Open Access Article
Open Access ArticleFew-shot molecular property optimization via a domain-specialized large language model
Yan
Guo†
 a,
Menglan
Luo†
b,
Wenbo
Zhang
c,
Peidong
Liu
a,
Jin
Liu
b,
Shudong
Huang
a,
Jiancheng
Lv
a,
Bowen
Ke
*b and
Xianggen
Liu
*ab
a,
Menglan
Luo†
b,
Wenbo
Zhang
c,
Peidong
Liu
a,
Jin
Liu
b,
Shudong
Huang
a,
Jiancheng
Lv
a,
Bowen
Ke
*b and
Xianggen
Liu
*ab
aCollege of Computer Science, Sichuan University, Chengdu 610065, China
bDepartment of Anesthesiology, Laboratory of Anesthesia and Critical Care Medicine, National-Local Joint Engineering Research Centre of Translational Medicine of Anesthesiology, Frontiers Science Center for Disease-related Molecular Network, West China Hospital, Sichuan University, Chengdu 610041, China
cSchool of Computer Science and Technology, Xidian University, Xi'an 710126, China
First published on 7th February 2026
Abstract
Large language models (LLMs) have revolutionized machine learning with their few-shot learning and reasoning capabilities, demonstrating impressive results in fields like natural language processing and computer vision. However, when applied to the domains of biology and chemistry, current LLMs face substantial limitations, particularly in capturing the nuanced relationships between the molecular structure and pharmacochemical properties. This challenge has constrained the application of few-shot learning for small-molecule generation and optimization in drug discovery. Here, we introduce DrugLLM, a novel LLM tailored specifically for molecular optimization. DrugLLM leverages Functional Group Tokenization (FGT), which effectively tokenizes molecules for LLM learning, achieving over 53% token compression compared to SMILES. Besides, we propose a new pre-training strategy that enables DrugLLM to iteratively predict and modify molecular structures based on a few prior modifications, aligning each modification toward optimizing a specified pharmacological property. In multiple computational experiments, DrugLLM achieved state-of-the-art performance in few-shot molecular generation, surpassing all the mainstream LLMs including GPT-4. Furthermore, by applying DrugLLM to optimize HCN2 inhibitors, two bioactive compounds were obtained and successfully validated through wet-lab experiments. These results highlight the robust potential of DrugLLM in accelerating the optimization of molecules and AI-driven drug discovery.
Introduction
Large language models (LLMs) have fundamentally transformed the machine learning paradigm by bridging cross-task comprehension gaps with minimal samples.1 These models demonstrate remarkable capabilities in generalization, generation, and reasoning across domains ranging from natural language processing to software synthesis.2 However, when applied to chemical and molecular science, where semantic meaning is encoded not in syntax, but in three-dimensional conformations and electronic configurations, even state-of-the-art LLMs reveal their inherent limitations.3This discrepancy becomes particularly pronounced in molecular optimization, a critical phase in drug discovery requiring iterative refinement of candidate molecules to meet complex pharmacokinetic and pharmacodynamic criteria.4,5 While traditional computational approaches like density functional theory6 and molecular dynamics7 provide physically grounded insights, they suffer from prohibitive computational costs and frequently diverge from real-world development scenarios. On the other hand, data-driven methodologies, including deep generative models (DGM),8 offer potential for accelerated solutions but typically demand tens of thousands of labeled molecules, which remain scarce for most target properties. For instance, gradient-based optimization of DGM typically requires exceeding 104 training samples9–11 and traps in local minima of chemical space. Furthermore, cross-domain transfer learning struggles due to the fractured geometry of molecular representation spaces,12 exacerbated by data scarcity. Thus, the field currently lacks systems capable of emulating experienced medicinal chemists in inferring high-order structure–property relationships from limited data (typically ≤10 examples).13
This study posits that large language models (LLMs) hold the key to this conundrum through an underappreciated reasoning isomorphism. While SMILES/SELFIES linearizations have been treated as syntactic formalisms for validity-centric generation,14 we reveal their latent potential as cognitive manifolds. Recent theoretical advancements demonstrate that Transformer-based LLMs implement implicit meta-gradient descent through in-context learning.15 This mechanism closely parallels the human chemist's ability to internalize relationships between structural perturbations and property changes, treating them as composable reasoning primitives. By recasting optimization as a sequence-to-sequence task with tripartite prompts (problem definition, exemplars, and solution), this study presents DrugLLM, a framework that enables LLMs to disentangle domain-invariant optimization logic from target-specific methodologies.
In DrugLLM, we introduce a functional group tokenization (FGT) strategy that represents molecules as semantically meaningful subunits (e.g., [COOH] for carboxyl groups), achieving a 53.27% reduction in sequence length and improving representation consistency for structurally related compounds. Building upon this foundation, we further develop the next modification prediction (NMP) paradigm—a molecular-level learning framework inspired by the iterative reasoning of medicinal chemists. By formulating molecular optimization as a sequence of context-dependent structural modifications, NMP aligns large language model reasoning with fundamental biochemical principles, enabling property-guided molecular design without handcrafted domain priors.
We evaluated DrugLLM on 24 optimization tasks, encompassing properties such as the water–octanol partition coefficient (Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), solubility, synthetic accessibility, and topological polar surface area (TPSA), and 20 biological activities across individual targets. Extensive computational comparisons revealed that DrugLLM outperforms all existing generative algorithms in terms of few-shot property optimization success rates. Our model achieves optimization performance comparable to graph neural networks (GNNs) trained on 34
P), solubility, synthetic accessibility, and topological polar surface area (TPSA), and 20 biological activities across individual targets. Extensive computational comparisons revealed that DrugLLM outperforms all existing generative algorithms in terms of few-shot property optimization success rates. Our model achieves optimization performance comparable to graph neural networks (GNNs) trained on 34![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 labeled compounds,9 while requiring only 8 contextual examples and demonstrating a 4000-fold reduction in required training data. Notably, in the discovery of HCN channel inhibitors, DrugLLM identified two novel scaffolds (IC50 = 2.24 µM and 2.70 µM) that were undetectable via traditional virtual screening methods. These results demonstrate that DrugLLM democratizes rapid-response drug development confronting new properties and provides a blueprint for language model-driven innovation across drug discovery and materials science.
000 labeled compounds,9 while requiring only 8 contextual examples and demonstrating a 4000-fold reduction in required training data. Notably, in the discovery of HCN channel inhibitors, DrugLLM identified two novel scaffolds (IC50 = 2.24 µM and 2.70 µM) that were undetectable via traditional virtual screening methods. These results demonstrate that DrugLLM democratizes rapid-response drug development confronting new properties and provides a blueprint for language model-driven innovation across drug discovery and materials science.
Results
The DrugLLM framework
The focus of this work is to train a large language model that can capture the structure–activity relationship (SAR) of small molecules. SMILES16 is a well-known chemical language in the form of a line notation for describing the structure of molecules. Sometimes, two nearly identical molecules can possess markedly different canonical SMILES strings, thereby complicating the task of language modeling. To this end, this paper proposes LLM-oriented Functional Group Tokenization (FGT). As shown in Fig. 1, FGT employs unique string identifiers to represent distinct structural groups. These identifiers are interconnected through numerical position information on either side of a slash. FGT enables the model to discern molecular strings by considering structural groups as singular entities, thus diminishing the token count. Crucially, FGT reduces the representation variances in SMILES caused by minor structural modifications (Fig. S1). This is achieved because FGT decomposes the molecule into functional fragments and aligns them in a canonical core-to-periphery order, establishing a single, unambiguous representation that better reflects structural similarity. This streamlines the molecular assembly logic and eases the model's recognition task.The FGT construction process involves segmenting a molecule into its connected structural fragments. These fragments are then mapped to unique identifiers from a predefined dictionary to generate the FGT string. The group-based tokenization and interconnection recording makes FGT strings efficient in encoding the molecular structure. Statistics indicate that the average sequence length of FGT on the ZINC dataset is 17.86 ± 5.66, significantly shorter than the average length of SMILES (38.22 ± 7.16), resulting in a compression of 53.27% in the sequence length. The shorter encoding length and the sequential nature of the representation in FGT lay a good foundation for learning large language models. To ensure the practical utility of FGT, we conducted an extensive round-trip reconstruction analysis (SMILES → FGT → SMILES) across multiple datasets. The results, summarized in Table S6, demonstrate that FGT maintains exceptional structural fidelity, achieving a reconstruction success rate of 99.97% on 10 million molecules from the ZINC database and 98.41% on complex macrocyclic datasets from the Macrocycle-DB. This high fidelity ensures that DrugLLM operates on a stable and reversible chemical representation space.
FGT demonstrates significant advantages in vocabulary efficiency and molecular coverage over commonly used fragmentation schemes. With a vocabulary of only 4796 tokens, FGT achieves near-complete molecular (99.95%) coverage (Table S2). In contrast, RECAP17 requires over 500![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 tokens to cover 64.75% of molecules, while BRICS,18 despite high molecular coverage (98.34%), relies on a much larger vocabulary of 32
000 tokens to cover 64.75% of molecules, while BRICS,18 despite high molecular coverage (98.34%), relies on a much larger vocabulary of 32![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 874 tokens. This functional-group-centric decomposition preserves chemically meaningful substructures and hierarchical motifs, benefiting downstream machine learning tasks. While FGT does not explicitly enforce retrosynthetic constraints, its representational consistency and expressiveness highlight its advantages over reaction-based methods.
874 tokens. This functional-group-centric decomposition preserves chemically meaningful substructures and hierarchical motifs, benefiting downstream machine learning tasks. While FGT does not explicitly enforce retrosynthetic constraints, its representational consistency and expressiveness highlight its advantages over reaction-based methods.
Crucially, the design philosophy of FGT diverges from these retrosynthetic methods: while RECAP and BRICS prioritize synthetic feasibility, their extremely sparse vocabularies and low coverage create a “data sparsity” bottleneck that is detrimental to training generative LLMs. In contrast, FGT is a representation-centric scheme that treats functional groups as “reasoning primitives.” This approach allows DrugLLM to focus on the logical relationship between structural perturbations and property shifts, rather than long-range atom-level dependencies found in SMILES. FGT's compact vocabulary ensures that each token appears with sufficient frequency during pre-training, providing a stable and interpretable manifold for in-context SAR reasoning. This is not achievable by reaction-based or purely atom-based methods.
Furthermore, we evaluated the scalability of FGT for complex structural classes, such as macrocyclic molecules. Experiments on the Macformer dataset19 (5551 macrocycles) and the Macrocycle-DB20 (50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 653 macrocycles) showed that the vocabulary growth follows a sub-linear trend (see Fig. S5). Specifically, for over 50
653 macrocycles) showed that the vocabulary growth follows a sub-linear trend (see Fig. S5). Specifically, for over 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 macrocycles, the FGT vocabulary remained manageable at 5799 tokens. This efficiency stems from the structural redundancy in chemical space, where macrocycles often share recurring scaffolds.
000 macrocycles, the FGT vocabulary remained manageable at 5799 tokens. This efficiency stems from the structural redundancy in chemical space, where macrocycles often share recurring scaffolds.
Another challenge in training LLMs for molecule generation and optimization lies in the training paradigm. Current strategies for molecule generation include molecular graph-text translation,21 molecule encoding–decoding,22etc. Since the above objectives involve limited SAR data (such as molecular properties), generative models can hardly capture the fundamental relationship between the molecular structure and biological properties. In this work, we adopt the next modification prediction (NMP) paradigm as the overall learning framework of DrugLLM, where the model predicts the next molecular modification from one molecule to another. This paradigm is realized in training via an auto-regressive modification prediction (RMP) objective, where each modification is generated token by token based on FGT, attending to previous tokens and cross-context patterns (Fig. 1). Specifically, each molecular modification is represented as a sequence of tokens, and multiple modifications are organized into a sentence. Sentences focusing on the same molecular property form a paragraph, whose property is described in natural language at the beginning. For instance, if the first three sentences describe an increase in the number of hydrogen bond acceptors, all subsequent sentences in that paragraph also target the same property. In this way, paragraph contents are concentrated, allowing DrugLLM to generate each token auto-regressively based on previous contexts. Moreover, paragraphs are independent, encompassing diverse molecular properties, which enables DrugLLM to perform in-context learning (a form of few-shot learning) for molecular optimization.
However, there are few related datasets available for training DrugLLM. In this work, we collected the tabular form of the molecule datasets from the ZINC database23 and the ChEMBL platform,24,25 and converted them into the corresponding sentences and paragraphs of molecule modifications. In total, we collected over 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 modification paragraphs and 180
000 modification paragraphs and 180![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 molecules to build the training dataset (Table S1). The dataset involves over 10
000 molecules to build the training dataset (Table S1). The dataset involves over 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 different molecular properties or activities, such as the count of hydrogen bond acceptors and affinity to the GABAA receptor. The dataset was then split into the training and validation sets with a ratio of 9
000 different molecular properties or activities, such as the count of hydrogen bond acceptors and affinity to the GABAA receptor. The dataset was then split into the training and validation sets with a ratio of 9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. Considering that the few-shot learning capability of machine learning models arises from their exposure to a sufficient variety of training tasks, a large number of diverse paragraphs helps DrugLLM capture the intrinsic nature of molecule design in a few-shot fashion.
1. Considering that the few-shot learning capability of machine learning models arises from their exposure to a sufficient variety of training tasks, a large number of diverse paragraphs helps DrugLLM capture the intrinsic nature of molecule design in a few-shot fashion.
DrugLLM is based on the Transformer architecture.26 The vocabulary was built by generating functional group tokens via FGT, followed by byte pair encoding (BPE)27 to create a compact and efficient token set for training. DrugLLM is a large-scale model trained from scratch to generate molecular modification paragraphs. From a machine learning perspective, each paragraph provides contextual information describing the few-shot molecular optimization process. After large-scale training, DrugLLM is able to perform few-shot molecular optimization without further fine-tuning.
Few-shot molecular optimization toward physicochemical properties
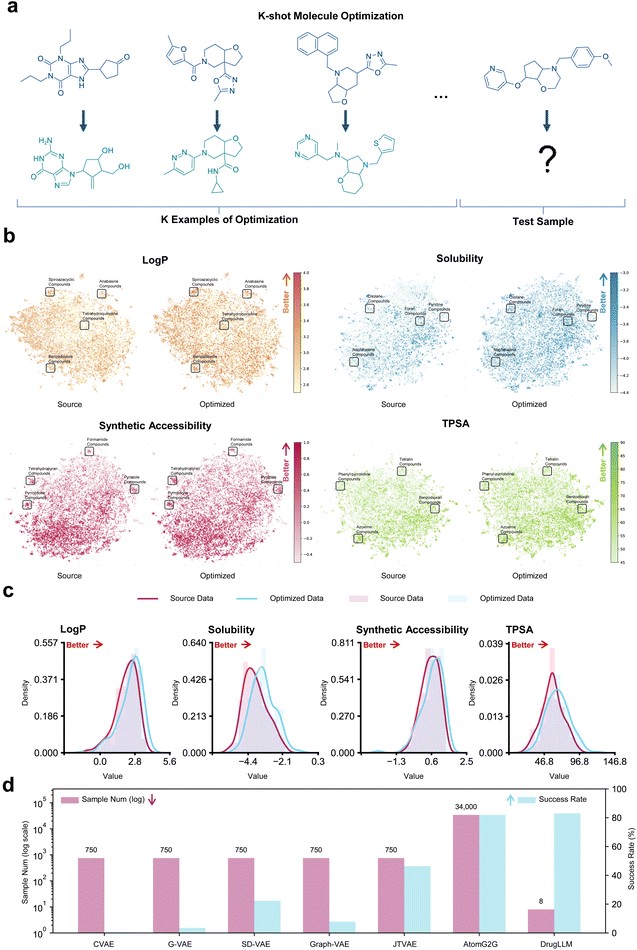
To evaluate the capacity of DrugLLM in terms of few-shot molecular optimization, we adopted several molecular properties that were not used in training based on the following assessment criteria (Fig. 2a). Given K pairs of example modifications and a molecule to be optimized, the model was asked to generate a new molecule that not only maintains structural similarity to the given molecule but also exhibits superior properties (either increased or decreased, as guided by the preceding examples). We selected four physicochemical properties that were not included in the training and validation sets as the test tasks, including the water–octanol partition coefficient (Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), solubility, synthetic accessibility, and topological polar surface area (TPSA), with 15
P), solubility, synthetic accessibility, and topological polar surface area (TPSA), with 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 testing samples for each property.
000 testing samples for each property.
We adopted Uniform Manifold Approximation and Projection (UMAP) to visualize the molecular space and qualitatively evaluate the optimization ability of DrugLLM (Fig. 2b). Specifically, we set the property in the context to increase instead of both increase and decrease for ease of evaluation. Then we validated the property changes in the molecules generated by DrugLLM. We observed that there was a high degree of consistency between the distributions of the optimized molecules (right) and the source molecules (left), indicating the diversity of the model generation. Despite similar distributions, the property values of the generated molecules were consistently higher than the original ones (reflected by the darker color map). Also, the distribution shift toward property improvement via Kernel Density Estimation (KDE) further underscores the powerful optimization capability of DrugLLM (Fig. 2c). Compared with Graph-VAE, JTVAE, and AtomG2G (Fig. 2d), we observed that these supervised methods require up to a 4000 times larger data scale to achieve a similar optimization performance to DrugLLM.
To quantitatively analyze the optimization capacity of DrugLLM, we compared it with several previous state-of-the-art molecule generators, including the junction tree-based variational auto-encoder (JTVAE),9 the variational junction tree neural network (VJTNN),28 and the scaffold-based molecule generator (MoLeR).29 Unlike these baselines, which were designed primarily as general-purpose molecular generators for de novo molecule creation, our focus is on few-shot molecular optimization, aiming to improve specific molecular properties while preserving core scaffolds. For fair comparison, we used the officially released pre-trained models and adapted them to perform few-shot optimization tasks. We also included a random optimization control implemented by random sampling based on the latent space of JTVAE (see the Implementations of the competitive baselines section in theSI Notes). The quality of the generated molecules was assessed based on their success rate and molecular similarity. The success rate represents the proportion of generated molecules that adhere to the property tendency of modifications (i.e., increment or decrement). To avoid the context bias of the generators, the input contexts described the increment or decrement of the property with a balanced proportion.
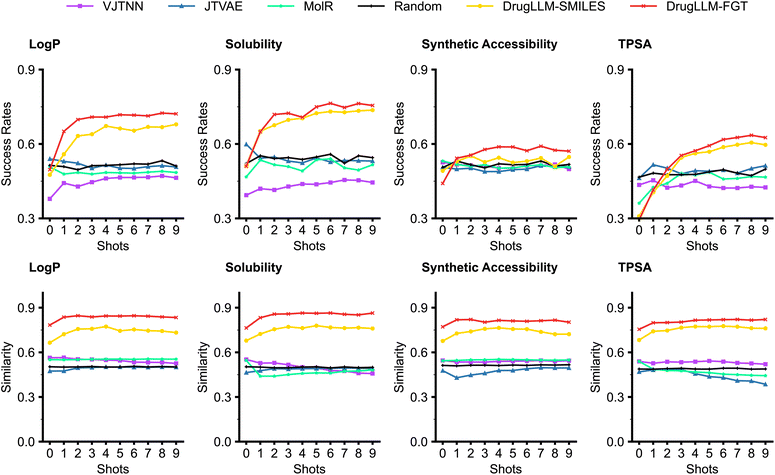
As shown in Fig. 3, we first report the performance of few-shot optimization with respect to the Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P value. We noted that the three baseline molecule generators, namely JTVAE, VJTNN, and MoLeR, obtained a success rate of about 0.50, which is similar to a random generation. In contrast, DrugLLM exhibits a progressive improvement in few-shot molecular optimization, with the success rate of the generated molecules increasing incrementally to 0.72 as the number of shots increases. Performance comparisons on molecular solubility, synthetic accessibility, and TPSA are similar and consistent. When it comes to similarity, it is typically more challenging to optimize a molecule with fewer modifications (i.e., higher similarity). Despite this, DrugLLM maintains a higher success rate even with increased generation similarity, underscoring its superior performance in the few-shot optimization. Furthermore, we noticed that FGT-based DrugLLM (denoted as DrugLLM-FGT) also significantly outperforms the DrugLLM that utilizes SMILES encodings (i.e., DrugLLM-SMILES), highlighting the benefits of FGT in the training of LLMs. Additional analyses of DrugLLM's generative quality across molecular optimization tasks can be found in the SI Notes (see the Evaluation of generative quality in molecular optimization section).
P value. We noted that the three baseline molecule generators, namely JTVAE, VJTNN, and MoLeR, obtained a success rate of about 0.50, which is similar to a random generation. In contrast, DrugLLM exhibits a progressive improvement in few-shot molecular optimization, with the success rate of the generated molecules increasing incrementally to 0.72 as the number of shots increases. Performance comparisons on molecular solubility, synthetic accessibility, and TPSA are similar and consistent. When it comes to similarity, it is typically more challenging to optimize a molecule with fewer modifications (i.e., higher similarity). Despite this, DrugLLM maintains a higher success rate even with increased generation similarity, underscoring its superior performance in the few-shot optimization. Furthermore, we noticed that FGT-based DrugLLM (denoted as DrugLLM-FGT) also significantly outperforms the DrugLLM that utilizes SMILES encodings (i.e., DrugLLM-SMILES), highlighting the benefits of FGT in the training of LLMs. Additional analyses of DrugLLM's generative quality across molecular optimization tasks can be found in the SI Notes (see the Evaluation of generative quality in molecular optimization section).
Few-shot molecular optimization towards biological activities
Since DrugLLM shows impressive few-shot optimization capacity in physicochemical properties, we next validated its effectiveness in the biological activities of molecules, which is more challenging due to the complex mechanisms in living systems. The molecules produced by DrugLLM are usually novel and not recorded in the ChEMBL database. Unlike the physicochemical properties mentioned above, biological activities (e.g., the Ki value on streptokinase A) are difficult to estimate using chemical or physical rules, and the time and cost of wet-lab experiments hinder large-scale evaluation. To address this, we employed learnable machine learning-based property predictors to approximate biological activities. Specifically, we utilized ChemProp, which is a message-passing neural network model that represents molecules as graphs.30 The specific training procedures, dataset partitioning, and validation protocols for these biological activity predictors are detailed in theSI Notes (see the Implementations of the biological activity predictors section). These predictors achieved strong correlations with experimental measurements (Pearson correlation ≥ 0.75, see Table S5), enabling reliable evaluation of generated molecules. For each property, the optimization direction (increase or decrease) can be explicitly specified through natural language instructions. The success rate measures the proportion of generated molecules that exhibit a property change aligning with this specified direction, relative to the reference molecule. The evaluation test set was balanced, equally split between tasks targeting property increases and decreases. Importantly, the twenty target biological activities used for testing were not included in the DrugLLM training set, allowing an unbiased assessment of DrugLLM's performance on previously unseen biological targets.When testing on these more difficult properties, the three generator baselines failed to obtain meaningful improvement (Table 1). All the baselines performed similarly to the random generator, indicating that these molecule generators still struggle to capture the modification rules underlying the limited examples. As for DrugLLM, it significantly outperforms the other baselines by a large margin in most of the test properties. In particular, for the bioassay target CHEMBL1963814 (Ki property), DrugLLM achieved a success rate of 0.76. Note that these test properties are not observable in the training of DrugLLM. Although the success rates obtained by DrugLLM are still not high enough, these attempts are the first steps in optimizing the biological activities in a few-shot manner. These results demonstrate that DrugLLM is able to figure out the intrinsic rules of molecule modifications given a limited number of examples of an unknown molecular property.
| Bioassay target | Property | Random | JTVAE | VJTNN | MoLeR | DrugLLM |
|---|---|---|---|---|---|---|
| CHEMBL1794496 | AC50 | 0.35 | 0.59 | 0.51 | 0.47 | 0.70 |
| CHEMBL2354301 | AC50 | 0.50 | 0.51 | 0.46 | 0.49 | 0.67 |
| CHEMBL1613983 | EC50 | 0.48 | 0.36 | 0.45 | 0.30 | 0.66 |
| CHEMBL1738500 | EC50 | 0.44 | 0.58 | 0.50 | 0.24 | 0.72 |
| CHEMBL1614183 | IC50 | 0.24 | 0.33 | 0.21 | 0.29 | 0.71 |
| CHEMBL1963888 | IC50 | 0.35 | 0.21 | 0.32 | 0.39 | 0.68 |
| CHEMBL4296185 | Inhibition | 0.55 | 0.47 | 0.49 | 0.51 | 0.67 |
| CHEMBL4296190 | Inhibition | 0.58 | 0.56 | 0.52 | 0.48 | 0.74 |
| CHEMBL1613886 | Potency | 0.35 | 0.40 | 0.44 | 0.42 | 0.61 |
| CHEMBL1614481 | Potency | 0.43 | 0.36 | 0.33 | 0.41 | 0.64 |
| CHEMBL1963722 | K i | 0.55 | 0.57 | 0.41 | 0.54 | 0.72 |
| CHEMBL1963723 | K i | 0.51 | 0.50 | 0.53 | 0.50 | 0.63 |
| CHEMBL1963727 | K i | 0.48 | 0.53 | 0.44 | 0.54 | 0.60 |
| CHEMBL1963788 | K i | 0.43 | 0.44 | 0.42 | 0.48 | 0.59 |
| CHEMBL1963790 | K i | 0.49 | 0.54 | 0.57 | 0.57 | 0.65 |
| CHEMBL1963807 | K i | 0.53 | 0.60 | 0.55 | 0.59 | 0.74 |
| CHEMBL1963814 | K i | 0.54 | 0.59 | 0.48 | 0.52 | 0.76 |
| CHEMBL1963835 | K i | 0.58 | 0.55 | 0.42 | 0.51 | 0.69 |
| CHEMBL1964107 | K i | 0.53 | 0.54 | 0.38 | 0.52 | 0.66 |
| CHEMBL1964119 | K i | 0.47 | 0.58 | 0.39 | 0.55 | 0.76 |
Zero-shot molecular optimization
Previous experiments demonstrated that DrugLLM can efficiently learn molecular modification rules and generate new molecules with the desired properties. In this section, we explore zero-shot molecular optimization, which involves generating molecules with improved properties according to natural language instructions, without observing specific training instances for those combinations. In this experimental setting, we assume that when DrugLLM is trained on a large collection of individual properties and their compositions, it can generalize to optimize molecules toward unseen combinations of properties, following the compositional zero-shot learning paradigm.31,32For example, the optimization of each individual property (e.g., Quantitative Estimation of Drug-likeness (QED) and FractionCSP3) is included in the training set, whereas the joint optimization of QED and FractionCSP3 does not appear in the training data and is used for zero-shot evaluation. Accordingly, we adopted six such optimization tasks that are absent from the DrugLLM training set as test tasks. Based on this setting, we constructed a test set containing over 6000 instructions, with 1000 instructions per optimization task. Generated molecules are evaluated using Python scripts based on the RDKit library. For these composite tasks, we define the optimization success rate as the percentage of generated molecules where both properties are simultaneously optimized according to the directions specified in the natural language instruction.
Zero-shot molecular optimization presents a significant challenge for language models, which is twofold. On the one hand, learning the mapping between semantics (instructions) and molecular properties from a general corpus is inherently difficult. On the other hand, biological data correlating structures with properties are often insufficient due to the lengthy time and high costs associated with wet-lab experiments.
To evaluate DrugLLM's zero-shot capabilities under these challenges, we compared it against several state-of-the-art general-purpose and domain-specific LLMs. The baseline models include general-purpose LLMs (ChatGPT3.5,1 GPT-4,33 and ChatGLM34) as well as two domain-specific biomedical language models, Meditron35 and BioMedLM,36 which are pretrained on large-scale biomedical corpora. Other general-purpose LLMs such as LLaMA37 were excluded as they were unable to generate valid SMILES strings.
As a result, we observed that ChatGLM struggled to provide appropriate molecules on all the zero-shot molecular optimization tasks (Table 2), with most generations outputting duplicated molecules identical to the input ones. In addition, ChatGPT3.5, GPT-4, Meditron, and BioMedLM were able to understand the instructions and optimize some of the given molecules, but the success rates remain relatively low. In contrast, DrugLLM improves the optimization success rates by significant margins compared with the other LLMs, indicating a superior capacity for instruction understanding and molecular optimization.
| Method | ChatGLM | ChatGPT3.5 | GPT-4 | Meditron | BioMedLM | DrugLLM |
|---|---|---|---|---|---|---|
| QED & FractionCSP3 | 0.02 | 0.11 | 0.20 | 0.33 | 0.15 | 0.40 |
| QED & # H-bond acceptors | 0.03 | 0.15 | 0.20 | 0.18 | 0.07 | 0.47 |
| QED & # rotatable bonds | 0.06 | 0.15 | 0.10 | 0.37 | 0.16 | 0.59 |
| # H-bond donors & FractionCSP3 | 0.03 | 0.30 | 0.40 | 0.39 | 0.14 | 0.60 |
| # H-bond donors & # H-bond acceptors | 0.04 | 0.19 | 0.43 | 0.23 | 0.07 | 0.55 |
| # H-bond donors & # rotatable bonds | 0.04 | 0.19 | 0.05 | 0.43 | 0.13 | 0.61 |
Applications of DrugLLM to the discovery of novel HCN2 inhibitors
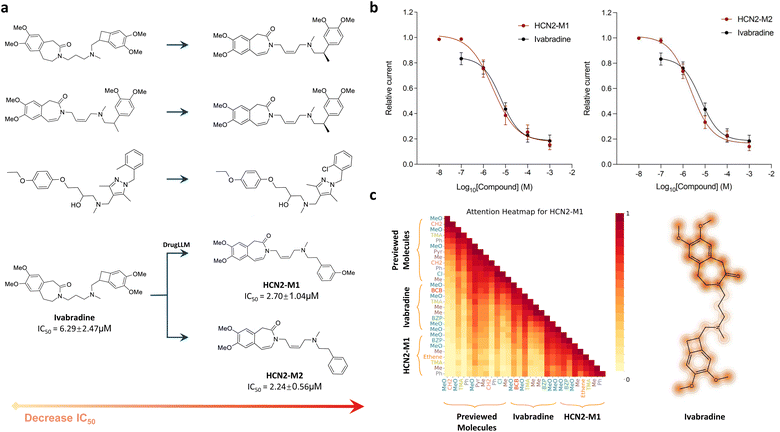
To verify the potential utility of DrugLLM in drug discovery, we applied DrugLLM to generate new molecules targeting hyperpolarization-activated cyclic nucleotide-gated channel 2 (HCN2). HCN2 is one of the four family members of HCN channels (HCN1-4) and is activated by hyperpolarizing potentials.38,39 Currently, the HCN2 subtype has been identified as a promising therapeutic target for treating chronic pain.40 However, research on the development of HCN2 inhibitors remains limited. Ivabradine, a broad-spectrum HCN inhibitor approved for the treatment of angina, offers a potential lead structure for the development of more effective small-molecule analgesics with a more favorable safety profile.41Here, we demonstrated the ability of DrugLLM to optimize ivabradine to have a better biological activity targeting HCN2. First, we formulated the optimization of ivabradine as a few-shot molecular optimization process, where DrugLLM learned from historical optimization examples and modifies the given molecule. We collected bioactivity data of validated HCN2 inhibitors as supporting examples from the recent studies.42,43 Then, these data were arranged into three pairs of molecules where the modification of each pair describes the decrease in the IC50 values to HCN2 (Fig. 4a). We fed these modifications together with ivabradine into DrugLLM and DrugLLM generated a series of new molecules (Table S7) that are expected to possess a lower IC50 to HCN2 than ivabradine.
Since DrugLLM performs molecular optimization rather than de novo generation, these generated molecules naturally retain high structural similarity to the input compound while exploring diverse substituent modifications. All molecules were generated directly by DrugLLM without manual editing. The model then automatically ranked these candidates based on their generation likelihood (sequence probability).
For wet-lab validation, two molecules (HCN2-M1 and HCN2-M2) were selected by chemistry experts from the top-ranked candidates. This selection was guided primarily by synthetic accessibility and practical considerations, ensuring that the chosen molecules could be feasibly synthesized. This approach ensured that the candidate generation and ranking are model-driven, while human expertise is applied to the final selection for practical validation.
For comparison, we independently applied these two molecules and ivabradine to perform patch-clamp experiments on the human HCN2 isoform heterologously expressed in HEK293 cells. Detailed protocols for these experiments are available in the SI Notes (Patch clamp experiments section). As expected, the in vitro testing results revealed that HCN2-M1 and HCN2-M2 showed lower IC50 values than ivabradine, both outperforming ivabradine with IC50 values approximately three times lower (Fig. 4b). Furthermore, we investigated the attention activations of DrugLLM when generating HCN2-M1 and HCN2-M2 as shown in Fig. 4c. We observed that DrugLLM had high attention on the methoxy group and the benzazepin moiety of ivabradine, which are indeed regarded as key groups for HCN2 interactions.44 Overall, these results suggest that the generated molecules possess stronger inhibitory potency against HCN2 and validate the effectiveness of DrugLLM in realistic drug discovery.
Discussion
In this study, we introduce a computational task named few-shot molecular optimization. Given a molecule of interest, the task involves generating new molecules that adhere to the optimization rules underlying a few modification examples. Although various few-shot learning tasks have been proposed and investigated in computational biology,45–47 few studies consider this specific optimization challenge. The difficulty of few-shot molecular optimization lies in requiring the model to capture abstract rules from a small set of examples and apply them to new molecules. This implicitly demands the ability to model complex “structure–effect–metabolism–toxicity” relationships. To address these challenges, we developed DrugLLM, a model designed specifically for few-shot molecular optimization.DrugLLM is a large language model (LLM) built on a large-scale textual corpus spanning a wide variety of small molecules and biological domains. Recently, general-purpose LLMs such as ChatGPT3.5,1 Alpaca,48 and ChatGLM34 have shown remarkable capabilities in natural language generation. However, they are designed for general use and lack the specialized knowledge required for pharmaceutical science. While several LLMs for the biomedical field exist, such as BioGPT49 and DrugGPT,50 they primarily focus on generating natural language (i.e., biomedical text), leaving open the question of how LLMs can understand the underlying language of chemistry and biology, particularly in a few-shot setting.
This study presents the first attempt to build an LLM specifically for few-shot molecule generation and optimization. Based on tabular data related to molecular properties and biological activities, we built a large-scale textual corpus formatted as sequences of molecule modifications. DrugLLM is trained to predict the next molecule based on historical modifications in an autoregressive manner. In extensive computational experiments, we observed that DrugLLM surpassed all competitive methods (including GPT-4) in optimizing new molecules in the few-shot setting across over 24 properties and biological activities. These results demonstrate the substantial enhancement in efficacy achieved by our methodology, highlighting the potential of DrugLLM as a powerful computational tool in drug discovery.
An important characteristic observed in DrugLLM's outputs is the tendency to preserve the core molecular scaffold of the input molecule during optimization, even though no explicit structural constraints are enforced. We attribute this emergent behavior primarily to two factors. First, the FGT representation, which encodes molecules starting from the core (often the dominant ring system) outwards, inherently prioritizes the central structure in the sequence representation. Second, the training data predominantly consist of modification trajectories where structural changes between consecutive molecules are typically small and located at the periphery. The combination of this core-first representation and the nature of the training examples implicitly guides the model to learn modification patterns that maintain the central scaffold while optimizing properties, mirroring common practices in medicinal chemistry lead optimization.
To further understand why DrugLLM achieves these emergent behaviors, we highlight the design philosophy of FGT, which is fundamentally representation-centric and distinct from retrosynthetic fragmentation methods like RECAP or BRICS. While the latter prioritizes formal synthetic feasibility, it often leads to extremely sparse vocabularies (e.g., exceeding 500![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 tokens) that cover only a fraction of drug-like chemical space, creating a “data sparsity” bottleneck for LLM training. In contrast, FGT treats structural groups as “reasoning primitives,” optimizing for a compact yet expressive vocabulary (4796 tokens) that ensures each motif is seen with sufficient frequency during pre-training. By providing an intermediate, sequential representation that reduces sequence length by 53.27% compared to SMILES, FGT effectively bridges symbolic chemistry with the sequence-modeling strengths of Transformers. This approach mitigates representation variance and long-range dependency issues, allowing DrugLLM to disentangle domain-invariant optimization logic from target-specific motifs—a capability that reaction-based or purely atom-based methods struggle to achieve.
000 tokens) that cover only a fraction of drug-like chemical space, creating a “data sparsity” bottleneck for LLM training. In contrast, FGT treats structural groups as “reasoning primitives,” optimizing for a compact yet expressive vocabulary (4796 tokens) that ensures each motif is seen with sufficient frequency during pre-training. By providing an intermediate, sequential representation that reduces sequence length by 53.27% compared to SMILES, FGT effectively bridges symbolic chemistry with the sequence-modeling strengths of Transformers. This approach mitigates representation variance and long-range dependency issues, allowing DrugLLM to disentangle domain-invariant optimization logic from target-specific motifs—a capability that reaction-based or purely atom-based methods struggle to achieve.
An additional benefit of this domain-specific design is the reduction of generative “hallucinations,” i.e., chemically implausible or invalid molecules. By generating at the functional-group level with predefined structural and semantic constraints, and anchoring modifications around known leads in the few-shot setting, DrugLLM maintains chemical validity. All outputs are further sanitized using RDKit to filter invalid structures. While this does not entirely eliminate rare edge cases or complex inconsistencies, these mechanisms collectively ensure that DrugLLM generates high-quality, chemically reasonable molecules.
In our computational experiments, DrugLLM demonstrated state-of-the-art optimization performance across 20 biological targets based on large-scale ChEMBL bioassay data, providing a statistically robust evaluation of the model's performance on these datasets. To further assess real-world applicability, we conducted wet-lab validation on two HCN2 inhibitor candidates (HCN2-M1 and HCN2-M2) as a proof-of-concept (PoC). Both candidates exhibited improved inhibitory potency against HCN2 compared to ivabradine. Although the experimental scope is limited, these PoC results demonstrate that DrugLLM can effectively guide experimental candidate selection under realistic constraints.
Despite these advantages, this study has several limitations. First, DrugLLM's zero-shot molecular optimization capability remains relatively elementary; while it can optimize molecules guided by simple instructions, complex constraints such as protein structures are still challenging. Second, the current model handles either a single property or simple combinations of two properties, and optimizing more complex multi-property objectives simultaneously remains difficult. Third, the wet-lab validation, limited to two HCN2 inhibitors, provides only a preliminary assessment of real-world performance; comprehensive investigation of synthetic routes, mechanisms of action, and preclinical development was beyond the scope of this study. Future work will aim to systematically expand wet-lab validation, develop more sophisticated multi-property optimization strategies, and explore the progression of promising candidates toward preclinical development.
Materials and methods
Data collection and preparation
To train and analyze the DrugLLM model, we constructed a large-scale dataset from the ZINC23 and ChEMBL24,25 datasets. ZINC is a free database that contains more than 230 million purchasable compounds in ready-to-dock, 3D formats. We filtered the drug-like molecules from ZINC and obtained 4.5 million molecules. ChEMBL is a comprehensive repository for bioactive compounds with their properties. We gathered bioactivity data from the ChEMBL database with the corresponding web resource client, totaling approximately 180 million molecules. Following the preprocessing pipeline in Stanley et al.,45 we excluded all compounds that were not drug-like molecules. A standard cleaning and canonicalization procedure was applied to the filtered compounds. All of the molecules were represented by FGT strings and labeled with specific properties. To facilitate property comparisons between two molecules, we only considered property categories with real numbers. Therefore, we obtained a large-scale dataset that comprised thousands of data tables, each table corresponding to hundreds of molecules measured by the same property.Based on the collected tabular data, we then transformed them into meaningful textual sentences and paragraphs. In particular, we regarded the modification between two molecules with similar structures as a sentence and multiple cases of molecular modifications as a paragraph. In the meantime, we stipulated that the molecular modifications in a paragraph should describe the same property changes. In other words, if the first two cases of molecule modifications indicated an increase in solubility, we ensured that the remaining sentences of this paragraph were all about the solubility improvement.
To ensure that modifications were captured between molecules sharing core structural features, we implemented a heuristic clustering algorithm based on molecular scaffolds. Specifically, given a pool of molecules with their associated properties, we first clustered the molecules based on scaffold-level structural similarity. Molecular similarity was computed using RDKit fingerprints with the Dice similarity metric,51 which effectively captures scaffold-level relationships. A molecule was assigned to a cluster if its similarity to the corresponding cluster center exceeded 0.60. This cutoff, consistent with settings used in JTVAE experiments,9 helped maintain scaffold consistency within each cluster while allowing peripheral variations. Cluster centers were initially chosen randomly, and the number of centers was dynamically increased until all molecules were assigned, resulting in a scaffold-level grouping across the dataset. This structural grouping facilitated the subsequent step of selecting molecule pairs within clusters that exhibited consistent property changes (e.g., solubility increase) to form the modification sentences and paragraphs described above.
Apart from the modifications regarding a single property, we also considered the combinations of multiple properties, which were mainly involved in the simple molecular properties that can be calculated by Python scripts. In total, we collected about 24.6 million modification paragraphs and 184.7 million molecules to build the training dataset. The dataset involves over 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 different molecular properties, activities, and compositions. In addition to the FGT strings of molecules, we also added descriptions of the property optimizations at the beginning of each paragraph to build the relationship between the molecule structures and the semantic meaning of the properties.
000 different molecular properties, activities, and compositions. In addition to the FGT strings of molecules, we also added descriptions of the property optimizations at the beginning of each paragraph to build the relationship between the molecule structures and the semantic meaning of the properties.
Functional group tokenization (FGT)
The construction of an FGT string involved segmenting a molecule into chemically meaningful structural fragments and recording their connectivity. The decomposition process iteratively removed peripheral fragments from the molecule while ensuring that the remaining structure stayed connected. This continued until only a central core fragment remained, which was often the largest fused ring system within the molecule. The resulting FGT string began with this core fragment, followed by the removed fragments attached sequentially using “/” as a separator, in the reverse order of their removal (effectively core-to-periphery).This core-first alignment establishes a single, unambiguous starting point for the sequence, reducing encoding variability compared to starting from potentially multiple equivalent peripheral points. It embeds a natural structural hierarchy, with the core structure preceding peripheral fragments, thereby aiming to enhance interpretability and provide a stable input representation for machine learning models. A detailed algorithmic specification of FGT, together with formal definitions of all variables used in Algorithms S1 and S2, is provided in the SI Notes (see the section Implementation details of functional group tokenization (FGT)).
The FGT framework was implemented in three key stages:
(1) Dictionary construction: we first leveraged the extensive molecular data resources available in the training dataset. Ring structures were identified, and rings sharing one or more atoms (including those connected via external double/triple bonds, forming conjugated systems) were merged into fused ring systems. For non-ring regions, non-ring C–C single bonds were selectively broken to isolate side-chain fragments, while other bonds (e.g., C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, C–N, C–S, and multiple bonds) were generally kept intact within fragments. This approach allowed complete decomposition of a molecule into chemically meaningful fragments, assigned each unique fragment (after canonicalization) a string identifier, and constructed a comprehensive fragment dictionary.
O, C–N, C–S, and multiple bonds) were generally kept intact within fragments. This approach allowed complete decomposition of a molecule into chemically meaningful fragments, assigned each unique fragment (after canonicalization) a string identifier, and constructed a comprehensive fragment dictionary.
(2) Molecular encoding: the encoding logic was designed as a deterministic, rule-based decomposition process, as detailed in Algorithm S1. To ensure the practical injectivity and reversibility of the FGT scheme, the algorithm transformed a molecular graph into a unique token sequence through a canonical graph traversal. The process began by identifying structural fragments, including fused ring systems and side-chain motifs. The decomposition iteratively removed peripheral fragments while maintaining the connectivity of the remaining molecular graph, prioritizing terminal groups in each step. A significant challenge in fragment-based tokenization is the instability of atom indexing, which often shifts during the SMILES canonicalization of isolated fragments. To resolve this and ensure a consistent mapping, FGT utilized a multi-dimensional local atomic environment descriptor based on Breadth-First Search (BFS). This descriptor captured the intrinsic topological properties around the attachment points layer-by-layer. By recording these BFS-based identifiers to locate attachment atoms instead of relying solely on volatile atom indices, the algorithm could reliably re-identify corresponding atoms across the encoding and decoding stages. This process was repeated until only a single core structural group remained, establishing a canonical core-to-periphery order. Finally, the core and recorded fragments were mapped to their unique dictionary identifiers and progressively integrated using “/” as a separator.
(3) Molecular decoding: the decoding process reversed the encoding procedure, reconstructing the molecular structure from its FGT string as described in Algorithm S2. Starting from the initial core fragment specified in the string, each subsequent structural group listed after a “/” was reattached to the current structure at the position re-identified by the recorded BFS-based descriptors. This process continued until all groups were spliced back, thereby restoring the original molecular topology. The reversibility of this process ensured both structural integrity and faithful chemical representation.
The FGT representation aims to enhance encoding consistency and provide a compact, canonical, and chemically interpretable input space for the learning algorithm described in the next section.
Next modification prediction (NMP) paradigm
DrugLLM introduces the next modification prediction (NMP) paradigm, a molecular-level learning framework that emulates the iterative design process of medicinal chemists. NMP defines the overall learning paradigm, in which the model predicts the next molecular modification from one molecule to another.This paradigm is implemented during training with an autoregressive modification prediction (RMP) objective. Under RMP, the model learns to generate the FGT string representation of the next target molecule token by token. This generation is conditioned on the sequence of preceding molecules (represented by their FGT strings) within a specific property optimization trajectory provided as context. The RMP objective thus follows the standard autoregressive decoding strategy common in Transformer-based language models.26,52 In essence, NMP sets the molecular-level predictive goal, while RMP provides the token-level autoregressive mechanism to achieve it during training and inference.
The NMP framework aims to implicitly capture biochemical principles directly from the patterns in modification data, reducing the reliance on explicit domain knowledge encoding. When implemented using the FGT representation, as done in DrugLLM, the NMP paradigm allows formulating molecular optimization as an interpretable and context-aware sequence generation task. This approach thereby bridges symbolic chemical representation with the powerful sequence modeling capabilities of large-scale language models.
Implementation of DrugLLM
DrugLLM was built upon the Transformer decoder architecture. Specifically, we adopted a model consisting of 32 layers and 32 attention heads to extract contextual information from the input sequences. The hidden dimension was set to 4096. In total, DrugLLM comprised 7 billion parameters, all of which were updated during pre-training.Similar to other large language models, the core training objective of DrugLLM is to predict the next token in a sequence in an autoregressive manner. Formally, a training sequence x in the DrugLLM dataset is constructed by concatenating an optimization instruction o (describing the desired property change) with a sequence representing a trajectory of molecules m, given by:
| x = [o,m1,m2,…,mn] | (1) |
During the training process, DrugLLM learned to approximate the probability of the next token xt given the context of the previous tokens (x1, …, xt−1). This autoregressive objective (referred to as RMP in the previous section) was computed by maximizing the likelihood:
| P(xt|x1,x2,…,xt−1) = DrugLLM(x1,x2,…,xt−1) | (2) |
This training paradigm enabled the model to learn the patterns underlying sequential molecule modifications represented as token sequences, which directly supports its application in molecular optimization. The training process employed the AdamW optimizer with a learning rate of 3 × 10−5, utilizing a cosine annealing schedule and a linear warm-up phase of 1000 steps. We utilized mixed-precision training (fp16) and ZeRO optimizer stage 3 for efficient large-scale training. The model was trained for three epochs over the entire dataset.
For inference, DrugLLM leverages its learned autoregressive generation capability.
Few-shot molecular optimization: given a context consisting of an optimization instruction (o), a few example molecule modifications represented as a sequence (m1, …, mK), and a query molecule (mo), the model generates the optimized molecule (mg) token-by-token by predicting the sequence following the input [o, m1, …, mK, mo]. The next generated molecule is the optimized molecule.
Zero-shot molecular optimization: the model takes only the natural language description of the optimization task (o) and the query molecule (mo) as input, i.e., [o, mo], to generate the optimized molecule (mg).
Region-constrained generation: furthermore, DrugLLM supports region-constrained generation at inference time without requiring retraining. By providing a partial FGT sequence corresponding to the query molecule where certain fragments are fixed as an immutable prefix, users can condition the generation. The model then autoregressively completes the sequence, effectively applying modifications only to the unconstrained (non-prefix) regions. This allows for localized and structurally controlled molecular editing. An illustrative example of such region-constrained modification is shown in Fig. S4.
Author contributions
X. L. and Y. G. designed and performed the experiments, interpreted the results, and wrote the manuscript. M. L. primarily conducted the wet-lab experiments. W. Z. and P. L. assisted with the computational experiments. J. Liu contributed to data interpretation and experimental procedures. S. H. assisted with research design and data interpretation. B. K. and J. Lv supported the experimental setup and offered guidance throughout the study.Conflicts of interest
There are no conflicts to declare.Data availability
The pre-training datasets used in this study are publicly accessible at the following links: ZINC database (https://zinc15.docking.org) and ChEMBL database (https://www.ebi.ac.uk/chembldb). The preprocessed training data used for DrugLLM, along with the wet-lab experimental results supporting this study, are available at https://osf.io/f6yqn.The source code for DrugLLM is available on GitHub at https://github.com/ziyanglichuan/DrugLLM. The pre-trained model can be accessed via the Hugging Face platform at https://huggingface.co/ziyanglichuan/DrugLLM.
Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5sc08859c.
Acknowledgements
This work was supported by the Science and Technology Major Project of Sichuan Province (2024ZDZX0003), the National Key R&D Program of China (2024YFB3312503), the Natural Science Foundation of Sichuan Province (2024NSFTD0048), and the State Key Laboratory of Advanced Nuclear Energy Technology in Nuclear Power Institute of China (STSW-0224-0202-08-01). We also acknowledge the support of Sichuan Province Engineering Technology Research Center of Broadband Electronics Intelligent Manufacturing.Notes and references
- T. Brown, B. Mann, N. Ryder, M. Subbiah, J. D. Kaplan, P. Dhariwal, A. Neelakantan, P. Shyam, G. Sastry and A. Askell, Language models are few-shot learners, NeurIPS, 2020, pp. 1877–1901 Search PubMed.
- Y. Li, D. Choi, J. Chung, N. Kushman, J. Schrittwieser, R. Leblond, T. Eccles, J. Keeling, F. Gimeno and A. Dal Lago, Competition-level code generation with alphacode, Science, 2022, 378, 1092–1097 CrossRef CAS PubMed.
- M. Yuksekgonul, F. Bianchi, J. Boen, S. Liu, P. Lu, Z. Huang, C. Guestrin and J. Zou, Optimizing generative AI by backpropagating language model feedback, Nature, 2025, 639, 609–616 CrossRef CAS PubMed.
- A. V. Sadybekov and V. Katritch, Computational approaches streamlining drug discovery, Nature, 2023, 616, 673–685 CrossRef CAS PubMed.
- X. Liu, Z. Xue, M. Luo, B. Ke and J. Lv, Anesthetic drug discovery with computer-aided drug design and machine learning, Anesthesiol. Perioper. Sci., 2024, 2, 7 CrossRef.
- W. Kohn and L. J. Sham, Self-consistent equations including exchange and correlation effects, Phys. Rev., 1965, 140, A1133 CrossRef.
- M. Karplus and J. A. McCammon, Molecular dynamics simulations of biomolecules, Nat. Struct. Biol., 2002, 9, 646–652 CrossRef CAS PubMed.
- J.-L. Yu, C. Zhou, X.-L. Ning, J. Mou, F.-B. Meng, J.-W. Wu, Y.-T. Chen, B.-D. Tang, X.-G. Liu and G.-B. Li, Knowledge-guided diffusion model for 3D ligand-pharmacophore mapping, Nat. Commun., 2025, 16, 2269 CrossRef CAS PubMed.
- W. Jin, R. Barzilay and T. Jaakkola, Junction tree variational autoencoder for molecular graph generation, ICML, 2018, pp. 2323–2332 Search PubMed.
- M. J. Kusner, B. Paige and J. M. Hernández-Lobato, Grammar variational autoencoder, ICML, 2017, pp. 1945–1954 Search PubMed.
- Z. Xue, C. Sun, W. Zheng, J. Lv and X. Liu, TargetSA: adaptive simulated annealing for target-specific drug design, Bioinformatics, 2025, 41, btae730 CrossRef CAS PubMed.
- F. Zhuang, Z. Qi, K. Duan, D. Xi, Y. Zhu, H. Zhu, H. Xiong and Q. He, A comprehensive survey on transfer learning, Proc. IEEE, 2020, 109, 43–76 Search PubMed.
- S. Chen, O. Zhang, C. Jiang, H. Zhao, X. Zhang, M. Chen, Y. Liu, Q. Su, Z. Wu and X. Wang, Deep lead optimization enveloped in protein pocket and its application in designing potent and selective ligands targeting LTK protein, Nat. Mach. Intell., 2025, 1–11 Search PubMed.
- Z. Wu, O. Zhang, X. Wang, L. Fu, H. Zhao, J. Wang, H. Du, D. Jiang, Y. Deng and D. Cao, Leveraging language model for advanced multiproperty molecular optimization via prompt engineering, Nat. Mach. Intell., 2024, 1–11 Search PubMed.
- J. Von Oswald, E. Niklasson, E. Randazzo, J. Sacramento, A. Mordvintsev, A. Zhmoginov and M. Vladymyrov, Transformers learn in-context by gradient descent, ICML, 2023, pp. 35151–35174 Search PubMed.
- D. Weininger, SMILES, a chemical language and information system. 1. introduction to methodology and encoding rules, J. Chem. Inf. Comput. Sci., 1988, 28, 31–36 CrossRef CAS.
- X. Q. Lewell, D. B. Judd, S. P. Watson and M. M. Hann, Recap retrosynthetic combinatorial analysis procedure: a powerful new technique for identifying privileged molecular fragments with useful applications in combinatorial chemistry, J. Chem. Inf. Comput. Sci., 1998, 38, 511–522 CrossRef CAS PubMed.
- J. Degen, C. Wegscheid-Gerlach, A. Zaliani and M. Rarey, On the art of compiling and using’drug-like’chemical fragment spaces, ChemMedChem, 2008, 3, 1503 CrossRef CAS PubMed.
- Y. Diao, D. Liu, H. Ge, R. Zhang, K. Jiang, R. Bao, X. Zhu, H. Bi, W. Liao and Z. Chen, Macrocyclization of linear molecules by deep learning to facilitate macrocyclic drug candidates discovery, Nat. Commun., 2023, 14, 4552 CrossRef CAS PubMed.
- M. Jiang, T. Liu, M. Hussain, Y. Luo, R. Zheng, T. Hou, X. Lu and Y. Zhou, Macrocycle-DB: a comprehensive database for macrocycle-based drug discovery, Nucleic Acids Res., 2025, gkaf1107 Search PubMed.
- H. Zhao, S. Liu, M. Chang, H. Xu, J. Fu, Z. Deng, L. Kong and Q. Liu, Gimlet: A unified graph-text model for instruction-based molecule zero-shot learning, Adv. Neural Inform. Process. Syst., 2023, 36, 5850–5887 Search PubMed.
- J. Lim, S. Ryu, J. W. Kim and W. Y. Kim, Molecular generative model based on conditional variational autoencoder for de novo molecular design, J. Cheminf., 2018, 10, 1–9 Search PubMed.
- Z. Wu, B. Ramsundar, E. N. Feinberg, J. Gomes, C. Geniesse, A. S. Pappu, K. Leswing and V. Pande, MoleculeNet: A benchmark for molecular machine learning, Chem. Sci., 2018, 9, 513–530 RSC.
- D. Mendez, A. Gaulton, A. P. Bento, J. Chambers, M. De Veij, E. Félix, M. P. Magariños, J. F. Mosquera, P. Mutowo and M. Nowotka, ChEMBL: Towards direct deposition of bioassay data, Nucleic Acids Res., 2019, 47, D930–D940 CrossRef CAS PubMed.
- M. Davies, M. Nowotka, G. Papadatos, N. Dedman, A. Gaulton, F. Atkinson, L. Bellis and J. P. Overington, ChEMBL web services: Streamlining access to drug discovery data and utilities, Nucleic Acids Res., 2015, 43, W612–W620 CrossRef CAS PubMed.
- A. Vaswani, N. Shazeer, N. Parmar, J. Uszkoreit, L. Jones, A. N. Gomez, Ł. Kaiser and I. Polosukhin, Attention is All you Need, NeurIPS, 2017, vol. 30 Search PubMed.
- R. Sennrich, B. Haddow and A. Birch, Neural Machine Translation of Rare Words with Subword Units, ACL, 2016, vol. 1, pp. 1715–1725.
- W. Jin, K. Yang, R. Barzilay and T. Jaakkola, Learning multimodal graph-to-graph translation for molecular optimization, ICLR, 2018 Search PubMed.
- K. Maziarz, H. Jackson-Flux and P. Cameron, Learning to extend molecular scaffolds with structural motifs, ICLR, 2021, pp. 1–10 Search PubMed.
- K. Yang, K. Swanson, W. Jin, C. Coley, P. Eiden, H. Gao, A. Guzman-Perez, T. Hopper, B. Kelley and M. Mathea, Analyzing learned molecular representations for property prediction, J. Chem. Inf. Model., 2019, 59, 3370–3388 CrossRef CAS PubMed.
- Y. Atzmon, F. Kreuk, U. Shalit and G. Chechik, A causal view of compositional zero-shot recognition, Adv. Neural Inf. Process. Syst., 2020, 33, 1462–1473 Search PubMed.
- Q. Wang, L. Liu, C. Jing, H. Chen, G. Liang, P. Wang and C. Shen, Learning conditional attributes for compositional zero-shot learning, Proc. IEEE/CVF Conf. Comput. Vis. Pattern Recognit., 2023, 11197–11206 Search PubMed.
- G. Le Mens, B. Kovács, M. T. Hannan and G. Pros, Uncovering the semantics of concepts using GPT-4, Proc. Natl. Acad. Sci., 2023, 120, e2309350120 CrossRef CAS PubMed.
- Z. Du, Y. Qian, X. Liu, M. Ding, J. Qiu, Z. Yang and J. Tang, GLM: General Language Model Pretraining with Autoregressive Blank Infilling, Proc. 60th Annu. Meet. Assoc. Comput. Linguist., 2022, 320–335 Search PubMed.
- Z. Chen, A. H. Cano, A. Romanou, A. Bonnet, K. Matoba, F. Salvi, M. Pagliardini, S. Fan, A. Köpf and A. Mohtashami, Meditron-70b: Scaling medical pretraining for large language models, arXiv, 2023, DOI:10.48550/arXiv.2311.16079.
- E. Bolton, A. Venigalla, M. Yasunaga, D. Hall, B. Xiong, T. Lee, R. Daneshjou, J. Frankle, P. Liang and M. Carbin, Biomedlm: A 2.7 b parameter language model trained on biomedical text, arXiv, 2024, DOI:10.48550/arXiv.2403.18421.
- H. Touvron, T. Lavril, G. Izacard, X. Martinet, M.-A. Lachaux, T. Lacroix, B. Rozière, N. Goyal, E. Hambro and F. Azhar, LLama: Open and efficient foundation language models, arXiv, 2023, DOI:10.48550/arXiv.2302.13971.
- U. B. Kaupp and R. Seifert, Molecular diversity of pacemaker ion channels, Annu. Rev. Physiol., 2001, 63, 235–257 CrossRef CAS PubMed.
- M. Biel, A. Schneider and C. Wahl, Cardiac HCN channels: structure, function, and modulation, Trends Cardiovasc. Med., 2002, 12, 206–213 CrossRef CAS PubMed.
- E. C. Emery, G. T. Young, E. M. Berrocoso, L. Chen and P. A. McNaughton, HCN2 ion channels play a central role in inflammatory and neuropathic pain, Science, 2011, 333, 1462–1466 CrossRef CAS PubMed.
- K. Swedberg, M. Komajda, M. Böhm, J. S. Borer, I. Ford, A. Dubost-Brama, G. Lerebours and L. Tavazzi, Ivabradine and outcomes in chronic heart failure (SHIFT): a randomised placebo-controlled study, Lancet, 2010, 376, 875–885 CrossRef CAS PubMed.
- M. Melchiorre, M. Del Lungo, L. Guandalini, E. Martini, S. Dei, D. Manetti, S. Scapecchi, E. Teodori, L. Sartiani and A. Mugelli, Design, synthesis, and preliminary biological evaluation of new isoform-selective f-current blockers, J. Med. Chem., 2010, 53, 6773–6777 CrossRef CAS PubMed.
- S.-J. Chen, Y. Xu, Y.-M. Liang, Y. Cao, J.-Y. Lv, J.-X. Pang and P.-Z. Zhou, Identification and characterization of a series of novel HCN channel inhibitors, Acta Pharmacol. Sin., 2019, 40, 746–754 CrossRef CAS PubMed.
- A. Bucchi, M. Baruscotti, M. Nardini, A. Barbuti, S. Micheloni, M. Bolognesi and D. DiFrancesco, Identification of the molecular site of ivabradine binding to HCN4 channels, PLoS One, 2013, 8, e53132 CrossRef CAS PubMed.
- M. Stanley, J. F. Bronskill, K. Maziarz, H. Misztela, J. Lanini, M. Segler, N. Schneider and M. Brockschmidt, FS-Mol: A Few-Shot Learning Dataset of Molecules, NeurIPS, 2021 Search PubMed.
- J. Ma, S. H. Fong, Y. Luo, C. J. Bakkenist, J. P. Shen, S. Mourragui, L. F. Wessels, M. Hafner, R. Sharan and J. Peng, Few-shot learning creates predictive models of drug response that translate from high-throughput screens to individual patients, Nat. Cancer, 2021, 2, 233–244 CrossRef CAS PubMed.
- C. Zhang, B. Jia, Y. Zhu and S.-C. Zhu, Human-level few-shot concept induction through minimax entropy learning, Sci. Adv., 2024, 10, eadg2488 CrossRef PubMed.
- K. Maeng, A. Colin and B. Lucia, Alpaca: Intermittent execution without checkpoints, Proc. ACM Program. Lang., 2017, 1, 1–30 CrossRef.
- R. Luo, L. Sun, Y. Xia, T. Qin, S. Zhang, H. Poon and T.-Y. Liu, BioGPT: Generative pre-trained transformer for biomedical text generation and mining, Briefings Bioinf., 2022, 23, bbac409 CrossRef PubMed.
- Y. Li, C. Gao, X. Song, X. Wang, Y. Xu and S. Han, DrugGPT: A GPT-based Strategy for Designing Potential Ligands Targeting Specific Proteins, bioRxiv, 2023, DOI:10.1101/2023.06.29.543848.
- S. Riniker and G. A. Landrum, Similarity maps-a visualization strategy for molecular fingerprints and machine-learning methods, J. Cheminf., 2013, 5, 43 CAS.
- A. Radford, J. Wu, R. Child, D. Luan, D. Amodei and I. Sutskever, Language models are unsupervised multitask learners, OpenAI Technical Report, 2019, https://cdn.openai.com/better-language-models/language_models_are_unsupervised_multitask_learners.pdf Search PubMed.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |