Open Access Article
Open Access ArticleUnlocking azobenzene isomerization mechanisms via an LLM agent-driven workflow integrating simulation, experiment, and machine learning
Yixi Shen†
a,
Ledu Wang†a,
Yan Huang†*a,
Xiaolong Zhang† a,
Meng Huanga,
Huirong Lia,
Jing Hea,
Aoran Caia,
Yang Wanga,
Pieter E. S. Smithb,
Jun Jiang
a,
Meng Huanga,
Huirong Lia,
Jing Hea,
Aoran Caia,
Yang Wanga,
Pieter E. S. Smithb,
Jun Jiang *ac,
Zhuoying Zhu*a and
Linjiang Chen
*ac,
Zhuoying Zhu*a and
Linjiang Chen *a
*a
aState Key Laboratory of Precision and Intelligent Chemistry, Hefei National Research Center for Physical Sciences at the Microscale, School of Chemistry and Materials Science, University of Science and Technology of China, Hefei 230026, China. E-mail: hyang@ustc.edu.cn; jiangj1@ustc.edu.cn; zyzhuq@ustc.edu.cn; linjiangchen@ustc.edu.cn
bHefei JiShu Quantum Technology Co., Ltd, Hefei 230026, China
cHefei National Laboratory, University of Science and Technology of China, Hefei, 230026, China
First published on 25th March 2026
Abstract
Bridged azobenzene derivatives are key photo-responsive molecular switches. However, probing and interpreting their microscopic Z ↔ E isomerization mechanisms remain challenging as isolated spectroscopic and computational efforts struggle to establish clear structure–spectrum relationships. We report an integrated, large-language-model (LLM) agent-driven workflow that links literature-guided planning, ab initio molecular dynamics (AIMD) sampling, density functional theory spectral calculations, robotic infrared/Raman measurements, and interpretable machine learning for structural–spectral analysis of bridged azobenzenes. Central to the analysis is an attention-based convolutional neural network (ATT-CNN) that predicts the C–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–C dihedral angle directly from vibrational spectra with r = 0.99 and MAE = 5°. Attention maps highlight mechanistically informative bands and support holistic (non-marker-dependent) interpretation; transfer learning extends performance across chemical environments and experimental datasets. LLM agents formulated the research plan and coordinated automated simulations and measurements, whereas neural-network architecture design, training, and comparative benchmarking were performed by human researchers to retain full flexibility for model exploration and ensure rigorous interpretation. To our knowledge, this is the first LLM-agent-planned and -orchestrated mechanistic study unifying literature synthesis, theory, experiment, and machine learning. The resulting strategy advances quantitative insight into azobenzene photoisomerization and provides a generalizable blueprint for AI-driven investigations of dynamic molecular systems.
N–C dihedral angle directly from vibrational spectra with r = 0.99 and MAE = 5°. Attention maps highlight mechanistically informative bands and support holistic (non-marker-dependent) interpretation; transfer learning extends performance across chemical environments and experimental datasets. LLM agents formulated the research plan and coordinated automated simulations and measurements, whereas neural-network architecture design, training, and comparative benchmarking were performed by human researchers to retain full flexibility for model exploration and ensure rigorous interpretation. To our knowledge, this is the first LLM-agent-planned and -orchestrated mechanistic study unifying literature synthesis, theory, experiment, and machine learning. The resulting strategy advances quantitative insight into azobenzene photoisomerization and provides a generalizable blueprint for AI-driven investigations of dynamic molecular systems.
Introduction
The reversible Z ↔ E photoisomerization of azobenzene derivatives is a foundational process in photochemistry, enabling the construction of molecular switches for a wide array of applications, such as responsive materials,1–6 photopharmacology,7–9 molecular machines,10,11 and optoelectronics.12,13 Within this versatile class, bridged azobenzenes—distinguished by a covalent tether between their phenyl rings—offer a set of especially attractive features, such as enhanced photoisomerization quantum yields, well-separated absorption bands for the Z and E isomers, and, in some cases, thermodynamically stabilized Z-isomers.14,15 These characteristics allow for more precise and robust switching behavior, which is essential for advanced photoresponsive systems.Despite decades of research, a comprehensive molecular-level understanding of the dynamic Z ↔ E isomerization process across the vast chemical space of bridged azobenzene derivatives and under realistic environmental conditions remains elusive.16–19 This challenge arises from several persistent limitations inherent to traditional approaches. Conventional studies often rely on painstaking, molecule-by-molecule experimental and computational investigations,20 which restrict the systematic exploration of chemical diversity and slow the rational design and optimization of new derivatives. Moreover, experimental efforts to capture the ultrafast dynamics of isomerization are further complicated by the inherently complex and often overlapping spectral features, the high sensitivity of these features to environmental factors, and the difficulty in directly interpreting subtle changes in molecular conformation or distinguishing transient intermediates.21,22
From a theoretical standpoint, the accurate simulation of photochemical dynamics in these systems requires a careful balance between computational precision and feasibility, especially when it comes to modeling non-adiabatic transitions,23,24 explicit solvent effects,24 and rare but critical conformational states.25,26 Moreover, there is often a disconnect between theoretical predictions and experimental measurements; these efforts are frequently pursued in parallel rather than in an integrated, high-throughput, or feedback-driven fashion. As a result, closing the loop between hypothesis, observation, and mechanistic understanding remains a formidable task. Together, these factors create a substantial bottleneck that limits both the development of fundamental insight and the accelerated application of photo-switchable azobenzene-based materials.
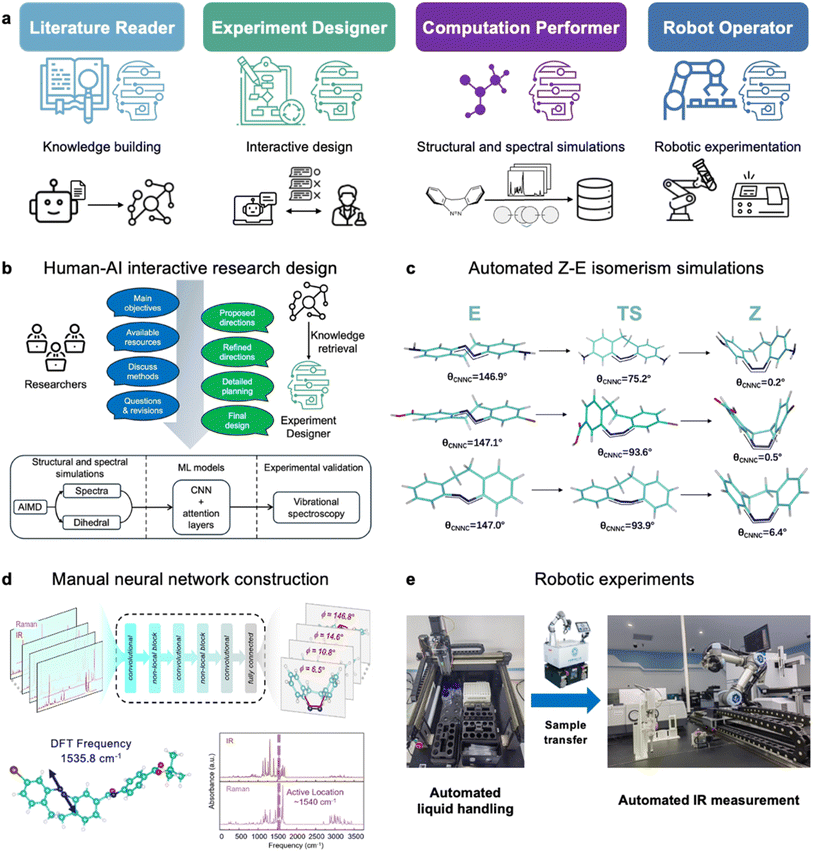
Recent advances in laboratory automation, robotics, and artificial intelligence (AI),27–32 most notably large language models (LLMs) and their agent systems—present a transformative opportunity to overcome these long-standing barriers. By integrating autonomous knowledge extraction, hypothesis generation, experimental design, high-throughput data acquisition, and interpretable machine learning, it is now possible to move beyond piecemeal strategies and toward data-driven, AI-augmented collaborative research. In this work, we developed and implemented an integrated workflow for the elucidation of Z ↔ E isomerization dynamics in bridged azobenzenes (Fig. 1), powered by a multi-agent-driven robotic AI chemist platform.31 Our workflow combines agent-guided computational simulation and spectral modeling, automated experimental spectroscopic measurements, and interpretable machine learning to enable automated, high-throughput structural analysis during photoisomerization. By systematically coordinating the actions of literature-mining, research-planning, data-generating, and machine-learning agents, our approach lays the groundwork for autonomous structure–function discovery and rational design of new molecular photoswitches. The results and methodology presented herein seek not only to advance the study of azobenzene photochemistry, but also to serve as a blueprint for autonomous, generalizable research platforms capable of accelerating discovery across the broader landscape of dynamic molecular systems. It should be noted that this study does not directly simulate the excited-state dynamics (such as the evolution prior to the conical intersection), which constitute the key mechanism governing the ultrafast photochemical step. Instead, it focuses on establishing a quantitative “structure–spectrum” relationship based on ground-state conformers and their vibrational spectra. This research paradigm not only provides a feasible pathway for high-throughput structural analysis of complex photochemical systems but also lays a methodological foundation for subsequent studies that integrate steady-state analysis with excited-state dynamics.
Results and discussion
Knowledge extraction and graph-based analysis
To establish a rigorous foundation for collaborative research planning, we deployed the Literature Reader agent to systematically extract knowledge from the scientific literature on azobenzene Z ↔ E isomerization. The agent analyzed 80 peer-reviewed articles—selected by human experts—encompassing recent advances in azobenzene photochemistry, mechanistic studies of isomerization pathways, spectroscopic characterization, and theoretical modeling. The selection spanned experimental reports of vibrational spectra, computational analyses of isomerization dynamics, and seminal works that define the conceptual framework for this field (see SI Data files for the full list of publications).The Literature Reader combined GraphRAG (Graph Retrieval-Augmented Generation) technique33 with state-of-the-art large language models, DeepSeek V3 (ref. 34) and DeepSeek R1 (ref. 35) to construct a comprehensive, hierarchical knowledge graph. These open-source models were selected to ensure cost-efficiency for token-intensive workflows and to facilitate reproducibility within the scientific community, avoiding reliance on proprietary ecosystems. This approach enabled multiscale parsing of the literature—capturing not only chemical entities and molecular structures, but also experimental conditions, spectroscopic features, theoretical methodologies, kinetic and thermodynamic data, and mechanistic hypotheses. The resulting knowledge graph was organized such that each publication formed a high-level node, with nested sub-nodes detailing molecular systems, experimental protocols, spectral observations, and computational outcomes. This architecture facilitated the rapid identification of knowledge gaps, precedents, and best practices relevant to our research objectives, mirroring the literature mining and knowledge integration framework previously developed for multi-domain catalyst discovery. To inform subsequent research planning, the Literature Reader was systematically queried to generate targeted reports synthesizing the key theoretical concepts and experimental methodologies in the field, providing a robust contextual foundation for intelligent experimental design.
Interactive human–LLM collaborative research planning
Building upon this structured knowledge base, the Research Designer agent initiated an interactive, dialogue-driven research planning process. Through a natural language conversational interface, human researchers articulated their scientific objectives—most notably, the challenge of developing a robust method for tracking the dynamic Z ↔ E isomerization of bridged azobenzene derivatives using experimentally accessible data. Leveraging both the chemical insights synthesized by the Literature Reader and its own scientific reasoning capabilities, the Research Designer agent engaged in stepwise analysis, systematically evaluating available methodologies and navigating alternative strategies. At each stage, the agent compared potential approaches, incorporated researcher-specified constraints, and provided tailored recommendations, ensuring that the final research plan was both comprehensive and optimally aligned with the project's goals. The following excerpts illustrate the collaborative process, where human requests and feedback guided the agent form a practical solution:User: “The idea of generating molecular ‘movie’ from a femtosecond Raman experiment is promising, however, conducting such experiments are resource-extensive, and we would like to know if there are promising methods to generate such trajectories via simulations”.
Agent: “Proposed Research Direction: High-Fidelity Nonadiabatic Molecular Dynamics Simulations… Rationale: While experimental ultrafast spectroscopy (e.g., FSRS) is powerful, its cost/time intensity can be mitigated by computational trajectory generation”.
User: “Are basic neural networks enough, or should we use advanced models like CNNs? … Since molecular vibrations exist, are individual bond lengths inappropriate…? Given our expertise and equipment, Raman spectra is our preferred spectral feature”.
Agent: “Refined Predictive Framework: Direct Dihedral Angle Prediction from Raman Spectra… Core Concept: Develop a 1D-CNN with Spectral Attention focused solely on predicting the bridge dihedral angle… This prioritizes quality over quantity”.
Consistent with our previous work on LLM-driven catalyst discovery, this collaborative process enabled effective fusion of knowledge across theoretical, spectroscopic, and experimental domains. The Research Designer recognized that achieving real-time, mechanistic insight into isomerization required bridging the gap between unobservable molecular parameters (such as the C–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–C dihedral angle) and directly measurable vibrational spectra. Thus, the agent proposed a strategy centered on constructing predictive, interpretable machine learning models capable of inferring structural coordinates from IR and Raman data, effectively translating complex spectroscopic signatures into actionable molecular information.
N–C dihedral angle) and directly measurable vibrational spectra. Thus, the agent proposed a strategy centered on constructing predictive, interpretable machine learning models capable of inferring structural coordinates from IR and Raman data, effectively translating complex spectroscopic signatures into actionable molecular information.
Computation–experimentation integration and automated workflow execution
Drawing on the comprehensive knowledge graph constructed by the Literature Reader, the Research Designer agent formulated a multi-pronged computational and experimental strategy to unravel the Z ↔ E isomerization dynamics of bridged azobenzenes. This agent-driven plan encompassed (1) automated ab initio molecular dynamics (AIMD) simulations to exhaustively sample the conformational landscape along the isomerization coordinate, (2) density functional theory (DFT) calculations to generate theoretical IR and Raman spectra for conformers—thus establishing a large, high-quality structure–spectra dataset, and (3) supervised machine learning, building deep neural network models, to quantitatively link vibrational spectra to the key C–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–C dihedral angle.
N–C dihedral angle.
The AIMD simulations were performed using the CP2K software package with the hybrid Gaussian plane wave method. The agent employed the PBE exchange–correlation functional augmented with Grimme's D3 dispersion correction and the DZVP basis set. Metadynamics was used with the dihedral angle as the collective variable to efficiently sample the reaction pathway within the canonical (NVT) ensemble, using a Nosé–Hoover thermostat. From these 5 ps trajectories (0.5 fs timestep), approximately 500 snapshots covering the complete Z-to-E transition were extracted for each system.
Subsequently, the infrared and Raman spectra for all sampled conformers were computed at the DFT level to generate the spectral dataset. All frequency calculations were carried out using Gaussian 16 with the B3LYP functional and the 6-31+G(d,p) basis set. The resulting harmonic frequencies were uniformly scaled, and spectral lines were broadened with a Lorentzian function (8 cm−1 FWHM) to produce continuous spectra suitable for model training.
The automated workflow produced a curated dataset of 4400 structure–spectrum pairs across 11 bridged azobenzene derivatives. With this high-fidelity dataset in hand, the workflow then shifted to the construction and training of interpretable machine-learning models to decode the spectral–structural relationship. To complement and validate the purely computational data, the workflow was extended to incorporate experimental measurements.
To ensure both robust validation and broad transferability, the workflow further incorporated systematic experimental measurements under multiple isomerization conditions, including both thermal and photoinduced processes, with the Experiment Designer and Robot Operator agents coordinating the planning and execution of high-throughput sample preparation and spectroscopic acquisition. This agent-driven integration enabled the capture of a comprehensive range of isomerization behaviors, resulting in a rigorously curated dataset for subsequent machine learning (ML) studies.
Combining automated data generation with human-led ML studies
The automated phase produced a comprehensive dataset of 4400 structure–spectrum pairs across 11 systems, a scale of data generation that would be prohibitively time-consuming to assemble manually. At this juncture, the workflow shifted to a human-led phase to transform raw data into mechanistic insight. Although LLM agents effectively synthesized literature, formulated the plan, and coordinated automated simulations and IR/Raman measurements, neural-network development demands human flexibility and accountability: selecting and developing architectures, constructing fair baselines, running reproducible hyperparameter studies, and, critically, interpreting results in a physically meaningful way. Accordingly, we specified and implemented the ATT-CNN and alternative models, established benchmarking protocols, and executed systematic attention analyses and ablation experiments that link spectral features to the C–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–C dihedral coordinate. This human–AI coordinated approach guaranteed that every stage of the workflow—from initial research conception and planning, through automated computation and simulation, to manual ML model development and high-throughput experimental validation—remained integrated, data-driven, and systematically executed. The following sections detail the specific computational, ML, and experimental advances that emerged from this multi-agent workflow.
N–C dihedral coordinate. This human–AI coordinated approach guaranteed that every stage of the workflow—from initial research conception and planning, through automated computation and simulation, to manual ML model development and high-throughput experimental validation—remained integrated, data-driven, and systematically executed. The following sections detail the specific computational, ML, and experimental advances that emerged from this multi-agent workflow.
The impact of neural network architecture on molecular structure prediction
C–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–C dihedral angles were calculated for the 4400 conformations of the 11 bridged azobenzene derivatives (Table S1) generated using AIMD. The IR and Raman spectra of these conformers were obtained via DFT calculations. The compiled dataset of spectrum-dihedral angle pairs was randomly divided into training and testing subsets in an 8
N–C dihedral angles were calculated for the 4400 conformations of the 11 bridged azobenzene derivatives (Table S1) generated using AIMD. The IR and Raman spectra of these conformers were obtained via DFT calculations. The compiled dataset of spectrum-dihedral angle pairs was randomly divided into training and testing subsets in an 8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
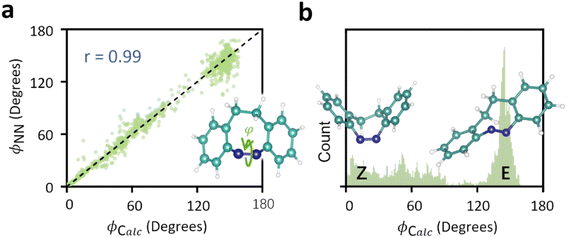
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 ratio. The overall performance of the attention-based convolutional neural network (ATT-CNN) achieved high accuracy, as evidenced by a 0.99 correlation (r) between the calculated dihedral angles (φCalc) and those predicted by the neural network (φNN) (Fig. 2a). As shown in Fig. 2b, the φCalc distribution spans a wide range, with dihedral angles of 6.4° (Z conformation) and 146.8° (E conformation) identified as relatively stable.
2 ratio. The overall performance of the attention-based convolutional neural network (ATT-CNN) achieved high accuracy, as evidenced by a 0.99 correlation (r) between the calculated dihedral angles (φCalc) and those predicted by the neural network (φNN) (Fig. 2a). As shown in Fig. 2b, the φCalc distribution spans a wide range, with dihedral angles of 6.4° (Z conformation) and 146.8° (E conformation) identified as relatively stable.
We compared ATT-CNN with a convolutional neural network without an attention layer (CNN), a traditional fully connected neural network (FCNN), a long short-term memory (LSTM) network, and a transformer network. Our results indicate that ATT-CNN best predicts the dihedral angle, yielding both the highest correlation with φCalc (r = 0.99) and the smallest mean absolute error (MAE) (MAE = 5.0°). Removal of the attention layer from ATT-CNN increased the MAE from 5.0° to 8.6° and decreased the correlation coefficient from 0.99 to 0.97, illustrating its importance to ATT-CNN's predictive performance. The MAEs for the LSTM, FCNN, and transformer networks were 7.3°, 10.3°, and 6.3°, respectively, and the correlations between φCalc and φNN for the LSTM, FCNN, and transformer networks were 0.96, 0.94, and 0.98, respectively.
The role of the attention mechanism in our machine learning model
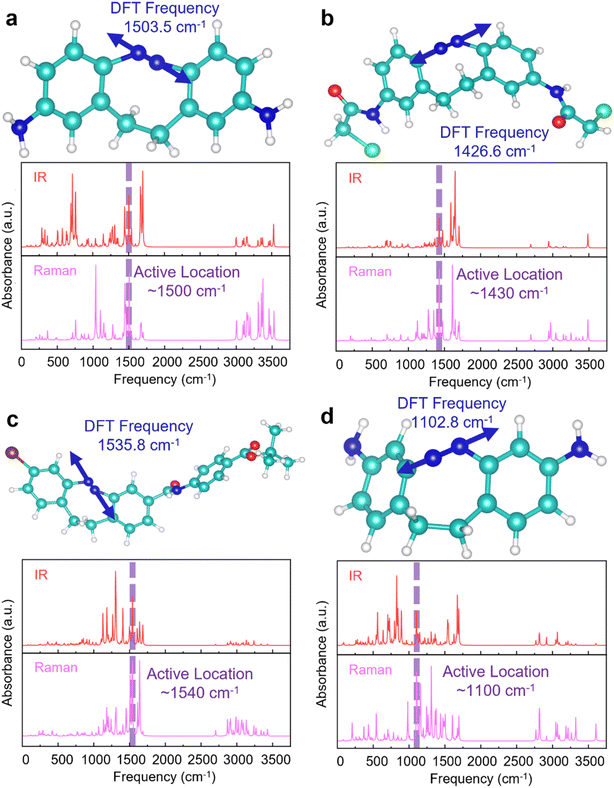
We examined the attention layer's weighting (Fig. S2–S5) of various regions of the simulated IR and Raman spectra to better understand its role in the ATT-CNN model. Note that this layer operates on ∼10 cm−1 segments of the simulated spectra due to the convolution and pooling that precedes it. The predominant DFT-calculated NN stretching frequency for C–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–C generally falls within the spectral region with the highest attention layer weight allocation, termed the active location (in cases where multiple NN stretches exist, the one with the largest amplitude of motion for the directly bonded nitrogens is considered the predominant NN stretch). Examples of the predominant NN stretching frequency coinciding with the active location are provided in Fig. 3a–d. Given the role of the C–N
N–C generally falls within the spectral region with the highest attention layer weight allocation, termed the active location (in cases where multiple NN stretches exist, the one with the largest amplitude of motion for the directly bonded nitrogens is considered the predominant NN stretch). Examples of the predominant NN stretching frequency coinciding with the active location are provided in Fig. 3a–d. Given the role of the C–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–C moiety in defining the dihedral angle, its NN stretching vibration is logically expected to be significant in angle prediction. Therefore, these examples underscore the utility of the attention layer's ability to pinpoint crucial spectral frequency regions.
N–C moiety in defining the dihedral angle, its NN stretching vibration is logically expected to be significant in angle prediction. Therefore, these examples underscore the utility of the attention layer's ability to pinpoint crucial spectral frequency regions.
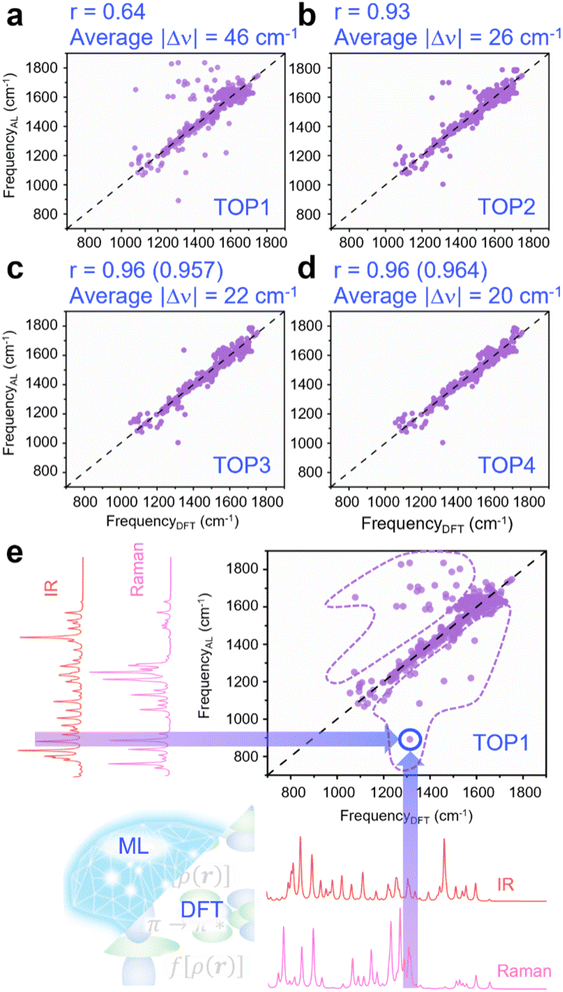
The relationship between the NN stretching frequency and the attention layer's weight distribution was confirmed through quantitative analysis (Fig. 4a–d). IR and Raman spectral regions were ranked based on the weights conferred by the attention layer: the highest-weighted region was named active location 1 (AL1), followed by active location 2 (AL2), active location 3 (AL3), and active location 4 (AL4). In our TOP1 analysis, the average frequency of AL1 was defined as FrequencyAL, and it was plotted against the DFT-derived NN stretching frequency for C–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–C (FrequencyDFT) (Fig. 4a). The TOP2 analysis involved designating the average frequency of AL1 and AL2 nearest to FrequencyDFT as FrequencyAL. Similarly, TOP3 analysis entailed choosing the closest frequency to FrequencyDFT from the averages of AL1, AL2, and AL3. Last, the TOP4 method extended this selection to include the nearest frequency from the combined averages of AL1 through AL4. In the TOP2, TOP3, and TOP4 analyses, FrequencyAL was then plotted against FrequencyDFT (Fig. 4b–d).
N–C (FrequencyDFT) (Fig. 4a). The TOP2 analysis involved designating the average frequency of AL1 and AL2 nearest to FrequencyDFT as FrequencyAL. Similarly, TOP3 analysis entailed choosing the closest frequency to FrequencyDFT from the averages of AL1, AL2, and AL3. Last, the TOP4 method extended this selection to include the nearest frequency from the combined averages of AL1 through AL4. In the TOP2, TOP3, and TOP4 analyses, FrequencyAL was then plotted against FrequencyDFT (Fig. 4b–d).
Although the correlation between the frequency of the NN stretch critical for dihedral angle prediction and the highest weighted spectral region frequency was only 0.64 (Fig. 4a), the correlation increased significantly when other highly weighted spectral regions were included (r > 0.9) (Fig. 4b–d). This is also reflected in the average absolute frequency difference between FrequencyAL and FrequencyDFT, which decreases from 46 cm−1 for TOP1 to 26 cm−1 for TOP2 and, as more highly weighted spectral regions are included, remains below 23 cm−1. These data confirm the link between the NN stretching frequency, vital for predicting the dihedral angle, and the spectral regions prioritized by the attention layer; this connection was previously illustrated with select examples in Fig. 3. The connection explains the enhanced accuracy of the ATT-CNN model that incorporates attention layers, as they effectively highlight the most informative spectral features for analysis.
Holistic spectral analysis enhances predictive performance
To investigate why the attention mechanism sometimes gives precedence to spectral regions distant from the NN stretching frequency, we examined data points lying far from the diagonal in Fig. 4a. In the TOP1 analysis, we identified 40 data points with large absolute differences between FrequencyAL and FrequencyDFT, categorizing them as “untreated data”; these are delineated by purple dashed lines in Fig. 4e. From the untreated data, we created five new datasets with AL1, AL2, AL3, AL4, and the NN stretching frequency spectral regions replaced by a sequence of 16 zeros. As a control, which we named “average SWZ” (Fig. S1), a sliding window method was used to systematically zero out 16 adjacent absorbance values across the spectral width spanning from 0 to 4000 cm−1.As illustrated in Fig. S1, this zeroing process was repeated at regular intervals of 4 cm−1 to ensure a comprehensive and unbiased assessment. We used each of these datasets to predict the bridged azobenzene derivative dihedral angle. The MAE of the average SWZ, 8.9°, is similar to that of the untreated data, 8.7°, suggesting that an indiscriminate zeroing out of 16 consecutive absorbance values generally has little impact on dihedral angle prediction. Notably, the maximum MAE of 11.2° was observed when the AL1 absorbance values were zeroed out, marking a 28% increase from the MAE of the untreated data. Zeroing out the NN stretch absorbance values led to an MAE of 9.9°, a comparatively smaller 14% increase in MAE.
These data indicate that, although the AL1 spectral region is far from the N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching frequency, it still plays a critical role in accurately predicting the dihedral angle. To further elucidate the physicochemical nature of these anomalous AL1 regions, we analyzed the 40 cases with the largest deviations (Fig. S6). The results show that the vibrational modes corresponding to these AL1 regions have clear chemical significance, such as C–N coupling, ring breathing vibrations, and ester C
N stretching frequency, it still plays a critical role in accurately predicting the dihedral angle. To further elucidate the physicochemical nature of these anomalous AL1 regions, we analyzed the 40 cases with the largest deviations (Fig. S6). The results show that the vibrational modes corresponding to these AL1 regions have clear chemical significance, such as C–N coupling, ring breathing vibrations, and ester C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, confirming that the attention mechanism can dynamically identify the most informative spectral features based on the molecular environment.
O stretching, confirming that the attention mechanism can dynamically identify the most informative spectral features based on the molecular environment.
Notably, although zeroing out the secondary attention regions (AL2–AL4) led to measurable increases in MAE (10.3°, 9.7°, and 9.4°, respectively), the predictive accuracy of the model remained largely intact. This demonstrates that ATT-CNN does not rigidly rely on a single spectral peak but is capable of capturing synergistic relationships between the primary feature (AL1) and other relevant spectral signals. These secondary features play an indispensable role in calibrating predictions, refining structural information, and providing necessary chemical context. Therefore, although AL1 contains the primary spectral–structure relationship, the inclusion of other attention-weighted regions remains essential for achieving high-precision predictions—this also reflects the model's effectiveness in integrating and utilizing complex spectral features.
The generalizability of our spectral analysis method
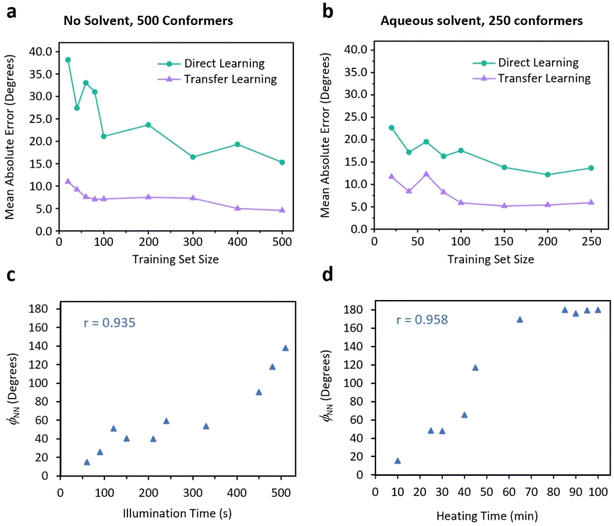
The generalizability of our spectral analysis method was evaluated using transfer learning. Our model, pretrained on a source domain comprising 4400 IR and Raman spectra from bridged azobenzene derivative conformers, was fine-tuned on two small-scale target domain datasets. The first dataset contained DFT-calculated spectra from 500 conformers of novel nitrogen-bridged diazocines (structurally distinct from the carbon-bridged diazocines in the training set). The second comprised spectra from 250 conformers in aqueous solution, with both datasets generated using ab initio molecular dynamics (AIMD). The first three layers of our model, which include convolutional and attention layers, perform general feature detection and were frozen in the fine-tune process while the remaining layers were adapted to the target domains. As a comparison, we directly trained ATT-CNN from scratch on the target domain datasets (all associated hyperparameters were kept consistent).The performance comparison in Fig. 5 show that transfer learning consistently surpasses direct learning. For the target domain consisting of 500 conformers (Fig. 5a), transfer learning on only 20 training examples resulted in a dihedral angle prediction with an MAE less than 12°, outperforming direct learning on 500 training samples. To validate ATT-CNN's applicability to experimental data, we used time-resolved IR and Raman spectroscopy to monitor dihedral angle changes in bridged azobenzene under 400 nm illumination and azobenzene under heating at 70 °C (Fig. S11, S13, S17 and S18). Since experimental data lacked explicit dihedral angle labels, we employed a semi-supervised learning approach: the model was first fine-tuned using a loss function that penalized deviations from the general increasing trend of predicted dihedral angles with illumination or heating time.
For bridged azobenzene, 19 experimental datasets were collected (9 for fine-tuning, 10 for testing); the refined model successfully captured the increasing trend in the testing set under illumination (Fig. 5c, r = 0.935). For azobenzene, 20 datasets were collected (10 for fine-tuning, 10 for testing); the refined model captured the increasing trend in the testing set under heating (Fig. 5d, r = 0.958). These results reflect the macroscopic structural evolution process based on ensemble-averaged spectra, verifying the model's capability to track changes in product distribution driven by light (or heat), and further demonstrate that ATT-CNN can generalize from theoretical training to experimental data with good robustness and generalizability.
Conclusions
In this work, we have established an integrated, agent-orchestrated, human–AI collaborative workflow for molecular structure elucidation, highlighted by the development and application of the ATT-CNN machine learning framework for IR and Raman spectral analysis. By applying this approach to bridged azobenzene derivatives, we achieved highly accurate predictions of the key C–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–C dihedral angle, with a mean absolute error as low as 5°. The attention mechanism within ATT-CNN reliably identifies and weights the most informative spectral features, enabling robust and interpretable structural assignments even amid common experimental variations or in the absence of traditional marker peaks.
N–C dihedral angle, with a mean absolute error as low as 5°. The attention mechanism within ATT-CNN reliably identifies and weights the most informative spectral features, enabling robust and interpretable structural assignments even amid common experimental variations or in the absence of traditional marker peaks.
Notably, the holistic learning approach of ATT-CNN allows it to extract structural information from complex spectral data without relying exclusively on any single feature, as demonstrated by successful predictions even when NN stretch regions were omitted. Further, transfer learning to new domains demonstrated the model's strong generalizability and adaptability, underscoring its potential as a broadly applicable tool for automated structure determination.
Altogether, our results lay the groundwork for a new approach of data-driven, interpretable, and human–AI collaborative workflows for real-time spectroscopic structure analysis. By harnessing both the accessibility and information-rich nature of vibrational spectroscopy, this strategy opens the door to accelerated mechanistic discovery in ultrafast dynamics, responsive materials, and beyond. Continued expansion of ATT-CNN's training and application to a wider chemical space will further enhance its power, paving the way for transformative advances in the automated interpretation of complex spectroscopic data across diverse chemical systems.
Finally, we acknowledge that the present workflow is grounded in ground-state simulations and steady-state spectroscopy. Although this approach does not capture the ultrafast excited-state dynamics underlying the initial photochemical step, it successfully establishes a robust mapping between readily accessible vibrational spectra and the key structural descriptor (the C–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–C dihedral angle) of the relaxed photoproducts. This enables the quantitative tracking of macroscopic isomerization progress and photostationary states, which is directly relevant for evaluating photoswitch performance and guiding molecular design. Future integration with time-resolved spectroscopy and nonadiabatic dynamics simulations could further extend the framework to probe the real-time photochemical trajectory.
N–C dihedral angle) of the relaxed photoproducts. This enables the quantitative tracking of macroscopic isomerization progress and photostationary states, which is directly relevant for evaluating photoswitch performance and guiding molecular design. Future integration with time-resolved spectroscopy and nonadiabatic dynamics simulations could further extend the framework to probe the real-time photochemical trajectory.
Author contributions
Yixi Shen, Ledu Wang, Yan Huang and Xiaolong Zhang contributed equally to this work. Jun Jiang, Linjiang Chen, Zhuoying Zhu and Yan Huang conceived and designed the study. Yixi Shen and Ledu Wang developed the code and performed the data analysis. Yixi Shen, Xiaolong Zhang and Jing He conducted the experiments. Yan Huang, Meng Huang and Huirong Li performed the simulations and calculations. Yang Wang contributed to data acquisition. Yixi Shen and Ledu Wang wrote the original draft. Linjiang Chen and Yan Huang reviewed and edited the manuscript. All authors read and approved the final manuscript.Conflicts of interest
The authors declare no competing interests.Data availability
The data supporting the findings of this study are available within the paper and its supplementary information (SI). Supplementary information: computational and data processing details, machine learning model architecture and parameter settings, attention weight distribution analysis, structural transformation and spectral analysis of bridged azobenzene and azobenzene under light and heating conditions, synthesis and characterization (1H NMR, MS, IR, Raman), LLM research dialogues, and supporting figures and tables. All code and scripts developed and used in this study are available at https://github.com/pic-ai-robotic-chemistry/azobenzenes-spectra. See DOI: https://doi.org/10.1039/d5sc08794e.Acknowledgements
The AI-driven experiments, simulations and model training were performed on the robotic AI-Scientist platform of Chinese Academy of Sciences (CAS). This work was financially supported by the Innovation Program for Quantum Science and Technology (2021ZD0303303), the National Key Research and Development Program of China (2018YFA0208603), the CAS Strategic Priority Research Program (Grant XDB0450302), the CAS Project for Young Scientists in Basic Research (YSBR-005), the National Natural Science Foundation of China (22573101, 22025304, 22033007, 22303091, 22201277), the Fundamental Research Funds for the Central Universities (WK2490250008) and the National Key Research and Development Program of China (2023YFA1508200). Z. Z and L. C. acknowledge the University of Science and Technology of China (USTC) Startup Programs for funding. The Hefei Advanced Computing Center is acknowledged for the provision of supercomputing services.References
- S. Hu, et al., Lighting Up Nonemissive Azobenzene Derivatives by Pressure, J. Am. Chem. Soc., 2024, 146, 28961–28972 CrossRef CAS PubMed.
- J. Gemen, et al., Disequilibrating azobenzenes by visible-light sensitization under confinement, Science, 2023, 381, 1357–1363 CrossRef CAS PubMed.
- S. Li, et al., Self-regulated non-reciprocal motions in single-material microstructures, Nature, 2022, 605, 76–83 CrossRef CAS PubMed.
- R. Lin, et al., Phenylazothiazoles as Visible-Light Photoswitches, J. Am. Chem. Soc., 2023, 145, 9072–9080 CrossRef CAS PubMed.
- M. C. Brand, et al., Photoresponsive Organic Cages-Computationally Inspired Discovery of Azobenzene-Derived Organic Cages, J. Am. Chem. Soc., 2024, 146, 30332–30339 CrossRef CAS.
- C. Chen, Y. Ji, H. Li, T. Song and H. Yu, Unusual photo-tunable mechanical transformation of azobenzene terminated aliphatic polycarbonate, Nat. Commun., 2025, 16, 2620 CrossRef CAS PubMed.
- H. Cheng, S. Zhang, J. Qi, X. Liang and J. Yoon, Advances in Application of Azobenzene as a Trigger in Biomedicine: Molecular Design and Spontaneous Assembly, Adv. Mater., 2021, 33, 2007290 CrossRef CAS.
- F. A. Jerca, V. V. Jerca and R. Hoogenboom, Advances and opportunities in the exciting world of azobenzenes, Nat. Rev. Chem., 2021, 6, 51–69 CrossRef PubMed.
- J. N. Martins, et al., Photoswitchable Calixarene Activators for Controlled Peptide Transport across Lipid Membranes, J. Am. Chem. Soc., 2023, 145, 13126–13133 CrossRef CAS PubMed.
- H. Shan, et al., Manipulating the Isomerization of a Tris-azobenzene Cage by Anion Binding, J. Am. Chem. Soc., 2025, 147, 14960–14965 CrossRef CAS PubMed.
- A. Ghosh, et al., Light-Powered Reversible Guest Release and Uptake from Zn4 L4 Capsules, J. Am. Chem. Soc., 2023, 145, 3828–3832 CrossRef CAS PubMed.
- F. Corrado, et al., Azobenzene-based optoelectronic transistors for neurohybrid building blocks, Nat. Commun., 2023, 14, 6760 CrossRef CAS PubMed.
- F. Cai, B. Yang, X. Lv, W. Feng and H. Yu, Mechanically mutable polymer enabled by light, Sci. Adv., 2022, 8, eabo1626 CrossRef CAS PubMed.
- M. Colaço, et al., Diazocines as Guests of Cucurbituril Macrocycles: Light-Responsive Binding and Supramolecular Catalysis of Thermal Isomerization, J. Am. Chem. Soc., 2025, 147, 734–745 CrossRef PubMed.
- P. Lentes, et al., Nitrogen Bridged Diazocines: Photochromes Switching within the Near-Infrared Region with High Quantum Yields in Organic Solvents and in Water, J. Am. Chem. Soc., 2019, 141, 13592–13600 CrossRef CAS PubMed.
- W. W. Paudler and A. G. Zeiler, Diazocine chemistry. VI. Aromaticity of 5,6-dihydrodibenzo[b,f][1,2]diazocine, J. Org. Chem., 1969, 34, 3237–3239 CrossRef CAS.
- R. Siewertsen, et al., Highly Efficient Reversible Z–E Photoisomerization of a Bridged Azobenzene with Visible Light through Resolved S1 (nπ*) Absorption Bands, J. Am. Chem. Soc., 2009, 131, 15594–15595 CrossRef CAS PubMed.
- P. Lentes, et al., Nitrogen Bridged Diazocines: Photochromes Switching within the Near-Infrared Region with High Quantum Yields in Organic Solvents and in Water, J. Am. Chem. Soc., 2019, 141, 13592–13600 CrossRef CAS PubMed.
- J. Ewert, et al., Photoswitchable Diazocine-Based Estrogen Receptor Agonists: Stabilization of the Active Form inside the Receptor, J. Am. Chem. Soc., 2022, 144, 15059–15071 CrossRef CAS PubMed.
- J. Ramos-Soriano, et al., Bridged Azobenzene Exhibits Fully Reversible Photocontrolled Binding to a G-Quadruplex DNA/Duplex Junction, JACS Au, 2025, 5c00532, DOI:10.1021/jacsau.5c00532.
- E. M. M. Tan, et al., Fast photodynamics of azobenzene probed by scanning excited-state potential energy surfaces using slow spectroscopy, Nat. Commun., 2015, 6, 5860 CrossRef.
- A. Nenov, et al., UV-Light-Induced Vibrational Coherences: The Key to Understand Kasha Rule Violation in trans -Azobenzene, J. Phys. Chem. Lett., 2018, 9, 1534–1541 CrossRef CAS PubMed.
- H. Zhou, et al., Bridged Azobenzene Enables Dynamic Control of Through-Space Charge Transfer for Photochemical Conversion, J. Phys. Chem. Lett., 2021, 12, 3868–3874 CrossRef CAS PubMed.
- R. Liang, First-Principles Nonadiabatic Dynamics Simulation of Azobenzene Photodynamics in Solutions, J. Chem. Theory Comput., 2021, 17, 3019–3030 CrossRef CAS.
- D. Keefer, et al., Imaging conical intersection dynamics during azobenzene photoisomerization by ultrafast X-ray diffraction, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2022037118 CrossRef CAS PubMed.
- D. R. Yarkony, Diabolical conical intersections, Rev. Mod. Phys., 1996, 68, 985–1013 CrossRef CAS.
- M. Abolhasani and E. Kumacheva, The rise of self-driving labs in chemical and materials sciences, Nat. Synth., 2023, 2, 483–492 CrossRef CAS.
- D. A. Boiko, R. MacKnight, B. Kline and G. Gomes, Autonomous chemical research with large language models, Nature, 2023, 624, 570–578 CrossRef CAS PubMed.
- T. Dai, et al., Autonomous mobile robots for exploratory synthetic chemistry, Nature, 2024, 635, 890–897 CrossRef PubMed.
- Y. Ruan, et al., An automatic end-to-end chemical synthesis development platform powered by large language models, Nat. Commun., 2024, 15, 10160 CrossRef CAS PubMed.
- T. Song, et al., A Multiagent-Driven Robotic AI Chemist Enabling Autonomous Chemical Research On Demand, J. Am. Chem. Soc., 2025, 147, 12534–12545 CrossRef CAS PubMed.
- G. Tom, et al., Self-Driving Laboratories for Chemistry and Materials Science, Chem. Rev., 2024, 124, 9633–9732 CrossRef CAS.
- D. Edge et al., From Local to Global: A Graph RAG Approach to Query-Focused Summarization, arXiv, 2024, preprint, arxiv.2404.16130, DOI:10.48550/ARXIV.2404.16130.
- DeepSeek-AI et al., DeepSeek-V3 Technical Report, arXiv, 2024, preprint, arxiv.2412.19437, DOI:10.48550/ARXIV.2412.19437.
- DeepSeek-AI et al., DeepSeek-R1: Incentivizing Reasoning Capability in LLMs via Reinforcement Learning, arXiv, 2025, preprint, arxiv.2501.12948, DOI:10.48550/ARXIV.2501.12948.
Footnote |
| † These authors contributed equally: Y. S., L. W., Y. H., and X. Z. |
| This journal is © The Royal Society of Chemistry 2026 |