Open Access Article
Open Access ArticleStructural basis for substrate-dependent allostery in oxygen activation by a cytochrome P450 enzyme revealed by analysis at different temperatures
Matthew N.
Podgorski†
 a,
Daniel P.
McDougal†
a,
Daniel P.
McDougal†
 b,
Eleanor C.
Campbell
c,
John B.
Bruning
b and
Stephen G.
Bell
b,
Eleanor C.
Campbell
c,
John B.
Bruning
b and
Stephen G.
Bell
 *a
*a
aDepartment of Chemistry, University of Adelaide, Adelaide, South Australia 5005, Australia. E-mail: stephen.bell@adelaide.edu.au
bSchool of Biological Sciences, University of Adelaide, Adelaide, South Australia 5005, Australia
cAustralian Synchrotron, 800 Blackburn Rd, Clayton, Melbourne, Victoria 3168, Australia
First published on 18th February 2026
Abstract
Cytochrome P450 (CYP) enzymes are ubiquitous and important monooxygenases whose archetypal reaction is to insert an oxygen atom from dioxygen into unactivated carbon–hydrogen bonds. They require the orchestrated delivery of electrons as well as protons from the solvent. The latter is controlled through an “acid–alcohol pair” of residues located above the heme though the precise details of proton delivery are unresolved. Here, using variable-temperature X-ray crystallography and all-atom molecular dynamics simulations of the bacterial CYP199A4 enzyme we demonstrate that the conformation of the acidic residue D251, of the “acid–alcohol” pair is allosterically coupled to the heme and the substrate. In general, and in common with other CYPs, the side chain of D251 favours the ‘out’ of the active site orientation. In this enzyme this overcomes incompatibility with hydrophobic residues. This side chain can rotate into the active site, and this is allosterically coupled to the presence of a distal heme ligand and other structural changes at the E-helix, C-terminal loop and on the proximal side of the heme. These and other structural changes can be related to differences in water molecule access to and egress from the distal side of the heme, which would facilitate proton delivery during the catalytic cycle. Comparison of the different environments of the side chain of D251 in CYP199A4 with the equivalent acidic residue in other diverse CYP enzymes suggest that there may not be a ‘universal’ model for proton transfer in CYP enzymes, but that allosteric effects and transient interactions are critically important.
Introduction
In nature an impressive number of diverse chemical reactions are performed by the cytochrome P450 superfamily (CYP) of enzymes.1,2 The origin of the name “P450” is their 450-nm absorption peak when carbon monoxide is coordinated to the ferrous heme-thiolate cofactor.3 As of 2019, >800![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 CYPs have been discovered, and this number will now surpass one million.4–6 Hydroxylation of C–H bonds is the most common activity catalysed by these enzymes, though they also catalyse many other types of reactions including C–C bond cleavage, desaturation, epoxidation, sulfoxidation, and N,O,S-dealkylation.2,7–12
000 CYPs have been discovered, and this number will now surpass one million.4–6 Hydroxylation of C–H bonds is the most common activity catalysed by these enzymes, though they also catalyse many other types of reactions including C–C bond cleavage, desaturation, epoxidation, sulfoxidation, and N,O,S-dealkylation.2,7–12
The catalytic cycle of CYP enzymes commences when a substrate enters the active site via a ligand access channel on the distal side of the heme.13,14 Upon ligand binding, the enzymes are generally more stable in a closed conformation and hydrophobic substrates usually eject the heme-bound water ligand switching the ferric heme iron from a 6-coordinate low-spin state (S = 1/2) to a 5-coordinate high-spin state (S = 5/2) (Scheme S1).15 The expulsion of the heme-bound aqua ligand and additional active site water molecules is proposed to occur through other channels in the enzyme.16 These changes alter the coordination environment and spin state of the heme, and permit more facile reduction of this cofactor to the ferrous state (Scheme S1).17,18 For most CYPs, these electrons are derived from NAD(P)H nicotinamide cofactors and delivered one at a time to the heme via electron transfer proteins.19 Next, the dioxygen (O2) ligand binds to the reduced heme and this ligand points towards a dioxygen-binding groove within the I-helix. This step is followed by delivery of the second electron to generate a ferric-peroxo anion (Scheme S1). Protonation of the distal oxygen generates the ferric-hydroperoxy intermediate, compound 0. These intermediates can also participate in some reactions such as certain C–C cleavage reactions.12,20 For most CYP reactions a second protonation triggers release of a molecule of H2O and formation of the active oxidant. This is a ferryl-oxo porphyrin radical cation, referred to as compound I (Cpd I),21 which hydroxylates organic molecules via Groves' radical rebound mechanism.22–24 The delivery of protons to the active site is essential for enzymatic activity, but given the diversity of sequence and function within this enzyme superfamily the precise molecular mechanism of this process and whether it is conserved across CYP enzymes is unresolved.
What is known is that in most instances, the activation of dioxygen to generate Cpd I (Scheme S1), including the delivery of the protons, is orchestrated by two active site residues located above the heme in the I-helix dubbed the “acid–alcohol pair”.25 This pair of residues, commonly either an aspartate or glutamate, and a threonine (or alternatively serine), have been extensively studied in the model, camphor hydroxylase, enzyme P450cam (CYP101A1).25–27 More recently, the role of the acid–alcohol pair in the dioxygen-binding groove has also been further studied, in the CYP199A4 enzyme from the bacterium Rhodopseudomonas palustris HaA2, which catalyses the efficient O-demethylation of 4-methoxybenzoic acid.28 In both P450cam and CYP199A4, the acid–alcohol pair comprise the D251 and T252 residues (Fig. S1a).25,28 However, the environment surrounding D251 and between P450cam and CYP199A4 is markedly different in sequence and in the types of residues which interact with the acid group.28 In both P450cam and CYP199A4 the threonine (T252) alcohol residue stabilises the hydroperoxy intermediate (Cpd 0) (Fig. S1b).27–29 If the threonine is replaced with a small hydrophobic residue the reducing equivalents are channelled into increased H2O2 production (the peroxide uncoupling pathway; Scheme S1) rather than substrate oxidation.28–32
There is compelling evidence that the D251 residues of P450cam and CYP199A4 are important for proton delivery.29 In these, and in several other CYP enzymes, if the acidic residue is mutated to a neutral asparagine, the rate of the catalytic cycle is typically slowed down by two orders of magnitude.27–29,31,33 This is consistent with a role in delivering protons from the bulk solvent to activate the dioxygen. However, in the X-ray crystal structures of these enzymes the aspartate side chain is located outside of the active site, thus it is presumed to rotate into the active site at the required steps of the catalytic cycle (Fig. S1c).27,34–36 However, an inward orientation of the acidic residue is rarely observed in crystal structures of CYP enzymes. For example, in P450cam, the aspartate residue D251 is held outside of the heme through salt bridge interactions with lysine and arginine residues (Lys178 and Arg186) and is restricted from moving into the active site.28,37 A notable exception is the cyano-complex of camphor-bound P450cam–Pdx complex in which the D251 side chain exists in two conformations: one pointing in and the other out of the active site (PDB ID: 6NBL).35,38 In P450cam allosteric effects on redox partner binding are proposed to generate a more open structure breaking these salt bridge interactions and enabling motion of the acidic residue (Fig. S2).34,35,38–41 This is in agreement with the strong effector role of the ferredoxin in this system, a distinctive feature of this CYP enzyme.35,39 Alternative studies have suggested that the D251 residue of P450cam may not be responsible for directly supplying protons to the active site, but that during the catalytic cycle water molecules enter the active site and deliver the required protons (Fig. S3).27,29,31 In comparison, X-ray crystal structures of CYP199A4 show that D251 is also located outside of the active site and points into a water channel which extends to the protein surface (Fig. S4).28,42 In contrast to P450cam, D251 is not immobilised by salt bridge interactions. In crystal structures of CYP199A4 with the D251N mutant, the side chain was observed to adopt alternate rotamer conformations, with some pointing into the active site and the flexibility of the side chain would be consistent with a role for the residue in this position in proton delivery (Fig. S4; PDB code: 7TQM and PDB code: 5KDY).20,28
The method of crystallisation for CYP199A4 is robust, and several examples of high-resolution crystal structures, of wild-type (WT) and mutants, with different substrates/inhibitors bound are available.11,20,42–46 X-ray crystallography is usually performed at cryogenic temperatures (100 K) to offer protection against rapid radiation damage. However, ligand occupancy and alternative conformations of residues may not be visible in crystals at cryogenic temperatures.47,48 These may be more readily assessed at higher temperatures and with all-atom molecular dynamics (MD) simulations.49–52 Here, we use variable temperature crystallography of CYP199A4 with different substrates combined with all-atom MD simulations to investigate and provide insight into the dynamics of the enzyme. These analyses focus on the D251 residue and have significant implications for proton delivery in this important and ubiquitous superfamily of enzymes.
Experimental
Materials and methods
3-Methylaminobenzoic acid (97% pure) was sourced from Sigma-Aldrich and 4-methoxybenzoic acid (>99% pure) was purchased from TCI (Tokyo Chemical Industry). Ethanol and DMSO were from Chem-Supply, and NVH immersion oil was from Cargille Laboratories. Tris and Bis–Tris were obtained from Astral Scientific, PEG 3350 was from Sigma-Aldrich, potassium chloride was from Chem-Supply, DTT was from Apollo Scientific, ammonium sulfate was from Ajax Finechem, kanamycin monosulfate was from Gold Biotechnology, and IPTG (isopropyl ß-D-1-thiogalactopyranoside) was from VWR. Crystals were mounted onto MicroLoops or MicroMounts from MiTeGen. UV-vis spectroscopy was performed using an Agilent Cary 60 UV-vis spectrophotometer.Production and purification of CYP199A4
The CYP199A4 enzyme was produced and purified according to a procedure similar to that published previously.28,53 The final purity ratio (Reinheitszahl (RZ) = A419/A280) of the CYP199A4 enzyme was ∼2.1. The enzyme was filter-sterilized and stored in ∼50% glycerol at −20 °C.X-ray crystallography
The previously published procedure was followed to obtain crystal structures of each CYP199A4-substrate combination.20,43,46,54 Glycerol and salt were first removed from the CYP199A4 stock solution using a PD-10 desalting column (Cytiva), with Tris–HCl buffer (50 mM, pH 7.4) used as the eluent. The concentration of CYP199A4 was determined by UV-vis spectroscopy using the Soret band extinction coefficient reported in the literature (ε419 = 119 mM−1 cm−1).42 The CYP enzyme was subsequently concentrated to ∼40 mg mL−1 (∼900 µM) via ultrafiltration, using a centrifugal device containing a 10 kDa MWCO (molecular weight cut-off) membrane. The concentrated protein was then saturated with substrate (3–3.2 mM), which was added from a 100 mM stock solution in DMSO/EtOH. The hanging-drop vapor-diffusion method was used to grow crystals of each CYP199A4-ligand complex, using 24-well crystallisation trays.44 Each drop (consisting of 1.2 µL of the protein–substrate complex mixed with 1.2 µL of the reservoir solution) was suspended on a coverslip and equilibrated at 16 °C against 500 µL of reservoir solution. Each well was sealed with Glisseal high vacuum grease. The reservoir solution consisted of 100 mM Bis–Tris buffer (pH 5.00, 5.25, 5.50 or 5.75), 200 mM magnesium acetate and 20, 23, 26, 29, 30.5 or 32% PEG (polyethylene glycol) 3350.44 Crystals grew rapidly under these conditions (<24 hours) and were typically left for several weeks before harvesting.Crystals were mounted onto MicroLoops or MicroMounts, cryoprotected in NVH oil, and flash-cooled in liquid N2 for transportation to the Australian synchrotron. All X-ray diffraction data was collected at the MX2 beamline.55,56 The X-ray wavelength was 0.9537 Å. Initially, diffraction data for each CYP199A4 crystal was collected at 100 K. The temperature of each crystal was subsequently raised in 50 K increments to 200 K, and a dataset was collected at each temperature (once the temperature of the cryojet had stabilised). If the crystal diffracted well, datasets were also collected at 250 and 300 K.
The order of the variable-temperature experiment was reversed with a crystal of CYP199A4 bound to 3-methylaminobenzoic acid. Initially, diffraction data was collected at 200 K, and the temperature of the crystal was then reduced in 50 K increments to 100 K, and a dataset was collected at each temperature (data is provided in the SI). In these crystal structures (PDB entries: 9PLS, 9PMA and 9PMC), a small difference map density peak was visualized near the heme. This could correspond to a low occupancy water. However, addition of a water molecule in the structure and subsequent refinement led to a very low occupancy water at that position which did not justify modelling.
The diffraction data was automatically processed on the MX2 beamline using xdsme57 and AIMLESS58 and the phase problem was then overcome using the molecular replacement method. The program PhaserMR57,59 was used for the molecular replacement step, and a previously reported structure of CYP199A4 (PDB ID: 5UVB)45 with ligands and water molecules deleted was used as the search model. Coordinates and restraints for the substrate were obtained using Phenix eLBOW.60 Refinement of the structure was performed using phenix.refine.61 Where the mFo − DFc difference map indicated that adjustments should be made to the structure (e.g., addition of the substrate, heme, the chloride ion,42 and alternative protein conformations), these changes were made using Coot.62 This was followed by further rounds of refinement and manual adjustments. Positive density in the mFo − DFc difference map revealed the presence of a second conformation of D251 (in which the side chain had rotated into the active site) in certain crystal structures at 100 K, but this second conformation disappeared at higher temperatures. The occupancies of the substrate, heme-bound aqua ligand and alternative protein conformations were all refined. The restraints used for the Fe–S and Fe–O bond lengths were 2.3 ± 10 Å and 2.2 ± 10 Å. MolProbity63 was used to assess the quality of each model, and the final model was then deposited into the Protein Data Bank (PDB; https://www.rcsb.org/). The PDB code and statistics for each structure are presented in the SI. Because each crystal was used to collect multiple datasets and radiation damage may therefore be an issue, each structure was examined in Coot for any obvious evidence of radiation damage (e.g., disappearance of electron density due to decarboxylation of aspartate/glutamate side chains).51,64 2mFo − DFc composite omit maps were generated using the program phenix.composite_omit_map in the Phenix software suite, using the default “simple” method. Feature-enhanced maps were also generated using Phenix.65–67 Images of the final structures were generated using PyMOL.68,69 The structures were analysed using UCSF Chimera.70
Molecular dynamics simulations
Variable temperature all-atom molecular dynamics (MD) simulations were performed using GROMACS to complement the variable temperature crystallography experiments.68 In brief, the simulations were performed in triplicate at 100 K, 150 K, 200 K and 300 K for 3-methlyaminobenzoic acid and 4-methoxybenzoic acid substrates. All simulations began from the starting coordinates which were that of the 100 K crystal structures; the ‘in’ conformation of the D251 side chain was used for the 3-methylaminobenzoic acid simulations, and all crystallographic waters were retained for both substrates, as well as the Cl-ion. Amber parameters for the ferric heme and axial cysteine ligand were borrowed from Shahrokh et al. and the parameters for the substrates were prepared using Antechamber; hydrogens were added to the protein using the H++ server (pH 7.0) and the complex was assembled using tleap housed in the AmberTools23 software package.71–73 The complex was placed in the centre of a cubic box and solvated with the TIP3P water model and charge neutralised with the addition of Na+ ions.74 Input files were generated for use in GROMACS, and energy minimisation was performed using the steepest descent algorithm (energy minimisation was performed once for use in all temperature simulations). At this point, each system was equilibrated with 1 ns NVT restrained simulation using PME electrostatics, Berendsen thermostat coupling, Lincs constraint algorithm (hydrogens); followed by 1 ns NPT restrained simulation using PME electrostatics, Berendsen thermostat, Lincs constraint algorithm (hydrogens) and Parrinello–Rahman pressure coupling.75–78 The substrate and heme, along with protein, were restrained during system equilibration. The restraints were then released and 1 µs production simulations were performed in triplicate (totalling 24 µs simulation time); frames were saved every 20 ps and a 2 fs timestep was used. The simulations were performed using Nvidia A100 GPUs on the University of Adelaide Phoenix HPC cluster. Analyses of the simulations were conducted using MDTraj (stride = 5 i.e., frames every 100 ps were analysed for memory efficiency) and k-means clustering using Scikit-learn (https://scikit-learn.org).79Results
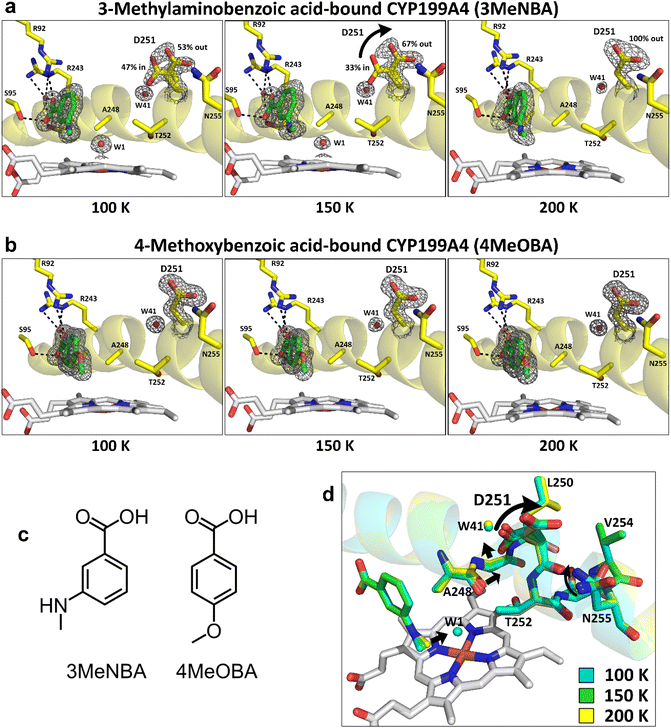
We first assessed the available structures of CYP199A4 in the Protein Data Bank (PDB) and searched for alternate conformations of the side chain of the D251 residue (Fig. S5 and S6). This residue points out of the active site into a solvent channel in the structure with 4-methoxybenzoic acid and in most other crystal structures (Fig. S4). However, we observed differences in the X-ray crystal structures of CYP199A4 with 4-(furan-2-yl)benzoic acid, 4-(1H-1,2,4-triazol-1-yl)benzoic acid and 3-methylaminobenzoic acid (Fig. S5 and S6, PDB codes: 4DO1, 6PRR, 7TRT, and 7N14).42,53,54,80 In each of these X-ray crystal structures the electron density of the D251 residue demonstrates that it can rotate into the active site of the enzyme. Based on these available structures we chose to further characterise the crystal structures of CYP199A4 with 3-methylaminobenzoic acid ligand at different temperatures. This was compared to the CYP199A enzyme with 4-methoxybenzoic acid, which is the physiological substrate for the enzyme and is O-demethylated. When 4-methoxybenzoic acid (4MeOBA) binds to CYP199A4, it displaces the heme-bound aqua ligand and induces a virtually complete spin-state shift from low-spin to ≥95% high-spin (HS). In contrast, 3-methylaminobenzoic acid (3MeNBA)-bound CYP199A4 is predominantly low-spin (only ∼10% high-spin). This is in agreement the heme environments observed in the X-ray structures. While 3-methylaminobenzoic acid binds in an inhibitory manner it can be oxidised via N-demethylation by the CYP199A4 enzyme.53In crystallo movement of D251 in CYP199A4 bound to 3-methylaminobenzoic acid
X-ray diffraction data was collected at 100 K for a crystal of CYP199A4 bound to 3-methylaminobenzoic acid (Table S1). The solved crystal structure was similar to the previously determined structure (PDB code: 6PRR) (Fig. S7, S8, and Table S2) and the D251 side chain is present in two conformations. In one conformation (∼50% occupancy), the D251 side chain faces out of the active site into the solvent channel (Fig. 1, S6 and S9). In the other conformation, the D251 side chain has rotated into the active site (∼50% occupancy). The temperature of this crystal was then raised in 50 K increments, and a dataset was collected at each temperature up to 200 K (no solvable structural data were obtained above 200 K). As the temperature is raised to 150 and 200 K, the D251 side chain rotates out of the active site (Fig. 1, S10, S11 and Table S3). At 200 K, the D251 side chain points into the solvent channel (100% occupancy) and the electron density for the second conformation pointing into the active site disappears (Fig. S9–S14).In the conformation with the D251 side chain in the active site, it interacts with an ordered water molecule, Wat41 (3.2 Å) which is located within the I-helix, and the backbone NHs of D251 (2.9 Å) and T252 (2.9 Å) (Fig. S15a). In the conformation in which the D251 side chain points out of the active site into the solvent channel, it interacts with the side chain of N255 (3.4 Å) and water molecules in the channel (Fig. S15a). The side chain of N255 is also present in two conformations at 100 K with occupancies of 64% (interacting with D251) and 36% (Fig. S15a, d, and Table S3). At higher temperatures (200 K), when the D251 side chain has rotated out of the active site, the electron density of N255 has 100% occupancy for the conformation that interacts with D251. The other residues in the active site did not move significantly, the electron density of Wat41 is retained, and the overall structure of the I-helix was not disrupted by the movement of D251 suggesting that the side chain of this aspartate residue has a higher degree of mobility compared to other residues and even the substrate (Fig. S16).
As the temperature increases the other significant change within the active site is related to the heme-bound water ligand. In the 100 K crystal structure, a high-occupancy (∼90% occupancy) heme-bound aqua ligand is present (Fig. 1 and Table S4). This water ligand is hydrogen-bonded to the nitrogen of 3-methylaminobenzoic acid (2.9 Å). However, the electron density associated with this heme-bound aqua ligand decreases in the crystal structures at 150 K and 200 K (Fig. 1 and Table S4). In the 200 K structure the electron density suggests that this aqua ligand is absent. This could be due to gradual photoreduction of the heme iron in the X-ray beam to the five-coordinate ferrous (Fe2+) form,81,82 which would result in the heme-bound water ligand being released.81–83 Another possibility is that the water is loosely bound. Increasing the temperature may result in disappearance of the electron density as the water ligand would be expected to more mobile at higher temperatures. The UV-vis absorbance spectrum at room temperature suggests the predominance of a 6-coordinate ferric heme species.53 As the heme-bound aqua ligand disappears, the methylamino substituent of the substrate rotates slightly towards the heme (Fig. S16 and S17). When the aqua ligand is present the carbonyl of the A248 residue forms a hydrogen bond with it (O–O distance: 3.2 Å; Fig. S17) and the iron is more in the plane of the porphyrin ring (Table S5, and Fig. S18).
We assessed if our crystal structure of CYP199A4 with 3-methylaminobenzoic acid had suffered from radiation damage during data collection as the temperature is raised. Acidic aspartate and glutamate residues are susceptible to decarboxylation, and methionine residues may undergo cleavage of the C–S bond.84 Tyrosine residues may also may lose their OH group (Fig. S19). Comparison of the electron density maps for selected Asp/Glu residues in the three structures demonstrated that these residues do not appear to have undergone obvious decarboxylation (Tables S6–S8). To investigate whether the electron density of the heme-bound aqua ligand was disappearing due to the increase in temperature or due to X-ray induced photoreduction of the heme iron, we reversed the experiment by first collecting a dataset at high temperature (200 K). We then reduced the temperature and collected subsequent datasets at 150 K and 100 K (Tables S9, S10, and Fig. S20–S23). In the first dataset collected at 200 K, no heme-bound aqua ligand was present and the D251 side chain pointed entirely out of the active site into the solvent channel (Fig. S21). When the temperature was reduced to 150 K and 100 K, electron density for the heme-bound aqua ligand did not appear and the side chain of D251 did not rotate into the active site (Fig. S22 and S23). For comparison, the active site water molecule equivalent to Wat41, which interacts with D251 when it points into the active site, was again present in the structures at all temperatures in both sets of experiments (Fig. S21–S23, and Table S3). Based on these results we propose that the aqua ligand may disappear due to X-ray-induced photoreduction of the heme to generate the five-coordinate ferrous form.81
To further investigate how the position of the D251 residue is related to these phenomena we did variable temperature crystallography with CYP199A4 bound to 4-phenoxybenzoic acid (Tables S11–S15, and Fig. S24–S28).85 The crystal structure of CYP199A4 with this ligand contains a tightly bound heme-bound aqua ligand which interacts with the ether oxygen of 4-phenoxybenzoic acid. In the X-ray crystal structure at 100 K D251 was present in two conformations with occupancies of 27% (into the active site) and 73% (out of the active site). As the temperature was raised to 150 K and 200 K the electron density of D251 indicated it was only in the orientation which points out of the active site. At these elevated temperatures the heme aqua ligand was maintained (70% at 100 K and 59% at 200 K. Table S13). This infers that the movement of the D251 side chain is due to the increase in temperature and not X-ray-induced photoreduction.
Variable-temperature structures of 4-methoxybenzoic acid-bound CYP199A4
We next repeated these experiments for crystals of CYP199A4 bound with the physiological substrate 4-methoxybenzoic acid to assess the relationship between D251 conformation and substrate. Previously reported X-ray crystal structures of CYP199A4 determined with this substrate show the side chain of D251 pointing out of the active site into the solvent channel and interacting with N255.28 The crystal structure of wild-type CYP199A4 in complex with 4-methoxybenzoic acid was solved at temperatures of 100, 150, 200 and 300 K with each dataset being collected using the same crystal (Fig. 1 and S29–S38, and Tables S16–S18). The resolution of the 100 K structure (1.26 Å) was higher than that of the previously determined structure (see Fig. S36, S37, Tables S19 and S20). The resolution decreased (1.26–1.50 Å at 100–200 K and 2.49 Å at 300 K) as the temperature was raised and the crystal was repeatedly irradiated (Fig. S29–S32, and Table S16). However, at each temperature, a single conformation of both D251 and N255 are observed (Fig. S34). Importantly, no heme ligated aqua ligand was observed at any temperature. The electron density maps were visually inspected for evidence of radiation damage e.g., decarboxylation of Asp/Glu residues (Tables S18, S21 and S22). At 300 K, the electron density did not completely disappear for the majority of the Asp/Glu side chains examined but the electron density was poorer particularly for residue D251, compared to nearby residues such as F182 (Fig. S35) which could be due to increased mobility. Negative density regions appeared in the difference map, which may be due to the lower resolution or radiation damage, or both (Table S18). There was no evidence of movement of the D251 side chain into the active site as the temperature of the crystal of 4-methoxybenzoic acid-bound CYP199A4 was raised (Fig. S34).The variable-temperature X-ray crystallography of CYP199A4 with the 3-methylaminobenzoic acid bound demonstrated that the D251 side chain is flexible in this system and can rotate in and out of the active site. This agrees with the proposed role of D251, in assisting in delivering protons to the active site from bulk solvent during the catalytic cycle. However, D251 was stable in the ‘out’ position in the X-ray crystal structures of CYP199A4 with 4-methoxybenzoic acid and did not move as the temperature was modified, suggesting that dynamics of D251 and neighboring residues in the I-helix could be substrate-dependent. To investigate this further, as well as identifying potential sources of protons from pathways of solvent flux, we turned to all-atom MD simulations.
Variable temperature molecular dynamics simulations
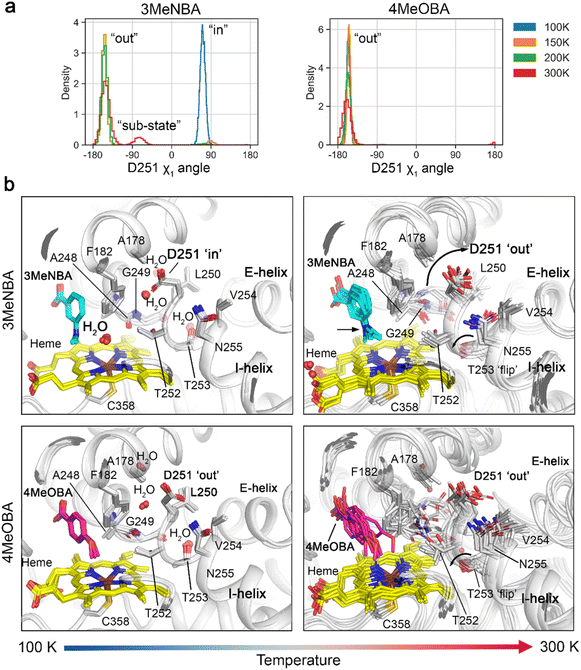
All-atom MD simulations, 1 µs, were designed and performed in triplicate to replicate the variable temperature X-ray crystallography experiments (totalling 24 µs). Each simulation was undertaken using the starting coordinates of the 100 K crystal structures of CYP199A4 with 3-methylaminobenzoic acid and 4-methoxybenzoic acid, respectively. For the 3-methylaminobenzoic acid experiments, the ‘in’ rotamer of the D251 side chain was selected as the starting conformation. The temperature was maintained at 100 K or raised to 150 K, 200 K and 300 K with appropriate system equilibration for each temperature increase. The MD simulations revealed that temperature has a profound effect on local and tertiary protein dynamics. As the temperature increased, the CYP199A4 backbone and residues became progressively more dynamic (Fig. S39 and S40). We found that the two substrates, 3-methylaminobenzoic acid and 4-methoxybenzoic acid, differentially affect the dynamics of the enzyme. For example, the oxygen-binding groove of the I-helix adjacent to the substrate and E-helix are markedly more dynamic or disordered when 4-methoxybenzoic acid is the substrate as compared to 3-methylaminobenzoic acid. In the cryogenic 3-methylaminobenzoic acid crystal structures (100 K and 150 K), a heme-bound aqua ligand is present which forms an additional hydrogen bond with the carbonyl of A248 (Fig. S15). 4-Methoxybenzoic acid displaces this aqua ligand, abolishing a link between the heme-centre and the I-helix (Fig. S37). The χ1 dihedral angle of the D251 residue was analysed to measure the different conformational states of the side chain as temperature was increased (Fig. 2).In the 3-methylaminobenzoic acid simulations at 100 K, the D251 side chain remained stable in the ‘in’ conformation. Here it rests against A178 and F182; these interactions would be expected to be unfavourable but may be better tolerated under cryogenic and ultra-low energy conditions. When the temperature was raised to 150 K and above the side chain switched to the ‘out’ conformation. This switch is directly coupled to the presence (“in”) or absence (“out”) of the aqua ligand bound to the heme centre which when lost, increases the motion of the I-helix around the oxygen-binding groove (Fig. 2). Once in the ‘out’ conformation, the D251 side chain remained stable cradled within a ring of neighbouring hydrophobic and non-polar residues including L175, A178, G179, F182, L50, V254 and L396. At 300 K it was observed to sample a ‘minor’ conformational sub-state in the 3-methylaminobenzoic acid simulations (Fig. 2). Importantly, across all simulations once D251 was in the ‘out’ conformation the side chain was not observed to rotate into the active site. To rotate inward, the acidic side chain must overcome a significant barrier imposed by the A178 and F182 residues and to do so, would likely require a global conformational change.
The simulation data further showed that in all 4-methoxybenzoic acid simulations, D251 recapitulated the crystallographic conformation facing outward into the solvent channel, remaining highly stable and did not sample a minor alternative substate (Fig. 2). In these MD simulations, the absence of the aqua ligand when 4-methoxybenzoic acid is bound correlated with an increase in mobility of the I-helix (Fig. S40 and S41).
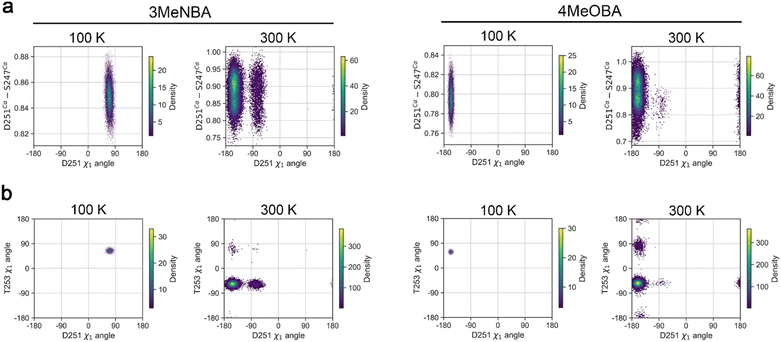
While the MD simulations show that the I-helix is more dynamic when 4-methoxybenzoic acid is the substrate, the position of the side chain of D251 is more stable when compared to what is found with 3-methylaminobenzoic acid. In the 3-methylaminobenzoic acid simulations, D251 and N255 only hydrogen bond transiently but in the simulations with 4-methoxybenzoic acid a weak hydrogen bonding interaction is maintained throughout. To explore any potential implications for proton delivery, we analysed the distances between the Oδ1 atom of D251 to the Nδ2 of N255 and the Cγ of T252; the width of the oxygen-binding groove between Cα atoms of S247 and D251; and the dihedral angles of the T252, T253 and N255 residue (Fig. 3, S42 and S43).
The greater motion of D251 in the 3-methylaminobenzoic acid simulations was also apparent from the distance between D251 and T252 which fluctuates as the temperature increases. This distance was stable when 4-methoxybenzoic acid was bound, but more significant differences were observed at 300 K coinciding with conformational change of the oxygen-binding groove. When D251 switches to the ‘out’ conformation the oxygen-binding groove widens by approximately 0.1 nm, and this increases the solvent flux surrounding the I-helix and the loss of water molecules from the active site (Fig. S44).
In the 300 K MD simulations of 4-methoxybenzoic acid, T252 was observed to flip with the OH group projecting upward toward D251. While this was a relatively rare event in the course of the simulations, when with an inward rotation of protonated D251, the OH of T252 could be in a position to receive a proton (Fig. S45).
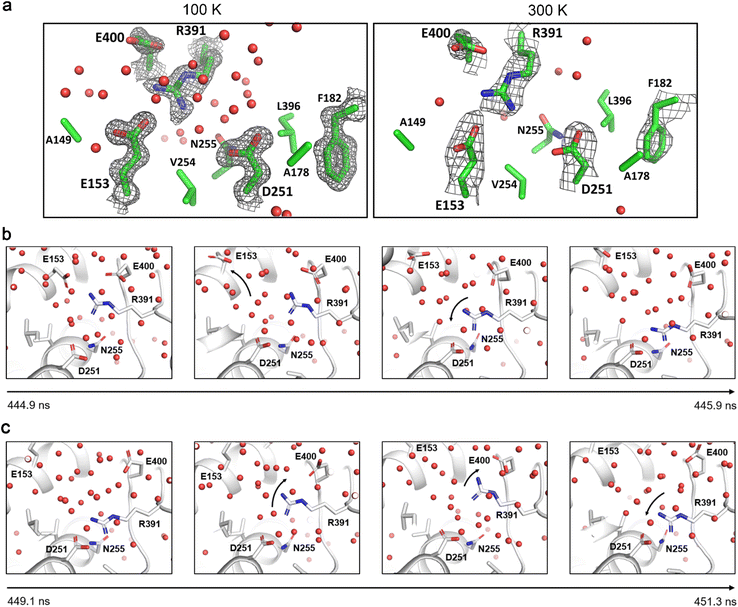
We next analysed the whole structure and identified a network of water molecules and salt bridges linking the D251 residue to E153 (in the E-helix), R391 and E400 (both in the C-terminal loop) (Fig. 4). In the X-ray crystal structures, the electron density for each residue and the water are well defined in all of the 3-methylaminobenzoic acid and 4-methoxybenzoic acid-bound forms at ≤ 200 K. However, they are less clear in the 300 K 4-methoxybenzoic acid structure (Fig. S32 and S46). Importantly, the water network is present in the 100 K and 150 K 3-methylaminobenzoic acid structures in which D251 is observed in both the ‘in’ and the ‘out’ states, but the location differs when compared to those in the 4-methoxybenzoic acid-bound structures (Fig. S46).
Analysis of the Cα-atom RMSF data showed the E-helix, which contains the E153 residue, is highly dynamic in the 300 K 4-methoxybenzoic acid simulations. In one of the 4-methoxybenzoic acid simulations the salt bridge between E153 and R391 is broken upon destabilisation of the E-helix, which is initiated by L250 of the I-helix clashing with the residues P156, L157 and F160. This liberates R391 to form an intermediary interaction with E400 before switching its position to point into the solvent channel and form a more long-lived salt bridge with D251 (Fig. 4). Several nanoseconds later, the interaction between D251 and R391 is broken and the sidechain of R391 transiently interacts with E400, before dissociating and re-forming a longer-lived salt bridge with D251 (Fig. 4). This switching of the salt bridge network was observed in one of the three replicates and occurred over approximate 1–2 ns timescales. Notably, this switching of the network and longer-lived salt bridge formation with D251 is also observed in one of the 300 K 3-methylaminobenzoic acid replicates. In all the other 4-methoxybenzoic acid and 3-methylaminobenzoic acid replicates the salt bridge between E153 and R391 also breaks frequently, which would allow the free amide to transfer the proton to D251 via water molecules rather than directly interacting with D251 (Fig. S47–S49). Thus, these interactions could provide a proton-relay system which is not observed crystallographically.
Our data show that D251 and the putative proton-relay are allosterically coupled to substrate via movement of the I-helix and E-helix. The conformational changes in the oxygen-binding groove, particularly when 4-methoxybenzoic acid is bound, introduces clashes between L250 on the I-helix and P156, L157 and F160 on the E-helix, resulting in destabilisation and an increased likelihood of breaking the E153 and R391 salt bridge. Interestingly, the analysis also revealed that when the I-helix undergoes conformational change at 300 K – for both substrates – the side chain of T253 is flipped inwards toward the heme (Fig. 2 and S44). When this change occurs the T253 residue is in a similar orientation to that observed in the X-ray crystal structure for the equivalent V253 residue in P450cam (PDB ID: 1DZ4).15 This change in position of T253 breaks a hydrogen bond network with a single water molecule and the E367 residue of the L-helix. This changes the position of the E367 side chain which flips to form a hydrogen bond with Y155 of the E-helix. In some instances, the hydrogen bond between the E367 carboxyl and Y155 phenol is broken, the latter then hydrogen bonds with the E367 backbone carbonyl instead. These changes result in the creation of a new solvent channel between the heme and the protein surface between Y155 and V159, correlating with the large increases in RMSF of the D-helix (Fig. S41 and S44). This channel could be involved in the expulsion of water from the active site or even be an alternative pathway for water molecules to enter and possibly deliver protons.
Together, these findings indicate that the I-helix participates in long-range allostery with other regions of the CYP199A4 enzyme, likely increasing the efficiency of proton delivery to and solvent efflux from the active site. While the solvent channel is created in both 3-methylaminobenzoic acid and 4-methoxybenzoic acid simulations, correlating to conformational switching of D251 and T253, the allostery between the I-helix and the E-helix, resulting in initiation of the proton relay occurs in a substrate-dependent manner. These findings could explain the differences in catalytic efficiency of the enzyme with closely related substrates.53
Discussion
X-ray crystallography experiments with CYP199A4 revealed alternate ‘in’ and ‘out’ conformations of D251 under cryogenic conditions when 3-methylaminobenzoic acid was the substrate. The different conformations were coupled to the presence of the distal heme-aqua ligand. Partial rotation of D251 into the active site has also been observed in crystal structures of CYP199A4 with certain ligands (products and inhibitors) bound that do not displace the heme-aqua ligand (PDB ID: 8VKF, 8VOT, 6PRR, 7TRT, and 7N14).53,54,80,86,87 Variable temperature experiments demonstrated that when the temperature was raised to 200 K, D251 switched completely to the ‘out’ orientation and the electron density of the heme-aqua ligand disappeared. In the X-ray crystal structures of 4-methoxybenzoic acid the D251 side chain was in the ‘out’ position at all temperatures. In the ‘out’ orientation the D251 side chain interacts with N255 and is also surrounded by ring of hydrophobic and non-polar residues. In fact, the D251 side chain shields A178 and F182 from solvent in the ‘out’ orientation. When D251 is oriented in the ‘in’ conformation the acidic carboxyl rests against the hydrophobic and non-polar side chains of A178 and F182 and the N255 side chain rotates away from its other position by ∼90°.28 Recent studies demonstrate when the bulky F182 residue is replaced with alanine, glycine or threonine that the position of D251 can rotate into the active site (PDB ID: 8WS4, 9DY9, 9DY5, 8HGB, and 8HGC).88–90When the temperature of each crystal was raised there was the loss, or a reduction, in the electron density of the heme-bound aqua ligand (if present). We concluded that this was most likely due to gradual X-ray-induced photoreduction of the ferric heme-iron to the five-coordinate ferrous form.81,82 Reduction of the heme would result in release of the aqua ligand.91 An alternative reason for this observation could be that the low-spin six-coordinate ferric species is favoured at lower temperatures. This would be in agreement with EPR measurements which are also undertaken at low temperatures and which show a lower proportion of high-spin ferric heme when compared to UV-vis absorbance experiments undertaken at ambient temperatures.92 We only observe the D251 side chain pointing into the active site when there was electron density associated with an aqua ligand bound to the heme iron. In crystal structures of WT CYP199A4 bound to 4-methoxybenzoic acid that completely displaces the heme-bound aqua ligand we detected no evidence of the side chain of D251 pointing into the active site. The reduction of 5-coordinate high-spin ferric CYPs is significantly easier than in the 6-coordinate low-spin state.20 It is important to note here that the D251 residue would be required to deliver protons after dioxygen has bound to ferrous heme and that the presence of this heme ligand may facilitate this process.
In both sets of X-ray structures the electron density associated with the substrate and most other active site residues of CYP199A4 did not significantly change as the temperature was raised. This implies that the side chain of D251 has a higher degree of mobility than the other active site residues. This movement is consistent with the proposed role of D251, which is to transfer protons from external solvent to iron-bound dioxygen and the ability to move in and out of the active site would facilitate this process.27 In the X-ray crystal structures the overall structure of the I-helix was not greatly changed by the motion of D251.28 There was a subtle shift in secondary structure of the I-helix around the oxygen-binding groove that removes a weak hydrogen bond between the heme-aqua ligand and carbonyl of A248. Substrate dependent changes in the I-helix structure were observed in the MD simulations at 300 K, and these were linked to other structural changes throughout the enzyme. In the simulations of CYP199A4 starting with a 5-coordinate heme and 4-methoxybenzoic acid-bound, allostery was observed between the I-helix and E-helix, resulting in breaking and restructuring of the putative proton relay between E153, D251, R391 and E400. In the simulations with 3-methylaminobenzoic acid the presence of a distal water ligand to the heme resulted in less pronounced allostery between the I-helix and E-helix and only resulted in the transient breaking of the salt bridges in this network.
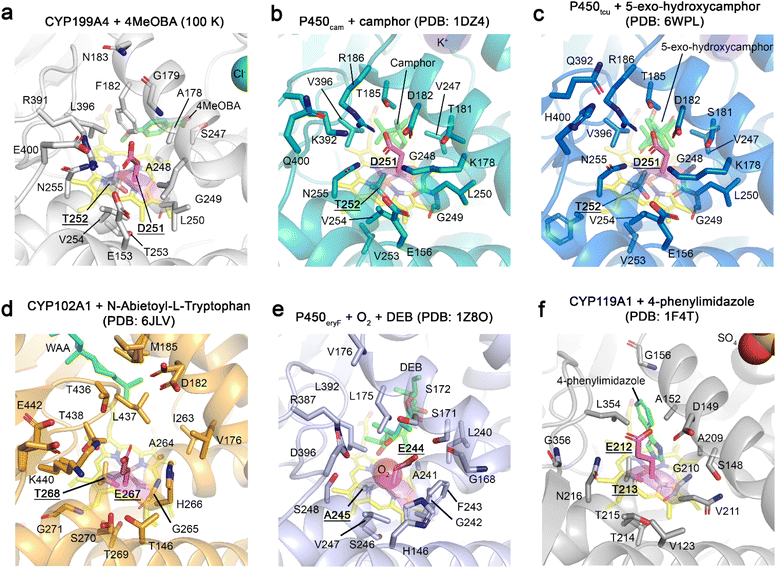
In P450cam – the prototypical model system from which D251's hypothesised role in proton delivery was first proposed – the equivalent residues surrounding D251 are different to those of CYP199A4. In the ‘out’ orientation D251 is clamped between K178 and R186 by salt bridge interactions. The binding of the redox partner has been demonstrated to help break these interactions. In addition, in P450cam the hydrophobic residues observed in CYP199A4 are substituted for polar and charged hydrophilic residues (Fig. 5). Notably, F182 and A178 are both substituted for threonines in P450cam. A survey of other structurally characterised CYP enzymes highlights that this region varies greatly (Fig. 5). In the MD simulations the D251 has a negative charge and this seemed to favour the movement of the side chain to the ‘out’ orientation. In certain X-ray structures of CYP199A4 in which D251 has been mutated to a neutral asparagine a higher proportion of the ‘in’ side chain orientation was observed (Fig. S4).
In CYP199A4 the interaction of D251 with N255 would be easier to break than the equivalent salt bridge network of interactions found in P450cam. This would explain why CYP199A4 and other CYPs do not have such a restrictive requirement for the physiological redox partner that P450cam has for putidaredoxin (the effector role). It is proposed that in P450cam the binding of the redox partner in this system opens up the conformation of the enzyme enabling breaking of the salt network and freeing up of the aspartate residue for proton delivery.36 These changes that we observed in CYP199A4 when the physiological substrate is bound could allow for D251 in CYP199A4 to switch between ‘in’ and ‘out’ conformations and allow solvent flow to the heme. In P450cam changes in the position of the side chain of D251 are observed after conformational changes introduced by redox-partner binding.
The conformational changes to the I-helix oxygen-binding groove and neighbouring residues, particularly T253, resulted in formation of a new solvent channel passing from the active site, under the I-helix to the surface under the D-helix in the MD simulations. Previously, a solvent channel, harbouring multiple ordered waters, passing below the acid–alcohol pair and oxygen-binding groove in P450cam was postulated to be an alternate pathway for proton delivery or water entry/exit. The amino acid at residue position 253 of CYP199A4 is a threonine, instead of the valine of P450cam, and this forms a coordinated hydrogen bonding network with a single ordered water and conserved E367 (Fig. S44). In the X-ray crystal structures of P450cam no such solvent channel is observable and T253 interacts with this single water molecule in all the CYP199A4 structures. In P450cam (PDB: 1DZ4), the side chains of V253 and the conserved glutamate residues have different orientations, and two water molecules occupy this space. The MD simulations of both substrate-bound forms of CYP199A4 showed movements in the position of T253 and E367 residues. These changes opened up a transient solvent channel above and below the oxygen-binding groove of the I-helix ultimately exiting near the D-helix, which could facilitate water exit or entry.
Conclusions
Variable-temperature X-ray crystallography and all-atom molecule dynamics simulations were used to demonstrate that the D251 and other residues within the oxygen-binding groove of the I-helix of CYP199A4 are flexible. The side chain of D251 can rotate in and out of the active site which, agrees with its proposed role in facilitating proton delivery to the active site from bulk solvent during the catalytic cycle. In both X-ray crystal structures and MD simulation of CYP199A4, D251 favours the ‘out’ of the active site orientation to overcome chemical incompatibility with A178 and F182. Its orientation was also coupled to the presence or absence of the 6th distal heme ligand. There was a correlation between the presence of the heme-bound water ligand and the position of the D251 side chain in CYP199A4 and the level of allosteric conformational changes induced within the I-helix and throughout the enzyme. Overall, these changes around the oxygen-binding groove act as a substrate-dependent allosteric switch resulting in structural changes throughout the enzyme. This enables side chain interactions to change and water molecules to approach the distal side of the heme assisting in proton delivery when required during the catalytic cycle. These changes may have a role in enabling solvent efflux from the active site of the enzyme via a channel in the proximal side of the heme. A comparison of the results and the highly diverse neighbouring environment of the acidic D251 residue in CYP199A4 and P450cam (and other CYP enzymes) suggest that there may not be a ‘universal’ model for proton transfer in CYP enzymes, but that allosteric effects, transient salt bridges and other interactions are likely be important. The further investigation of these enzymes using a combination of other techniques, e.g. NMR combined with room temperature serial crystallography, mutagenesis and more advanced MD simulations will provide further information on oxygen activation and proton transfer.Author contributions
M. N. P. carried out the protein purification, crystal structure determination and analysis with assistance from E. C. C. and J. B. B. D. P. McD. did the MD simulations and analysis. M. N. P., D. P. McD. and S. G. B. wrote the manuscript draft. E. C. C. and J. B. B. provided supervision on aspects of the project. S. G. B. conceived and had overall supervision of the project and critical manuscript review.Conflicts of interest
There are no conflicts to declare.Abbreviations
| CYP | Cytochrome P450 |
| NAD(P)H | Reduced nicotinamide adenine dinucleotide (phosphate) |
| Cpd I (compound I) | The ferryl-oxo heme radical cation |
| CYP or P450 | Cytochrome P450 enzyme |
| CYP199A4 | A cytochrome P450 enzyme from Rhodopseudomonas palustris strain HaA2 |
| H2O2 | Hydrogen peroxide |
| P450cam (CYP101A1) | A camphor-hydroxylating cytochrome P450 enzyme from Pseudomonas putida |
| P450cin (CYP176A1) | A 1,8-cineole-hydroxylating P450 enzyme |
| PDB | Protein Data Bank |
| Pdx (putidaredoxin) | The [2Fe–2S] ferredoxin that delivers electrons to P450cam from Pseudomonas putida |
| WT | Wild type |
| MD | Molecular dynamics |
| PME | Particle mesh ewald |
| NVT | Constant number of particles, volume and temperature |
| NPT | Constant number of particles, pressure and temperature |
| fs | Femtosecond |
| ps | Picosecond |
| ns | Nanosecond |
| µs | Microsecond |
| GPU | Graphics processing unit |
Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: further analysis of the X-ray crystal structure and MD simulations; PDB validation reports and CIF files. See DOI: https://doi.org/10.1039/d5sc07539d.Variable-temperature X-ray crystal structures have been deposited into the Protein Data Bank (https://www.rcsb.org/) (PDB accession codes: 9DOE, 9MJF, 9MJJ, 9MJK, 9MIM, 9MIO, 9MJE, 9PLS, 9PMA, 9PMC, 9ZEG, 9ZEH, and 9ZEI).93a–m
Acknowledgements
This research was undertaken using the MX2 beamline at the Australian Synchrotron, part of ANSTO, and made use of the Australian Cancer Research Foundation (ACRF) detector. Computing resources were provided by the University of Adelaide Phoenix HPC cluster. We acknowledge Dr Alicia Kirk (University of Queensland) for aiding in preparing MD input files. This research was supported by an AINSE Ltd. Postgraduate Research Award (PGRA), an Australian Government Research Training Program Scholarship and a Constance Fraser PhD Scholarship from the University of Adelaide to M. N. P. D. P. McD. was supported by an Australian Government Research Training Program Scholarship and NHMRC funded postgraduate Scholarship.Notes and references
- P. S. Coelho, E. M. Brustad, A. Kannan and F. H. Arnold, Science, 2013, 339, 307–310 CrossRef CAS PubMed.
- P. R. Ortiz de Montellano, in Cytochrome P450: Structure, Mechanism, and Biochemistry, ed. P. R. Ortiz de Montellano, Springer International Publishing, Cham, 2015, pp. 111–176 Search PubMed.
- D. J. Cook, J. D. Finnigan, K. Cook, G. W. Black and S. J. Charnock, in Advances in Protein Chemistry and Structural Biology, ed. C. Z. Christov, Academic Press, 2016, vol. 105, pp. 105–126 Search PubMed.
- F. P. Guengerich, Nat. Rev. Drug Discovery, 2002, 1, 359–366 Search PubMed.
- D. C. Lamb, A. H. Follmer, J. V. Goldstone, D. R. Nelson, A. G. Warrilow, C. L. Price, M. Y. True, S. L. Kelly, T. L. Poulos and J. J. Stegeman, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 12343–12352 Search PubMed.
- D. R. Nelson, Biochim. Biophys. Acta, 2018, 1866, 141–154 CrossRef CAS PubMed.
- X. He, M. J. Cryle, J. J. D. Voss and P. R. O. de Montellano, J. Biol. Chem., 2005, 280, 22697–22705 CrossRef CAS PubMed.
- L. T. Burka, A. Thorsen and F. P. Guengerich, J. Am. Chem. Soc., 1980, 102, 7615–7616 CrossRef CAS.
- F. P. Guengerich, Chem. Res. Toxicol., 2001, 14, 611–650 Search PubMed.
- T. C. Pochapsky, N. Wong, Y. Zhuang, J. Futcher, M.-E. Pandelia, D. R. Teitz and A. M. Colthart, Biochim. Biophys. Acta, 2018, 1866, 126–133 CrossRef CAS PubMed.
- T. Coleman, M. N. Podgorski, M. L. Doyle, J. M. Scaffidi-Muta, E. C. Campbell, J. B. Bruning, J. J. De Voss and S. G. Bell, J. Inorg. Biochem., 2023, 244, 112234 CrossRef CAS PubMed.
- J. C. Miller, J. H. Z. Lee, M. A. McLean, R. R. Chao, I. S. J. Stone, T. L. Pukala, J. B. Bruning, J. J. De Voss, M. A. Schuler, S. G. Sligar and S. G. Bell, J. Am. Chem. Soc., 2023, 145, 9207–9222 CrossRef CAS PubMed.
- V. Cojocaru, P. J. Winn and R. C. Wade, Biochim. Biophys. Acta, 2007, 1770, 390–401 Search PubMed.
- R. C. Wade, P. J. Winn and I. Schlichting, Sudarko, J. Inorg. Biochem., 2004, 98, 1175–1182 CrossRef CAS PubMed.
- I. Schlichting, J. Berendzen, K. Chu, A. M. Stock, S. A. Maves, D. E. Benson, R. M. Sweet, D. Ringe, G. A. Petsko and S. G. Sligar, Science, 2000, 287, 1615–1622 Search PubMed.
- T. I. Oprea, G. Hummer and A. E. Garcia, Proc. Natl. Acad. Sci. U. S. A., 1997, 94, 2133–2138 Search PubMed.
- M. J. Honeychurch, H. A. O. Hill and L.-L. Wong, FEBS Lett., 1999, 451, 351–353 CrossRef CAS PubMed.
- I. G. Denisov and S. G. Sligar, in Cytochrome P450: Structure, Mechanism, and Biochemistry, ed. P. R. Ortiz de Montellano, Springer International Publishing, Cham, 2015, pp. 69–109 Search PubMed.
- F. Hannemann, A. Bichet, K. M. Ewen and R. Bernhardt, Biochim. Biophys. Acta, Gen. Subj., 2007, 1770, 330–344 CrossRef CAS PubMed.
- M. N. Podgorski, T. Coleman, L. R. Churchman, J. B. Bruning, J. J. De Voss and S. G. Bell, Chem.–Eur. J., 2022, 28, e202202428 CrossRef CAS PubMed.
- J. Rittle and M. T. Green, Science, 2010, 330, 933–937 CrossRef CAS PubMed.
- M. R. Sarkar, S. D. Houston, G. P. Savage, C. M. Williams, E. H. Krenske, S. G. Bell and J. J. De Voss, J. Am. Chem. Soc., 2019, 141, 19688–19699 CrossRef CAS PubMed.
- X. Huang and J. T. Groves, J. Biol. Inorg. Chem., 2017, 22, 185–207 CrossRef CAS PubMed.
- J. T. Groves and G. A. McClusky, J. Am. Chem. Soc., 1976, 98, 859–861 CrossRef CAS.
- A. R. Modi and J. H. Dawson, in Monooxygenase, Peroxidase and Peroxygenase Properties and Mechanisms of Cytochrome P450, ed. E. G. Hrycay and S. M. Bandiera, Springer International Publishing, Cham, 2015, pp. 63–81 Search PubMed.
- T. L. Poulos and R. Raag, FASEB J., 1992, 6, 674–679 CrossRef CAS PubMed.
- T. L. Poulos, Chem. Rev., 2014, 114, 3919–3962 CrossRef CAS PubMed.
- T. Coleman, J. E. Stok, M. N. Podgorski, J. B. Bruning, J. J. De Voss and S. G. Bell, J. Biol. Inorg. Chem., 2020, 25, 583–596 CrossRef CAS PubMed.
- S. Nagano and T. L. Poulos, J. Biol. Chem., 2005, 280, 31659–31663 CrossRef CAS PubMed.
- M. Imai, H. Shimada, Y. Watanabe, Y. Matsushima-Hibiya, R. Makino, H. Koga, T. Horiuchi and Y. Ishimura, Proc. Natl. Acad. Sci. U. S. A., 1989, 86, 7823–7827 CrossRef CAS PubMed.
- J. E. Stok, S. Yamada, A. J. Farlow, K. E. Slessor and J. J. De Voss, Biochim. Biophys. Acta, 2013, 1834, 688–696 Search PubMed.
- K. D. Dubey, B. Wang, M. Vajpai and S. Shaik, Chem. Sci., 2017, 8, 5335–5344 RSC.
- R. Davydov, T. M. Makris, V. Kofman, D. E. Werst, S. G. Sligar and B. M. Hoffman, J. Am. Chem. Soc., 2001, 123, 1403–1415 CrossRef CAS PubMed.
- T. L. Poulos and A. H. Follmer, Acc. Chem. Res., 2022, 55, 373–380 Search PubMed.
- A. H. Follmer, S. Tripathi and T. L. Poulos, J. Am. Chem. Soc., 2019, 141, 2678–2683 CrossRef CAS PubMed.
- S. Tripathi, H. Li and T. L. Poulos, Science, 2013, 340, 1227–1230 CrossRef CAS PubMed.
- J. A. Amaya, D. Batabyal and T. L. Poulos, Biochemistry, 2020, 59, 2896–2902 CrossRef CAS PubMed.
- I. Ugur and P. Chandrasekhar, Proteins:Struct., Funct., Bioinf., 2020, 88, 558–572 CrossRef CAS PubMed.
- S. P. Skinner, A. H. Follmer, M. Ubbink, T. L. Poulos, J. J. Houwing-Duistermaat and E. Paci, Biochemistry, 2021, 60, 2932–2942 CrossRef CAS PubMed.
- A. H. Follmer, M. Mahomed, D. B. Goodin and T. L. Poulos, J. Am. Chem. Soc., 2018, 140, 16222–16228 CrossRef CAS PubMed.
- J. A. Gable, T. L. Poulos and A. H. Follmer, J. Am. Chem. Soc., 2023, 145, 4254–4265 CrossRef CAS PubMed.
- S. G. Bell, W. Yang, A. B. H. Tan, R. Zhou, E. O. D. Johnson, A. Zhang, W. Zhou, Z. Rao and L.-L. Wong, Dalton Trans., 2012, 41, 8703–8714 RSC.
- M. N. Podgorski, J. S. Harbort, T. Coleman, J. E. Stok, J. A. Yorke, L. L. Wong, J. B. Bruning, P. V. Bernhardt, J. J. De Voss, J. R. Harmer and S. G. Bell, Biochemistry, 2020, 59, 1038–1050 Search PubMed.
- T. Coleman, S. H. Wong, M. N. Podgorski, J. B. Bruning, J. J. De Voss and S. G. Bell, ACS Catal., 2018, 8, 5915–5927 CrossRef CAS.
- T. Coleman, D. Z. Doherty, T. Zhang, M. N. Podgorski, R. Qiao, J. H. Z. Lee, J. B. Bruning, J. J. De Voss, W. Zhou and S. G. Bell, Chem.–Asian J., 2022, 17, e202200986 CrossRef CAS PubMed.
- T. Coleman, A. M. Kirk, R. R. Chao, M. N. Podgorski, J. S. Harbort, L. R. Churchman, J. B. Bruning, P. V. Bernhardt, J. R. Harmer, E. H. Krenske, J. J. De Voss and S. G. Bell, ACS Catal., 2021, 11, 1995–2010 CrossRef CAS.
- T. Mehlman, J. T. Biel, S. M. Azeem, E. R. Nelson, S. Hossain, L. E. Dunnett, N. G. Paterson, A. Douangamath, R. Talon, D. Axford, H. Orins, F. von Delft and D. A. Keedy, eLife, 2023, 12, e84632 CrossRef CAS PubMed.
- S. Y. C. Bradford, L. El Khoury, Y. Ge, M. Osato, D. L. Mobley and M. Fischer, Chem. Sci., 2021, 12, 11275–11293 RSC.
- E. F. Garman and R. L. Owen, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2006, 62, 32–47 CrossRef PubMed.
- E. Garman, Curr. Opin. Struct. Biol., 2003, 13, 545–551 CrossRef CAS PubMed.
- K. L. Shelley and E. F. Garman, Nat. Commun., 2022, 13, 1314 CrossRef CAS PubMed.
- J. S. Fraser, H. van den Bedem, A. J. Samelson, P. T. Lang, J. M. Holton, N. Echols and T. Alber, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 16247–16252 CrossRef CAS PubMed.
- M. N. Podgorski, T. Coleman, R. R. Chao, J. J. De Voss, J. B. Bruning and S. G. Bell, J. Inorg. Biochem., 2020, 203, 110913 Search PubMed.
- M. N. Podgorski, A. B. Keto, T. Coleman, J. B. Bruning, J. J. De Voss, E. H. Krenske and S. G. Bell, Chem.–Eur. J., 2023, 29, e202301371 CrossRef CAS PubMed.
- N. P. Cowieson, D. Aragao, M. Clift, D. J. Ericsson, C. Gee, S. J. Harrop, N. Mudie, S. Panjikar, J. R. Price, A. Riboldi-Tunnicliffe, R. Williamson and T. Caradoc-Davies, J. Synchrotron Radiat., 2015, 22, 187–190 Search PubMed.
- D. Aragão, J. Aishima, H. Cherukuvada, R. Clarken, M. Clift, N. P. Cowieson, D. J. Ericsson, C. L. Gee, S. Macedo, N. Mudie, S. Panjikar, J. R. Price, A. Riboldi-Tunnicliffe, R. Rostan, R. Williamson and T. T. Caradoc-Davies, J. Synchrotron Radiat., 2018, 25, 885–891 Search PubMed.
- M. D. Winn, C. C. Ballard, K. D. Cowtan, E. J. Dodson, P. Emsley, P. R. Evans, R. M. Keegan, E. B. Krissinel, A. G. W. Leslie, A. McCoy, S. J. McNicholas, G. N. Murshudov, N. S. Pannu, E. A. Potterton, H. R. Powell, R. J. Read, A. Vagin and K. S. Wilson, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2011, 67, 235–242 CrossRef CAS PubMed.
- P. R. Evans and G. N. Murshudov, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2013, 69, 1204–1214 CrossRef CAS PubMed.
- A. J. McCoy, R. W. Grosse-Kunstleve, P. D. Adams, M. D. Winn, L. C. Storoni and R. J. Read, J. Appl. Crystallogr., 2007, 40, 658–674 CrossRef CAS PubMed.
- N. W. Moriarty, R. W. Grosse-Kunstleve and P. D. Adams, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2009, 65, 1074–1080 CrossRef CAS PubMed.
- P. V. Afonine, R. W. Grosse-Kunstleve, N. Echols, J. J. Headd, N. W. Moriarty, M. Mustyakimov, T. C. Terwilliger, A. Urzhumtsev, P. H. Zwart and P. D. Adams, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2012, 68, 352–367 CrossRef CAS PubMed.
- P. Emsley, B. Lohkamp, W. G. Scott and K. Cowtan, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2010, 66, 486–501 CrossRef CAS PubMed.
- V. B. Chen, W. B. Arendall III, J. J. Headd, D. A. Keedy, R. M. Immormino, G. J. Kapral, L. W. Murray, J. S. Richardson and D. C. Richardson, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2010, 66, 12–21 CrossRef CAS PubMed.
- R. B. G. Ravelli and S. M. McSweeney, Structure, 2000, 8, 315–328 CrossRef CAS PubMed.
- P. V. Afonine, N. W. Moriarty, M. Mustyakimov, O. V. Sobolev, T. C. Terwilliger, D. Turk, A. Urzhumtsev and P. D. Adams, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2015, 71, 646–666 CrossRef CAS PubMed.
- T. C. Terwilliger, R. W. Grosse-Kunstleve, P. V. Afonine, N. W. Moriarty, P. D. Adams, R. J. Read, P. H. Zwart and L.-W. Hung, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2008, 64, 515–524 CrossRef CAS PubMed.
- D. Liebschner, P. V. Afonine, M. L. Baker, G. Bunkoczi, V. B. Chen, T. I. Croll, B. Hintze, L.-W. Hung, S. Jain, A. J. McCoy, N. W. Moriarty, R. D. Oeffner, B. K. Poon, M. G. Prisant, R. J. Read, J. S. Richardson, D. C. Richardson, M. D. Sammito, O. V. Sobolev, D. H. Stockwell, T. C. Terwilliger, A. G. Urzhumtsev, L. L. Videau, C. J. Williams and P. D. Adams, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2019, 75, 861–877 CrossRef CAS PubMed.
- D. Van Der Spoel, E. Lindahl, B. Hess, G. Groenhof, A. E. Mark and H. J. Berendsen, J. Comput. Chem., 2005, 26, 1701–1718 CrossRef CAS PubMed.
- The PyMOL Molecular Graphics System, Version 2.4.1, Schrödinger, LLC Search PubMed.
- E. F. Pettersen, T. D. Goddard, C. C. Huang, G. S. Couch, D. M. Greenblatt, E. C. Meng and T. E. Ferrin, J. Comput. Chem., 2004, 25, 1605–1612 CrossRef CAS PubMed.
- D. A. Case, H. M. Aktulga, K. Belfon, D. S. Cerutti, G. A. Cisneros, V. W. D. Cruzeiro, N. Forouzesh, T. J. Giese, A. W. Götz, H. Gohlke, S. Izadi, K. Kasavajhala, M. C. Kaymak, E. King, T. Kurtzman, T. S. Lee, P. Li, J. Liu, T. Luchko, R. Luo, M. Manathunga, M. R. Machado, H. M. Nguyen, K. A. O'Hearn, A. V. Onufriev, F. Pan, S. Pantano, R. Qi, A. Rahnamoun, A. Risheh, S. Schott-Verdugo, A. Shajan, J. Swails, J. Wang, H. Wei, X. Wu, Y. Wu, S. Zhang, S. Zhao, Q. Zhu, T. E. Cheatham 3rd, D. R. Roe, A. Roitberg, C. Simmerling, D. M. York, M. C. Nagan and K. M. Merz Jr, J. Chem. Inf. Model., 2023, 63, 6183–6191 CrossRef CAS PubMed.
- J. C. Gordon, J. B. Myers, T. Folta, V. Shoja, L. S. Heath and A. Onufriev, Nucleic Acids Res., 2005, 33, W368–W371 CrossRef CAS PubMed.
- K. Shahrokh, A. Orendt, G. S. Yost and T. E. Cheatham 3rd, J. Comput. Chem., 2012, 33, 119–133 CrossRef CAS PubMed.
- W. L. Jorgensen, J. Chandrasekhar, J. D. Madura, R. W. Impey and M. L. Klein, J. Chem. Phys., 1983, 79, 926–935 CrossRef CAS.
- B. Hess, H. Bekker, H. J. C. Berendsen and J. G. E. M. Fraaije, J. Comput. Chem., 1997, 18, 1463–1472 CrossRef CAS.
- T. Darden, D. York and L. Pedersen, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS.
- H. J. C. Berendsen, J. P. M. Postma, W. F. van Gunsteren, A. DiNola and J. R. Haak, J. Chem. Phys., 1984, 81, 3684–3690 CrossRef CAS.
- M. Parrinello and A. Rahman, J. Appl. Phys., 1981, 52, 7182–7190 Search PubMed.
- R. T. McGibbon, K. A. Beauchamp, M. P. Harrigan, C. Klein, J. M. Swails, C. X. Hernández, C. R. Schwantes, L. P. Wang, T. J. Lane and V. S. Pande, Biophys. J., 2015, 109, 1528–1532 CrossRef CAS PubMed.
- M. N. Podgorski, T. Coleman, P. D. Giang, C. R. Wang, J. B. Bruning, P. V. Bernhardt, J. J. De Voss and S. G. Bell, Inorg. Chem., 2022, 61, 236–245 CrossRef CAS PubMed.
- T. Beitlich, K. Kühnel, C. Schulze-Briese, R. L. Shoeman and I. Schlichting, J. Synchrotron Radiat., 2007, 14, 11–23 CrossRef CAS PubMed.
- V. Pfanzagl, J. H. Beale, H. Michlits, D. Schmidt, T. Gabler, C. Obinger, K. Djinović-Carugo and S. Hofbauer, J. Biol. Chem., 2020, 295, 13488–13501 CrossRef CAS PubMed.
- A. A. Trofimov, K. M. Polyakov, V. A. Lazarenko, A. N. Popov, T. V. Tikhonova, A. V. Tikhonov and V. O. Popov, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2015, 71, 1087–1094 CrossRef CAS PubMed.
- E. Garman, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2010, 66, 339–351 CrossRef CAS PubMed.
- T. Coleman, J. Z. H. Lee, A. M. Kirk, D. Z. Doherty, M. N. Podgorski, D. K. Pinidiya, J. B. Bruning, J. J. De Voss, E. H. Krenske and S. G. Bell, Chem.–Eur. J., 2022, 28, e202201895 CrossRef CAS PubMed.
- J. H. Z. Lee, T. Coleman, M. A. McLean, M. N. Podgorski, E. F. Hayball, I. S. J. Stone, J. B. Bruning, F. Whelan, J. J. De Voss, S. G. Sligar and S. G. Bell, ACS Catal., 2024, 14, 8958–8971 CrossRef CAS PubMed.
- M. N. Podgorski, L. Martínez-Castro, J. B. Bruning, E. C. Campbell, J.-D. Maréchal and S. G. Bell, ACS Catal., 2025, 15, 2867–2884 CrossRef CAS.
- P. Zhao, Y. Jiang, Q. Wang, J. Chen, F. Yao and Z. Cong, Chem. Sci., 2024, 15, 8062–8070 RSC.
- M. N. Podgorski and S. G. Bell, ACS Catal., 2025, 15, 5191–5210 CrossRef CAS.
- P. Zhao, F. Kong, Y. Jiang, X. Qin, X. Tian and Z. Cong, J. Am. Chem. Soc., 2023, 145, 5506–5511 CrossRef CAS PubMed.
- I. G. Denisov, T. M. Makris, S. G. Sligar and I. Schlichting, Chem. Rev., 2005, 105, 2253–2278 CrossRef CAS PubMed.
- J. S. Harbort, J. J. De Voss, J. E. Stok, S. G. Bell and J. R. Harmer, in Future Directions in Metalloprotein and Metalloenzyme Research, ed. G. Hanson and L. Berliner, Springer International Publishing, Cham, 2017, pp. 103–142 Search PubMed.
- (a) M. N. Podgorski and S. G. Bell, 9DOE, 2025, DOI:10.2210/pdb9doe/pdb; (b) M. N. Podgorski and S. G. Bell, 9MJF, 2025, DOI:10.2210/pdb9mjf/pdb; (c) M. N. Podgorski and S. G. Bell, 9MJJ, 2025, DOI:10.2210/pdb9mjj/pdb; (d) M. N. Podgorski and S. G. Bell, 9MJK, 2025, DOI:10.2210/pdb9mjk/pdb; (e) M. N. Podgorski and S. G. Bell, 9MIM, 2025, DOI:10.2210/pdb9mim/pdb; (f) M. N. Podgorski and S. G. Bell, 9MIO, 2025, DOI:10.2210/pdb9mio/pdb; (g) M. N. Podgorski and S. G. Bell, 9MJE, 2025, DOI:10.2210/pdb9mje/pdb; (h) M. N. Podgorski and S. G. Bell, 9PLS, 2025, DOI:10.2210/pdb9pls/pdb; (i) M. N. Podgorski and S. G. Bell, 9PMA, 2025, DOI:10.2210/pdb9pma/pdb; (j) M. N. Podgorski and S. G. Bell, 9PMC, 2025, DOI:10.2210/pdb9pmc/pdb; (k) M. N. Podgorski and S. G. Bell, 9ZEG, 2025, DOI:10.2210/pdb9zeg/pdb; (l) M. N. Podgorski and S. G. Bell, 9ZEH, 2025, DOI:10.2210/pdb9zeh/pdb; (m) M. N. Podgorski and S. G. Bell, 9ZEI, 2025, DOI:10.2210/pdb9zei/pdb.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |