Open Access Article
Open Access ArticleExposure single-cell metabolomics mass spectrometry reveals HFPO-DA toxicity mechanisms
Yuanxing
Liu†
a,
Wenmei
Zhang†
a,
Hanyu
Yuan
a,
Tong
Pei
a,
Tian
Chen
a,
Ke
Gao
b,
Denghui
Guo
a,
Xianfa
Yang
c,
Naihe
Jing
*c,
Guangsheng
Guo
a and
Xiayan
Wang
 *a
*a
aState Key Laboratory of Materials Low-Carbon Recycling, Center of Excellence for Environmental Safety and Biological Effects, Department of Chemistry, College of Chemistry and Life Science, Beijing University of Technology, Beijing 100124, China. E-mail: xiayanwang@bjut.edu.cn
bKey Laboratory of Beijing on Regional Air Pollution Control, Department of Environmental Science, Beijing University of Technology, Beijing 100124, China
cGuangzhou National Laboratory, Guangzhou 510005, China. E-mail: jing_naihe@gzlab.ac.cn
First published on 3rd December 2025
Abstract
Hexafluoropropylene oxide-dimer acid (HFPO-DA) is widely used in food packaging bags, surfactants, lubricants, etc. As a new substitute for the persistent organic pollutant perfluorooctanoic acid (PFOA), it has been proven to have potential health and environmental risks similar to other perfluorinated compounds. However, studies based on the average levels of many cells or tissues cannot accurately reflect the toxic mechanism of HFPO-DA. Here, we construct a high-throughput exposure single-cell metabolomics mass spectrometry analysis platform based on intact living-cell electrolaunching ionization MS analysis, optimize the detection conditions of the platform, and use it to investigate the effects of a wide range of HFPO-DA concentrations on the metabolism of mouse embryonic stem cells (E14TG2a cells). We evaluate the effect of HFPO-DA exposure on the viability of E14TG2a cells using the CCK8 method. The results showed that exposure to HFPO-DA at a concentration below 1 mmol L−1 within 24 hours had no significant effect on the viability of E14TG2a cells, while exposure above 1 mmol L−1 led to significant differences in cell viability. Comprehensive single-cell metabolic analysis revealed that even environmental concentrations of HFPO-DA exposure that did not affect cell viability could affect cellular metabolic changes, and the levels of some metabolites showed a non-monotonic dose–response relationship with HFPO-DA. HFPO-DA exposure interfered with metabolic pathways such as nitrogen metabolism, arginine biosynthesis, and arginine and proline metabolism, thereby affecting the homeostasis of basic biological metabolic processes such as nucleotide metabolism and amino acid metabolism in E14TG2a cells. This study indicates that although HFPO-DA has relatively low cytotoxicity, it can still interfere with the metabolic processes of E14TG2a cells, suggesting that the safety of HFPO-DA needs further evaluation.
Introduction
Hexafluoropropylene oxide dimer acid (HFPO-DA) is an industrial alternative to the persistent organic pollutant perfluorooctanoic acid (PFOA), and is widely used in lubricants, food packaging, fluorinated polymers, and electroplating applications.1–3 As a per- and polyfluoroalkyl substance (PFAS), the structural properties make it resistant to degradation, enable long-distance transmissibility, and contribute to its chemical stability, raising concerns about potential environmental and health risks.4–7 HFPO-DA has become widely distributed in various environmental media such as water and soil.3,8,9 In 2024, Jensen et al. detected HFPO-DA concentrations as high as 229 ng L−1 in groundwater near the Cape Fear River in North Carolina.9 Additionally, studies have identified PFAS contamination in agricultural soils of China's Fuxin region, with concentrations ranging from 57.36 to 1271.06 pg g−1, where HFPO-DA emerged as the predominant pollutant.10 Moreover, 35 air samples from Huantai County, China, showed higher indoor dust concentrations of PFOA, HFPO-DA, and HFPO-TA than other PFASs.11Pollutants spread through environmental media (water, soil, and air) and are directly contacted or absorbed by organisms. Subsequently, they are gradually enriched through the food chain and finally accumulate in organisms, causing harm.12–14 For example, the concentration of PFAS in fish in the northern Bohai Sea reached 262.92 ng g−1. Among them, the concentration of HAPO-DA increased significantly with the increase of individual fish size and lipid content, further increasing the health risk.15 Furthermore, toxicological studies have confirmed that HFPO-DA possesses various toxicities, including hepatotoxicity, neurotoxicity, and reproductive and developmental toxicity.16,17 For example, exposure to PFOA and its alternatives (such as HFPO-DA and HFPO-TA) at concentrations of 5–500 µg L−1 disrupted embryonic development in zebrafish, leading to altered heart rate, changes in enzyme activity, and differential gene expression. Specifically, HFPO-DA was found to impair neurodevelopment and lipid homeostasis, suggesting that these alternatives may pose risks comparable to or even distinct from those of PFOA.18 However, these studies are based on the average levels of a large number of cells or tissues and do not accurately reflect the distribution characteristics and effects of HFPO-DA at the cellular level. Additionally, because of the averaging of rare-cell information in population-based analyses, and issues with the sensitivity of detection techniques, most endpoints of HFPO-DA exposure studies show changes in cell morphology or function after such exposure occurs.19,20 On the other hand, current exposure studies on HFPO-DA are mostly conducted at concentrations higher than environmental levels, and for low-dose, long-term exposure scenarios, changes at the population level may be overlooked, which poses a challenge to the comprehensive assessment of the potential risk of environmental pollutants.
Single-cell analysis provides in-depth resolution of cellular characteristics, enabling the investigation of cellular heterogeneity, tracking of cell lineages, identification of rare cell types, and understanding of dynamic changes of cells under specific conditions.21–25 However, the sensitivity requirements for metabolite detection are elevated due to the vast number of metabolites within single cells, their extremely low concentrations, significant differences in component levels, and the inability to use amplification techniques for detection.23 Mass spectrometry (MS) has become a preferred method for detecting trace substances in complex biological samples in recent years, not only providing molecular weight information and chemical or biological information of metabolites, but also achieving high-sensitivity, simultaneous multi-component detection, and superior qualitative capabilities in single-cell metabolic analysis.26–28 Various single-cell metabolite detection methods based on mass spectrometry have been developed, including single-probe mass spectrometry, single-cell analysis with probe ESI-MS for the detection of metabolites at cellular and subcellular levels,29 label-free mass cytometry,30,31 organic mass cytometry,27 and intact living-cell electrolaunching ionization MS (ILCEI-MS).32 Additionally, some studies have assessed the accumulation of environmental pollutants in organisms and single-cell heterogeneities on a single-cell level.33,34 For example, Li et al. constructed a chemo-selective single-cell metabolomics analysis platform for quantitative, in-depth, and minimally destructive characterization of cis-diol metabolites during embryo development from the single oocyte level to the tailbud-stage embryos, and to evaluate their spatiotemporal behavior.35 Deng et al. constructed a single-cell mass cytometry system to employ the accumulation behaviors and heterogeneities of perfluorooctanesulfonic acid (PFOS) in zebrafish primary organ cells.33 Nevertheless, these studies focus on the detection of trace pollutants in cells, and few reports exist on the toxicological mechanisms of environmental dose pollutants and their molecular-level impacts.
In this study, we constructed a high-throughput exposure single-cell metabolomics mass spectrometry platform based on intact living-cell electrolaunching ionization MS analysis (ILCEI-MS) to investigate the metabolic effects of the environmental pollutant HFPO-DA on embryonic stem cells (E14TG2a cells) across a wide concentration range (including environmentally relevant levels). After optimizing parameters such as single-cell detection time, mass spectrometry detection voltage, ion transfer tube temperature, and cell driving pressure, we detected the metabolites of E14TG2a cells exposed to different concentrations of HFPO-DA (0–10 mmol L−1) and analyzed their metabolic changes. We also used the CCK8 method to evaluate the inhibitory effects of various concentrations of HFPO-DA on the proliferation of E14TG2a cells. Based on metabolic analysis, we assessed the metabolic pathways involved in different concentration levels of differential metabolites and found that the pyrimidine metabolism, purine metabolism, nitrogen metabolism, arginine biosynthesis, and arginine and proline metabolism pathways were associated with HFPO-DA exposure.
Results and discussion
Exposure single-cell metabolomics mass spectrometry platform
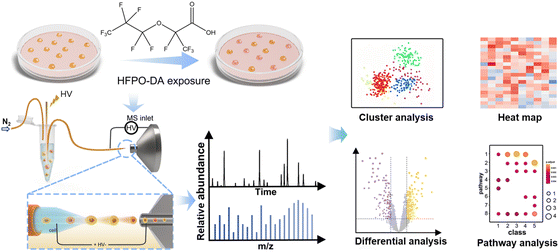
ILCEI-MS is a technology capable of simultaneously achieving online dispersion and undiluted mass spectrometric sampling of single cells, offering high sensitivity and throughput.32 We constructed an exposure single-cell metabolomics mass spectrometry analysis platform based on the established ILCEI-MS technology (Fig. 1) to explore the mechanism of HFPO-DA exposure through metabolomics at the single-cell level. To ensure compatibility of solvents with mass spectrometric detection while maintaining the osmotic pressure and physiological state of the cells to the greatest extent, we resuspended the cells in a 40 mmol L−1 ammonium formate solution. The cells were injected into a capillary with an inner diameter of 16 µm using high-purity nitrogen, and were electrolaunching-ionized at the emitter tip after being dispersed and requeued in the capillary. The obtained single-cell MS data were processed through a single-cell metabolomics analysis platform developed by our group. Ion information from the samples was obtained through background subtraction, normalization, and peak alignment. Further, nonlinear dimension reduction analysis was employed to visualize the distribution of metabolite molecular features in a two-dimensional space. The distribution of metabolite molecular features in the two-dimensional space reveals the differences in the overall metabolic profile of E14TG2a cells upon exposure. Differences between two or multiple groups of data are assessed using t-tests or variance analysis to screen for significantly altered characteristic metabolite ions. Subsequently, the small molecule metabolome database is confirmed by non-targeted scanning DIA for differential metabolites. Finally, pathway enrichment analysis and topological analysis were conducted to study the metabolic pathways affected by significantly different metabolites, with changes in metabolites depicted using violin plots.Effect of ion transfer tube temperature
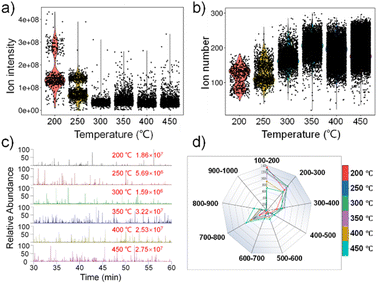
We investigated the impact of the ion transfer tube temperature on the single-cell detection of E14TG2a cells. The results are shown in Fig. 2. When the temperature was between 200 °C and 250 °C, the ion intensity was significantly higher than that of other groups (Fig. 2a), and the number of ions detected from a single cell differed slightly (Fig. 2b). This is because of the low temperature, which makes it difficult for the ammonium formate solution carried by the cell to evaporate quickly when it enters the ion transport tube, and the cell fragmentation is incomplete, resulting in low ionization efficiency of the cell fragments. Fig. 2c shows the extracted ion chromatogram (EIC) of single-cell detection of E14TG12a at different temperatures. When the temperature was lower than 300 °C, the number of detected single cells and the signal intensity were significantly lower. The radar chart shows that when the ion transfer tube temperature was 300 °C (green line) and 350 °C (purple line), the number of ions detected in different segments was relatively high (Fig. 2d). To obtain as much ion information as possible while the cells remain in good condition, both 300 °C and 350 °C are good choices. Considering both the ion intensity and the number of ions from a single cell, 350 °C was finally selected as the experimental condition for subsequent experiments in this study.Effect of ion source voltage
A voltage was applied to the tip of the mass emitter and the port of the ion transfer tube, forming a stable electric field, and then realizing the electric emission of living single cells. However, the voltage can affect the ionization efficiency and ion beam stability of mass spectrometry detection. Therefore, we evaluated the influence of different voltages (1.5–2.5 kV) on the single-cell detection of E14TG2a cells, as shown in Fig. S1. When the voltage was 1.75 kV, the ion intensity (Fig. S1a), the number of detected ions (Fig. S1b), and the cell detection signal (Fig. S1c) were all higher than those under other voltage conditions. The radar chart (Fig. S1d) shows that ion detection in different m/z ranges can be selectively achieved by adjusting the voltage. Among them, the number of ions in each m/z ratio segment at 1.75 kV was better than that under other conditions. Therefore, in this study, an injection voltage of 1.75 kV was finally selected as the experimental condition for subsequent experiments.Effect of driving pressure
Single cells were injected into the capillary using high-purity nitrogen, so the driving pressure affected the injection speed of the cells. We evaluated the influence of different driving pressures on single-cell detection, as shown in Fig. S2. There were no differences in the single-cell ion signal intensity (Fig. S2a) and the number of ions (Fig. S2b) under different driving conditions, indicating that they were less affected by the driving pressure. The EIC (Fig. S2c) compared the effects of samples with the same volume and density under different driving pressures. It was found that the lower the driving pressure, the better the single-cell detection signal and the greater the number of detected cells. However, we evaluated cell availability at different pressures (Table S1) and found that the higher the driving pressure, the higher the cell collection rate. In addition, by comparing the number of ion distributions in different m/z ranges (Fig. S2d), it was found that when the driving pressure was 400 psi, the number of ions obtained in each m/z range was more evenly distributed. Therefore, 400 psi was selected as the driving pressure for the samples in this study.Effect of mass spectrometry detection time
We evaluated the clustering, the ion signal intensity, and the ion detection number of single cells for 180 min continuous detection using 30 min as a time detection unit. The UMAP plot (Fig. S3a) shows that there were relatively small overall differences among the cells detected within 180 minutes, which is reflected in the smaller coordinate range of the two dimensions. It was found that the clustering of the cells was more concentrated within 60 min of continuous detection. After 60 min, the differences gradually increased and were positively correlated with the detection time. This difference might be due to metabolic changes caused by the cells being continuously immersed in ammonium formate. In addition, there was no significant difference in the signal intensity of ions detected continuously for 180 min (Fig. S3b). In comparison, the number of ions detected for more than 90 min (Fig. S3c) increased significantly. This could be because continuous cell injection led to the accumulation of un-ionized cell debris in the instrument, increasing background ions. Therefore, we kept the detection duration within 60 min in subsequent experiments.Effect of HFPO-DA on cell activity
We used the CCK-8 method to evaluate the impact of different concentrations of HFPO-DA on the viability of E14TG2a cells after 24 hours of exposure, with the results shown in Fig. S4. Compared to the control group, the cell viability of the exposure groups showed a decrease, with lower cell viability at higher exposure concentrations. When the concentration of HFPO-DA was below 1 mmol L−1, there was little effect on cell viability and no significant difference, and the cell survival rate was higher than 90%. There was a significant difference in cell viability at a concentration of 1 mmol L−1 of HFPO-DA (p < 0.05). For exposure concentrations higher than 1 mmol L−1, the cell survival rate was less than 50%, and the difference in viability was significant (p < 0.01). Furthermore, the concentration of HFPO-DA in the environment varies depending on the environmental medium. Surface water or groundwater is usually at the ng L−1 level, soil is at the pg g−1 to ng g−1 level, and in organisms it can reach the ng g−1 to µg g−1 level.9,10,36 It has been reported that the concentrations of HFPO-DA in environmental and biological samples range from 48.59 to 5850 ng L−1,9,37,38 which corresponds to 0.14 nmol L−1 to 17.7 nmol L−1. Here, we simulate the environmental concentration exposure with 10 nmol L−1 HFPO-DA, and the result shows that there is no toxicity on E14TG2a at the environmental concentration of HFPO-DA.Visual analysis of E14TG2a cells exposed to HFPO-DA
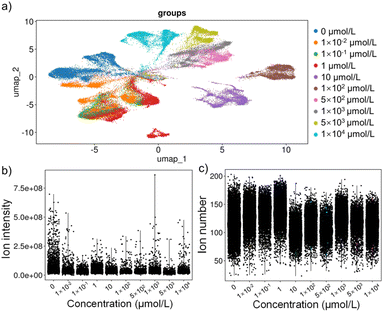
To intuitively demonstrate the effect of different concentrations of HFPO-DA exposure on the metabolism of E14TG2a cells, we performed a UMAP clustering analysis on all detected metabolite-related ions, as shown in Fig. 3a. The UMAP analysis revealed distinct metabolic profiles between the exposure and the control group, with clear clustering and separation observed between groups along the UMAP1 or UMAP2 dimensions. The separation from the control group became more pronounced as the exposure concentration increased. The clustering of exposure to environmental concentrations of HFPO-DA also showed a dispersion relative to the control group in both dimensions. Additionally, there were significant differences between and within groups at different exposure concentrations, indicating that HFPO-DA exposure led to significant metabolic disorders in E14TG2a cells, which are difficult to detect in population-level cell analyses. This highlights the value of studying the exposure mechanisms of environmental pollutants at the single-cell level.We performed UMAP analysis to cluster E14TG2a cells exposed to different concentrations of HFPO-DA (Fig. S5). The results clearly demonstrate distinct distribution patterns in the metabolic profiles between the exposed groups and the control group. Notably, even the unexposed control cells formed multiple subpopulations, which can be attributed to the inherent heterogeneity of the E14TG2a cell line. Rather than being a completely uniform population, E14TG2a consists of a mixture of undifferentiated pluripotent stem cells, early differentiated cells, and cells at various stages of pluripotency. HFPO-DA-exposed cells exhibited additional subpopulation structures, suggesting that HFPO-DA exposure may induce distinct metabolic states. Furthermore, cells exposed to different concentrations of HFPO-DA showed multiple distribution trends, indicating that the E14TG2a cell population does not respond synchronously to HFPO-DA exposure. Instead, there exists a metabolically highly altered state, which may hold significant implications for understanding the compound's toxicity mechanisms.
To further evaluate ILCEI-MS for single-cell metabolic profiling, the signals from E14TG2a cells are displayed in Fig. S6. The average number of detected cells was 32 per minute, featuring a high throughput for single-cell MS detection. Moreover, we used violin plots to compare the ion intensity information detected in different samples (Fig. 3b) and the number of ions detected in individual cells (Fig. 3c). Under different exposure concentrations, the average total ion intensity in individual cells remained relatively stable, and the differences in the number of ions detected in individual cells were also minor.
Differential analysis of E14TG2a cells exposed to HFPO-DA
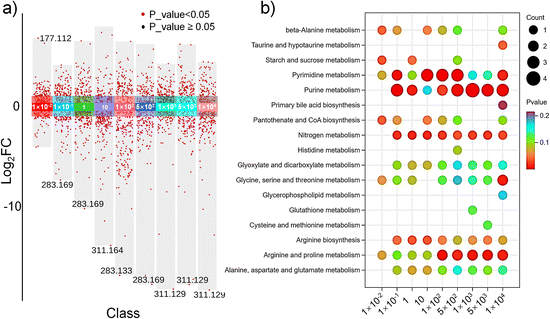
The mass spectra of E14TG2a cells before and after exposure to HFPO-DA are shown in Fig. S7. Compared with unexposed cells, there were multiple differences in ionic strength after HFPO-DA exposure, such as m/z 311.16 and m/z 219.03, which were at high levels in E14TG2a cells that were not exposed to HFPO-DA. To better understand the metabolic changes in E14TG2a cells exposed to HFPO-DA, we used the ANOVA method to compare the ion expression in different HFPO-DA exposure groups (Fig. 4a). As the exposure concentration increased, the number of differentially expressed ions significantly increased, and the significance of these differences became more pronounced. By comparing the differential ions with the m/z (mass deviation < 5 ppm) and MS2 information of standardized compounds from the mzCloud and HMDB databases, we identified 168 significantly differentially expressed metabolites (Table S2). The MS2 information of the differential metabolites was obtained by analyzing the lysate of population cells using traditional metabolomics methods. The heatmap depicts the expression of some differentially expressed metabolites in all detected single cells exposed to HFPO-DA (Fig. S8), with the color key indicating the expression levels.Based on the identified significantly differentially expressed metabolites, we utilized MetaboAnalyst and the MetWare Metabolism Cloud Platform to investigate the impact of HFPO-DA exposure on the metabolic pathways of E14TG2a cells. It was revealed that HFPO-DA exposure primarily disrupted 17 metabolic pathways, including pyrimidine metabolism, purine metabolism, nitrogen metabolism, arginine biosynthesis, and arginine and proline metabolism (Fig. 4b). Among 17 metabolic pathways, glycine, serine, and threonine metabolism, arginine biosynthesis, arginine and proline metabolism, alanine, aspartate, and glutamate metabolism are part of amino acid metabolism, nitrogen metabolism, and glyoxylate and dicarboxylate metabolism, which are supplementary to amino acid metabolism. Purine metabolism and pyrimidine metabolism are part of nucleotide metabolic pathways. Even under concentrations of HFPO-DA exposure with no apparent cytotoxicity (<1 mmol L−1), we observed significant changes in E14TG2a cellular metabolism. Among them, environmental concentration (10 nmol L−1) exposure mainly interfered with five metabolic pathways, including beta-alanine metabolism, starch and sucrose metabolism, and pyrimidine metabolism. These pathways are all related to the supply of energy for cells. Compared to the environmental concentration of HFPO-DA exposure, higher HFPO-DA concentration exposure disrupted even more metabolic pathways, primarily including purine metabolism, nitrogen metabolism, and arginine biosynthesis. The significance of metabolic differences in purine metabolism and nitrogen metabolism was similar across different groups. Arginine biosynthesis showed more pronounced significant differences at lower exposure concentrations. Additionally, the significance of differences in arginine and proline metabolism increased with higher exposure concentrations, indicating that the metabolic pathway was significantly affected by HFPO-DA.
Metabolic pathway analysis of E14TG2a cells exposed to HFPO-DA
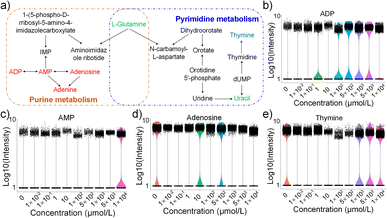
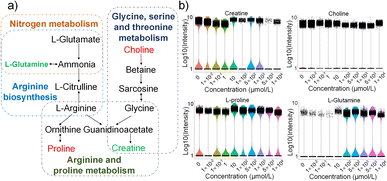
According to the KEGG database, we drew the metabolic pathway maps related to nucleotides and amino acids in E14TG2a cells exposed to HFPO-DA. Fig. 5a depicts the impact of HFPO-DA exposure on nucleotide metabolism in E14TG2a cells. It mainly affects the purine and pyrimidine metabolic pathways by interfering with ADP, AMP, adenosine, uracil, and thymine levels. Fig. 5b–e are violin plots showing the content levels of characteristic metabolites in different exposure groups. The difference analysis results (Table S3) show that E14TG2a cells were exposed to HFPO-DA, and the details are shown in Table S4 to Table S12. The screening criteria for differential metabolites were a P-value <0.05 and a Fold Change >1.5. ADP, AMP, and adenosine exhibit increased levels. The level of thymine is significantly reduced only when exposed to 10–500 µmol L−1 of HFPO-DA, and the level of uracil shows a non-monotonic change. Nucleotides are the basic building blocks of nucleic acids, and nucleotide metabolism is central to all living systems and plays an important role in the transmission of genetic information and energy.39 ADP, AMP, and adenosine are metabolites of adenine nucleotides and play an important role in energy metabolism, nucleotide synthesis, and signal transduction. They maintain cellular energy homeostasis and normal functions through mutual transformation and synergistic effects, serving as indispensable molecules for life activities. Uracil and thymine are important nucleic acid bases. In addition, uracil can be converted to thymine by metabolic pathways;40,41 this process is necessary to maintain DNA synthesis and repair. Exposure to HFPO-DA causes changes in the levels of those metabolites that may affect signaling, energy regulation, and proliferation of E14TG2a cells.HFPO-DA mainly interferes with the pathways related to arginine biosynthesis, arginine and proline metabolism, alanine, aspartate and glutamate metabolism, glycine, serine and threonine metabolism, glyoxylate and dicarboxylate metabolism, and nitrogen metabolism in E14TG2a cells by affecting creatine, choline, proline, and glutamine (Fig. 6a). Violin plots show the levels of relevant metabolites in different exposure groups (Fig. 6b). It was found that there was a non-monotonic change trend between the content of some metabolites and the HFPO-DA concentration. For example, both creatine and glutamine showed a trend of first decrease and then increase. Compared with the control group, the contents of creatine and glutamine were down-regulated when the HFPO-DA exposure concentration was less than 10 µmol L−1, and gradually up-regulated when the exposure concentration was higher than 10 µmol L−1.
Amino acids play important roles in protein synthesis, maintenance of cellular redox balance, and signal transduction, and amino acid metabolism is the basis for maintaining normal growth, development, and tissue repair of the organism, and relates to physiological processes such as energy metabolism and neurotransmitter synthesis.42–44 Proline and glutamine play roles in protein synthesis, energy metabolism, and antioxidant defense. Furthermore, glutamine is a vital amino acid in the body and plays a crucial regulatory role at both gene and protein levels, influencing various cell-specific processes. It is integral to metabolism, serving as an oxidative fuel, a precursor for gluconeogenesis, and a substrate for lipogenesis. Additionally, glutamine supports cell integrity by promoting cell survival and proliferation, highlighting its multifaceted importance in cellular function and homeostasis.45–47 Creatine and choline are derivatives of amino acids and play important roles in energy metabolism and nerve conduction.48,49 Exposure to HFPO-DA interferes with the levels of amino acids and their derivatives in E14TG2a cells, affecting the cellular amino acid metabolic homeostasis, energy status, and stability of the cells.
Vandenberg noted that non-monotonic dose responses are widely observed across multiple levels—from cells to humans—and can be triggered by various substances such as nutrients, pharmaceuticals, and vitamins. The intensity of these responses may be influenced by the substance's mechanism of action, the specific biological system under study, and internal feedback mechanisms.50 In this study, it was found that metabolites such as uracil and glutamine have non-monotonic dose responses, which play a significant role in the synthesis of DNA and proteins, as well as in cell proliferation and energy metabolism, highlighting the complexity of the toxic effect of HFPO-DA on E14TG2a cells. Our findings suggest that non-monotonic dose responses may be a common feature of perfluoroalkyl substances (PFAS). This is exemplified by studies showing sex-specific non-monotonic dose responses to HFPO-DA in locomotion and brain gene expression over the lifespan of Drosophila melanogaster,51 as well as by the non-monotonic cytotoxic effects of PFOA and PFOS in prostate cancer cells.52
Conclusion
In summary, we constructed a high-throughput exposed single-cell metabolomics mass spectrometry analysis platform and optimized the detection conditions. It was determined that better detection information could be obtained when the ion transfer tube temperature was 350 °C, the detection voltage was 1.75 kV, the cell driving pressure was 400 psi, and the detection time was within 60 min. This platform was used to study the effects of HFPO-DA, a substitute for the persistent organic pollutant PFOA, on the metabolism of embryonic stem cells (E14TG2a cells). The results of the CCK8 assay showed that exposure to HFPO-DA at concentrations below 1 mmol L−1, including environmentally relevant levels, did not affect the viability of E14TG2a cells. In contrast, exposure above 1 mmol L−1 had a significant difference. Comprehensive single-cell metabolic analysis revealed that even non-cytotoxic HFPO-DA exposure could affect cellular metabolic changes. HFPO-DA exposure interfered with 17 metabolic pathways, including nitrogen metabolism, arginine biosynthesis, and arginine and proline metabolism, thereby affecting the homeostasis of basic biological metabolic processes such as nucleotide metabolism and amino acid metabolism in E14TG2a cells. In addition, the levels of some metabolites showed non-monotonic dose responses to HFPO-DA. The result indicates that low-dose HFPO-DA can still interfere with the biological processes of E14TG2a cells without obvious cytotoxicity, suggesting that the safety of HFPO-DA needs further evaluation. This study demonstrates the potential of single-cell metabolic analysis in pollutant exposure studies and provides new ideas and research schemes for the safety assessment of pollutants.Experimental
Cell line, reagents, and materials
Mouse embryonic stem cells (ES-E14TG2a) were a kind gift from Jinsong Li's laboratory. Mouse embryonic stem cells complete growth medium (SCSP-603) was purchased from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). Sodium pyruvate, Dulbecco's Modified Eagle's Medium (DMEM), penicillin-streptomycin (PS), 0.05% trypsin-EDTA, dimethyl sulfoxide (DMSO), and phosphate-buffered saline (PBS) were purchased from Gibco. Mass spectrometry grade water, methanol, formic acid, ammonium formate, ammonia solution, acetonitrile, isopropanol, and ion calibration solution were all purchased from Thermo Fisher Scientific (USA). MEK inhibitor (Mirdametinib) and GSK-3α/β inhibitor (Laduviglusib) were purchased from MedChemExpress (USA). HFPO-DA (purity > 97%) was purchased from Titan Scientific Co., Ltd (Shanghai, China). Hydrofluoric acid (HF) was purchased from Tianjin Fuchen Chemical Reagent Factory (China). The cell counting kit-8 (CCK-8) was purchased from Dojindo Beijing Co., Ltd (China). Polyimide-coated quartz capillaries were purchased from Yongnian Ruifeng Chromatography Devices Co., Ltd. The EC-C18 chromatographic column was purchased from Agilent (USA).Cell culture and treatment
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 g, 20 min) to obtain the supernatant and dried in a vacuum centrifugal concentrator (CV600, Beijing JM Technology Co., Ltd). The dried sample was redissolved in 5% (v/v) methanol–water and stored for use.
000 g, 20 min) to obtain the supernatant and dried in a vacuum centrifugal concentrator (CV600, Beijing JM Technology Co., Ltd). The dried sample was redissolved in 5% (v/v) methanol–water and stored for use.
Narrow-bore constant-inner-diameter emitter preparation
A capillary with an outer diameter (O.D.) of 360 µm, an inner diameter (I.D.) of 16 µm, and a total length of 40 cm was intercepted to prepare an emitter. One end of the capillary was etched to form a narrow-bore tip with a constant inner diameter using the method of gravity-assisted sleeving etching established by our group.53 The prepared emitters were stored in a dust-free environment for use.Intact living-cell electrolaunching ionization MS analysis
The ILCEI-MS system consists of three parts: single-cell injection, dispersion, and MS detection. Cell injection is composed of a pressure chamber and a high-purity nitrogen cylinder. The cell suspension was loaded into 200 µL centrifuge tubes and placed in a pressure chamber. Nitrogen with a pressure of 400 psi drove the cell suspension into the sampling end of the capillary for injection. The etched capillary end is installed at the front of the mass spectrometer with a tip of approximately 5 mm from the entrance of the ion transfer tube. A DC voltage is applied to the solution, and the cells are electrolaunching-ionized at the emitter tip after being dispersed and requeued in a capillary. Single-cell signal detection was performed using a high-resolution mass spectrometer (LTQ Orbitrap XL, Thermo Fisher Scientific, San Jose, CA, USA). In addition, to reduce the detection error of the instrument, the instrument was calibrated before the experiment. The MS parameters were set as follows: in the positive ion full MS scan, the AGC target was 1 × 105, the resolution was 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, the maximum inject time was 50 ms, the temperature of the ion transport tube was 350 °C, the m/z scan range was 100–1000, and the voltage was 1.75 kV.
000, the maximum inject time was 50 ms, the temperature of the ion transport tube was 350 °C, the m/z scan range was 100–1000, and the voltage was 1.75 kV.
LC–MS/MS analysis
The analysis was performed using an ultra-high performance liquid chromatography (UHPLC) system (Ultimate 3000, Thermo Fisher Scientific) equipped with an Eclipse Plus C18 (150 × 2.1 mm, 1.8 µm) column; column thermostat was set at 40 °C. Mobile phase “C” consists of water containing 0.01% formic acid, and mobile phase “D” consists of methanol containing 0.1% formic acid. The chromatographic separation was performed by the following gradient: 0–3 min isocratic 5% “D”, 3–10 min linear gradient 5–95% “D”, 10–14 min isocratic 95% “D”, 14–15 min linear gradient 95–5% “D”, 15–18 min isocratic 5% “D”. The flow rate was 300 µL min−1 and the injection volume was 3 µL. The LC system was coupled with a high-resolution mass spectrometer (Orbitrap Eclipse, Thermo Scientific). Mass spectrometry data acquisition was conducted using the Xcalibur software, and the source and mass spectrum parameters are as follows: Spray voltage 3.4 kV for positive polarity, the ion transfer tube temperature was 350 °C, resolution was 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, the m/z scan range was 100–1000, the AGC Target was set to standard, and the MIT was 22 ms. For MS2 scans, the AGC target was 5 × 104, the MIT was 45 ms, and the collision energy for high-energy collision-induced dissociation (HCD) ranged from 10 to 90 eV.
000, the m/z scan range was 100–1000, the AGC Target was set to standard, and the MIT was 22 ms. For MS2 scans, the AGC target was 5 × 104, the MIT was 45 ms, and the collision energy for high-energy collision-induced dissociation (HCD) ranged from 10 to 90 eV.
Single-cell metabolomics data analysis
Single-cell signal screening and analysis were performed on a single-cell analysis platform developed by our research group. We took the cell membrane's main component of phosphatidylcholine (PC 34![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, m/z 760.58) as the screening criteria for the single-cell signal, subtracting the same batch of cell-free background solution signal, and the cell intensities at least 3 × 104 after subtracting the background solution signals of the same batch without cells. In addition, ions with a signal-to-noise ratio greater than 3 times and occurrence frequency greater than 10% were extracted. Uniform Manifold Approximation and Projection (UMAP) clustering analysis was conducted using R language-based programs to study the overall differences in metabolic profiles between the HFPO-DA exposure group and the control group. Differential analysis of cells was performed using the MetWare Cloud platform (https://cloud.metware.cn), and significantly different metabolite ions were screened and displayed in a volcano plot. We used the online mzCloud database (https://www.mzcloud.org), the Human Metabolome Database (HMDB), and Compound Discoverer 3.3 (ThermoFisher Scientific) software to qualitatively identify the metabolite ions through accurate m/z and fragmentation information to mark characteristic ions. Then, we used the MetWare Cloud platform, the online metabolomics analysis tool MetaboAnalyst 6.0 (http://www.metaboanalyst.ca), and the Kyoto Encyclopedia of Genes and Genomes (KEGG) to investigate the metabolic pathways involved in differential metabolites, and then analyzed the molecular mechanism of the effect of HFPO-DA on E14TG2a cells.
1, m/z 760.58) as the screening criteria for the single-cell signal, subtracting the same batch of cell-free background solution signal, and the cell intensities at least 3 × 104 after subtracting the background solution signals of the same batch without cells. In addition, ions with a signal-to-noise ratio greater than 3 times and occurrence frequency greater than 10% were extracted. Uniform Manifold Approximation and Projection (UMAP) clustering analysis was conducted using R language-based programs to study the overall differences in metabolic profiles between the HFPO-DA exposure group and the control group. Differential analysis of cells was performed using the MetWare Cloud platform (https://cloud.metware.cn), and significantly different metabolite ions were screened and displayed in a volcano plot. We used the online mzCloud database (https://www.mzcloud.org), the Human Metabolome Database (HMDB), and Compound Discoverer 3.3 (ThermoFisher Scientific) software to qualitatively identify the metabolite ions through accurate m/z and fragmentation information to mark characteristic ions. Then, we used the MetWare Cloud platform, the online metabolomics analysis tool MetaboAnalyst 6.0 (http://www.metaboanalyst.ca), and the Kyoto Encyclopedia of Genes and Genomes (KEGG) to investigate the metabolic pathways involved in differential metabolites, and then analyzed the molecular mechanism of the effect of HFPO-DA on E14TG2a cells.
Author contributions
Y. L. and W. Z. conceived the research. Y. L. conducted the majority of experiments. W.Z. wrote the paper. Y. L., W. Z., and H. Y. analyzed the data. T. P., T. C., and D. G. provided guidance on experiments. X. Y., N. J., and G.G. supervised the research. W. Z., N. J., and X. W. contributed to conceptualization, supervision, writing – review and editing, funding acquisition, and resources.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: the influence of ion source voltage, cell driving pressure, and MS detection time on cell detection, cell viability, metabolite heatmaps, cell flux, and metabolite lists. See DOI: https://doi.org/10.1039/d5sc07089a.Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (22127805 and 22206008), and the Beijing Outstanding Young Scientist Program (BJJWZYJH01201910005017).References
- Z. Y. Wang, I. T. Cousins, M. Scheringer and K. Hungerbühler, Fluorinated alternatives to long-chain perfluoroalkyl carboxylic acids (PFCAs), perfluoroalkane sulfonic acids (PFSAs) and their potential precursors, Environ. Int., 2013, 60, 242–248 Search PubMed.
- L. Zhang, M. Wang, M. Zhang and D. Yang, er- and polyfluoroalkyl substances in Chinese surface waters: A review, Ecotoxicol. Environ. Saf., 2023, 262, 115178 Search PubMed.
- R. A. Brase, E. J. Mullin and D. C. Spink, Legacy and Emerging Per- and Polyfluoroalkyl Substances: Analytical Techniques, Environmental Fate, and Health Effects, Int. J. Mol. Sci., 2021, 22, 995 CrossRef CAS.
- E. E. Harrison and M. L. Waters, Detection and differentiation of per- and polyfluoroalkyl substances (PFAS) in water using a fluorescent imprint-and-report sensor array, Chem. Sci., 2023, 14, 928–936 RSC.
- J. K. Kang, M. G. Kim, S. B. Kim, S. Jeong and J. E. Oh, Comparative study on Perfluoro(2-methyl-3-oxahexanoic) acid removal by quaternary ammonium functionalized silica gel and granular activated carbon from batch and column experiments and molecular simulation-based interpretation, Sci. Total Environ., 2024, 926, 171753 CrossRef CAS PubMed.
- H. Joerss, Z. Xie, C. C. Wagner, W.-J. Von Appen, E. M. Sunderland and R. Ebinghaus, Transport of Legacy Perfluoroalkyl Substances and the Replacement Compound HFPO-DA through the Atlantic Gateway to the Arctic Ocean-Is the Arctic a Sink or a Source?, Environ. Sci. Technol., 2020, 54, 9958–9967 CrossRef CAS PubMed.
- Z. H. Yang, Q. F. Zhuo, W. L. Wang, S. T. Guo, J. F. Chen, Y. L. Li, S. H. Lv, G. Yu and Y. F. Qiu, Fabrication and characterizations of Zn-doped SnO2-Ti4O7 anode for electrochemical degradation of hexafluoropropylene oxide dimer acid and its homologues, J. Hazard. Mater., 2023, 455, 131605 CrossRef CAS PubMed.
- Y. Li, Z. Niu and Y. Zhang, Occurrence of legacy and emerging poly- and perfluoroalkyl substances in water: A case study in Tianjin (China), Chemosphere, 2022, 287, 132409 CrossRef CAS.
- C. R. Jensen, D. P. Genereux, D. K. Solomon, D. R. U. Knappe and T. E. Gilmore, Forecasting and hindcasting PFAS concentrations in groundwater discharging to streams near a PFAS production facility, Environ. Sci. Technol., 2024, 58, 17926–17936 CrossRef CAS.
- Z. Xing, G. G. Wang, S. H. Liu, H. Y. Chen, X. Dong, H. X. Wang and Y. Liu, Legacy and emerging per- and polyfluoroalkyl substances (PFASs) in agricultural soils affected by fluorochemical manufacturing facilities, North China: Occurrence, region-specific distribution, substitution trend and source appointment, J. Hazard. Mater., 2024, 474(10), 134770 CrossRef CAS.
- X. Feng, X. Chen, Y. Yang, L. Yang, Y. Zhu, G. Shan, L. Zhu and S. Zhang, External and internal human exposure to PFOA and HFPOs around a mega fluorochemical industrial park, China: Differences and implications, Environ. Int., 2021, 157, 106824 Search PubMed.
- Z. Zhao, L. Yue, H. Qiao, Y. Li, X. Cheng, X. Hua, T. Lin, Q. Li and H. Sun, Perfluoroalkyl acids in dust on residential indoor/outdoor window glass in Chinese cities: occurrence, composition, and toddler exposure, Environ. Sci. Pollut. Res., 2022, 29, 13881–13892 CrossRef CAS PubMed.
- S. Feng, X. Lu, K. Ouyang, G. Su, Q. Li, B. Shi and J. Meng, Environmental occurrence, bioaccumulation and human risks of emerging fluoroalkylether substances: Insight into security of alternatives, Sci. Total Environ., 2024, 922, 171151 CrossRef CAS PubMed.
- L. Chen, D. Chen, S. Zhou, J. Lin, Y. Liu, X. Huang, Q. Lin, J. L. Morel, Z. Ni, S. Wang and R. Qiu, New insights into the accumulation, transport, and distribution mechanisms of hexafluoropropylene oxide homologues, important alternatives to perfluorooctanoic acid, in Lettuce (Lactuca sativa L.), Environ. Sci. Technol., 2023, 57, 9702–9712 CrossRef CAS PubMed.
- Z. F. Xiu, N. Zheng, Q. R. An, C. C. Chen, Q. Y. Lin, X. Q. Li, S. J. Wang, L. Y. Peng, Y. Y. Li, H. C. Zhu, S. Y. Sun and S. Wang, Tissue-specific distribution and fatty acid content of PFAS in the northern Bohai Sea fish: Risk-benefit assessment of legacy PFAS and emerging alternatives, J. Hazard. Mater., 2024, 480, 136024 CrossRef CAS PubMed.
- E. Ivantsova, V. Lopez-Scarim, A. Sultan, C. English, A. Biju, C. L. Souders Ii, N. E. Padillo-Anthemides, I. Konig and C. J. Martyniuk, Evidence for neurotoxicity and oxidative stress in zebrafish embryos/larvae treated with HFPO-DA ammonium salt (GenX), Environ. Toxicol. Pharmacol., 2023, 104, 104315 CrossRef CAS.
- S. Gong, F. McLamb, D. Shea, J. P. Vu, M. F. Vasquez, Z. Feng, K. Bozinovic, K. K. Hirata, R. M. Gersberg and G. Bozinovic, Toxicity assessment of hexafluoropropylene oxide-dimer acid on morphology, heart physiology, and gene expression during zebrafish (Danio rerio) development, Environ. Sci. Pollut. Res., 2023, 30, 32320–32336 CrossRef CAS.
- Y. Wang, S. Jiang, B. Wang, X. Chen and G. Lu, Comparison of developmental toxicity induced by PFOA, HFPO-DA, and HFPO-TA in zebrafish embryos, Chemosphere, 2023, 311, 136999 CrossRef CAS PubMed.
- L. Kashobwe, F. Sadrabadi, L. Brunken, A. Coelho, T. M. Sandanger, A. Braeuning, T. Buhrke, M. Öberg, T. Hamers and P. E. G. Leonards, Legacy and alternative perand polyfluoroalkyl substances (PFAS) alter the lipid profile of HepaRG cells, Toxicology, 2024, 506, 153862 CrossRef CAS PubMed.
- E. Marques, M. Pfohl, W. Wei, G. Tarantola, L. Ford, O. Amaeze, J. Alesio, S. Ryu, X. Jia, H. Zhu, G. D. Bothun and A. Slitt, Replacement per- and polyfluoroalkyl substances (PFAS) are potent modulators of lipogenic and drug metabolizing gene expression signatures in primary human hepatocytes, Toxicol. Appl. Pharmacol., 2022, 442, 115991 CrossRef CAS PubMed.
- S. Qin, X. Zhang, Y. Zhang, D. Miao, W. Wei and Y. Bai, Multi-dimensional bio mass cytometry: simultaneous analysis of cytoplasmic proteins and metabolites on single cells, Chem. Sci., 2025, 16, 3187–3197 RSC.
- R. Zenobi, Single-cell metabolomics: analytical and biological perspectives, Science, 2013, 342, 1243259 CrossRef CAS PubMed.
- X. Sun, Y. Yu, K. Qian, J. Wang and L. Huang, Recent progress in mass spectrometry-based single-cell metabolic analysis, Small Methods, 2023, 8, 2301317 CrossRef.
- Y. Lei, R. Tang, J. Xu, W. Wang, B. Zhang, J. Liu, X. Yu and S. Shi, Applications of single-cell sequencing in cancer research: progress and perspectives, J. Hematol. Oncol., 2021, 14, 91 CrossRef PubMed.
- S. Cheng, C. Cao, Y. Qian, H. Yao, X. Gong, X. Dai, Z. Ouyang and X. Ma, High-throughput single-cell mass spectrometry enables metabolic network analysis by resolving phospholipid C=C isomers, Chem. Sci., 2024, 15, 6314–6320 RSC.
- G. Zhu, Y. Shao, Y. Liu, T. Pei, L. Li, D. Zhang, G. Guo and X. Wang, Single-cell metabolite analysis by electrospray ionization mass spectrometry, TrAC, Trends Anal. Chem., 2021, 143, 116351 CrossRef CAS.
- S. J. Qin, Y. Zhang, M. Y. Shi, D. Y. Miao, J. S. Lu, L. Wen and Y. Bai, In-depth organic mass cytometry reveals differential contents of 3-hydroxybutanoic acid at the single-cell level, Nat. Commun., 2024, 15, 4387 CrossRef CAS PubMed.
- M. Zhuang, Z. Hou, P. Chen, G. Liang and G. Huang, Introducing charge tag via click reaction in living cells for single cell mass spectrometry, Chem. Sci., 2020, 11, 7308–7312 RSC.
- X. Gong, Y. Zhao, S. Cai, S. Fu, C. Yang, S. Zhang and X. Zhang, Single cell analysis with probe ESI-mass spectrometry: detection of metabolites at cellular and subcellular levels, Anal. Chem., 2014, 86, 3809–3816 CrossRef CAS PubMed.
- H. Yao, H. Zhao, X. Pan, X. Zhao, J. Feng, C. Yang, S. Zhang and X. Zhang, Discriminating leukemia cellular heterogeneity and screening metabolite biomarker candidates using label-free mass cytometry, Anal. Chem., 2021, 93, 10282–10291 CrossRef CAS PubMed.
- Z. Shen, H. Zhao, H. Yao, X. Pan, J. Yang, S. Zhang, G. Han and X. Zhang, Dynamic metabolic change of cancer cells induced by natural killer cells at the single-cell level studied by label-free mass cytometry, Chem. Sci., 2022, 13, 1641–1647 RSC.
- Y. Shao, Y. Zhou, Y. Liu, W. Zhang, G. Zhu, Y. Zhao, Q. Zhang, H. Yao, H. Zhao, G. Guo, S. Zhang, X. Zhang and X. Wang, Intact living-cell electrolaunching ionization mass spectrometry for single-cell metabolomics, Chem. Sci., 2022, 13, 8065–8073 RSC.
- J. Deng, X. Zeng, C. He, D. Zhong, Y. Wu, N. Liu, T. Luan and Y. Yang, Exploring the accumulation behavior and heterogeneity of perfluorooctanesulfonic acid in Zebrafish primary organ cells by single-cell mass cytometry, Anal. Chem., 2023, 95, 13750–13755 CrossRef CAS PubMed.
- J. W. Deng, J. L. Xie, C. Wang, Y. H. Wu, T. A. Luan and Y. Y. Yang, Inner-wall coated nanopipette microextraction for quantitative analysis of per- and polyfluoroalkyl substances in single cells using mass spectrometry, Anal. Chem., 2024, 96, 1391–1396 CrossRef CAS PubMed.
- P. F. Li, S. Gao, W. T. Qu, Y. Li and Z. Liu, Chemo-selective single-cell metabolomics reveals the spatiotemporal behavior of exogenous pollutants during Xenopus Laevis embryogenesis, Adv. Sci., 2024, 11, e2305401 CrossRef PubMed.
- F. Zhang, L. Liu, J. Hu, H. Fu, H. Li, J. Chen, C. Yang, Q. Guo, X. Liang, L. Wang, Y. Guo, J. Dai, N. Sheng and J. Wang, Accumulation and glucocorticoid signaling suppression by four emerging perfluoroethercarboxylic acids based on animal exposure and cell testing, Environ. Int., 2023, 178, 108092 CrossRef CAS PubMed.
- S. Wang, T. Liu, X. Qian, H. Wang, M. Li, X. Wang, S. Wei and H. Chen, Microbial plankton responses to perfluoroalkyl acids and their alternatives in the aquatic environment, J. Hazard. Mater., 2023, 441, 129980 CrossRef CAS.
- T. C. Guillette, J. McCord, M. Guillette, M. E. Polera, K. T. Rachels, C. Morgeson, N. Kotlarz, D. R. U. Knappe, B. J. Reading, M. Strynar and S. M. Belcher, Elevated levels of per- and polyfluoroalkyl substances in Cape Fear River Striped Bass (Morone saxatilis) are associated with biomarkers of altered immune and liver function, Environ. Int., 2020, 136, 105358 Search PubMed.
- D. Armenta-Medina, L. Segovia and E. Perez-Rueda, Comparative genomics of nucleotide metabolism: a tour to the past of the three cellular domains of life, BMC Genomics, 2014, 15, 800 CrossRef PubMed.
- X. E. Zhang, C. Liu, J. Dai, Y. Yuan, C. Gao, Y. Feng, B. Wu, P. Wei, C. You, X. Wang and T. Si, Enabling technology and core theory of synthetic biology, Sci. China:Life Sci., 2023, 66, 1742–1785 CrossRef PubMed.
- H. E. Krokan, F. Drablos and G. Slupphaug, Uracil in DNA - occurrence, consequences and repair, Oncogene, 2002, 21, 8935–8948 Search PubMed.
- L. J. Van Winkle, Amino acid transport and metabolism regulate early embryo development: species differences, clinical significance, and evolutionary implications, Cells, 2021, 10, 3154 CrossRef CAS PubMed.
- Z. N. Ling, Y. F. Jiang, J. N. Ru, J. H. Lu, B. Ding and J. Wu, Amino acid metabolism in health and disease, Signal Transduction Targeted Ther., 2023, 8, 345 CrossRef PubMed.
- B. Kelly and E. L. Pearce, Amino assests: how amino acids support immunity, Cell Metab., 2020, 32, 154–175 CrossRef CAS PubMed.
- R. Curi, P. Newsholme, J. Procopio, C. Lagranha, R. Gorjao and T. C. Pithon-Curi, Glutamine, gene expression, and cell function, Front. Biosci., 2007, 12, 344–357 CrossRef CAS PubMed.
- R. Curi, C. J. Lagranha, S. Q. Doi, D. F. Sellitti, J. Procopio, T. C. Pithon-Curi, M. Corless and P. Newsholme, Molecular mechanisms of glutamine action, J. Cell. Physiol., 2005, 204, 392–401 CrossRef CAS PubMed.
- B. Wang, G. Wu, Z. Zhou, Z. Dai, Y. Sun, Y. Ji, W. Li, W. Wang, C. Liu, F. Han and Z. Wu, Glutamine and intestinal barrier function, Amino Acids, 2014, 47, 2143–2154 Search PubMed.
- F. Luckose, M. C. Pandey and K. Radhakrishna, Effects of amino acid derivativeson physical, mental, and physiological activities, Crit. Rev. Food Sci. Nutr., 2015, 55, 1793–1807 CrossRef CAS PubMed.
- P. Garbati, A. Salis, E. Adriano, A. Galatini, G. Damonte, M. Balestrino and E. Millo, A new method to synthesize creatine derivatives, Amino Acids, 2013, 45, 821–833 CrossRef CAS PubMed.
- L. N. Vandenberg, in Endocrine Disruption and Human Health, ed. P. D. Darbre, Academic Press, 2nd edn, 2022, pp. 141–163, DOI:10.1016/B978-0-12-821985-0.00006-2.
- J. P. Vu, F. McLamb, Z. Feng, L. Griffin, S. Gong, D. Shea, M. A. Szuch, S. Scott, R. M. Gersberg and G. Bozinovic, Locomotion and brain gene expression exhibit sex-specific non-monotonic dose-response to HFPO-DA during Drosophila melanogaster lifespan, Neurotoxicology, 2023, 96, 207–221 CrossRef CAS PubMed.
- G. Gałęzowska, J. Rogowska and J. Antosiewicz, Drug interactions with perfluorooctanoic acid and perfluorooctane sulfonate in cytotoxic activity against prostate cancer – in vitro studies, J. Environ. Sci., 2026, 159, 88–96 CrossRef PubMed.
- R. Li, Y. Shao, Y. Yu, X. Wang and G. Guo, Pico-HPLC system integrating an equal inner diameter femtopipette into a 900 nm I.D. porous layer open tubular column, Chem. Commun., 2017, 53, 4104–4107 RSC.
Footnote |
| † Y. L. and W. Z. contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |