Open Access Article
Open Access ArticleLarge-scale protein conformational transitions revealed by weighted ensemble simulations and EPR
Shramana
Palit†
a,
Darian T.
Yang†
 a,
Xiaowei
Bogetti
ab,
Anthony T.
Bogetti
a,
Xiaowei
Bogetti
ab,
Anthony T.
Bogetti
 ab,
Olivia
Wood
a,
Sunil
Saxena
ab,
Olivia
Wood
a,
Sunil
Saxena
 *a and
Lillian T.
Chong
*a and
Lillian T.
Chong
 *a
*a
aDepartment of Chemistry, University of Pittsburgh, Pittsburgh PA 15260, USA. E-mail: ltchong@pitt.edu; sksaxena@pitt.edu
bLaufer Center for Physical and Quantitative Biology, Stony Brook University, Stony Brook, NY 11794, USA
First published on 26th March 2026
Abstract
Large-amplitude protein conformational changes are essential for cellular processes. Electron paramagnetic resonance (EPR) spectroscopy can detect such conformational changes by measuring distances between site-directed spin labels, but lacks atomistic detail. While standard molecular dynamics (MD) simulations provide atomistic details, the relevant timescales are often inaccessible. Here, we combine weighted ensemble path sampling with EPR distances to capture long-timescale protein conformational transitions in an unbiased manner. We simulated hundreds of pathways for a large-scale conformational transition between the closed and open states of the lysine/arginine/ornithine binding protein (LAOBP). Furthermore, we identified key residue-level interactions that distinguish the two states. Selective mutagenesis of these residues leads to stabilization of the open state of the protein. This approach integrates sparse EPR distances with atomistic simulations, revealing hidden protein states that evolve on milliseconds to seconds timescales.
Introduction
Many proteins undergo large-scale conformational changes to drive essential cellular processes, including signal transduction, enzyme catalysis, and molecular transport. These structural transitions enable a single protein to engage with different partners at distinct times or locations.1–4 Conformational changes may be triggered by ligand binding or by interactions with other substrates, proteins, and DNA.2,5–7 Understanding the mechanisms of these binding-induced conformational changes is critical for understanding how biological processes are regulated.Electron paramagnetic resonance (EPR) spectroscopy combined with site-directed spin labeling8–15 has emerged as a powerful method for probing protein conformational dynamics.1,16–23 Pulsed dipolar EPR techniques24–28 are particularly informative, as they provide not only average spin–spin distances, but also full distance distributions, which report on the relative populations of alternate states and the degree of conformational flexibility. A key limitation, however, is that such experiments are performed on flash-frozen protein ensembles. Thus, the measurements typically preclude time resolution, although recently time-resolution has been reported by utilizing microsecond freeze-hyperquenching.21 Moreover, EPR provides only sparse distance distributions, making computational modeling essential for interpreting these measurements into ensembles of atomistic structures.
Standard molecular dynamics (MD) simulations can capture protein motions at the atomic level on the microsecond time scale but are unable to sample large-scale protein conformational transitions on the milliseconds to seconds timescale. As an alternative, more rapid and approximate methods have been developed to generate protein structures that are consistent with EPR distances. These methods include (i) elastic network modeling methods, such as Multiscale Modeling of Macromolecules (MMM), to model conformational transitions;29,30 (ii) atomistic simulation methods that apply external forces such as restrained molecular dynamics31 and Modeling Employing Limited Data (MELD);32 (iii) atomistic simulation methods that alter the free energy landscape, such as metadynamics33 and simulated annealing;34–37 and (iv) atomistic structure prediction methods, including RosettaEPR38 and more recent deep-learning-guided approaches (e.g., AlphaFold2)39 that incorporate EPR distance restraints.40
In contrast, the weighted ensemble (WE)41,42 strategy is particularly powerful for sampling pathways and rates of rare events, including large-scale conformational transitions in proteins43–45 and protein–protein binding.46 Its efficiency comes from focusing simulation efforts on the transitions between stable states rather than the stable states themselves.42 In a WE simulation, multiple MD trajectories are run in parallel and assigned statistical weights. The algorithm then proceeds iteratively in two steps: (i) dynamics are propagated for a short time interval τ, and (ii) trajectories are resampled based on progress—typically along a chosen coordinate (the progress coordinate) that is partitioned into bins. During resampling, trajectories that occupy new bins are replicated (with weights split accordingly among child trajectories), while oversampled trajectories within a bin are occasionally terminated (with weights merged with those of surviving trajectories). This procedure ensures even coverage of the free energy landscape. Crucially, trajectory weights are rigorously tracked throughout, maintaining a statistically unbiased distribution with a total probability of one. Because weights are independent of the progress coordinate, both the coordinate and bin positions can be adjusted during a WE simulation.
The key advantage of WE sampling is its ability to generate unbiased pathways and direct estimates of rates—all without the use of any external biasing forces or alterations of the energy landscape. The main caveat of the WE strategy is the requirement of a proper progress coordinate to monitor the slowest relevant motion(s). Recently, we applied the WE strategy based on insights from EPR-measured distance distributions47 to explore the conformational landscape of a homo-dimeric protein, human glutathione S-transferase A1-1 (hGSTA1-1). The progress coordinate was designed to capture a large difference in EPR-measured distances for two alternate conformational states.47 Specifically, this progress coordinate monitored the dissociation of the α9 helix in one of the two subunits using a root-mean-squared deviation (RMSD) metric. In addition to sampling atomistic structures corresponding to both states, the WE simulations produced an ensemble of transition pathways for the interconversion of the states.
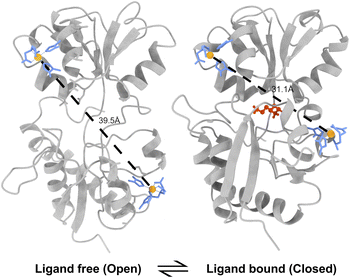
In the current work, we focus on the closed-to-open transition between two alternate states of a monomeric protein, lysine/arginine/ornithine binding protein (LAOBP), an extracellular receptor that mediates the uptake of nutrients such as amino acids.48,49 LAOBP is an ideal model system for studying large conformational transitions due to the availability of high-resolution crystal structures for its closed, ligand-bound and open, ligand-free states (Fig. 1).50 Due to the long timescales of the closed-to-open transition, previous simulation studies of LAOBP have either employed external biasing forces51 or relied on a large set of short conventional MD simulations.52 Using the WE strategy with the EPR distance directly as a progress coordinate, we simulated 381 closed-to-open pathways of the ligand-free LAOBP protein within 29 days on eight A100 GPUs. These simulations employed a two-dimensional WE progress coordinate that simultaneously monitored the long-range distance between EPR spin labels and tracked the protein's opening angle. We further identified key interactions that distinguish the closed and open states. We show that mutagenesis of these residues leads to stabilization of the open state. Together, these results establish a WE-based simulation workflow that integrates EPR-measured distances without introducing any bias in the dynamics and provides a general framework for studying large-scale protein conformational changes in other dynamic protein systems.
Results and discussion
In this work, we used the distance between EPR spin labels in LAOBP as a progress coordinate in weighted ensemble (WE) simulations to sample the transition pathway between the closed and open states of the protein. While there is a wide variety of spin labels used in EPR to study conformational changes in proteins,8,12,14,53,54 we chose the Cu(II)–nitrilotriacetic acid (NTA) spin label. In this spin label, Cu(II) coordinates with two strategically mutated histidine residues at i, i + 4 for the α helix, and i, i + 2 for the β sheet.14,55,56 The rigidity imparted by the coordination of two histidines to a Cu(II) ion enables distance measurements up to five times greater precision than traditional labels, making the label ideal to incisively resolve conformational changes.14,57EPR experiments
Choice of labeling sites
To determine potential spin label sites, we used the Multiscale Modeling of Macromolecules (MMM) method to scan potential Cu(II)-NTA labeling sites in the ligand-free (open) and ligand-bound (closed) structures of the LAOBP. This method predicts potential labeling sites by using rotamer libraries to select the most probable conformations of spin labels at various sites in the protein structure. We then filtered down the choice of labeling sites to pairs of sites that are (i) located at secondary structures (i.e., α-helix or β-sheet), (ii) solvent exposed in both the open and closed states, and (iii) yield measurable differences in Cu(II)–Cu(II) distances and Cα–Cα distances between the two states. The predicted distances were calculated using both the MMM29,30,58 and chiLife methods.59,60In total, there are 15 site pairs that are potentially suitable for describing the conformational changes (Table S1 and Fig. S2). To further narrow down the number of site pairs, we explored the ability of each pair to differentiate between the closed and open states of LAOBP using MD starting from both the ligand-free and the lysine-bound states. Calculated distance distributions between the Cα atoms of the 15 site pairs discussed above are presented in the SI (Fig. S1). Out of the 15 site pairs, sites I56/K60 to sites D144/S148 show the largest peak separation (∼9 Å), with minimal overlap between the two states. Sites I56/K60 and D144/S148 are also close in sequence to prior DEER experiments using Gd(III)-based labels.61 The I56H/K60H/D144H/S148H construct was therefore chosen for the EPR studies.
CW-EPR and ESEEM to ascertain labeling
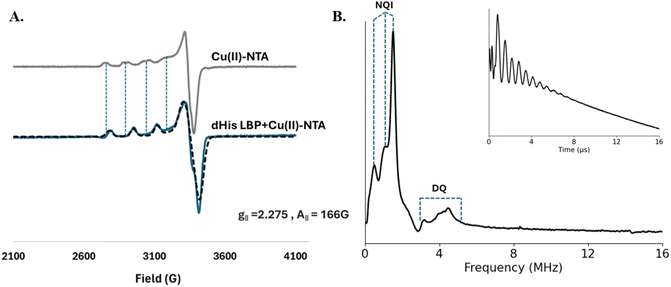
The double histidine (dHis) mutant (I56H/K60H/D144H/S148H) of LAOBP was expressed and labeled with Cu(II)-NTA as described in the methods section. Continuous-wave (CW) -EPR experiments were then performed to ensure proper labeling of the I56H/K60H/D144H/S148H mutant of LAOBP. Fig. 2A shows the CW-EPR spectra of free Cu(II)-NTA (grey) and dHis-protein bound Cu(II)-NTA (blue). There are clear spectral changes observed when the Cu(II)-label binds to the protein. These shifts in peak positioning are evident in the low field region and are indicated by vertical dashed lines. The spectrum of LAOBP-bound Cu(II)-NTA was simulated using only one component, and the simulated spectrum is overlaid in black dashed in Fig. 2A. The g‖ and A‖ values were 2.275 and 166 G, respectively, which is consistent with reported values for dHis-Cu(II) coordination.55,62,63 Additionally, no free Cu(II)-NTA component was observed in the protein data, which suggests complete labeling of the dHis sites on LAOBP.Next, to further confirm the binding of Cu(II)-NTA specifically to the sites on LAOBP, we performed pulsed electron spin echo envelope modulation (ESEEM) experiments. ESEEM64,65 experiments probe the interaction between the central Cu(II) ion and the non-coordinating nitrogen of the histidine residue. The inset to Fig. 2B shows the ESEEM signal obtained for dHis-LAOBP bound to Cu(II)-NTA. The signal has clear modulations characteristic of histidine coordination. The ESEEM spectrum, shown in Fig. 2B, contains three distinct nuclear quadrupolar interaction peaks below 2 MHz and a broad double quantum peak from 4–6 MHz.66–72 These peaks are indicative of Cu(II) coordinating to histidine sites. Taken together, CW-EPR and ESEEM suggest complete and specific coordination of Cu(II)-NTA to the target sites on LAOBP.
Distance measurements
To probe the different conformational states of LAOBP, we performed double electron–electron resonance (DEER) experiments. These experiments were performed at Q-Band (∼34 GHz) to obtain Cu(II)–Cu(II) distance constraints on LAOBP with and without the lysine ligand. Fig. 3 shows the background-corrected DEER data and corresponding distance distributions on LAOBP. The data was collected at two optimal field positions73,74 as indicated in the inset to Fig. 3, and then summed to ensure orientational effects are mitigated.75–77 The primary signals and spectra are provided in the SI (Fig. S3A and S3B). In the absence of the ligand, the time trace shows shallow modulations (top panel) and the presence of two frequency components. Analysis of this trace yields a bimodal distance distribution with a peak at 3.5 nm with a standard deviation of 0.26 nm and another at 4.6 nm with a standard deviation of 0.18 nm. This data is noteworthy, as prior crystallographic studies on LAOBP only resolved one conformation in the absence of the ligand.50 This conformation is consistent with the 4.6 nm distance. Here, solution EPR results resolve the complete conformational ensemble, and the results suggest that the protein samples two distinct conformations in the ligand-free state. The rigidity of the Cu(II) spin label is particularly crucial here, as the measurement clearly resolves the bimodal distribution within the open state of LAOBP. On the other hand, complete saturation of the protein with lysine (1.5 lysine: protein) yields a DEER signal (bottom trace) with three periods of well-resolved modulations, which suggests that the protein has a more limited range of conformational flexibility. Indeed, the distance distribution (c.f. right bottom panel) shows a single narrow distance of 3.3 nm with a standard deviation of 0.15 nm. Taken together, this data suggests that in the absence of lysine, the protein samples two dominant conformations, and the binding of the ligand leads to a selection of the more rigid and compact conformation.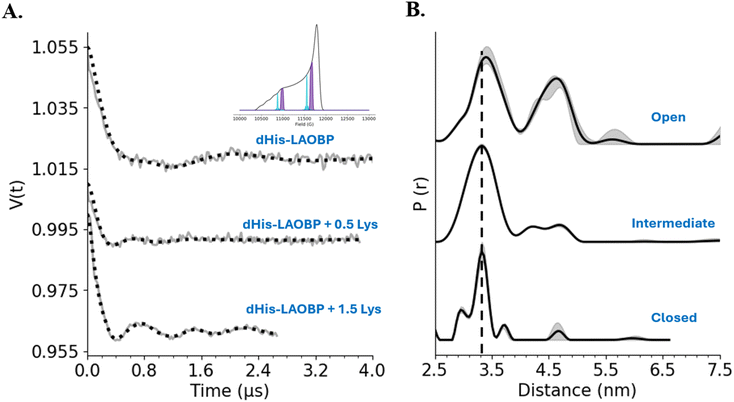 | ||
| Fig. 3 (A) Background-corrected DEER signal for LAOBP. The top panel shows the trace for the ligand-free state of the protein. The middle trace shows the time domain signal for an intermediate state with a sub-stoichiometric amount of ligand. The bottom time trace is for the ligand-bound state. The inset shows the Field Swept Electron Spin Echo (FS-ESE) spectrum, indicating the two positions where DEER measurement was performed. (B) Corresponding distance distributions obtained from DEERAnalysis.71 The distribution for the open state is broad and bimodal. Thus, the protein adopts a wide range of conformations in this state. The distribution for the closed state is narrow with a standard deviation of 0.15 nm. The grey shaded regions show the uncertainty in distance analysis due to the variation in the fitting of the background function in the DEER signal. | ||
We also acquired data at a sub-stoichiometric concentration of lysine (0.5 lysine: protein). This data is shown in the middle panel of Fig. 3. The modulations in the time domain signal are more pronounced than the ligand-free state but shallower than the ligand-bound state. As anticipated, the peak at 4.6 nm is significantly attenuated. Similarly, the peak at 3.3 nm is preserved but is broader than the closed state, with a standard deviation of 0.20 nm. This shift in the distance distribution suggests that the protein adopts a mixture of conformations, consistent with a partially bound intermediate state distribution. The DEER data were also analyzed using DEERNet78 and Comparative Deer Analyzer.79,80 The distributions obtained are shown in Fig. S4, and these distributions are consistent with the results shown in Fig. 3B. Data on wildtype LAOBP was also collected to ensure there were no contributions to these distance distributions from non-specific coordination of the Cu(II) label. The DEER signal for this sample, shown in Fig. S5, does not contain any modulations.
Weighted ensemble simulations
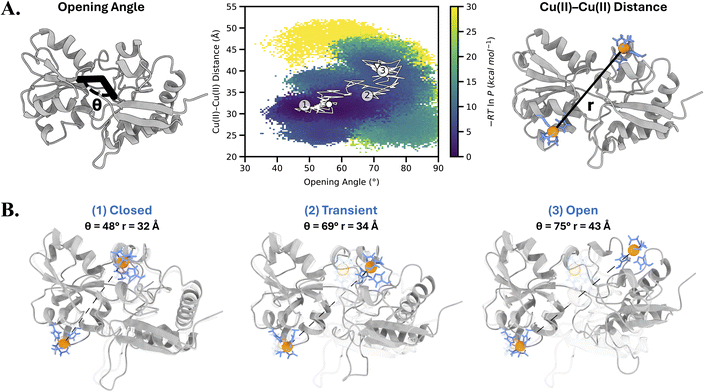
While the standard simulations failed to capture transitions, they provided initial definitions of the stable states. To focus the computing power on the transition between the ligand-bound and ligand-free states of LAOBP rather than stable states, we ran weighted ensemble (WE) simulations. We started the WE simulations from the equilibrated structure of the ligand-bound (holo) state with the ligand removed. WE simulations were carried out along a two-dimensional progress coordinate consisting of the Cu(II)–Cu(II) distance and the opening angle. Fig. 4A (left) shows the definition of the opening angle, θ. This angle provides a quantitative measure of how widely the lobes are oriented relative to one another, and more details of the definition are provided in the methods section. Fig. 4A (right) shows the position of the Cu(II)-NTA labels modeled on the protein for the simulation.
This scheme of using a two-dimensional progress coordinate is different from the prior work on a detoxification enzyme,47 where a one-dimensional progress coordinate based on the combined heavy atom root mean square deviation of two α9 helices (one from each monomer) after alignment on the rest of the protein dimer was sufficient, as the protein was a homo dimer. Since LAOBP is not a homodimer, a one-dimensional coordinate based solely on the Cu(II)–Cu(II) distance was insufficient to drive transitions between states in the WE simulation. We therefore added the tracking of the opening angle as a second dimension to the progress coordinate. The opening angle is a suitable choice for substrate-binding proteins, which typically undergo opening or twisting motions during conformational changes. In the case of LAOBP, the opening angle of the stable states differed significantly. The MD equilibrated structure of the ligand-bound state has an average Cu(II)–Cu(II) distance and opening angle of 33 Å and 50°, respectively. For the ligand-free state, these values are 43 Å and 85°, respectively. Using WE simulations, we successfully captured 381 continuous pathways from the closed to the open states of LAOBP (Fig. 4). The middle panel of Fig. 4A shows a two-dimensional probability distribution of all five WE simulations as a function of the opening angle and Cu(II)–Cu(II) distance. The distribution reveals the ensemble of conformational pathways sampled during the transition. The stable states are defined by high-probability basins, and the connecting probability density delineates several distinct pathways for the transition. A representative pathway for the transition between the two states is traced in white in Fig. 4, along with different conformational snapshots of LAOBP sampled along the representative pathway. Structures 1, 2, and 3 represent conformations characteristic of the closed state, transient intermediate, and open state, respectively. Although the transient (semi-closed) state has a Cu(II)–Cu(II) distance similar to that of the closed state, it is structurally distinct from the ligand-bound closed conformation, showing a markedly different opening angle. This transient state likely corresponds to the intermediate we previously detected by EPR using a sub-stoichiometric ratio of the lysine ligand. A movie of the continuous pathway from the WE simulation is shown in Movie S1. During the WE simulation, the Cu(II)–Cu(II) distance changes from 32 Å to 45 Å, which is consistent with the EPR results (Fig. 3). This result was confirmed by five independent WE simulations (Fig. S8). This concordance with experimental EPR results is particularly notable because the WE simulations were not restrained to match the EPR distances.
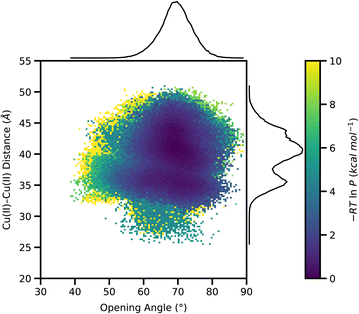
To access the stability of the open state sampled from the WE simulations, we selected five end structures from each WE simulation and initiated 25, 600-ns standard MD simulations from each structure (totaling to an aggregate simulation time of 15 µs). The probability distribution of all conformations sampled in the standard MD simulations, plotted as a function of Cu(II)–Cu(II) distance and opening angle, is shown in Fig. 5 and S9. From Fig. 5, we note that after 600 ns of simulation time, the simulations remained mostly in the ligand-free state originally sampled during the WE simulations. The open state displays a range of Cu(II)–Cu(II) distances and opening angles, as reflected in the distribution, suggesting intrinsic flexibility within the open-state ensemble. The marginal plots in Fig. 5, show the one-dimensional normalized probability distribution of each progress coordinate, where the Cu(II)–Cu(II) distance shows a bimodal distribution consistent with our EPR distance distribution data.
While WE simulations can be used to estimate rates between stable states, obtaining converged estimates would require substantially greater sampling of both the closed-to-open and open-to-closed transitions. We leave this calculation for future work, when experimentally measured values for these rates become available for validation. Beyond the LAOBP open and closed states and the transient intermediates connecting them, we also sampled structures that fall outside of the primary pathway shown in Fig. 4. These conformations exhibit either a large opening angle with a short Cu(II)–Cu(II) distance or conversely, a small opening angle with a long Cu(II)–Cu(II) distance (Fig. S11).
Collectively, our simulations show that the two stable states have a Cu(II)–Cu(II) distance of 32 Å and 45 Å. This agrees with our EPR data (Fig. 3), which also suggests that in the LAOBP ligand-free state, there are two different conformational states. This is not unexpected, as previous studies with substrate-binding proteins have shown that they can be in a semi-closed or closed state even in the absence of ligand.76,81–85 Together, our data suggest that EPR distances can be used as a reliable progress coordinate for driving the sampling of conformational transitions and simulating atomistically detailed pathways of large-scale protein conformational transitions.
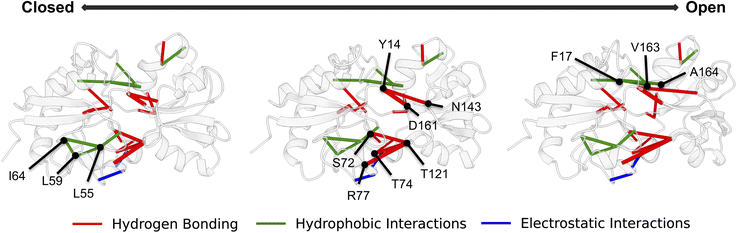
We identified residue T121 as a key anchor point for the main hydrogen-bond network, predominantly interacting with T74, R77, and S72. The LAOBP domain–domain interface is further stitched together by an H-bonding network between Y14, D161, and N143, along with hydrophobic interactions between F17, V163, and A164. We also identified Y14 as the anchor point for a secondary hydrogen-bonding network, predominately interacting with D161 and N143. The most dominant set of hydrophobic interactions was between residues, I64, L59, and L55, which underwent modest rearrangements near the EPR spin label. From both our EPR and simulation data, both open and closed conformations are accessible in the absence of ligand. This result suggests a conformational selection mechanism rather than an induced fit mechanism where the protein changes conformation upon ligand binding.
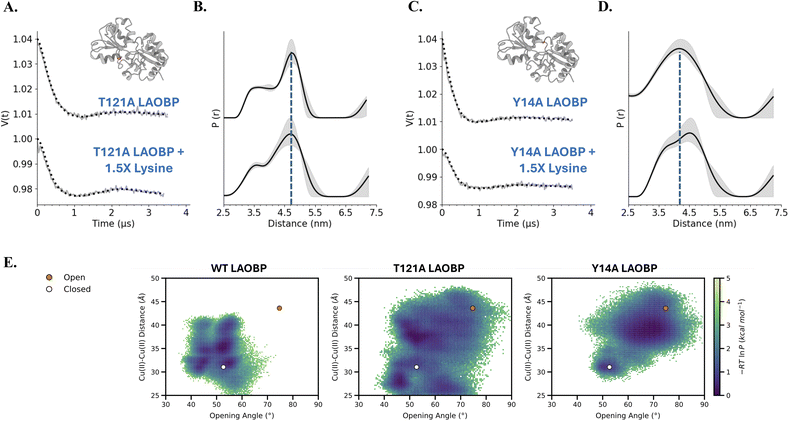
To test the effect of mutating residues critical to the interdomain hydrogen-bond network that guides the conformational transition, we expressed, purified, and spin-labeled two mutants of LAOBP: T121A and Y14A. Fig. 7A shows the background-corrected DEER signal for the T121A mutant in the absence and presence of ligand. The primary signals are shown in the SI (Fig. S12). Fig. 7B shows the corresponding distance distributions. Interestingly, even in the ligand-free state, the T121A mutation introduces some changes in the conformational ensemble. When compared to the WT data shown in Fig. 3A and B, the conformational ensemble has shifted towards a larger average distance value. Importantly, in the presence of 1.5 equivalents of lysine, there is no significant change in the time domain signal or the distance distribution. This data contrasts with the WT protein, where lysine binding led to a clear compaction of the conformational ensemble. This result demonstrates that substitution of threonine at the 121 position alone is sufficient to bias the conformational equilibrium toward the open state. Therefore, ligand-induced closure of the protein is prevented.
Time domain DEER data and distance distribution for the Y14A mutant are shown in Fig. 7C and D, respectively. For the Y14A mutant, we obtain a broad distribution ranging from 30–53 Å. As observed for T121A, the addition of lysine does not produce any observable shift in the distance distribution. Tyrosine 14 serves as a central hinge within the interdomain hydrogen-bond network that drives ligand-induced closure. Disruption of this residue destabilizes the closed state, effectively biasing the protein toward a broad open conformation.
To further assess the effects of the T121A and Y14A variants, we carried out conventional MD simulations of each variant starting from the closed, ligand-removed state (Fig. 7E). While the WT LAOBP remains in the initial closed state throughout the simulations, both the T121A and Y14A variants show a substantially greater preference towards the open state. In particular, the Y14A variant favors a Cu(II)–Cu(II) distance distribution centered around 40 Å, which is remarkably consistent with the EPR-measured distances. In contrast, the T121A variant samples a broader and more heterogeneous conformational distribution, also consistent with our EPR results. Together, these differences between the LAOBP T121A and Y14A variants likely arise from disruptions in the hydrogen-bonding networks that separate the two states, which we previously identified, thereby directly supporting the results of our interaction analysis.
Conclusions
In this work, we characterize the large-scale conformational transition from the closed to open states of the lysine/arginine/ornithine binding protein (LAOBP) by integrating EPR spectroscopy with weighted ensemble (WE) simulations. EPR provided sparse yet highly informative distance distributions that captured the dynamic nature of LAOBP, while WE simulations complemented these data with atomistic details of the transition process, which was inaccessible to standard MD due to the millisecond timescale of the exchange. Using WE, we sampled multiple continuous transition pathways between the open and closed states in the absence of ligand, providing direct structural insights into the structures underlying the experimentally observed distance distributions. Our analysis revealed that the transition is primarily mediated by an inter-domain hydrogen-bonding network centered around residue T121 and Y14, with lysine binding potentially stabilizing the closed state through a conformational selection mechanism, and substitution of T121 or Y14 residues to alanine shifts the conformational equilibrium toward the open state. Together, these findings highlight the power of combining sparse EPR distance restraints with enhanced-sampling simulations to resolve protein conformational exchange processes at atomic resolution. We anticipate that this integrative strategy will be broadly applicable to other proteins undergoing large-scale conformational changes, yielding mechanistic insights that are both experimentally grounded and atomically detailed.Experimental
Protein purification
The double histidine mutant (I56H/K60H/D144H/S148H) of LAOBP and WT LAOBP was expressed in BL21 (DE3) E. coli cells using a previously described procedure with the pET12a vector, which contains the sequence. The cell pellets were resuspended in 20 mM Tris-HCl buffer (pH 8.5) containing 5 mM NaCl and 0.12% Triton X-100, and then lysed through sonication. The lysate was centrifuged at 5000 rpm for 30 minutes at 4 °C, and the supernatant containing LAOBP was subjected to purification. Initial purification was carried out using a HiTrap Q HP anion exchange column, and the protein was eluted in the flow-through and checked using 12% SDS-PAGE. To dialyze and remove ligands (amino acids) that may have bound during expression and purification, the protein had to be washed before the size exclusion column. The protein was unfolded to remove bound ligands by treatment with 5 mM Tris buffer (pH 8.5) containing 2 M guanidinium chloride (GdnHCl), followed by centrifugation using a 10 kDa filter. This centrifugation step was repeated five times to ensure effective ligand removal. Refolding was then induced by repeated buffer exchange with denaturant-free buffer (sodium phosphate buffer at pH 6.5) and centrifugation (at least five cycles) to remove GdnHCl completely. The refolded protein was analysed through LC-MS, and the data is shown in Fig. S13. There is a single sharp peak with the mass corresponding to the apo protein. This procedure of unfolding and refolding the protein has been described in prior work with LAOBP.61,87,88 The refolded protein was subsequently purified further by passing it through a size-exclusion column. For the EPR experiments, 100 µM LAOBP was labeled with 200 µM Cu(II)-NTA in MOPS buffer at pH 7.4 using procedures outlined before.89 Mutants T121A and Y14A dHis LAOBP were expressed in BL21 (DE3) E. coli cells using the pET3a vector. The same purification protocol described above was followed, along with the refolding and folding procedure (5 times each) to dialyze any bound ligands. CW-EPR and ESEEM experiments were performed on the mutants to ensure proper labeling with Cu(II)-NTA and the data is provided in Fig. S14.CW EPR and ESEEM experiments
CW-EPR experiments were performed on a Bruker E580 X-band (∼9.4 GHz) FT/CW spectrometer. The temperature for all experiments was controlled using an Oxford ITC503 temperature controller with an Oxford ER 4118CF gas flow cryostat. CW-EPR spectra were recorded using a Bruker ER4118X-MD5 resonator at 80 K. The data were collected with a 4 G modulation amplitude, 100 kHz modulation frequency, 10.24 ns time constant, 20.48 ms conversion time, and 30 dB attenuation. Each spectrum was centered at 3100 G with a 2000 G sweep range over 1024 points. The resulting spectra were background-subtracted, area-normalized, and simulated using EasySpin.90The ESEEM experiments were performed at X-Band using the Bruker ER4118X-MD5 resonator at 18 K. A three-pulse sequence of [(π/2)–τ–(π/2)–T–(π/2)–echo] with a four-step phase cycling was employed. The length of the π/2 pulses were 10 ns. The separation, τ, was set to 132 ns, and an initial value of 264 ns was used for T. This value of T was incremented by 16 ns steps for a total of 1024 points. Measurements were taken at the magnetic field corresponding to the maximum intensity in the field-swept echo-detected spectrum of each sample. The acquired ESEEM traces were phased, background-corrected, zero-filled with an additional 1024 points, and fast Fourier transformed to obtain the frequency spectra. The magnitude spectrum is shown.
DEER experiments
Q-band distance measurements were carried out on a Bruker ElexSys FT/CW spectrometer equipped with a Bridge12 resonator, Bruker SpinJet AWG, and a 300 W amplifier. A four-pulse DEER pulse sequence was used: (π/2)ν1–τ1–(π)ν1–T–(π)ν2–τ2–(π)ν1–τ2 echo. Observer π/2 and π pulses of 8 ns and 16 ns were applied, while the π pump pulse was set to be a frequency-swept chirp of 100 ns (−300 to −100 MHz relative to the observer frequency). The measurement for the lysine-bound sample was collected for a total of 129 points with a 20 ns step size. For the ligand-free sample, the pump pulse was set to be a 200-ns chirp pulse, and the data was acquired over 187 points with a step size of 24 ns. The data was analyzed using DEERAnalysis91 and DEERNet.78Standard MD simulations
Heavy-atom coordinates for the open and closed conformations of LAOBP were extracted from crystal structures of the ligand-free protein (PDB code: 2LAO) and the lysine-bound conformation (PDB code: 1LST) of LAOBP.50 For the dHis LAOBP mutant (56H/60H/144H/148H), the initial structure was created through computational mutagenesis. The I56, K60, D144, and S148 sites were mutated to histidine. Appropriate rotamers of dHis were selected in PyMOL for incorporation of Cu(II)-NTA. The resulting dHis-Cu(II)-NTA complex was then aligned with the dHis sites of the proteins.All simulations of the EPR spin-labeled LAOBP protein were carried out using the AMBER ff19SB force field92 for the protein and the OPC water model.93 The protein was immersed in a periodic, cubic box of solvent with a minimum 12 Å distance between the protein and the box edge. Sodium and chloride ions were added to neutralize the system using Joung and Cheatham ion parameters.94 Simulations were performed with the GPU-accelerated PMEMD module of the AMBER21 software package.95–97 Energy minimization was first applied to the solvated systems with harmonic position restraints on the protein backbone, and gradually reduced from 20, 10, 5, and 1 to 0 kcal (mol−1 Å2) over 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps. The energy-minimized structures were then equilibrated at a constant temperature of 298 K and constant volume for 20 ps with harmonic position restraints on the heavy atoms with a force constant of 1 kcal (mol−1 Å2).98 Each system was subsequently equilibrated for 1 ns with the same heavy-atom restraints and then without restraints at constant temperature (298 K) and constant pressure of 1 atm. Temperature was controlled by a Langevin thermostat with a collision frequency of 1.0 ps−1, and pressure was maintained at 1 atm using a Monte Carlo barostat with pressure changes attempted every 0.2 ps. To enable a 2-fs time step, all bonds to hydrogens were constrained to their equilibrium values using the SHAKE algorithm.99 Short-range nonbonded interactions were truncated at 10 Å and long-range electrostatic interactions were treated with the particle mesh Ewald method.100 Five independent 200-ns MD simulations were performed on the spin-labeled 56H/60H/144H/148H mutant of LAOBP. To access the stability of WE-sampled conformations of the LAOBP open state, 25, 600-ns MD simulations were initiated from each conformation using the same settings as described above. For the T121A and Y14A variants of LAOBP, each system was modified using ChimeraX101 starting from the closed state, ligand-removed LAOBP structure (PDB code: 1LST) and equilibrated as described above. For each variant, five independent replicas were then carried out for 600-ns each.
000 steps. The energy-minimized structures were then equilibrated at a constant temperature of 298 K and constant volume for 20 ps with harmonic position restraints on the heavy atoms with a force constant of 1 kcal (mol−1 Å2).98 Each system was subsequently equilibrated for 1 ns with the same heavy-atom restraints and then without restraints at constant temperature (298 K) and constant pressure of 1 atm. Temperature was controlled by a Langevin thermostat with a collision frequency of 1.0 ps−1, and pressure was maintained at 1 atm using a Monte Carlo barostat with pressure changes attempted every 0.2 ps. To enable a 2-fs time step, all bonds to hydrogens were constrained to their equilibrium values using the SHAKE algorithm.99 Short-range nonbonded interactions were truncated at 10 Å and long-range electrostatic interactions were treated with the particle mesh Ewald method.100 Five independent 200-ns MD simulations were performed on the spin-labeled 56H/60H/144H/148H mutant of LAOBP. To access the stability of WE-sampled conformations of the LAOBP open state, 25, 600-ns MD simulations were initiated from each conformation using the same settings as described above. For the T121A and Y14A variants of LAOBP, each system was modified using ChimeraX101 starting from the closed state, ligand-removed LAOBP structure (PDB code: 1LST) and equilibrated as described above. For each variant, five independent replicas were then carried out for 600-ns each.
Weighted ensemble simulations
To enhance the conformational sampling of LAOBP, weighted ensemble (WE) simulations were run starting from the closed, ligand-removed state obtained after standard MD equilibration, where the conformational transition could be more readily induced by ligand removal. All WE simulations were carried out using the open-source WESTPA 2.0 software package.102 In WE sampling, a progress coordinate is partitioned into bins, and multiple short trajectories are run in parallel. At fixed time intervals τ, a resampling procedure is applied that replicates or prunes trajectories to maintain a target number of trajectories per bin. Throughout the simulation, trajectory weights are rigorously tracked to ensure that their sum remains normalized to one, ensuring no statistical bias occurs during trajectory resampling.WE simulations employed a two-dimensional progress coordinate defined by (i) the distance between the two Cu(II) ions and (ii) an opening angle that describes the relative motion of the two lobes surrounding the copper-binding site. The opening angle was calculated by first determining centers of mass (COMs) for three structural regions: the bottom lobe (residues 9–18, 29, 49–55, 66–73, 76, 82, 87–88, 193–195), the top lobe (residues 115–123, 140–143, 145, 158–163, 189–190), and the loop region connecting the lobes (residues 89 and 191). A plane was defined using the COM of the bottom lobe together with the Cα atoms of residues 89 and 191. The opening angle was then computed as the angle between the vector from the loop COM to the top lobe COM and the normal vector to the plane defined by the bottom lobe and the two loop residues. This angle provides a quantitative measure of how widely the lobes are splayed relative to one another.
Using the minimal adaptive binning (MAB) scheme,103 six adaptive bins were placed along each of the two dimensions of the progress coordinate. A resampling time interval τ of 50 ps was employed to maintain a target number of four trajectories per bin. Five independent WE replicates were performed, each for 200 WE iterations to generate a maximum trajectory length of 10 ns.
Author contributions
S. P., D. T. Y., X. B., A. T. B., S. S. and L. T. C. designed research; S. P., D. T. Y., X. B., A. T. B., and O. R. W. performed research; S. P., D. T. Y., X. B., A. T. B., O. R. W., S. S. and L. T. C. analyzed data; S. P., D. T. Y., X. B., A. T. B., O. R. W., S. S. and L. T. C. wrote the paper.Conflicts of interest
The authors declare the following competing financial interest(s): LTC serves on the scientific advisory board of OpenEye, Cadence Molecular Sciences.Data availability
All primary EPR data is provided in the supplementary information (SI). WE simulation and analysis scripts are available on GitHub (https://github.com/westpa/user_submitted_scripts). The WE simulation trajectories are available upon request due to large file size limitations. Supplementary information is available. See DOI: https://doi.org/10.1039/d5sc08971a.Acknowledgements
This research was supported by the National Science Foundation (NSF MCB-2112871). This research was supported in part by the University of Pittsburgh Center for Research Computing and Data, RRID:SCR_022735, through the resources provided. Specifically, this work used the H2P cluster, which is supported by NSF award number OAC-2117681.Notes and references
- C. Altenbach, A. K. Kusnetzow, O. P. Ernst, K. P. Hofmann and W. L. Hubbell, High-resolution distance mapping in rhodopsin reveals the pattern of helix movement due to activation, Proc. Natl. Acad. Sci., 2008, 105(21), 7439–7444, DOI:10.1073/pnas.0802515105.
- N. C. Horton and J. J. Perona, Crystallographic snapshots along a protein-induced DNA-bending pathway, Proc. Natl. Acad. Sci., 2000, 97(11), 5729–5734, DOI:10.1073/pnas.090370797.
- L. C. Cantley, C. T. Carilli, R. L. Smith and D. Perlman, Conformational Changes of Na, K-ATPase Necessary for Transport, In Current Topics in Membranes and Transport, ed. F. Bronner and A. Kleinzeller, Academic Press, 1983, pp. 315–322, DOI:10.1016/S0070-2161(08)60582-1, https://www.sciencedirect.com/science/chapter/bookseries/abs/pii/S0070216108605821.
- D. D. F. Loo, B. A. Hirayama, E. M. Gallardo, J. T. Lam, E. Turk and E. M. Wright, Conformational changes couple Na+ and glucose transport, Proc. Natl. Acad. Sci., 1998, 95(13), 7789–7794, DOI:10.1073/pnas.95.13.7789.
- A. K. Kusnetzow, C. Altenbach and W. L. Hubbell, Conformational States and Dynamics of Rhodopsin in Micelles and Bilayers, Biochemistry, 2006, 45(17), 5538–5550, DOI:10.1021/bi060101v.
- O. Delalande, S. Sacquin-Mora and M. Baaden, Enzyme Closure and Nucleotide Binding Structurally Lock Guanylate Kinase, Biophys. J., 2011, 101(6), 1440–1449, DOI:10.1016/j.bpj.2011.07.048.
- K. Arora and C. L. Brooks, Large-scale allosteric conformational transitions of adenylate kinase appear to involve a population-shift mechanism, Proc. Natl. Acad. Sci., 2007, 104(47), 18496–18501, DOI:10.1073/pnas.0706443104.
- D. L. Farrens, C. Altenbach, K. Yang, W. L. Hubbell and H. G. Khorana, Requirement of Rigid-Body Motion of Transmembrane Helices for Light Activation of Rhodopsin, Science, 1996, 274(5288), 768–770, DOI:10.1126/science.274.5288.768.
- H. S. Mchaourab, M. A. Lietzow, K. Hideg and W. L. Hubbell, Motion of Spin-Labeled Side Chains in T4 Lysozyme. Correlation with Protein Structure and Dynamics, Biochemistry, 1996, 35(24), 7692–7704, DOI:10.1021/bi960482k.
- W. L. Hubbell, H. S. Mchaourab, C. Altenbach and M. A. Lietzow, Watching proteins move using site-directed spin labeling, Structure, 1996, 4(7), 779–783, DOI:10.1016/S0969-2126(96)00085-8.
- P. Widder, J. Schuck, D. Summerer and M. Drescher, Combining site-directed spin labeling in vivo and in-cell EPR distance determination, Phys. Chem. Chem. Phys., 2020, 22(9), 4875–4879, 10.1039/C9CP05584C.
- D. Goldfarb, Gd3+ spin labeling for distance measurements by pulse EPR spectroscopy, Phys. Chem. Chem. Phys., 2014, 16(21), 9685–9699, 10.1039/C3CP53822B.
- J. J. Jassoy, A. Berndhäuser, F. Duthie, S. P. Kühn, G. Hagelueken and O. Schiemann, Versatile Trityl Spin Labels for Nanometer Distance Measurements on Biomolecules In Vitro and within Cells, Angew. Chem., Int. Ed., 2017, 56(1), 177–181, DOI:10.1002/anie.201609085.
- T. F. Cunningham, M. R. Putterman, A. Desai, W. S. Horne and S. Saxena, The Double-Histidine Cu2+-Binding Motif: A Highly Rigid, Site-Specific Spin Probe for Electron Spin Resonance Distance Measurements, Angew. Chem., Int. Ed., 2015, 54(21), 6330–6334, DOI:10.1002/anie.201501968.
- S. Palit, Z. Hasanbasri, N. A. Moriglioni, J. Casto and S. Saxena, Rational Design of a Cu(II) Spin Label Improves the Sensitivity of Distance Measurements, J. Phys. Chem. Lett., 2025, 16(39), 10256–10264, DOI:10.1021/acs.jpclett.5c02221.
- K. Barth, S. Hank, P. E. Spindler, T. F. Prisner, R. Tampé and B. Joseph, Conformational Coupling and trans-Inhibition in the Human Antigen Transporter Ortholog TmrAB Resolved with Dipolar EPR Spectroscopy, J. Am. Chem. Soc., 2018, 140(13), 4527–4533, DOI:10.1021/jacs.7b12409.
- O. Dalmas, P. Sompornpisut, F. Bezanilla and E. Perozo, Molecular mechanism of Mg2+-dependent gating in CorA, Nat. Commun., 2014, 5(1), 3590, DOI:10.1038/ncomms4590.
- E. G. B. Evans, J. L. W. Morgan, F. DiMaio, W. N. Zagotta and S. Stoll, Allosteric conformational change of a cyclic nucleotide-gated ion channel revealed by DEER spectroscopy, Proc. Natl. Acad. Sci., 2020, 117(20), 10839–10847, DOI:10.1073/pnas.1916375117.
- E. G. B. Evans, M. J. Pushie, K. A. Markham, H. W. Lee and G. L. Millhauser, Interaction between Prion Protein's Copper-Bound Octarepeat Domain and a Charged C-Terminal Pocket Suggests a Mechanism for N-Terminal Regulation, Structure, 1993, 24(7), 1057–1067, DOI:10.1016/j.str.2016.04.017.
- T. Schmidt, C. D. Schwieters and G. M. Clore, Spatial domain organization in the HIV-1 reverse transcriptase p66 homodimer precursor probed by double electron-electron resonance EPR, Proc. Natl. Acad. Sci., 2019, 116(36), 17809–17816, DOI:10.1073/pnas.1911086116.
- T. Hett, T. Zbik, S. Mukherjee, H. Matsuoka, W. Bönigk and D. Klose, et al., Spatiotemporal Resolution of Conformational Changes in Biomolecules by Combining Pulsed Electron–Electron Double Resonance Spectroscopy with Microsecond Freeze-Hyperquenching, J. Am. Chem. Soc., 2021, 143(18), 6981–6989, DOI:10.1021/jacs.1c01081.
- T. Schmidt, D. Wang, J. Jeon, C. D. Schwieters and G. M. Clore, Quantitative Agreement between Conformational Substates of Holo Calcium-Loaded Calmodulin Detected by Double Electron–Electron Resonance EPR and Predicted by Molecular Dynamics Simulations, J. Am. Chem. Soc., 2022, 144(27), 12043–12051, DOI:10.1021/jacs.2c02201.
- A. M. Stewart, M. Shanmugam, R. J. Kutta, N. S. Scrutton, J. E. Lovett and S. Hay, Combined Pulsed Electron Double Resonance EPR and Molecular Dynamics Investigations of Calmodulin Suggest Effects of Crowding Agents on Protein Structures, Biochemistry, 2022, 61(17), 1735–1742, DOI:10.1021/acs.biochem.2c00099.
- A. D. Milov, A. B. Ponomarev and T. YuD, Electron-electron double resonance in electron spin echo: Model biradical systems and the sensitized photolysis of decalin, Chem. Phys. Lett., 1984, 110(1), 67–72, DOI:10.1016/0009-2614(84)80148-7.
- P. P. Borbat and J. H. Freed, Multiple-quantum ESR and distance measurements, Chem. Phys. Lett., 1999, 313(1), 145–154, DOI:10.1016/S0009-2614(99)00972-0.
- G. Jeschke, DEER Distance Measurements on Proteins, Annu. Rev. Phys. Chem., 2012, 63(63), 419–446, DOI:10.1146/annurev-physchem-032511-143716.
- S. Milikisyants, F. Scarpelli, M. G. Finiguerra, M. Ubbink and M. Huber, A pulsed EPR method to determine distances between paramagnetic centers with strong spectral anisotropy and radicals: the dead-time free RIDME sequence, J. Magn. Reson., 1997, 201(1), 48–56, DOI:10.1016/j.jmr.2009.08.008.
- M. Pannier, S. Veit, A. Godt, G. Jeschke and H. W. Spiess, Dead-time free measurement of dipole-dipole interactions between electron spins, J. Magn. Reson., 1997, 142(2), 331–340, DOI:10.1006/jmre.1999.
- G. M. M. M. Jeschke, A toolbox for integrative structure modeling, Protein Sci., 2018, 27(1), 76–85, DOI:10.1002/pro.3269.
- G. Jeschke, MMM: Integrative ensemble modeling and ensemble analysis, Protein Sci., 2021, 30(1), 125–135, DOI:10.1002/pro.3965.
- Y. Qi, J. Lee, X. Cheng, R. Shen, S. M. Islam and B. Roux, CHARMM-GUI DEER facilitator for spin-pair distance distribution calculations and preparation of restrained-ensemble molecular dynamics simulations, J. Comput. Chem., 2020, 41, 415–420, DOI:10.1002/jcc.26032.
- J. L. MacCallum, A. Perez and K. A. Dill, Determining protein structures by combining semireliable data with atomistic physical models by Bayesian inference, Proc. Natl. Acad. Sci., 2015, 112(22), 6985–6990, DOI:10.1073/pnas.1506788112.
- A. Laio and M. Parrinello, Escaping free-energy minima, Proc. Natl. Acad. Sci., 2002, 99(20), 12562–12566, DOI:10.1073/pnas.202427399.
- F. Marinelli and J. D. Faraldo-Gómez, Ensemble-Biased Metadynamics: A Molecular Simulation Method to Sample Experimental Distributions, Biophys. J., 2015, 108(12), 2779–2782, DOI:10.1016/j.bpj.2015.05.024.
- E. J. Hustedt, F. Marinelli, R. A. Stein, J. D. Faraldo-Gómez and H. S. Mchaourab, Confidence Analysis of DEER Data and Its Structural Interpretation with Ensemble-Biased Metadynamics, Biophys. J., 2018, 115(7), 1200–1216, DOI:10.1016/j.bpj.2018.08.008.
- C. D. Schwieters, J. J. Kuszewski, N. Tjandra and G. Marius Clore, The Xplor-NIH NMR molecular structure determination package, J. Magn. Reson., 2003, 160(1), 65–73, DOI:10.1016/S1090-7807(02)00014-9.
- C. C. Jao, B. G. Hegde, J. Chen, I. S. Haworth and R. Langen, Structure of membrane-bound α-synuclein from site-directed spin labeling and computational refinement, Proc. Natl. Acad. Sci., 2008, 105(50), 19666–19671, DOI:10.1073/pnas.0807826105.
- S. J. Hirst, N. Alexander, H. S. Mchaourab and J. Meiler, RosettaEPR: An integrated tool for protein structure determination from sparse EPR data, J Struct Biol, 2011,(3), 506–514, DOI:10.1016/j.jsb.2010.10.013.
- T. Wu, R. A. Stein, T. Y. Kao, B. Brown and H. S. Mchaourab, Modeling protein conformational ensembles by guiding AlphaFold2 with Double Electron Electron Resonance (DEER) distance distributions, Nat. Commun., 2025, 16(1), 7107, DOI:10.1038/s41467-025-62582-4.
- X. Bogetti and S. Saxena, Integrating Electron Paramagnetic Resonance Spectroscopy and Computational Modeling to Measure Protein Structure and Dynamics, ChemPlusChem, 2024, 89(1), e202300506, DOI:10.1002/cplu.202300506.
- G. A. Huber and S. Kim, Weighted-ensemble Brownian dynamics simulations for protein association reactions, Biophys. J., 1996, 70(1), 97–110, DOI:10.1016/S0006-3495(96)79552-8.
- L. T. Chong, A. S. Saglam and D. M. Zuckerman, Path-sampling strategies for simulating rare events in biomolecular systems, Curr. Opin. Struct. Biol., 2017, 43, 88–94, DOI:10.1016/j.sbi.2016.11.019.
- D. T. Yang, L. T. Chong and A. M. Gronenborn, Illuminating an invisible state of the HIV-1 capsid protein CTD dimer using 19F NMR and weighted ensemble simulations, Proc. Natl. Acad. Sci., 2025, 122(8), e2420371122, DOI:10.1073/pnas.2420371122.
- X. Bogetti, A. Bogetti, J. Casto, G. Rule, L. Chong and S. Saxena, Direct observation of negative cooperativity in a detoxification enzyme at the atomic level by Electron Paramagnetic Resonance spectroscopy and simulation, Protein Sci., 2023, 32(10), e4770, DOI:10.1002/pro.4770.
- T. Sztain, S. H. Ahn, A. T. Bogetti, L. Casalino, J. A. Goldsmith and E. Seitz, et al., A glycan gate controls opening of the SARS-CoV-2 spike protein, Nat. Chem., 2021, 13(10), 963–968, DOI:10.1038/s41557-021-00758-3.
- A. S. Saglam and L. T. Chong, Protein–protein binding pathways and calculations of rate constants using fully-continuous, explicit-solvent simulations, Chem. Sci., 2019, 10(8), 2360–2372, 10.1039/C8SC04811H.
- X. Bogetti, A. Bogetti, J. Casto, G. Rule, L. Chong and S. Saxena, Direct observation of negative cooperativity in a detoxification enzyme at the atomic level by Electron Paramagnetic Resonance spectroscopy and simulation, Protein Sci., 2023, 32(10), e4770, DOI:10.1002/pro.4770.
- G. H. Scheepers, J. A. Lycklama a Nijeholt and B. Poolman, An updated structural classification of substrate-binding proteins, FEBS Lett., 2016, 590(23), 4393–4401, DOI:10.1002/1873-3468.12445.
- R. Tam and M. H. Saier, Structural, functional, and evolutionary relationships among extracellular solute-binding receptors of bacteria, Microbiol. Rev., 1993, 57(2), 320–346, DOI:10.1128/mr.57.2.320-346.1993.
- B. H. Oh, J. Pandit, C. H. Kang, K. Nikaido, S. Gokcen and G. F. Ames, et al., Three-dimensional structures of the periplasmic lysine/arginine/ornithine-binding protein with and without a ligand, J. Biol. Chem., 1993, 268(15), 11348–11355 CrossRef CAS PubMed.
- D. A. Silva, L. Domínguez-Ramírez, A. Rojo-Domínguez and A. Sosa-Peinado, Conformational dynamics of L-lysine, L-arginine, L-ornithine binding protein reveals ligand-dependent plasticity, Proteins: Struct., Funct., Bioinf., 2011, 79(7), 2097–2108, DOI:10.1002/prot.23030.
- H. X. Kondo, N. Okimoto, G. Morimoto and M. Taiji, Free-Energy Landscapes of Protein Domain Movements upon Ligand Binding, J. Phys. Chem. B, 2011, 115(23), 7629–7636, DOI:10.1021/jp111902t.
- N. Fleck, C. Heubach, T. Hett, S. Spicher, S. Grimme and O. Schiemann, Ox-SLIM: Synthesis of and Site-Specific Labelling with a Highly Hydrophilic Trityl Spin Label, Chem. – Eur. J., 2021, 27(16), 5292–5297, DOI:10.1002/chem.202100013.
- Z. Hasanbasri, K. Singewald, T. D. Gluth, B. Driesschaert and S. Saxena, Cleavage-resistant Protein Labeling with Hydrophilic Trityl Enables Distance Measurements In-Cell, J. Phys. Chem. B, 2021, 125(20), 5265–5274, DOI:10.1021/acs.jpcb.1c02371.
- S. Ghosh, M. J. Lawless, G. S. Rule and S. Saxena, The Cu2+-nitrilotriacetic acid complex improves loading of α-helical double histidine site for precise distance measurements by pulsed ESR, J. Magn. Reson., 2018, 286, 163–171, DOI:10.1016/j.jmr.2017.12.005.
- M. J. Lawless, S. Ghosh, T. F. Cunningham, A. Shimshi and S. Saxena, On the use of the Cu2+–iminodiacetic acid complex for double histidine based distance measurements by pulsed ESR, Phys. Chem. Chem. Phys., 2017, 19(31), 20959–20967, 10.1039/C7CP02564E.
- H. Sameach, S. Ghosh, L. Gevorkyan-Airapetov, S. Saxena and S. Ruthstein, EPR Spectroscopy Detects Various Active State Conformations of the Transcriptional Regulator CueR, Angew. Chem., Int. Ed., 2019, 58(10), 3053–3056, DOI:10.1002/anie.201810656.
- S. Ghosh, S. Saxena and G. Jeschke, Rotamer Modelling of Cu(II) Spin Labels Based on the Double-Histidine Motif, Appl. Magn. Reson., 2018, 49(11), 1281–1298, DOI:10.1007/s00723-018-1052-8.
- Z. Hasanbasri, M. H. Tessmer, S. Stoll and S. Saxena, Modeling of Cu(II)-based protein spin labels using rotamer libraries, Phys. Chem. Chem. Phys., 2024, 26(8), 6806–6816, 10.1039/D3CP05951K.
- M. H. Tessmer and S. Stoll, chiLife: An open-source Python package for in silico spin labeling and integrative protein modeling, PLoS Comput. Biol., 2023, 19(3), e1010834, DOI:10.1371/journal.pcbi.1010834.
- J. A. Kaczmarski, M. C. Mahawaththa, A. Feintuch, B. E. Clifton, L. A. Adams and D. Goldfarb, et al., Altered conformational sampling along an evolutionary trajectory changes the catalytic activity of an enzyme, Nat. Commun., 2020, 11(1), 5945, DOI:10.1038/s41467-020-19695-9.
- K. Singewald, J. A. Wilkinson, Z. Hasanbasri and S. Saxena, Beyond structure: Deciphering site-specific dynamics in proteins from double histidine-based EPR measurements, Protein Sci., 2022, 31(7), e4359, DOI:10.1002/pro.4359.
- A. Gamble Jarvi, J. Casto and S. Saxena, Buffer effects on site directed Cu2+-labeling using the double histidine motif, J. Magn. Reson., 1997, 320, 106848, DOI:10.1016/j.jmr.2020.106848.
- L. G. Rowan, E. L. Hahn and W. B. Mims, Electron-Spin-Echo Envelope Modulation, Phys. Rev., 1965, 137(1A), A61–A71, DOI:10.1103/PhysRev.137.A61.
- J. McCracken, P. R. Desai, N. J. Papadopoulos, J. J. Villafranca and J. Peisach, Electron spin-echo studies of the copper(II) binding sites in dopamine .beta.-hydroxylase, Biochemistry, 1988, 27(11), 4133–4137, DOI:10.1021/bi00411a034.
- J. McCracken, J. Peisach, C. E. Cote, M. A. McGuirl and D. M. Dooley, Pulsed EPR studies of the semiquinone state of copper-containing amine oxidases, J. Am. Chem. Soc., 1992, 114(10), 3715–3720, DOI:10.1021/ja00036a021.
- J. McCracken, S. Pember, S. J. Benkovic, J. J. Villafranca, R. J. Miller and J. Peisach, Electron spin-echo studies of the copper binding site in phenylalanine hydroxylase from Chromobacterium violaceum, J. Am. Chem. Soc., 1988, 110(4), 1069–1074, DOI:10.1021/ja00212a012.
- F. Jiang, J. McCracken and J. Peisach, Nuclear quadrupole interactions in copper(II)-diethylenetriamine-substituted imidazole complexes and in copper(II) proteins, J. Am. Chem. Soc., 1990, 112(25), 9035–9044, DOI:10.1021/ja00181a002.
- B. K. Shin and S. Saxena, Substantial Contribution of the Two Imidazole Rings of the His13−His14 Dyad to Cu(II) Binding in Amyloid-β(1−16) at Physiological pH and Its Significance, J. Phys. Chem. A, 2011, 115(34), 9590–9602, DOI:10.1021/jp200379m.
- B. K. Shin and S. Saxena, Direct Evidence That All Three Histidine Residues Coordinate to Cu(II) in Amyloid-β1−16, Biochemistry, 2008, 47(35), 9117–9123, DOI:10.1021/bi801014x.
- K. I. Silva, B. C. Michael, S. J. Geib and S. Saxena, ESEEM Analysis of Multi-Histidine Cu(II)-Coordination in Model Complexes, Peptides, and Amyloid-β, J. Phys. Chem. B, 2014, 118(30), 8935–8944, DOI:10.1021/jp500767n.
- J. Casto, S. Palit and S. Saxena, PELDOR to the Metal: Cu(II)-Based Labels Put a New Spin on Distance Measurements, Appl. Magn. Reson., 2024, 55, 889–922, DOI:10.1007/s00723-024-01658-8.
- Z. Hasanbasri and S. Saxena, Orientational Selectivity in Pulsed-EPR Does Not Have to be Complicated, Appl. Magn. Reson., 2024, 55(1), 61–78, DOI:10.1007/s00723-023-01594-z.
- Z. Hasanbasri, N. A. Moriglioni and S. Saxena, Efficient sampling of molecular orientations for Cu(II)-based DEER on protein labels, Phys. Chem. Chem. Phys., 2023, 25(19), 13275–13288, 10.1039/D3CP00404J.
- A. Gamble Jarvi, X. Bogetti, K. Singewald, S. Ghosh and S. Saxena, Going the dHis-tance: Site-Directed Cu2+ Labeling of Proteins and Nucleic Acids, Acc. Chem. Res., 2021, 54(6), 1481–1491, DOI:10.1021/acs.accounts.0c00761.
- X. Bogetti, S. Ghosh, A. Gamble Jarvi, J. Wang and S. Saxena, Molecular Dynamics Simulations Based on Newly Developed Force Field Parameters for Cu2+ Spin Labels Provide Insights into Double-Histidine-Based Double Electron–Electron Resonance, J. Phys. Chem. B, 2020, 124(14), 2788–2797, DOI:10.1021/acs.jpcb.0c00739.
- A. Gamble Jarvi, K. Ranguelova, S. Ghosh, R. T. Weber and S. Saxena, On the Use of Q-Band Double Electron–Electron Resonance To Resolve the Relative Orientations of Two Double Histidine-Bound Cu2+ Ions in a Protein, J. Phys. Chem. B, 2018, 122(47), 10669–10677, DOI:10.1021/acs.jpcb.8b07727.
- S. G. Worswick, J. A. Spencer, G. Jeschke and I. Kuprov, Deep neural network processing of DEER data, Sci. Adv., 2018, 4(8), eaat5218, DOI:10.1126/sciadv.aat5218.
- H. Russell, R. Cura and J. E. Lovett, DEER Data Analysis Software: A Comparative Guide, Front. Mol. Biosci., 2022, 9 DOI:10.3389/fmolb.2022.915167.
- G. Jeschke, ComparativeDeerAnalyzer 2.0, 2022 Search PubMed.
- C. Tang, C. D. Schwieters and G. M. Clore, Open-to-closed transition in apo maltose-binding protein observed by paramagnetic NMR, Nature, 2007, 449(7165), 1078–1082, DOI:10.1038/nature06232.
- M. H. Seo, J. Park, E. Kim, S. Hohng and H. S. Kim, Protein conformational dynamics dictate the binding affinity for a ligand, Nat. Commun., 2014, 5(1), 3724, DOI:10.1038/ncomms4724.
- B. Selmke, P. P. Borbat, C. Nickolaus, R. Varadarajan, J. H. Freed and W. E. Trommer, Open and Closed Form of Maltose Binding Protein in Its Native and Molten Globule State As Studied by Electron Paramagnetic Resonance Spectroscopy, Biochemistry, 2018, 57(38), 5507–5512, DOI:10.1021/acs.biochem.8b00322.
- B. C. H. Chu, D. I. Chan, T. DeWolf, X. Periole and H. J. Vogel, Molecular dynamics simulations reveal that apo-HisJ can sample a closed conformation, Proteins: Struct., Funct., Bioinf., 2014, 82(3), 386–398, DOI:10.1002/prot.24396.
- J. Glaenzer, M. F. Peter, G. H. Thomas and G. Hagelueken, PELDOR Spectroscopy Reveals Two Defined States of a Sialic Acid TRAP Transporter SBP in Solution, Biophys. J., 2017, 112(1), 109–120, DOI:10.1016/j.bpj.2016.12.010.
- R. M. Crean, J. S. G. Slusky, P. M. Kasson and S. C. L. Kamerlin, KIF—Key Interactions Finder: A program to identify the key molecular interactions that regulate protein conformational changes, J. Chem. Phys., 2023, 158(14), 144114, DOI:10.1063/5.0140882.
- J. Banda-Vázquez, S. Shanmugaratnam, R. Rodríguez-Sotres, A. Torres-Larios, B. Höcker and A. Sosa-Peinado, Redesign of LAOBP to bind novel l-amino acid ligands, Protein Sci., 2018, 27(5), 957–968, DOI:10.1002/pro.3403.
- R. Vergara, T. Berrocal, E. I. Juárez Mejía, S. Romero-Romero, I. Velázquez-López and N. O. Pulido, et al., Thermodynamic and kinetic analysis of the LAO binding protein and its isolated domains reveal non-additivity in stability, folding and function, FEBS J., 2023, 290(18), 4496–4512, DOI:10.1111/febs.16819.
- K. Singewald, J. A. Wilkinson and S. Saxena, Copper Based Site-directed Spin Labeling of Proteins for Use in Pulsed and Continuous Wave EPR Spectroscopy, Bio-protocol, 2021, 11(24), e4258, DOI:10.21769/BioProtoc.4258.
- S. Stoll and A. Schweiger, EasySpin, a comprehensive software package for spectral simulation and analysis in EPR, J. Magn. Reson., 2006, 178(1), 42–55, DOI:10.1016/j.jmr.2005.08.013.
- G. Jeschke, V. Chechik, P. Ionita, A. Godt, H. Zimmermann and J. E. Banham, et al., DeerAnalysis2006 –A Comprehensive Software Package for Analyzing Pulsed ELDOR Data, Appl. Magn. Reson., 2007, 30, 473–498, DOI:10.1007/BF03166213.
- C. Tian, K. Kasavajhala, K. A. A. Belfon, L. Raguette, H. Huang and A. N. Migues, et al., ff19SB: Amino-Acid-Specific Protein Backbone Parameters Trained against Quantum Mechanics Energy Surfaces in Solution, J. Chem. Theory Comput., 2020, 16(1), 528–552, DOI:10.1021/acs.jctc.9b00591.
- S. Izadi, R. Anandakrishnan and A. V. Onufriev, Building Water Models: A Different Approach, J. Phys. Chem. Lett., 2014, 5(21), 3863–3871, DOI:10.1021/jz501780a.
- I. S. Joung and T. E. I. Cheatham, Determination of Alkali and Halide Monovalent Ion Parameters for Use in Explicitly Solvated Biomolecular Simulations, J. Phys. Chem. B, 2008, 112(30), 9020–9041, DOI:10.1021/jp8001614.
- D. A. Case, T. E. Cheatham, T. Darden, H. Gohlke, R. Luo and K. M. Merz, et al., The Amber biomolecular simulation programs, J. Comput. Chem., 2005, 26(16), 1668–1688, DOI:10.1002/JCC.20290.
- A. W. Götz, M. J. Williamson, D. Xu, D. Poole, S. Le Grand and R. C. Walker, Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. generalized born, J. Chem. Theory Comput., 2012, 8(5), 1542–1555, DOI:10.1021/ct200909j.
- R. Salomon-Ferrer, A. W. Götz, D. Poole, S. Le Grand and R. C. Walker, Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. Explicit solvent particle mesh ewald, J. Chem. Theory Comput., 2013, 9(9), 3878–3888, DOI:10.1021/ct400314y.
- H. J. C. Berendsen, J. P. M. Postma, W. F. Van Gunsteren, A. DiNola and J. R. Haak, Molecular dynamics with coupling to an external bath, J. Chem. Phys., 1984, 81(8), 3684–3690, DOI:10.1063/1.448118.
- J. P. Ryckaert, G. Ciccotti and H. J. C. Berendsen, Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes, J. Comput. Phys., 1977, 23(3), 327–341, DOI:10.1016/0021-9991(77)90098-5.
- U. Essmann, L. Perera, M. L. Berkowitz, T. Darden, H. Lee and L. G. Pedersen, A smooth particle mesh Ewald method, J. Chem. Phys., 1995, 103, 234505, DOI:10.1063/1.470117.
- E. C. Meng, T. D. Goddard, E. F. Pettersen, G. S. Couch, Z. J. Pearson, J. H. Morris and T. E. Ferrin, UCSF ChimeraX: Tools for structure building and analysis, Protein Sci., 2023, 32(11), e4792 CrossRef CAS PubMed.
- J. D. Russo, S. Zhang, J. M. G. Leung, A. T. Bogetti, J. P. Thompson and A. J. DeGrave, et al., WESTPA 2.0: High-Performance Upgrades for Weighted Ensemble Simulations and Analysis of Longer-Timescale Applications, J. Chem. Theory Comput., 2022, 1c01154, DOI:10.1021/ACS.JCTC.1C01154.
- P. A. Torrillo, A. T. Bogetti, L. T. Chong and A. Minimal, Adaptive Binning Scheme for Weighted Ensemble Simulations, J. Phys. Chem. A, 2021, 125(7), 1642–1649, DOI:10.1021/acs.jpca.0c10724.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |