanopores, Appl. Surf. Sci., 2024, 674, 160937 Search PubMed.
- H. Liu, X. Huang, Y. Wang, B. Kuang and W. Li, Nanowire-assisted electrochemical perforation of graphene oxide nanosheets for molecular separation, Nat. Commun., 2024, 15, 164 Search PubMed.
- T. Lee and B.-S. Kim, Two-dimensional designer nanochannels for controllable ion transport in graphene oxide nanomembranes with tunable sheet dimensions, ACS Appl. Mater. Interfaces, 2020, 12, 13116–13126 CrossRef CAS PubMed.
- Y. Wu, C.-F. Fu, Q. Huang, P. Zhang, P. Cui, J. Ran, J. Yang and T. Xu, 2D heterostructured nanofluidic channels for enhanced desalination performance of graphene oxide membranes, ACS Nano, 2021, 15, 7586–7595 CrossRef CAS PubMed.
- D. R. Dreyer, S. Park, C. W. Bielawski and R. S. Ruoff, The chemistry of graphene oxide, Chem. Soc. Rev., 2010, 39, 228–240 Search PubMed.
- B. S. Bohra, P. Singh, A. Rana, H. Sharma, T. Arya, M. Pathak, A. Chaurasia, S. Rana and N. G. Sahoo, Specific functionalized graphene oxide-based vitrimer epoxy nanocomposites for self-healing applications, Compos. Sci. Technol., 2023, 241, 110143 CrossRef CAS.
- Q. Sheng, C. Wang, X. Li, H. Qin, M. Ye, Y. Xiong, X. Wang, X. Li, M. Lan and J. Li, Highly efficient separation of methylated peptides utilizing selective complexation between lysine and 18-Crown-6, Anal. Chem., 2020, 92, 15663–15670 Search PubMed.
- N. G. Hartel, B. Chew, J. Qin, J. Xu and N. A. Graham, Deep protein methylation profiling by combined chemical and immunoaffinity approaches reveals novel PRMT1 targets, Mol. Cell. Proteomics, 2019, 18, 2149–2164 Search PubMed.
- Z. Li, Q. Wang, Y. Wang, K. Wang, Z. Liu, W. Zhang and M. Ye, An efficient approach based on basic strong cation exchange chromatography for enriching methylated peptides with high specificity for methylproteomics analysis, Anal. Chim. Acta, 2021, 1161, 338467 Search PubMed.
- H. G. Chin, M. Pradhan, P. O. Estève, D. Patnaik, T. C. Evans and S. Pradhan, Sequence specificity and role of proximal amino acids of the histone H3 tail on catalysis of murine G9A lysine 9 histone H3 methyltransferase, Biochemistry, 2005, 44, 12998–13006 Search PubMed.
- J. Lee, J. Chen, P. Sarkar, M. Xue, R. J. Hooley and W. Zhong, Monitoring the crosstalk between methylation and phosphorylation on histone peptides with host-assisted capillary electrophoresis, Anal. Bioanal. Chem., 2020, 412, 6189–6198 CrossRef CAS PubMed.
- C. Poulard, L. M. Noureddine, L. Pruvost and M. Le Romancer, Structure, activity, and function of the protein lysine methyltransferase G9a, Life, 2021, 11, 1082 CrossRef CAS PubMed.
- F. Casciello, F. Al-Ejeh, G. Kelly, D. J. Brennan, S. F. Ngiow, A. Young, T. Stoll, K. Windloch, M. M. Hill and M. J. Smyth, G9a drives hypoxia-mediated gene repression for breast cancer cell survival and tumorigenesis, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 7077–7082 Search PubMed.
- M. Ivan, K. Kondo, H. Yang, W. Kim, J. Valiando, M. Ohh, A. Salic, J. M. Asara, W. S. Lane and W. G. Kaelin Jr, HIFα targeted for VHL-mediated destruction by proline hydroxylation: implications for O2 sensing, Science, 2001, 292, 464–468 Search PubMed.
- J. Kang, S. H. Shin, H. Yoon, J. Huh, H. W. Shin, Y. S. Chun and J. W. Park, FIH is an oxygen sensor in ovarian cancer for G9a/GLP-driven epigenetic regulation of metastasis-related genes, Cancer Res., 2018, 78, 1184–1199 Search PubMed.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |