Open Access Article
Open Access ArticleMachine learning-assisted screening of small-molecule drugs for suppressing protein aggregation and ROS generation based on ECL and CV dual-mode signals amplified by DNA
Jiaqi
Shi
a,
Wenxiu
Zhu
a,
Jingyu
Yan
a,
Cong
Xiao
a,
Huiqin
Yao
 b,
Lingchen
Meng
b,
Lingchen
Meng
 b,
Hongyun
Liu
b,
Hongyun
Liu
 *a and
Lanqun
Mao
*a and
Lanqun
Mao
 *a
*a
aKey Laboratory of Radiopharmaceuticals, Ministry of Education, College of Chemistry, Beijing Normal University, Beijing 100875, Peoples' Republic of China. E-mail: liuhongyun@bnu.edu.cn; lqmao@bnu.edu.cn
bSchool of Public Health, Key Laboratory of Environmental Factors and Chronic Disease Control, Ningxia Medical University, Yinchuan 750004, People's Republic of China
First published on 2nd February 2026
Abstract
Screening of small-molecule drugs to suppress both protein aggregation and reactive oxygen species (ROS) generation is critical for developing therapies for neurodegenerative diseases (NDs). However, existing methods are limited to characterizing only a single pathological feature (either protein aggregation or ROS generation) in a single measurement. Herein, taking α-synuclein (α-Syn) as the template protein, we developed a dual-mode electrochemical sensing platform for concurrently monitoring protein aggregation and ROS generation characteristics. A gold electrode functionalized with α-Syn via self-assembled monolayers (SAMs) was constructed as the sensing platform, realizing both ordered α-Syn immobilization and monitoring of metal ion (e.g., Cu(II))-driven aggregation. This was accomplished by synchronously recording the electrochemiluminescence (ECL) and cyclic voltammetry (CV) dual responses of the tris(2,2′-bipyridine) ruthenium(II) (Ru(bpy)32+) reporter in a single integrated assay. The catalysis of DNA oxidation by Ru(bpy)32+ enables the amplification of ECL and CV dual-mode signals, which increased the detection sensitivity for both aggregation and ROS generation accompanied by the α-Syn − Cu(II) complex. Machine learning algorithms were then utilized to analyze ECL and CV responses of small molecules with known drug effects. This analysis culminated in the development of a linear discriminant analysis (LDA) screening model, which enabled the assessment of drug efficacy against the two pathological features. The predictive capability of the screening model was verified through transmission electron microscopy (TEM), cell viability and intracellular aggregation studies. This model was further successfully applied to assess two previously unexplored small molecules: diethylenetriaminepentaacetic dianhydride (DTPA) and deferiprone. Collectively, this dual-mode sensing platform, integrating DNA-amplified monitoring of protein aggregation and ROS generation, enables the robust establishment of a machine learning-assisted small-molecule drug screening model, offering a novel approach for the in vitro characterization of protein-related pathological features.
Introduction
Protein misfolding and aggregation represent central pathological mechanisms underlying numerous neurodegenerative diseases (NDs). For example, the misfolding and aggregation of α-synuclein (α-Syn) have been identified as key drivers of synucleinopathies, including Parkinson's disease (PD) and multiple system atrophy (MSA).1–3 The pathological consequences of protein aggregation extend beyond direct cellular toxicity to the disruption of key biological pathways, including autophagy–lysosomal systems and mitochondrial function, thereby exacerbating neurodegeneration.3 Emerging evidence has demonstrated that α-Syn aggregation triggers endoplasmic reticulum stress and oxidative damage,3,4 most probably from reactive oxygen species (ROS) generation through redox-active metals,5 metallo-enzymes,6 or nonenzymatic redox activity7 after proteins assemble. Since protein aggregation and ROS generation processes influence each other reciprocally,5–7 concurrent monitoring of their levels would provide a more comprehensive evaluation of the pathological state and more precise screening of drugs, especially small-molecule drugs for the prevention and treatment of NDs.4,8,9Currently, the methods developed for studying protein–small-molecule interactions include fluorescence spectroscopy,8,10 circular dichroism,11 transmission electron microscopy,12 surface plasmon resonance,13 isothermal titration calorimetry,14 electrochemical techniques15–17 and computational simulations.18 However, these techniques, which typically rely on single-mode signals, still exhibit notable limitations, such as a tendency toward false-positive results, prolonged detection periods, a fundamental challenge in balancing analytical accuracy with detection efficiency, and crucially, the difficulty in simultaneously detecting protein aggregation and ROS production.10 Thus, developing a multimodal detection platform capable of rapidly resolving the dynamic interplay between small molecules and protein aggregates/ROS generation is paramount for advancing the precision diagnostics and therapeutic development of NDs.
Electrochemiluminescence (ECL) is an analytical technique that combines electrochemical methods with chemiluminescence, integrating the advantages of both electrochemical and chemiluminescence approaches, such as high sensitivity, a low background signal, and a wide linear range. It has been widely applied in the field of biosensing.19,20 For example, ECL technology has been employed for the highly sensitive detection of AD-related β-amyloid (Aβ) peptides with aptasensors.21 The integration of ECL with simultaneously acquired cyclic voltammetry (CV) signals was performed for the detection of nucleic acids,22 antibiotics,23 and glucose,24 driving advances in intelligent sensing and biocomputing.25 This dual-mode strategy not only increases detection sensitivity but also enables multiplexed target analysis through potentially resolved signals. However, to date, the dynamic current response of CV and the static optical signals of ECL have not been combined to study small molecules associated with protein aggregation in complex biological samples.
Owing to its inherent biocompatibility and redox activity, DNA has emerged as a versatile biomaterial for signal amplification in ECL and CV assays.24,26–28 Specifically, guanine (G) residues within the DNA strand can be catalytically oxidized by ruthenium bipyridine (Ru(bpy)32+), yielding significant enhancements in both ECL and CV signals.26 During this catalytic process, CV and ECL signals sensitively reflect the DNA damage induced by ROS, enabling real-time monitoring of the ROS generated via Fenton reactions.27,29,30 This dynamic responsiveness has inspired the development of smart biosensors31 and further enabled the creation of a dual-mode logic system for biocomputing and information encryption using the ECL intensity and CV peak current.32 To the best of our knowledge, no prior studies have explored the use of natural DNA for electrochemical signal enhancement in the context of protein aggregation and small-molecule interactions.
In this study, ordered polypeptide interfaces were immobilized on a gold electrode using self-assembled monolayers (SAMs) to construct a dual-mode biosensor with natural DNA as a sensitizer for screening of small-molecule drugs. Taking α-Syn as an example, the pathological phenomenon of protein aggregation promoted by Cu(II) and the corresponding process of ROS production were monitored by simultaneously acquiring ECL and CV signals amplified by DNA in solution. Furthermore, linear discriminant analysis (LDA) was applied to analyze the dual signals, establishing a screening model for successfully categorizing 9 small molecules into 4 categories according to their known medicinal effects. The model was validated by TEM, UV spectroscopy and cell toxicity. Furthermore, an mCherry-α-Syn cell model was generated through plasmid transfection to establish physiological relevance. The established screening model was ultimately applied to predict and validate the pharmacological effects of two small-molecule drug candidates, diethylenetriaminepentaacetic dianhydride (DTPA) and deferiprone, providing a new technical platform for the early diagnosis of NDs and drug screening.
Experimental section
Preparation of α-Syn-modified electrodes
Prior to use, the gold electrodes were sequentially polished with alumina powders of 1.0, 0.3, and 0.05 µm, followed by sequential ultrasonic treatment in deionized water, ethanol, and deionized water again for 5 min each. The electrodes were subsequently immersed in “piranha solution” (prepared by mixing concentrated sulfuric acid and hydrogen peroxide in a 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 volume ratio) for 10 min to remove organic contaminants and increase hydrophilicity. Caution: piranha solution should be handled with extreme care, and only small volumes should be prepared at any time. Finally, clean electrodes were obtained by ultrasonication in deionized water and ethanol for 3 min each. Then the treated Au electrodes were incubated with 25 µL of an ethanol solution containing 65 µM 11-mercaptoundecanoic acid (MUA) and 170 µM 6-mercapto-1-hexanol (MCH) for 8 h to functionalize the surface with a near-monolayer of carboxyl groups (see the SI).33 This was followed by activation of the carboxyl groups with 400 mM N-(3-dimethylaminopropyl)-N′-ethylcarbodiimide hydrochloride (EDC) and 100 mM N-hydroxysuccinimide (NHS) for 20 min. Subsequently, 25 µL of a 50 µM α-Syn solution was applied to the electrode surface and incubated for 2 h. The unreacted sites were blocked with 0.5 M ethanolamine (EA) for 30 min, resulting in α-Syn-modified electrodes. The electrodes were further incubated with a 50 µM Cu(II) solution for 2 h to obtain α-Syn − Cu(II)-modified electrodes, which were used to simulate the effects of drugs on the metal-induced alterations of α-Syn (Scheme 1).
1 volume ratio) for 10 min to remove organic contaminants and increase hydrophilicity. Caution: piranha solution should be handled with extreme care, and only small volumes should be prepared at any time. Finally, clean electrodes were obtained by ultrasonication in deionized water and ethanol for 3 min each. Then the treated Au electrodes were incubated with 25 µL of an ethanol solution containing 65 µM 11-mercaptoundecanoic acid (MUA) and 170 µM 6-mercapto-1-hexanol (MCH) for 8 h to functionalize the surface with a near-monolayer of carboxyl groups (see the SI).33 This was followed by activation of the carboxyl groups with 400 mM N-(3-dimethylaminopropyl)-N′-ethylcarbodiimide hydrochloride (EDC) and 100 mM N-hydroxysuccinimide (NHS) for 20 min. Subsequently, 25 µL of a 50 µM α-Syn solution was applied to the electrode surface and incubated for 2 h. The unreacted sites were blocked with 0.5 M ethanolamine (EA) for 30 min, resulting in α-Syn-modified electrodes. The electrodes were further incubated with a 50 µM Cu(II) solution for 2 h to obtain α-Syn − Cu(II)-modified electrodes, which were used to simulate the effects of drugs on the metal-induced alterations of α-Syn (Scheme 1).
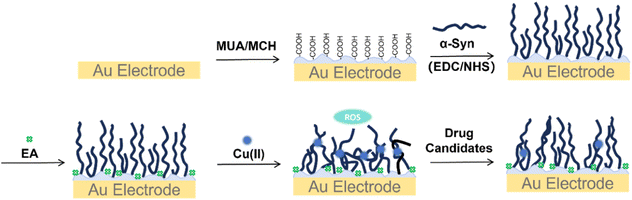 | ||
| Scheme 1 Schematic diagram of the step-by-step modification of the surface of a gold electrode with α-Syn via covalent bonding with the self-assembled monolayer of MUA. | ||
Results and discussion
Characterization of α-Syn-modified electrodes
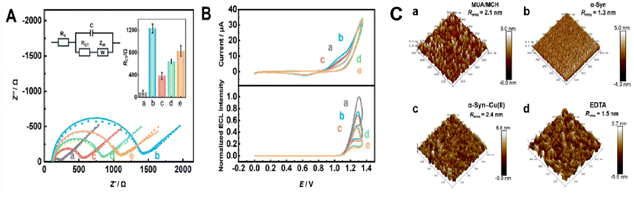
The successful assembly of α-Syn on the gold electrode surface was first characterized using electrochemical impedance spectroscopy (EIS). The changes in the state of the gold electrode surface during the modification process were reflected by the charge transfer resistance (RCT).34 Using Fe(CN)63−/4− as an electroactive probe, the electrodes at different stages of the modification process were tested via EIS at pH 7.4 (Fig. 1A). Owing to the hydrophobicity of the long chains of MUA and MCH and the electrostatic repulsion between –COO− and Fe(CN)63−/4−, the MUA/MCH layer hinders the diffusion of the probe, leading to an increase in RCT values (Fig. 1A, curve b). After EDC/NHS treatment, the negatively charged –COO− residue at the end of MUA was replaced by succinimide ester, partially eliminating the electrostatic repulsion and thereby increasing the redox current in the CV curve and reducing the RCT value (Fig. 1A, curve c).35 When the poorly conductive peptide α-Syn was further covalently linked to the MUA surface activated by EDC/NHS, the RCT value increased (Fig. 1A, curve d). The use of ethanolamine (EA) to cover other active sites resulted in a further increase in the RCT values (Fig. 1A, curve e).During the modification of the electrode, both the CV and ECL signals, with Ru(bpy)32+ as the probe (Fig. 1B), demonstrated similar trends to the results obtained by EIS (Fig. 1A), except for the modification process of MUA/MCH. Owing to the electrostatic attraction between the –COO− residue on the MUA/MCH layer and Ru(bpy)32+ in the solution, this step slightly increased the CV signal but did not significantly affect the ECL intensity (E) (Fig. 1B, curves a and b).
In addition, atomic force microscopy (AFM) was used to characterize the successful assembly of α-Syn-modified electrodes. MUA/MCH were evenly distributed on the Au-coated silicon wafer surface, with a root mean square roughness (Rrms) of 2.1 nm (Fig. 1C, panel a), which decreased to 1.3 nm upon α-Syn immobilization (Fig. 1C, panel b), consistent with the literature for the immobilization of flexible-conformation proteins.36 Subsequent treatment with Cu(II) induced α-Syn aggregation on the electrode surface, leading to the formation of irregular protein aggregates and an increase in Rrms to 2.4 nm (Fig. 1C, panel c). Upon addition of EDTA, the chelation of Cu(II) partially reversed the aggregation, resulting in a reduced Rrms of 1.5 nm (Fig. 1C, panel d). The AFM topography results visually demonstrated the dynamic Cu(II)-mediated aggregation and EDTA induced disaggregation process. All these results suggested the successful immobilization of α-Syn on the electrode surface, verifying the feasibility of the modification scheme. In the following experiments, the prepared MUA/MCH/α-Syn/EA films at the Au electrodes are termed α-Syn-modified electrodes for concision.
DNA amplification behaviors shown by CV and ECL signals
After modification, both the CV and ECL signals of Ru(bpy)32+ in the solution for the α-Syn-modified electrode decreased (Fig. 1B curve e). To achieve moderate and observable signals, natural salmon DNA with intrinsic electrochemical inertness was added to the electrolyte solution to increase the strength of both the ECL and the CV.26,37–39 In the absence of DNA, Ru(bpy)32+ demonstrated an ECL signal at 1.2 V40,41 accompanying the corresponding electrochemical reaction (Fig. S1A and S1B, curve a and Fig. S1C reactions 1 ∼ 4). Upon the addition of 0.3 mg·mL−1 DNA to the electrolyte after optimization, the oxidation peak current (Ipa) of Ru(bpy)32+ at 1.05 V![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 42 for the α-Syn electrodes clearly increased (Fig. S1A, curve b). A distinct increase was also found for the ECL signal of Ru(bpy)32+(Fig. S1B, curve b). These results were characteristic of the electrocatalytic behavior of Ru(bpy)32+ in the presence of DNA.26,31,32
42 for the α-Syn electrodes clearly increased (Fig. S1A, curve b). A distinct increase was also found for the ECL signal of Ru(bpy)32+(Fig. S1B, curve b). These results were characteristic of the electrocatalytic behavior of Ru(bpy)32+ in the presence of DNA.26,31,32
Ru(III) in ruthenium(III) tris(bipyridine) (Ru(bpy)33+), which has strong oxidizing properties, can react with guanine (G) in DNA and further mediate CV/ECL signal amplification.26,27,29,30 The principle was as follows:32 Ru(bpy)32+ is first electrochemically oxidized into Ru(bpy)33+ at the electrodes (Fig. S1C reaction 1). The product Ru(bpy)33+ then oxidizes G groups of DNA (DNA-G) to DNA-GOx and returns to the original form Ru(bpy)32+, completing an electrocatalytic cycle (Fig. S1C reaction 5). Ru(bpy)33+ might further oxidize DNA-GOx and become excited-state Ru(bpy)32+* (Fig. S1C reaction 6), which emits luminescence at 610 nm and decays to the ground state of Ru(bpy)32+, forming a luminescent catalytic cycle (Fig. S1C reaction 4).31,39 With DNA in the solution, the Ipa value of CV was amplified by approximately 1.7 times, and the ECL intensity was amplified by nearly 1.3 times (Fig. S1), similar to what was reported in the literature.32 Thus, in the following experiments, 0.3 mg·mL−1 DNA was used in all the tests to increase the detection sensitivity.
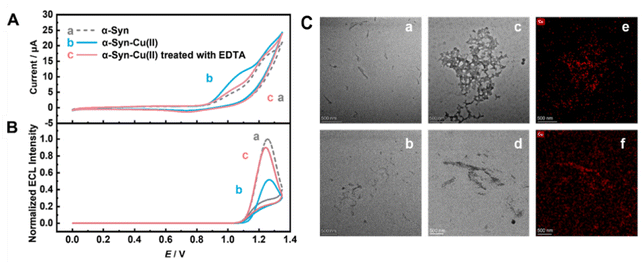
Aggregation behaviors of α-Syn interfaces with Cu(II)
Trace metal ions are naturally present in the brain; however, the dysregulation of metal homeostasis can affect the protein structure and oxidative stress, leading to severe neurodegenerative diseases.43 Cu(II) is a quintessential example, as its overabundance can exacerbate the risk of PD by inducing conformational changes in α-Syn.44–46 In this study, Cu(II) was selected as a factor to promote α-Syn aggregation on the electrode surface under physiological conditions and the production of ROS, aiming to shorten the time required to study the interaction between α-Syn interfaces and small molecules. After incubation with Cu(II), opposite changes in the signals of ECL and CV were observed (Fig. 2A and B). This section provides a detailed discussion of the aggregation behavior of α-Syn under Cu(II) induction, as reflected in ECL and CV signals, respectively.Furthermore, the extent to which Cu(II) promoted the aggregation of α-Syn on the electrode surface was also dependent on the concentration of Cu(II) (cCu(II)). The ECL intensity gradually decreased with increasing cCu(II) content after incubation of the α-Syn-modified electrodes in Cu(II) for 2 h (Fig. S2B and S2D). However, when cCu(II) reached 50 µM, further increases in the concentration did not result in a significant change in signal intensity. This might be because of the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding ratio between α-Syn and Cu(II).49 Thus, during the following experiments, α-Syn-modified electrodes were incubated in a 50 µM Cu(II) solution for 2 h to prepare α-Syn − Cu(II) interfaces.
1 binding ratio between α-Syn and Cu(II).49 Thus, during the following experiments, α-Syn-modified electrodes were incubated in a 50 µM Cu(II) solution for 2 h to prepare α-Syn − Cu(II) interfaces.
To further validate the reliability of using the ECL intensity to reflect α-Syn aggregation on electrode surfaces under the influence of Cu(II), we employed TEM to observe the morphological changes in α-Syn before and after the Cu(II) interaction. Compared with monomeric α-Syn before Cu(II) treatment (Fig. 2C, panel a) and its self-aggregation in the blank control for 24 h (Fig. 2C, panel b), Cu(II)-treated α-Syn formed more prominent irregular, amorphous large aggregates, namely, α-Syn − Cu(II) complexes (Fig. 2C, panel c). Corresponding high-angle annular dark-field scanning transmission electron microscopy (HAADF-STEM) images revealed distinct bright spots indicating copper distribution within the aggregates (Fig. 2C, panel e), demonstrating significantly elevated copper concentrations in these aggregates. This confirmed the participation of copper in α-Syn aggregation and verified that variations in the ECL intensity could reflect α-Syn aggregation states under the influence of Cu(II).
To investigate the specific cause of the opposite trends in the CV and ECL signals during the aggregation process of α-Syn, different concentrations of H2O2 were introduced into the electrolyte. With increasing concentrations of H2O2, the CV curves revealed that Ipa of Ru(bpy)32+ at approximately 1.05 V increased for the α-Syn-modified electrode (Fig. S4A and S4C), whereas the ECL intensity decreased gradually (Fig. S4B and S4D). This result indicated that both the CV and ECL signals of the α-Syn − Cu(II)-modified electrodes demonstrated trends analogous to those of the α-Syn-modified electrodes without Cu(II) but with H2O2 present in the test solution. In the ECL system, O2 or radicals resulting from the decomposition of H2O2 may function as a quencher for the excited-state Ru(bpy)32+*, thereby attenuating the ECL intensity50,51 and producing Ru(bpy)32+, which increases the Ipa of CVs.50
According to the above results, we reasoned that when α-Syn was incubated with Cu(II), the resulting α-Syn − Cu(II) complexes on the electrode surface might have produced oxidizing substances with functions analogous to those of H2O2, i.e., ROS.
To test our hypothesis, we employed the same α-Syn-modified electrode as the working electrode in a blank buffer solution to measure peak current changes before and after Cu(II) incubation via CV. Compared with the α-Syn-modified electrode alone, the α-Syn − Cu(II)-modified electrode exhibited a pair of quasi-reversible CV peaks between 0 and 0.2 V vs. Ag/AgCl in the air-equilibrated pH 7.4 buffer (Fig. S5A). According to previous studies,52,53 the peak at 0.08 V vs. Ag/AgCl was attributed to copper bound to α-Syn, thus representing the “α-Syn − Cu(II)/α-Syn − Cu(I)” redox couple. Since the potential of Syn − Cu(I) oxidation to α-Syn − Cu(II) was slightly lower than that of O2 reduction to H2O2 (O2/H2O2, 0.10 V vs. Ag/AgCl) (Fig. S5C),54–56 and the reaction between α-Syn − Cu(I) and dissolved oxygen to generate ROS was thermodynamically favorable.15,52,53
Moreover, under physiological conditions, various reducing agents or antioxidants with redox potentials lower than that of α-Syn − Cu(II), such as extracellular ascorbic acid (AA), intracellular glutathione (GSH),57 and membrane-bound redox species associated with the mitochondrial electron transport chain (ETC), such as cytochrome c, can reduce α-Syn − Cu(II) to α-Syn − Cu(I), leading to oxidative stress or ROS production.58,59 Changes in the concentrations of these substances often accompany the pathological progression of PD.60
Therefore, these experiments not only demonstrated that Cu(II) could induce α-Syn aggregation at the interface but also revealed that the α-Syn − Cu(II) interfaces could generate ROS with effects similar to those of H2O2in vitro, resulting in increased CV signals and decreased ECL signals (Fig. 2). This viewpoint was further corroborated by the CV experiments (Fig. S5B and S6). Specifically, the CV signal at 1.05 V was significantly diminished when the voltage in the test did not reach the redox potential of Cu (Fig. S6, curve c), or with N2-purged in the solution (Fig. S5B), or when the amount of dissolved O2 in the solution decreased (Fig. S6, curve b).
ROS are usually generated during protein aggregation, especially when aggregation is induced by metal ions.58,59 To verify that the generation of ROS could be monitored using our designed interfaces, we used edaravone (PMP), a typical radical scavenger that can remove oxygen-containing ROS (such as OH˙, O2−˙, and H2O2)61 and does not interfere with the electrochemical signals under our experimental conditions.62 H2O2 was first used to test the elimination function of PMP. After the introduction of PMP under the same conditions as those of the α-Syn-modified electrode without Cu(II), the Ipa of the CVs for Ru(bpy)32+ decreased, but the ECL intensity increased (Fig. S7). That is, PMP eventually compensated for the consequences of the addition of H2O2 to both the ECL and CV signals, suggesting that in the presence of PMP, ROS could be eliminated to a certain extent. After optimization, 0.1 mM PMP was used to characterize the production of ROS and further exclude the effects of ROS during protein aggregation under the influence of metal ions.
Disaggregation behavior of α-Syn − Cu(II) modified electrodes with EDTA
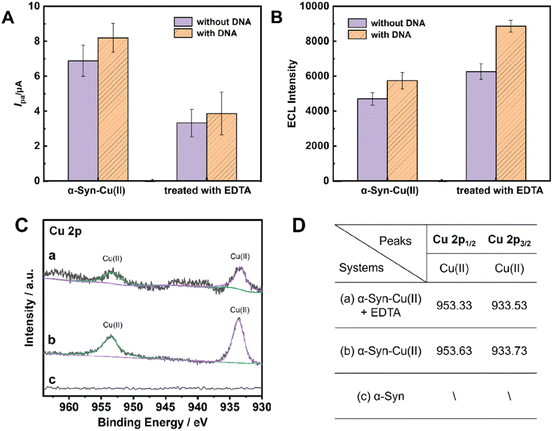
The addition of ethylenediaminetetraacetic acid (EDTA),63,64 a small-molecule drug widely recognized for mitigating protein-Cu(II) aggregation through chelation, resulted in aggregate dispersion and size reduction (Fig. 2C, panel e). Compared with that of the α-Syn − Cu(II) aggregates, the corresponding HAADF-STEM images revealed an increased background copper content (Fig. 2C, panel f), suggesting that EDTA effectively hindered the formation of some α-Syn − Cu(II) aggregates. This process likely reduced the steric hindrance of the aggregates on the electrode surfaces, which was subsequently reflected in the recovery of the electrochemical signals (Fig. 2, curve c, Fig. 3A and B).The DNA introduced in the above description not only enhanced the electrochemical signal at the protein-modified electrode interfaces, facilitating observation of the drug-induced disaggregation process, but also amplified the dual-mode signal changes caused by the chelation of Cu(II) by small-molecule candidates (e.g., EDTA) (Fig. 3A and B). Specifically, the reduction in the CV signal was amplified by 1.2 times compared to that without DNA (Fig. 3A), whereas the increase in the ECL signal was amplified by 1.9 times (Fig. 3B).
X-ray photoelectron spectroscopy (XPS) analysis revealed that after EDTA treatment (Fig. 3C, curve a), the characteristic Cu(II) peak at 933.53 eV significantly decreased, indicating that EDTA removes some Cu(II) from α-Syn − Cu(II) aggregates by forming a spatially stable Cu-EDTA complex.65,66 The XPS data support our earlier inference on the basis of CV and ECL observations (Fig. 2) that certain small molecules could counteract the influence of Cu(II) on α-Syn aggregation.
Thus, the protein interfaces we constructed could be effectively used to monitor protein aggregation/disaggregation and the production/removal of ROS on the basis of ECL and CV signals.
Changes in the ECL and CV signals of α-Syn − Cu(II)-modified electrodes under small-molecule treatment
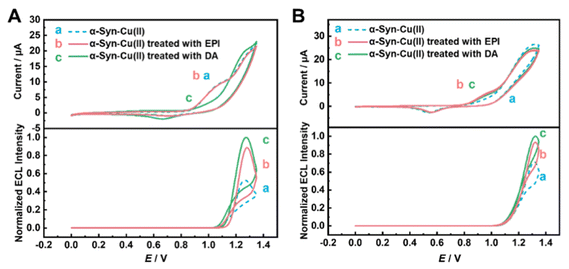
To investigate the effects of potential small-molecule drugs with disaggregation or ROS removal properties on the α-Syn − Cu(II)-modified electrode, simultaneous CV and ECL tests were conducted (Fig. 4). Epinephrine (EPI) and dopamine (DA) are typical α-Syn disaggregators, and their effects on α-Syn disaggregation have been extensively studied.67–71 This work used EPI and DA as small-molecule drug templates to study the alleviation of α-Syn − Cu(II) aggregation and ROS generation processes by monitoring ECL and CV signals.After 2 h of incubation with EPI (Fig. 4A, top, curve b), the Ipa of the CV for the α-Syn − Cu(II)-modified electrode was very similar to that without EPI treatment (Fig. 4A, top, curve a) but still significantly greater than that without Cu(II) (Fig. 2A, curve a). In contrast, the ECL results showed that after EPI treatment, the ECL signal of the α-Syn − Cu(II) electrode clearly increased (Fig. 4A, bottom, curve b). The results revealed that EPI could partially restore the changes in the ECL signal caused by Cu(II) but had no significant effect on the CV Ipa, suggesting that EPI could disaggregate protein aggregates but not remove ROS. If EPI could remove ROS, the corresponding results should demonstrate the restoration of the ECL signal but a decrease in Ipa. Based on the previous results (Fig. 2 and S7), Cu(II) might induce α-Syn aggregation while simultaneously generating ROS. The increase in the CV signal due to ROS generation appeared to outweigh the decrease caused by aggregation, leading to an overall increase in the CV signal (Fig. 2A, curves a and b). Therefore, if ROS removal indeed plays a significant role, the CV signals should consequently decrease.
To support our above inferences, the signal changes were recorded again in the presence of PMP, where the effects of ROS were eliminated (Fig. 4B). After EPI treatment, the ECL intensity of the α-Syn − Cu(II)-modified electrode significantly recovered, accompanied by a slight increase in the Ipa of the CV signal (Fig. 4B, curve b). Here, we suggest that EPI alleviated the aggregation of α-Syn − Cu(II), increasing the extent of the structure of α-Syn − Cu(II) on the electrode surface, thereby allowing Ru(bpy)32+ to reach the electrode surface more easily and react. This led to the recovery of both the Ipa and ECL intensities, demonstrating the efficacy of EPI in disaggregating proteins on the electrode surface, which was consistent with previous reports.71
Similarly, when DA was used instead of EPI to treat the α-Syn − Cu(II)-modified electrode, the Ipa of CV significantly decreased, whereas the ECL intensity dramatically recovered (Fig. 4A, curves a and c). However, in the presence of PMP (Fig. 4B, curve c), the effect of DA on the signals of the α-Syn − Cu(II)-modified electrode was similar to that of EPI. This suggests that, in addition to having a similar disaggregation effect to that of EPI, DA might also have ROS removal properties related to ROS scavenging, which aligns with previous reports.67,71,72
These experimental results demonstrated that monitoring ECL and CV signal changes at α-Syn − Cu(II)-modified electrodes before and after small-molecule treatment enabled the determination of whether a small molecule alleviates α-Syn − Cu(II) aggregation and/or scavenged ROS. Therefore, by analyzing the dual CV and ECL signal dynamics of α-Syn − Cu(II)-modified electrodes upon treatment with candidate small-molecule drugs, their potential efficacy in mitigating both α-Syn − Cu(II) aggregation and ROS generation could be systematically evaluated.
Construction of a small-molecule drug efficacy screening model based on LDA
The above experimental results also demonstrated that the dual CV and ECL signals obtained simultaneously from the α-Syn-modified electrode were influenced by both the cross-effects of protein aggregation and the oxidative stress induced by Cu(II) in α-Syn. Specifically, upon protein aggregation, both the ECL signal and the CV oxidation peak current decreased synchronously. In contrast, ROS caused opposing trends in the ECL and CV signals: ECL intensity diminishes, while the CV Ipa increases.To quantify these relationships, linear discriminant analysis (LDA), a robust machine learning technique,73 is proposed in this work. LDA could reveal hidden correlations between electrochemical signals and protein aggregation/ROS generation occurring at the α-Syn − Cu(II) interfaces through feature weighting and dimensionality reduction visualization and further enable the development of a predictive screening model for PD-targeting small molecules.
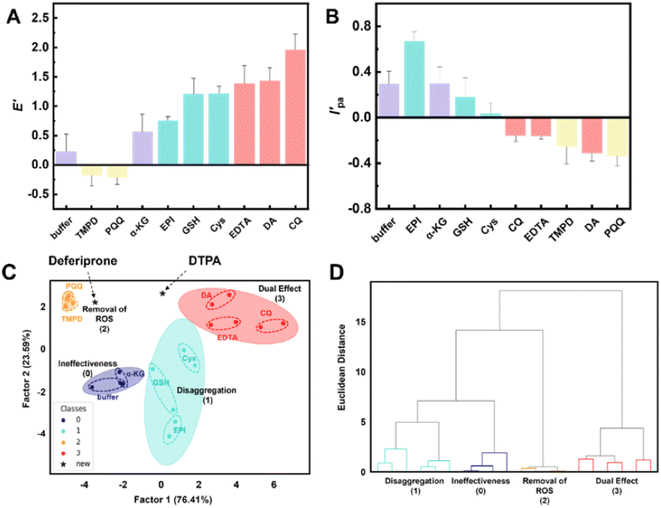
Initially, a calibration set was established by selecting 9 small molecules reported to have depolymerization activity and/or antioxidant activity, or neither (Table S1). Using the same method as described in the above experiments, tests were conducted and the corresponding ECL/CV signals were obtained simultaneously (Fig. 5A and B). The rate of change in the ECL intensity (E′) before and after drug treatment was used to measure the degree of change in the ECL intensity, which was defined as (E1 − E0)/E0. E0 and E1 correspond to the ECL signal intensity (E) of the α-Syn − Cu(II)-modified electrode before and after coincubation with the test drugs for 2 h, respectively. Similarly, the change rate of the oxidation peak current intensity (I'pa) of the CV curve at 1.05 V was used to measure the degree of change, defined as (Ipa1 − Ipa0)/Ipa0. Ipa0 and Ipa1 correspond to the oxidation peak current intensity (Ipa) values of the α-Syn − Cu(II)-modified electrode before and after coincubation with the test drug for 2 h, respectively.
Using E′ and I′pa as inputs, LDA was employed to perform classification calculations through different linear combinations, yielding two discriminant factors: Factor 1 and Factor 2 (Table S2). These factors represent discriminant directions (dimensions) obtained from various linear combinations of E′ and I′pa with maximal intergroup separation. Factor 1 and Factor 2 accounted for 76.41% and 23.59% of the contribution, respectively, with their combined contributions exceeding 90%. These findings indicate that these two factors are key discriminant elements in the screening model that capture nearly all the discriminant information in the data and are highly representative of its classification features.
The receiver operating characteristic (ROC) curve was further used to evaluate the classification performance of the LDA algorithm. The ROC curve demonstrated an ideal diagnostic outcome, with the true positive rate (TPR) = 1, the false positive rate (FPR), and the area under the curve (AUC) = 1 (Fig. S8). Therefore, for this drug screening model, two dimensions were sufficient to effectively distinguish between different groups without relying on additional dimensions.
Using the two discriminant factors as the horizontal and vertical coordinates, a two-dimensional scatter plot of the LDA classification results was generated (Fig. 5C). Nine small molecules were classified into 4 groups with maximum intergroup separation, where 2–3 small-molecule drugs (each represented by two replicate experimental data points) in each group were clustered together with the minimum intragroup distance after LDA calculation. These groups corresponded to 4 different drug efficacies: ineffective (0), disaggregation (1), removal of ROS (2), and dual effect (3).
Additionally, in Fig. 5C, the cluster for “removal of ROS (2)” exhibited a greater distance from the other groups, suggesting that the model had greater discriminative power for mitigating ROS drugs. In summary, LDA effectively distinguished among the 4 types of drug efficacy, highlighting its utility in the ECL/CV dual-signal platform established in this study for screening and classifying drugs that alleviate protein disaggregation and/or suppress ROS generation.
Hierarchical cluster analysis (HCA) is an unsupervised pattern recognition clustering method that constructs hierarchical clustering structures on the basis of Euclidean distance similarity features.74 As illustrated in the HCA dendrogram (Fig. 5D), small molecules with shorter Euclidean distances clustered together, and the 4 distinct small-molecule drug effects were clearly differentiated without any overlap. These results demonstrate that the HCA-based drug screening platform has excellent discriminative ability.
Primary molecular-level validation of the drug screening results
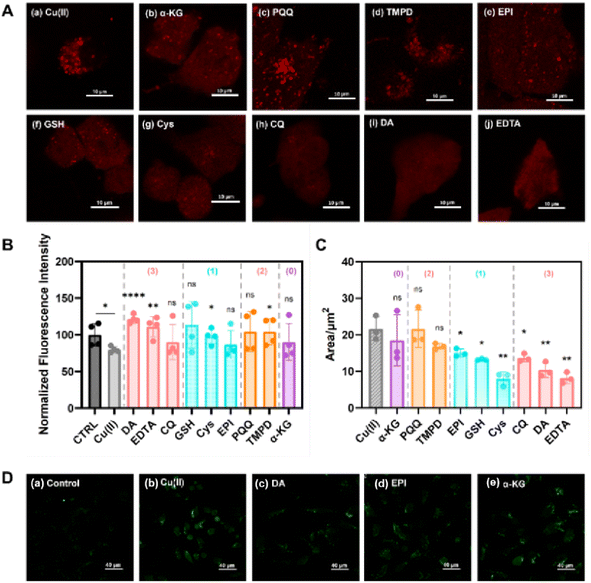
To evaluate the reliability of the drug efficacy classification results from our LDA-based screening model, several representative small-molecule drugs were chosen for direct TEM observation of their mitigating effects on α-Syn − Cu(II) aggregate formation.When α-Syn was coincubated with Cu(II) for 24 h (Fig. 6B), significantly larger aggregates (∼2 µm) and more amorphous structures were observed than when α-Syn self-aggregated for 24 h (Fig. 6A). After treatment with drugs classified as having protein-disaggregating effects using our screening model (e.g., chloroquine (CQ) and DA), distinct changes in the protein aggregation state were observed, with either reduced aggregation/smaller aggregate sizes (Fig. 6C) or dispersed fibrillar structures (Fig. 6D). For candidates predicted by the model to have minimal disaggregation effects (e.g., PQQ), the treated α-Syn − Cu(II) complexes showed similar morphologies to the samples incubated with only Cu(II), with no significant disaggregation (Fig. 6E). These TEM observations consistently matched our screening model's classification results, supporting the reliability of the model in assessing the ability of drugs to mitigate α-Syn − Cu(II) aggregation.
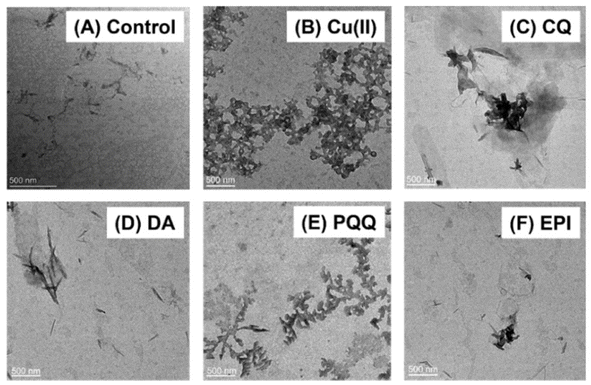 | ||
| Fig. 6 TEM images of (A) α-Syn and (B) α-Syn − Cu(II) aggregates after treatment with (C) CQ, (D) DA, (E) PQQ and (F) EPI. | ||
To support the reliability of the model in assessing the ability of drugs to mitigate ROS, UV spectroscopy of ascorbic acid (AA) was adopted. Under the action of common biological reductants such as AA, α-Syn − Cu(II) can catalyze the incomplete reduction of O2 to form ROS, including superoxide anions and hydrogen peroxide.52,75–78 The removal of Cu(II) from α-Syn − Cu(II) could prevent the generation of ROS. Therefore, the electrode interfaces could also serve as a platform for evaluating the effects of therapeutic drugs on mitigating ROS generation.
The observed drug performance in mitigating ROS generation experiments matched the classification results from our screening model.
Validation of the drug screening results in the intracellular environment
To validate the Cu(II)-induced α-Syn aggregation observed on electrodes in a cellular context, we engineered a HeLa cell line stably expressing mCherry-α-Syn. Confocal microscopy confirmed successful expression, with uniform red fluorescence (587 nm excitation) indicating soluble mCherry-α-Syn in transfected cells (Fig. S10).Consistent with electrode experiments, Cu(II) treatment (15 µM) triggered time-dependent formation of fluorescent puncta, reflecting intracellular α-Syn aggregation; it was reported that Cu(II) can induce significant α-Syn aggregation in cells under similar conditions.79 Quantitative analysis revealed maximal aggregate formation at 24 h (Fig. S11), beyond which minimal additional changes occurred. This timeframe was thus selected for subsequent studies.
To further validate the reliability of the previously established drug screening model, efficacy assessments were performed using an efficient, reproducible, and practical HeLa cell line with stable expression of mCherry-α-Syn through plasmid transfection.79 The protective effects of candidate drugs against cytotoxicity induced by mCherry-α-Syn − Cu(II) aggregates were assessed using the MTT assay to measure cell viability under 3 conditions: before Cu(II) treatment, after Cu(II) treatment alone, and after cotreatment with Cu(II) and small-molecule drugs for 24 h. When mCherry-α-Syn cells cultured in a complete medium without Cu(II) or drugs were used as the control group (CTRL) (Fig. 7B), we observed that the mCherry-α-Syn − Cu(II) group presented significantly reduced cell viability (79.6% ± 4.1%). Treatment with the dual-function small-molecule drugs (DA and EDTA) identified in our screening model, which resulted in both disaggregation and removal of ROS, resulted in marked recovery of cell viability. Specifically, compared with the Cu(II)-only group, the mCherry-α-Syn + Cu(II) + DA and mCherry-α-Syn + Cu(II) + EDTA groups presented increased viability, with values of 121.6 ± 5.7% and 111.3 ± 13.8%, respectively (Fig. 7B, group (3)), indicating statistically significant differences.
Confocal microscopy revealed that, compared with the control with Cu(II) only (Fig. 7A, panel a), dual-function drugs (CQ, DA, and EDTA) substantially reduced both the size and number of bright mCherry-α-Syn − Cu(II) aggregate puncta (Fig. 7A, panels h–j), with aggregate areas decreasing by 36.7, 52.1, and 60.2%, respectively (Fig. 7C). These results suggest that these compounds function as dual-action therapeutics capable of both disaggregating mCherry-α-Syn − Cu(II) complexes and removing the effects of ROS. In contrast, under identical treatment conditions, PQQ exhibited weaker efficacy. Microscopy analysis (Fig. 7A, panel c) revealed that while the PQQ-treated cells displayed brighter and more dispersed fluorescent puncta than the control cells (Fig. 7A, panel a), the total aggregate area remained largely unchanged. This observation aligns with the classification of PQQ as a single-function drug possessing only anti-ROS activity without significant protein-disaggregating effects, a pharmacological profile similar to that of its analog TMPD.
Furthermore, we employed the 2′,7′-dichlorodihydrofluorescein diacetate (DCFH-DA) probe to assess the removal of ROS or the inhibitory effects of different categories of small-molecule drugs (Fig. 7D). Confocal microscopy revealed that the Cu(II)-treated group exhibited significant fluorescence enhancement, indicating markedly elevated ROS levels. Notably, treatment with DA, which was predicted by our model to possess anti-ROS activity (Fig. 7D, panel c), resulted in substantial attenuation of the fluorescence signal intensity to levels comparable to those of the untreated control group (Fig. 7D, panel a). In contrast, compounds predicted by the model to lack ROS removal efficacy (EPI and α-KG) demonstrated weaker ROS scavenging capacity (Fig. 7D, panels d and e). These experimental results showed excellent agreement with the predictions from our intelligent screening platform, thereby further validating the reliability of the model.
Collectively, these experimental findings provide strong evidence supporting the reliability of our drug screening model's classification results.
Application of the drug screening model for the prediction and evaluation of potential small-molecule drugs
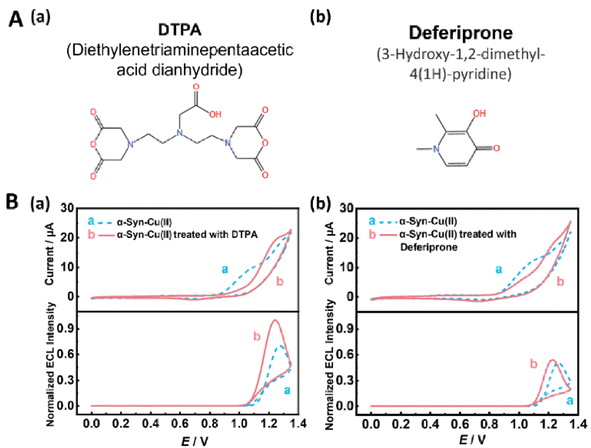
As a polydentate ligand, DTPA binds with high affinity to Cu(II),80,81 potentially inhibiting protein aggregation and ROS generation by competitively sequestering Cu(II) and blocking its interaction with α-Syn. In contrast, deferiprone—a bidentate ligand capable of crossing the blood–brain barrier—primarily targets Fe3+/Cu2+ homeostasis both intracellularly and extracellularly.82–84 Although it has demonstrated clinical potential for PD treatment,85 its precise molecular mechanisms remain incompletely understood. These characteristics provide both a theoretical rationale and a practical reason for researching these two compounds.In this study, DTPA and deferiprone were selected as model compounds for drug screening. Their electrochemical properties were first characterized on our screening platform, followed by efficacy classification using the screening model and preliminary pharmacological validation. We applied our developed LDA algorithm-based drug screening model for α-Syn to classify the pharmacological effects of candidate small-molecule drugs with unknown efficacy. Specifically, for the Cu(II)-treated α-Syn-modified electrodes, we further co-incubated them with either DTPA or deferiprone. The electrodes were then tested before and after small-molecule treatment to obtain the corresponding ECL/CV signals.
Analysis of the synchronously acquired ECL/CV signals revealed that both small-molecule treatments reduced the Ipa to approximately 1.05 V in the CV measurements, with DTPA treatment resulting in a more pronounced reduction trend. With respect to the ECL signals, the two candidate drugs exhibited divergent effects: DTPA treatment increased (restored) the ECL signals, whereas deferiprone had no significant effect on the ECL intensity (Fig. 8B). When the derived parameters E′ and I'pa were input into our drug screening model, the LDA scatter plot demonstrated distinct clustering patterns (Fig. 5C). Deferiprone showed closer proximity to the “removal of ROS (2)” category, whereas DTPA was positioned closer to the “dual effect (3)” cluster. This spatial distribution might correspond to their pharmacological profiles: deferiprone might primarily exhibit anti-ROS efficacy, whereas DTPA likely possesses dual functionality.
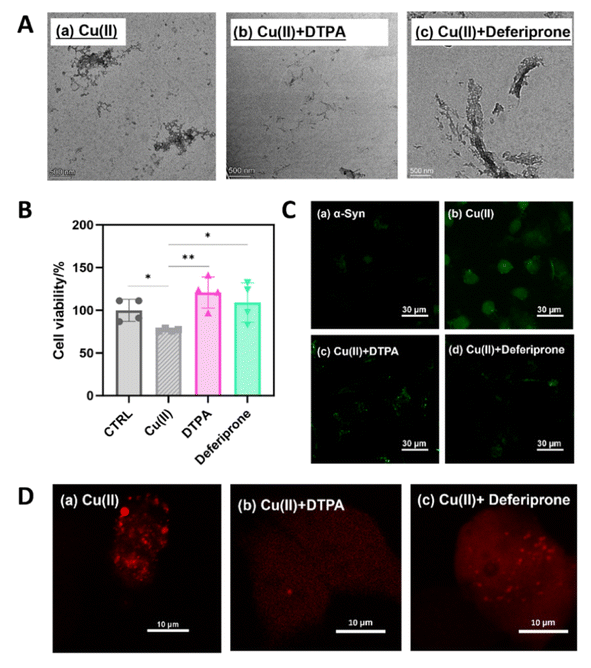
To confirm that the small-molecule drugs DTPA and deferiprone, identified through our pharmacodynamic screening model, exhibit efficacies consistent with the model predictions, we employed TEM and an mCherry-α-Syn cell line to evaluate the ability of these drugs to mitigate aggregation and remove ROS. Compared with incubation with Cu(II) alone (Fig. 9A, panel a), addition of DTPA significantly reduced protein aggregation, resulting in smaller aggregate sizes (Fig. 9A, panel b). In contrast, incubation with Cu(II) and deferiprone altered the protein aggregation morphology but did not substantially alleviate aggregation (Fig. 9A, panel c).
Next, cells expressing mCherry-α-Syn were incubated with either 15 µM Cu(II) alone or 15 µM Cu(II) + 15 µM small-molecule drugs. We first evaluated the effects of both compounds on cytotoxicity (Fig. 9B). The results demonstrated that DTPA and deferiprone markedly restored cell viability. Compared with the Cu(II)-only control group (76.7 ± 1.7%), viability increased to 120.8 ± 18.2% and 109.1 ± 23.0%, respectively, ranking among the highest-performing compounds in our screening.
Confocal microscopy revealed that both DTPA and deferiprone reduced the area and number of bright mCherry-α-Syn aggregate foci in cells compared with those in the untreated control, with DTPA exhibiting superior efficacy (Fig. 9D). This observation aligns with the positions of the DTPA and deferiprone in the LDA scatter plot (Fig. 5C). Notably, DTPA clustered closer to the “dual effect (3)” group, indicating that our screening model effectively distinguished its anti-aggregation potency from that of deferiprone.
Additionally, intracellular ROS scavenging/inhibition by these drugs was assessed using a DCFH-DA probe (Fig. 9C). Confocal imaging revealed significantly less fluorescence in both treatment groups than in the Cu(II)-exposed groups, confirming the suppression of ROS production.
Conclusions
This study developed an innovative dual-mode signal platform for the simultaneous screening of small-molecule drugs that target both protein aggregation and ROS generation. By monitoring dual pathological features of α-Syn aggregation through synchronously acquired ECL and CV signals in a single test and employing machine learning algorithms (LDA/HCA) as analytical aids, this antibody-free system successfully achieved intelligent drug efficacy classification using α-Syn interfaces. The platform demonstrated marked efficiency, reducing the analysis time from several days to mere hours while enabling real-time monitoring of both the progression of protein aggregation and the production of ROS. After cellular-level validation, the model was applied to preliminarily assess and subsequently confirm the therapeutic effects of two previously unreported small-molecule compounds (DTPA and deferiprone) in alleviating metal-induced α-Syn aggregation and accompanying ROS production.This integration of electrochemical sensing with machine learning established a rapid and cost-effective screening tool, highlighting the synergistic pathogenic role of metal ions and oxidative stress in neurodegenerative pathologies. The platform's unique ability to simultaneously track dual pathological features in a single measurement represents a significant advance over conventional single-parameter detection methods. Its modular design could be readily adapted to study other pathological marker proteins (such as amyloid-β peptide and tau in Alzheimer's disease or huntingtin in Huntington's disease), offering broad application prospects in the field of NDs.
Author contributions
H. Liu and L. Mao guided the project. J. Shi, W. Zhu, J. Yu, and C. Xiao conceived the idea and designed and performed the experiments. J. Shi and W. Zhu analyzed the data and drew the figures. All authors prepared the manuscript and provided their suggestions.Conflicts of interest
The authors declare no competing financial interest.Data availability
The authors confirm that the data supporting the findings of this study are available within the article and as its supplementary information (SI). Supplementary information: the SI contain experimental procedures, characterization data, and supporting figures and tables. See DOI: https://doi.org/10.1039/d5sc08658b.Acknowledgements
The authors greatly appreciate the funding from the Key Research and Development Program of Ningxia Province of China (2023BEG03020 and 2023BEG02031) and Natural Science Foundation of Ningxia Province of China (2023AAC02039 and 2024AAC03265).References
- A. Abeliovich, Y. Schmitz, I. Fariñas, D. Choi-Lundberg, W.-H. Ho, P. E. Castillo, N. Shinsky, J. M. G. Verdugo, M. Armanini, A. Ryan, M. Hynes, H. Phillips, D. Sulzer and A. Rosenthal, Neuron, 2000, 25, 239–252 CrossRef CAS PubMed.
- M. Shahnawaz, A. Mukherjee, S. Pritzkow, N. Mendez, P. Rabadia, X. Liu, B. Hu, A. Schmeichel, W. Singer, G. Wu, A.-L. Tsai, H. Shirani, K. P. R. Nilsson, P. A. Low and C. Soto, Nature, 2020, 578, 273–277 CrossRef CAS PubMed.
- C. R. Fields, N. Bengoa-Vergniory and R. Wade-Martins, Front. Mol. Neurosci., 2019, 12, 299 CrossRef CAS PubMed.
- Y. Huang, J. Wen, L.-M. Ramirez, E. Gümüşdil, P. Pokhrel, V. H. Man, H. Ye, Y. Han, Y. Liu, P. Li, Z. Su, J. Wang, H. Mao, M. Zweckstetter, S. Perrett, S. Wu and M. Gao, Nat. Commun., 2023, 14, 5444 CrossRef CAS PubMed.
- L. Macomber and J. A. Imlay, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 8344–8349 Search PubMed.
- K. J. Barnham, C. L. Masters and A. I. Bush, Nat. Rev. Drug Discovery, 2004, 3, 205–214 Search PubMed.
- M. W. Chen, X. Ren, X. Song, N. Qian, Y. Ma, W. Yu, L. Yang, W. Min, R. N. Zare and Y. Dai, J. Am. Chem. Soc., 2025, 147, 8267–8279 CrossRef CAS PubMed.
- K. A. Murray, C. J. Hu, H. Pan, J. Lu, R. Abskharon, J. T. Bowler, G. M. Rosenberg, C. K. Williams, G. Elezi, M. Balbirnie, K. F. Faull, H. V. Vinters, P. M. Seidler and D. S. Eisenberg, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2217835120 CrossRef CAS PubMed.
- P. M. Seidler, K. A. Murray, D. R. Boyer, P. Ge, M. R. Sawaya, C. J. Hu, X. Cheng, R. Abskharon, H. Pan, M. A. DeTure, C. K. Williams, D. W. Dickson, H. V. Vinters and D. S. Eisenberg, Nat. Commun., 2022, 13, 5451 CrossRef CAS PubMed.
- Y. Wang, M. Lv, W. Zhu, J. Yan, J. Shi, Z. Qin, X. Zhang, H. Yao, L. Meng, H. Liu and L. Mao, Anal. Chem., 2025, 97, 27805–27814 CrossRef CAS PubMed.
- K. Matsuo, H. Hiramatsu, K. Gekko, H. Namatame, M. Taniguchi and R. W. Woody, J. Phys. Chem. B, 2014, 118, 2785–2795 CrossRef CAS PubMed.
- M. Kim, J. Kang, M. Lee, J. Han, G. Nam, E. Tak, M. S. Kim, H. J. Lee, E. Nam, J. Park, S. J. Oh, J.-Y. Lee, J.-Y. Lee, M.-H. Baik and M. H. Lim, J. Am. Chem. Soc., 2020, 142, 8183–8193 CrossRef CAS PubMed.
- K. Liu, Y. Tao, Q. Zhao, W. Xia, X. Li, S. Zhang, Y. Yao, H. Xiang, C. Han, L. Tan, B. Sun, D. Li, A. Li and C. Liu, Proc. Natl. Acad. Sci. U. S. A., 2024, 121, e2321633121 CrossRef CAS PubMed.
- Y. Qi, X. Xu, H. Dong, Z. Yang and P. Wang, Food Biophys., 2023, 18, 71–81 CrossRef.
- D. Pramanik, K. Sengupta, S. Mukherjee, S. G. Dey and A. Dey, J. Am. Chem. Soc., 2012, 134, 12180–12189 CrossRef CAS PubMed.
- S. Li and K. Kerman, Anal. Chem., 2019, 91, 3818–3826 CrossRef CAS PubMed.
- Y. Zhang, W. Ji, S. Zhang, N. Gao, T. Xu, X. Wang and M. Zhang, Angew. Chem., Int. Ed., 2022, 61, e202111853 CrossRef CAS PubMed.
- M. Asadbegi and A. Shamloo, J. Chem. Inf. Model., 2021, 61, 1383–1401 CrossRef CAS PubMed.
- J. Yuan, T. Li, X.-B. Yin, L. Guo, X. Jiang, W. Jin, X. Yang and E. Wang, Anal. Chem., 2006, 78, 2934–2938 Search PubMed.
- J. Ni, B. Yang, L. Liu, X. Dai, W. Yang, Q. Wang, X. Chen, Z. Song and Z. Lin, Anal. Chem., 2024, 96, 12577–12583 Search PubMed.
- H. Qin, X. Gao, X. Yang, W. Cao and S. Liu, Biosens. Bioelectron., 2019, 141, 111438 Search PubMed.
- X. Feng, N. Gan, H. Zhang, T. Li, Y. Cao, F. Hu and Q. Jiang, Biosens. Bioelectron., 2016, 75, 308–314 Search PubMed.
- W. Lian, J. Liang, L. Shen, Y. Jin and H. Liu, Biosens. Bioelectron., 2018, 100, 326–332 CrossRef CAS PubMed.
- M. M. Richter, Chem. Rev., 2004, 104, 3003–3036 CrossRef CAS PubMed.
- J. Liang, M. Li, Y. Lu, H. Yao and H. Liu, Biosens. Bioelectron., 2018, 118, 44–50 Search PubMed.
- H. Wei, Y. Du, J. Kang and E. Wang, Electrochem. Commun., 2007, 9, 1474–1479 Search PubMed.
- I. Bist, B. Song, I. M. Mosa, T. E. Keyes, A. Martin, R. J. Forster and J. F. Rusling, ACS Sens., 2016, 1, 272–278 Search PubMed.
- Q.-M. Feng, Y.-H. Guo, J.-J. Xu and H.-Y. Chen, Biosens. Bioelectron., 2018, 1, 571–576 CrossRef PubMed.
- B. Song, M. Shen, D. Jiang, S. Malla, I. M. Mosa, D. Choudhary and J. F. Rusling, Analyst, 2016, 141, 5722–5729 Search PubMed.
- Y. Zhang, H. Zhang and N. Hu, Biosens. Bioelectron., 2008, 23, 1077–1082 CrossRef CAS PubMed.
- T. Yang, J. Fu, S. Zheng, H. Yao, Y. Jin, Y. Lu and H. Liu, Biosens. Bioelectron., 2018, 108, 62–68 CrossRef CAS PubMed.
- S. Liu, M. Li, X. Yu, C.-Z. Li and H. Liu, Chem. Commun., 2015, 51, 13185–13188 Search PubMed.
- J. Tong, M. Xiao, K. Wang, Z. Zhao, Y. Chen, Y. Liu, T. Qing, X. Liu and Z. Zhang, ACS Nano, 2025, 19, 11358–11370 CrossRef CAS PubMed.
- C. Xiao, J. Shi, J. Yan, R. Xiao, Y. Wang, H. Yao, L. Meng, H. Liu and L. Mao, Anal. Chem., 2025, 97, 10328–10336 Search PubMed.
- H. Zhu, J. Yan and A. Revzin, Colloids Surf., B, 2008, 64, 260–268 CrossRef CAS PubMed.
- Y. Wang, F. Zhai, Y. Hasebe, H. Jia and Z. Zhang, Bioelectrochemistry, 2018, 122, 174–182 Search PubMed.
- S. Zhang, Y. Ding and H. Wei, Molecules, 2014, 19, 11933–11987 Search PubMed.
- W. Lian, R. Xiao, J. Li, H. Yao and H. Liu, Sens. Acutators Rep., 2021, 3, 100054 Search PubMed.
- W. Lian, X. Yu, L. Wang and H. Liu, J. Phys. Chem. C, 2015, 119, 20003–20010 CrossRef CAS.
- P. Ceroni, G. Bergamini and V. Balzani, Angew. Chem., Int. Ed., 2009, 48, 8516–8518 CrossRef CAS PubMed.
- M. Su and S. Liu, Anal. Biochem., 2010, 402, 1–12 Search PubMed.
- D. H. Johnston, K. C. Glasgow and H. H. Thorp, J. Am. Chem. Soc., 1995, 117, 8933–8938 Search PubMed.
- A. Binolfi, R. M. Rasia, C. W. Bertoncini, M. Ceolin, M. Zweckstetter, C. Griesinger, T. M. Jovin and C. O. Fernández, J. Am. Chem. Soc., 2006, 128, 9893–9901 Search PubMed.
- A. Binolfi, L. Quintanar, C. W. Bertoncini, C. Griesinger and C. O. Fernández, Coord. Chem. Rev., 2012, 256, 2188–2201 Search PubMed.
- M. D. Tuttle, G. Comellas, A. J. Nieuwkoop, D. J. Covell, D. A. Berthold, K. D. Kloepper, J. M. Courtney, J. K. Kim, A. M. Barclay, A. Kendall, W. Wan, G. Stubbs, C. D. Schwieters, V. M. Y. Lee, J. M. George and C. M. Rienstra, Nat. Struct. Mol. Biol., 2016, 23, 409–415 Search PubMed.
- A. Carija, F. Pinheiro, J. Pujols, I. C. Brás, D. F. Lázaro, C. Santambrogio, R. Grandori, T. F. Outeiro, S. Navarro and S. Ventura, Redox Biol., 2019, 22, 101135 Search PubMed.
- R. Tan, Y. Wang, X. Mi, H. Li and Y. Tu, Sens. Actuators, B, 2022, 352, 131065 Search PubMed.
- Z. Chen, Y. Li, H. Qin, X. Yang and W. Cao, Anal. Methods, 2022, 14, 1739–1746 Search PubMed.
- N. González, T. Arcos-López, A. König, L. Quintanar, M. Menacho Márquez, T. F. Outeiro and C. O. Fernández, J. Neurochem., 2019, 150, 507–521 Search PubMed.
- C.-Y. Wang and H.-J. Huang, Anal. Chim. Acta, 2003, 498, 61–68 CrossRef CAS.
- S. Liu, Q. Li, H. Yang, P. Wang, X. Miao and Q. Feng, Biosens. Bioelectron., 2022, 196, 113744 Search PubMed.
- C. Wang, L. Liu, L. Zhang, Y. Peng and F. Zhou, Biochemistry, 2010, 49, 8134–8142 Search PubMed.
- D. Jiang, L. Men, J. Wang, Y. Zhang, S. Chickenyen, Y. Wang and F. Zhou, Biochemistry, 2007, 46, 9270–9282 CrossRef CAS PubMed.
- D. L. Nelson, A. A. Hoskins and M. M. Cox, in Principles of biochemistry, Macmillan International Higher Education, New York, NY, edn. 8th edn, 2021, ch. 13, pp. 3000–3001 Search PubMed.
- B. E. Conway, Electrochemical Data, New York: Printed in New York, 1952 Search PubMed.
- G. Dryhurst, K. M. Kadish, F. Scheller and R. Renneberg, in Biological Electrochemistry, ed. G. Dryhurst, K. M. Kadish, F. Scheller and R. Renneberg, Academic Press, 1982, pp. 256–278 Search PubMed.
- J. Sian, D. T. Dexter, A. J. Lees, S. Daniel, Y. Agid, F. Javoy-Agid, P. Jenner and C. D. Marsden, Ann. Neurol., 1994, 36, 348–355 CrossRef CAS PubMed.
- Y. Okita, A. N. Rcom-H'cheo-Gauthier, M. Goulding, R. S. Chung, P. Faller and D. L. Pountney, Front. Neurosci., 2017, 11, 114 CrossRef PubMed.
- Y. Li, C. Yang, S. Wang, D. Yang, Y. Zhang, L. Xu, L. Ma, J. Zheng, R. B. Petersen, L. Zheng, H. Chen and K. Huang, Int. J. Biol. Macromol., 2020, 163, 562–573 CrossRef CAS PubMed.
- A. Kumar, D. Ganini and R. P. Mason, Mol. Neurodegener., 2016, 11, 70 CrossRef PubMed.
- Y. Inokuchi, S. Imai, Y. Nakajima, M. Shimazawa, M. Aihara, M. Araie and H. Hara, J. Pharmacol. Exp. Ther., 2009, 329, 687–698 CrossRef CAS PubMed.
- H. R. Lotfi Zadeh Zhad, M. H. Banitaba, M. H. Aghabeigi Roozbahani and S. S. Hosseiny Davarani, ECS Electrochem. Lett., 2012, 1, G4–G6 CrossRef.
- C. Wang, H.-L. Liu, Y.-Q. Li, J. Cao, B. Zheng, X.-H. Xia and F. Feng, Electrochem. Commun., 2016, 66, 25–28 Search PubMed.
- S. Li, M. Noroozifar and K. Kerman, Electrochim. Acta, 2021, 388, 138534 Search PubMed.
- L. Zhang, B. Wu, G. Zhang, Y. Gan and S. Zhang, Chem. Eng. J., 2019, 358, 1218–1226 Search PubMed.
- Y. Mu, X. Tong, Y. Guan, Q. Yu, W. Ren, L. Tian, H. Pei, S. Zhang, L. Yang, H. Li, L. Zhang and J.-P. Zou, Environ. Sci. Technol., 2025, 59, 968–977 Search PubMed.
- K. A. Conway, J.-C. Rochet, R. M. Bieganski and P. T. Lansbury, Science, 2001, 294, 1346–1349 Search PubMed.
- L. Breydo, J. W. Wu and V. N. Uversky, Biochim. Biophys. Acta, 2012, 1822, 261–285 CrossRef CAS PubMed.
- A. F. Fischer and K. M. Matera, Neurochem. Res., 2015, 40, 1341–1349 CrossRef CAS PubMed.
- J. Li, M. Zhu, A. B. Manning-Bog, D. A. Di Monte and A. L. Fink, FASEB J., 2004, 18, 962–964 CrossRef CAS PubMed.
- T. Zohoorian-Abootorabi, A. A. Meratan, S. Jafarkhani, V. Muronetz, T. Haertlé and A. A. Saboury, Sci. Rep., 2023, 13, 5275 Search PubMed.
- W. Poewe, K. Seppi, C. M. Tanner, G. M. Halliday, P. Brundin, J. Volkmann, A.-E. Schrag and A. E. Lang, Nat. Rev. Dis. Primers, 2017, 3, 17013 CrossRef PubMed.
- S. Zhao, B. Zhang, J. Yang, J. Zhou and Y. Xu, Nat. Rev. Methods Primers, 2024, 4, 70 CrossRef CAS.
- Y. Shen, Y. Huang, P. Zhang, B. Guo, H. Jiang, C. Tan and Y. Jiang, ACS Appl. Bio Mater., 2020, 3, 5639–5643 Search PubMed.
- V. Borghesani, B. Alies and C. Hureau, Eur. J. Inorg. Chem., 2018, 2018, 7–15 Search PubMed.
- C. Esmieu, G. Ferrand, V. Borghesani and C. Hureau, Chem. - Eur. J., 2021, 27, 1777–1786 Search PubMed.
- D. Pramanik, C. Ghosh, S. Mukherjee and S. G. Dey, Coord. Chem. Rev., 2013, 257, 81–92 CrossRef CAS.
- K. Reybier, S. Ayala, B. Alies, J. V. Rodrigues, S. Bustos Rodriguez, G. La Penna, F. Collin, C. M. Gomes, C. Hureau and P. Faller, Angew. Chem., 2016, 128, 1097–1101 Search PubMed.
- S. Ray, N. Singh, R. Kumar, K. Patel, S. Pandey, D. Datta, J. Mahato, R. Panigrahi, A. Navalkar, S. Mehra, L. Gadhe, D. Chatterjee, A. S. Sawner, S. Maiti, S. Bhatia, J. A. Gerez, A. Chowdhury, A. Kumar, R. Padinhateeri, R. Riek, G. Krishnamoorthy and S. K. Maji, Nat. Chem., 2020, 12, 705–716 Search PubMed.
- J. Wang, H. Guan, Q. Liang and M. Ding, Composites, Part B, 2020, 198, 108248 Search PubMed.
- U. Muller-Eberhard, M. E. Erlandson, H. E. Ginn and C. H. Smith, Blood, 1963, 22, 209–217 Search PubMed.
- Z. I. Cabantchik, A. Munnich, M. B. Youdim and D. Devos, Front. Pharmacol, 2013, 4, 167 Search PubMed.
- Y.-S. Sohn, A.-M. Mitterstiller, W. Breuer, G. Weiss and Z. I. Cabantchik, Br. J. Pharmacol., 2011, 164, 406–418 Search PubMed.
- A. M. Fredenburg, R. K. Sethi, D. D. Allen and R. A. Yokel, Toxicology, 1996, 108, 191–199 Search PubMed.
- S. Ayton, M. Woodward, K. A. Ellis, Y. Y. Lim, P. T. Maruff, P. M. Desmond, O. Salvado, A. Fazlollahi, C. C. Rowe, L. Churilov, A. I. Bush and D. T. Investigators, Alzheimer's Dementia, 2020, 16, e044107 CrossRef.
| This journal is © The Royal Society of Chemistry 2026 |