Open Access Article
Open Access ArticleTime-ordered-expression mRNA (TOE mRNA) for melanoma RNA vaccines
Xiangdong
Zhang†
 ab,
Xucong
Teng†
ab,
Xucong
Teng†
 ac,
Yicong
Dai†
ac,
Ningqiang
Gong†
ac,
Qiushuang
Zhang
b,
Difei
Hu
b,
Yuncong
Wu
b,
Hongwei
Hou
*a and
Jinghong
Li
ac,
Yicong
Dai†
ac,
Ningqiang
Gong†
ac,
Qiushuang
Zhang
b,
Difei
Hu
b,
Yuncong
Wu
b,
Hongwei
Hou
*a and
Jinghong
Li
 *abc
*abc
aBeijing Life Science Academy, Beijing 102209, China. E-mail: jhli@mail.tsinghua.edu.cn; houhw@blsa.com.cn
bNew Cornerstone Science Laboratory, Department of Chemistry, Key Laboratory of Bioorganic Phosphorus Chemistry & Chemical Biology, Tsinghua University, Beijing 100084, China
cCenter for BioAnalytical Chemistry, Hefei National Laboratory of Physical Science at Microscale, University of Science and Technology of China, Hefei 230026, China
First published on 26th January 2026
Abstract
Modified mRNA technology has transformed vaccine development by enabling rapid and precise antigen production. The incorporation of adjuvants may further enhance innate immune activation, thereby improving the efficacy of mRNA vaccines. However, inappropriate vaccine kinetics leading to excessive activation of the innate immune system can inhibit the mTORC1 pathway and impair antigen mRNA translation, ultimately limiting vaccine potency. Herein, we present the development of a time-ordered-expression mRNA (TOE mRNA) based on ADAR-mediated A-to-I base editing. Upon cellular entry, the TOE mRNA initiates immediate antigen translation, while the adjuvant encoded within the same mRNA is translated approximately 12 hours post-antigen expression. This delayed adjuvant expression ensures sustained activation of the mTORC1 pathway and robust antigen expression, effectively overcoming the limitations imposed by suboptimal vaccine kinetics. We demonstrate that vaccines utilizing TOE mRNA encoding a tumor neoantigen (for normal translation) and an IL-12 adjuvant (with delayed translation) elicit significantly enhanced antitumor immune responses. TOE mRNA technology represents a promising platform for advancing next-generation mRNA vaccines with improved efficacy.
Introduction
Vaccines serve as pivotal tools in advancing tumor therapy and preventing infectious diseases.1–3 mRNA vaccines, exemplified by the Moderna4 and BioNTech5 SARS-CoV-2 vaccines, represent a landmark advancement in vaccine technology due to their exceptional efficacy (95%), scalable manufacturing processes, and adaptability for targeting mutant strains. Recent investigations have highlighted that the integration of immune adjuvants can enhance both the potency and durability of immune responses by stimulating innate immunity-related pathways, including complement activation,6 toll-like receptor (TLR) signaling,7 and dendritic cell (DC) maturation.8 However, the premature administration of excessive concentrations of immune adjuvants may lead to diminished antigen translation9–11 and accelerated antigen degradation,12 thereby constraining the potential for further optimization of vaccine efficacy.13–15Recent advances have revealed that immune adjuvants can impair vaccine efficacy via multiple mechanisms,16 including the induction of cellular apoptosis,17 hyperactivation of innate immunity,18 suppression of the mTORC1 pathway, and induction of systemic toxicity.19 These insights highlight the importance of delaying adjuvant expression in tumor immunotherapy to maintain Th1 polarization and preserve CD8+ T-cell functionality.7 As a result, achieving a harmonized kinetic balance between innate immune activation and robust antigen expression has become a key challenge in optimizing mRNA vaccine efficacy.20,21 However, current strategies for modulating vaccine kinetics—such as nanomaterial-based delivery systems21–23 or co-delivery of adjuvant-encoding mRNAs24—are associated with notable limitations. For instance, biodegradable polymers and hydrogels require intricate fabrication processes and exhibit variable degradation profiles,25 while lipid nanoparticle (LNP)-mediated co-delivery often induces premature innate immune activation due to simultaneous antigen/adjuvant expression.24 Critically, no existing approach enables intrinsic temporal control over antigen/adjuvant expression through programmable mRNA translation regulation.
Herein, we present the development of a time-ordered-expression mRNA (TOE mRNA) system, wherein the mRNA region encoding the desired antigen is translated immediately upon cellular entry, while another region encoding the adjuvant on the same mRNA undergoes delayed translation. This temporal separation enables its application in tumor vaccines. To achieve this, an endogenous-mRNA-sensing sequence harboring a stop codon (UAG) was inserted between the antigen-encoding and adjuvant-encoding regions. Upon mRNA entry into cells, the antigen sequence is promptly translated. The adjuvant sequence translation is subsequently delayed by approximately 12 hours due to the requirement for adenosine-to-inosine (A-to-I) base editing at the stop codon mediated by adenosine deaminases acting on RNA (ADARs). This strategy effectively circumvents the inhibition of antigen mRNA translation caused by innate immune activation triggered by premature adjuvant expression. Moreover, the delayed adjuvant expression facilitates antigen processing and presentation, thereby ensuring robust vaccine efficacy. We first validated and optimized the TOE mRNA system and confirmed that it enhances immune responses in dendritic cells (DCs). Furthermore, we demonstrated the efficacy of TOE mRNA vaccines in melanoma immunotherapy and metastasis inhibition. This approach represents a promising platform for advancing next-generation mRNA vaccines.
Results
Design and optimization of the TOE mRNA
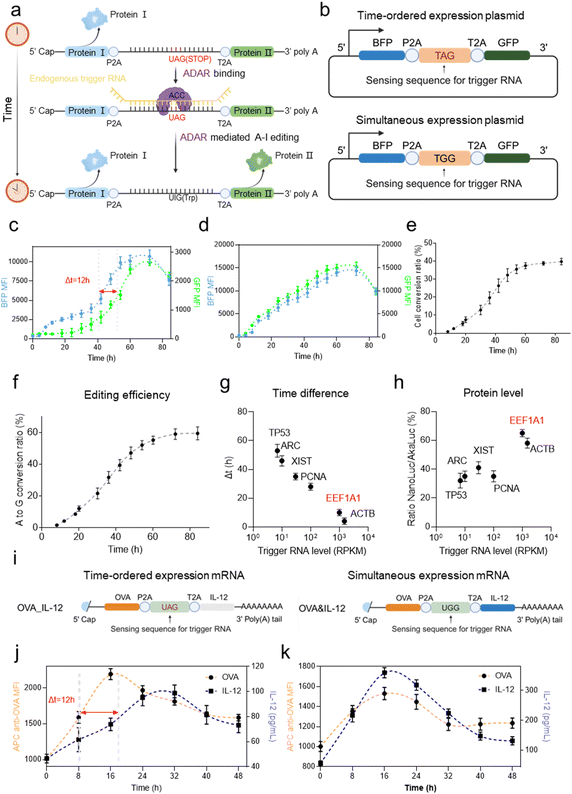
The ADAR family comprises three members: ADAR1, ADAR2, and ADAR3. ADAR1 edits a broad range of double-stranded RNAs (dsRNAs) and plays a key role in immunomodulation and the maintenance of genomic stability.26 In contrast, ADAR2 specializes in the precise editing of a limited number of critical sites,27 while ADAR3 is generally considered to lack significant editing activity.28 This study leverages the editing activity of ADAR1 for our TOE mRNA design.ADAR1 catalyses A-to-I conversion on stretches of base-paired dsRNAs, and the cellular machinery subsequently recognizes inosine as guanosine (G).29 This process is ubiquitous in animal cells30 and can be designed to target specific mRNA sites. Here, we wanted to utilize this mechanism to design a TOE mRNA encoding an antigen and an adjuvant to achieve delayed adjuvant expression (Fig. 1a). The TOE mRNA system comprises three segments: an upstream protein-coding sequence, an endogenous-mRNA-sensing sequence, and a downstream protein-coding sequence. To avoid the middle sequence extending to the open reading frame of the upstream and downstream proteins and potentially affecting the folding of the encoded proteins, we excised the polypeptide encoded by the middle sequence from the upstream and downstream proteins through P2A and T2A self-cleaving peptides.31–33 In eukaryotic systems,34 these two types of 2A peptides mediate rapid35 and efficient self-cleavage31 of fusion proteins during mRNA translation via a ribosome skipping mechanism.32
The sensing sequence in the TOE mRNA system is designed using a modular strategy to achieve temporally ordered protein expression. It is engineered to form a complementary duplex with specific cellular trigger RNAs (200–300 nucleotides), introducing a central A-to-C mismatch at the TAG stop codon to activate ADAR-mediated A-to-I editing. This conversion transforms the stop codon (UAG) into a tryptophan codon (UGG), enabling downstream protein translation. The upstream and downstream coding regions are separated by self-cleaving P2A/T2A peptides to ensure efficient post-translational processing of the proteins. The sensing sequence lacks ATG start codons and avoids complex secondary structures, thereby promoting stable binding to trigger RNAs while preventing unintended translation. By harnessing endogenous RNA editing dynamics, this design introduces a programmable delay in adjuvant expression following antigen translation, addressing the critical challenge of balancing innate immune activation and antigen production in mRNA vaccines.
To validate the hypothesis that the TOE mRNA system delays the expression of a model protein, we initially constructed a TOE mRNA encoding blue fluorescent protein (BFP) in the upstream segment and green fluorescent protein (GFP) in the downstream segment (Fig. 1b) and introduced this TOE mRNA into cells via plasmid transfection. Fluorescence imaging revealed that BFP expression was initiated approximately 8 h post-transfection, whereas GFP expression was delayed, commencing approximately 16 h later (Fig. S1a). However, upon mutating the stop codon TAG to TGG within the sensing sequence, the temporal difference between BFP and GFP expression disappeared (Fig. S1b). Flow cytometry analysis further confirmed the time-ordered expression of BFP and GFP mediated by the TOE mRNA system in cells (Fig. 1c, d and S1c). Additionally, we first characterized A-to-I editing efficiency by flow cytometry, quantifying the percentage of GFP+ cells within the BFP+ population (Fig. 1e). Subsequent Sanger sequencing confirmed A-to-I/G editing at the target site, which converted the stop codon TAG to the tryptophan codon TGG, with the editing efficiency reaching 59.4% (Fig. 1f and S2). This combined analysis showed sufficient editing to ensure abundant downstream protein expression. Moreover, we directly transfected TOE mRNAs encoding AkaLuc (normal translation) and NanoLuc (delayed translation) luciferases and observed their time-ordered expression, demonstrating the versatility of the TOE mRNA system for regulating the temporal expression of other proteins (Fig. S3a and b).
After confirming the feasibility of the TOE mRNA system, we next aimed to optimize both the time gap between upstream and downstream protein expression and the yield of downstream protein production. To achieve this, we designed several sensing sequences tailored to various endogenous trigger RNAs (Fig. S3c). Our findings revealed that a higher abundance of endogenous trigger RNA within cells resulted in a shorter time gap (Fig. 1g) and increased downstream protein production (Fig. 1h). Specifically, Eef1a1 mRNA induced a 12 hours time gap and achieved the highest level of downstream protein expression, making it the most favorable trigger RNA candidate (Fig. 1g and h). Moreover, we performed trigger RNA perturbation detection and found that TOE mRNA transfection did not alter the normal expression of endogenous trigger RNAs, indicating that the system operates without perturbing their physiological expression patterns (Fig. S4). Additionally, detection of delivered mRNA levels in cells at different time points post-transfection showed that the binding of endogenous trigger RNA to TOE mRNA does not affect its stability during A-to-I base editing (Fig. S5). Furthermore, we established HEK293FT cell lines with either ADAR1 overexpression or knockdown (Fig. S6a–c) and demonstrated the critical role of ADAR1 in regulating the TOE mRNA system. Specifically, elevated ADAR1 enzyme expression in HEK293FT cells accelerated the base editing process, thereby reducing the temporal delay between upstream and downstream protein expression. Conversely, ADAR1 knockdown abolished base editing and eliminated downstream protein production entirely (Fig. S6d). Given the ubiquitous presence of ADAR1 across diverse immune cell types (Fig. S6e), the TOE mRNA system appears highly adaptable for use in mRNA vaccines. Subsequently, we engineered a TOE mRNA encoding ovalbumin (OVA, a model antigen) and the cytokine IL-12 (Fig. 1i) and confirmed its ability to sequentially express antigens and cytokine adjuvants (Fig. 1j and k). Additionally, western blot analysis indicated the expression of both proteins flanking the 2A peptide in the aforementioned three cell models, verifying that the self-cleaving activity of the 2A peptide mediated the generation of two distinct protein products within the cells (Fig. S7). Collectively, these results indicate that our optimized TOE mRNA system holds significant potential for enhancing mRNA vaccine kinetics.
The TOE mRNA system enhances antigen expression via delayed adjuvant expression
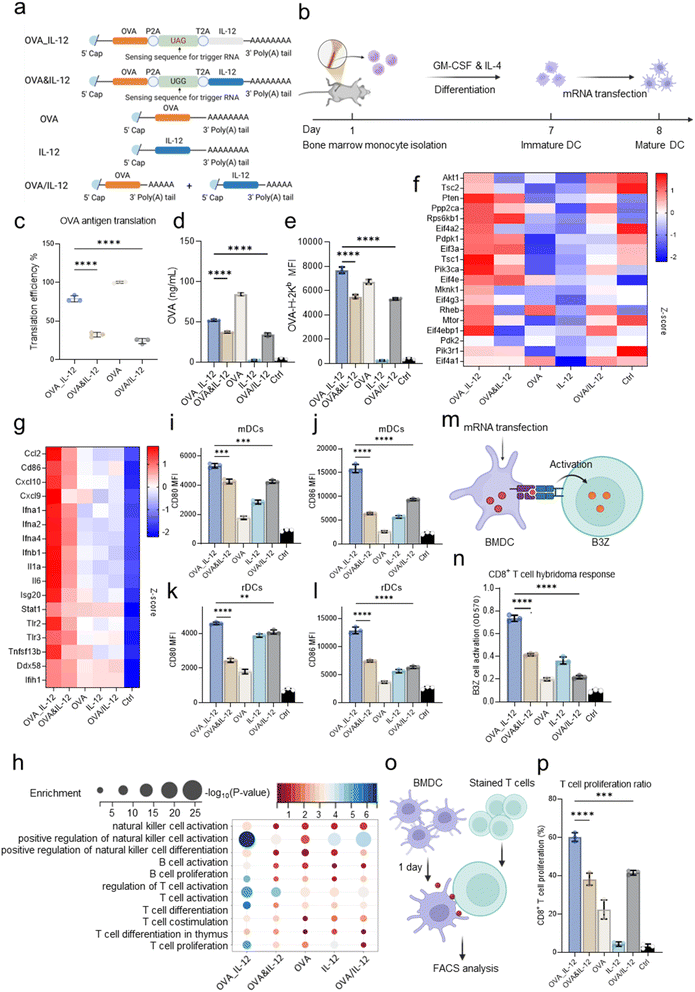
After developing the TOE mRNA for the OVA antigen and the IL-12 adjuvant, we investigated the effects of this TOE mRNA-based vaccine on DC activation and maturation. To this end, we constructed lipid nanoparticles (LNPs) encapsulating the TOE mRNAs encoding OVA (normal expression) and IL-12 (delayed expression) and constructed LNPs encapsulating an mRNA simultaneously encoding OVA and IL-12 (OVA&IL-12, with UAG replaced by UGG in the TOE mRNA of OVA_IL-12), LNPs encapsulating mRNAs encoding OVA or IL-12 individually, and LNPs encapsulating equimolar mixtures of mRNAs encoding OVA and IL-12 (OVA/IL-12) (Fig. 2a) as control groups. We used these LNPs to treat bone marrow-derived dendritic cells (BMDCs) (Fig. 2b) and subsequently evaluated the effect of this TOE mRNA in BMDCs (Fig. S8). Compared with those in the OVA&IL-12 and OVA/IL-12 groups, the OVA_IL-12 TOE mRNA group showed markedly improved OVA antigen expression (Fig. 2c), increased OVA secretion (Fig. 2d) and, consequently, elevated levels of OVA antigenic peptide (SIINFEKL)-MHC I complexes on the surface of the BMDCs (Fig. 2e). The mTORC1 pathway regulates protein synthesis in immune cells through 4E-BPs and eIF4E,10,36 affecting IRF7 translation,37 type I interferon production, and T-cell differentiation.38 Through RNA-seq analysis, we demonstrated that the OVA_IL-12 TOE mRNA activated the mTORC1 pathway, as characterized by increased expression levels of crucial genes such as Tsc1, Pik3ca, and Eif4ebp1 (Fig. 2f), which potentially contributed to improved antigen expression in the cells. In contrast, owing to the expression of the adjuvant, the mTORC1 pathway was inhibited by the innate immune response in the OVA&IL-12 group, the OVA/IL-12 group and, especially, the IL-12 group. Consistently, western blot (WB) analysis revealed that the OVA_IL-12 TOE mRNA group exhibited significantly increased phosphorylation levels of 4E-BP1 (p-4E-BP1, Fig. S9a) and S6K1 (p-S6K1, Fig. S9b), which are well-characterized downstream effectors of mTORC1. This directly confirms mTORC1 pathway activation, aligning with our RNA-seq results showing upregulated expression of key regulatory genes. To further mechanistically validate the role of the mTORC1 pathway in TOE mRNA-mediated antigen expression enhancement, we treated BMDCs with rapamycin (an mTORC1-specific inhibitor)39 prior to OVA_IL-12 TOE mRNA transfection. The results showed that rapamycin pretreatment completely abolished the improved OVA antigen expression (Fig. S10a) and elevated OVA secretion (Fig. S10b), and the increased levels of SIINFEKL-MHC I complexes on BMDCs (Fig. S10c) induced by OVA_IL-12 TOE mRNA. This finding directly demonstrates that mTORC1 activation is indispensable for the TOE mRNA-mediated enhancement of antigen presentation.Next, we evaluated whether the TOE mRNA vaccine could induce the activation of bone marrow-derived dendritic cells (BMDCs). RT-qPCR analysis revealed that the expression levels of cytokines Tnf, Il1b, and Tlr2, as well as key immune sensors Ddx58 and Ifih1, were significantly upregulated 24 h after TOE mRNA transfection (Fig. S11a). This upregulation of Ddx58 and Ifih1 indicates that ADAR-mediated RNA editing triggers the activation of these dsRNA-sensing pathways (Fig. S11a). The Pkr gene plays a critical role in translational control, particularly during viral infection or cellular stress, as it phosphorylates the eIF2α subunit to inhibit protein synthesis.40,41 Notably, the expression level of the translational repressor Pkr remained unchanged (Fig. S11a), indicating that the dsRNA formed by the complementary pairing of the TOE mRNA and triggering RNA activates interferon production without inducing translational arrest or RNA degradation. At the protein level, BMDCs treated with the TOE mRNA vaccine exhibited markedly higher secretion of inflammatory cytokines, including IFN-γ, TNF-α, IL-10, and IL-12p70, as well as the chemokine MCP-1, compared to those treated with OVA&IL-12 or OVA/IL-12 vaccines. These findings support the stronger activation of BMDCs induced by the TOE mRNA vaccine (Fig. S11b).
We subsequently evaluated the overall transcriptomic changes following LNP transfection using RNA-seq. Our findings revealed that TOE mRNA transfection significantly upregulated key genes associated with immune activation (Fig. 2g), thereby enhancing the enrichment of critical biological processes, including immune cell activation (Fig. 2h), cytokine secretion, immune cell chemotaxis, and signal transduction (Fig. S12a). Additionally, the TOE mRNA vaccine markedly activated genes involved in cytokine–cytokine receptor interactions, Th1 and Th2 cell differentiation, and the TNF signaling pathway (Fig. S12b).
At the cellular function level, the expression of costimulatory molecules CD80 and CD86 on the surface of BMDCs was significantly enhanced after TOE mRNA treatment (Fig. 2i–l and S13). Furthermore, BMDCs treated with the TOE mRNA vaccine demonstrated substantially improved MHC class I antigen presentation compared to those treated with OVA&IL-12 or OVA/IL-12 vaccines, as evidenced by a BMDC and B3Z hybridoma cell culture assay (Fig. 2m and n) and an OT-I CD8+ T-cell proliferation assay (Fig. 2o and p).
Collectively, these results highlight the notable synergistic immune-enhancing effects of time-ordered antigen and adjuvant expression mediated by the TOE mRNA system, surpassing those achieved through the delivery of individual antigen and adjuvant mRNA components.
TOE mRNA vaccines initiate robust innate and adaptive immune responses in vivo
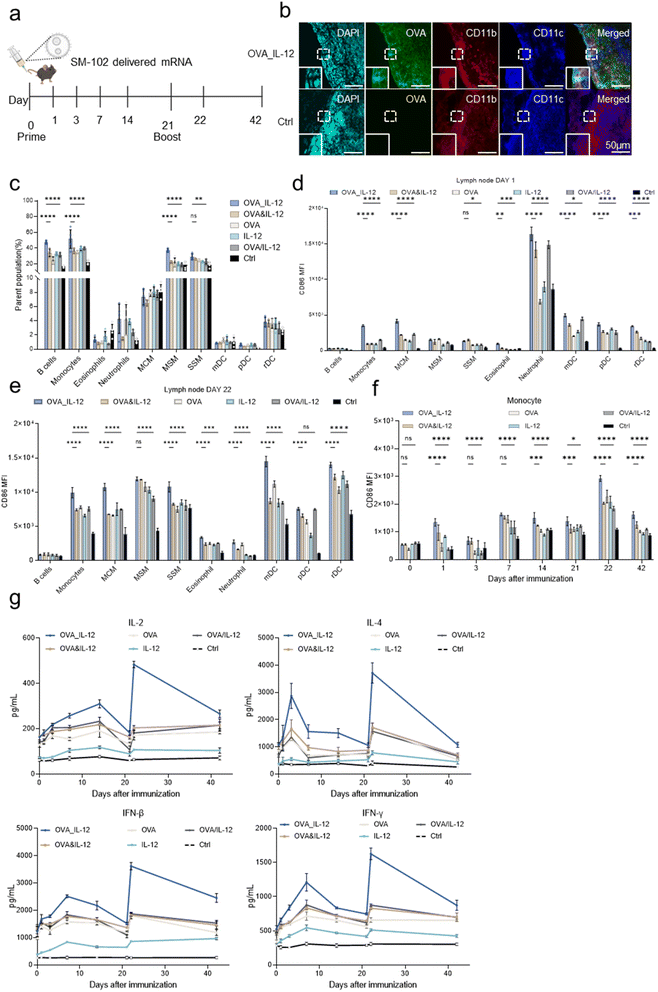
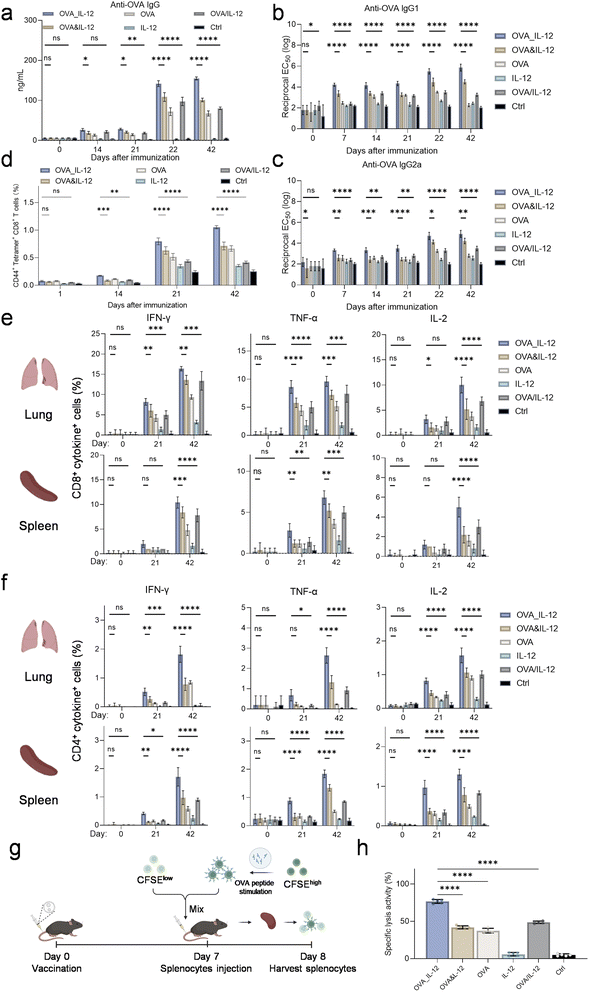
After confirming that the TOE mRNA vaccine elicited a strong immune response in bone marrow-derived dendritic cells (BMDCs), we proceeded to evaluate its ability to induce potent cellular and humoral immune responses in mice. A prime-boost immunization strategy was employed, with intravenous administration of the vaccines on days 0 and 21 (Fig. 3a). Consistent with its known biodistribution profile, SM-102 LNPs predominantly deliver encapsulated mRNA to the liver,42 with substantial accumulation also observed in lymph nodes and other peripheral tissues.43,44 The SM-102 lipid nanoparticle (LNP) formulation used in the Moderna COVID-19 vaccine was adopted for TOE mRNA vaccine encapsulation,45 and this LNP, which had been fully characterized in terms of particle size (Fig. S14a), PDI (Fig. S14b), and zeta potential (Fig. S14c) and thus exhibited excellent performance, exhibited a high encapsulation efficiency for all the mRNA (Fig. S14d). First, we continuously monitored the plasma levels of two proteins via ELISA to validate the in vivo feasibility of the TOE mRNA design under complex physiological conditions (Fig. S15). Subsequently, we assessed the biodistribution of the mRNA vaccine by examining OVA expression in draining lymph nodes (dLNs) 24 h post-vaccination (Fig. 3b). Next, we analysed innate immune cell populations in the dLN, spleen, and lung using flow cytometry (Fig. S16). Compared with the OVA&IL-12 simultaneous expression and OVA/IL-12 coexpression groups, the TOE mRNA vaccine group exhibited a significant increase in the number of various innate immune cells, particularly monocytes capable of antigen presentation, antigen-recognizing B cells, and macrophages responsible for antigen uptake (including marginal cord macrophages (MCMs), marginal sinus macrophages (MSMs), and subcapsular sinus macrophages (SSMs)) within the first day after primary immunization (Fig. 3c). This trend was also observed in the spleen (Fig. S17a) and lungs (Fig. S17b). Furthermore, an increase in CD86 expression was noted across all macrophage and dendritic cell (DC) subsets in the dLN (Fig. 3d) and spleen (Fig. S18a), consistent with the increased proportions of these cells. Following secondary immunization, there was a general increase in the proportions of innate immune cell subpopulations in the spleen (∼1.5-fold) and lungs (∼1.3-fold) compared with primary immunization (Fig. S17a and b). Notably, macrophage and DC subpopulations in the spleen exhibited nearly two times greater activation than those in the OVA&IL-12 or OVA/IL-12 groups (Fig. S17a). The secondary immune activation effect of the TOE mRNA vaccine was approximately 2-fold greater than that in the OVA&IL-12 group and 1.8-fold greater than that in the OVA/IL-12 group (Fig. S18a). Additionally, CD86 expression levels in the dLNs significantly increased after secondary immunization (Fig. 3d and e). Importantly, the activation levels of several immune cell types, including monocytes, SSMs, and mDCs, were markedly higher in the TOE mRNA group than in the OVA&IL-12 (1.34–1.66-fold) and OVA/IL-12 (1.32–1.72-fold) treatment groups (Fig. 3e). Moreover, TOE mRNA-inoculated mice consistently demonstrated robust activation of CD86 expression in monocytes, mDCs, and rDCs (Fig. 3f and S18b). Additionally, we found that TOE mRNA vaccine immunization greatly enhanced the release of cytokines, including IL-2, IL-4, IFN-β, and IFN-γ, by immune cells compared with the other groups (Fig. 3g). Collectively, these results indicate that vaccination with TOE mRNA leads to more efficient antigen uptake and processing by innate immune cells, subsequent activation of a diverse array of innate immune cells, robust induction of inflammatory cytokine release, and more effective triggering of innate immune responses in mice.We further investigated whether the TOE mRNA vaccine could induce adaptive immune responses. First, we assessed the production of OVA-specific antibodies following vaccination and observed a substantial increase in the levels of OVA-specific IgG antibodies (Fig. 4a), including IgG1 (Fig. 4b) and IgG2a (Fig. 4c), 14 days after primary immunization. This elevated antibody level was maintained for up to 21 days post-primary immunization. Additionally, secondary immunization on day 21 resulted in an over 5-fold increase in the antibody response (Fig. 4a–c). Notably, OVA-specific IgA antibody levels exhibited a similar upward trend (Fig. S18c), indicating that the TOE mRNA vaccine elicited a robust humoral immune response. We further analysed antigen-specific T-cell responses in the spleen and lung after primary (day 21) and secondary immunization (day 42) using flow cytometry (Fig. S19). The TOE mRNA vaccine induced an increase in the proportion of SIINFEKL tetramer-positive CD8+ T cells after both primary and secondary immunization (Fig. 4d and S20a). Moreover, the TOE mRNA vaccine also enhanced antigen-specific CD4+ and CD8+ T-cell responses, as evidenced by significantly increased secretion of IFN-γ, TNF-α, and IL-2 by these cells (Fig. 4e and f). Furthermore, histological examination of H&E-stained images of major organs revealed no signs of tissue damage or inflammation in vaccinated mice compared with healthy controls, indicating the excellent biocompatibility of the TOE mRNA vaccine (Fig. S20b). To further confirm the absence of off-target toxicity, we assessed key serum biomarkers of hepatic (ALT, AST), muscular (LDH, creatine kinase), and renal (urea, cystatin C) injury in vaccinated mice. As shown in Fig. S21, there were no significant differences in these biomarkers between the TOE mRNA vaccine groups and healthy controls, indicating that the vaccine did not induce off-target toxicity in major organs. Finally, we evaluated whether TOE mRNA vaccine-induced immune responses could lead to target cell killing (Fig. 4g). The TOE mRNA vaccine achieved the highest OVA-specific cell killing rate (76.7%), which was 1.86-fold higher than that of the OVA&IL-12 group and 1.57-fold higher than that of the OVA/IL-12 group (Fig. 4h). These results demonstrate the significant advantage of the TOE mRNA vaccine in stimulating both cellular and humoral immune responses in vivo.
LNPs encapsulating the TOE mRNA system elicit strong tumour-specific immune responses in vivo
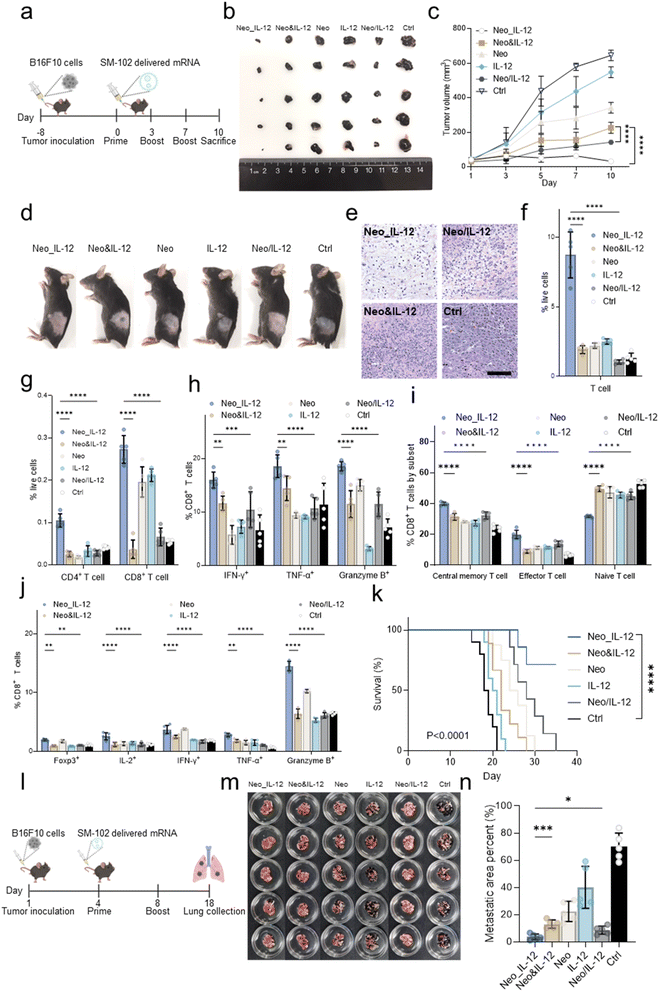
Having established the potent immunotherapeutic effects of TOE mRNA vaccines, we subsequently explored their potential for directly targeting and treating malignant tumours in vivo. Mouse B16F10 cells were subcutaneously injected to establish the tumour model, and the TOE mRNA encoding Pbk-Actn4 (a neoantigen identified in the B16F10 cell line)46 and an IL-12 adjuvant (delayed expression) were used to formulate the mRNA melanoma vaccine (Fig. 5a). Tumour-bearing mice were administered LNPs encapsulating the TOE mRNA (Neo_IL-12), LNPs encapsulating simultaneously expressed mRNAs (Neo&IL-12), LNPs encapsulating the neoantigen mRNA (Neo), LNPs encapsulating the IL-12 mRNA, LNPs encapsulating equimolar mixtures of mRNAs encoding neoantigens and IL-12 (Neo/IL-12), or saline on days 0 and 21. Tumour growth and mouse body weight were monitored. Administration of the TOE mRNA melanoma vaccine resulted in the most potent inhibition of tumour growth, leading to the smallest tumour volume in the TOE mRNA vaccine-treated group compared with the other groups (Fig. 5b–d). Histological analysis further confirmed the therapeutic efficacy of the TOE mRNA melanoma vaccine in restraining tumour cell proliferation (Fig. 5e).Furthermore, flow cytometry was performed to analyse the immune cell status in the tumours, spleens, and lymph nodes (Fig. S22). Within the tumour microenvironment, the TOE mRNA melanoma vaccine significantly increased T-cell infiltration (Fig. 5f), particularly in terms of CD4+ and CD8+ T-cell population proportions (Fig. 5g). Additionally, markedly increased populations of CD8+ T cells secreting IFN-γ, TNF-α, and granzyme B were detected (Fig. 5h). Phenotyping of immune cell populations in the spleen and lymph nodes revealed that injection of the TOE mRNA melanoma vaccine not only expanded effector CD8+ T cells (CD44+ CD62L−) and central memory CD8+ T cells (CD44+ CD62L+) but also enhanced the production of IL-2, IFN-γ, TNF-α, and granzyme B by CD8+ T cells (Fig. 5i, j, S23a and b). Importantly, the robust anti-tumour immune response triggered by the TOE mRNA melanoma vaccine not only effectively suppressed primary tumour growth and extended the survival of tumour-bearing mice (Fig. 5k) but also prevented lung metastasis of melanoma (Fig. 5l–n and S23c). These results indicate that the TOE mRNA tumour vaccine enhances immune cell infiltration into tumour tissue and promotes the anti-tumour activity of T cells, thereby effectively inhibiting tumour growth and preventing metastasis.
Discussion
The emergence of mRNA vaccine technology has ushered in a new era for vaccinology, providing a rapid and versatile platform for the development of novel cancer therapeutics and solutions to infectious diseases. However, a critical challenge in mRNA vaccine design lies in achieving an optimal balance between immune response stimulation via adjuvants and antigen expression for the induction of adaptive immunity. Prior studies have demonstrated that the inclusion of immune adjuvants can suppress antigen translation and accelerate mRNA degradation, thereby limiting the overall efficacy of vaccines.47–50 Unlike conventional strategies that rely on complex delivery systems, we developed a TOE mRNA-based approach to address this issue by leveraging ADAR-mediated base editing to delay adjuvant expression until after antigen translation is completed. This ensures the resolution of the inherent trade-off between robust innate immune activation and efficient antigen expression. Our strategy not only sustains high levels of antigen production but also elicits a potent innate immune response, as evidenced by enhanced DC activation and the subsequent induction of both cellular and humoral immunity in vivo. The TOE mRNA system signifies a paradigm shift toward more efficient, cost-effective, and clinically translatable mRNA vaccine design.TOE mRNA technology provides several distinct advantages over conventional mRNA vaccines involving separate administration of the antigen and adjuvant (e.g. OVA/IL-12 and Neo/IL-12 groups). First, the time-ordered expression of antigens and adjuvants (with a precise 12 hours delay) enables the optimization of immune responses, thereby enhancing vaccine efficacy. Unlike separate administration, which is prone to spatial–temporal mismatch or simultaneous expression-induced mTORC1 pathway inhibition, our TOE mRNA system ensures coordinated, cell-autonomous action. Our findings demonstrate that, based on the general mechanism of antigen-mediated immune activation followed by adjuvant-induced amplification of the immune response,51,52 the TOE mRNA vaccine elicits stronger immune responses compared with traditional mRNA formulations. Specifically, it achieves an OVA-specific target cell killing rate of 76.7% (1.54-fold higher than the OVA/IL-12 group's 49.8%) (Fig. 4h), reduces melanoma tumor volume to 22.3% of the Neo/IL-12 group's size 10 days post-inoculation (Fig. 5c), achieves long-term survival in 71.4% of mice (persistent survival vs. all deaths by day 30 in the Neo/IL-12 group) (Fig. 5k), and suppresses the lung metastasis area to only 3.7% (57% lower than the Neo/IL-12 group's 8.6%) (Fig. 5n). This is attributed to the precise timing of adjuvant expression, which enhances antigen processing and presentation without compromising initial translation efficiency. Second, our approach minimizes the risk of systemic toxicity and adverse effects associated with excessive innate immune activation. Unlike traditional translation regulation methods that rely on nanomaterial-based delivery systems24 (with unstable efficacy) or exogenous stimulus-responsive elements,53,54 this study achieves kinetic control exclusively through mRNA engineering, thereby bypassing complex material construction. This method eliminates the need for nanocarrier systems or external triggers, enabling programmable kinetic regulation via intrinsic mRNA sequence design. Third, the TOE mRNA strategy offers notable convenience. With a single mRNA, it enables one-time transfection for programmed expression of the antigen and adjuvant, unlike separate administration of the antigen and adjuvant, which requires independent optimization of dosages, timelines, and carriers. This simplifies workflows, reduces costs, and facilitates production scalability, making it more advantageous than separate administration. Finally, the flexibility of TOE mRNA design enables the rapid development of vaccines targeting emerging pathogens or tumour-specific antigens, thereby preserving the inherent advantages of mRNA vaccines.
Despite these promising results, further studies are necessary to comprehensively evaluate the long-term safety of TOE mRNA vaccines. Such investigations should include exploring their applicability to a broader spectrum of antigens and adjuvants, as well as advancing towards clinical evaluation. Moreover, the interplay between TOE mRNA vaccines and the IFN-γ/mTOR signalling axis in macrophages offers additional insights into the potential mechanisms for optimizing these vaccines.55 Further exploration of the interactions between TOE mRNA vaccines and other regulatory pathways, such as cGAS/STING and inflammatory mediators,56 could provide valuable information for enhancing vaccine-induced immune responses.
In conclusion, our study presents a transformative platform for designing mRNA vaccines that addresses the limitations of conventional mRNA vaccines. By precisely regulating the timing of adjuvant translation, this innovative platform enhances vaccine efficacy while maintaining safety, thereby paving the way for the development of advanced therapeutic and prophylactic interventions.
Author contributions
The individual contributions are as follows: Xiangdong Zhang: conceptualization, methodology, investigation, writing – original draft. Xucong Teng: methodology, investigation, formal analysis, validation. Yicong Dai: investigation, data curation, formal analysis, visualization. Ningqiang Gong: methodology, software, validation, data curation. Qiushuang Zhang: investigation, resources. Difei Hu: resources, writing – review & editing. Yuncong Wu: resources, project administration. Hongwei Hou: supervision, funding acquisition, writing – review & editing. Jinghong Li: supervision, funding acquisition, project administration, writing – review & editing. All authors have reviewed and approved the final manuscript.Conflicts of interest
The authors declare no conflicts of interest.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: reagents, nucleic acid sequence information, experimental methods, Sanger sequencing electrophoregrams, RNA-seq profiles, western blot images, fluorescence imaging and flow cytometry data. See DOI: https://doi.org/10.1039/d5sc07482g.Acknowledgements
This work was financially supported by the New Cornerstone Science Foundation, the Natural Science Foundation of Anhui Province (2508085QH286 to X. T.), the Fundamental Research Funds for the Central Universities (WK9990250175 to X. T.), and the Beijing Life Science Academy Initiative Scientific Research Program (No. 2023000CA0050, No. 2024100CA0080, No. 2024101RPIA02, No. 2023100CC0230, No. 2023100CC0240 and No. 2023100CC0250). We thank Bin Yu (Core Facility, Center of Biomedical Analysis, Tsinghua University) for technical support with flow cytometry analysis.References
- N. Chaudhary, D. Weissman and K. A. Whitehead, mRNA vaccines for infectious diseases: principles, delivery and clinical translation, Nat. Rev. Drug Discovery, 2021, 20, 817–838 CrossRef.
- C. Liu, Q. Shi, X. Huang, S. Koo, N. Kong and W. Tao, mRNA-based cancer therapeutics, Nat. Rev. Cancer, 2023, 23, 526–543 CrossRef.
- G. Zhang, T. Tang, Y. Chen, X. Huang and T. Liang, mRNA vaccines in disease prevention and treatment, Signal Transduction Targeted Ther., 2023, 8, 365 CrossRef PubMed.
- L. R. Baden, H. M. E. Sahly, B. Essink, K. Kotloff, S. Frey, R. Novak, D. Diemert, S. A. Spector, N. Rouphael, C. B. Creech, J. McGettigan, S. Khetan, N. Segall, J. Solis, A. Brosz, C. Fierro, H. Schwartz, K. Neuzil, L. Corey, P. Gilbert, H. Janes, D. Follmann, M. Marovich, J. Mascola, L. Polakowski, J. Ledgerwood, B. S. Graham, H. Bennett, R. Pajon, C. Knightly, B. Leav, W. Deng, H. Zhou, S. Han, M. Ivarsson, J. Miller and T. Zaks, Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine, N. Engl. J. Med., 2021, 384, 403–416 CrossRef.
- F. P. Polack, S. J. Thomas, N. Kitchin, J. Absalon, A. Gurtman, S. Lockhart, J. L. Perez, G. P. Marc, E. D. Moreira, C. Zerbini, R. Bailey, K. A. Swanson, S. Roychoudhury, K. Koury, P. Li, W. V. Kalina, D. Cooper, R. W. Frenck, L. L. Hammitt, Ö. Türeci, H. Nell, A. Schaefer, S. Ünal, D. B. Tresnan, S. Mather, P. R. Dormitzer, U. Şahin, K. U. Jansen and W. C. Gruber, Safety and Efficacy of the BNT162b2 mRNA Covid-19 Vaccine, N. Engl. J. Med., 2020, 383, 2603–2615 CrossRef PubMed.
- B. Li, A. Y. Jiang, I. Raji, C. Atyeo, T. M. Raimondo, A. G. R. Gordon, L. H. Rhym, T. Samad, C. MacIsaac, J. Witten, H. Mughal, T. M. Chicz, Y. Xu, R. P. McNamara, S. Bhatia, G. Alter, R. Langer and D. G. Anderson, Enhancing the immunogenicity of lipid-nanoparticle mRNA vaccines by adjuvanting the ionizable lipid and the mRNA, Nat. Biomed. Eng., 2025, 9, 167–184 CrossRef PubMed.
- X. Han, M.-G. Alameh, K. Butowska, J. J. Knox, K. Lundgreen, M. Ghattas, N. Gong, L. Xue, Y. Xu, M. Lavertu, P. Bates, J. Xu, G. Nie, Y. Zhong, D. Weissman and M. J. Mitchell, Adjuvant lipidoid-substituted lipid nanoparticles augment the immunogenicity of SARS-CoV-2 mRNA vaccines, Nat. Nanotechnol., 2023, 18, 1105–1114 CrossRef PubMed.
- B. Brook, V. Duval, S. Barman, L. Speciner, C. Sweitzer, A. Khanmohammed, M. Menon, K. Foster, P. Ghosh, K. Abedi, J. Koster, E. Nanishi, L. R. Baden, O. Levy, T. VanCott, R. Micol and D. J. Dowling, Adjuvantation of a SARS-CoV-2 mRNA vaccine with controlled tissue-specific expression of an mRNA encoding IL-12p70, Sci. Transl. Med., 2024, 16, eadm8451 CrossRef PubMed.
- F. Salerno, S. Engels, M. van den Biggelaar, F. P. J. van Alphen, A. Guislain, W. Zhao, D. L. Hodge, S. E. Bell, J. P. Medema, M. von Lindern, M. Turner, H. A. Young and M. C. Wolkers, Translational repression of pre-formed cytokine-encoding mRNA prevents chronic activation of memory T cells, Nat. Immunol., 2018, 19, 828–837 CrossRef.
- C. A. Piccirillo, E. Bjur, I. Topisirovic, N. Sonenberg and O. Larsson, Translational control of immune responses: from transcripts to translatomes, Nat. Immunol., 2014, 15, 503–511 CrossRef PubMed.
- X. Su, Y. Yu, Y. Zhong, E. G. Giannopoulou, X. Hu, H. Liu, J. R. Cross, G. Rätsch, C. M. Rice and L. B. Ivashkiv, Interferon-γ regulates cellular metabolism and mRNA translation to potentiate macrophage activation, Nat. Immunol., 2015, 16, 838–849 CrossRef PubMed.
- A. De Beuckelaer, C. Pollard, S. Van Lint, K. Roose, L. Van Hoecke, T. Naessens, V. K. Udhayakumar, M. Smet, N. Sanders, S. Lienenklaus, X. Saelens, S. Weiss, G. Vanham, J. Grooten and S. De Koker, Type I Interferons Interfere with the Capacity of mRNA Lipoplex Vaccines to Elicit Cytolytic T Cell Responses, Mol. Ther., 2016, 24, 2012–2020 Search PubMed.
- N. Pardi, M. J. Hogan, F. W. Porter and D. Weissman, mRNA vaccines — a new era in vaccinology, Nat. Rev. Drug Discovery, 2018, 17, 261–279 Search PubMed.
- K. Karikó, H. Muramatsu, F. A. Welsh, J. Ludwig, H. Kato, S. Akira and D. Weissman, Incorporation of Pseudouridine Into mRNA Yields Superior Nonimmunogenic Vector With Increased Translational Capacity and Biological Stability, Mol. Ther., 2008, 16, 1833–1840 CrossRef PubMed.
- S. Xu, K. Yang, R. Li and L. Zhang, mRNA Vaccine Era—Mechanisms, Drug Platform and Clinical Prospection, Int. J. Mol. Sci., 2020, 21, 6582 CrossRef PubMed.
- J. Deckers, T. Anbergen, A. M. Hokke, A. de Dreu, D. P. Schrijver, K. de Bruin, Y. C. Toner, T. J. Beldman, J. B. Spangler, T. F. A. de Greef, F. Grisoni, R. van der Meel, L. A. B. Joosten, M. Merkx, M. G. Netea and W. J. M. Mulder, Engineering cytokine therapeutics, Nat. Rev. Bioeng., 2023, 1, 286–303 Search PubMed.
- T. Kawasaki and T. Kawai, Toll-Like Receptor Signaling Pathways, Front. Immunol., 2014, 5, 461 Search PubMed.
- K. D. Nance and J. L. Meier, Modifications in an Emergency: The Role of N1-Methylpseudouridine in COVID-19 Vaccines, ACS Cent. Sci., 2021, 7, 748–756 CrossRef PubMed.
- E. A. Chiocca, J. S. Yu, R. V. Lukas, I. H. Solomon, K. L. Ligon, H. Nakashima, D. A. Triggs, D. A. Reardon, P. Wen, B. M. Stopa, A. Naik, J. Rudnick, J. L. Hu, P. Kumthekar, B. Yamini, J. Y. Buck, N. Demars, J. A. Barrett, A. B. Gelb, J. Zhou, F. Lebel and L. J. N. Cooper, Regulatable interleukin-12 gene therapy in patients with recurrent high-grade glioma: Results of a phase 1 trial, Sci. Transl. Med., 2019, 11, eaaw5680 Search PubMed.
- J. Lee, M. C. Woodruff, E. H. Kim and J.-H. Nam, Knife's edge: Balancing immunogenicity and reactogenicity in mRNA vaccines, Exp. Mol. Med., 2023, 55, 1305–1313 Search PubMed.
- D. Bitounis, E. Jacquinet, M. A. Rogers and M. M. Amiji, Strategies to reduce the risks of mRNA drug and vaccine toxicity, Nat. Rev. Drug Discovery, 2024, 23, 281–300 Search PubMed.
- P. S. Kowalski, A. Rudra, L. Miao and D. G. Anderson, Delivering the Messenger: Advances in Technologies for Therapeutic mRNA Delivery, Mol. Ther., 2019, 27, 710–728 Search PubMed.
- M. A. Islam, J. Rice, E. Reesor, H. Zope, W. Tao, M. Lim, J. Ding, Y. Chen, D. Aduluso, B. R. Zetter, O. C. Farokhzad and J. Shi, Adjuvant-pulsed mRNA vaccine nanoparticle for immunoprophylactic and therapeutic tumor suppression in mice, Biomaterials, 2021, 266, 120431 CrossRef PubMed.
- B. Brook, V. Duval, S. Barman, L. Speciner, C. Sweitzer, A. Khanmohammed, M. Menon, K. Foster, P. Ghosh, K. Abedi, J. Koster, E. Nanishi, L. R. Baden, O. Levy, T. VanCott, R. Micol and D. J. Dowling, Adjuvantation of a SARS-CoV-2 mRNA vaccine with controlled tissue-specific expression of an mRNA encoding IL-12p70, Sci. Transl. Med., 2024, 16(757), eadm8451 Search PubMed.
- N. Chen, M. M. Johnson, M. A. Collier, M. D. Gallovic, E. M. Bachelder and K. M. Ainslie, Tunable degradation of acetalated dextran microparticles enables controlled vaccine adjuvant and antigen delivery to modulate adaptive immune responses, J. Controlled Release, 2018, 273, 147–159 CrossRef.
- L. P. Keegan, K. Hajji and M. A. O'Connell, Adenosine Deaminase Acting on RNA (ADAR) Enzymes: A Journey from Weird to Wondrous, Acc. Chem. Res., 2023, 56, 3165–3174 Search PubMed.
- L. P. Keegan, A. Gallo and M. A. O'Connell, The many roles of an RNA editor, Nat. Rev. Genet., 2001, 2, 869–878 Search PubMed.
- C.-X. Chen, D.-S. C. Cho, Q. Wang, F. Lai, K. C. Carter and K. Nishikura, A third member of the RNA-specific adenosine deaminase gene family, ADAR3, contains both single- and double-stranded RNA binding domains, RNA, 2000, 6, 755–767 Search PubMed.
- K. Nishikura, A-to-I editing of coding and non-coding RNAs by ADARs, Nat. Rev. Mol. Cell Biol., 2016, 17, 83–96 CrossRef PubMed.
- M. H. Tan, Q. Li, R. Shanmugam, R. Piskol, J. Kohler, A. N. Young, K. I. Liu, R. Zhang, G. Ramaswami, K. Ariyoshi, A. Gupte, L. P. Keegan, C. X. George, A. Ramu, N. Huang, E. A. Pollina, D. S. Leeman, A. Rustighi, Y. P. S. Goh, F. Aguet, K. G. Ardlie, B. B. Cummings, E. T. Gelfand, G. Getz, K. Hadley, R. E. Handsaker, K. H. Huang, S. Kashin, K. J. Karczewski, M. Lek, X. Li, D. G. MacArthur, J. L. Nedzel, D. T. Nguyen, M. S. Noble, A. V. Segrè, C. A. Trowbridge, T. Tukiainen, N. S. Abell, B. Balliu, R. Barshir, O. Basha, A. Battle, G. K. Bogu, A. Brown, C. D. Brown, S. E. Castel, L. S. Chen, C. Chiang, D. F. Conrad, N. J. Cox, F. N. Damani, J. R. Davis, O. Delaneau, E. T. Dermitzakis, B. E. Engelhardt, E. Eskin, P. G. Ferreira, L. Frésard, E. R. Gamazon, D. Garrido-Martín, A. D. H. Gewirtz, G. Gliner, M. J. Gloudemans, R. Guigo, I. M. Hall, B. Han, Y. He, F. Hormozdiari, C. Howald, H. Kyung Im, B. Jo, E. Yong Kang, Y. Kim, S. Kim-Hellmuth, T. Lappalainen, G. Li, X. Li, B. Liu, S. Mangul, M. I. McCarthy, I. C. McDowell, P. Mohammadi, J. Monlong, S. B. Montgomery, M. Muñoz-Aguirre, A. W. Ndungu, D. L. Nicolae, A. B. Nobel, M. Oliva, H. Ongen, J. J. Palowitch, N. Panousis, P. Papasaikas, Y. Park, P. Parsana, A. J. Payne, C. B. Peterson, J. Quan, F. Reverter, C. Sabatti, A. Saha, M. Sammeth, A. J. Scott, A. A. Shabalin, R. Sodaei, M. Stephens, B. E. Stranger, B. J. Strober, J. H. Sul, E. K. Tsang, S. Urbut, M. van de Bunt, G. Wang, X. Wen, F. A. Wright, H. S. Xi, E. Yeger-Lotem, Z. Zappala, J. B. Zaugg, Y.-H. Zhou, J. M. Akey, D. Bates, J. Chan, L. S. Chen, M. Claussnitzer, K. Demanelis, M. Diegel, J. A. Doherty, A. P. Feinberg, M. S. Fernando, J. Halow, K. D. Hansen, E. Haugen, P. F. Hickey, L. Hou, F. Jasmine, R. Jian, L. Jiang, A. Johnson, R. Kaul, M. Kellis, M. G. Kibriya, K. Lee, J. B. Li, Q. Li, X. Li, J. Lin, S. Lin, S. Linder, C. Linke, Y. Liu, M. T. Maurano, B. Molinie, S. B. Montgomery, J. Nelson, F. J. Neri, M. Oliva, Y. Park, B. L. Pierce, N. J. Rinaldi, L. F. Rizzardi, R. Sandstrom, A. Skol, K. S. Smith, M. P. Snyder, J. Stamatoyannopoulos, B. E. Stranger, H. Tang, E. K. Tsang, L. Wang, M. Wang, N. Van Wittenberghe, F. Wu, R. Zhang, C. R. Nierras, P. A. Branton, L. J. Carithers, P. Guan, H. M. Moore, A. Rao, J. B. Vaught, S. E. Gould, N. C. Lockart, C. Martin, J. P. Struewing, S. Volpi, A. M. Addington, S. E. Koester, A. R. Little, L. E. Brigham, R. Hasz, M. Hunter, C. Johns, M. Johnson, G. Kopen, W. F. Leinweber, J. T. Lonsdale, A. McDonald, B. Mestichelli, K. Myer, B. Roe, M. Salvatore, S. Shad, J. A. Thomas, G. Walters, M. Washington, J. Wheeler, J. Bridge, B. A. Foster, B. M. Gillard, E. Karasik, R. Kumar, M. Miklos, M. T. Moser, S. D. Jewell, R. G. Montroy, D. C. Rohrer, D. R. Valley, D. A. Davis, D. C. Mash, A. H. Undale, A. M. Smith, D. E. Tabor, N. V. Roche, J. A. McLean, N. Vatanian, K. L. Robinson, L. Sobin, M. E. Barcus, K. M. Valentino, L. Qi, S. Hunter, P. Hariharan, S. Singh, K. S. Um, T. Matose, M. M. Tomaszewski, L. K. Barker, M. Mosavel, L. A. Siminoff, H. M. Traino, P. Flicek, T. Juettemann, M. Ruffier, D. Sheppard, K. Taylor, S. J. Trevanion, D. R. Zerbino, B. Craft, M. Goldman, M. Haeussler, W. J. Kent, C. M. Lee, B. Paten, K. R. Rosenbloom, J. Vivian, J. Zhu, A. Chawla, G. Del Sal, G. Peltz, A. Brunet, D. F. Conrad, C. E. Samuel, M. A. O'Connell, C. R. Walkley, K. Nishikura, J. B. Li and G. T. Consortium, D. A. Laboratory, G. Coordinating Center—Analysis Working, G. Statistical Methods groups—Analysis Working, G. g. Enhancing, N. I. H. C. Fund, Nih/Nci, Nih/Nhgri, Nih/Nimh, Nih/Nida, N. Biospecimen Collection Source Site—R. Biospecimen Collection Source Site—V. Biospecimen Core Resource—B. Brain Bank Repository—University of Miami Brain Endowment, M. Leidos Biomedical—Project, E. Study, I. Genome Browser Data, Visualization—Ebi, I. Genome Browser Data and U. o. C. S. C. Visualization—Ucsc Genomics Institute, Dynamic landscape and regulation of RNA editing in mammals, Nature, 2017, 550, 249–254 CrossRef PubMed.
- P. Sharma, F. Yan, V. A. Doronina, H. Escuin-Ordinas, M. D. Ryan and J. D. Brown, 2A peptides provide distinct solutions to driving stop-carry on translational recoding, Nucleic Acids Res., 2012, 40, 3143–3151 CrossRef PubMed.
- M. L. L. Donnelly, G. Luke, A. Mehrotra, X. Li, L. E. Hughes, D. Gani and M. D. Ryan, Analysis of the aphthovirus 2A/2B polyprotein ‘cleavage’ mechanism indicates not a proteolytic reaction, but a novel translational effect: a putative ribosomal ‘skip’, J. Gen. Virol., 2001, 82, 1013–1025 CrossRef.
- J. Chng, W. Tianhua, N. Rui, L. Ally, H. K. Meng, S. C. L. Ho, G. Peter, B. Xuezhi and Y. and Yang, Cleavage efficient 2A peptides for high level monoclonal antibody expression in CHO cells, mAbs, 2015, 7, 403–412 CrossRef PubMed.
- J. H. Kim, S.-R. Lee, L.-H. Li, H.-J. Park, J.-H. Park, K. Y. Lee, M.-K. Kim, B. A. Shin and S.-Y. Choi, High Cleavage Efficiency of a 2A Peptide Derived from Porcine Teschovirus-1 in Human Cell Lines, Zebrafish and Mice, PLoS One, 2011, 6, e18556 CrossRef.
- T. Z. Khatib, A. Osborne, S. Yang, Z. Ali, W. Jia, I. Manyakin, K. Hall, R. Watt, P. S. Widdowson and K. R. Martin, Receptor-ligand supplementation via a self-cleaving 2A peptide–based gene therapy promotes CNS axonal transport with functional recovery, Sci. Adv., 2021, 7, eabd2590 CrossRef PubMed.
- C. C. Thoreen, L. Chantranupong, H. R. Keys, T. Wang, N. S. Gray and D. M. Sabatini, A unifying model for mTORC1-mediated regulation of mRNA translation, Nature, 2012, 485, 109–113 CrossRef PubMed.
- R. Colina, M. Costa-Mattioli, R. J. O. Dowling, M. Jaramillo, L.-H. Tai, C. J. Breitbach, Y. Martineau, O. Larsson, L. Rong, Y. V. Svitkin, A. P. Makrigiannis, J. C. Bell and N. Sonenberg, Translational control of the innate immune response through IRF-7, Nature, 2008, 452, 323–328 CrossRef PubMed.
- M. S. Lee, B. Kim, G. T. Oh and Y.-J. Kim, OASL1 inhibits translation of the type I interferon–regulating transcription factor IRF7, Nat. Immunol., 2013, 14, 346–355 CrossRef PubMed.
- M.-K. Ge, C. Zhang, N. Zhang, P. He, H.-Y. Cai, S. Li, S. Wu, X.-L. Chu, Y.-X. Zhang, H.-M. Ma, L. Xia, S. Yang, J.-X. Yu, S.-Y. Yao, X.-L. Zhou, B. Su, G.-Q. Chen and S.-M. Shen, The tRNA-GCN2-FBXO22-axis-mediated mTOR ubiquitination senses amino acid insufficiency, Cell Metab., 2023, 35, 2216–2230e2218 CrossRef PubMed.
- A. J. Sadler and B. R. G. Williams, in Interferon: The 50th Anniversary, ed. P. M. Pitha, Springer Berlin Heidelberg, Berlin, Heidelberg, 2007, pp. 253–292, DOI:10.1007/978-3-540-71329-6_13.
- M. A. García, J. Gil, I. Ventoso, S. Guerra, E. Domingo, C. Rivas and M. Esteban, Impact of Protein Kinase PKR in Cell Biology: from Antiviral to Antiproliferative Action, Microbiol. Mol. Biol. Rev., 2006, 70, 1032–1060 CrossRef PubMed.
- X. Han, M.-G. Alameh, N. Gong, L. Xue, M. Ghattas, G. Bojja, J. Xu, G. Zhao, C. C. Warzecha, M. S. Padilla, R. El-Mayta, G. Dwivedi, Y. Xu, A. E. Vaughan, J. M. Wilson, D. Weissman and M. J. Mitchell, Fast and facile synthesis of amidine-incorporated degradable lipids for versatile mRNA delivery in vivo, Nat. Chem., 2024, 16, 1687–1697 CrossRef PubMed.
- X. Han, J. Xu, Y. Xu, M.-G. Alameh, L. Xue, N. Gong, R. El-Mayta, R. Palanki, C. C. Warzecha, G. Zhao, A. E. Vaughan, J. M. Wilson, D. Weissman and M. J. Mitchell, In situ combinatorial synthesis of degradable branched lipidoids for systemic delivery of mRNA therapeutics and gene editors, Nat. Commun., 2024, 15, 1762 CrossRef PubMed.
- J. Yoon, E. Fagan, M. Jeong and J.-H. Park, In Situ Tumor-Infiltrating Lymphocyte Therapy by Local Delivery of an mRNA Encoding Membrane-Anchored Anti-CD3 Single-Chain Variable Fragment, ACS Nano, 2024, 18, 32401–32420 CrossRef PubMed.
- L. Zhang, K. R. More, A. Ojha, C. B. Jackson, B. D. Quinlan, H. Li, W. He, M. Farzan, N. Pardi and H. Choe, Effect of mRNA-LNP components of two globally-marketed COVID-19 vaccines on efficacy and stability, npj Vaccines, 2023, 8, 156 CrossRef PubMed.
- J. C. Castle, S. Kreiter, J. Diekmann, M. Löwer, N. van de Roemer, J. de Graaf, A. Selmi, M. Diken, S. Boegel, C. Paret, M. Koslowski, A. N. Kuhn, C. M. Britten, C. Huber, Ö. Türeci and U. Sahin, Exploiting the Mutanome for Tumor Vaccination, Cancer Res., 2012, 72, 1081–1091 CrossRef PubMed.
- A. Batista-Duharte, D. T. Martínez and I. Z. Carlos, Efficacy and safety of immunological adjuvants. Where is the cut-off?, Biomed. Pharmacother., 2018, 105, 616–624 Search PubMed.
- S. G. Reed, M. T. Orr and C. B. Fox, Key roles of adjuvants in modern vaccines, Nat. Med., 2013, 19, 1597–1608 CrossRef PubMed.
- C. Cohet, R. van der Most, V. Bauchau, R. Bekkat-Berkani, T. M. Doherty, A. Schuind, F. Tavares Da Silva, R. Rappuoli, N. Garçon and B. L. Innis, Safety of AS03-adjuvanted influenza vaccines: A review of the evidence, Vaccine, 2019, 37, 3006–3021 CrossRef PubMed.
- B. Pulendran, P. S. Arunachalam and D. T. O'Hagan, Emerging concepts in the science of vaccine adjuvants, Nat. Rev. Drug Discovery, 2021, 20, 454–475 CrossRef PubMed.
- S. Jhunjhunwala, C. Hammer and L. Delamarre, Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion, Nat. Rev. Cancer, 2021, 21, 298–312 CrossRef PubMed.
- T. Zhao, Y. Cai, Y. Jiang, X. He, Y. Wei, Y. Yu and X. Tian, Vaccine adjuvants: mechanisms and platforms, Signal Transduction Targeted Ther., 2023, 8, 283 Search PubMed.
- K. E. Darrah and A. Deiters, Translational control of gene function through optically regulated nucleic acids, Chem. Soc. Rev., 2021, 50, 13253–13267 RSC.
- A. Estévez-Torres, C. Crozatier, A. Diguet, T. Hara, H. Saito, K. Yoshikawa and D. Baigl, Sequence-independent and reversible photocontrol of transcription/expression systems using a photosensitive nucleic acid binder, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 12219–12223 CrossRef PubMed.
- Z. Wang, J. C. Valera, X. Zhao, Q. Chen and J. Silvio Gutkind, mTOR co-targeting strategies for head and neck cancer therapy, Cancer Metastasis Rev., 2017, 36, 491–502 CrossRef PubMed.
- L. Amurri, B. Horvat and M. Iampietro, Interplay between RNA viruses and cGAS/STING axis in innate immunity, Front. Cell. Infect. Microbiol., 2023, 13, 1172739 CrossRef PubMed.
Footnote |
| † X. Z., X. T., Y. D. and N. G. contributed equally to this paper. |
| This journal is © The Royal Society of Chemistry 2026 |