Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Computational redesign and directed evolution of a lanthanide-dependent photoredox enzyme for enantioselective diol cleavage
Florian
Leiss-Maier
 ,
Joshua
Behringer
,
Joshua
Behringer
 ,
Ghulam
Mustafa
,
Ghulam
Mustafa
 ,
Anna
Heider
,
Anna
Heider
 ,
Rahel
Mühlhofer
,
Rahel
Mühlhofer
 ,
Andreas S.
Klein
,
Andreas S.
Klein
 ,
Michael
Groll
and
Cathleen
Zeymer
,
Michael
Groll
and
Cathleen
Zeymer
 *
*
Center for Functional Protein Assemblies & Department of Bioscience, TUM School of Natural Sciences, Technical University of Munich (TUM), 85748 Garching, Germany. E-mail: cathleen.zeymer@tum.de
First published on 17th March 2026
Abstract
De novo designed metalloenzymes and photoenzymes are a valuable addition to the biocatalytic toolbox. We previously introduced PhotoLanZymes (PLZ), a family of lanthanide-dependent photoredox enzymes that enable radical carbon–carbon bond cleavages of diol substrates upon Ce(III/IV) binding and visible-light irradiation. While rational optimization increased their catalytic activity and photostability, the first generation of PLZ variants was limited by slow lanthanide binding and a lack of enantioselectivity. Here, we demonstrate that coupling computational redesign with directed evolution provides an effective strategy to overcome these limitations. First, we reduced the cavity size to enhance substrate interactions with the protein's active site, which facilitated initial enantiocontrol. Simultaneously, the AI-guided redesign approach improved the lanthanide binding kinetics. We then performed directed evolution to selectively accelerate the photocatalytic turnover for one of the substrate enantiomers, yielding a PLZ variant with markedly improved enantioselectivity. These results underscore the value of integrating AI-guided protein design with laboratory evolution to obtain stereoselective de novo metalloenzymes and photoenzymes.
Introduction
The computational design of proteins has seen remarkable improvements in recent years, in particular after the implementation of deep learning-based methods.1–3 This increase in design accuracy has yielded high affinity binders4–6 and tailor-made protein scaffolds.7,8 However, the computational design of catalytically active enzymes remains challenging, as it necessitates the precise arrangement of catalytic residues in three-dimensional space with sufficient conformational flexibility.9–11In an alternative approach to generate artificial enzymes with new-to-nature activity, protein scaffolds can be equipped with cofactors that catalyze challenging reactions inside of the chiral protein environment, thereby enabling enantioselective transformations. Options to endow protein scaffolds with catalytic function comprise photosensitizers, to produce artificial photoenzymes,12–15 and metal cofactors, to generate artificial metalloenzymes. The latter make up a big portion of the de novo enzymes known to date and include a range of potent hybrid catalysts for a variety of applications.16 Frequently used metal cofactors encompass transition metals customary to both nature and synthetic chemistry, such as copper17,18 and iron,19–21 but also abiological metals like titanium22 have been used in the past. Here, we focus on lanthanide ions, a currently underexplored class of enzyme cofactors known to catalyze a broad range of reactions, including photoredox transformations.23–26
Our group has recently reported a de novo designed photoenzyme that leverages lanthanide ions as photoredox cofactors to catalyze radical reactions upon visible-light irradiation.27 The enzyme called PhotoLanZyme (PLZ) uses a datively coordinated Ce(III/IV) ion to catalyze the photoinduced C–C bond cleavage of diol substrates to yield carbonyl products. The protein scaffold consists of a de novo TIM barrel domain coordinating the lanthanide ion with four glutamate residues and a dimeric ferredoxin domain that acts as a lid above the active site.28 Induced by photoexcitation, the central Ce(IV) is proposed to engage in ligand-to-metal charge transfer (LMCT) with its diol substrates to catalyze radical diol cleavages, as described previously.29–31 Rational engineering of the initial design PLZ1.0 generated PLZ1.4, a variant with optimized photostability and reduced off-target metal binding. This resulted in improved yields for a variety of diol substrates.27 However, a major limitation was the enzyme's lack of enantioselectivity, which we set out to tackle in this work. Our goal was thus to redesign and evolve PLZ to enable the kinetic resolution of racemic mixtures of diol substrates.
Here we describe a computational redesign strategy that yielded a PLZ variant with an adjusted cavity size, resulting in a 100-fold higher kcat/KM for the model substrate hydrobenzoin and initial enantioselectivity for a slightly larger diol substrate. Directed evolution of this improved photoenzyme could further increase the enantioselectivity. A crystal structure of the final variant validated the success of our redesign strategy. These results highlight that the combination of computational and experimental optimization is a promising strategy to improve activity and selectivity of de novo enzymes.
Results and discussion
Computational redesign to adjust the size of the substrate binding cavity
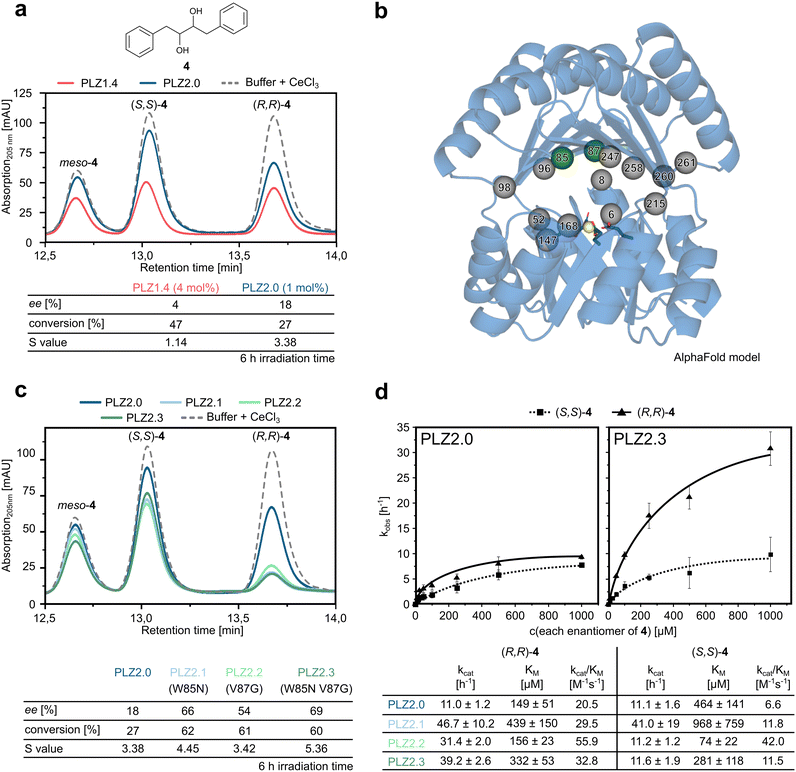
To improve activity and selectivity of PLZ1.4, the previously reported and rationally engineered cerium-dependent photoenzyme,27 we first analyzed its structural features. We hypothesized that the active site cavity was too large to efficiently pre-orient diol substrates near the catalytically active cerium ion and thus to discriminate between stereoisomers. Therefore, we set out to computationally redesign and optimize PLZ1.4 in order to reduce the size of the binding pocket. We first deleted the four flexible linker regions originally connecting the ferredoxin domain and the TIM barrel and then manually lowered the lid domain by 3 Å to generate a smaller cavity. This adjusted scaffold structure was used as an input for protein redesign using AI-guided tools. The connecting linkers were re-introduced by RFdiffusion and the newly generated domain–domain interface was optimized using partial diffusion.32 Following this step, we generated sequences for the obtained fold using LigandMPNN,33 taking into account the desired position of the diol substrate. Here, the four coordinating glutamate residues remained fixed in their position to assure high affinity for lanthanide cofactors (Fig. 1a and S1). After structure prediction, first using ESMfold34 and then Alphafold2 (ref. 35) in Colabfold,36 the designs with the highest confidence levels (predicted local distance difference test, pLDDT) were selected. These structures were manually evaluated by comparing cavity size, shape, and domain-specific prediction confidence. Off-target metal binding sites were identified using BioMetAll37 and removed by mutagenesis (E to Q, D to N, and C to S mutations). Finally, two loops with low pLDDT scores in the ferredoxin domains were shortened to improve domain rigidity and model confidence. The resulting protein, called PLZ2.0, has 35% sequence identity to PLZ1.4 and is shortened by 14 residues in total (12 residues less due to linker remodeling and 2 residues less in the ferredoxin domains). It retains the overall fold, but with a significantly reduced cavity volume (Fig. 1a and S1). | ||
| Fig. 1 PLZ redesign and catalytic efficiency. (a) Structural models of PLZ1.4 (red, left) and PLZ2.0 (blue, right), showing the reduction in cavity size after computational redesign (depicted as internal protein surface). Right: the cerium ion is shown in yellow, coordinating glutamate residues are shown as sticks. (b) Proposed mechanism of the photoinduced cerium-dependent radical diol cleavage as reported previously in organic solvent.30 LMCT: ligand-to-metal charge transfer. HAT: hydrogen atom transfer. (c) Comparison of yields obtained for model substrates 1 and 3 using PLZ2.0 and PLZ1.4 with 5 mol% catalyst loading after 24 h irradiation at 410–420 nm. (d) Michaelis–Menten kinetics for substrate (R,R)-1 using the photoenzymes PLZ1.4 and PLZ2.0, in red and blue, respectively. | ||
PLZ2.0 shows improved catalytic efficiency
Even though the scaffold redesign process only intended to decrease the cavity size, AI-guided sequence redesign produced additional benefits: first, PLZ2.0 showed significantly faster lanthanide binding kinetics than PLZ1.4. When following the specific tryptophan-enhanced Tb(III) luminescence as a binding readout, PLZ1.4 needed more than two hours at room temperature to reach the maximum signal,27 while for PLZ2.0 complete metal binding was observed within 5 minutes (Fig. S2). This may be due to an improved conformational equilibrium. In a previous study, we found that MPNN-based sequence redesign stabilized the binding-competent conformation of the original PLZ1.0 scaffold by introducing additional molecular interactions in the dimer interface.38 We hypothesize that a similar effect is also responsible for the accelerated lanthanide binding here, when redesigning for a smaller cavity. Importantly, nanomolar lanthanide affinity was retained in the redesign process, as shown by titration experiments (Fig. S3–S6). A knockout variant (PLZ2.0_KO), in which the coordinating glutamate residues (E31, E147, E193, E309) were replaced by glutamines, did not show any lanthanide binding in the active site (Fig. S2).Next, we tested the catalytic activity of PLZ2.0 in cerium-dependent photoinduced diol cleavage reactions. The proposed mechanism of the equivalent reaction in organic solvent proceeds via the coordination of the diol to Ce(IV), followed by visible light-induced LMCT (Fig. 1b).30 The resulting alkoxy radical can subsequently undergo β-scission, producing a carbonyl product and a carbon-centered radical. This radical can further undergo hydrogen atom transfer (HAT) to produce the second carbonyl product. Finally, Ce(III) is proposed to be re-oxidized to Ce(IV) by atmospheric oxygen, as recently observed in organic solvents.39 While mechanistic details of the photoenzymatic transformation remain unknown, UV/Vis absorption spectra support that the Ce(IV)-bound protein can be excited at around 400 nm (Fig. S7).
Indeed, reducing the size of the active site cavity showed a beneficial outcome in our test reactions. Using our previously reported standard reaction conditions with 5 mol% catalyst loading and irradiation for 24 h at 410–420 nm, we observed quantitative diol cleavage for the model substrate hydrobenzoin (R,R)-1 to benzaldehyde 2, compared to 80% yield for PLZ1.4. Even the smaller and less reactive diol (R)-3 yielded 65% of product 2, which marked a significant increase compared to the 20% yield observed with PLZ1.4 (Fig. 1c). The knock-out variant PLZ2.0_KO showed very low background activity, attributed to only trace amounts of lanthanides binding to the protein surface (Fig. S8). This may also be a beneficial side effect of the redesign process, in which unspecific metal binding was further reduced.
To assess whether the reduced cavity size restricted access for larger substrates, we performed docking with a range of diols of increasing size using Glide.40 Applying a blind docking mode, where no binding cavity was pre-defined, we observed that the experimentally confirmed substrates 1 and 4 as well as di-methylated variants can enter the active site (Fig. S9). Larger diols were repeatedly excluded, indicating a stringent size selectivity.
We then measured Michaelis–Menten kinetics of PLZ1.4 and PLZ2.0 using substrate (R,R)-1. Strikingly, the KM value was lowered more than 200 fold, from 2900 ± 400 µM for PLZ1.4 to 13 ± 1 µM for PLZ2.0, while kcat slightly decreased by a factor of two (Fig. 1d). The resulting 100-fold improvement in catalytic efficiency can thus be attributed exclusively to an optimized substrate binding behavior, which is consistent with the hypothesis that the smaller cavity would allow a better preorganization of the diol for the reaction. Additionally, a change in conformational flexibility of the inter-domain linkers may improve substrate positioning in the active site.41–43 Replacing the four highly flexible tri-glycine linkers connecting the TIM barrel to the ferredoxin lid by shorter and more rigid sequences (DTD, QG, DQD, and LG, respectively) could have further enhanced substrate binding by reducing twisting motions previously observed for similar scaffolds.38 Encouraged by these results, we decided to investigate whether the newly designed active site could also discriminate between individual diol substrate enantiomers.
The smaller cavity facilitates initial enantiocontrol
We next compared the enantioselectivity of the redesigned photoenzyme PLZ2.0 with that of the parent variant PLZ1.4, using both hydrobenzoin (1) and the slightly bulkier diol substrate 4. Because of the higher catalytic activity of PLZ2.0, catalyst loading was reduced to 1 mol% in these experiments. While no pronounced selectivity was observed for 1 (Fig. S10), promising initial stereocontrol was achieved for 4, where the redesigned PLZ2.0 could discriminate between the (R,R) and (S,S) enantiomer (Fig. 2a). When analyzing the unreacted substrate 4 by chiral HPLC after 6 h irradiation time, we found an enantiomeric excess (ee) of 18% at 27% conversion, which translates into a selectivity factor (S) of 3.4—substantially higher than the value of 1.1 observed for PLZ1.4. This advancement in selectivity demonstrated that PLZ2.0 could potentially be optimized for the kinetic resolution of racemic diol substrates. These results established PLZ2.0 as an improved catalyst and thus a promising starting point for directed evolution.Directed evolution improves the enantioselectivity
We screened site-saturation libraries of residues lining the active site of PLZ2.0 (Fig. 2b). In a first round we randomized positions L52, W85, V87, A98, Q168, and A247 using a reduced amino acid alphabet encoded by NDT codons (including amino acids D, R, Y, F, I, S, C, H, N, G, L, V). After expression in 96-well plate format, cells were lysed and the highly thermostable de novo protein was purified by a heat step (1 h at 80 °C), while remaining DNA was removed by precipitation with polythylenimine (Fig. S11). The clean, cerium(III)-loaded enzymes were irradiated in an LED photoreactor equipped with a custom cooling system for microtiter plates. All reactions were then analyzed by chiral HPLC to detect variants with improved enantioselectivity. We were able to identify two hits: W85N (=PLZ2.1) and V87G (=PLZ2.2). After large-scale purification of these enzymes we confirmed that the mutations increased both enantioselectivity and activity of the photoenzyme (Fig. 2c). Combining the two point mutations yielded PLZ2.3 (=PLZ2.0 W85N V87G). The double mutant showed further enhanced selectivity with an S value of 5.4. This improvement demonstrates that the PLZ family can be optimized both in selectivity and catalytic efficiency and therefore is a promising class of catalysts for kinetic resolution experiments. Screening of ten further single site-saturation libraries (see Fig. 2b) based on PLZ2.3 unfortunately did not yield any additional hits. We therefore decided to determine the kinetic parameters of all available enzyme variants towards the enantiomers of 4 to improve our understanding of the underlying mechanism of selectivity.For the Michaelis–Menten experiments, we also removed meso-4 from the substrate mixture (Fig. S13) to be able to better assess the individual kinetics for the (R,R) and (S,S) enantiomer. Compared to PLZ2.0, kcat and KM for (S,S)-4 remained in similar regimes for PLZ2.3, while for (R,R)-4 the kcat increased almost 4-fold (Fig. 2d). Despite being accompanied by a slight increase in KM for the (R,R)-enantiomer, this resulted in the improved selectivity observed for the double mutant.
We can therefore conclude that the two mutations selectively increased the reaction rate for only (R,R)-4, while hardly affecting the behavior of the enzyme towards (S,S)-4. When comparing the kinetic parameters to the single-mutant variants PLZ2.1 and PLZ2.2 it becomes apparent that PLZ2.1 exhibits a higher kcat than PLZ2.3, while PLZ2.2 shows an improved KM (Fig. 2d and S14). Although these effects are partially attenuated in the double mutant, PLZ2.3 exhibits the highest enantioselectivity, as judged by the difference in kcat/KM for (R,R)-4versus (S,S)-4, in line with our evolutionary focus on improved enantioselectivity. Nevertheless, the improvement between the single mutants and PLZ2.3 is modest and less than additive, indicating an interdependence between the individual point mutants. To quantify this effect, we performed a double-mutant cycle analysis using kcat/KM for (R,R)-4. The resulting interaction free energy (ΔΔG‡int ≈ +0.5 kcal mol−1) suggests antagonistic epistasis, consistent with the double mutant performing worse than predicted from the individual substitutions. Interestingly, when assessing the enantioselectivity of PLZ2.3 towards the original substrate 1, we observed a slight selectivity for the opposite (S,S)-enantiomer, although less pronounced (Fig. S15).
Next, we determined if the lanthanide binding affinity was impacted by the mutations in the active site of PLZ2.3. We transitioned to isothermal titration calorimetry (ITC), as the tryptophan antenna (W85) required for the sensitized terbium luminescence readout was no longer present. At pH 8.5, lanthanide hydrolysis44 impaired the measurements and a shift to lower pH was required. Slightly lower affinities due to partial protonation of the coordinating residues45 were thus expected. We determined cerium and terbium binding affinities between 50 nM and 350 nM (Fig. S4–S6) and thus concluded that the affinity for lanthanides did not drastically suffer from the introduced point mutations.
To assess potential photodamage during extended irradiation, protein samples containing CeCl3 with or without substrate were analyzed by SDS-PAGE and LC-MS after 6 hours of irradiation. SDS-PAGE revealed no evidence of protein crosslinking or degradation (Fig. S16). LC-MS showed a pattern of several +16 Da mass shifts for PLZ2.0 and PLZ2.3 (Fig. S17), consistent with oxidative modifications of residues such as Trp or Met, as also observed for PLZ1.4.27 In the presence of substrate 4, additional adduct peaks were observed. These effects were more pronounced for PLZ2.3 than for PLZ2.0, consistent with its higher turnover. Importantly, the unmodified protein remained the dominant species, and simple oxidation events are not expected to inactivate the enzyme significantly.
Finally, we performed a buffer screen to investigate the potential influence of buffer components on the photoreaction. Simple inorganic buffers such as phosphate and carbonate were unsuitable, as they form insoluble lanthanide salts. We thus tested a set of common Good's buffers and found that HEPES and MOPS performed equally well, while all others significantly decreased the activity (Fig. S12). Due to their various functional groups, including alcohol, amine, carboxylic acid, sulfonic acid, Good's buffers may interfere with redox chemistry,46 potentially serving as sacrificial reductants, HAT agents, or radical scavangers.47 Particularly relevant for our reaction is the reported formation of HEPES radicals in the presence of superoxide,48 which is generated in the proposed regeneration of Ce(IV) by molecular oxygen. The buffer may thus mediate the level of oxidative damage to the protein, but could also actively participate in the radical reaction mechanism. Furthermore, buffers containing diols or triols, such as Bicine, Tris, Tricine, may compete with the substrate, which is in agreement with the observed activity loss. Overall, it can be concluded that a fully innocent buffer system is not available, positive and negative effects are conceivable, and the choice of buffer was thus guided by optimal enzyme stability, activity, and stereoselectivity.
Structural analysis by X-ray crystallography and MD simulations
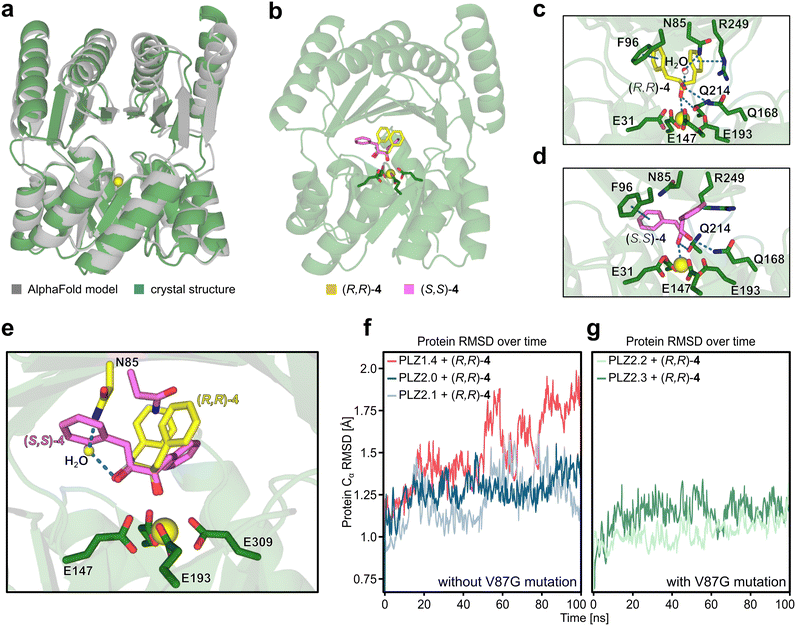
To further improve our understanding of the enzyme scaffold, we determined the crystal structure of PLZ2.3 bound to cerium(III) at a resolution of 2.1 Å (Fig. 3a). The experimental structure is in very good agreement with the AlphaFold3 model of our design output for PLZ2.0. The backbone RMSD of 1.59 Å originates from a slightly different orientation of both domains to each other. Also a template modeling (TM) score of 0.94 indicates a successful design.49 This is further supported by highly similar circular dichroism spectra of all PLZ variants (Fig. S18). As evident from the crystal structure, the size of the active site cavity is reduced compared to PLZ1.4 and the coordinated cerium(III) ion is located at the desired position.We next modeled PLZ1.4 using Boltz-2 (ref. 50) and generated models of PLZ2.0, PLZ2.1, and PLZ2.2 from the crystal structure of PLZ2.3. We docked the individual enantiomers of substrate 4 using Glide.40 We then performed molecular dynamics (MD) simulations to examine differences between the variants and rationalize the observed selectivity and epistasis (Fig. 3b–g). The cavity volume of the redesign was reduced as intended. Averaging the calculated pocket size over a 100 ns MD trajectory of the proteins with docked (R,R)-4 gave a volume of ca. 730 Å3 for PLZ2.3, compared to ca. 1640 Å3 for PLZ1.4, as calculated by MDpocket (Fig. S19).51 As a consequence, this enabled a more stable binding mode and distinct molecular interactions between protein and substrate. Comparison of (R,R)-4 and (S,S)-4 binding modes in PLZ2.3 indicates that both enantiomers position their hydroxyl groups similarly but differ in their aromatic ring orientation (Fig. 3b and e). While both retain hydrogen bonding to Q168 and Q214, as well as π–π stacking with F96, only the (R,R) enantiomer forms a N85-mediated hydrogen-bond network and a cation–π interaction with R249 during the simulation (Fig. 3c and d). These interactions are sterically inaccessible for the (S,S) enantiomer (Fig. 3e). The MD results thus indicate a full fixation of both hydroxyl groups and aromatic rings only for (R,R)-4. We therefore assume that N85 and R249 are crucial for the enantioselectivity of PLZ2.3. Additionally, distinct second-sphere interactions could influence the LMCT step, giving rise to slightly different activation barriers for the two enantiomers.
Furthermore, our MD simulations suggest that the stabilization of (R,R)-4 in the individual PLZ variants arises from distinct and partially mutually exclusive interaction networks, providing a structural rationale for the antagonistic epistasis observed in the double-mutant cycle analysis. PLZ2.0 shows a hydrogen bond between the substrate and Q214. Additionally, both phenyl rings of 4 are stabilized by cation–π interactions with R249 or π–π-interactions with W85 (Fig. S20). In PLZ2.1, (R,R)-4 is positioned through a transient water-mediated hydrogen-bond network involving N85, accompanied by a cation–π interaction between R249 and the substrate (Fig. S21). In PLZ2.2, the V87G mutation removes a steric restriction of L8, allowing it to shift inward and enabling a concerted loop rearrangement that positions Q6 for hydrogen bonding to the substrate (Fig. S24). This reorganization coincides with a pronounced stabilization of global RMSD over the MD trajectory, indicating increased scaffold stability, which is observed for both variants carrying the V87G mutation (Fig. 3f and g). While aromatic interactions with W85 are observed, PLZ2.2 lacks the R249 cation–π interaction (Fig. S22). In the double mutant PLZ2.3, W85 is mutated, and instead F96 engages in T-shaped π–π stacking with the substrate.52,53 In addition, (R,R)-4 forms hydrogen bonds to Q169 and Q214, a transient water-mediated interaction with N85, and a cation–π interaction with R249 (Fig. 3c and S23).
We attribute the non-additivity of the individual point mutations to the fact that the decisive hydrogen bond interactions for PLZ2.1 and PLZ2.2 are mutually exclusive: Q6 and the N85/Q169 network reside on opposite sides of the active site and cannot be engaged simultaneously (Fig. S25). Consequently, PLZ2.3 adopts the N85/Q169-based interaction pattern, explaining the lack of additive improvement in catalytic efficiency and the antagonistic epistasis observed in the double-mutant cycle analysis.
Overall, our results underline the successful protein redesign using AI-guided tools, in particular when comparing to PLZ1.4, which was based on a scaffold that originated from physics-based de novo design.28
Kinetic resolution of rac-4 using PLZ2.3
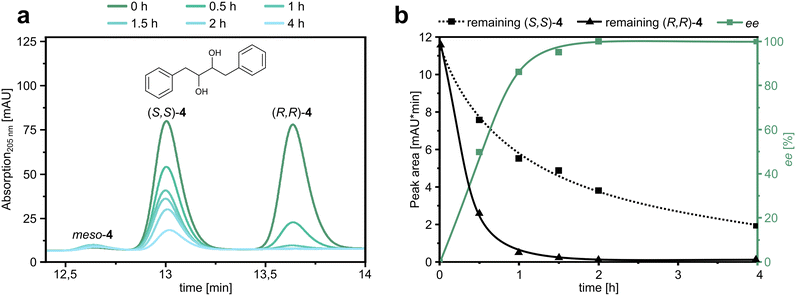
As a final experiment, we asked whether PLZ2.3 was selective enough to catalyze the kinetic resolution of rac-4. We irradiated a mixture of (R,R)-4 and (S,S)-4 in the presence of PLZ2.3 at a catalyst loading of 5 mol% and took samples at several time points to follow the consumption of the individual enantiomers (Fig. 4a). As expected, (R,R)-4 was consumed significantly faster than the (S,S)-enantiomer. After two hours, the ee of the remaining substrate approached 100%, as the (R,R)-enantiomer was completely consumed (Fig. 4b). The kinetic resolution experiment therefore yields enantiopure (S,S)-4 if the reaction is stopped after 2 h. However, due to the remaining enzymatic activity towards this disfavored enantiomer, the yields are much lower than practically desired.54 To increase the applicability in a scaled-up reaction, further improvement of the S value would be required. Nevertheless, this proof-of-concept experiment showed that the PLZ family of photoenzymes can be used in a stereoselective fashion.Conclusions
Photoenzymes are an emerging class of biocatalysts.55 Here we show that PhotoLanZymes – a family of lanthanide-dependent, de novo designed photoredox enzymes27 – can achieve enantioselective catalysis, which is a decisive feature for their future application. We combined AI-guided protein redesign and directed evolution to enhance both catalytic efficiency and enantiocontrol for a cerium-dependent radical C–C bond cleavage reaction that proceeds upon visible-light irradiation. The accessibility of powerful deep learning tools for protein design1–3 makes this a general approach to gain conformational stability and favorable dynamics in a de novo enzyme of interest, thereby generating viable starting points for directed evolution. Ongoing efforts concerning the PLZ family of photoredox enzymes focus on broadening their reaction scope to other lanthanide-dependent transformations, including radical C–C bond forming reactions.Author contributions
C. Z., A. S. K. and F. L. M. conceived the project. F. L. M. produced, evolved, and characterized the photoenzymes. J. B. performed the computational redesign and experiments for the revision. R. M. supported the filtering of initial design outputs. G. M. performed and evaluated the MD simulations. A. S. K. performed initial experiments with PLZ1.4. A. H. and M. G. performed protein crystallization and structure determination. F. L. M. and C. Z. wrote the manuscript. All authors have given approval to the final version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The crystal structure of PLZ2.3 has been deposited in the RCSB Protein Data Bank (PDB entry: 9SZU). All raw data files associated with this study will be openly available in the university's research data repository “mediaTUM” at https://doi.org/10.14459/2026mp1840493.Supplementary information (SI): SI figures, detailed procedures and results for all reported experiments, and data for compound characterization. See DOI: https://doi.org/10.1039/d5sc08010j.
Acknowledgements
This work was funded by an ERC Starting Grant (PhotoLanZyme, project number: 101039592). F. L. M. and R. M. received PhD fellowships of the German Academic Scholarship Foundation. A. H. received a PhD fellowship from “Stiftung der Deutschen Wirtschaft” (sdw). The project was further supported by the German Research Foundation (DFG, project number: 453748800). The authors are grateful to the staff members of the synchrotron beamline P13 of PETRAIII at DESY (Hamburg, Germany, proposal number MX-970) for technical support with crystallographic data collection and to the group of Martin Zacharias for access to the CPA in-house cluster to run MD simulations. The cluster infrastructure was funded by the German Research Foundation (DFG, project number: 439143547) via a Major Research Instrumentation Grant. We also acknowledge the Leibniz Supercomputing Center (LRZ) for providing computational resources and access to the Schrödinger software.Notes and references
- A. E. Chu, T. Lu and P.-S. Huang, Sparks of function by de novo protein design, Nat. Biotechnol., 2024, 42, 203–215 CrossRef CAS PubMed.
- T. Kortemme, De novo protein design - From new structures to programmable functions, Cell, 2024, 187, 526–544 CrossRef CAS PubMed.
- H. Y. Koh, Y. Zheng, M. Yang, R. Arora, G. I. Webb, S. Pan, L. Li and G. M. Church, AI-driven protein design, Nat. Rev. Bioeng., 2025, 3, 1034–1056 Search PubMed.
- S. Vázquez Torres, M. Benard Valle, S. P. Mackessy, S. K. Menzies, N. R. Casewell, S. Ahmadi, N. J. Burlet, E. Muratspahić, I. Sappington, M. D. Overath, E. Rivera-De-Torre, J. Ledergerber, A. H. Laustsen, K. Boddum, A. K. Bera, A. Kang, E. Brackenbrough, I. A. Cardoso, E. P. Crittenden, R. J. Edge, J. Decarreau, R. J. Ragotte, A. S. Pillai, M. Abedi, H. L. Han, S. R. Gerben, A. Murray, R. Skotheim, L. Stuart, L. Stewart, T. J. A. Fryer, T. P. Jenkins and D. Baker, De novo designed proteins neutralize lethal snake venom toxins, Nature, 2025, 639, 225–231 CrossRef PubMed.
- N. R. Bennett, B. Coventry, I. Goreshnik, B. Huang, A. Allen, D. Vafeados, Y. P. Peng, J. Dauparas, M. Baek, L. Stewart, F. Dimaio, S. De Munck, S. N. Savvides and D. Baker, Improving de novo protein binder design with deep learning, Nat. Commun., 2023, 14, 2625 CrossRef CAS PubMed.
- B. Liu, N. F. Greenwood, J. E. Bonzanini, A. Motmaen, J. Meyerberg, T. Dao, X. Xiang, R. Ault, J. Sharp, C. Wang, G. M. Visani, D. K. Vafeados, N. Roullier, A. Nourmohammad, D. A. Scheinberg, K. C. Garcia and D. Baker, Design of high-specificity binders for peptide–MHC-I complexes, Science, 2025, 389, 386–391 CrossRef CAS PubMed.
- S. Lee, R. D. Kibler, G. Ahn, Y. Hsia, A. J. Borst, A. Philomin, M. A. Kennedy, B. Huang, B. Stoddard and D. Baker, Four-component protein nanocages designed by programmed symmetry breaking, Nature, 2025, 638, 546–552 CrossRef CAS PubMed.
- K. Meador, R. Castells-Graells, R. Aguirre, M. R. Sawaya, M. A. Arbing, T. Sherman, C. Senarathne and T. O. Yeates, A suite of designed protein cages using machine learning and protein fragment-based protocols, Structure, 2024, 32, 751–765 CrossRef CAS PubMed.
- A. Lauko, S. J. Pellock, K. H. Sumida, I. Anishchenko, D. Juergens, W. Ahern, J. Jeung, A. Shida, A. Hunt, I. Kalvet, C. Norn, I. R. Humphreys, C. Jamieson, R. Krishna, Y. Kipnis, A. Kang, E. Brackenbrough, A. K. Bera, B. Sankaran, K. N. Houk and D. Baker, Computational design of serine hydrolases, Science, 2025, 388, eadu2454 CrossRef CAS PubMed.
- D. Listov, E. Vos, G. Hoffka, S. Y. Hoch, A. Berg, S. Hamer-Rogotner, O. Dym, S. C. L. Kamerlin and S. J. Fleishman, Complete computational design of high-efficiency Kemp elimination enzymes, Nature, 2025, 643, 1421–1427 CrossRef CAS PubMed.
- A. H.-W. Yeh, C. Norn, Y. Kipnis, D. Tischer, S. J. Pellock, D. Evans, P. Ma, G. R. Lee, J. Z. Zhang, I. Anishchenko, B. Coventry, L. Cao, J. Dauparas, S. Halabiya, M. Dewitt, L. Carter, K. N. Houk and D. Baker, De novo design of luciferases using deep learning, Nature, 2023, 614, 774–780 CrossRef CAS PubMed.
- J. S. Trimble, R. Crawshaw, F. J. Hardy, C. W. Levy, M. J. B. Brown, D. E. Fuerst, D. J. Heyes, R. Obexer and A. P. Green, A designed photoenzyme for enantioselective [2+2] cycloadditions, Nature, 2022, 611, 709–714 CrossRef CAS PubMed.
- K.-Y. Chen, H. Ming, H.-X. Wang, H.-Q. Wang and Z. Xiang, Genetic Incorporation of a Thioxanthone-Containing Amino Acid for the Design of Artificial Photoenzymes, Angew. Chem., Int. Ed., 2025, 64, e202419022 CrossRef CAS PubMed.
- X. Yang, J. Huang, J. Guo, S. Fang, Z. Wang, G. Wu, Y. Wu and F. Zhong, Bridging chemistry and biology for light-driven new-to-nature enantioselective photoenzymatic catalysis, Chem. Soc. Rev., 2025, 54, 5157–5188 RSC.
- N. Sun, J. Huang, J. Qian, T.-P. Zhou, J. Guo, L. Tang, W. Zhang, Y. Deng, W. Zhao, G. Wu, R.-Z. Liao, X. Chen, F. Zhong and Y. Wu, Enantioselective [2+2]-cycloadditions with triplet photoenzymes, Nature, 2022, 611, 715–720 CrossRef CAS PubMed.
- T. Vornholt, F. Leiss-Maier, W. J. Jeong, C. Zeymer, W. J. Song, G. Roelfes and T. R. Ward, Artificial metalloenzymes, Nat. Rev. Methods Primers, 2024, 4, 78 CrossRef CAS.
- F. Pirro, S. La Gatta, F. Arrigoni, A. Famulari, O. Maglio, P. Del Vecchio, M. Chiesa, L. De Gioia, L. Bertini, M. Chino, F. Nastri and A. Lombardi, A De Novo-Designed Type 3 Copper Protein Tunes Catechol Substrate Recognition and Reactivity, Angew. Chem., Int. Ed., 2023, 62, e202211552 CrossRef CAS PubMed.
- S. Mitra, D. Prakash, K. Rajabimoghadam, Z. Wawrzak, P. Prasad, T. Wu, S. K. Misra, J. S. Sharp, I. Garcia-Bosch and S. Chakraborty, De Novo Design of a Self-Assembled Artificial Copper Peptide that Activates and Reduces Peroxide, ACS Catal., 2021, 11, 10267–10278 CrossRef CAS PubMed.
- A. Lombardi, F. Pirro, O. Maglio, M. Chino and W. F. Degrado, De Novo Design of Four-Helix Bundle Metalloproteins: One Scaffold, Diverse Reactivities, Acc. Chem. Res., 2019, 52, 1148–1159 CrossRef CAS PubMed.
- D. W. Watkins, J. M. X. Jenkins, K. J. Grayson, N. Wood, J. W. Steventon, K. K. Le Vay, M. I. Goodwin, A. S. Mullen, H. J. Bailey, M. P. Crump, F. Macmillan, A. J. Mulholland, G. Cameron, R. B. Sessions, S. Mann and J. L. R. Anderson, Construction and in vivo assembly of a catalytically proficient and hyperthermostable de novo enzyme, Nat. Commun., 2017, 8, 358 CrossRef PubMed.
- I. Kalvet, M. Ortmayer, J. Zhao, R. Crawshaw, N. M. Ennist, C. Levy, A. Roy, A. P. Green and D. Baker, Design of Heme Enzymes with a Tunable Substrate Binding Pocket Adjacent to an Open Metal Coordination Site, J. Am. Chem. Soc., 2023, 145, 14307–14315 Search PubMed.
- A. Paredes, B. M. Loh, O. M. Peduzzi, A. J. Reig and K. M. Buettner, DNA Cleavage by a De Novo Designed Protein–Titanium Complex, Inorg. Chem., 2020, 59, 11248–11252 Search PubMed.
- A. Kawada, S. Mitamura and S. Kobayashi, Lanthanide trifluoromethanesulfonates as reusable catalysts: catalytic Friedel-Crafts acylation, J. Chem. Soc., Chem. Commun., 1993, 1157–1158 RSC.
- L. Yu, D. Chen and P. G. Wang, Aqueous aza Diels-Alder reactions catalyzed by lanthanide(III) trifluoro-methanesulfonates, Tetrahedron Lett., 1996, 37, 2169–2172 CrossRef CAS.
- Y. Qiao and E. J. Schelter, Lanthanide Photocatalysis, Acc. Chem. Res., 2018, 51, 2926–2936 CrossRef CAS PubMed.
- H. Pellissier, Recent developments in enantioselective lanthanide-catalyzed transformations, Coord. Chem. Rev., 2017, 336, 96–151 CrossRef CAS.
- A. S. Klein, F. Leiss-Maier, R. Mühlhofer, B. Boesen, G. Mustafa, H. Kugler and C. Zeymer, A De Novo Metalloenzyme for Cerium Photoredox Catalysis, J. Am. Chem. Soc., 2024, 146, 25976–25985 CrossRef CAS PubMed.
- S. J. Caldwell, I. C. Haydon, N. Piperidou, P.-S. Huang, M. J. Bick, H. S. Sjöström, D. Hilvert, D. Baker and C. Zeymer, Tight and specific lanthanide binding in a de novo TIM barrel with a large internal cavity designed by symmetric domain fusion, Proc. Natl. Acad. Sci. U. S. A, 2020, 117, 30362–30369 CrossRef CAS PubMed.
- Q. An, L. Chang, H. Pan and Z. Zuo, Ligand-to-Metal Charge Transfer (LMCT) Catalysis: Harnessing Simple Cerium Catalysts for Selective Functionalization of Inert C–H and C–C Bonds, Acc. Chem. Res., 2024, 57, 2915–2927 CrossRef CAS PubMed.
- J. Schwarz and B. König, Visible-light mediated C–C bond cleavage of 1,2-diols to carbonyls by cerium-photocatalysis, Chem. Commun., 2019, 55, 486–488 RSC.
- F. Juliá, Ligand-to-Metal Charge Transfer (LMCT) Photochemistry at 3d-Metal Complexes: An Emerging Tool for Sustainable Organic Synthesis, ChemCatChem, 2022, 14, e202200916 CrossRef.
- J. L. Watson, D. Juergens, N. R. Bennett, B. L. Trippe, J. Yim, H. E. Eisenach, W. Ahern, A. J. Borst, R. J. Ragotte, L. F. Milles, B. I. M. Wicky, N. Hanikel, S. J. Pellock, A. Courbet, W. Sheffler, J. Wang, P. Venkatesh, I. Sappington, S. V. Torres, A. Lauko, V. De Bortoli, E. Mathieu, S. Ovchinnikov, R. Barzilay, T. S. Jaakkola, F. Dimaio, M. Baek and D. Baker, De novo design of protein structure and function with RFdiffusion, Nature, 2023, 620, 1089–1100 CrossRef CAS PubMed.
- J. Dauparas, G. R. Lee, R. Pecoraro, L. An, I. Anishchenko, C. Glasscock and D. Baker, Atomic context-conditioned protein sequence design using LigandMPNN, Nat. Methods, 2025, 22, 717–723 CrossRef CAS PubMed.
- Z. Lin, H. Akin, R. Rao, B. Hie, Z. Zhu, W. Lu, N. Smetanin, R. Verkuil, O. Kabeli, Y. Shmueli, A. Dos Santos Costa, M. Fazel-Zarandi, T. Sercu, S. Candido and A. Rives, Evolutionary-scale prediction of atomic-level protein structure with a language model, Science, 2023, 379, 1123–1130 CrossRef CAS PubMed.
- J. Jumper, R. Evans, A. Pritzel, T. Green, M. Figurnov, O. Ronneberger, K. Tunyasuvunakool, R. Bates, A. Žídek, A. Potapenko, A. Bridgland, C. Meyer, S. A. A. Kohl, A. J. Ballard, A. Cowie, B. Romera-Paredes, S. Nikolov, R. Jain, J. Adler, T. Back, S. Petersen, D. Reiman, E. Clancy, M. Zielinski, M. Steinegger, M. Pacholska, T. Berghammer, S. Bodenstein, D. Silver, O. Vinyals, A. W. Senior, K. Kavukcuoglu, P. Kohli and D. Hassabis, Highly accurate protein structure prediction with AlphaFold, Nature, 2021, 596, 583–589 CrossRef CAS PubMed.
- M. Mirdita, K. Schütze, Y. Moriwaki, L. Heo, S. Ovchinnikov and M. Steinegger, ColabFold: making protein folding accessible to all, Nat. Methods, 2022, 19, 679–682 CrossRef CAS PubMed.
- J.-E. Sánchez-Aparicio, L. Tiessler-Sala, L. Velasco-Carneros, L. Roldán-Martín, G. Sciortino and J.-D. Maréchal, BioMetAll: Identifying Metal-Binding Sites in Proteins from Backbone Preorganization, J. Chem. Inf. Model., 2021, 61, 311–323 CrossRef PubMed.
- P. Wagner Egea, F. Delhommel, G. Mustafa, F. Leiss-Maier, L. Klimper, T. Badmann, A. Heider, I. Wille, M. Groll, M. Sattler and C. Zeymer, Modular protein scaffold architecture and AI-guided sequence optimization facilitate de novo metalloenzyme engineering, Structure, 2026, 34, 32–44 CrossRef CAS PubMed.
- R. K. Venkatraman, D. Ghosh, A. H. Tolba, T. I. Sølling, A. J. Orr-Ewing and A. M. El-Zohry, The Crucial Role of Solvent and Bound Oxygen on the Interplay of Cerium (III) and Cerium (IV) Chloride in a Photocatalytic Cycle for Organic Transformations, Angew. Chem., Int. Ed., 2025, 64, e202513444 CrossRef CAS PubMed.
- R. A. Friesner, J. L. Banks, R. B. Murphy, T. A. Halgren, J. J. Klicic, D. T. Mainz, M. P. Repasky, E. H. Knoll, M. Shelley, J. K. Perry, D. E. Shaw, P. Francis and P. S. Shenkin, Glide: A New Approach for Rapid, Accurate Docking and Scoring. 1. Method and Assessment of Docking Accuracy, J. Med. Chem., 2004, 47, 1739–1749 CrossRef CAS PubMed.
- H. Chen, B. Wu, T. Zhang, J. Jia, J. Lu, Z. Chen, Z. Ni and T. Tan, Effect of Linker Length and Flexibility on the Clostridium thermocellum Esterase Displayed on Bacillus subtilis Spores, Appl. Biochem. Biotechnol., 2017, 182, 168–180 CrossRef CAS PubMed.
- C. Sun, G. Li, H. Li, Y. Lyu, S. Yu and J. Zhou, Enhancing Flavan-3-ol Biosynthesis in Saccharomyces cerevisiae, J. Agric. Food Chem., 2021, 69, 12763–12772 CrossRef CAS PubMed.
- W. Wriggers, S. Chakravarty and P. A. Jennings, Control of protein functional dynamics by peptide linkers, J. Pept. Sci., 2005, 80, 736–746 CrossRef CAS PubMed.
- C. E. P. L. Brown, Hydrolysis of Metal Ions, Wiley-VCH Verlag GmbH & Co, KGaA, Germany, 2016 Search PubMed.
- B. M. Baker and K. P. Murphy, Evaluation of linked protonation effects in protein binding reactions using isothermal titration calorimetry, Biophys. J., 1996, 71, 2049–2055 CrossRef CAS PubMed.
- S. Bansal, M. Gori, J. A. Quaye, G. Gadda and B. Wang, Detection and Analysis of Reactive Oxygen Species (ROS): Buffer Components Are Not Bystanders, Anal. Chem., 2025, 97, 14931–14942 CrossRef CAS PubMed.
- B. R. Van Dyke and P. Saltman, Hemoglobin: A mechanism for the generation of hydroxyl radicals, Free Radical Biol. Med., 1996, 20, 985–989 CrossRef CAS PubMed.
- J. K. Grady, N. D. Chasteen and D. C. Harris, Radicals from “Good's” buffers, Anal. Biochem., 1988, 173, 111–115 CrossRef CAS PubMed.
- Y. Zhang and J. Skolnick, TM-align: a protein structure alignment algorithm based on the TM-score, Nucleic Acids Res., 2005, 33, 2302–2309 CrossRef CAS PubMed.
- S. Passaro, G. Corso, J. Wohlwend, M. Reveiz, S. Thaler, V. R. Somnath, N. Getz, T. Portnoi, J. Roy, H. Stark, D. Kwabi-Addo, D. Beaini, T. Jaakkola and R. Barzilay, Boltz-2: Towards Accurate and Efficient Binding Affinity Prediction, bioRxiv, preprint, 2025, DOI:10.1101/2025.06.14.659707.
- V. Le Guilloux, P. Schmidtke and P. Tuffery, Fpocket: An open source platform for ligand pocket detection, BMC Bioinf., 2009, 10, 168 CrossRef PubMed.
- Y. Zhao, J. Li, H. Gu, D. Wei, Y. C. Xu, W. Fu and Z. Yu, Conformational Preferences of π-π Stacking Between Ligand and Protein, Analysis Derived from Crystal Structure Data Geometric Preference of π-π Interaction, Interdiscipl. Sci. Comput. Life Sci., 2015, 7, 211–220 CrossRef CAS PubMed.
- R. Thakuria, N. K. Nath and B. K. Saha, The Nature and Applications of π–π Interactions: A Perspective, Cryst. Growth Des., 2019, 19, 523–528 CrossRef CAS.
- J. M. Keith, J. F. Larrow and E. N. Jacobsen, Practical Considerations in Kinetic Resolution Reactions, Adv. Synth. Catal., 2001, 343, 5–26 CrossRef CAS.
- M. A. Emmanuel, S. G. Bender, C. Bilodeau, J. M. Carceller, J. S. Dehovitz, H. Fu, Y. Liu, B. T. Nicholls, Y. Ouyang, C. G. Page, T. Qiao, F. C. Raps, D. R. Sorigué, S.-Z. Sun, J. Turek-Herman, Y. Ye, A. Rivas-Souchet, J. Cao and T. K. Hyster, Photobiocatalytic Strategies for Organic Synthesis, Chem. Rev., 2023, 123, 5459–5520 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |